Differentially Expressed microRNAs in MIA PaCa-2 and PANC-1 Pancreas Ductal Adenocarcinoma Cell Lines are Involved in Cancer Stem Cell Regulation
Abstract
1. Introduction
2. Results
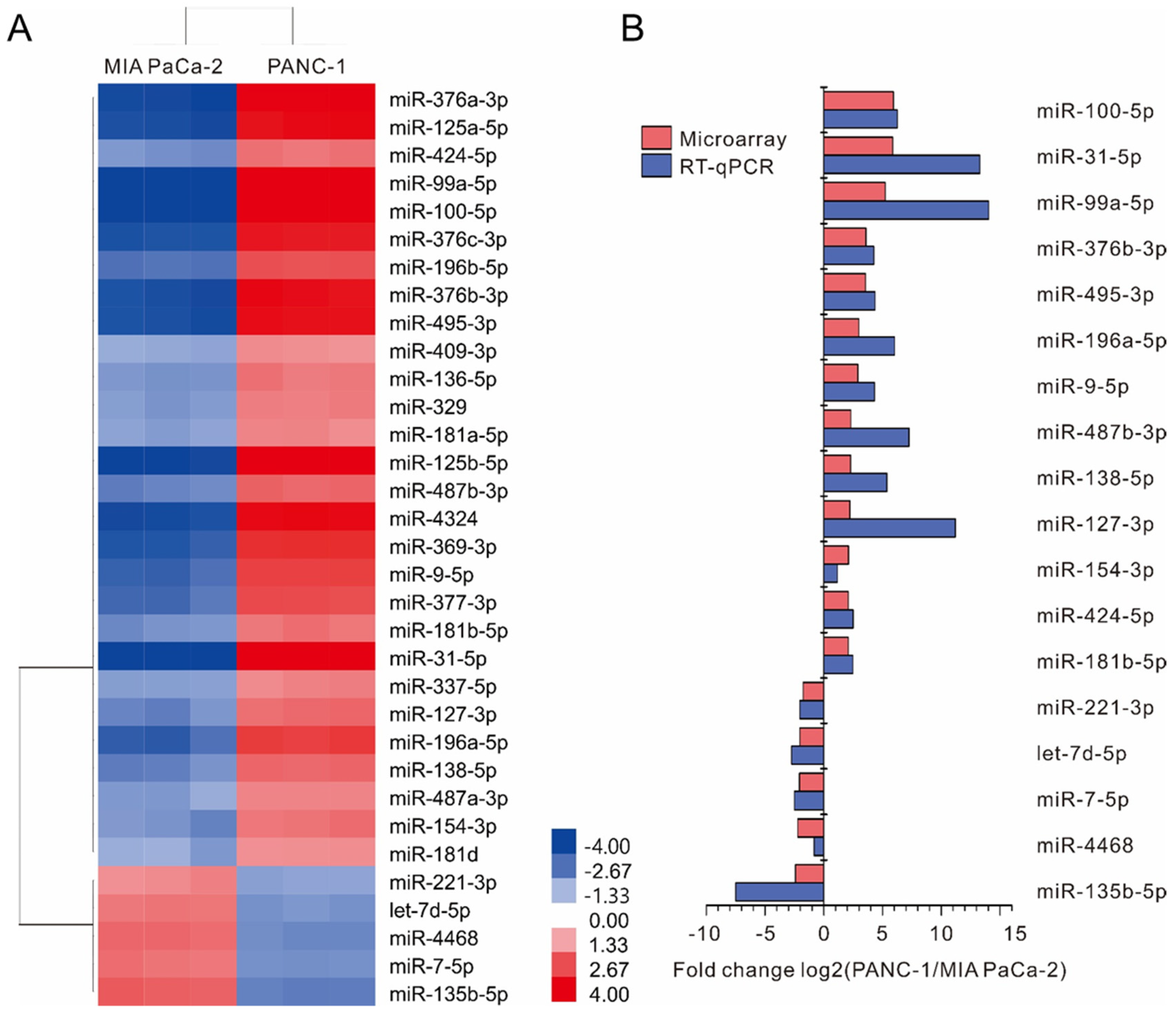
2.1. Screening of Differentially Expressed miRNAs between MIA PaCa-2 and PANC-1 Cell Lines
2.2. Validation of Differentially Expressed miRNAs by Real-Time RT-PCR
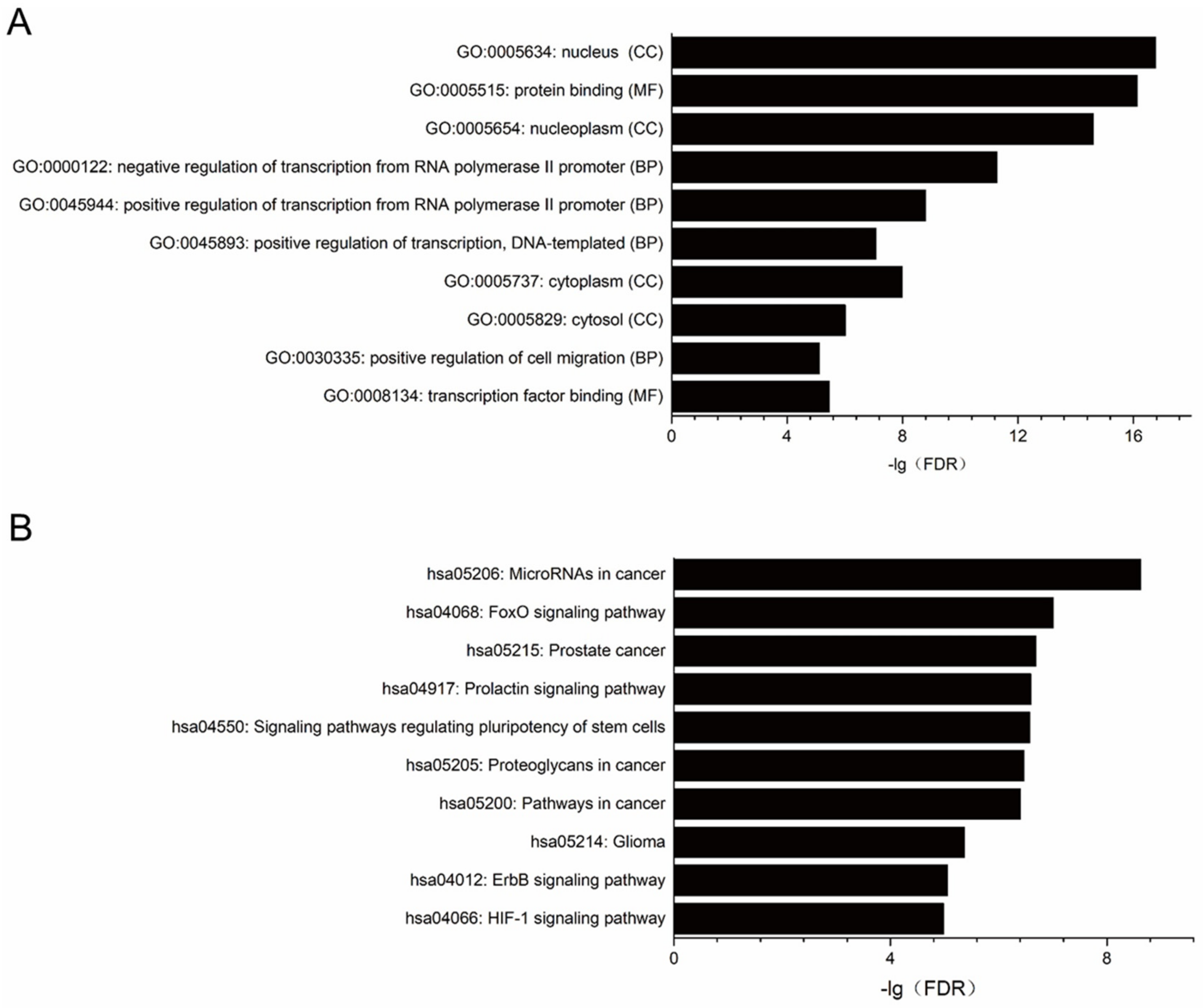
2.3. Target Prediction, Gene Ontology (GO) and Pathway Enrichment Analysis for miRNAs Significantly Changed in PDAC Cell Lines
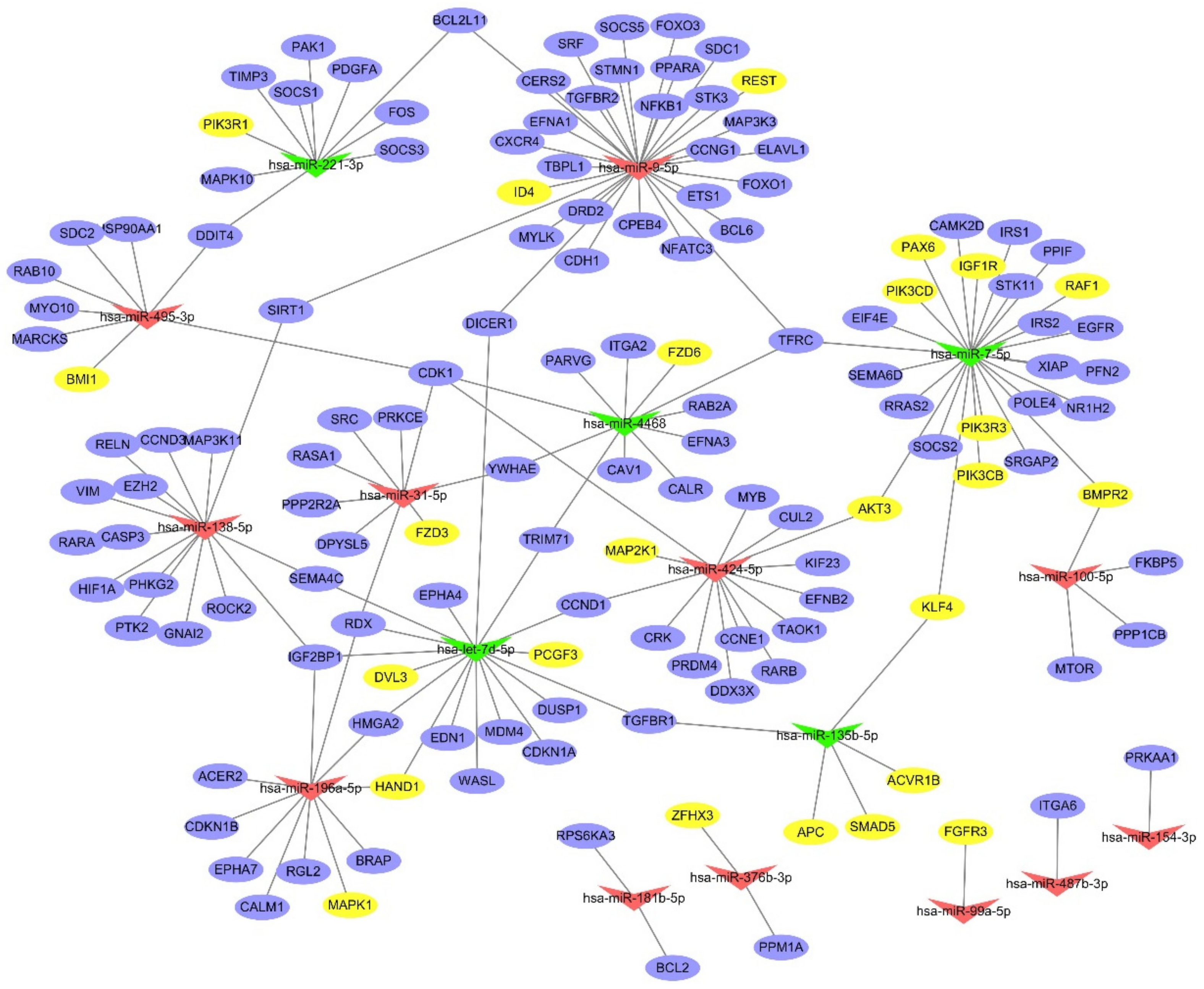
2.4. Interaction Network Analysis
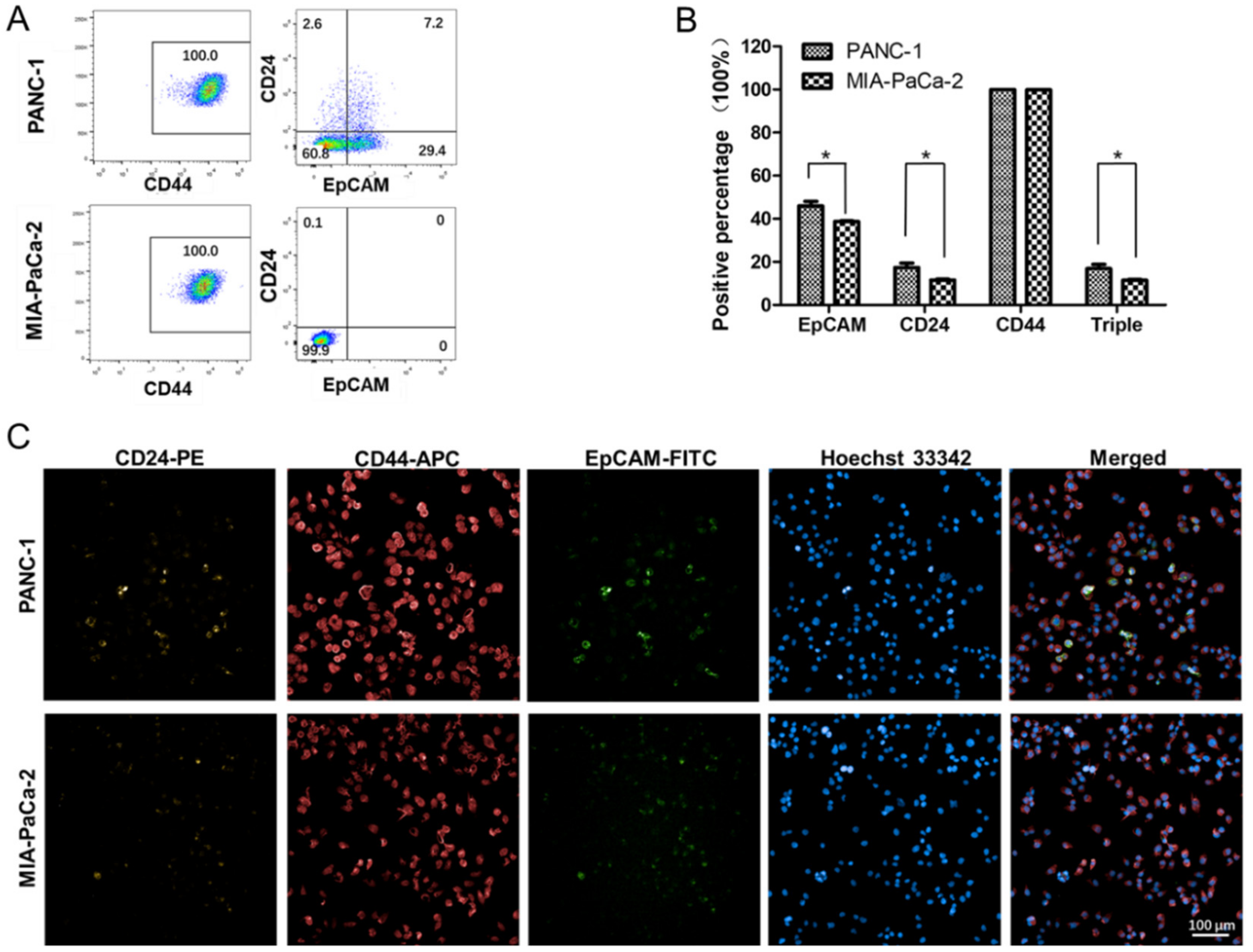
2.5. Differences in the Stem Cell Surface Markers Expression Percentage of PANC-1 and MIA-PaCa-2
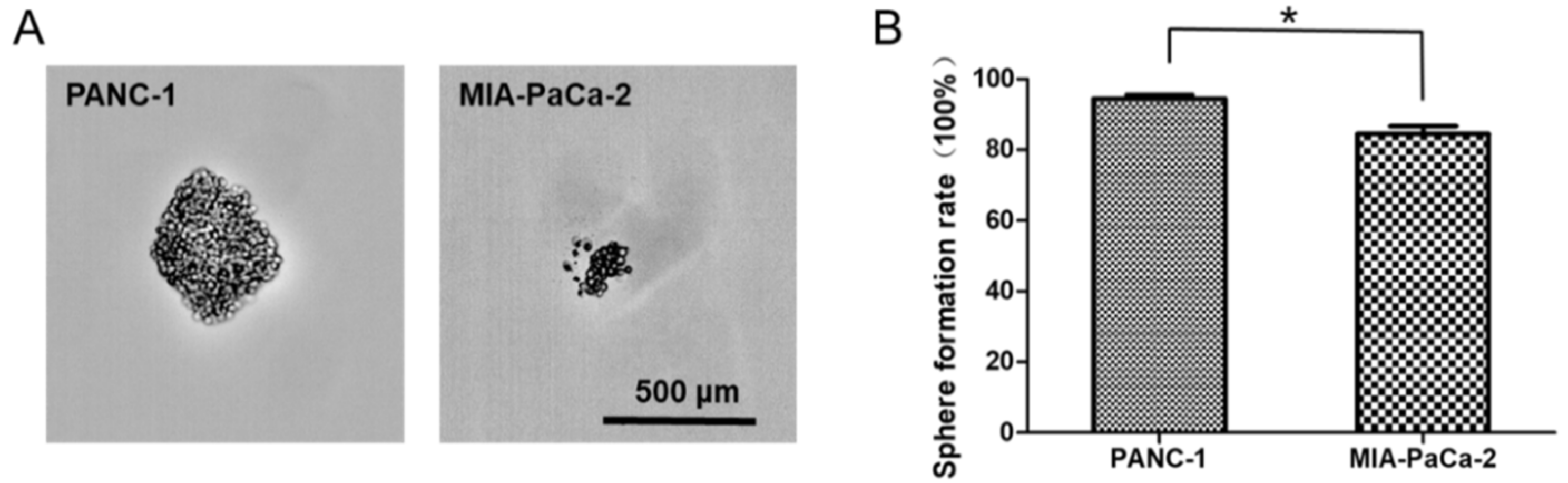
2.6. Differences in Capacity of Sphere Formation between PANC-1 and MIA-PaCa-2
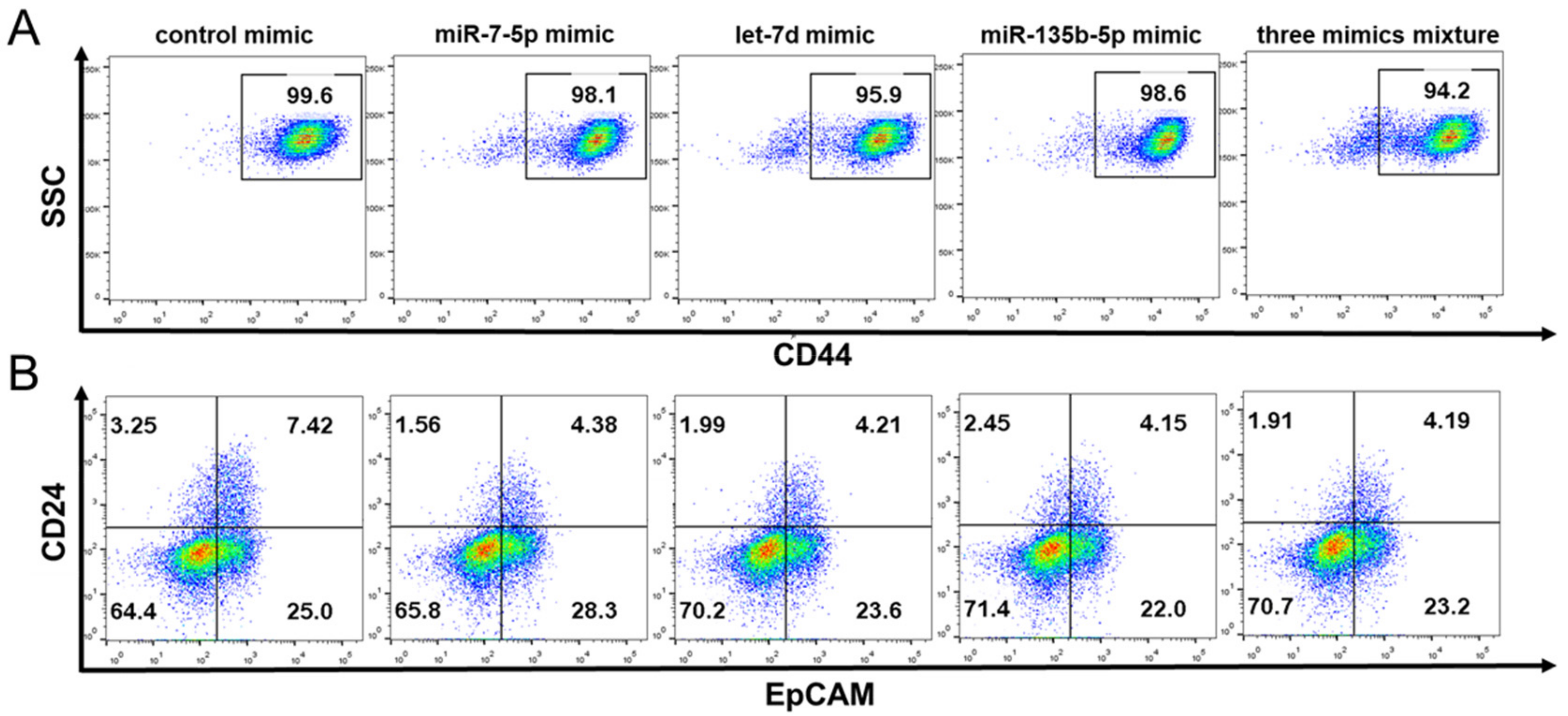
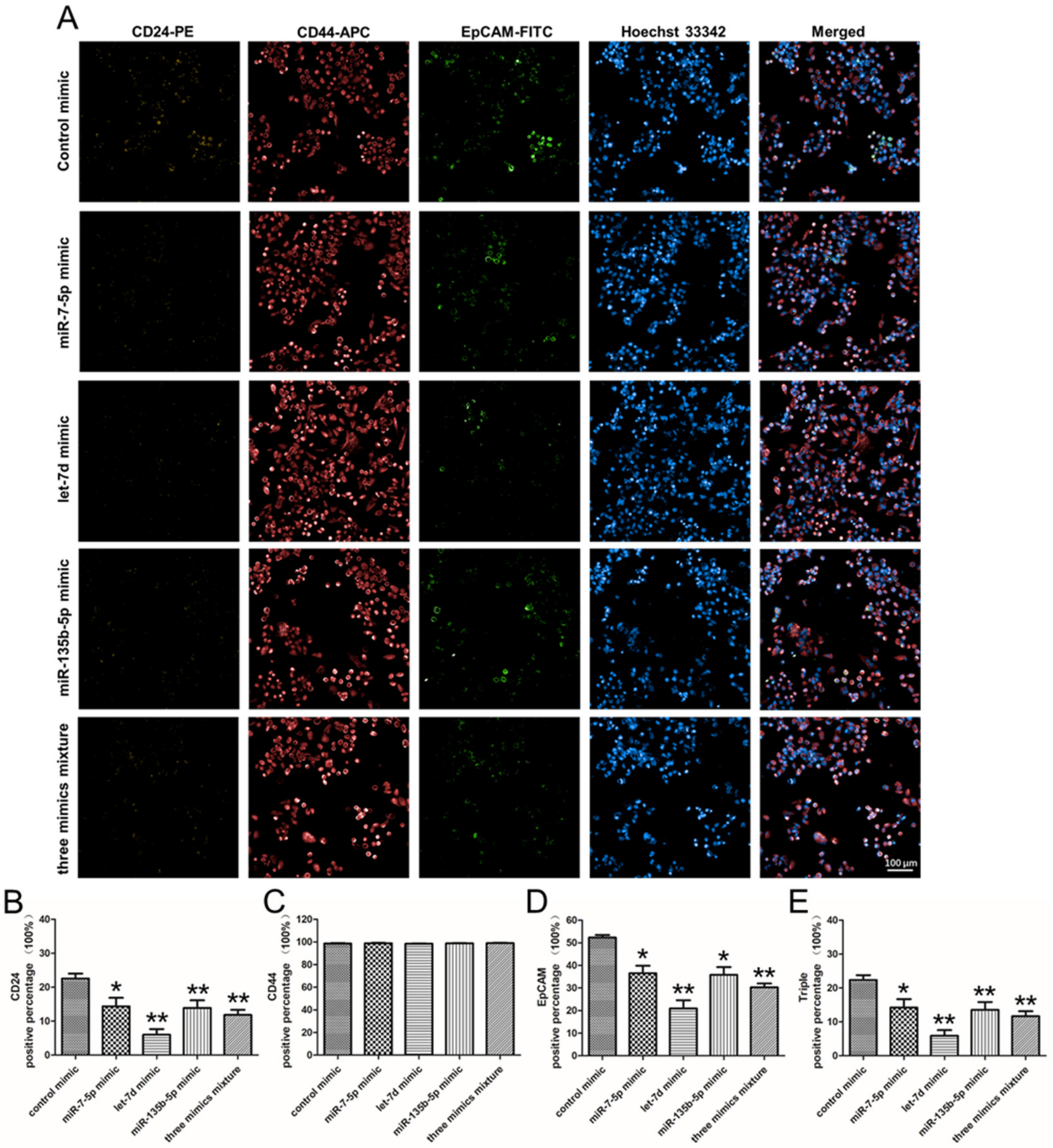
2.7. The Effect of miRNA Mimics on the Stem Cell Surface Markers Expression Percentage of PANC-1
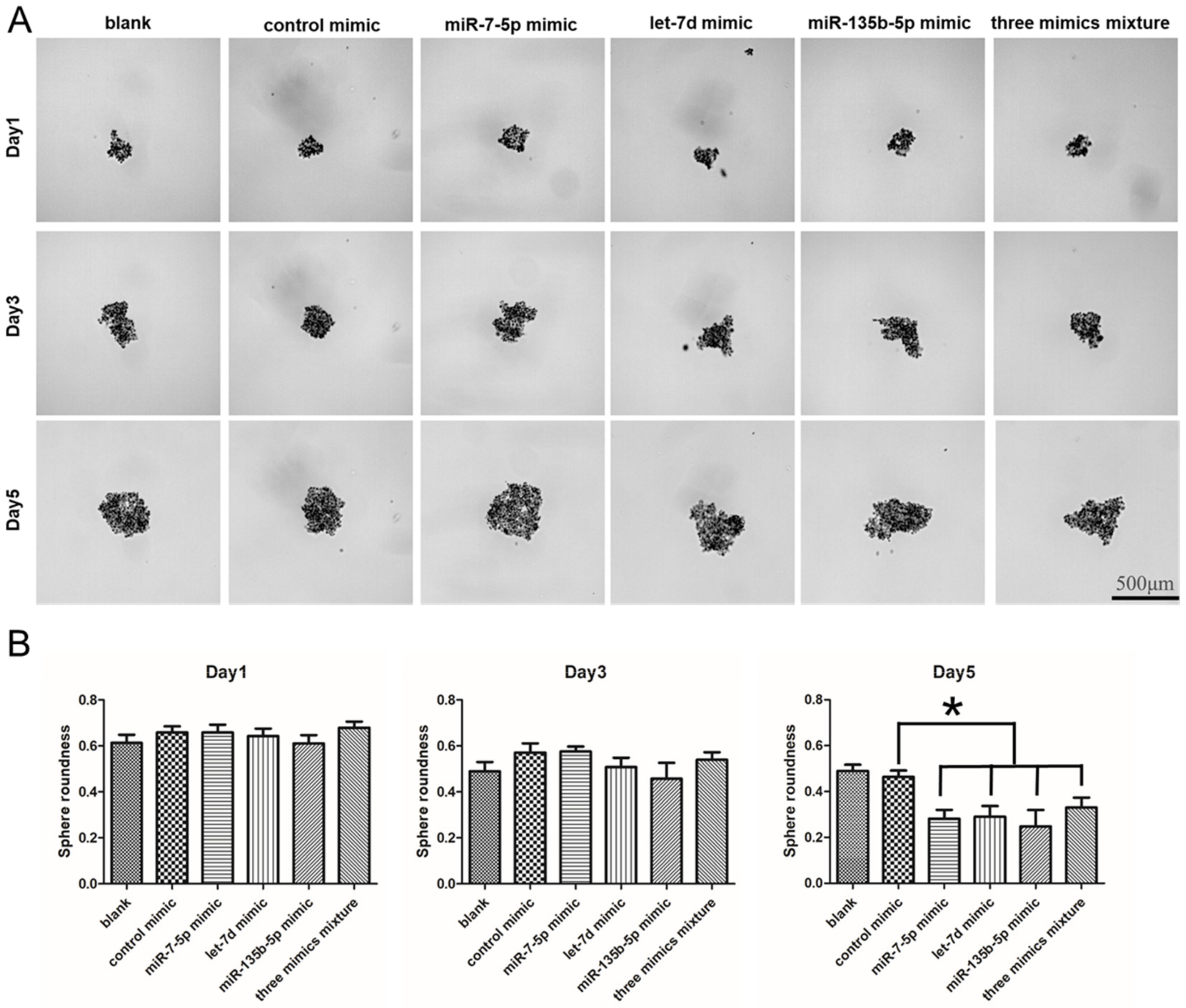
2.8. The Regulation of miRNA Mimics on the Sphere Formation Ability in PANC-1
3. Discussion
4. Materials and Methods
4.1. Cell Culture
4.2. Extraction of microRNA and Microarray Screening
4.3. miRNA Expression Analysis Using qRT-PCR
4.4. Bioinformatics
4.5. Flow Cytometry
4.6. Immunocytochemistry (ICC)
4.7. Sphere Formation Culture
4.8. miRNA Mimics Transfection
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| BP | biological process |
| BL | burkitt’s lymphoma |
| CC | cellular component |
| CRC | colorectal |
| CSC | cancer stem cell |
| DAVID | Database for Annotation, Visualization and Integrated Discovery |
| EGFR | epidermal growth factor receptor |
| EMT | epithelial-to-mesenchymal transition |
| GBM | glioblastoma multiforme |
| GO | Gene Ontology |
| HER2 | human epidermal growth factor receptor 2 |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| Klf4 | Kruppel-like factor 4 |
| NSCLC | non-small cell lung carcinoma |
| MF | molecular function |
| miRNA | small noncoding microRNA |
| Pak-1 | p21 activated kinase 1 |
| PDAC | pancreatic ductal adenocarcinoma |
| PDCD4 | programmed cell death 4 |
| SCID | Severe Combined Immunodeficiency |
| SD | standard deviation |
| SFM | sphere formation medium |
| Sox2 | stem cell factors Y–box 2 |
| TUBB3 | tubulin beta 3 class III |
| UT | Universal Tag |
| ZEB1 | zinc finger E-box binding homeobox 1 |
References
- Chiaravalli, M.; Reni, M.; O’Reilly, E.M. Pancreatic ductal adenocarcinoma: State-of-the-art 2017 and new therapeutic strategies. Cancer Treat. Rev. 2017, 60, 32–43. [Google Scholar] [CrossRef]
- Adamska, A.; Domenichini, A.; Falasca, M. Pancreatic Ductal Adenocarcinoma: Current and Evolving Therapies. Int. J. Mol. Sci. 2017, 18, 1338. [Google Scholar] [CrossRef]
- Ishiwata, T. Pancreatic Ductal Adenocarcinoma: Basic and Clinical Challenges for Better Prognosis. J. Carcinog. Mutagen. 2013, S9, 005. [Google Scholar] [CrossRef]
- Bailey, P.; Chang, D.K.; Nones, K.; Johns, A.L.; Patch, A.M.; Gingras, M.C.; Miller, D.K.; Christ, A.N.; Bruxner, T.J.C.; Quinn, M.C. Genomic analyses identify molecular subtypes of pancreatic cancer. Nature 2016, 531, 47–52. [Google Scholar] [CrossRef] [PubMed]
- Chiorean, E.G.; Coveler, A.L. Pancreatic cancer: Optimizing treatment options, new, and emerging targeted therapies. Drug Des. Dev. Ther. 2015, 9, 3529–3545. [Google Scholar] [CrossRef]
- Yonemori, K.; Kurahara, H.; Maemura, K.; Natsugoe, S. MicroRNA in pancreatic cancer. J. Hum. Genet. 2016, 62, 33. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.F.; Hannafon, B.N.; Ding, W.Q. microRNA regulation of human pancreatic cancer stem cells. Stem Cell Investig. 2017, 4, 5. [Google Scholar] [CrossRef]
- Bao, B.; Ali, S.; Ahmad, A.; Li, Y.; Banerjee, S.; Kong, D.; Aboukameel, A.; Mohammad, R.; Van Buren, E.; Azmi, A.S.; et al. Differentially expressed miRNAs in cancer-stem-like cells: Markers for tumor cell aggressiveness of pancreatic cancer. Stem Cells Dev. 2014, 23, 1947–1958. [Google Scholar] [CrossRef]
- Li, Y.; Laterra, J. Cancer stem cells: Distinct entities or dynamically regulated phenotypes? Cancer Res. 2012, 72, 576. [Google Scholar] [CrossRef]
- Wang, A.; Chen, L.; Li, C.; Zhu, Y. Heterogeneity in cancer stem cells. Cancer Lett. 2015, 357, 63–68. [Google Scholar] [CrossRef]
- Dorado, J.; Serrano, A.G.; Heeschen, C. Cancer Stem Cells in Pancreatic Cancer. Cancers 2010, 2, 1629. [Google Scholar]
- Chenwei, L.; Heidt, D.G.; Piero, D.; Burant, C.F.; Lanjing, Z.; Volkan, A.; Max, W.; Clarke, M.F.; Simeone, D.M. Identification of pancreatic cancer stem cells. Cancer Res. 2007, 130, 194–195. [Google Scholar]
- Rui, G.; Silva, H.C.; Carvalho, L.; Botelho, M.F.; Motapinto, A. MIA PaCa-2 and PANC-1 – pancreas ductal adenocarcinoma cell lines with neuroendocrine differentiation and somatostatin receptors. Sci. Rep. 2016, 6, 21648. [Google Scholar]
- Deer, E.L.; Jessica, G.H.; Coursen, J.D.; Shea, J.E.; Josephat, N.; Scaife, C.L.; Firpo, M.A.; Mulvihill, S.J. Phenotype and genotype of pancreatic cancer cell lines. Pancreas 2010, 39, 425–435. [Google Scholar] [CrossRef] [PubMed]
- Kent, O.A.; Mullendore, M.; Wentzel, E.A.; López-Romero, P.; Tan, A.C.; Alvarez, H.; West, K.; Ochs, M.F.; Hidalgo, M.; Arking, D.E.; et al. A resource for analysis of microRNA expression and function in pancreatic ductal adenocarcinoma cells. Cancer Biol. Ther. 2009, 8, 2013–2024. [Google Scholar] [CrossRef] [PubMed]
- Duan, D.; Zheng, K.X.; Shen, Y.; Cao, R.; Jiang, L.; Lu, Z.; Yan, X.; Li, J. Label-free high-throughput microRNA expression profiling from total RNA. Nucleic Acids Res. 2011, 39, e154. [Google Scholar] [CrossRef] [PubMed]
- Wang, A.; Chen, L.; Pu, K.; Zhu, Y. Identification of stem-like cells in non-small cell lung cancer cells with specific peptides. Cancer Lett. 2014, 351, 100–107. [Google Scholar] [CrossRef]
- Rothschild, S.I. microRNA therapies in cancer. Mol. Cell. Ther. 2014, 2, 7. [Google Scholar] [CrossRef]
- Christopher, A.F.; Kaur, R.P.; Kaur, G.; Kaur, A.; Gupta, V.; Bansal, P. MicroRNA therapeutics: Discovering novel targets and developing specific therapy. Perspect. Clin. Res. 2016, 7, 68–74. [Google Scholar]
- Candido, S.; Lupo, G.; Pennisi, M.; Basilel, M.S.; Anfuso, C.D.; Petralia, M.C.; Gattuso, G.; Vivarelli, S.; Spandidos, D.A.; Libra, M.; et al. The analysis of miRNA expression profiling datasets reveals inverse microRNA patterns in glioblastoma and Alzheimer’s disease. Oncol. Rep. 2019, 42, 911–922. [Google Scholar]
- Falzone, L.; Lupo, G.; La Rosa, G.R.; Crimi, S.; Anfuso, C.D.; Salemi, R.; Rapisarda, E.; Libra, M.; Candido, S. Identification of Novel MicroRNAs and Their Diagnostic and Prognostic Significance in Oral Cancer. Cancers 2019, 11, 610. [Google Scholar] [CrossRef] [PubMed]
- Felix, T.F.; Lopez Lapa, R.M.; de Carvalho, M.; Bertoni, N.; Tokar, T.; Oliveira, R.A.; Rodrigues, M.A.M.; Hasimoto, C.N.; Oliveira, W.K.; Pelafsky, L.; et al. MicroRNA modulated networks of adaptive and innate immune response in pancreatic ductal adenocarcinoma. PLoS ONE 2019, 14, e0217421. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Uzair-Ur-Rehman; Guo, Y.; Liang, H.; Cheng, R.; Yang, F.; Hong, Y.; Zhao, C.; Liu, M.; Yu, M.; et al. miR-181b functions as an oncomiR in colorectal cancer by targeting PDCDZ. Protein Cell 2016, 7, 722–734. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Lv, X.; Wang, X.; Wang, B.; Shao, X.; Huang, Y.; Shi, L.; Chen, Z.; Huang, J.; Huang, P. MiR-181b promotes chemoresistance in breast cancer by regulating Bim expression. Oncol. Rep. 2016, 35, 683–690. [Google Scholar] [CrossRef] [PubMed]
- Hafsi, S.; Candido, S.; Maestro, R.; Falzone, L.; Soua, Z.; Bonavida, B.; Spandidos, D.A.; Libra, M. Correlation between the overexpression of Yin Yang 1 and the expression levels of miRNAs in Burkitt’s lymphoma: A computational study. Oncol. Lett. 2016, 11, 1021–1025. [Google Scholar] [CrossRef] [PubMed]
- Boo, L.; Wan, Y.H.; Ali, N.M.; Yeap, S.K.; Ky, H.; Chan, K.G.; Yin, W.F.; Satharasinghe, D.A.; Liew, W.C.; Tan, S.W. Phenotypic and microRNA transcriptomic profiling of the MDA-MB-231 spheroid-enriched CSCs with comparison of MCF-7 microRNA profiling dataset. PeerJ 2017, 5, e3551. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Kelnar, K.; Vlassov, A.V.; Brown, D.; Wang, J.; Tang, D.G. Distinct microRNA expression profiles in prostate cancer stem/progenitor cells and tumor-suppressive functions of let-7. Cancer Res. 2012, 72, 3393–3404. [Google Scholar] [CrossRef]
- Nam, E.J.; Lee, M.; Yim, G.W.; Kim, J.H.; Kim, S.; Sang, W.K.; Kim, Y.T. MicroRNA profiling of a CD133+spheroid-forming subpopulation of the OVCAR3 human ovarian cancer cell line. BMC Med. Genom. 2012, 5, 18. [Google Scholar] [CrossRef]
- Jung, D.E.; Jing, W.; Taeyun, O.; Young, S.S. Differentially expressed microRNAs in pancreatic cancer stem cells. Pancreas 2011, 40, 1180–1187. [Google Scholar] [CrossRef]
- Tesfaye, A.A.; Azmi, A.S.; Philip, P.A. miRNA and Gene Expression in Pancreatic Ductal Adenocarcinoma. Am. J. Pathol. 2019, 189, 58–70. [Google Scholar] [CrossRef]
- Li, P.; Xu, Q.; Zhang, D.; Li, X.; Han, L.; Lei, J.; Duan, W.; Ma, Q.; Wu, Z.; Wang, Z. Upregulated miR-106a plays an oncogenic role in pancreatic cancer. FEBS Lett. 2014, 588, 705–712. [Google Scholar] [CrossRef]
- Tsukasa, K.; Ding, Q.; Miyazaki, Y.; Matsubara, S.; Natsugoe, S.; Takao, S. miR-30 family promotes migratory and invasive abilities in CD133+ pancreatic cancer stem-like cells. Hum. Cell 2016, 29, 130–137. [Google Scholar] [CrossRef]
- Zhou, W.; Li, Y.; Gou, S.; Xiong, J.; Wu, H.; Wang, C.; Yan, H.; Liu, T. MiR-744 increases tumorigenicity of pancreatic cancer by activating Wnt/β-catenin pathway. Oncotarget 2015, 6, 37557–37569. [Google Scholar] [CrossRef] [PubMed]
- Hasegawa, S.; Eguchi, H.; Nagano, H.; Konno, M.; Tomimaru, Y.; Wada, H.; Hama, N.; Kawamoto, K.; Kobayashi, S.; Nishida, N. MicroRNA-1246 expression associated with CCNG2-mediated chemoresistance and stemness in pancreatic cancer. Br. J. Cancer 2014, 111, 1572–1580. [Google Scholar] [CrossRef] [PubMed]
- Nwaeburu, C.C.; Bauer, N.; Zhao, Z.; Abukiwan, A.; Gladkich, J.; Benner, A.; Herr, I. Up-regulation of microRNA Let-7c by quercetin inhibits pancreatic cancer progression by activation of Numbl. Oncotarget 2016, 7, 58367–58380. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Diaz, P.C.; Hsiao, T.H.; Chang, J.C.; Yue, D.; Tan, M.C.; Chen, H.I.; Tomlinson, G.E.; Huang, Y.; Chen, Y.; Hung, J.Y. De-regulated microRNAs in pediatric cancer stem cells target pathways involved in cell proliferation, cell cycle and development. PLoS ONE 2013, 8, e61622. [Google Scholar] [CrossRef]
- Wang, H. FOXO Signaling Pathways as Therapeutic Targets in Cancer. Int. J. Biol. Sci. 2017, 13, 815–827. [Google Scholar]
- Zhang, L.; Cai, M.; Gong, Z.; Zhang, B.; Li, Y.; Guan, L.; Hou, X.; Li, Q.; Liu, G.; Xue, Z. Geminin facilitates FoxO3 deacetylation to promote breast cancer cell metastasis. J. Clin. Investig. 2017, 127, 2159. [Google Scholar] [CrossRef]
- Kumazoe, M.; Takai, M.; Hiroi, S.; Takeuchi, C.; Kadomatsu, M.; Nojiri, T.; Onda, H.; Bae, J.; Huang, Y.; Takamatsu, K. FOXO3/PGC-1β signaling axis is essential for cancer stem cell properties of pancreatic ductal adenocarcinoma. J. Biol. Chem. 2017, 292, 10813. [Google Scholar] [CrossRef]
- Ghasemi, R.; Rapposelli, I.G.; Capone, E.; Rossi, C.; Lattanzio, R.; Piantelli, M.; Sala, G.; Iacobelli, S. Dual targeting of ErbB-2/ErbB-3 results in enhanced antitumor activity in preclinical models of pancreatic cancer. Oncogenesis 2014, 3, e117. [Google Scholar] [CrossRef]
- Ziello, J.E.; Jovin, I.S.; Yan, H. Hypoxia-Inducible Factor (HIF)-1 regulatory pathway and its potential for therapeutic intervention in malignancy and ischemia. Yale J. Biol. Med. 2007, 80, 51–60. [Google Scholar] [PubMed]
- Horsham, J.L.; Ganda, C.; Kalinowski, F.C.; Brown, R.A.M.; Epis, M.R.; Leedman, P.J. MicroRNA-7: A miRNA with expanding roles in development and disease. Int. J. Biochem. Cell Biol. 2015, 69, 215–224. [Google Scholar] [CrossRef] [PubMed]
- Zhu, W.; Wang, Y.; Zhang, D.; Yu, X.; Leng, X. MiR-7-5p functions as a tumor suppressor by targeting SOX18 in pancreatic ductal adenocarcinoma. Biochem. Biophys. Res. Commun. 2018, 497, 963–970. [Google Scholar] [CrossRef] [PubMed]
- Kolenda, T.; Przybyła, W.; Teresiak, A.; Mackiewicz, A.; Lamperska, K.M. The mystery of let-7d – a small RNA with great power. Contemp. Oncol. 2014, 18, 293–301. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Che, X.; Yang, N.; Bai, Z.; Wu, Y.; Zhao, L.; Pei, H. miR-135b-5p Promotes migration, invasion and EMT of pancreatic cancer cells by targeting NR3C2. Biomed. Pharmacother. 2017, 96, 1341–1348. [Google Scholar] [CrossRef] [PubMed]
- Thiruvengadam, A.; Vijaya, R.; Fournier, K.F.; Huamin, W.; Lauren, M.; Abbruzzese, J.L.; Gallick, G.E.; Logsdon, C.D.; Mcconkey, D.J.; Woonyoung, C. Epithelial to mesenchymal transition contributes to drug resistance in pancreatic cancer. Cancer Res. 2009, 69, 5820–5828. [Google Scholar]
- Ma, C.; Huang, T.; Ding, Y.C.; Yu, W.; Wang, Q.; Meng, B.; Luo, S.X. MicroRNA-200c overexpression inhibits chemoresistance, invasion and colony formation of human pancreatic cancer stem cells. Int. J. Clin. Exp. Pathol. 2014, 8, 6533. [Google Scholar]
- Singh, S.; Chitkara, D.; Kumar, V.; Behrman, S.W.; Mahato, R.I. miRNA profiling in pancreatic cancer and restoration of chemosensitivity. Cancer Lett. 2013, 334, 211–220. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shen, Y.; Pu, K.; Zheng, K.; Ma, X.; Qin, J.; Jiang, L.; Li, J. Differentially Expressed microRNAs in MIA PaCa-2 and PANC-1 Pancreas Ductal Adenocarcinoma Cell Lines are Involved in Cancer Stem Cell Regulation. Int. J. Mol. Sci. 2019, 20, 4473. https://doi.org/10.3390/ijms20184473
Shen Y, Pu K, Zheng K, Ma X, Qin J, Jiang L, Li J. Differentially Expressed microRNAs in MIA PaCa-2 and PANC-1 Pancreas Ductal Adenocarcinoma Cell Lines are Involved in Cancer Stem Cell Regulation. International Journal of Molecular Sciences. 2019; 20(18):4473. https://doi.org/10.3390/ijms20184473
Chicago/Turabian StyleShen, Ye, Kefeng Pu, Kexiao Zheng, Xiaochuan Ma, Jingyi Qin, Li Jiang, and Jiong Li. 2019. "Differentially Expressed microRNAs in MIA PaCa-2 and PANC-1 Pancreas Ductal Adenocarcinoma Cell Lines are Involved in Cancer Stem Cell Regulation" International Journal of Molecular Sciences 20, no. 18: 4473. https://doi.org/10.3390/ijms20184473
APA StyleShen, Y., Pu, K., Zheng, K., Ma, X., Qin, J., Jiang, L., & Li, J. (2019). Differentially Expressed microRNAs in MIA PaCa-2 and PANC-1 Pancreas Ductal Adenocarcinoma Cell Lines are Involved in Cancer Stem Cell Regulation. International Journal of Molecular Sciences, 20(18), 4473. https://doi.org/10.3390/ijms20184473





