Maximum Topological Distances Based Indices as Molecular Descriptors for QSPR. 4. Modeling the Enthalpy of Formation of Hydrocarbons from Elements
Abstract
:1. Introduction
- a)
- Distance is a positive quantity, Dij ≥ 0, assigned to a pair of elements (points in an n-dimensional vector space).
- b)
- Dii = 0 ∀i = 1, 2, ......N; N = number of elements
- c)
- Distance does not depend on the direction of measurement, i.e. Dij = Dji
- d)
- Distance satisfies the triangular inequality, i.e. Dij ≤ Dik + Dkj
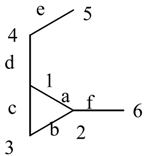
2. Basic Definitions
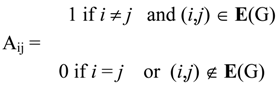

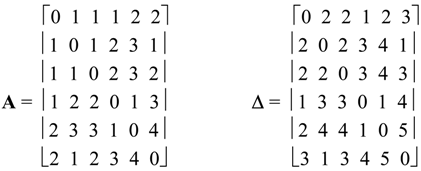
3. Results and Discussion
| Hydrocarbon | Nc | 0χ | 1χ | 2χ | M1 | M2 | W | H | J | MTI | W* | H* | J* | MTI* |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1. ethane | 2 | 2.000 | 2.000 | 0.000 | 2 | 2 | 1 | 1.000 | 1.000 | 4 | 1 | 1.000 | 1.000 | 4 |
| 2. propane | 3 | 2.707 | 2.828 | 1.414 | 6 | 8 | 4 | 2.500 | 1.633 | 16 | 4 | 2.500 | 1.633 | 16 |
| 3. butane | 4 | 3.414 | 3.828 | 2.000 | 10 | 16 | 10 | 4.333 | 1.975 | 38 | 10 | 4.333 | 1.975 | 38 |
| 4. 2-methylpropane | 4 | 3.577 | 3.464 | 3.464 | 12 | 18 | 9 | 4.500 | 2.324 | 36 | 9 | 4.500 | 2.324 | 36 |
| 5. pentane | 5 | 4.121 | 4.828 | 2.707 | 14 | 24 | 20 | 6.417 | 2.191 | 74 | 20 | 6.417 | 2.191 | 74 |
| 6. 2-methylbutane | 5 | 4.284 | 4.540 | 3.604 | 16 | 28 | 18 | 6.667 | 2.540 | 68 | 18 | 6.667 | 2.540 | 68 |
| 7. 2,2-dimethyl-propane | 5 | 4.500 | 4.000 | 6.000 | 20 | 32 | 16 | 7.000 | 3.024 | 64 | 16 | 7.000 | 3.024 | 64 |
| 8. hexane | 6 | 4.828 | 5.828 | 3.414 | 18 | 32 | 35 | 8.700 | 2.339 | 128 | 35 | 8.700 | 2.339 | 128 |
| 9. 2-methylpentane | 6 | 4.992 | 5.540 | 4.365 | 20 | 36 | 32 | 9.000 | 2.627 | 118 | 32 | 9.000 | 2.627 | 118 |
| 10. 3-methylpentane | 6 | 4.992 | 5.616 | 3.843 | 20 | 38 | 31 | 9.083 | 2.754 | 114 | 31 | 9.083 | 2.754 | 114 |
| 11. 2,2-dimethyl-butane | 6 | 5.207 | 5.121 | 5.828 | 24 | 44 | 28 | 9.500 | 3.168 | 106 | 28 | 9.500 | 3.168 | 106 |
| 12. 2,3-dimethyl-butane | 6 | 5.155 | 5.285 | 4.976 | 22 | 42 | 29 | 9.333 | 2.993 | 108 | 29 | 9.333 | 2.993 | 108 |
| 13. heptane | 7 | 5.536 | 6.828 | 4.121 | 22 | 40 | 56 | 11.150 | 2.447 | 204 | 56 | 11.150 | 2.447 | 204 |
| 14. 3-methylhexane | 7 | 5.699 | 6.616 | 4.604 | 24 | 46 | 50 | 11.617 | 2.832 | 182 | 50 | 11.617 | 2.832 | 182 |
| 15. 2,2-dimethyl-pentane | 7 | 5.914 | 6.121 | 6.621 | 28 | 52 | 46 | 12.083 | 3.154 | 170 | 46 | 12.083 | 3.154 | 170 |
| 16. 2,2,3-trime-thylbutane | 7 | 6.077 | 5.887 | 7.041 | 30 | 60 | 42 | 12.500 | 3.541 | 156 | 42 | 12.500 | 3.541 | 156 |
| 17. 3-methylheptane | 8 | 6.406 | 7.616 | 5.311 | 28 | 54 | 76 | 14.267 | 2.862 | 276 | 76 | 14.267 | 2.862 | 276 |
| 18. 4-methylheptane | 8 | 6.406 | 7.616 | 5.365 | 28 | 54 | 75 | 14.317 | 2.920 | 272 | 75 | 14.317 | 2.920 | 272 |
| 19. 2,2-dimethylhexane | 8 | 6.621 | 7.121 | 7.328 | 23 | 60 | 71 | 14.767 | 3.112 | 260 | 71 | 14.767 | 3.112 | 260 |
| 20. 2,3-dimethylhexane | 8 | 6.569 | 7.361 | 6.020 | 30 | 60 | 70 | 14.733 | 3.171 | 254 | 70 | 14.733 | 3.171 | 254 |
| 21. 2,4-dimethylhexane | 8 | 6.569 | 7.328 | 6.286 | 30 | 58 | 71 | 14.650 | 3.099 | 258 | 71 | 14.650 | 3.099 | 258 |
| 22. 2,5-dimethylhexane | 8 | 6.569 | 7.252 | 6.730 | 30 | 56 | 74 | 14.467 | 2.928 | 270 | 74 | 14.467 | 2.928 | 270 |
| 23. 3,3-dimethylhexane | 8 | 6.621 | 7.243 | 6.536 | 32 | 64 | 67 | 15.033 | 3.373 | 244 | 67 | 15.033 | 3.373 | 244 |
| 24. 1,3-dime-thylbenzene | 8 | 5.983 | 7.575 | 6.754 | 36 | 76 | 61 | 16.083 | 2.231 | 268 | 123 | 7.833 | 1.071 | 516 |
| 25. 1,2-dime-thylbenzene | 8 | 5.983 | 7.609 | 6.478 | 36 | 78 | 60 | 16.167 | 2.279 | 264 | 124 | 7.810 | 1.061 | 520 |
| 26. 1,4-dime-thylbenzene | 8 | 5.983 | 7.575 | 6.730 | 36 | 76 | 62 | 16.033 | 2.192 | 272 | 122 | 7.867 | 1.082 | 512 |
| 27.1-methyl-3-ethylbenzene | 9 | 6.690 | 8.651 | 7.092 | 40 | 86 | 88 | 19.150 | 2.232 | 373 | 164 | 10.295 | 1.153 | 677 |
| 28.1-methyl-4-ethylbenzene | 9 | 6.690 | 8.651 | 7.068 | 40 | 86 | 90 | 19.067 | 2.180 | 381 | 162 | 10.52 | 1.171 | 669 |
| 29. 1,2,3-trime-thylbenzene | 9 | 6.853 | 8.430 | 7.489 | 42 | 94 | 82 | 19.667 | 2.413 | 349 | 164 | 10.102 | 1.152 | 677 |
| 30. 1,2,4-trime-thylbenzene | 9 | 6.853 | 8.397 | 7.746 | 42 | 92 | 84 | 19.533 | 2.346 | 357 | 162 | 10.160 | 1.168 | 669 |
| 31. 1,3,5-triethyl-benzene | 9 | 6.853 | 8.363 | 8.045 | 42 | 90 | 84 | 19.500 | 2.341 | 357 | 162 | 10.150 | 1.167 | 669 |
| 32. 1,3-triethyl-benzene | 10 | 7.397 | 9.727 | 7.430 | 44 | 96 | 121 | 22.383 | 2.246 | 500 | 213 | 12.882 | 1.224 | 868 |
| 33. 1,4-diethyl-benzene | 10 | 7.397 | 9.727 | 7.406 | 44 | 96 | 125 | 22.243 | 2.174 | 516 | 209 | 12.981 | 1.254 | 852 |
| 34.1,2,3,4-tetrame-thylbenzene | 10 | 7.724 | 9.252 | 8.500 | 48 | 110 | 109 | 23.367 | 2.516 | 452 | 209 | 12.595 | 1.249 | 852 |
| 35. 1,2,3-triethyl-benzene | 12 | 8.975 | 11.658 | 8.679 | 54 | 124 | 190 | 30.233 | 2.524 | 760 | 338 | 18.192 | 1.343 | 1352 |
| 36. 1,2,4-triethyl-benzene | 12 | 8.975 | 11.625 | 8.848 | 54 | 122 | 198 | 29.876 | 2.413 | 792 | 330 | 18.365 | 1.379 | 1320 |
| 37. 1,3,5-triethyl-benzene | 12 | 8.975 | 11.591 | 9.060 | 54 | 120 | 198 | 29.800 | 2.405 | 792 | 330 | 18.339 | 1.377 | 1320 |
| 38.1,2,3,4-tetrae-thylbenzene | 14 | 10.55 | 13.556 | 10.116 | 64 | 150 | 287 | 38.543 | 2.674 | 1124 | 483 | 24.252 | 1.504 | 1908 |
| 39.1,2,3,5-tetrae-thylbenzene | 14 | 10.55 | 13.522 | 10.309 | 64 | 148 | 291 | 38.326 | 2.631 | 1140 | 479 | 24.326 | 1.517 | 1892 |
| 40.1,2,4,5-tetra-ethylbenzene | 14 | 10.55 | 13.522 | 10.289 | 64 | 148 | 295 | 38.186 | 2.592 | 1156 | 475 | 24.425 | 1.530 | 1876 |
| 41. 1-methylna-phthalene | 11 | 7.682 | 10.754 | 9.233 | 56 | 130 | 140 | 27.850 | 1.993 | 646 | 426 | 8.024 | 0.626 | 1898 |
| 42. 2-methylna-phthalene | 11 | 7.682 | 10.720 | 9.446 | 56 | 128 | 144 | 27.633 | 1.932 | 664 | 424 | 8.049 | 0.629 | 1890 |
| 43. 1-ethylna-phthalene | 12 | 8.389 | 11.830 | 9.615 | 60 | 140 | 182 | 31.533 | 1.987 | 816 | 518 | 10.444 | 0.662 | 2280 |
| 44. 2-ethylna-phthalene | 12 | 8.389 | 11.796 | 9.748 | 60 | 138 | 190 | 31.176 | 1.895 | 852 | 514 | 10.489 | 0.667 | 2264 |
| 45. 1,2-dimethyl-naphthalene | 12 | 8.552 | 11.575 | 10.244 | 62 | 146 | 178 | 31.917 | 2.027 | 800 | 516 | 10.166 | 0.664 | 2270 |
| 46. 1,3-dimethyl-naphthalene | 12 | 8.552 | 11.542 | 10.525 | 62 | 144 | 179 | 31.833 | 2.015 | 804 | 515 | 10.175 | 0.665 | 2266 |
| 47. 1,4-dimethyl-naphthalene | 12 | 8.552 | 11.575 | 10.288 | 62 | 146 | 176 | 32.000 | 2.055 | 790 | 516 | 10.161 | 0.664 | 2270 |
| 48. 1,5-dimethyl-naphthalene | 12 | 8.552 | 11.575 | 10.308 | 62 | 146 | 175 | 32.050 | 2.066 | 786 | 517 | 10.150 | 0.663 | 2274 |
| 49. 1,6-dimethyl-naphthalene | 12 | 8.552 | 11.542 | 10.501 | 62 | 144 | 180 | 31.783 | 2.003 | 808 | 514 | 10.186 | 0.667 | 2262 |
| 50. 1,7-dimethyl-naphthalene | 12 | 8.552 | 11.542 | 10.501 | 62 | 144 | 181 | 31.750 | 1.992 | 812 | 515 | 10.175 | 0.665 | 2266 |
| 51. 2,3-dimethyl-naphthalene | 12 | 8.552 | 11.542 | 10.461 | 62 | 144 | 182 | 31.700 | 1.976 | 818 | 514 | 10.191 | 0.667 | 2262 |
| 52. 2,6-dimethyl-naphthalene | 12 | 8.552 | 11.508 | 10.713 | 62 | 142 | 185 | 31.533 | 1.944 | 830 | 511 | 10.225 | 0.671 | 2250 |
| 53. 2,7-dimethyl-naphthalene | 12 | 8.552 | 11.508 | 10.713 | 62 | 142 | 186 | 31.510 | 1.936 | 834 | 512 | 10.211 | 0.669 | 2254 |
| 54. 1-propylna-phthalene | 13 | 9.096 | 12.830 | 10.376 | 64 | 148 | 236 | 35.110 | 1.940 | 1036 | 622 | 13.111 | 0.695 | 2712 |
| 55. 2-propylna-phthalene | 13 | 9.096 | 12.790 | 10.545 | 64 | 146 | 248 | 34.654 | 1.837 | 1090 | 616 | 13.173 | 0.702 | 2688 |
| 56. 2-ethyl-3-methylnaphthalene | 13 | 9.259 | 12.618 | 10.843 | 66 | 154 | 232 | 35.493 | 1.965 | 1020 | 616 | 12.714 | 0.701 | 2682 |
| 57. 2-ethyl-6-me-thylnaphthalene | 13 | 9.259 | 12.584 | 11.051 | 66 | 152 | 238 | 35.219 | 1.915 | 1044 | 610 | 12.776 | 0.708 | 2658 |
| 58. 2-ethyl-7-me-thylnaphthalene | 13 | 9.259 | 12.584 | 11.051 | 66 | 152 | 240 | 35.177 | 1.902 | 1052 | 612 | 12.751 | 0.706 | 2666 |
| 59. 1-butylna-phthalene | 14 | 9.803 | 13.830 | 11.083 | 68 | 156 | 303 | 38.646 | 1.876 | 1310 | 739 | 15.957 | 0.726 | 3198 |
| 60. 2-butylna-phthalene | 14 | 9.803 | 13.796 | 11.252 | 68 | 154 | 319 | 38.117 | 1.772 | 1382 | 731 | 16.033 | 0.734 | 3166 |
| Descriptors | First order | Second order | Third order | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Molecular set 1 | R | s | F | R | s | F | R | s | F |
| J | 0.77724 | 17.5335 | 89 | 0.80464 | 16.8340 | 52 | 0.88161 | 13.4999 | 65 |
| 0χ, M1 | 0.97498 | 6.1944 | 548 | 0.97904 | 5.8789 | 317 | 0.98079 | 5.7355 | 223 |
| 0χ, 1χ, M1 | 0.98938 | 4.0506 | 865 | 0.99807 | 1.8269 | 2279 | 0.99817 | 1.8316 | 1511 |
| Nc, 1χ, M1, H | 0.99210 | 3.4968 | 859 | 0.99675 | 2.4147 | 975 | 0.99728 | 2.3017 | 716 |
| Nc, 0χ, M1, H, MTI | 0.99637 | 2.3729 | 1479 | 0.99849 | 1.6802 | 1618 | 0.99941 | 1.1113 | 2470 |
| Molecular set 2 | |||||||||
| J* | 0.95134 | 8.5866 | 553 | 0.98210 | 5.3400 | 774 | 0.98346 | 5.1812 | 550 |
| 1χ, H* | 0.97910 | 5.6678 | 660 | 0.98620 | 4.7786 | 487 | 0.98733 | 4.6685 | 341 |
| M1, H*, J* | 0.99409 | 3.02624 | 1564 | 0.99652 | 2.4500 | 1263 | 0.99710 | 2.3052 | 952 |
| W, J, MTI, J* | 0.99719 | 2.0868 | 2438 | 0.99779 | 1.9920 | 1436 | 0.99891 | 1.4588 | 1790 |
| Nc, 0χ, M1, J, J* | 0.99906 | 1.2103 | 5715 | 0.99937 | 1.0811 | 3914 | 0.99959 | 0.9292 | 3534 |
| Molecule | ΔHof (exp) [45/ | ΔHof (theor)(1) | ΔHof (theor)(2) |
|---|---|---|---|
| 1. | -20.24 | -20.27 | -19.89 |
| 2. | -24.82 | -24.90 | -25.25 |
| 3. | -30.15 | -29.43 | -30.31 |
| 4. | -32.15 | -32.97 | -32.28 |
| 5. | -35.00 | -34.48 | -34.75 |
| 6. | -36.92 | -37.05 | -37.49 |
| 7. | -39.67 | -39.31 | -38.72 |
| 8. | -39.96 | -40.11 | -38.86 |
| 9. | -41.66 | -41.93 | -41.76 |
| 10. | -41.02 | -41.51 | -42.63 |
| 11. | -44.35 | -43.44 | -43.62 |
| 12. | -42.49 | -43.76 | -44.38 |
| 13. | -44.88 | -45.80 | -42.77 |
| 14. | -45.96 | -46.38 | -46.75 |
| 15. | -49.27 | -48.23 | -48.17 |
| 16. | -48.95 | -49.64 | -48.74 |
| 17. | -50.82 | -50.68 | -50.48 |
| 18. | -50.69 | -50.45 | -50.75 |
| 19. | -53.71 | -52.66 | -52.41 |
| 20 | -51.13 | -51.22 | -53.31 |
| 21. | -52.44 | -51.46 | -53.22 |
| 22. | -53.21 | -52.19 | -52.69 |
| 23. | -52.61 | -51.63 | -52.30 |
| 24. | 4.12 | 5.31 | 4.15 |
| 25. | 4.54 | 5.79 | 4.64 |
| 26. | 4.29 | 4.83 | 3.61 |
| 27. | -0.46 | -0.78 | -0.40 |
| 28. | -0.78 | -1.56 | -1.25 |
| 29. | -2.29 | -4.03 | -2.30 |
| 30. | -3.33 | -4.83 | -2.76 |
| 31. | -3.84 | -4.83 | -2.73 |
| 32. | -5.22 | -6.46 | -4.81 |
| 33. | -5.32 | -7.63 | -6.14 |
| 34. | -10.02 | -11.66 | -11.32 |
| 35. | -16.25 | -14.98 | -16.55 |
| 36. | -16.99 | -16.52 | -17.25 |
| 37. | -17.86 | -16.52 | -17.17 |
| 38. | -29.46 | -28.32 | -29.42 |
| 39. | -29.36 | -29.52 | -29.32 |
| 40. | -29.46 | -30.76 | -29.34 |
| 41. | 27.93 | 30.08 | 27.71 |
| 42. | 27.75 | 29.97 | 26.88 |
| 43. | 23.10 | 22.41 | 23.74 |
| 44. | 22.92 | 22.69 | 22.41 |
| 45. | 19.97 | 20.38 | 20.34 |
| 46. | 19.55 | 20.04 | 20.18 |
| 47. | 19.72 | 19.59 | 20.62 |
| 48. | 19.55 | 19.93 | 20.76 |
| 49. | 19.72 | 19.70 | 19.99 |
| 50. | 19.55 | 19.36 | 19.92 |
| 51. | 19.97 | 20.49 | 19.68 |
| 52. | 19.72 | 19.49 | 19.17 |
| 53. | 19.72 | 19.16 | 19.12 |
| 54. | 17.85 | 16.66 | 18.89 |
| 55. | 17.65 | 17.78 | 17.22 |
| 56. | 15.72 | 15.76 | 14.99 |
| 57. | 14.65 | 14.23 | 14.15 |
| 58. | 14.65 | 13.72 | 14.02 |
| 59. | 12.68 | 11.73 | 13.53 |
| 60. | 12.50 | 13.91 | 11.65 |
| Average absolute deviation | - | 0.76 | 0.62 |
4. Conclusions
References
- Balaban, A. T. (Ed.) Chemical Applications of Graph Theory; Academic Press: London, 1976.
- King, R. B. (Ed.) In Chemical Applications of Topology and Graph Theory; Studies in Physical and Theoretical Chemistry 28. Elsevier: Amsterdam, 1983.
- Balasubramanian, K. Chem. Rev. 1985, 85, 599.
- Trinajstic, N. Mathematics and Computational Concepts in Chemistry; Ellis Horwood: Chichester, 1986. [Google Scholar]
- King, R. B.; Rouvray, D. H. (Eds.) In Graph Theory and Topology in Chemistry; Studies in Physical and Theoretical Chemistry. Elsevier: Amsterdam, 1987.
- Lacher, L. C. (Ed.) In MATH/CHEM/COMP 1987; Studies in Physical and Theoretical Chemistry 54. Elsevier: Amsterdam, 1988.
- Rouvray, D. H. (Ed.) Computational Graph Theory; Nova Sci. Publ.: New York, 1990.
- Klein, D. J.; Randic, M. (Eds.) Mathematical Chemistry; VCH: Weinheim, 1990.
- Graovac, A. (Ed.) In MATH/CHEM/COMP 1988; Studies in Physical and Theoretical Chemistry 63. Elsevier: Amsterdam, 1989.
- Trinajstic, N. Chemical Graph Theory; CRC Press: Boca Raton, FL, 1992. [Google Scholar]
- Balaban, A. T.; Motoc, I.; Bonchev, D.; Mekenyan, O. Top. Curr. Chem. 1983, 114, 21.
- Randic, M. J. Math. Chem. 1992, 9, 97.
- Katritzky, A. R.; Gordeeva, E. V. J. Chem. Inf. Comput. Sci. 1993, 33, 835. [CrossRef]
- Basak, S. C.; Magnuson, V. R.; Niemi, J. G.; Regal, R. R.; Vetih, G. D. Math. Modell. 1986, 8, 300. [CrossRef]
- Randic, M. Croat. Chem. Acta 1993, 66, 289.
- Randic, M.; Trinajstic, N. J. Mol. Struct. THEOCHEM 1993, 284, 209. [CrossRef]
- Ivanciuc, O.; Ivanciuc, T.; Diudea, M. V. SAR/QSAR Environm. Res. 1997, 7, 63.
- Randic, M.; Kleiner, A. F.; DeAlba, L. M. J. Chem. Inf. Comput. Sci. 1994, 34, 277. [CrossRef]
- Harary, F. Graph Theory; Addison-Wesley: Reading, MA, 1969; p. 203. [Google Scholar]
- Buckely, F.; Harary, F. Distance in Graphs; Addison-Wesley: Reading, MA, 1990; pp. 213–214. [Google Scholar]
- Ivanciuc, O.; Balaban, A. T. Commun. Math. Chem. (MATCH) 1994, 30, 141.
- Amic, D.; Trinajstic, N. Croat. Chem. Acta 1995, 68, 53.
- Trinajstic, N.; Nikolic, S.; Lucic, B.; Amic, D.; Mihalic, Z. J. Chem. Inf. Comput. Sci. 1997, 37, 631.
- Rücker, G.; Rücker, C. J. Chem. Inf. Comput. Sci. 1998, 38, 710.
- Firpo, M.; Gavernet, L.; Castro, E. A.; Toropov, A. A. J. Mol. Struct . THEOCHEM 2000, 501-502, 419. [CrossRef]
- Castro, E. A.; Tueros, M.; Toropov, A. A. Comp. & Chem. 2000, 24, 571.
- Tueros, M.; Castro, E. A.; Toropov, A. A. J. Mol. Model. (in press)
- Wiener, H. J. Am. Chem. Soc. 1947, 69, 17, 1947, 69, 2336; J. Chem. Phys. 1947, 15, 766; J. Phys. Chem. 1948, 52, 425; 1948, 52, 1082.
- Plavsic, D; Nikolic, S.; Trinajstic, N.; Mihalic, Z. J. Math. Chem. 1993, 12, 235. [CrossRef]
- Schultz, H. P. J. Chem. Inf. Comput. Sci. 1989, 29, 237.
- Mueller, W. R.; Szymanski, K.; Knop, J. V.; Trinajstic, N. J. Chem. Inf. Comput. Sci. 1990, 30, 169. [CrossRef]
- Randic, M. J. Am. Chem. Soc. 1975, 97, 6609. [CrossRef]
- Balaban, A. T. Chem. Phys. 1982, 89, 399.
- Balaban, A. T. J. Chem. Inf. Comput. Sci. 1985, 25, 334. [CrossRef]
- Kier, L. B.; Hall, H. Molecular Connectivity in Structure-Activity Analysis; Research Studies Press: Ukm, Letchworth, Hertfordshire, 1986. [Google Scholar]
- Gutman, I.; Trinajstic, N. Chem. Phys. Lett. 1972, 17, 535.
- Gutman, I.; Ruscic, B.; Trinajstic, N.; Wilcox, Jr. C. F. J. Chem. Phys. 1975, 62, 3399.
- Toropov, A. A.; Toropova, A. P.; Ismailov, T. T.; Bonchev, D. J. Mol. Struct. THEOCHEM 1998, 424, 237.
- Dearden, J. C. Chem. Intell.Lab. Syst. 1994, 24, 77.
- Basak, S. C.; Niemi, G. J.; Veith, G. D. J. Math. Chem. 1991, 7, 243. [CrossRef]
- Randic, M.; Trinajstic, N. New J. Chem. 1994, 18, 179.
- Castro, E. A. Comput. Chem. 1997, 21, 305.
- Mercader, A.; Castro, E. A.; Toropov, A. A. Chem. Phys. Lett. 2000, 330, 612.
- Herndon, W. C. Chem. Phys.Lett. 1995, 234, 82.
- Stull, D. R.; et al. The Chemical Thermodynamics of Organic Compounds; Wiley: New York, 1969. [Google Scholar]
- Pedley, J. D.; et al. Thermochemical Data of Organic Compounds, 2nd. ed.; Chapman and Hall, 1986. [Google Scholar]
- Cox, J. D.; Pilcher, G. Thermochemistry of Organic and Organometallic Compounds; Academic Press: London, 1970. [Google Scholar]
- Pedley, J. D.; et al. Computer Analysed Thermochemical Data: Organic and Organometallic Compounds; University of Sussex: Sussex, 1977. [Google Scholar]
- Wagman, D.D.; et al. Chem. Ref. Data Suppl. 1982, No 222, 11.
- Benson, S. W. Thermochemical Kinetics, 2nd. ed.; Wiley: New York, 1987. [Google Scholar]
- Lowry, T. H.; Richardson, K. S. Mechanism and Theory in Organic Chemistry, 3rd. ed.; Harper & Row: New York, 1987. [Google Scholar]
- Habibollahzadeh, D.; et al. J. Comput. Chem. 1995, 16, 654. [CrossRef]
- Ibrahim, M. B.; Schleyer, P. v. R. J. Comput. Chem. 1985, 6, 157. [CrossRef]
- Yala, Z. J. Molec. Struct. THEOCHEM 1990, 207, 217. [CrossRef]
- Castro, E. A. J. Molec. Struct. THEOCHEM 1994, 304, 93. [CrossRef]
- Castro, E. A. J. Molec. Struct. THEOCHEM 1995, 339, 239. [CrossRef]
- Dewar, J. M. S.; Storch, D. M. J. Am. Chem. Soc. 1985, 107, 3898. [CrossRef]
- Castro, E. A. J. Chem. Soc. Pak. 1995, 17, 156.
- Vericat, C.; Castro, E. A. Commun. Math. Comp. Chem. MATCH 1996, 34, 167.
- Vericat, C.; Castro, E. A. Egyp. J. Chem. 1998, 41, 109.
- Lukovits, I. Croat. Chem. Acta 1995, 68, 873.
- Lukovits, I. J. Chem. Inf. Comput. Sci. 1996, 36, 65.
- Zhu, H. –Y.; Klein, D. J.; Lukovits, I. J. Chem. Inf. Comput. Sci. 1996, 36, 420. [CrossRef]
© 2001 by MDPI (http://www.mdpi.org), Basel, Switzerland.
Share and Cite
Mercader, A.; Castro, E.A.; Toropov, A.A. Maximum Topological Distances Based Indices as Molecular Descriptors for QSPR. 4. Modeling the Enthalpy of Formation of Hydrocarbons from Elements. Int. J. Mol. Sci. 2001, 2, 121-132. https://doi.org/10.3390/i2020121
Mercader A, Castro EA, Toropov AA. Maximum Topological Distances Based Indices as Molecular Descriptors for QSPR. 4. Modeling the Enthalpy of Formation of Hydrocarbons from Elements. International Journal of Molecular Sciences. 2001; 2(2):121-132. https://doi.org/10.3390/i2020121
Chicago/Turabian StyleMercader, Andrés, Eduardo A. Castro, and Andrey A. Toropov. 2001. "Maximum Topological Distances Based Indices as Molecular Descriptors for QSPR. 4. Modeling the Enthalpy of Formation of Hydrocarbons from Elements" International Journal of Molecular Sciences 2, no. 2: 121-132. https://doi.org/10.3390/i2020121




