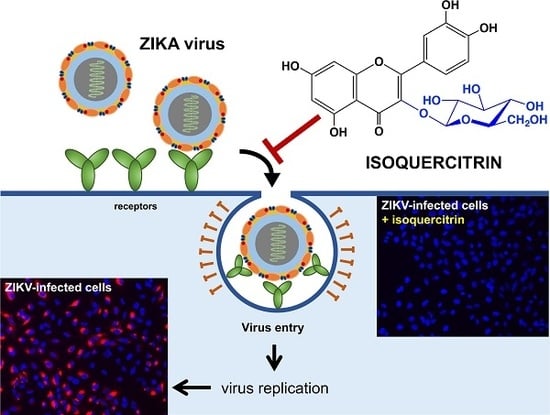
The Flavonoid Isoquercitrin Precludes Initiation of Zika Virus Infection in Human Cells
Abstract
:1. Introduction
2. Results and Discussion
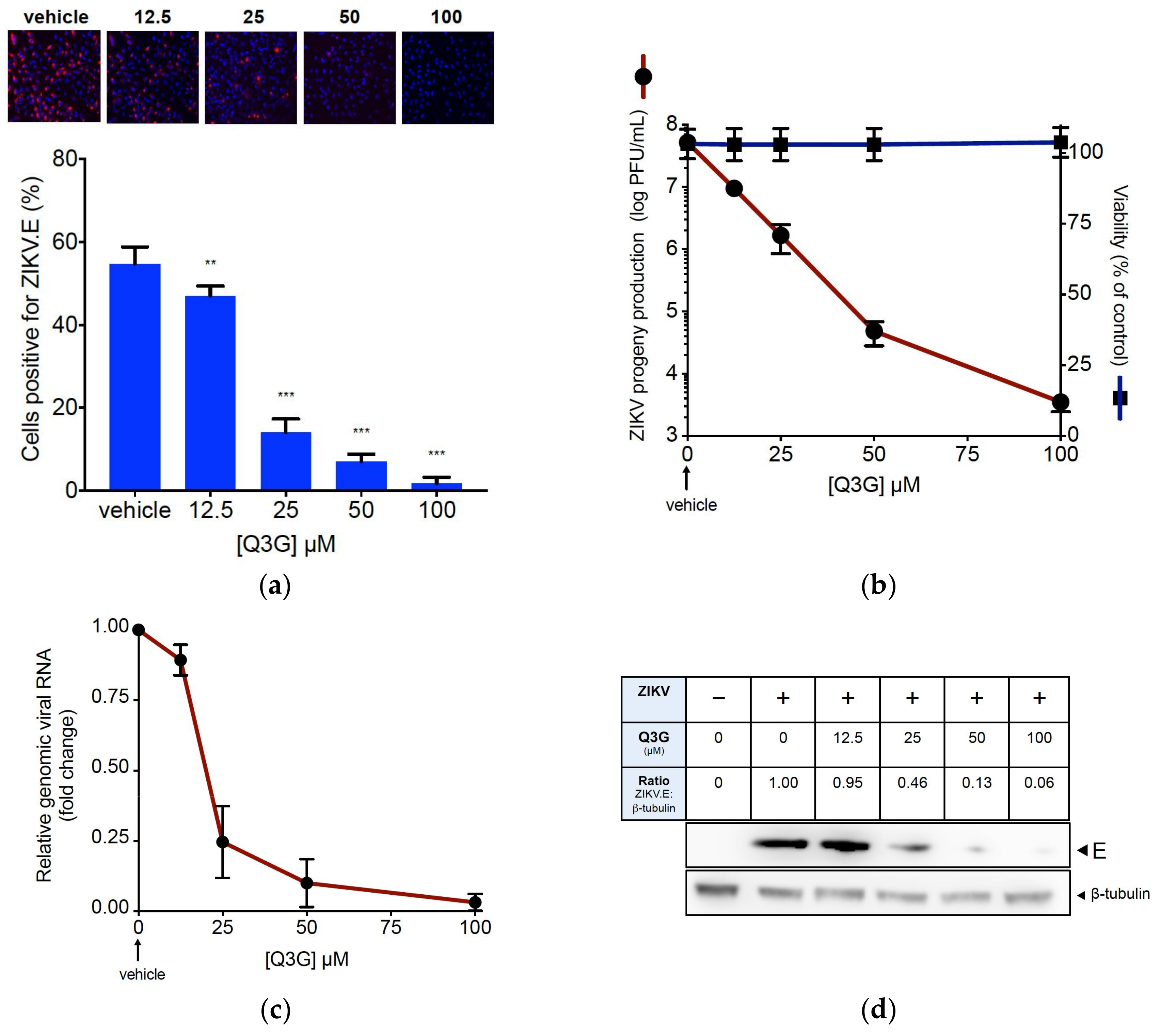
2.1. Q3G Precludes ZIKV Infection in Human Cells of Various Tissue Origin
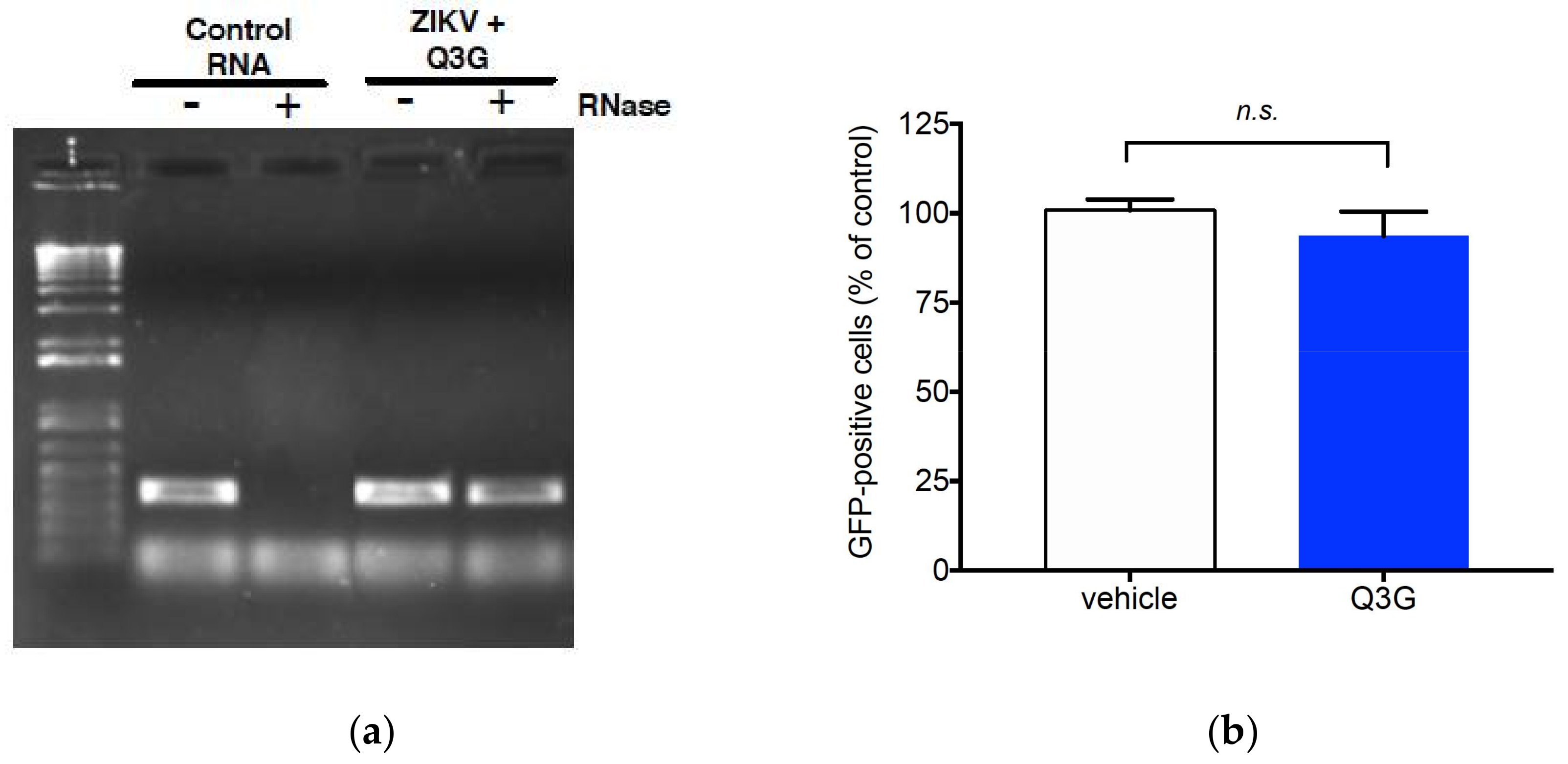
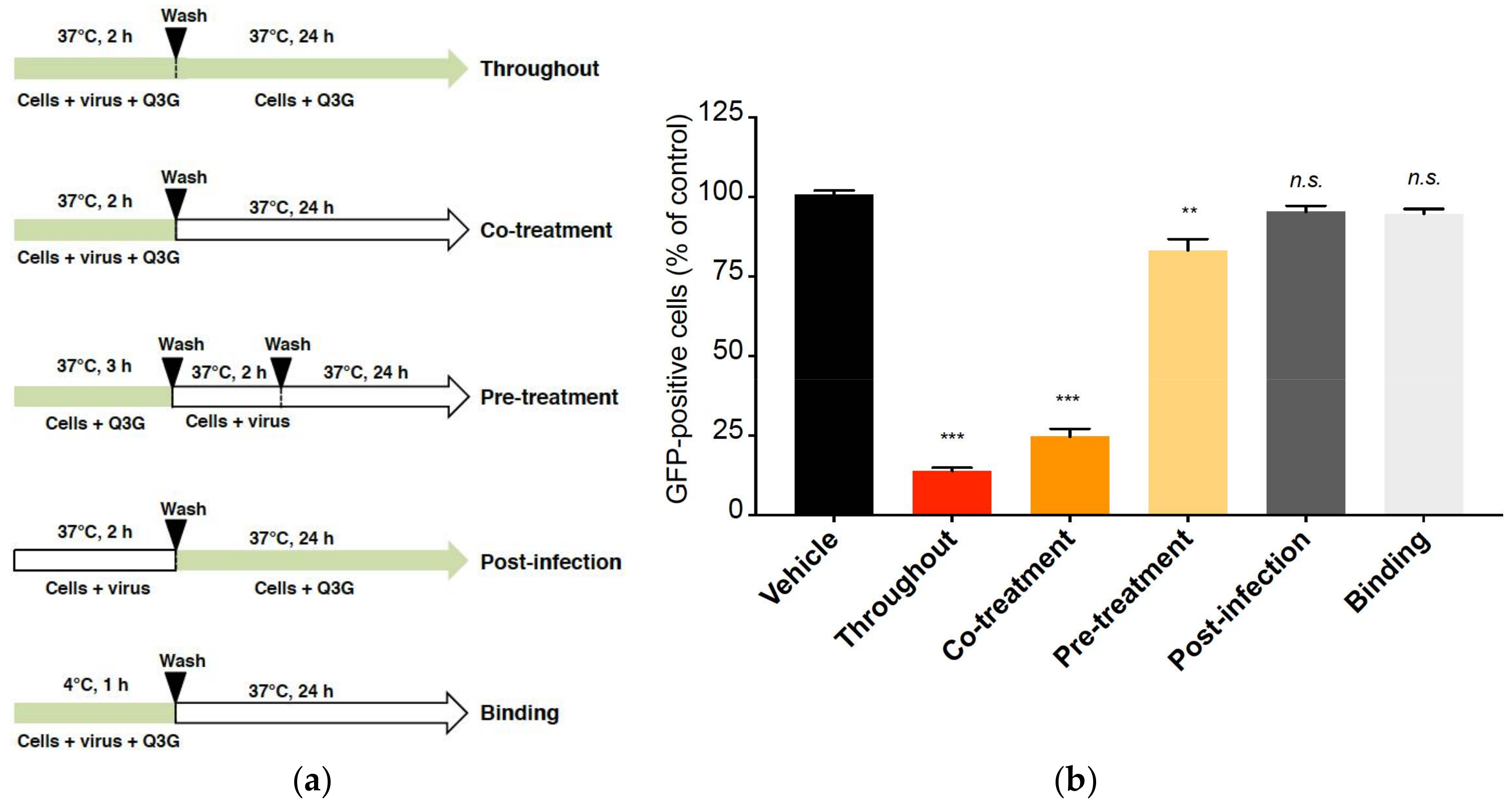
2.2. Q3G Targets Early Stages of ZIKV Infection
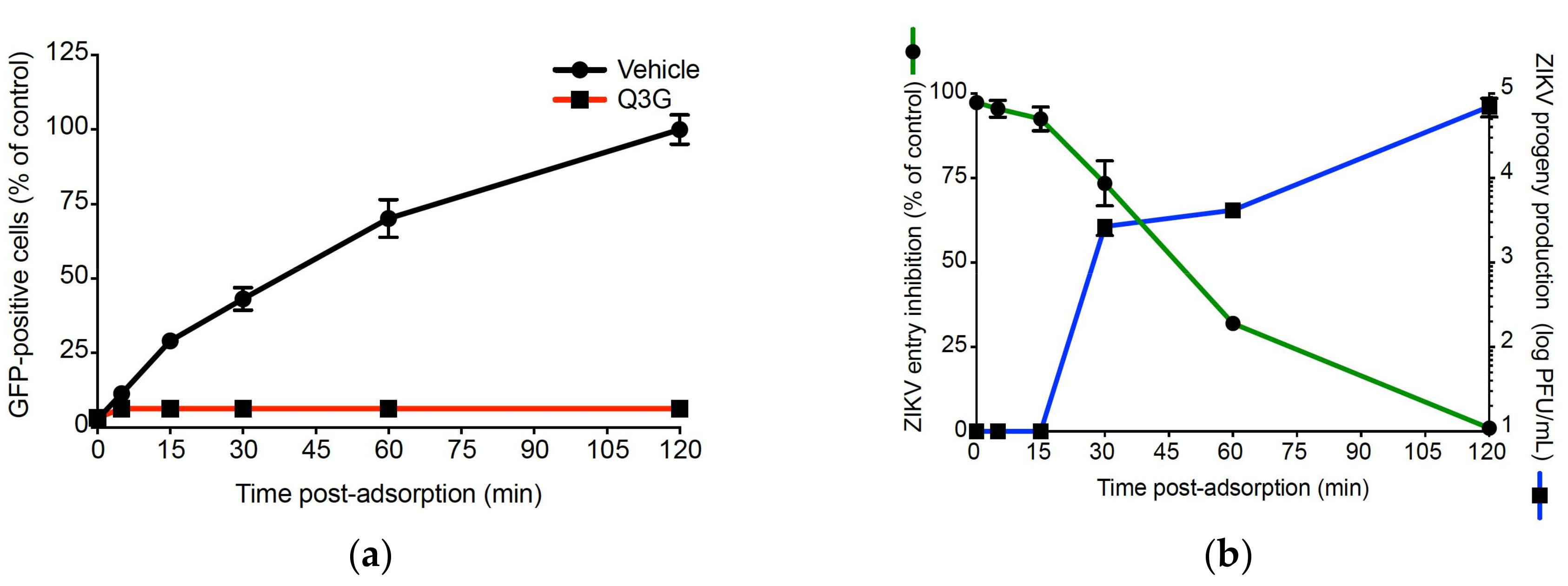
2.3. Q3G Inhibits ZIKV Internalisation in A549 Cells
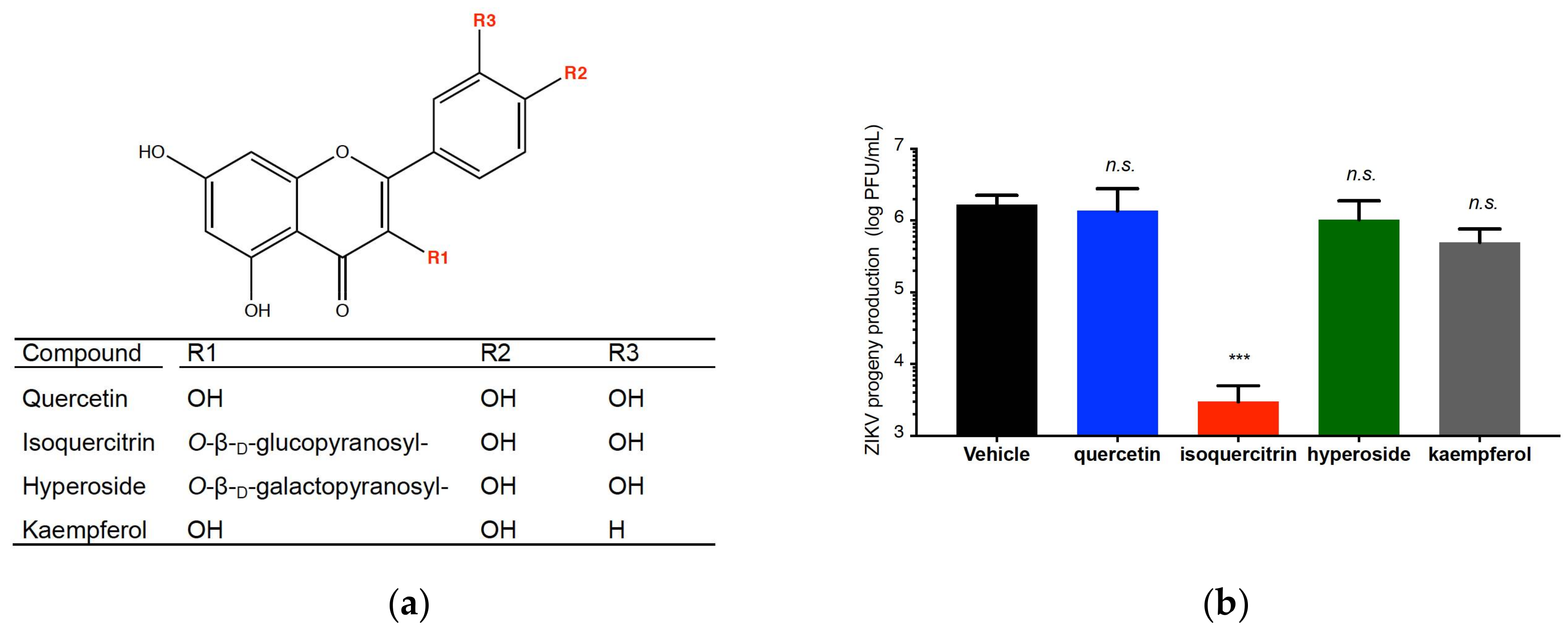
2.4. Quercetin, Hyperoside, and Kaempferol Are Inefficient against ZIKV
2.5. Concluding Remarks
3. Materials and Methods
3.1. Cells, Virus, and Reagents
3.2. Cytotoxicity Assay
3.3. Immunofluorescence and Flow Cytometry Assays
3.4. Virus Binding and Internalisation Assays
3.5. RNase Protection Assay
3.6. Viral Inactivation Assay
3.7. RT-qPCR
3.8. Western Blot Analysis
3.9. Data Analysis
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| A549 | Human lung epithelial cell line |
| DAPI | 4′,6-diamidino-2-phenylindole |
| ECL | Enhanced chemiluminescent substrate |
| EGCG | Epigallocatechin gallate |
| FACS | Fluorescence-activated cell sorting |
| GAPDH | Glyceraldehyde-3-phosphate dehydrogenase |
| GFP | Green fluorescent protein |
| HCV | Hepatitis C virus |
| Huh-7 | Human hepatoma cell line |
| IF | Immunofluorescence |
| mAB | Monoclonal antibody |
| MOI | Multiplicity of infection |
| p.i. | post infection |
| PBS | Phosphate-buffered saline |
| PFU | Plaque forming unit |
| Q3G | Quercetin-3-O-glucoside or isoquercitrin |
| RIPA | Radio immune precipitation assay |
| SH-SY5Y | Human neuroblastoma cell line |
| ZIKV | Zika virus |
| ΔΔCt | Cycle threshold |
References
- Chen, J.; Liang, Y.; Yi, P.; Xu, L.; Hawkins, H.K.; Rossi, S.L.; Soong, L.; Cai, J.; Menon, R.; Sun, J. Outcomes of Congenital Zika Disease Depend on Timing of Infection and Maternal-Fetal Interferon Action. Cell Rep. 2017, 21, 1588–1599. [Google Scholar] [CrossRef] [PubMed]
- Moreira-Soto, A.; Sarno, M.; Pedroso, C.; Netto, E.M.; Rockstroh, A.; Luz, E.; Feldmann, M.; Fischer, C.; Bastos, F.A.; Kummerer, B.M.; et al. Evidence for Congenital Zika Virus Infection from Neutralizing Antibody Titers in Maternal Sera, Northeastern Brazil. J. Infect. Dis. 2017, 216, 1501–1504. [Google Scholar] [CrossRef] [PubMed]
- D’Ortenzio, E.; Matheron, S.; Yazdanpanah, Y.; de Lamballerie, X.; Hubert, B.; Piorkowski, G.; Maquart, M.; Descamps, D.; Damond, F.; Leparc-Goffart, I. Evidence of Sexual Transmission of Zika Virus. N. Engl. J. Med. 2016, 374, 2195–2198. [Google Scholar] [CrossRef] [PubMed]
- Duggal, N.K.; Ritter, J.M.; Pestorius, S.E.; Zaki, S.R.; Davis, B.S.; Chang, G.J.; Bowen, R.A.; Brault, A.C. Frequent Zika Virus Sexual Transmission and Prolonged Viral RNA Shedding in an Immunodeficient Mouse Model. Cell Rep. 2017, 18, 1751–1760. [Google Scholar] [CrossRef] [PubMed]
- Govero, J.; Esakky, P.; Scheaffer, S.M.; Fernandez, E.; Drury, A.; Platt, D.J.; Gorman, M.J.; Richner, J.M.; Caine, E.A.; Salazar, V.; et al. Zika virus infection damages the testes in mice. Nature 2016, 540, 438–442. [Google Scholar] [CrossRef] [PubMed]
- Ma, W.; Li, S.; Ma, S.; Jia, L.; Zhang, F.; Zhang, Y.; Zhang, J.; Wong, G.; Zhang, S.; Lu, X.; et al. Zika Virus Causes Testis Damage and Leads to Male Infertility in Mice. Cell 2017, 168, 542. [Google Scholar] [CrossRef] [PubMed]
- Hasan, S.S.; Sevvana, M.; Kuhn, R.J.; Rossmann, M.G. Structural biology of Zika virus and other flaviviruses. Nat. Struct. Mol. Biol. 2018, 25, 13–20. [Google Scholar] [CrossRef] [PubMed]
- Lindenbach, B.D.; Rice, C.M. Molecular biology of flaviviruses. Adv. Virus Res. 2003, 59, 23–61. [Google Scholar] [PubMed]
- Meertens, L.; Labeau, A.; Dejarnac, O.; Cipriani, S.; Sinigaglia, L.; Bonnet-Madin, L.; Le Charpentier, T.; Hafirassou, M.L.; Zamborlini, A.; Cao-Lormeau, V.M.; et al. Axl Mediates ZIKA Virus Entry in Human Glial Cells and Modulates Innate Immune Responses. Cell Rep. 2017, 18, 324–333. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Garcia, M.D.; Mazzon, M.; Jacobs, M.; Amara, A. Pathogenesis of flavivirus infections: Using and abusing the host cell. Cell Host Microbe 2009, 5, 318–328. [Google Scholar] [CrossRef] [PubMed]
- Haddow, A.D.; Schuh, A.J.; Yasuda, C.Y.; Kasper, M.R.; Heang, V.; Huy, R.; Guzman, H.; Tesh, R.B.; Weaver, S.C. Genetic characterization of Zika virus strains: Geographic expansion of the Asian lineage. PLoS Negl. Trop. Dis. 2012, 6, e1477. [Google Scholar] [CrossRef] [PubMed]
- Giovanetti, M.; Milano, T.; Alcantara, L.C.; Carcangiu, L.; Cella, E.; Lai, A.; Lo Presti, A.; Pascarella, S.; Zehender, G.; Angeletti, S.; et al. Zika Virus spreading in South America: Evolutionary analysis of emerging neutralizing resistant Phe279Ser strains. Asian Pac. J. Trop. Med. 2016, 9, 445–452. [Google Scholar] [CrossRef] [PubMed]
- Adcock, R.S.; Chu, Y.K.; Golden, J.E.; Chung, D.H. Evaluation of anti-Zika virus activities of broad-spectrum antivirals and NIH clinical collection compounds using a cell-based, high-throughput screen assay. Antivir. Res. 2017, 138, 47–56. [Google Scholar] [CrossRef] [PubMed]
- Barrows, N.J.; Campos, R.K.; Powell, S.T.; Prasanth, K.R.; Schott-Lerner, G.; Soto-Acosta, R.; Galarza-Munoz, G.; McGrath, E.L.; Urrabaz-Garza, R.; Gao, J.; et al. A Screen of FDA-Approved Drugs for Inhibitors of Zika Virus Infection. Cell Host Microbe 2016, 20, 259–270. [Google Scholar] [CrossRef] [PubMed]
- Saiz, J.C.; Martin-Acebes, M.A. The Race to Find Antivirals for Zika Virus. Antimicrob. Agents Chemother. 2017, 61. [Google Scholar] [CrossRef] [PubMed]
- Julander, J.G.; Siddharthan, V.; Evans, J.; Taylor, R.; Tolbert, K.; Apuli, C.; Stewart, J.; Collins, P.; Gebre, M.; Neilson, S.; et al. Efficacy of the broad-spectrum antiviral compound BCX4430 against Zika virus in cell culture and in a mouse model. Antivir. Res. 2017, 137, 14–22. [Google Scholar] [CrossRef] [PubMed]
- Rausch, K.; Hackett, B.A.; Weinbren, N.L.; Reeder, S.M.; Sadovsky, Y.; Hunter, C.A.; Schultz, D.C.; Coyne, C.B.; Cherry, S. Screening Bioactives Reveals Nanchangmycin as a Broad Spectrum Antiviral Active against Zika Virus. Cell Rep. 2017, 18, 804–815. [Google Scholar] [CrossRef] [PubMed]
- Sacramento, C.Q.; de Melo, G.R.; de Freitas, C.S.; Rocha, N.; Hoelz, L.V.; Miranda, M.; Fintelman-Rodrigues, N.; Marttorelli, A.; Ferreira, A.C.; Barbosa-Lima, G.; et al. The clinically approved antiviral drug sofosbuvir inhibits Zika virus replication. Sci. Rep. 2017, 7, 40920. [Google Scholar] [CrossRef] [PubMed]
- Estoppey, D.; Lee, C.M.; Janoschke, M.; Lee, B.H.; Wan, K.F.; Dong, H.; Mathys, P.; Filipuzzi, I.; Schuhmann, T.; Riedl, R.; et al. The Natural Product Cavinafungin Selectively Interferes with Zika and Dengue Virus Replication by Inhibition of the Host Signal Peptidase. Cell Rep. 2017, 19, 451–460. [Google Scholar] [CrossRef] [PubMed]
- Lani, R.; Hassandarvish, P.; Shu, M.H.; Phoon, W.H.; Chu, J.J.; Higgs, S.; Vanlandingham, D.; Abu Bakar, S.; Zandi, K. Antiviral activity of selected flavonoids against Chikungunya virus. Antivir. Res. 2016, 133, 50–61. [Google Scholar] [CrossRef] [PubMed]
- Martin, K.W.; Ernst, E. Antiviral agents from plants and herbs: A systematic review. Antivir. Ther. 2003, 8, 77–90. [Google Scholar] [CrossRef] [PubMed]
- Dos Santos, A.E.; Kuster, R.M.; Yamamoto, K.A.; Salles, T.S.; Campos, R.; de Meneses, M.D.; Soares, M.R.; Ferreira, D. Quercetin and quercetin 3-O-glycosides from Bauhinia longifolia (Bong.) Steud. show anti-Mayaro virus activity. Parasit Vectors 2014, 7, 130. [Google Scholar] [CrossRef] [PubMed]
- Khachatoorian, R.; Arumugaswami, V.; Raychaudhuri, S.; Yeh, G.K.; Maloney, E.M.; Wang, J.; Dasgupta, A.; French, S.W. Divergent antiviral effects of bioflavonoids on the hepatitis C virus life cycle. Virology 2012, 433, 346–355. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.; Narayanan, S.; Chang, K.O. Inhibition of influenza virus replication by plant-derived isoquercetin. Antivir. Res. 2010, 88, 227–235. [Google Scholar] [CrossRef] [PubMed]
- Lim, H.J.; Nguyen, T.T.; Kim, N.M.; Park, J.S.; Jang, T.S.; Kim, D. Inhibitory effect of flavonoids against NS2B-NS3 protease of ZIKA virus and their structure activity relationship. Biotechnol. Lett. 2017, 39, 415–421. [Google Scholar] [CrossRef] [PubMed]
- Lulu, S.S.; Thabitha, A.; Vino, S.; Priya, A.M.; Rout, M. Naringenin and quercetin--potential anti-HCV agents for NS2 protease targets. Nat. Prod. Res. 2016, 30, 464–468. [Google Scholar] [CrossRef] [PubMed]
- Havsteen, B.H. The biochemistry and medical significance of the flavonoids. Pharmacol. Ther. 2002, 96, 67–202. [Google Scholar] [CrossRef]
- Verri, W.A.; Vicentini, F.T.M.C.; Baracat, M.M.; Georgetti, S.R.; Cardoso, R.D.R.; Cunha, T.M.; Ferreira, S.H.; Cunha, F.Q.; Fonseca, M.J.V.; Casagrande, R. Flavonoids as Anti-Inflammatory and Analgesic Drugs: Mechanisms of Action and Perspectives in the Development of Pharmaceutical Forms. In Studies in Natural Products Chemistry; Atta ur, R., Ed.; Elsevier: Amsterdam, The Netherlands, 2012; Volume 36, pp. 297–330. [Google Scholar]
- Abdal Dayem, A.; Choi, H.Y.; Kim, Y.B.; Cho, S.G. Antiviral effect of methylated flavonol isorhamnetin against influenza. PLoS ONE 2015, 10, e0121610. [Google Scholar] [CrossRef]
- Calland, N.; Sahuc, M.E.; Belouzard, S.; Pene, V.; Bonnafous, P.; Mesalam, A.A.; Deloison, G.; Descamps, V.; Sahpaz, S.; Wychowski, C.; et al. Polyphenols Inhibit Hepatitis C Virus Entry by a New Mechanism of Action. J. Virol. 2015, 89, 10053–10063. [Google Scholar] [CrossRef] [PubMed]
- Cotin, S.; Calliste, C.A.; Mazeron, M.C.; Hantz, S.; Duroux, J.L.; Rawlinson, W.D.; Ploy, M.C.; Alain, S. Eight flavonoids and their potential as inhibitors of human cytomegalovirus replication. Antivir. Res. 2012, 96, 181–186. [Google Scholar] [CrossRef] [PubMed]
- Frabasile, S.; Koishi, A.C.; Kuczera, D.; Silveira, G.F.; Verri, W.A., Jr.; Duarte Dos Santos, C.N.; Bordignon, J. The citrus flavanone naringenin impairs dengue virus replication in human cells. Sci. Rep. 2017, 7, 41864. [Google Scholar] [CrossRef] [PubMed]
- Grienke, U.; Richter, M.; Walther, E.; Hoffmann, A.; Kirchmair, J.; Makarov, V.; Nietzsche, S.; Schmidtke, M.; Rollinger, J.M. Discovery of prenylated flavonoids with dual activity against influenza virus and Streptococcus pneumoniae. Sci. Rep. 2016, 6, 27156. [Google Scholar] [CrossRef] [PubMed]
- Mounce, B.C.; Cesaro, T.; Carrau, L.; Vallet, T.; Vignuzzi, M. Curcumin inhibits Zika and chikungunya virus infection by inhibiting cell binding. Antivir. Res. 2017, 142, 148–157. [Google Scholar] [CrossRef] [PubMed]
- Sharma, N.; Murali, A.; Singh, S.K.; Giri, R. Epigallocatechin gallate, an active green tea compound inhibits the Zika virus entry into host cells via binding the envelope protein. Int. J. Biol. Macromol. 2017, 104, 1046–1054. [Google Scholar] [CrossRef] [PubMed]
- Wong, G.; He, S.; Siragam, V.; Bi, Y.; Mbikay, M.; Chretien, M.; Qiu, X. Antiviral activity of quercetin-3-beta-O-d-glucoside against Zika virus infection. Virol. Sin. 2017, 32, 545–547. [Google Scholar] [CrossRef] [PubMed]
- Frumence, E.; Roche, M.; Krejbich-Trotot, P.; El-Kalamouni, C.; Nativel, B.; Rondeau, P.; Misse, D.; Gadea, G.; Viranaicken, W.; Despres, P. The South Pacific epidemic strain of Zika virus replicates efficiently in human epithelial A549 cells leading to IFN-β production and apoptosis induction. Virology 2016, 493, 217–226. [Google Scholar] [CrossRef] [PubMed]
- Gadea, G.; Bos, S.; Krejbich-Trotot, P.; Clain, E.; Viranaicken, W.; El-Kalamouni, C.; Mavingui, P.; Despres, P. A robust method for the rapid generation of recombinant Zika virus expressing the GFP reporter gene. Virology 2016, 497, 157–162. [Google Scholar] [CrossRef] [PubMed]
- Bos, S.; Viranaicken, W.; Turpin, J.; El-Kalamouni, C.; Roche, M.; Krejbich-Trotot, P.; Despres, P.; Gadea, G. The structural proteins of epidemic and historical strains of Zika virus differ in their ability to initiate viral infection in human host cells. Virology 2018, 516, 265–273. [Google Scholar] [CrossRef] [PubMed]
- Visintini Jaime, M.F.; Redko, F.; Muschietti, L.V.; Campos, R.H.; Martino, V.S.; Cavallaro, L.V. In vitro antiviral activity of plant extracts from Asteraceae medicinal plants. Virol. J. 2013, 10, 245. [Google Scholar] [CrossRef] [PubMed]
- Valentova, K.; Vrba, J.; Bancirova, M.; Ulrichova, J.; Kren, V. Isoquercitrin: Pharmacology, toxicology, and metabolism. Food Chem. Toxicol. 2014, 68, 267–282. [Google Scholar] [CrossRef] [PubMed]
- Qiu, X.; Kroeker, A.; He, S.; Kozak, R.; Audet, J.; Mbikay, M.; Chretien, M. Prophylactic Efficacy of Quercetin 3-beta-O-d-Glucoside against Ebola Virus Infection. Antimicrob. Agents Chemother. 2016, 60, 5182–5188. [Google Scholar] [CrossRef] [PubMed]
- Hamel, R.; Dejarnac, O.; Wichit, S.; Ekchariyawat, P.; Neyret, A.; Luplertlop, N.; Perera-Lecoin, M.; Surasombatpattana, P.; Talignani, L.; Thomas, F.; et al. Biology of Zika Virus Infection in Human Skin Cells. J. Virol. 2015, 89, 8880–8896. [Google Scholar] [CrossRef] [PubMed]
- Talarico, L.B.; Pujol, C.A.; Zibetti, R.G.; Faria, P.C.; Noseda, M.D.; Duarte, M.E.; Damonte, E.B. The antiviral activity of sulfated polysaccharides against dengue virus is dependent on virus serotype and host cell. Antivir. Res. 2005, 66, 103–110. [Google Scholar] [CrossRef] [PubMed]
- Koishi, A.C.; Zanello, P.R.; Bianco, E.M.; Bordignon, J.; Nunes Duarte dos Santos, C. Screening of Dengue virus antiviral activity of marine seaweeds by an in situ enzyme-linked immunosorbent assay. PLoS ONE 2012, 7, e51089. [Google Scholar] [CrossRef] [PubMed]
- Lin, L.T.; Chen, T.Y.; Chung, C.Y.; Noyce, R.S.; Grindley, T.B.; McCormick, C.; Lin, T.C.; Wang, G.H.; Lin, C.C.; Richardson, C.D. Hydrolyzable tannins (chebulagic acid and punicalagin) target viral glycoprotein-glycosaminoglycan interactions to inhibit herpes simplex virus 1 entry and cell-to-cell spread. J. Virol. 2011, 85, 4386–4398. [Google Scholar] [CrossRef] [PubMed]
- Viranaicken, W.; Nativel, B.; Krejbich-Trotot, P.; Harrabi, W.; Bos, S.; El Kalamouni, C.; Roche, M.; Gadea, G.; Despres, P. ClearColi BL21(DE3)-based expression of Zika virus antigens illustrates a rapid method of antibody production against emerging pathogens. Biochimie 2017, 142, 179–182. [Google Scholar] [CrossRef] [PubMed]
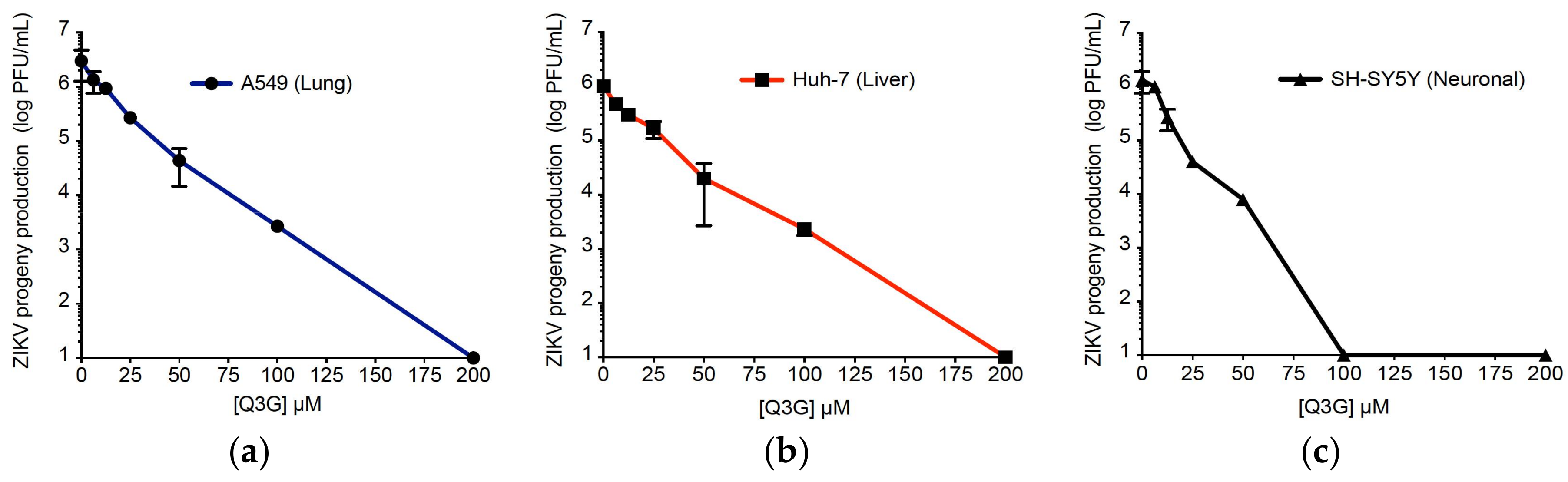
| Human Cell Lines | CC50 (µM) a | IC50 (µM) b | IC90 (µM) c | SI d |
|---|---|---|---|---|
| A549 | 551.2 ± 43.2 | 15.5 ± 2.3 | 32.0 ± 3.4 | 35.6 |
| Huh-7 | 326.8 ± 45.7 | 14.0 ± 3.8 | 50.0 ± 4.7 | 23.3 |
| SH-SY5Y | 582.2 ± 41.4 | 9.7 ± 1.2 | 15.0 ± 2.3 | 60.0 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gaudry, A.; Bos, S.; Viranaicken, W.; Roche, M.; Krejbich-Trotot, P.; Gadea, G.; Desprès, P.; El-Kalamouni, C. The Flavonoid Isoquercitrin Precludes Initiation of Zika Virus Infection in Human Cells. Int. J. Mol. Sci. 2018, 19, 1093. https://doi.org/10.3390/ijms19041093
Gaudry A, Bos S, Viranaicken W, Roche M, Krejbich-Trotot P, Gadea G, Desprès P, El-Kalamouni C. The Flavonoid Isoquercitrin Precludes Initiation of Zika Virus Infection in Human Cells. International Journal of Molecular Sciences. 2018; 19(4):1093. https://doi.org/10.3390/ijms19041093
Chicago/Turabian StyleGaudry, Arnaud, Sandra Bos, Wildriss Viranaicken, Marjolaine Roche, Pascale Krejbich-Trotot, Gilles Gadea, Philippe Desprès, and Chaker El-Kalamouni. 2018. "The Flavonoid Isoquercitrin Precludes Initiation of Zika Virus Infection in Human Cells" International Journal of Molecular Sciences 19, no. 4: 1093. https://doi.org/10.3390/ijms19041093
APA StyleGaudry, A., Bos, S., Viranaicken, W., Roche, M., Krejbich-Trotot, P., Gadea, G., Desprès, P., & El-Kalamouni, C. (2018). The Flavonoid Isoquercitrin Precludes Initiation of Zika Virus Infection in Human Cells. International Journal of Molecular Sciences, 19(4), 1093. https://doi.org/10.3390/ijms19041093