Molecular and Immunohistochemical Markers with Prognostic and Predictive Significance in Liver Metastases from Colorectal Carcinoma
Abstract
1. Introduction
2. Results
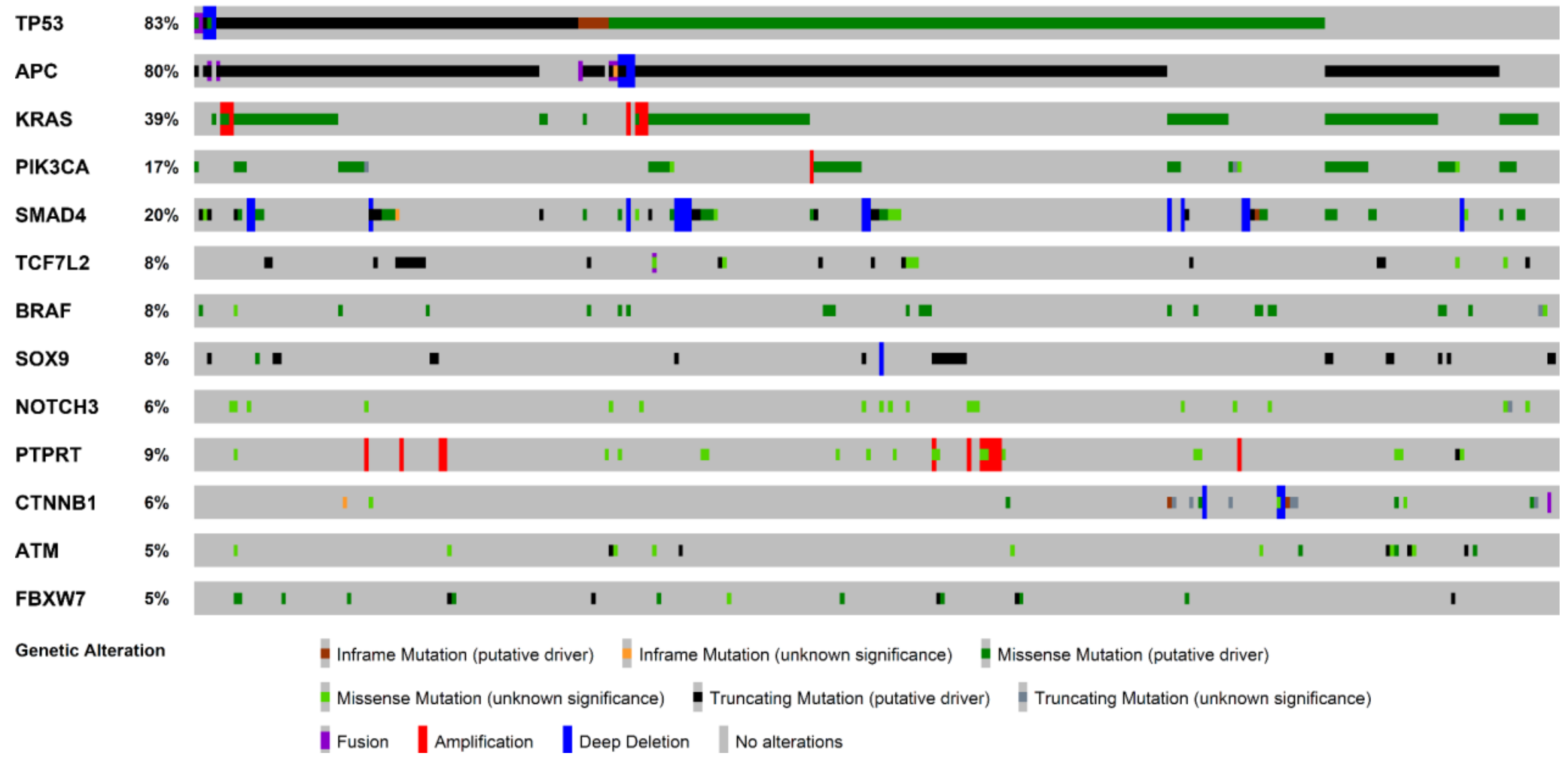
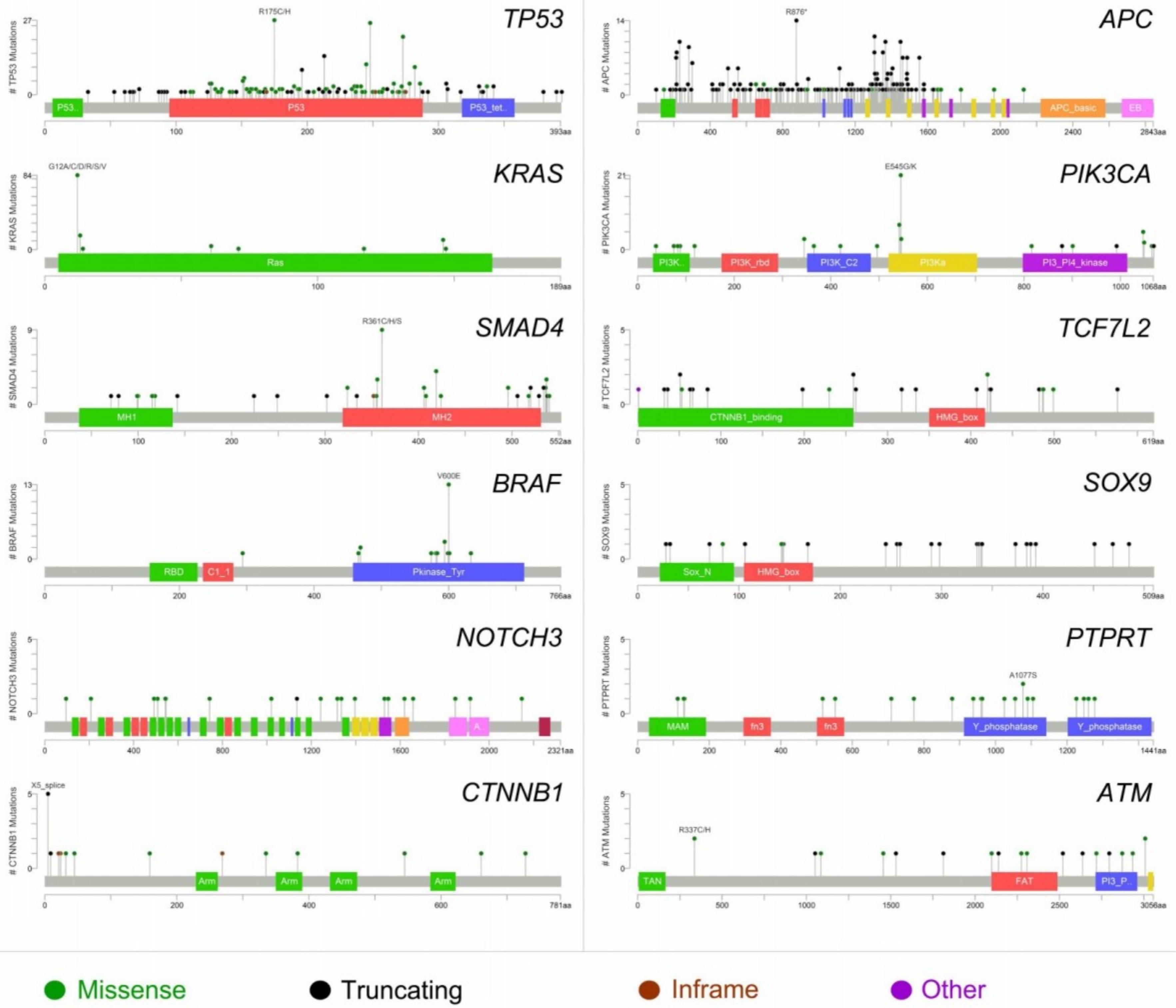
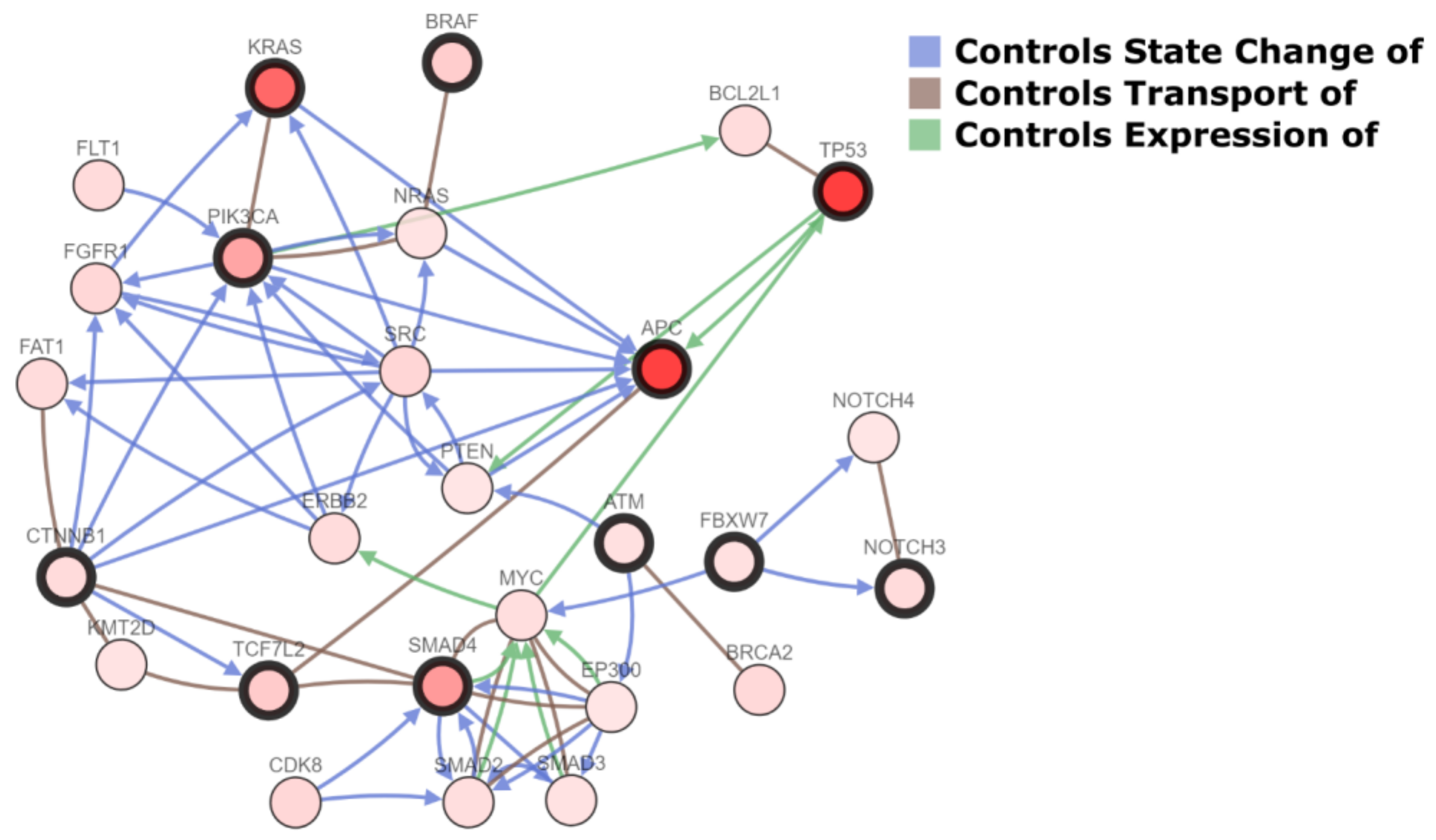
2.1. Molecular Landscape and Clonal Evolution Phenomena in CRCLM
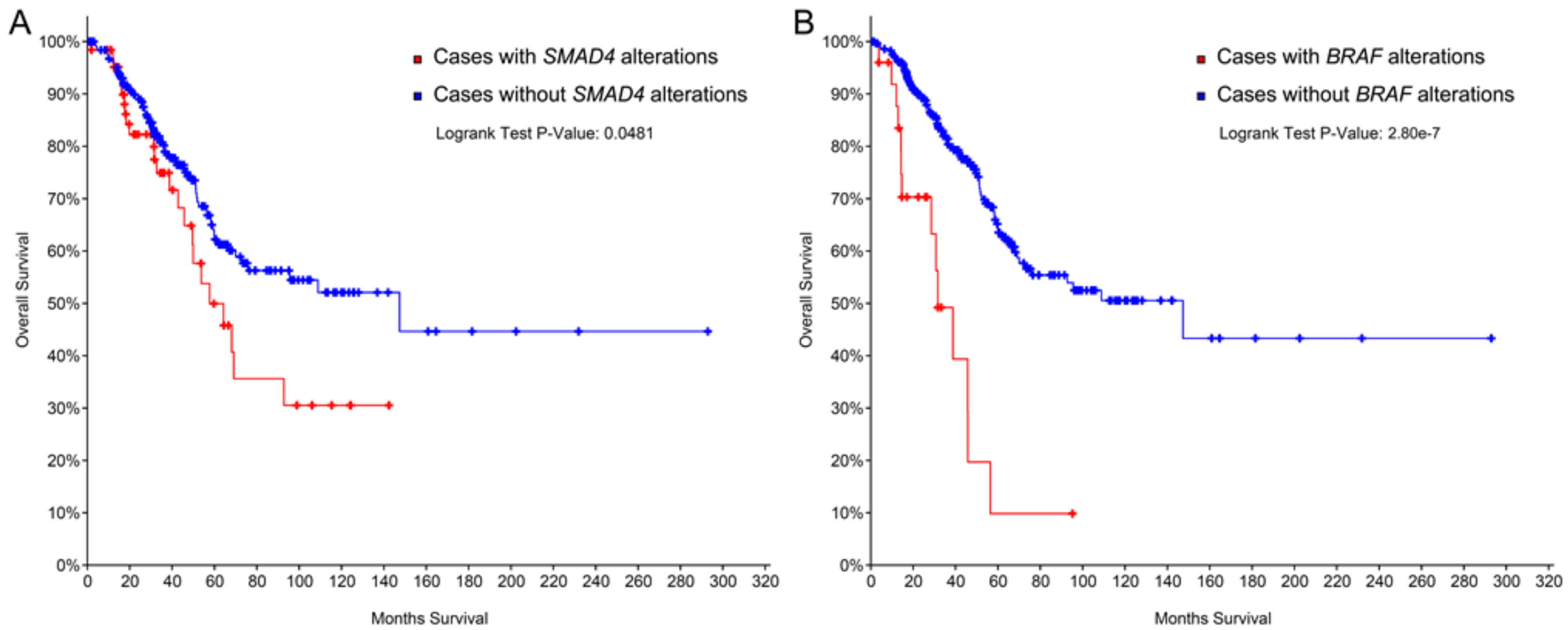
2.2. The Prognostic Role of SMAD4, KRAS, and BRAF.
2.3. miRNA Expression Profiling
2.4. EMX2
2.5. DYRK2
2.6. Chromosome 20p11 Gains
2.7. GADD45B
2.8. CD133+ CD54+ CD44+ Circulating Tumor Cells
2.9. TGF-Beta
2.10. TBL1XR1
2.11. SDF1
2.12. Galectin-3
2.13. FRalpha
2.14. Ki67
2.15. MMP7
2.16. CCL15
2.17. pIgR
2.18. Endoglin
2.19. Nek2
2.20. Beta-Catenin
2.21. PPARG
2.22. MACC1
2.23. MET
2.24. CA9
2.25. Beta-1 Integrin
2.26. MLH1/PMS2
2.27. Tenascin C
3. Discussion
4. Materials and Methods
4.1. Inclusion Criteria
4.2. Search Terms
4.3. Exclusion Criteria
Author Contributions
Funding
Conflicts of Interest
References
- Global Cancer Facts & Figures 3rd Edition. Available online: https://www.cancer.org/research/cancer-facts-statistics/global.html (accessed on 12 July 2018).
- Ervik, M.; Lam, F.; Ferlay, J.; Mery, L.; Soerjomataram, I.; Bray, F. Cancer Today. Available online: http://gco.iarc.fr/today (accessed on 12 July 2018).
- Torre, L.A.; Bray, F.; Siegel, R.L.; Ferlay, J.; Lortet-Tieulent, J.; Jemal, A. Global cancer statistics, 2012. CA Cancer J. Clin. 2015, 65, 87–108. [Google Scholar] [CrossRef] [PubMed]
- Siegel, R.; DeSantis, C.; Virgo, K.; Stein, K.; Mariotto, A.; Smith, T.; Cooper, D.; Gansler, T.; Lerro, C.; Fedewa, S.; et al. Cancer treatment and survivorship statistics. CA Cancer J. Clin. 2012, 62, 220–241. [Google Scholar] [CrossRef] [PubMed]
- Lynch, M.L.; Brand, M.I. Preoperative evaluation and oncologic principles of colon cancer surgery. Clin. Colon Rectal Surg. 2005, 18, 163–173. [Google Scholar] [CrossRef] [PubMed]
- Veen, T.; Soreide, K. Can molecular biomarkers replace a clinical risk score for resectable colorectal liver metastasis? World J. Gastrointest. Oncol. 2017, 9, 98–104. [Google Scholar] [PubMed]
- Yamashita, S.; Chun, Y.S.; Kopetz, S.E.; Vauthey, J.N. Biomarkers in colorectal liver metastases. Br. J. Surg. 2018, 105, 618–627. [Google Scholar] [PubMed]
- Akgul, O.; Cetinkaya, E.; Ersoz, S.; Tez, M. Role of surgery in colorectal cancer liver metastases. World J. Gastroenterol. 2014, 20, 6113–6122. [Google Scholar] [CrossRef] [PubMed]
- Zarour, L.R.; Anand, S.; Billingsley, K.G.; Bisson, W.H.; Cercek, A.; Clarke, M.F.; Coussens, L.M.; Gast, C.E.; Geltzeiler, C.B.; Hansen, L.; et al. Colorectal cancer liver metastasis: Evolving paradigms and future directions. Cell. Mol. Gastroenterol. Hepatol. 2017, 3, 163–173. [Google Scholar] [CrossRef] [PubMed]
- Mogensen, M.B.; Rossing, M.; Ostrup, O.; Larsen, P.N.; Heiberg Engel, P.J.; Jorgensen, L.N.; Hogdall, E.V.; Eriksen, J.; Ibsen, P.; Jess, P.; et al. Genomic alterations accompanying tumour evolution in colorectal cancer: Tracking the differences between primary tumours and synchronous liver metastases by whole-exome sequencing. BMC Cancer 2018, 18, 752. [Google Scholar] [CrossRef] [PubMed]
- Polyak, K. Tumor heterogeneity confounds and illuminates: A case for darwinian tumor evolution. Nat. Med. 2014, 20, 344–346. [Google Scholar] [CrossRef] [PubMed]
- Ng, C.K.; Pemberton, H.N.; Reis-Filho, J.S. Breast cancer intratumor genetic heterogeneity: Causes and implications. Expert Rev. Anticancer Ther. 2012, 12, 1021–1032. [Google Scholar] [CrossRef] [PubMed]
- Fumagalli, C.; Bianchi, F.; Raviele, P.R.; Vacirca, D.; Bertalot, G.; Rampinelli, C.; Lazzeroni, M.; Bonanni, B.; Veronesi, G.; Fusco, N.; et al. Circulating and tissue biomarkers in early-stage non-small cell lung cancer. Ecancermedicalscience 2017, 11, 717. [Google Scholar] [CrossRef] [PubMed]
- Ercoli, G.; Lopez, G.; Ciapponi, C.; Corti, C.; Despini, L.; Gambini, D.; Runza, L.; Blundo, C.; Sciarra, A.; Fusco, N. Building up a high-throughput screening platform to assess the heterogeneity of her2 gene amplification in breast cancers. J. Vis. Exp. 2017, 13, 233–236. [Google Scholar] [CrossRef] [PubMed]
- Fusco, N.; Bosari, S. Her2 aberrations and heterogeneity in cancers of the digestive system: Implications for pathologists and gastroenterologists. World J. Gastroenterol. 2016, 22, 7926–7937. [Google Scholar] [CrossRef] [PubMed]
- Fusco, N.; Guerini-Rocco, E.; Del Conte, C.; Pellegrini, C.; Bulfamante, G.; Di Nuovo, F.; Romagnoli, S.; Bosari, S. Her2 in gastric cancer: A digital image analysis in pre-neoplastic, primary and metastatic lesions. Mod. Pathol. 2013, 26, 816–824. [Google Scholar] [CrossRef] [PubMed]
- Diaz, Z.; Aguilar-Mahecha, A.; Paquet, E.R.; Basik, M.; Orain, M.; Camlioglu, E.; Constantin, A.; Benlimame, N.; Bachvarov, D.; Jannot, G.; et al. Next-generation biobanking of metastases to enable multidimensional molecular profiling in personalized medicine. Mod. Pathol. 2013, 26, 1413–1424. [Google Scholar] [PubMed]
- Lopez-Gomez, M.; Moreno-Rubio, J.; Suarez-Garcia, I.; Cejas, P.; Madero, R.; Casado, E.; Jimenez, A.; Sereno, M.; Gomez-Raposo, C.; Zambrana, F.; et al. Smad4 and ts expression might predict the risk of recurrence after resection of colorectal liver metastases. Clin. Transl. Oncol. 2015, 17, 133–138. [Google Scholar] [CrossRef] [PubMed]
- Huemer, F.; Thaler, J.; Piringer, G.; Hackl, H.; Pleyer, L.; Hufnagl, C.; Weiss, L.; Greil, R. Sidedness and TP53 mutations impact os in anti-EGFR but not anti-VEGF treated mCRC—An analysis of the KRAS registry of the AGMT (arbeitsgemeinschaft medikamentose tumortherapie). BMC Cancer 2018, 18, 11. [Google Scholar] [CrossRef] [PubMed]
- Toth, C.; Sukosd, F.; Valicsek, E.; Herpel, E.; Schirmacher, P.; Renner, M.; Mader, C.; Tiszlavicz, L.; Kriegsmann, J. Expression of ERCC1, RRM1, TUBB3 in correlation with apoptosis repressor arc, DNA mismatch repair proteins and p53 in liver metastasis of colorectal cancer. Int. J. Mol. Med. 2017, 40, 1457–1465. [Google Scholar] [CrossRef] [PubMed]
- Pilat, N.; Grunberger, T.; Langle, F.; Mittlbock, M.; Perisanidis, B.; Kappel, S.; Wolf, B.; Starlinger, P.; Kuhrer, I.; Muhlbacher, F.; et al. Assessing the TP53 marker type in patients treated with or without neoadjuvant chemotherapy for resectable colorectal liver metastases: A p53 research group study. Eur. J. Surg. Oncol. 2015, 41, 683–689. [Google Scholar] [CrossRef] [PubMed]
- Viana Lde, S.; Affonso, R.J., Jr.; Silva, S.R.; Denadai, M.V.; Matos, D.; Salinas de Souza, C.; Waisberg, J. Relationship between the expression of the extracellular matrix genes SPARC, SPP1, FN1, ITGA5 and ITGAV and clinicopathological parameters of tumor progression and colorectal cancer dissemination. Oncology 2013, 84, 81–91. [Google Scholar] [CrossRef] [PubMed]
- Melucci, E.; Cosimelli, M.; Carpanese, L.; Pizzi, G.; Izzo, F.; Fiore, F.; Golfieri, R.; Giampalma, E.; Sperduti, I.; Ercolani, C.; et al. Decrease of survivin, p53 and Bcl-2 expression in chemorefractory colorectal liver metastases may be predictive of radiosensivity radiosensivity after radioembolization with yttrium-90 resin microspheres. J. Exp. Clin. Cancer Res. 2013, 32, 13. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.T.; Lee, J.; Park, S.H.; Park, J.O.; Lim, H.Y.; Kang, W.K.; Kim, J.Y.; Kim, Y.H.; Chang, D.K.; Rhee, P.L.; et al. Clinical impact of microsatellite instability in colon cancer following adjuvant folfox therapy. Cancer Chemother. Pharmacol. 2010, 66, 659–667. [Google Scholar] [CrossRef] [PubMed]
- Jones, R.P.; Brudvik, K.W.; Franklin, J.M.; Poston, G.J. Precision surgery for colorectal liver metastases: Opportunities and challenges of omics-based decision making. Eur. J. Surg. Oncol. 2017, 43, 875–883. [Google Scholar] [CrossRef] [PubMed]
- Lahti, S.J.; Xing, M.; Zhang, D.; Lee, J.J.; Magnetta, M.J.; Kim, H.S. Kras status as an independent prognostic factor for survival after yttrium-90 radioembolization therapy for unresectable colorectal cancer liver metastases. J. Vasc. Interv. Radiol. 2015, 26, 1102–1111. [Google Scholar] [CrossRef] [PubMed]
- Kidess-Sigal, E.; Liu, H.E.; Triboulet, M.M.; Che, J.; Ramani, V.C.; Visser, B.C.; Poultsides, G.A.; Longacre, T.A.; Marziali, A.; Vysotskaia, V.; et al. Enumeration and targeted analysis of KRAS, BRAF and PIK3CA mutations in CTCs captured by a label-free platform: Comparison to ctDNA and tissue in metastatic colorectal cancer. Oncotarget 2016, 7, 85349–85364. [Google Scholar] [CrossRef] [PubMed]
- Isella, C.; Mellano, A.; Galimi, F.; Petti, C.; Capussotti, L.; De Simone, M.; Bertotti, A.; Medico, E.; Muratore, A. Macc1 mRNA levels predict cancer recurrence after resection of colorectal cancer liver metastases. Ann. Surg. 2013, 257, 1089–1095. [Google Scholar] [CrossRef] [PubMed]
- Cbioportal for Cancer Genomics. Available online: http://www.cbioportal.org/public-portal/ (accessed on 20 September 2018).
- Korphaisarn, K.; Morris, V.K.; Overman, M.J.; Fogelman, D.R.; Kee, B.K.; Raghav, K.P.S.; Manuel, S.; Shureiqi, I.; Wolff, R.A.; Eng, C.; et al. Fbxw7 missense mutation: A novel negative prognostic factor in metastatic colorectal adenocarcinoma. Oncotarget 2017, 8, 39268–39279. [Google Scholar] [CrossRef] [PubMed]
- Sclafani, F.; Rimassa, L.; Colombo, P.; Destro, A.; Stinco, S.; Lutman, F.R.; Carnaghi, C.; Beretta, G.; Zanello, A.; Roncalli, M.; et al. An exploratory biomarker study in metastatic tumors from colorectal cancer patients treated with bevacizumab. Int. J. Biol. Mark. 2015, 30, e73–e80. [Google Scholar] [CrossRef] [PubMed]
- Kawakami, M.; Yamaguchi, T.; Takahashi, K.; Matsumoto, H.; Yasutome, M.; Horiguchi, S.; Hayashi, Y.; Funata, N.; Mori, T. Assessment of SMAD4, p53, and Ki-67 alterations as a predictor of liver metastasis in human colorectal cancer. Surg. Today 2010, 40, 245–250. [Google Scholar] [CrossRef] [PubMed]
- Boulay, J.L.; Mild, G.; Lowy, A.; Reuter, J.; Lagrange, M.; Terracciano, L.; Laffer, U.; Herrmann, R.; Rochlitz, C. SMAD4 is a predictive marker for 5-fluorouracil-based chemotherapy in patients with colorectal cancer. Br. J. Cancer 2002, 87, 630–634. [Google Scholar] [CrossRef] [PubMed]
- Richman, S.D.; Chambers, P.; Seymour, M.T.; Daly, C.; Grant, S.; Hemmings, G.; Quirke, P. Intra-tumoral heterogeneity of KRAS and BRAF mutation status in patients with advanced colorectal cancer (ACRC) and cost-effectiveness of multiple sample testing. Anal. Cell Pathol. 2011, 34, 61–66. [Google Scholar] [CrossRef]
- Karagkounis, G.; Torbenson, M.S.; Daniel, H.D.; Azad, N.S.; Diaz, L.A., Jr.; Donehower, R.C.; Hirose, K.; Ahuja, N.; Pawlik, T.M.; Choti, M.A. Incidence and prognostic impact of KRAS and BRAF mutation in patients undergoing liver surgery for colorectal metastases. Cancer 2013, 119, 4137–4144. [Google Scholar] [CrossRef] [PubMed]
- Tsilimigras, D.I.; Ntanasis-Stathopoulos, I.; Bagante, F.; Moris, D.; Cloyd, J.; Spartalis, E.; Pawlik, T.M. Clinical significance and prognostic relevance of KRAS, BRAF, PI3K and TP53 genetic mutation analysis for resectable and unresectable colorectal liver metastases: A systematic review of the current evidence. Surg. Oncol. 2018, 27, 280–288. [Google Scholar] [CrossRef] [PubMed]
- Cohen, R.; Buhard, O.; Cervera, P.; Hain, E.; Dumont, S.; Bardier, A.; Bachet, J.B.; Gornet, J.M.; Lopez-Trabada, D.; Kaci, R.; et al. Clinical and molecular characterisation of hereditary and sporadic metastatic colorectal cancers harbouring microsatellite instability/DNA mismatch repair deficiency. Eur. J. Cancer 2017, 86, 266–274. [Google Scholar] [CrossRef] [PubMed]
- Tosi, F.; Magni, E.; Amatu, A.; Mauri, G.; Bencardino, K.; Truini, M.; Veronese, S.; De Carlis, L.; Ferrari, G.; Nichelatti, M.; et al. Effect of KRAS and BRAF mutations on survival of metastatic colorectal cancer after liver resection: A systematic review and meta-analysis. Clin. Colorectal Cancer 2017, 16, e153–e163. [Google Scholar] [CrossRef] [PubMed]
- Pikoulis, E.; Margonis, G.A.; Andreatos, N.; Sasaki, K.; Angelou, A.; Polychronidis, G.; Pikouli, A.; Riza, E.; Pawlik, T.M.; Antoniou, E. Prognostic role of BRAF mutations in colorectal cancer liver metastases. Anticancer Res. 2016, 36, 4805–4811. [Google Scholar] [CrossRef] [PubMed]
- Osumi, H.; Shinozaki, E.; Suenaga, M.; Matsusaka, S.; Konishi, T.; Akiyoshi, T.; Fujimoto, Y.; Nagayama, S.; Fukunaga, Y.; Ueno, M.; et al. Ras mutation is a prognostic biomarker in colorectal cancer patients with metastasectomy. Int. J. Cancer 2016, 139, 803–811. [Google Scholar] [CrossRef] [PubMed]
- Passiglia, F.; Bronte, G.; Bazan, V.; Galvano, A.; Vincenzi, B.; Russo, A. Can KRAS and BRAF mutations limit the benefit of liver resection in metastatic colorectal cancer patients? A systematic review and meta-analysis. Crit. Rev. Oncol. Hematol. 2016, 99, 150–157. [Google Scholar] [CrossRef] [PubMed]
- Schirripa, M.; Bergamo, F.; Cremolini, C.; Casagrande, M.; Lonardi, S.; Aprile, G.; Yang, D.; Marmorino, F.; Pasquini, G.; Sensi, E.; et al. BRAF and RAS mutations as prognostic factors in metastatic colorectal cancer patients undergoing liver resection. Br. J. Cancer 2015, 112, 1921–1928. [Google Scholar] [CrossRef] [PubMed]
- Laszlo, L. Predictive and prognostic factors in the complex treatment of patients with colorectal cancer. Magy. Onkol. 2010, 54, 383–394. [Google Scholar] [CrossRef] [PubMed]
- Vaira, V.; Fedele, G.; Pyne, S.; Fasoli, E.; Zadra, G.; Bailey, D.; Snyder, E.; Faversani, A.; Coggi, G.; Flavin, R.; et al. Preclinical model of organotypic culture for pharmacodynamic profiling of human tumors. Proc. Natl. Acad. Sci. USA 2010, 107, 8352–8356. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Ou, Q.; Braciak, T.A.; Chen, G.; Oduncu, F.S. MicroRNA-192-5p is a predictive biomarker of survival for Stage IIIB colon cancer patients. Jpn. J. Clin. Oncol. 2018, 48, 619–624. [Google Scholar] [CrossRef] [PubMed]
- Forno, I.; Ferrero, S.; Russo, M.V.; Gazzano, G.; Giangiobbe, S.; Montanari, E.; Del Nero, A.; Rocco, B.; Albo, G.; Languino, L.R.; et al. Deregulation of mir-34b/sox2 predicts prostate cancer progression. PLoS ONE 2015, 10, e0130060. [Google Scholar] [CrossRef] [PubMed]
- Bleau, A.M.; Redrado, M.; Nistal-Villan, E.; Villalba, M.; Exposito, F.; Redin, E.; de Aberasturi, A.L.; Larzabal, L.; Freire, J.; Gomez-Roman, J.; et al. Mir-146a targets c-met and abolishes colorectal cancer liver metastasis. Cancer Lett. 2018, 414, 257–267. [Google Scholar] [CrossRef] [PubMed]
- Ling, H.; Pickard, K.; Ivan, C.; Isella, C.; Ikuo, M.; Mitter, R.; Spizzo, R.; Bullock, M.; Braicu, C.; Pileczki, V.; et al. The clinical and biological significance of mir-224 expression in colorectal cancer metastasis. Gut 2016, 65, 977–989. [Google Scholar] [CrossRef] [PubMed]
- Kingham, T.P.; Nguyen, H.C.B.; Zheng, J.; Konstantinidis, I.T.; Sadot, E.; Shia, J.; Kuk, D.; Zhang, S.; Saltz, L.; D’Angelica, M.I.; et al. MicroRNA-203 predicts human survival after resection of colorectal liver metastasis. Oncotarget 2017, 8, 18821–18831. [Google Scholar] [PubMed]
- Torres, S.; Garcia-Palmero, I.; Bartolome, R.A.; Fernandez-Acenero, M.J.; Molina, E.; Calvino, E.; Segura, M.F.; Casal, J.I. Combined miRNA profiling and proteomics demonstrates that different miRNAs target a common set of proteins to promote colorectal cancer metastasis. J. Pathol. 2017, 242, 39–51. [Google Scholar] [PubMed]
- Li, W.; Chang, J.; Tong, D.; Peng, J.; Huang, D.; Guo, W.; Zhang, W.; Li, J. Differential microRNA expression profiling in primary tumors and matched liver metastasis of patients with colorectal cancer. Oncotarget 2017, 8, 35783–35791. [Google Scholar] [CrossRef] [PubMed]
- Ji, D.; Chen, Z.; Li, M.; Zhan, T.; Yao, Y.; Zhang, Z.; Xi, J.; Yan, L.; Gu, J. MicroRNA-181a promotes tumor growth and liver metastasis in colorectal cancer by targeting the tumor suppressor wif-1. Mol. Cancer 2014, 13, 86. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.; Chen, H.; Zhou, D.; Li, D.; Bai, R.; Zheng, S.; Ge, W. MicroRNA-9 up-regulation is involved in colorectal cancer metastasis via promoting cell motility. Med. Oncol. 2012, 29, 1037–1043. [Google Scholar] [CrossRef] [PubMed]
- Tsukamoto, M.; Iinuma, H.; Yagi, T.; Matsuda, K.; Hashiguchi, Y. Circulating exosomal microRNA-21 as a biomarker in each tumor stage of colorectal cancer. Oncology 2017, 92, 360–370. [Google Scholar] [CrossRef] [PubMed]
- Fukushima, Y.; Iinuma, H.; Tsukamoto, M.; Matsuda, K.; Hashiguchi, Y. Clinical significance of microRNA-21 as a biomarker in each dukes’ stage of colorectal cancer. Oncol. Rep. 2015, 33, 573–582. [Google Scholar] [CrossRef] [PubMed]
- Shibuya, H.; Iinuma, H.; Shimada, R.; Horiuchi, A.; Watanabe, T. Clinicopathological and prognostic value of microRNA-21 and microRNA-155 in colorectal cancer. Oncology 2010, 79, 313–320. [Google Scholar] [CrossRef] [PubMed]
- Aykut, B.; Ochs, M.; Radhakrishnan, P.; Brill, A.; Hocker, H.; Schwarz, S.; Weissinger, D.; Kehm, R.; Kulu, Y.; Ulrich, A.; et al. Emx2 gene expression predicts liver metastasis and survival in colorectal cancer. BMC Cancer 2017, 17, 555. [Google Scholar]
- Ito, D.; Yogosawa, S.; Mimoto, R.; Hirooka, S.; Horiuchi, T.; Eto, K.; Yanaga, K.; Yoshida, K. Dual-specificity tyrosine-regulated kinase 2 is a suppressor and potential prognostic marker for liver metastasis of colorectal cancer. Cancer Sci. 2017, 108, 1565–1573. [Google Scholar] [CrossRef] [PubMed]
- Mekenkamp, L.J.; Haan, J.C.; Koopman, M.; Vink-Borger, M.E.; Israeli, D.; Teerenstra, S.; Ylstra, B.; Meijer, G.A.; Punt, C.J.; Nagtegaal, I.D. Chromosome 20p11 gains are associated with liver-specific metastasis in patients with colorectal cancer. Gut 2013, 62, 94–101. [Google Scholar] [PubMed]
- Zhao, Z.; Gao, Y.; Guan, X.; Liu, Z.; Jiang, Z.; Liu, X.; Lin, H.; Yang, M.; Li, C.; Yang, R.; et al. Gadd45b as a prognostic and predictive biomarker in Stage II colorectal cancer. Genes 2018, 9, 361. [Google Scholar] [CrossRef] [PubMed]
- Kashihara, H.; Shimada, M.; Kurita, N.; Iwata, T.; Sato, H.; Kozo, Y.; Higashijima, J.; Chikakiyo, M.; Nishi, M.; Matsumoto, N. CD133 expression is correlated with poor prognosis in colorectal cancer. Hepatogastroenterology 2014, 61, 1563–1567. [Google Scholar] [PubMed]
- Fang, C.; Fan, C.; Wang, C.; Huang, Q.; Meng, W.; Yu, Y.; Yang, L.; Peng, Z.; Hu, J.; Li, Y.; et al. CD133+CD54+CD44+ circulating tumor cells as a biomarker of treatment selection and liver metastasis in patients with colorectal cancer. Oncotarget 2016, 7, 77389–77403. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Sheng, Y.; Guan, M. Co-expression of stem cell genes CD133 and CD44 in colorectal cancers with early liver metastasis. Surg. Oncol. 2012, 21, 103–107. [Google Scholar] [CrossRef] [PubMed]
- Ernst, A.; Aigner, M.; Nakata, S.; Engel, F.; Schlotter, M.; Kloor, M.; Brand, K.; Schmitt, S.; Steinert, G.; Rahbari, N.; et al. A gene signature distinguishing CD133hi from CD133-colorectal cancer cells: Essential role for EGR1 and downstream factors. Pathology 2011, 43, 220–227. [Google Scholar] [CrossRef] [PubMed]
- Fang, C.; Fan, C.; Wang, C.; Huang, Q.; Meng, W.; Yu, Y.; Yang, L.; Hu, J.; Li, Y.; Mo, X.; et al. Prognostic value of CD133+CD54+CD44+ circulating tumor cells in colorectal cancer with liver metastasis. Cancer Med. 2017, 6, 2850–2857. [Google Scholar] [CrossRef] [PubMed]
- Villalba, M.; Evans, S.R.; Vidal-Vanaclocha, F.; Calvo, A. Role of TGF-β in metastatic colon cancer: It is finally time for targeted therapy. Cell Tissue Res. 2017, 370, 29–39. [Google Scholar] [CrossRef] [PubMed]
- Brunen, D.; Willems, S.M.; Kellner, U.; Midgley, R.; Simon, I.; Bernards, R. TGF-β: An emerging player in drug resistance. Cell Cycle 2013, 12, 2960–2968. [Google Scholar] [PubMed]
- Liu, H.; Xu, Y.; Zhang, Q.; Li, K.; Wang, D.; Li, S.; Ning, S.; Yang, H.; Shi, W.; Liu, Z.; et al. Correlations between TBL1XR1 and recurrence of colorectal cancer. Sci. Rep. 2017, 7, 44275. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Xu, Y.; Zhang, Q.; Yang, H.; Shi, W.; Liu, Z.; Li, K.; Gong, Z.; Ning, S.; Li, S.; et al. Prognostic significance of TBL1XR1 in predicting liver metastasis for early stage colorectal cancer. Surg. Oncol. 2017, 26, 13–20. [Google Scholar] [CrossRef] [PubMed]
- Nagasawa, T. Cxcl12/sdf-1 and cxcr4. Front. Immunol. 2015, 6, 301. [Google Scholar] [CrossRef] [PubMed]
- Amara, S.; Chaar, I.; Khiari, M.; Ounissi, D.; Weslati, M.; Boughriba, R.; Hmida, A.B.; Bouraoui, S. Stromal cell derived factor-1 and cxcr4 expression in colorectal cancer promote liver metastasis. Cancer Biomark 2015, 15, 869–879. [Google Scholar] [CrossRef] [PubMed]
- Song, L.; Tang, J.W.; Owusu, L.; Sun, M.Z.; Wu, J.; Zhang, J. Galectin-3 in cancer. Clin. Chim. Acta 2014, 431, 185–191. [Google Scholar] [CrossRef] [PubMed]
- Arfaoui-Toumi, A.; Kria-Ben Mahmoud, L.; Ben Hmida, M.; Khalfallah, M.T.; Regaya-Mzabi, S.; Bouraoui, S. Implication of the galectin-3 in colorectal cancer development (about 325 tunisian patients). Bull. Cancer 2010, 97, E1–E8. [Google Scholar] [PubMed]
- Kelemen, L.E. The role of folate receptor alpha in cancer development, progression and treatment: Cause, consequence or innocent bystander? Int. J. Cancer 2006, 119, 243–250. [Google Scholar] [CrossRef] [PubMed]
- D’Angelica, M.; Ammori, J.; Gonen, M.; Klimstra, D.S.; Low, P.S.; Murphy, L.; Weiser, M.R.; Paty, P.B.; Fong, Y.; Dematteo, R.P.; et al. Folate receptor-alpha expression in resectable hepatic colorectal cancer metastases: Patterns and significance. Mod. Pathol. 2011, 24, 1221–1228. [Google Scholar] [PubMed]
- Scholzen, T.; Gerdes, J. The ki-67 protein: From the known and the unknown. J. Cell. Physiol. 2000, 182, 311–322. [Google Scholar] [CrossRef]
- Eefsen, R.L.; Engelholm, L.; Willemoe, G.L.; Van den Eynden, G.G.; Laerum, O.D.; Christensen, I.J.; Rolff, H.C.; Hoyer-Hansen, G.; Osterlind, K.; Vainer, B.; et al. Microvessel density and endothelial cell proliferation levels in colorectal liver metastases from patients given neo-adjuvant cytotoxic chemotherapy and bevacizumab. Int. J. Cancer 2016, 138, 1777–1784. [Google Scholar] [CrossRef] [PubMed]
- Yadav, L.; Puri, N.; Rastogi, V.; Satpute, P.; Ahmad, R.; Kaur, G. Matrix metalloproteinases and cancer—Roles in threat and therapy. Asian Pac. J. Cancer Prev. 2014, 15, 1085–1091. [Google Scholar] [CrossRef] [PubMed]
- Fang, Y.J.; Lu, Z.H.; Wang, G.Q.; Pan, Z.Z.; Zhou, Z.W.; Yun, J.P.; Zhang, M.F.; Wan, D.S. Elevated expressions of MMP7, TROP2, and survivin are associated with survival, disease recurrence, and liver metastasis of colon cancer. Int. J. Colorectal Dis. 2009, 24, 875–884. [Google Scholar] [CrossRef] [PubMed]
- Itatani, Y.; Kawada, K.; Fujishita, T.; Kakizaki, F.; Hirai, H.; Matsumoto, T.; Iwamoto, M.; Inamoto, S.; Hatano, E.; Hasegawa, S.; et al. Loss of SMAD4 from colorectal cancer cells promotes CCL15 expression to recruit CCR1+ myeloid cells and facilitate liver metastasis. Gastroenterology 2013, 145, 1064–1075.e1011. [Google Scholar] [CrossRef] [PubMed]
- Asano, M.; Komiyama, K. Polymeric immunoglobulin receptor. J. Oral Sci. 2011, 53, 147–156. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Ye, P.; Bi, T.; Teng, L.; Xiang, C.; Wang, H.; Li, Y.; Jin, K.; Mou, X. Colorectal polymeric immunoglobulin receptor expression is correlated with hepatic metastasis and poor prognosis in colon carcinoma patients with hepatic metastasis. Hepatogastroenterology 2014, 61, 652–659. [Google Scholar] [PubMed]
- Rosen, L.S.; Gordon, M.S.; Robert, F.; Matei, D.E. Endoglin for targeted cancer treatment. Curr. Oncol. Rep. 2014, 16, 365. [Google Scholar] [PubMed]
- Mitselou, A.; Galani, V.; Skoufi, U.; Arvanitis, D.L.; Lampri, E.; Ioachim, E. Syndecan-1, epithelial–mesenchymal transition markers (E-cadherin/β-catenin) and neoangiogenesis-related proteins (PCAM-1 and endoglin) in colorectal cancer. Anticancer Res. 2016, 36, 2271–2280. [Google Scholar] [PubMed]
- Fang, Y.; Zhang, X. Targeting NEK2 as a promising therapeutic approach for cancer treatment. Cell Cycle 2016, 15, 895–907. [Google Scholar] [CrossRef] [PubMed]
- Neal, C.P.; Fry, A.M.; Moreman, C.; McGregor, A.; Garcea, G.; Berry, D.P.; Manson, M.M. Overexpression of the NEK2 kinase in colorectal cancer correlates with beta-catenin relocalization and shortened cancer-specific survival. J. Surg. Oncol. 2014, 110, 828–838. [Google Scholar] [CrossRef] [PubMed]
- MacDonald, B.T.; Tamai, K.; He, X. Wnt/β-catenin signaling: Components, mechanisms, and diseases. Dev. Cell 2009, 17, 9–26. [Google Scholar] [CrossRef] [PubMed]
- Pancione, M.; Forte, N.; Sabatino, L.; Tomaselli, E.; Parente, D.; Febbraro, A.; Colantuoni, V. Reduced β-catenin and peroxisome proliferator-activated receptor-gamma expression levels are associated with colorectal cancer metastatic progression: Correlation with tumor-associated macrophages, cyclooxygenase 2, and patient outcome. Hum. Pathol. 2009, 40, 714–725. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Cheng, H.; Liu, Y.; Yu, W.; Zhang, G.; Chen, B.; Yu, Z.; Hu, S. Prognostic value of nuclear β-catenin overexpression at invasive front in colorectal cancer for synchronous liver metastasis. Ann. Surg. Oncol. 2011, 18, 1553–1559. [Google Scholar] [CrossRef] [PubMed]
- Jones, J.R.; Barrick, C.; Kim, K.A.; Lindner, J.; Blondeau, B.; Fujimoto, Y.; Shiota, M.; Kesterson, R.A.; Kahn, B.B.; Magnuson, M.A. Deletion of ppargamma in adipose tissues of mice protects against high fat diet-induced obesity and insulin resistance. Proc. Natl. Acad. Sci. USA 2005, 102, 6207–6212. [Google Scholar] [CrossRef] [PubMed]
- Ren, B.; Zakharov, V.; Yang, Q.; McMahon, L.; Yu, J.; Cao, W. Macc1 is related to colorectal cancer initiation and early-stage invasive growth. Am. J. Clin. Pathol. 2013, 140, 701–707. [Google Scholar] [CrossRef] [PubMed]
- Bottaro, D.P.; Rubin, J.S.; Faletto, D.L.; Chan, A.M.; Kmiecik, T.E.; Vande Woude, G.F.; Aaronson, S.A. Identification of the hepatocyte growth factor receptor as the c-met proto-oncogene product. Science 1991, 251, 802–804. [Google Scholar] [CrossRef] [PubMed]
- Van den Eynden, G.G.; Bird, N.C.; Majeed, A.W.; Van Laere, S.; Dirix, L.Y.; Vermeulen, P.B. The histological growth pattern of colorectal cancer liver metastases has prognostic value. Clin. Exp. Metastasis 2012, 29, 541–549. [Google Scholar] [CrossRef] [PubMed]
- Hynes, R.O. Integrins: Versatility, modulation, and signaling in cell adhesion. Cell 1992, 69, 11–25. [Google Scholar] [CrossRef]
- Vassos, N.; Rau, T.; Merkel, S.; Feiersinger, F.; Geppert, C.I.; Sturzl, M.; Hohenberger, W.; Croner, R.S. Prognostic value of β1 integrin expression in colorectal liver metastases. Int. J. Clin. Exp. Pathol. 2014, 7, 288–300. [Google Scholar] [PubMed]
- Richman, S. Deficient mismatch repair: Read all about it (review). Int. J. Oncol. 2015, 47, 1189–1202. [Google Scholar] [CrossRef] [PubMed]
- Larsen, N.B.; Heiberg Engel, P.J.; Rasmussen, M.; Rasmussen, L.J. Differential expression of hmlh1 in sporadic human colorectal cancer tumors and distant metastases. Apmis 2009, 117, 839–848. [Google Scholar] [CrossRef] [PubMed]
- Murakami, T.; Kikuchi, H.; Ishimatsu, H.; Iino, I.; Hirotsu, A.; Matsumoto, T.; Ozaki, Y.; Kawabata, T.; Hiramatsu, Y.; Ohta, M.; et al. Tenascin c in colorectal cancer stroma is a predictive marker for liver metastasis and is a potent target of mir-198 as identified by microRNA analysis. Br. J. Cancer 2017, 117, 1360–1370. [Google Scholar] [CrossRef] [PubMed]
- Leggett, B.; Whitehall, V. Role of the serrated pathway in colorectal cancer pathogenesis. Gastroenterology 2010, 138, 2088–2100. [Google Scholar] [CrossRef] [PubMed]
- Voutsina, A.; Tzardi, M.; Kalikaki, A.; Zafeiriou, Z.; Papadimitraki, E.; Papadakis, M.; Mavroudis, D.; Georgoulias, V. Combined analysis of KRAS and PIK3CA mutations, met and pten expression in primary tumors and corresponding metastases in colorectal cancer. Mod. Pathol. 2013, 26, 302–313. [Google Scholar] [CrossRef] [PubMed]
- Baretti, M.; Personeni, N.; Destro, A.; Santoro, A.; Rimassa, L. Emergence of kras-mutation in liver metastases after an anti-EGFR treatment in patient with colorectal cancer: Are we aware of the therapeutic impact of intratumor heterogeneity? Cancer Biol. Ther. 2018, 19, 659–663. [Google Scholar] [CrossRef] [PubMed]
- McGregor, M.; Price, T.J. Panitumumab in the treatment of metastatic colorectal cancer, including wild-type RAS, KRAS and NRAS MCRC. Future Oncol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Guinney, J.; Dienstmann, R.; Wang, X.; de Reynies, A.; Schlicker, A.; Soneson, C.; Marisa, L.; Roepman, P.; Nyamundanda, G.; Angelino, P.; et al. The consensus molecular subtypes of colorectal cancer. Nat. Med. 2015, 21, 1350. [Google Scholar] [CrossRef] [PubMed]
- Thanki, K.; Nicholls, M.E.; Gajjar, A.; Senagore, A.J.; Qiu, S.; Szabo, C.; Hellmich, M.R.; Chao, C. Consensus Molecular Subtypes of Colorectal Cancer and their Clinical Implications. Int. Biol. Biomed. J. 2017, 3, 105–111. [Google Scholar] [PubMed]
- Zhang, L.; Pickard, K.; Jenei, V.; Bullock, M.D.; Bruce, A.; Mitter, R.; Kelly, G.; Paraskeva, C.; Strefford, J.; Primrose, J.; et al. Mir-153 supports colorectal cancer progression via pleiotropic effects that enhance invasion and chemotherapeutic resistance. Cancer Res. 2013, 73, 6435–6447. [Google Scholar] [CrossRef] [PubMed]
- Qian, Z.; Zhang, G.; Song, G.; Shi, J.; Gong, L.; Mou, Y.; Han, Y. Integrated analysis of genes associated with poor prognosis of patients with colorectal cancer liver metastasis. Oncotarget 2017, 8, 25500–25512. [Google Scholar] [CrossRef] [PubMed]
- Patrussi, L.; Baldari, C.T. The cxcl12/cxcr4 axis as a therapeutic target in cancer and HIV-1 infection. Curr. Med. Chem. 2011, 18, 497–512. [Google Scholar] [CrossRef] [PubMed]
- Altwerger, G.; Bonazzoli, E.; Bellone, S.; Egawa-Takata, T.; Menderes, G.; Pettinella, F.; Bianchi, A.; Riccio, F.; Feinberg, J.; Zammataro, L.; et al. In vitro and in vivo activity of imgn853, an antibody-drug conjugate targeting folate receptor alpha linked to dm4, in biologically aggressive endometrial cancers. Mol. Cancer Ther. 2018. [Google Scholar] [CrossRef] [PubMed]
- Fang, Y.; Kong, Y.; Xi, J.; Zhu, M.; Zhu, T.; Jiang, T.; Hu, W.; Ma, M.; Zhang, X. Preclinical activity of mbm-5 in gastrointestinal cancer by inhibiting NEK2 kinase activity. Oncotarget 2016, 7, 79327–79341. [Google Scholar] [CrossRef] [PubMed]
- Del Gobbo, A.; Pellegrinelli, A.; Gaudioso, G.; Castellani, M.; Zito Marino, F.; Franco, R.; Palleschi, A.; Nosotti, M.; Bosari, S.; Vaira, V.; et al. Analysis of NSCLC tumour heterogeneity, proliferative and 18F-FDG pet indices reveals ki67 prognostic role in adenocarcinomas. Histopathology 2016, 68, 746–751. [Google Scholar] [CrossRef] [PubMed]
- Goasguen, N.; de Chaisemartin, C.; Brouquet, A.; Julie, C.; Prevost, G.P.; Laurent-Puig, P.; Penna, C. Evidence of heterogeneity within colorectal liver metastases for allelic losses, mRNA level expression and in vitro response to chemotherapeutic agents. Int. J. Cancer 2010, 127, 1028–1037. [Google Scholar] [CrossRef] [PubMed]
- Turtoi, A.; Blomme, A.; Debois, D.; Somja, J.; Delvaux, D.; Patsos, G.; Di Valentin, E.; Peulen, O.; Mutijima, E.N.; De Pauw, E.; et al. Organized proteomic heterogeneity in colorectal cancer liver metastases and implications for therapies. Hepatology 2014, 59, 924–934. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lopez, G.; Boggio, F.; Ferrero, S.; Fusco, N.; Del Gobbo, A. Molecular and Immunohistochemical Markers with Prognostic and Predictive Significance in Liver Metastases from Colorectal Carcinoma. Int. J. Mol. Sci. 2018, 19, 3014. https://doi.org/10.3390/ijms19103014
Lopez G, Boggio F, Ferrero S, Fusco N, Del Gobbo A. Molecular and Immunohistochemical Markers with Prognostic and Predictive Significance in Liver Metastases from Colorectal Carcinoma. International Journal of Molecular Sciences. 2018; 19(10):3014. https://doi.org/10.3390/ijms19103014
Chicago/Turabian StyleLopez, Gianluca, Francesca Boggio, Stefano Ferrero, Nicola Fusco, and Alessandro Del Gobbo. 2018. "Molecular and Immunohistochemical Markers with Prognostic and Predictive Significance in Liver Metastases from Colorectal Carcinoma" International Journal of Molecular Sciences 19, no. 10: 3014. https://doi.org/10.3390/ijms19103014
APA StyleLopez, G., Boggio, F., Ferrero, S., Fusco, N., & Del Gobbo, A. (2018). Molecular and Immunohistochemical Markers with Prognostic and Predictive Significance in Liver Metastases from Colorectal Carcinoma. International Journal of Molecular Sciences, 19(10), 3014. https://doi.org/10.3390/ijms19103014





