Clinical Significance of Measuring Global Hydroxymethylation of White Blood Cell DNA in Prostate Cancer: Comparison to PSA in a Pilot Exploratory Study
Abstract
1. Introduction
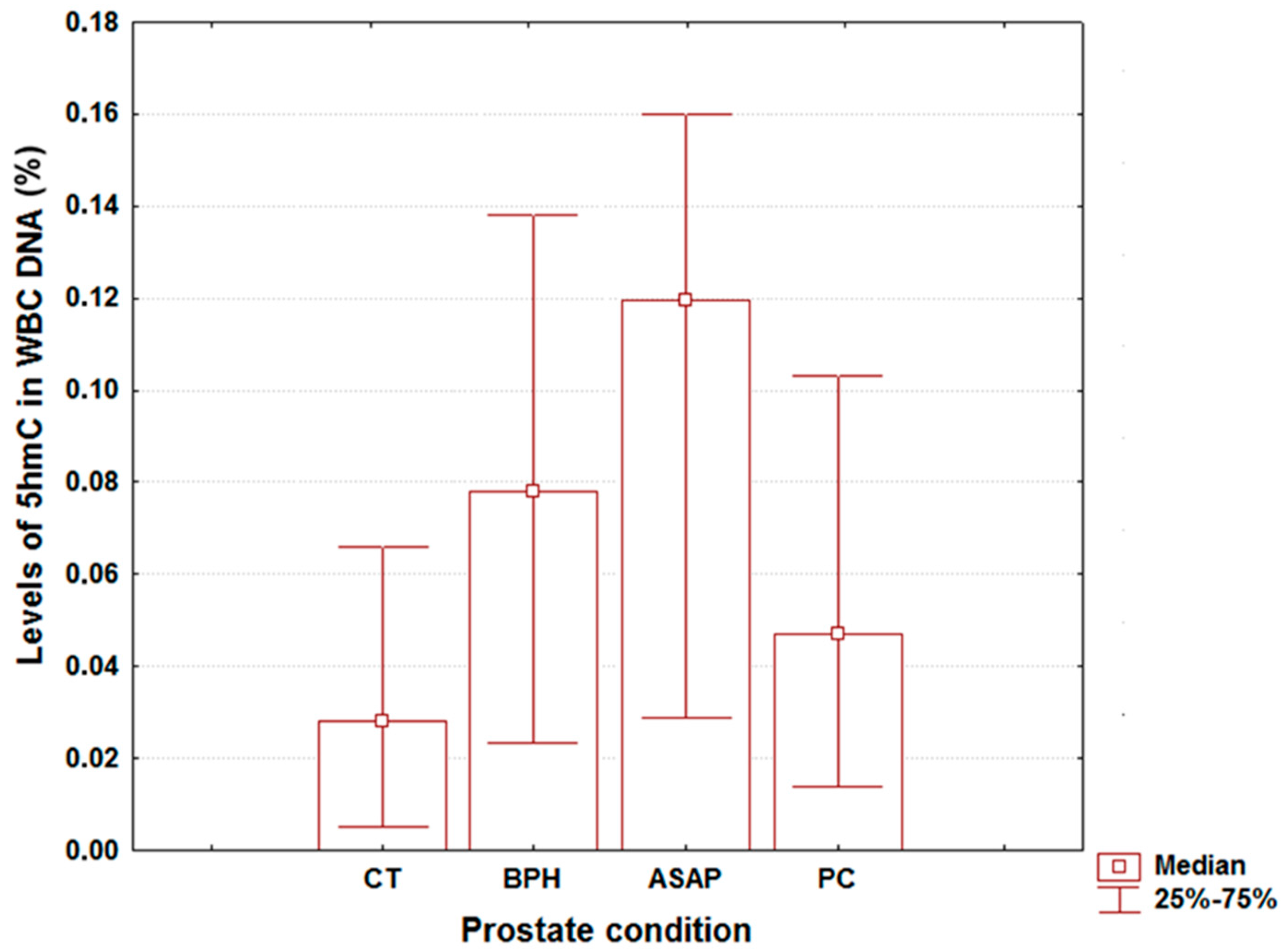
2. Results
3. Discussion
4. Materials and Methods
4.1. Study Population
4.2. Genomic DNA Extraction
4.3. Quantification of Global DNA Methylation in WBC
4.4. Statistical Analysis
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Attard, G.; Parker, G.; Eeles, R.A.; Schröder, F.; Tomlins, S.A.; Tannock, I.; Drake, C.G.; de Bono, J.S. Prostate cancer. Lancet 2016, 387, 70–82. [Google Scholar] [CrossRef]
- Center, M.M.; Jemal, A.; Lortet-Tieulent, J.; Ward, E.; Ferlay, J.; Brawley, O.; Bray, F. International variation in prostate cancer incidence and mortality rates. Eur. Urol. 2012, 61, 1079–1092. [Google Scholar] [CrossRef] [PubMed]
- McVary, K.T. BPH: Epidemiology and comorbidities. Am. J. Manag. Care 2006, 12, S122–S128. [Google Scholar] [PubMed]
- Esserman, L.J.; Thompson, I.M.; Reid, B.; Nelson, P.; Ransohoff, D.F.; Welch, H.G.; Hwang, S.; Berry, D.A.; Kinzler, K.W.; Black, W.C.; et al. Addressing over diagnosis and over treatment in cancer: A prescription for change. Lancet Oncol. 2014, 15, e234–e242. [Google Scholar] [CrossRef]
- Schröder, F.H.; Hugosson, J.; Roobol, M.J.; Tammela, T.L.J.; Ciatto, S.; Nelen, V.; Kwiatkowski, M.; Lujan, M.; Lilja, H.; Zappa, M.; et al. Screening and prostate-cancer mortality in a randomized European study. N. Eng. J. Med. 2009, 360, 1320–1328. [Google Scholar] [CrossRef] [PubMed]
- Lin, K.; Lipsitz, R.; Miller, T.; Janakiraman, S.U.S. Preventive Services Task Force. Benefits and harms of prostate-specific antigen screening for prostate cancer: An evidence update for the U.S. Preventive Services Task Force. Ann. Int. Med. 2008, 149, 192–199. [Google Scholar] [CrossRef] [PubMed]
- Heidenreich, A.; Bellmunt, J.; Bolla, M.; Joniau, S.; Mason, M.; Matveev, V.; Mottet, N.; Schmid, H.P.; van der Kwast, T.; Wiegel, T.; et al. EAU guidelines on prostate cancer. Part 1: Screening, diagnosis, and treatment of clinically localised disease. Eur. Urol. 2011, 59, 61–71. [Google Scholar] [CrossRef] [PubMed]
- Chiam, K.; Ricciardelli, C.; Bianco-Miotto, T. Epigenetic biomarkers in prostate cancer: Current and future uses. Cancer Lett. 2014, 342, 248–256. [Google Scholar] [CrossRef] [PubMed]
- Tirado-Magallanes, R.; Rebbani, K.; Lim, R.; Pradhan, S.; Benoukraf, T. Whole genome DNA methylation: Beyond genes silencing. Oncotarget 2017, 8, 5629–5637. [Google Scholar] [CrossRef] [PubMed]
- Branco, M.R.; Ficz, G.; Reik, W. Uncovering the role of 5-hydroxymethylcytosine in the epigenome. Nat. Rev. Genet. 2011, 13, 7–13. [Google Scholar] [CrossRef] [PubMed]
- Hoque, M.O. DNA methylation changes in prostate cancer: Current developments and future clinical implementation. Expert Rev. Mol. Diagn. 2009, 9, 243–257. [Google Scholar] [CrossRef] [PubMed]
- Strand, S.H.; Orntoft, T.F.; Sorensen, K.D. Prognostic DNA methylation markers for prostate cancer. Int. J. Mol. Sci. 2014, 15, 16544–16576. [Google Scholar] [CrossRef] [PubMed]
- Kirby, M.K.; Ramaker, R.C.; Roberts, B.S.; Lasseigne, B.N.; Gunther, D.S.; Burwell, T.C.; Davis, N.S.; Gulzar, Z.G.; Absher, D.M.; Cooper, S.J.; et al. Genome-wide DNA methylation measurements in prostate tissues uncovers novel prostate cancer diagnostic biomarkers and transcription factor binding patterns. BMC Cancer 2017, 17, 273. [Google Scholar] [CrossRef] [PubMed]
- He, W.; Bishop, K.S. The potential use of cell-free-circulating-tumor DNA as a biomarker for prostate cancer. Expert Rev. Mol. Diagn. 2016, 16, 839–852. [Google Scholar] [CrossRef] [PubMed]
- Kamdar, S.N.; Ho, L.T.; Kron, K.J.; Isserlin, R.; van der Kwast, T.; Zlotta, A.R.; Fleshner, N.E.; Bader, G.; Bapat, B. Dynamic interplay between locus-specific DNA methylation and hydroxymethylation regulates distinct biological pathways in prostate carcinogenesis. Clin. Epigenet. 2016, 8, 32. [Google Scholar] [CrossRef] [PubMed]
- Spans, L.; Van den Broeck, T.; Smeets, E.; Prekovic, S.; Thienpont, B.; Lambrechts, D.; Karnes, R.J.; Erho, N.; Alshalalfa, M.; Davicioni, E.; et al. Genomic and epigenomic analysis of high-risk prostate cancer reveals changes in hydroxymethylation and TET1. Oncotarget 2016, 7, 24326–24338. [Google Scholar] [CrossRef] [PubMed]
- Luo, J.H.; Ding, Y.; Chen, R.; Michalopoulos, G.; Nelson, J.; Yu, Y.P. Genome-wide methylation analysis of prostate tissues reveals global methylation patterns of prostate cancer. Am. J. Pathol. 2013, 182, 2028–2036. [Google Scholar] [CrossRef] [PubMed]
- FitzGerald, L.M.; Naeem, H.; Makalic, E.; Schmidt, D.F.; Dowty, J.G.; Joo, J.E.; Jung, C.H.; Bassett, J.K.; Dugue, P.A.; Chung, J.; et al. Genome-wide measures of peripheral blood DNA methylation and prostate cancer risk in a prospective nested case-control study. Prostate 2017, 77, 471–478. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Choi, J.Y.; Lee, K.; Sung, H.; Park, S.K.; Oze, I.; Pan, K.F.; You, W.C.; Chen, Y.X.; Fang, J.Y.; et al. DNA methylation in peripheral blood: A potential biomarker for cancer molecular epidemiology. J. Epidemiol. 2012, 22, 384–394. [Google Scholar] [CrossRef] [PubMed]
- Kawachi, M.H.; Bahnson, R.R.; Barry, M.; Busby, J.E.; Carroll, P.R.; Carter, H.B.; Catalona, W.J.; Cookson, M.S.; Epstein, J.I.; Etzioni, R.B.; et al. NCCN clinical practice guidelines in oncology: Prostate cancer early detection. J. Natl. Compr. Cancer Netw. 2010, 8, 240–262. [Google Scholar] [CrossRef]
- Cash, H.L.; Tao, L.; Yuan, J.M.; Marsit, C.J.; Housemank, E.A.; Xiang, Y.B.; Gao, Y.T.; Nelson, H.H.; Kelsey, K.T. LINE-1 hypomethylation is associated with bladder cancer risk among non-smoking Chinese. Int. J. Cancer 2011, 130, 1151–1159. [Google Scholar] [CrossRef] [PubMed]
- Hou, L.; Wang, H.; Sartori, S.; Gawron, A.; Gawron, A.; Lissowska, J.; Bollati, V.; Tarantini, L.; Zhang, F.F.; Zatonski, W.; et al. Blood leukocyte DNA hypomethylation and gastric cancer risk in a high-risk Polish population. Int. J. Cancer 2010, 127, 1866–1874. [Google Scholar] [CrossRef] [PubMed]
- Wilhelm, C.S.; Kelsey, K.T.; Butler, R.; Plaza, S.; Gagne, L.; Zens, M.S.; Andrew, A.S.; Morris, S.; Nelson, H.H.; Schned, A.R.; et al. Implications of LINE1 methylation for bladder cancer risk in women. Clin. Cancer Res. 2010, 16, 1682–1689. [Google Scholar] [CrossRef] [PubMed]
- Choi, J.Y.; James, S.R.; Link, P.A.; McCann, S.E.; Hong, C.C.; Davis, W.; Nesline, M.K.; Ambrosone, C.B.; Karpf, A.R. Association between global DNA hypomethylation in leukocytes and risk of breast cancer. Carcinogenesis 2009, 3, 1889–1918. [Google Scholar] [CrossRef] [PubMed]
- Lim, U.; Flood, A.; Choi, S.W.; Albanes, D.; Cross, A.J.; Schatzkin, A.; Sinha, R.; Katki, H.A.; Cash, B.; Schoenfeld, P.; et al. Genomic methylation of leukocyte DNA in relation to colorectal adenoma among asymptomatic women. Gastroenterology 2008, 134, 47–55. [Google Scholar] [CrossRef] [PubMed]
- Moore, L.E.; Pfeiffer, R.M.; Poscablo, C.; Real, F.X.; Kogevinas, M.; Silverman, D.; García-Closas, R.; Chanock, S.; Tardón, A.; Serra, C.; et al. Genomic DNA hypomethylation as a biomarker for bladder cancer susceptibility in the Spanish Bladder Cancer Study: A case-control study. Lancet Oncol. 2008, 9, 359–366. [Google Scholar] [CrossRef]
- Hsiung, D.; Marsit, C.; Houseman, E.; Eddy, K.; Furniss, C.S.; McClean, M.D.; Kelsey, K.T. Global DNA methylation level in whole blood as a biomarker in head and neck squamous cell carcinoma. Cancer Epidemiol. Biomark. Prev. 2007, 16, 108–114. [Google Scholar] [CrossRef] [PubMed]
- Pufulete, M.; Al-Ghnaniem, R.; Leather, A.J.M.; Appleby, P.; Gout, S.; Terry, C.; Emery, P.W.; Sanders, T.A. Folate status, genomic DNA hypomethylation and risk of colorectal adenoma and cancer: A case control study. Gastroenterology 2003, 124, 1240–1248. [Google Scholar] [CrossRef]
- Gilat, N.; Tabachnik, T.; Shwartz, A.; Shahal, T.; Torchinsky, D.; Michaeli, Y.; Nifker, G.; Zirkin, S.; Ebenstein, Y. Single-molecule quantification of 5-hydroxymethylcytosine for diagnosis of blood and colon cancers. Clin. Epigenet. 2017, 9, 70. [Google Scholar] [CrossRef] [PubMed]
- Chowdhury, B.; Cho, I.L.; Hahn, N.; Joseph Irudayaraj, J. Quantification of 5-methylcytosine, 5-hydroxymethylcytosine and 5-carboxylcytosine from the blood of cancer patients by an Enzyme-based Immunoassay. Anal. Chim. Acta 2014, 852, 212–217. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Guerra, M.; Zheng, Y.; Osorio-Yanez, C.; Zhong, J.; Chervona, Y.; Wang, S.; Chang, D.; McCracken, J.P.; Díaz, A.; Bertazzi, P.A.; et al. Effects of particulate matter exposure on blood 5-hydroxymethylation: Results from the Beijing truck driver air pollution study. Epigenetics 2015, 10, 633–642. [Google Scholar] [CrossRef] [PubMed]
- Tellez-Plaza, M.; Tang, W.Y.; Shang, Y.; Umans, J.G.; Francesconi, K.A.; Goessler, W.; Ledesma, M.; Leon, M.; Laclaustra, M. Association of global DNA methylation and global DNA hydroxymethylation with metals and other exposures in human blood DNA samples. Environ. Health Perspect. 2014, 122, 946–954. [Google Scholar] [CrossRef] [PubMed]
- Cihan, Y.B.; Arslan, A.; Ergul, M.A. Subtypes of white blood cells in patients with prostate cancer or benign prostatic hyperplasia and healthy individuals. Asian Pac. J. Cancer Prev. 2013, 14, 4779–4783. [Google Scholar] [CrossRef] [PubMed]
- Fujita, K.; Imamura, R.; Tanigawa, G.; Nakagawa, M.; Hayashi, T.; Kishimoto, N.; Hosomi, M.; Yamaguchi, S. Low serum neutrophil count predicts a positive prostate biopsy. Prostate Cancer Prostatic Dis. 2012, 15, 386–390. [Google Scholar] [CrossRef] [PubMed]
- Krušlin, B.; Tomas, D.; Džombeta, T.; Milković-Periša, M.; Ulamec, M. Inflammation in prostatic hyperplasia and carcinoma—basic scientific approach. Front. Oncol. 2017, 1, 9. [Google Scholar] [CrossRef] [PubMed]
- Terry, M.B.; Delgado-Cruzata, L.; Vin-Raviv, N.; Wu, H.C.; Santella, R.M. DNA methylation in white blood cells: Association with risk factors in epidemiologic studies. Epigenetics 2011, 6, 828–837. [Google Scholar] [CrossRef] [PubMed]
- Haffner, M.C.; Chaux, A.; Meeker, A.K.; Esopi, D.M.; Gerber, J.; Pellakuru, L.G.; Toubaji, A.; Argani, P.; Iacobuzio-Donahue, C.; Nelson, W.G.; et al. Global 5-hydroxymethylcytosine content is significantly reduced in tissue stem/progenitor cell compartments and in human cancers. Oncotarget 2011, 2, 627–637. [Google Scholar] [CrossRef] [PubMed]
- Nestor, C.E.; Ottaviano, R.; Reddington, J.; Sproul, D.; Reinhardt, D.; Dunican, D.; Katz, E.; Dixon, J.M.; Harrison, D.J.; Meehan, R.R. Tissue type is a major modifier of the 5-hydroxymethylcytosine content of human genes. Genome Res. 2012, 22, 467–477. [Google Scholar] [CrossRef] [PubMed]
- Sfanos, K.S.; Hempel, H.A.; De Marzo, A.M. The role of inflammation in prostate cancer. Adv. Exp. Med. Biol. 2014, 816, 153–181. [Google Scholar] [PubMed]
- Smeets, E.; Lynch, A.G.; Prekovic, S.; van den Broeck, T.; Moris, L.; Helsen, C.; Joniau, S.; Claessens, F.; Massie, C.E. The role of TET-mediated DNA hydroxymethylation in prostate cancer. Mol. Cell. Endocrinol. 2017. [Google Scholar] [CrossRef] [PubMed]
- Feng, J.; Wang, Q.; Li, G.; Zeng, X.; Kuang, S.; Li, X.; Yue, Y. TET1-mediated different transcriptional regulation in prostate cancer. Int. J. Clin. Exp. Med. 2015, 8, 203–211. [Google Scholar] [PubMed]
- Yang, H.; Liu, Y.; Bai, F.; Zhang, J.Y.; Ma, S.H.; Liu, J.; Xu, Z.H.; Zhu, H.G.; Ling, Z.Q.; Ye, D.; et al. Tumor development is associated with decrease of TET gene expression and 5-methylcytosine hydroxylation. Oncogene 2013, 32, 663–669. [Google Scholar] [CrossRef] [PubMed]
- Strand, S.H.; Hoyer, S.; Lynnerup, A.S.; Haldrup, C.; Storebjerg, T.M.; Borre, M.; Orntoft, T.F.; Sorensen, K.D. High levels of 5-hydroxymethylcytosine (5hmC) is an adverse predictor of biochemical recurrence after prostatectomy in ERG-negative prostate cancer. Clin. Epigenet. 2015, 7, 111. [Google Scholar] [CrossRef] [PubMed]
- Zelic, R.; Fiano, V.; Grasso, C.; Zugna, D.; Pettersson, A.; Gillio-Tos, A.; Merletti, F.; Richiardi, L. Global DNA hypomethylation in prostate cancer development and progression: A systematic review. Prostate Cancer Prostatic Dis. 2014, 1, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Jin, S.G.; Jiang, Y.; Qiu, R.; Rauch, T.A.; Wang, Y.; Schackert, G.; Krex, D.; Lu, Q.; Pfeifer, G.P. 5-Hydroxymethylcytosine is strongly depleted in human cancers but its levels do not correlate with IDH1 mutations. Cancer Res. 2011, 71, 7360–7365. [Google Scholar] [CrossRef] [PubMed]
- Chowdhury, B.; Cho, I.H.; Irudayaraj, J. Technical advances in global DNA methylation analysis in human cancers. J. Biol. Eng. 2017, 11, 10. [Google Scholar] [CrossRef] [PubMed]
- Kurdyukov, S.; Bullock, M. DNA methylation analysis: Choosing the right method. Biology 2016, 5, 3. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Liu, M. Distribution of 5-Hydroxymethylcytosine in different human tissues. J. Nucleic Acid 2011, 2, 325–330. [Google Scholar] [CrossRef] [PubMed]
- Thompson, I.M.; Pauler, D.K.; Goodman, P.J.; Tangen, C.M.; Lucia, M.S.; Parnes, H.L.; Minasian, L.M.; Ford, L.G.; Lippman, S.M.; Crawford, E.D.; et al. Prevalence of prostate cancer among men with a prostate-specific antigen level ≤ 4.0 ng per milliliter. N. Engl. J. Med. 2004, 350, 2239–2246. [Google Scholar] [CrossRef] [PubMed]
- Hoffman, R.M. Screening for prostate cancer. N. Eng. J. Med. 2011, 365, 2013–2019. [Google Scholar] [CrossRef] [PubMed]
- Ankerst, D.P.; Thompson, I.M. Sensitivity and specificity of prostate-specific antigen for prostate cancer detection with high rates of biopsy verification. Arch. Ital. Urol. Androl. 2006, 78, 125–129. [Google Scholar] [PubMed]
- Lilja, H.; Ulmert, D.; Vickers, A.J. Prostate-specific antigen and prostate cancer: Prediction, detection and monitoring. Nat. Rev. Cancer 2008, 8, 268–278. [Google Scholar] [CrossRef] [PubMed]
- Adhyam, M.; Gupta, A.K. A review on the clinical utility of PSA in cancer prostate. Indian J. Surg. Oncol. 2012, 2, 120–129. [Google Scholar] [CrossRef] [PubMed]
- Wu, T.; Giovannucci, E.; Welge, J.; Mallick, P.; Tang, W.Y.; Ho, S.M. Measurement of GSTP1 promoter methylation in body fluids may complement PSA screening: A meta-analysis. Br. J. Cancer 2011, 105, 65–73. [Google Scholar] [CrossRef] [PubMed]
- Hertzog, M.A. Considerations in determining sample size for pilot studies. Res. Nurs. Health 2008, 31, 180–191. [Google Scholar] [CrossRef] [PubMed]
- Leon, A.C.; Davis, L.L.; Kraemer, H.C. The role and interpretation of pilot studies in clinical research. J. Psychiatr. Res. 2011, 45, 626–629. [Google Scholar] [CrossRef] [PubMed]
- Chow, S.C. Controversial Statistical Issues in Clinical Trials; CRC Press: Boca Raton, FL, USA, 2011. [Google Scholar]
- Altman, D.G. The cost of dichotomising continuous variables. BMJ 2006, 332, 1080. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.C.; Delgado-Cruzata, L.; Flom, J.D.; Kappil, M.; Ferris, J.S.; Liao, Y.; Santella, M.R.; Terry, M.B. Global methylation profiles in DNA from different blood cell types. Epigenetics 2011, 6, 76–85. [Google Scholar] [CrossRef] [PubMed]
| Prostate Pathology | Age (Years) | PSA (ng/mL) |
|---|---|---|
| CT (control, n = 9) | 63 (61, 68) | 0.89 (0.29, 1.53) |
| BPH (n = 40) | 70 (60, 75) | 2.05 (0.52, 5.33) |
| ASAP (n = 10) | 68 (62, 79) | 7.61 (6.44, 9.39) |
| PCa (n = 30) | 72 (67, 76) | 34.01 (12.81, 116.02) |
| Prostate Pathology | DNA Hydroxymethylation Class | PSA Class | 5hmc-PSA Correlation | |||
|---|---|---|---|---|---|---|
| Low | High | Low | Intermediate | High | ||
| CT (n = 9) | 77.78% (7) | 22.22% (2) | 88.89% (8) | 11.11% (1) | 0.00% (0) | 0.18 (0.62) |
| BPH (n = 40) | 42.50% (17) | 57.50% (23) | 60.00% (24) | 35.00% (14) | 5.00% (2) | 0.22 (0.15) |
| ASAP (n = 10) | 30.00% (3) | 70.00% (7) | 10.00% (1) | 70.00% (7) | 20.00% (2) | 0.12 (0.73) |
| PCa (n = 30) | 66.67% (18) | 33.33% (9) | 3.70% (1) | 7.41% (2) | 88.89% (4) | 0.23 (0.24) |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Grelus, A.; Nica, D.V.; Miklos, I.; Belengeanu, V.; Ioiart, I.; Popescu, C. Clinical Significance of Measuring Global Hydroxymethylation of White Blood Cell DNA in Prostate Cancer: Comparison to PSA in a Pilot Exploratory Study. Int. J. Mol. Sci. 2017, 18, 2465. https://doi.org/10.3390/ijms18112465
Grelus A, Nica DV, Miklos I, Belengeanu V, Ioiart I, Popescu C. Clinical Significance of Measuring Global Hydroxymethylation of White Blood Cell DNA in Prostate Cancer: Comparison to PSA in a Pilot Exploratory Study. International Journal of Molecular Sciences. 2017; 18(11):2465. https://doi.org/10.3390/ijms18112465
Chicago/Turabian StyleGrelus, Alin, Dragos V. Nica, Imola Miklos, Valerica Belengeanu, Ioan Ioiart, and Cristina Popescu. 2017. "Clinical Significance of Measuring Global Hydroxymethylation of White Blood Cell DNA in Prostate Cancer: Comparison to PSA in a Pilot Exploratory Study" International Journal of Molecular Sciences 18, no. 11: 2465. https://doi.org/10.3390/ijms18112465
APA StyleGrelus, A., Nica, D. V., Miklos, I., Belengeanu, V., Ioiart, I., & Popescu, C. (2017). Clinical Significance of Measuring Global Hydroxymethylation of White Blood Cell DNA in Prostate Cancer: Comparison to PSA in a Pilot Exploratory Study. International Journal of Molecular Sciences, 18(11), 2465. https://doi.org/10.3390/ijms18112465




