Molecular Alliance of Lymantria dispar Multiple Nucleopolyhedrovirus and a Short Unmodified Antisense Oligonucleotide of Its Anti-Apoptotic IAP-3 Gene: A Novel Approach for Gypsy Moth Control
Abstract
1. Introduction
2. Results
2.1. Nucleopolyhedrovirus (LdMNPV)-Free Gypsy Moth Larvae Grown From Eggs in the Laboratory May Not Be Sufficiently Sensitive to the Antisense oligoRING Fragment
2.1.1. LdMNPV Contamination Exclusion
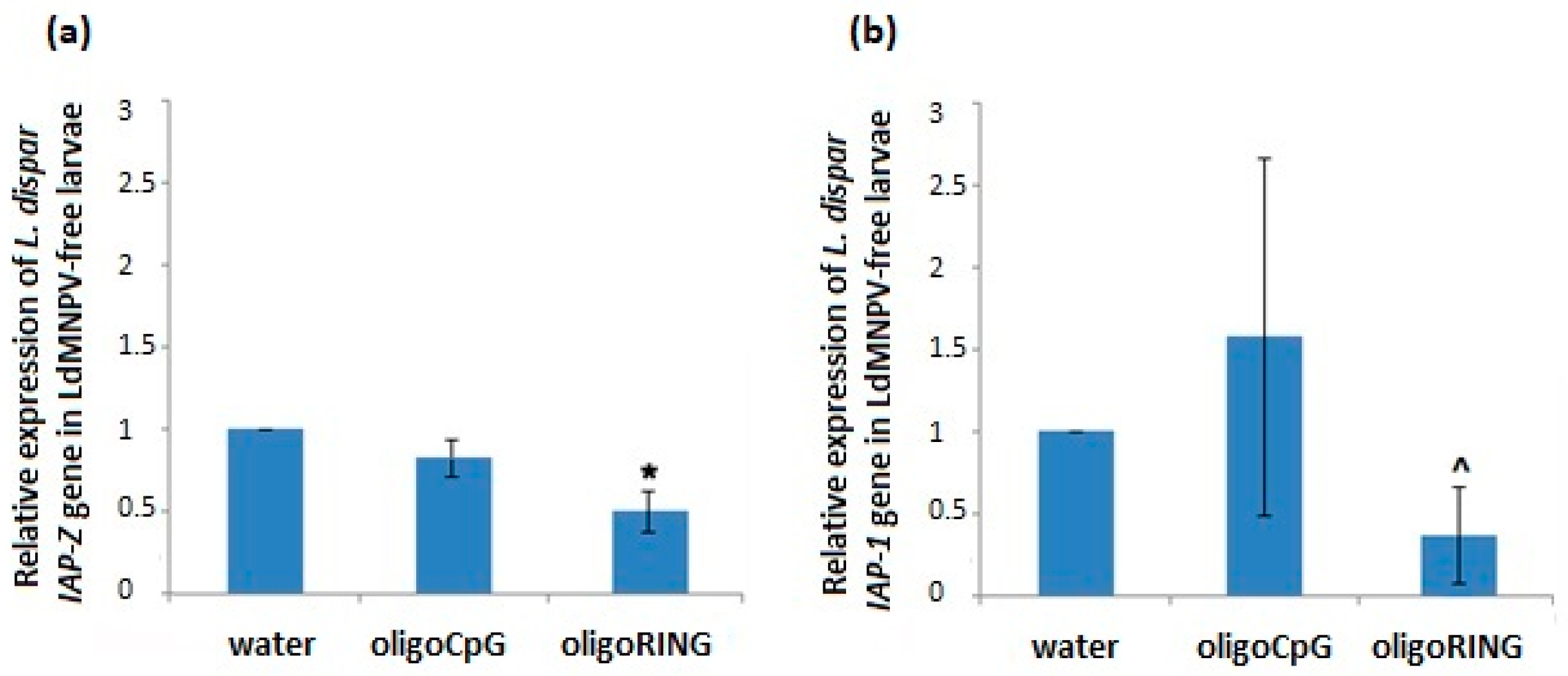
2.1.2. The Effect of the DNA oligonucleotides (oligoDNAs) on the LdMNPV-Free Gypsy Moth Larvae
2.1.3. Search for the oligoRING (Really Interesting New Gene Oligonucleotides) Target mRNA
2.2. Topical Application of oligoRING from the LdMNPV IAP-3 Gene Increases the Mortality Rate of LdMNPV-Infected Gypsy Moth Larvae “That Which Is Falling Should also Be Pushed”
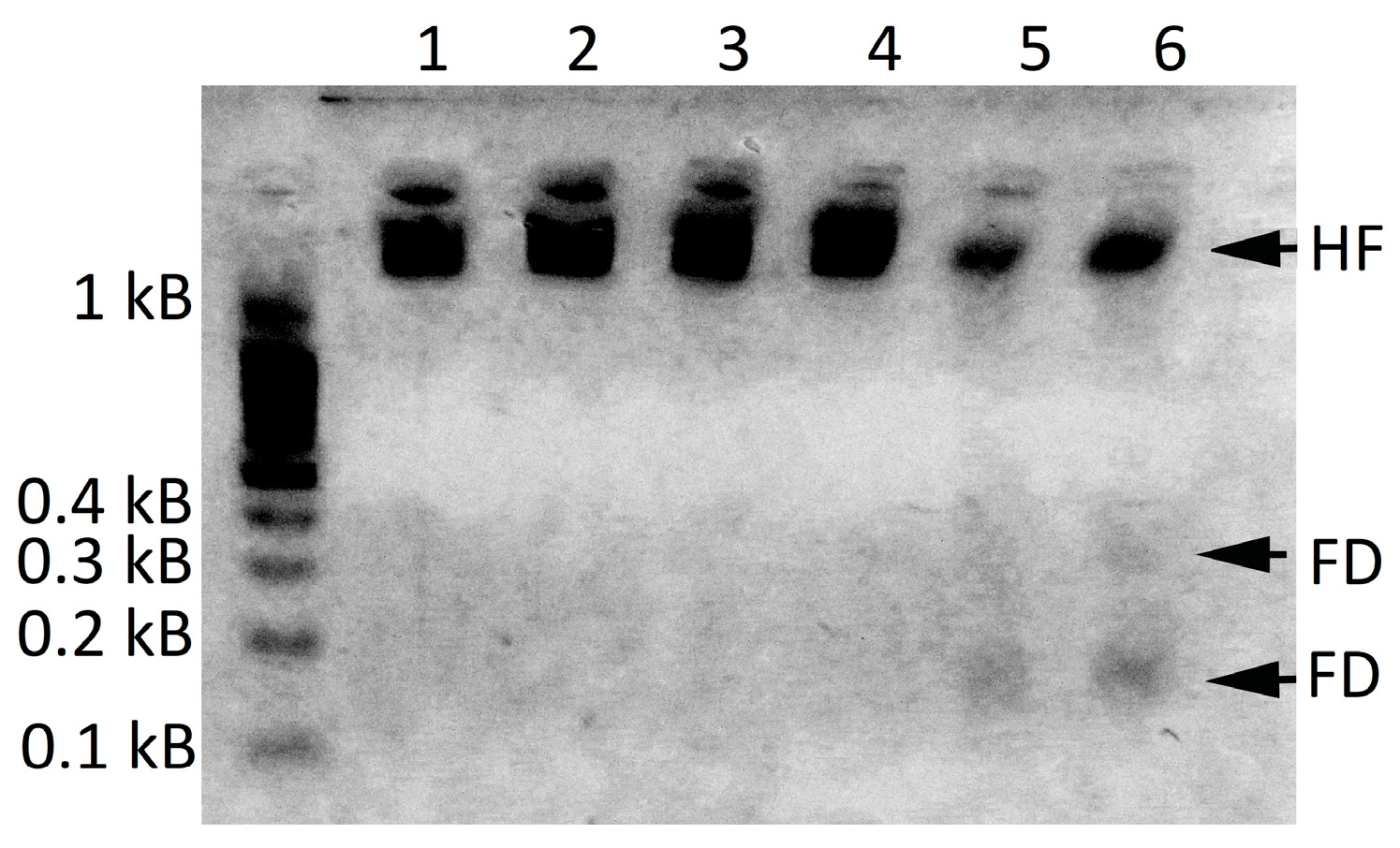
2.2.1. Evidence of Successful Infection with LdMNPV
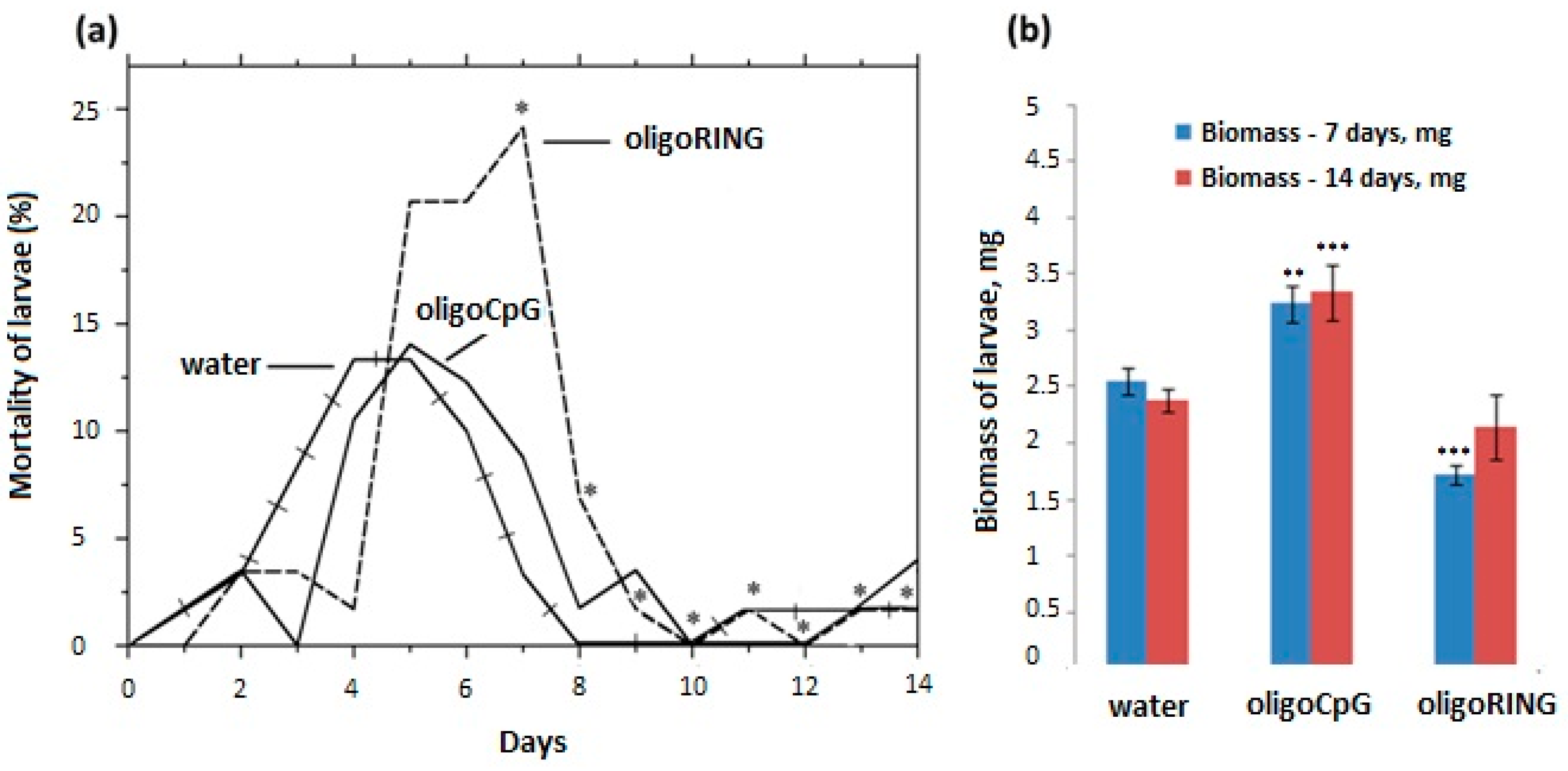
2.2.2. Topical Treatment with the Antisense oligoRING Fragment Leads to Decreased Rate of Biomass Accumulation and an Increased Mortality Rate in LdMNPV-Infected Gypsy Moth Larvae
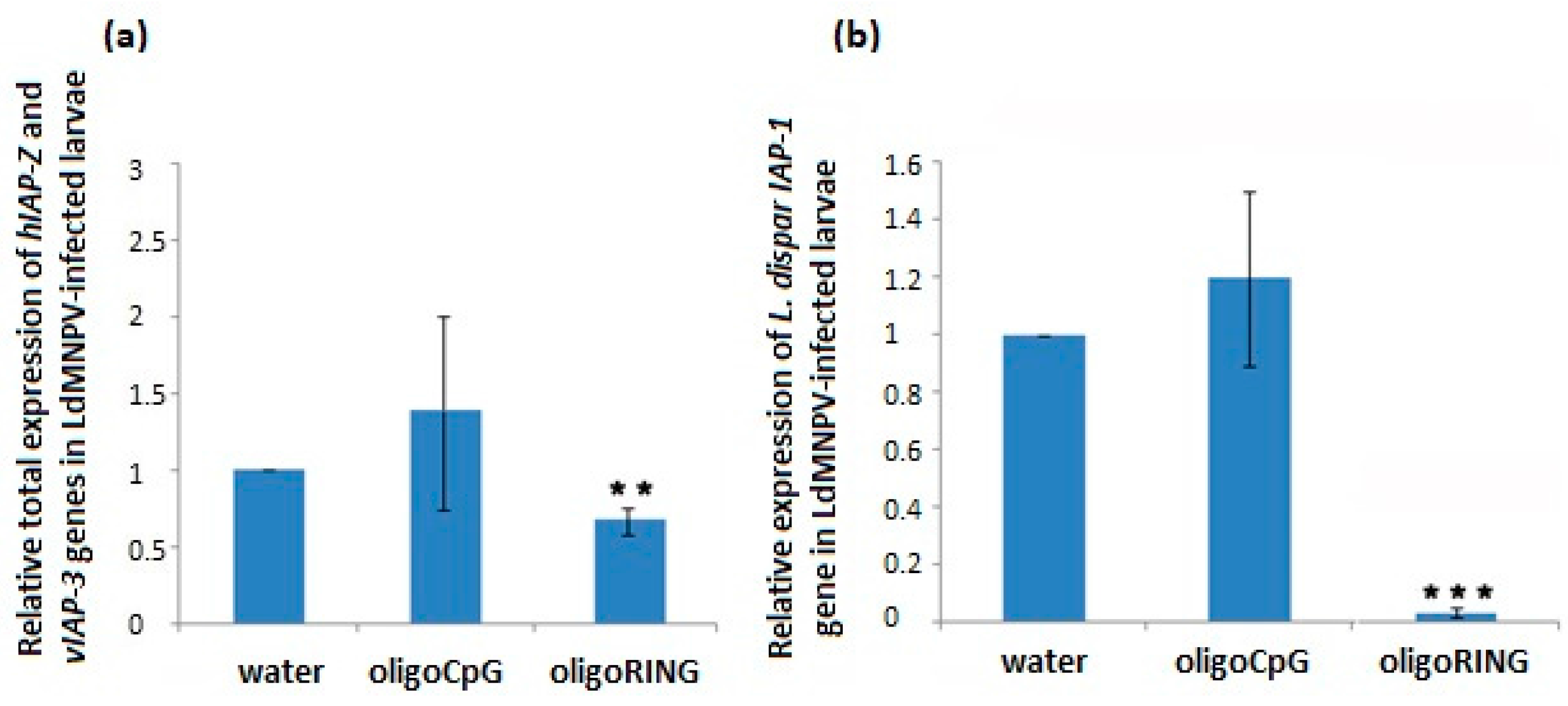
2.3. The oligoRING Significantly Decreases Expression of the Host IAP-1 Gene and Triggers Total Down-Regulation of Baculovirus IAP-3 and Host IAP-Z Genes in LdMNPV-Infected Gypsy Moth Larvae
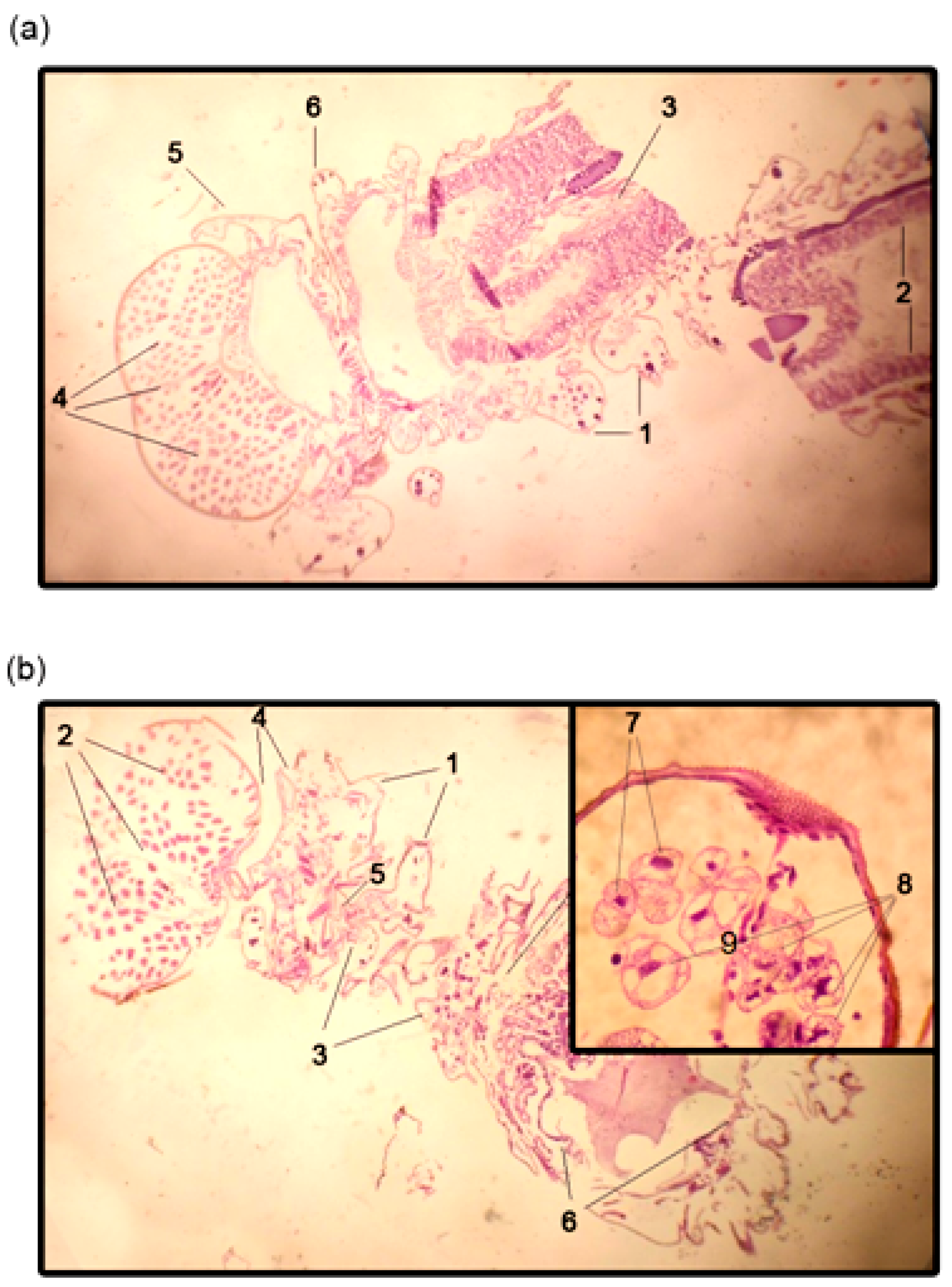
2.4. Evidence for the Development of Apoptosis in oligoRING-Treated Insect Cells
3. Discussion
4. Materials and Methods
4.1. Origin of L. dispar Larvae
4.2. Insect Rearing
4.3. Sequences of the Applied oligoDNAs Fragments
4.4. OligoDNA Treatment of Lymantria Dispar Larvae
4.5. Infection of Lymantria Dispar Larvae with LdMNPV
4.6. Detection of the LdMNPV Infection in L. dispar by PCR
4.7. Search for L. dispar mRNA Homologous to the LdMNPV IAP-3 Gene
4.8. Quantification of Lymantria Dispar IAP-Z and IAP-1 Gene Expression
4.9. Histological Studies
4.10. Apoptosis Detection
4.11. Statistical Analyses
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Correction Statement
References
- Gu, L.; Knipple, D.C. Recent advances in RNA interference research in insects: Implications for future insect pest management strategies. Crop. Protect. 2013, 45, 36–40. [Google Scholar] [CrossRef]
- Zotti, M.J.; Smagghe, G. RNAi technology for insect management and protection of beneficial insects from diseases: Lessons, challenges and risk assessments. Neotrop. Entomol. 2015, 44, 197–213. [Google Scholar] [CrossRef] [PubMed]
- Terenius, O.; Papanicolaou, A.; Garbutt, J.S.; Eleftherianos, I.; Huvenne, H.; Kanginakudru, S.; Albrechtsen, M.; An, C.; Aymeric, J.L.; Barthel, A.; et al. RNA interference in Lepidoptera: An overview of successful and unsuccessful studies and implications for experimental design. J. Insect Physiol. 2011, 57, 231–245. [Google Scholar] [CrossRef] [PubMed]
- Lundgren, J.G.; Duan, J.J. RNAi-based insecticidal crops: Potential effects on nontargetspecies. Bioscience 2013, 63, 657–665. [Google Scholar] [CrossRef]
- Oberemok, V.V. Method of elimination of phyllophagousinsects from order Lepidoptera. UA Patent 36445, 19 May 2008. [Google Scholar]
- Wang, Y.; Zhang, H.; Li, H.; Miao, X. Second-generation sequencing supply an effective way to screen RNAi targets in large scale for potential application in pest insect control. PLoS ONE 2011, 6, e18644. [Google Scholar] [CrossRef] [PubMed]
- Thorp, H.H. The importance of being r: Greater oxidative stability of RNA compared with DNA. Chem. Biol. 2000, 7, R33–R36. [Google Scholar] [CrossRef]
- Oberemok, V.V.; Skorokhod, O.A. Single-stranded DNA fragments of insect-specific nuclear polyhedrosis virus act as selective DNA insecticides for gypsy moth control. Pest. Biochem. Physiol. 2014, 113, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Oberemok, V.V.; Laikova, K.V.; Zaitsev, A.S.; Gushchin, V.A.; Skorokhod, O.A. The RING for gypsy moth control: Topical application of fragment of its nuclear polyhedrosis virus anti-apoptosis gene as insecticide. Pest. Biochem. Physiol. 2016, 131, 32–39. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, M.; Yamada, H.; Ito, H.; Kobayashi, M. Baculovirus IAP1 induces caspase-dependent apoptosis in insect cells. J. Gen. Virol. 2011, 92, 2654–2663. [Google Scholar] [CrossRef] [PubMed]
- Duckett, C.S.; Nava, V.E.; Gedrich, R.W.; Clem, R.J.; Van Dongen, J.L.; Gilfillan, M.C.; Shiels, H.; Hardwick, J.M.; Thompson, C.B. A conserved family of cellular genes related to the baculovirus iap gene and encoding apoptosis inhibitors. EMBO J. 1996, 11, 2685–2694. [Google Scholar]
- Srinivasula, S.M.; Ashwell, J.D. IAPs: What’s in a name? Mol. Cell 2008, 30, 123–135. [Google Scholar] [CrossRef] [PubMed]
- Kuzio, J.; Pearson, M.N.; Harwoo, S.H.; Funk, C.J.; Evans, J.T.; Slavicek, J.M.; Rohrmann, G.F. Sequence and analysis of the genome of a baculovirus pathogenic for Lymantria Dispar. Virology 1999, 253, 17–34. [Google Scholar] [CrossRef] [PubMed]
- Cerio, R.J.; Vandergaast, R.; Friesen, P.D. Host insect inhibitor-of-apoptosis SfIAP functionally replaces baculovirus IAP but is differentially regulated by its N-terminal leader. J. Virol. 2010, 84, 11448–11460. [Google Scholar] [CrossRef] [PubMed]
- Rumble, J.M.; Duckett, C.S. Diverse functions within the IAP family. J. Cell. Sci. 2008, 121, 3505–3507. [Google Scholar] [CrossRef] [PubMed]
- Dias, N.; Stein, C.A. Antisense oligonucleotides: Basic concepts and mechanisms. Mol. Cancer Ther. 2002, 1, 347–355. [Google Scholar] [PubMed]
- Toth, P.P. Antisense therapy and emerging applications for the management of dyslipidemia. J. Clin. Lipidol. 2011, 5, 441–449. [Google Scholar] [CrossRef] [PubMed]
- Schultz, S.J.; Champoux, J.J. RNase H activity: Structure, specificity, and function in reverse transcription. Virus Res. 2008, 134, 86–103. [Google Scholar] [CrossRef] [PubMed]
- Mayr, J.; Grijalvo, S.; Bachl, J.; Pons, R.; Eritja, R.; DíazDíaz, D. Transfection of Antisense Oligonucleotides Mediated by Cationic Vesicles Based on Non-Ionic Surfactant and Polycations Bearing Quaternary Ammonium Moieties. Int. J. Mol. Sci. 2017, 18, 1139. [Google Scholar] [CrossRef] [PubMed]
- Imler, J.L.; Zheng, L. Biology of Toll receptors: Lessons from insects and mammals. J. Leukoc. Biol. 2004, 75, 18–26. [Google Scholar] [CrossRef] [PubMed]
- Tkachev, A.V. Pyrethroid insecticides as analogs of the defensive chemicals of plants. Soros. Obraz. Zh. 2004, 8, 56–63. [Google Scholar]
- Evers, M.M.; Toonen, L.J.A.; van Roon-Mom, W.M.C. Antisense oligonucleotides in therapy for neurodegenerative disorders. Adv. Drug Deliv. Rev. 2015, 87, 90–103. [Google Scholar] [CrossRef] [PubMed]
- Oberemok, V.V. Proof of transovarial transmission of Lymantria disparNucleopolyhedrovirus (Fam. Baculoviridae) with the RAPD-PCR method. Zh. Obshch. Biol. 2008, 69, 397–400. [Google Scholar] [PubMed]
- Rabalski, L.; Krejmer-Rabalska, M.; Skrzecz, I.; Wasag, B.; Szewczyk, B. An alphabaculovirus isolated from dead Lymantria dispar larvae shows high genetic similarity to baculovirus previously isolated from Lymantria monacha—An example of adaptation to a new host. J. Invertebr. Pathol. 2016, 139, 56–66. [Google Scholar] [CrossRef] [PubMed]
- Swevers, L.; VandenBroeck, J.; Smagghe, G. The possible impact of persistent virus infection on the function of the RNAi machinery in insects: A hypothesis. Front. Physiol. 2013, 4, 319. [Google Scholar] [CrossRef] [PubMed]
- Hajek, A.E. Pathology and epizootiology of Entomophaga maimaiga infections in forest Lepidoptera. Microbiol. Mol. Biol. Rev. 1999, 63, 814–835. [Google Scholar] [PubMed]
- Matassov, D.; Kagan, T.; Leblanc, J.; Sikorska, M.; Zakeri, Z. Apoptosis methods and protocols. In Measurement of Apoptosis by DNA Fragmentation; Brady, H.M., Ed.; Humana Press: New York, NY, USA, 2004; pp. 1–17. [Google Scholar]
- Elmore, S. Apoptosis: A review of programmed cell death. Toxicol. Pathol. 2007, 35, 495–516. [Google Scholar] [CrossRef] [PubMed]
- Oberemok, V.V.; Laikova, K.V.; Zaitsev, A.S.; Nyadar, P.M.; Gninenko, Y.I.; Gushchin, V.A.; Makarov, V.V.; Agranovsky, A.A. Topical treatment of LdMNPV-infected gypsy moth caterpillars with 18 nucleotides long antisense fragment from LdMNPV IAP3 gene triggers higher levels of apoptosis in infected cells and mortality of the pest. J. Plant Protect. Res. 2017, 57, 18–24. [Google Scholar] [CrossRef]
- Clarke, T.E.; Clem, R.J. Insect defenses against virus infection: The role of apoptosis. Int. Rev. Immun. 2003, 22, 401–424. [Google Scholar] [CrossRef]
- Mitchell, J.K.; Friesen, P.D. Baculoviruses modulate a proapoptotic DNA damage response to promote virus multiplication. J. Virol. 2012, 86, 13542–13553. [Google Scholar] [CrossRef] [PubMed]
- Simchuk, A.P.; Oberemok, V.V.; Ivashov, A.V. Genetics of interactions among moths, their host plants and enemies in Crimean oak forests, and its perspective for their control. In Moths: Types, Ecological Significance and Control; Cauterruccio, L., Ed.; Nova Science Publisher: New York, NY, USA, 2012; pp. 187–205. ISBN 10:1614706263. [Google Scholar]
- Vandergaast, R.; Mitchell, J.K.; Byers, N.M.; Friesen, P.D. Insect inhibitor-of-apoptosis (IAP) proteins are negatively regulated by signal-induced N-terminal degrons absent within viral IAP proteins. J. Virol. 2015, 89, 4481–4493. [Google Scholar] [CrossRef] [PubMed]
- Agol, V.I. How viruses cause disease. Soros’s Educ. Mag. 1997, 9, 27. [Google Scholar]
- Clem, R.J.; Passarelli, L.A. Baculoviruses: Sophisticated pathogens of insects. PLoS Pathog. 2013, 9, e1003729. [Google Scholar] [CrossRef] [PubMed]
- Оberemok, V.V.; Laikova, K.V.; Zaitsev, A.S.; Gushchin, V.A.; Skorokhod, O.A. Data for increase of Lymantria dispar male survival after topical application of single-stranded RING domain fragment of IAP-3 gene of its nuclear polyhedrosis virus. Data Brief 2016, 7, 514–517. [Google Scholar] [CrossRef] [PubMed]
- Oberemok, V.V.; Laikova, K.V.; Zaitsev, A.S.; Nyadar, P.M.; Shumskykh, M.N.; Gninenko, Y.I. DNA insecticides based on IAP3 gene fragments of cabbage looper and gypsy moth nuclear polyhedrosis viruses show selectivity for non-target insects. Arch. Biol. Sci. 2015, 67, 785–792. [Google Scholar] [CrossRef]
- Mutah, N.P.; Adesoji, A.T. DNA insecticides: The lethal potency of LdMNPV IAP-2 gene antisense oligonucleotides in pre-infected gypsy moth (Lymantria dispar L.) larvae. Int. J. Pest Manag. 2017. [Google Scholar] [CrossRef]
- Zhang, Z.; Schwartz, S.; Wagner, L.; Miller, W. A greedy algorithm for aligning DNA sequences. J. Comput. Biol. 2000, 7, 203–214. [Google Scholar] [CrossRef] [PubMed]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef] [PubMed]

| Experimental Groups | χ2 Values | Total Number of Larvae in Two Compared Groups | |
|---|---|---|---|
| LdMNPV-free | oligoCpG vs. water control | 1.11 | 150 |
| oligoRING vs. water control | 0.08 | 151 | |
| LdMNPV-infected | oligoCpG vs. water control | 0.05 | 117 |
| oligoRING vs. water control | 9.63 ** | 118 | |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oberemok, V.V.; Laikova, K.V.; Zaitsev, A.S.; Shumskykh, M.N.; Kasich, I.N.; Gal’chinsky, N.V.; Bekirova, V.V.; Makarov, V.V.; Agranovsky, A.A.; Gushchin, V.A.; et al. Molecular Alliance of Lymantria dispar Multiple Nucleopolyhedrovirus and a Short Unmodified Antisense Oligonucleotide of Its Anti-Apoptotic IAP-3 Gene: A Novel Approach for Gypsy Moth Control. Int. J. Mol. Sci. 2017, 18, 2446. https://doi.org/10.3390/ijms18112446
Oberemok VV, Laikova KV, Zaitsev AS, Shumskykh MN, Kasich IN, Gal’chinsky NV, Bekirova VV, Makarov VV, Agranovsky AA, Gushchin VA, et al. Molecular Alliance of Lymantria dispar Multiple Nucleopolyhedrovirus and a Short Unmodified Antisense Oligonucleotide of Its Anti-Apoptotic IAP-3 Gene: A Novel Approach for Gypsy Moth Control. International Journal of Molecular Sciences. 2017; 18(11):2446. https://doi.org/10.3390/ijms18112446
Chicago/Turabian StyleOberemok, Volodymyr V., Kateryna V. Laikova, Aleksei S. Zaitsev, Maksym N. Shumskykh, Igor N. Kasich, Nikita V. Gal’chinsky, Viktoriya V. Bekirova, Valentin V. Makarov, Alexey A. Agranovsky, Vladimir A. Gushchin, and et al. 2017. "Molecular Alliance of Lymantria dispar Multiple Nucleopolyhedrovirus and a Short Unmodified Antisense Oligonucleotide of Its Anti-Apoptotic IAP-3 Gene: A Novel Approach for Gypsy Moth Control" International Journal of Molecular Sciences 18, no. 11: 2446. https://doi.org/10.3390/ijms18112446
APA StyleOberemok, V. V., Laikova, K. V., Zaitsev, A. S., Shumskykh, M. N., Kasich, I. N., Gal’chinsky, N. V., Bekirova, V. V., Makarov, V. V., Agranovsky, A. A., Gushchin, V. A., Zubarev, I. V., Kubyshkin, A. V., Fomochkina, I. I., Gorlov, M. V., & Skorokhod, O. A. (2017). Molecular Alliance of Lymantria dispar Multiple Nucleopolyhedrovirus and a Short Unmodified Antisense Oligonucleotide of Its Anti-Apoptotic IAP-3 Gene: A Novel Approach for Gypsy Moth Control. International Journal of Molecular Sciences, 18(11), 2446. https://doi.org/10.3390/ijms18112446









