1. Introduction
Since increasing interleukin (IL)-17 concentrations were observed in autoimmune diseases, increased attention has led to a precise definition of the role of human Th17 cells and their products in the process of ongoing immune response involved tissues [
1,
2,
3]. The role of Th17 cells and Th17-associated cytokines in the pathogenesis of rheumatoid arthritis (RA) is now widely recognized. Th17 cells, which are the dominant effector T cell involved in the induction of autoimmune chronic tissue by promoting the immune response, are much more effective in inducing inflammation than any other Th1 cell recognized thus far and is therefore the “main culprit” of autoimmune diseases [
4]. Th17 cell produce several cytokines including IL-17, IL-17F, IL-6, IL-21, IL-22 and tumor necrosis factor-alpha (TNF-α) that play a key role not only in the RA but also in the pathogenesis of experimental autoimmune encephalomyelitis (EAE) and collagen induced arthritis (CIA), and they induces dendritic cells (DCs) to produce IL-12 and interferon (IFN-c) [
5,
6,
7,
8,
9,
10]. Th17 cell present in the joint may create a positive feedback loop leading to the continuous activation of T cells, which is a critical event in the generation of autoimmunity [
11].
Although Th17 cells express numerous markers, human
RORc2 (nuclear hormone retinoic acid receptor-related orphan receptor variant 2; a short isoform of
RORc gene, encoded by the human
RORc gene located on chromosome 1q21–q23) ortholog of mice
RORγt, is a master transcriptional factor that can drive Th17 differentiation in both humans and mouse, respectively [
12,
13,
14,
15,
16]. Overexpression of
RORc2 by inducing IL-17, IL-26, TCR and CCR6 stimulates a wide range of phenotypic and functional programming during Th17 cells differentiation [
17,
18]. Knockdown of transcription factor RORc cause high Foxp3 levels and reduces expression of pro-inflammatory cytokines such as IL-1β, IL-6, IL-17, IFN-γ and TNF-α, suggesting that the role of
RORc2 in Th17 cells differentiation involves not only in induction of Th17 characteristics genes, but also suppression of Treg cells specific programs [
18,
19,
20].
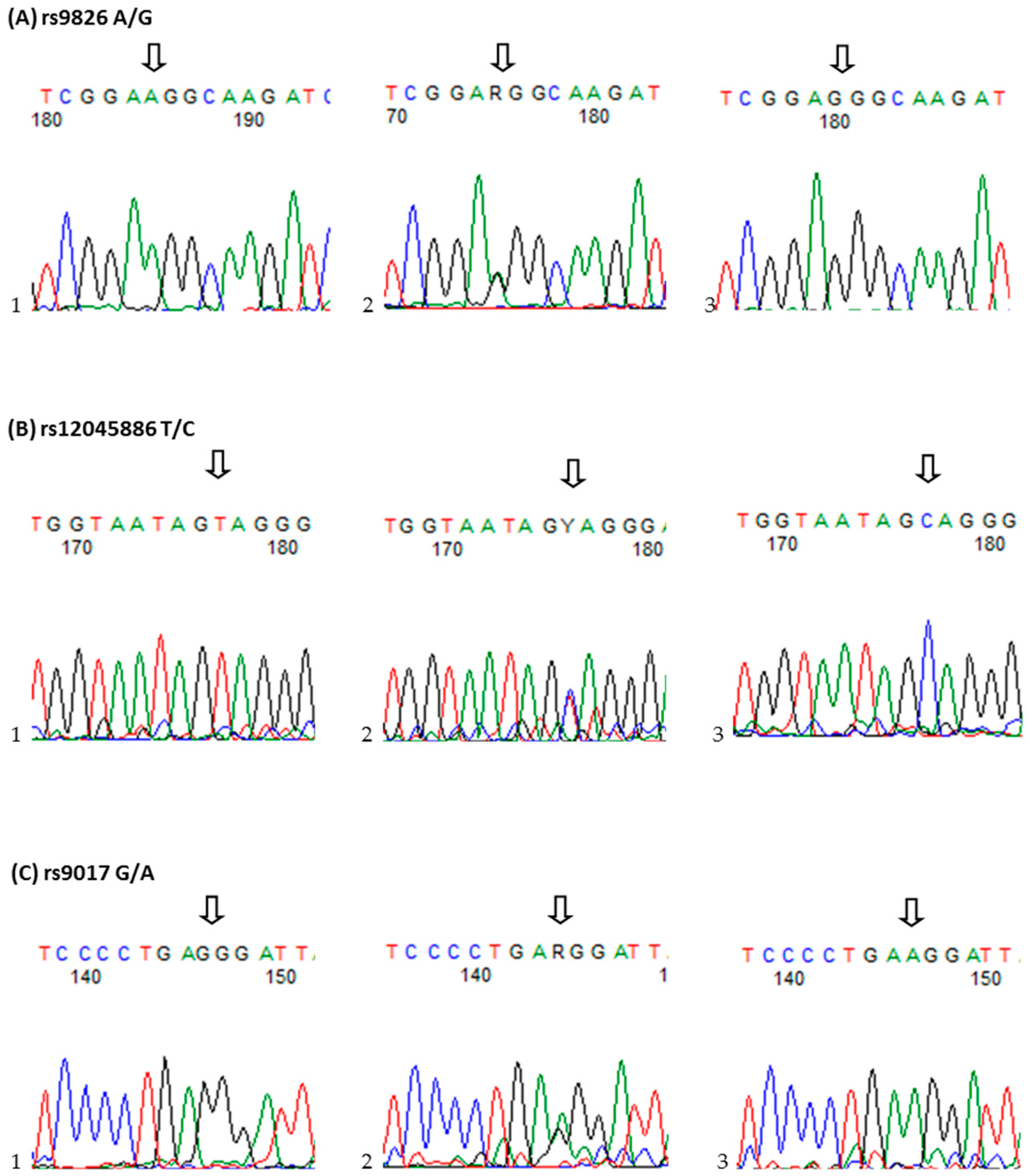
The research of the Th17 cells development may help better understand its role in the pathogenesis of RA. However, there was still no too much study in this field. In this study, we hypothesized that the RORc2 gene is responsible for Th17 cells and IL-17-producing Treg cells differentiation may also be a strong molecular candidate for rheumatoid arthritis severity and/or susceptibility. To test this hypothesis, we examined, for the first time, three candidate single nucleotide polymorphisms (SNPs) in the RORc2 gene, rs9826 A/G, rs12045886 T/C and rs9017 G/A, and RORc protein expression and determined their possible association with susceptibility to and clinical phenotype of RA in Polish population.
3. Discussion
Over the last decade, numerous studies have disclosed that both Th1 cells as well as Th17 cells are involved in the pathogenesis of RA, where higher frequency of Th17 cells had been found in the arthritic synovium [
21,
22]. In addition to RA, it has been presented that Th17 cells are also involved in the pathogenesis of other autoimmune diseases through the promotion of tissue inflammation, the mobilization of the innate or adaptive immune system and secretion of the pro-inflammatory cytokines and chemokines [
21,
23,
24]. Because RORc2 function as a lineage-specifying transcriptional factor of Th17 cells and play a key role in development and maintenance of these cells [
12,
19,
25,
26],
RORc2 gene may represent a candidate gene for autoimmune diseases. However, not too much is known about the function of
RORc2 genetic variants in autoimmune diseases, including RA. The
RORc2 gene polymorphisms have been analysis only in a Behcet’s disease (BD) [
27], type 2 diabetes (T2DM) [
15] and secondary lymphedema [
27]. This is why analysis of polymorphisms within the
RORc2 gene may help to uncover their correlations with some biochemical and laboratory findings. Liao D.
et al. [
27] in the sole report exploring
RORc genetic variants in BD studied the 25 selected taqSNPs and copy number variants (CNVs) in region of
RORc gene. They concluded that the high RORc CNV was associated with susceptibility to BD and that the RORc mRNA expression levels were increased in patients with high
RORc copy number. The study of Wang H.
et al. [
15] demonstrated that from 11 examined SNPs, in African American individuals and in Caucasian individuals from northern European, only the Ala454Gly variant was three-fold more common among African American patients with T2DM than in healthy subjects. Furthermore, Newman B.
et al. [
20] found that
RORc tagSNP rs11801866 located at the 5′ end of
RORc gene was associated with susceptibility to secondary lymphedema and together with rs12128071 are predicted to affect transcription factor binding sites.
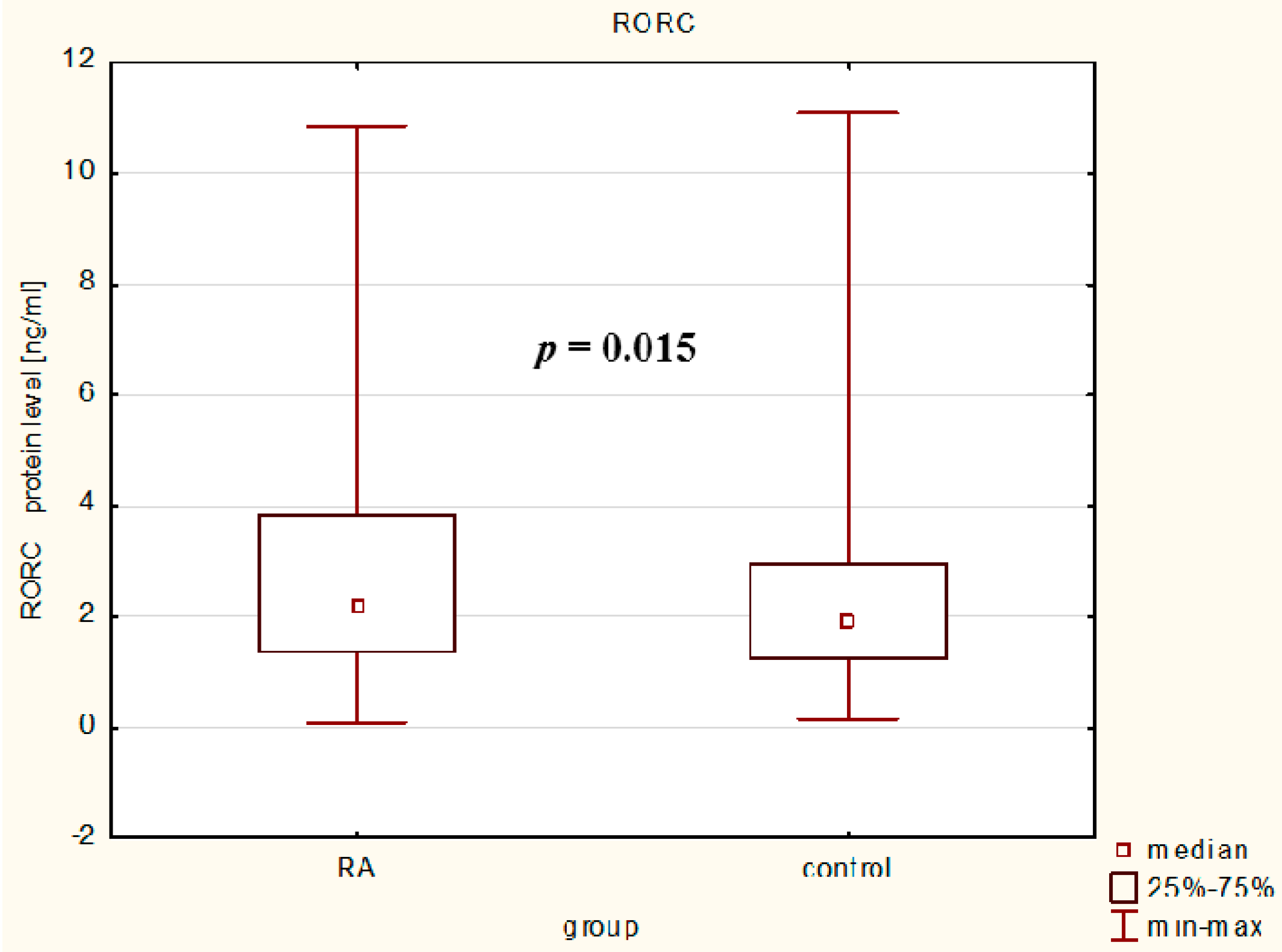
In our study, we observed that serum RORc levels were higher in RA patients than in controls reflecting the ongoing inflammatory process in rheumatoid arthritis and it confirms and extends the hypothesis that Th17 enrichment occurs in inflamed joints [
13]. Some previous reports have presented the RORc mRNA expression [
13,
18,
20,
23,
28], but our study is the first that exploring the RORc protein levels in the serum of RA patients. Our results indicate that RA patients with higher median value of RORc protein levels have higher disease activity and in our opinion, probably an increased number of Th17 cells than RA patients with lower median value of RORc protein levels and controls. As shown in earlier studies, active RA patients had a significantly higher frequency of peripheral Th17 cells and RORc mRNA expression than inactive RA patients and healthy subjects suggesting a potential predominant role of Th17 cells in the inflammatory process and chronic progression in RA [
21,
29]. We conclude that by induction of Th17-characteristics genes, suppression of regulatory T cell-specific program and controlling pro-inflammatory cytokines/chemokines expression, RORc might play an essential role in the regulation of inflammatory processes and therefore would represent an innovative therapeutic regiment for the balance between immunity and tolerance.
Considering an important role of the RORc2 in the development of the Th17/Treg cells and differences in genetic predispositions between populations [
13,
18,
19,
30], we decided to carry out an analysis of selected
RORc2 genetic variants in relation to RA. To the best our knowledge, this study is the first in which the associations between not only serum RORc levels, but also
RORc2 gene SNPs and haplotypes have been investigated in a cohort of patients with rheumatoid arthritis. For the analysis we chose the rs9826 A/G and rs9017 G/A polymorphisms located in the 3′UTR, and rs12045886 T/C polymorphism located in an intronic region. SNPs located in the 3′UTR region of the
RORc2 gene may interfere with mRNA translation and stability through effects on polyadenylation and regulatory miRNA-mRNA and protein-mRNA interactions, thus affecting the level of protein expression. However, this needs to be verified in further studies.
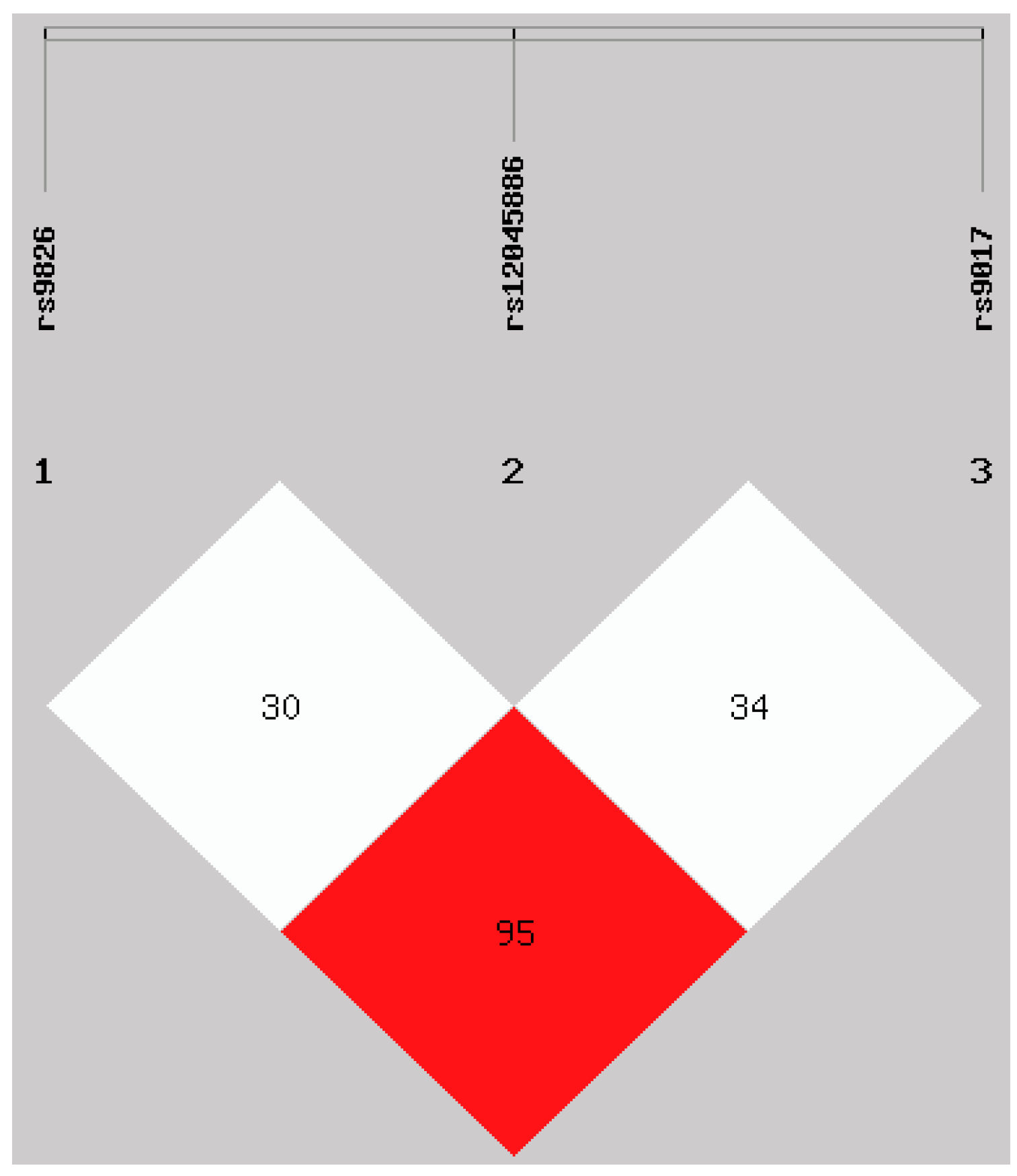
Interestingly, when we studied each of the polymorphisms separately, our results have shown no association with the susceptibility to RA in Polish population. A relatively small population size might have prevented from seeing the association. The proportion of genetic contribution of certain polymorphic locus to RA susceptibility may be influenced by other local environmental and/or joint-specific genetic factors most of which were postulated to be involved in cell signaling. Moreover, when we examined all the polymorphisms to create haplotypes, our study showed that a protective effect of the ATA haplotype against the risk of rheumatoid arthritis in our populations, suggesting that the impact of gene on disease risk may not limit to single SNP. Additionally, the results derived from our study are the first, which showing haplotype-based association analysis for these three genetic variants in the RORc2 gene.
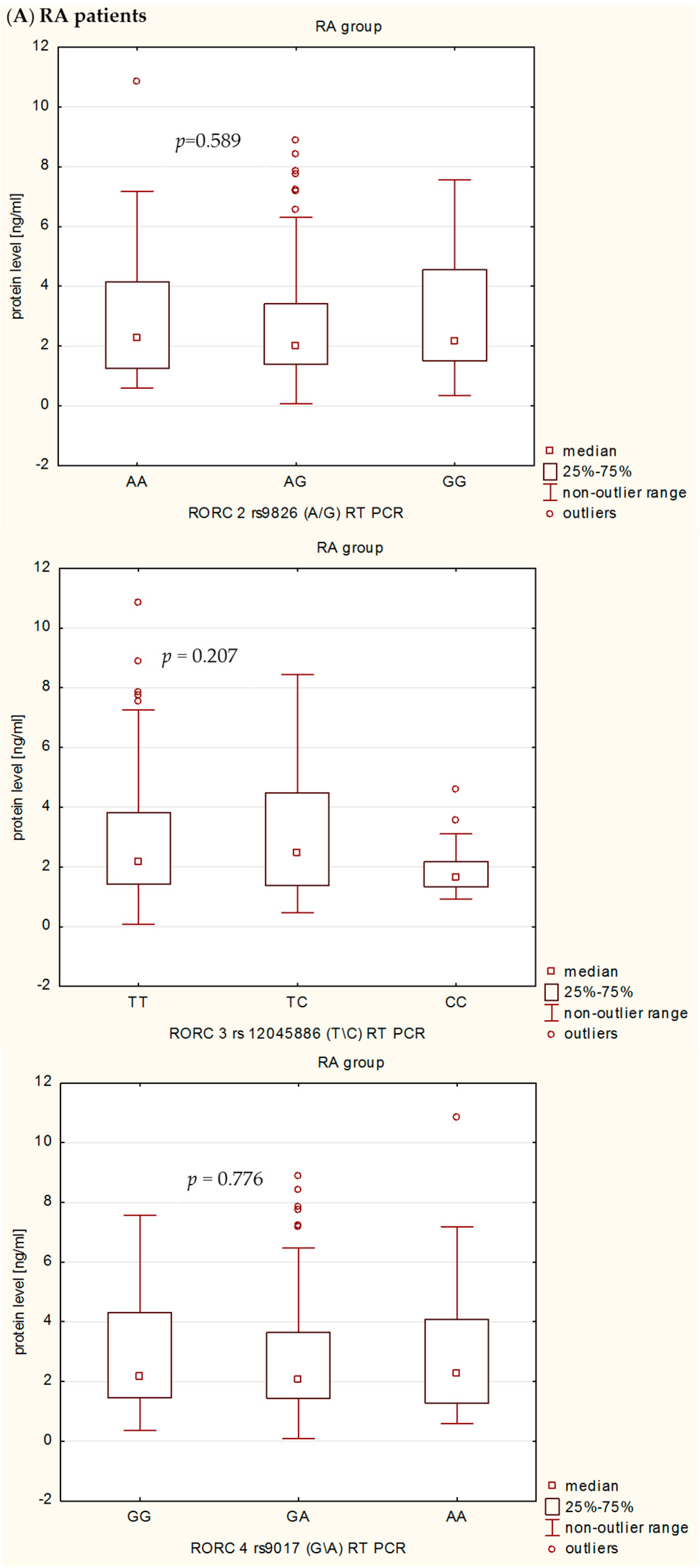
In the present study, we also investigated the influence of rs9826 A/G, rs12045886 T/C and rs9017 G/A RORc2 gene polymorphisms on clinical phenotype and RORc serum levels in RA patients. Our detailed genotype-phenotype analysis indicated that without Bonferroni correction: (i) the RORc2 rs9826 A/G polymorphisms under recessive model ((AA + AG) vs. GG) was associated with the disease duration of RA, median value of CRP, VAS score, and creatinine level; (ii) the rs12045886 T variant was associated with a significantly higher median value of PLT and lower number of women; and (iii) the carriers of the combined rs9017 GA + AA genotype were a higher median value of CRP, VAS, HAQ and DSA-28 score. However, after Bonferroni correction, there was an association only between RORc2 rs9826 A/G and rs9017 G/A SNPs and median value of CRP. In addition, we can also observed that RA patients with the RORc2 rs9017A allele had more advanced disease than rs9017 G allele carriers, suggesting that this polymorphism might be associated with higher disease activity. We also found evidence for association of the RORc2 rs9826 A/G, rs12045886 T/C and rs9017 G/A polymorphisms with difference in RORc serum levels between RA patients and controls. RORc serum levels were significantly higher in RA patients with rs9826AA, rs12045886TT and TC, and rs9017AA genotypes compared to healthy subjects with the same genotypes. Moreover, RA patients with rs9017AA genotype had a tendency to higher RORc levels in serum than RA patients with rs9017GG or GA genotypes. The expression level of the transcriptional factors within the target cells is a reflection of the transcriptional processes, whereas in the serum reflects passively the undergoing regulation. Moreover, we believe that analysis of transcriptional factors level in the serum may reflect pathological processes such as cell damage. In our opinion, RORc might increase with disease progression as their serum levels would depend much more on the number and activity of Th17 cells, which are typical pro-inflammatory cells promoting the induction of autoimmune tissue inflammation and plays a key role in the development of autoimmune arthritis, than on the sole genotype. Despite this observation, we infer that the analyzed polymorphisms might still affect RORc2 mRNA and protein intracellular expression. However, further studies with larger sample size will be needed to validate the genetic effects of the RORc2 gene polymorphisms on RA. Moreover, we believe that our results could be helpful to clarify the role of RORc in the pathogenesis of RA, which it might be a candidate factor consistent with the severity of disease.
4. Materials and Methods
4.1. Study Participants
In total, 591 patients with established RA and 341 unrelated healthy controls were included in this study. Blood samples were obtained from the patients recruited from the National Institute of Geriatrics, Rheumatology and Rehabilitation in Warsaw and Pomeranian Medical University in Szczecin, Poland. All patients had a diagnosis of RA, and met the RA classification criteria published by the American College of Rheumatology in 1987. Clinical and biochemical data were collected from patient’s files and questionnaires after signed informed consent and are presented in
Table 8.
The control group (217 females and 124 males, age between 18 and 63 years) consisted of healthy volunteers who showed no clinical and/or laboratory signs of any autoimmune disease. They were selected randomly from blood bank donors. Patients and healthy subjects were from the same geographical area and they had the same socioeconomic status. For the present study, we selected a representative sample of the admixed urban Polish population.
The study was reviewed and approved by the Research Ethics Committee of the Institute of Rheumatology and Pomeranian Medical University.
4.2. DNA Extraction
Genomic DNA from RA patients and healthy subjects was extracted from whole blood using the standard isothiocynate guanidine (GTC) extraction method and/or the QIAamp DNA Blood Mini Kit (Qiagen, Hilden, Germany).
4.3. Single-Nucleotide Polymorphisms Selection and Genotyping
The three RORc2 (NC_000001.11) SNPs studied, rs9826 A/G, rs12045886 T/C and rs9017 G/A, were selected on the basis of HapMap data release 24/phase II, November 2008, NCBI build 36, dbSNP b126.Genotyping was analyzed using TaqMan SNP genotyping assay (Applied Biosystems, Foster City, CA, USA): C_9624797_20 (rs9826), C_11260074_10 (rs12045886), and C_9624790_20 (rs9017). The reaction was performed in 10 µL volumes on StepOne real-time PCR system in RotorGene 6000 Real-Time PCR system with the fallowing amplification protocol: denaturation at 95 °C for 10 min, followed by 40 cycles of denaturation at 92 °C for 15 s, and annealing and extension at 60 °C for 1 min.
4.4. Serum Levels of RORC Determination
Blood samples were obtained from patients and controls, centrifuged and stored at −86 °C until analysis. Serum RORc levels (ng/mL) were determined by enzyme-linked immunosorbent assay kits (ELISA: Wuhan EIAab Science Co., Ltd., Wuhan, China), according to the manufacturer’s instructions. The minimum level of detection for RORc was 0.15 ng/mL. Plates were read at an absorbance of 450 nm on Elx800 reader (BIO-TEK Instruments, Winooski, VT, USA).
4.5. Statistical Analysis
Deviations from Hardy–Weinberg equilibrium expectations were determining using the software available at Institute of Human Genetics, Germany. To evaluate the extent of linkage disequilibrium (LD), D′ and
r2 between pairs of polymorphisms was quantified using the SHEsis software [
31,
32]. Logistic regression under three genetic models (codominant, dominant and recessive) was used to estimate the odds ratio (OR) and 95% confidence interval (CI) for the risk genotype. A probability value of
p < 0.017 (according to Bonferroni correction) was considered significant. Normality of the distribution of continuous variables was assessed using the Shapiro–Wilk test. The correlation between genetic variants and phenotype of RA were compared by: (1) Fisher exact test (depending on expected values) or χ
2 test or χ
2 test with Yates’ correction for categorical variables; and (2) Mann–Whitney
U test for continuous variable. Continuous variables are expressed as the median and interquartile range (IQR), whereas categorical variables as number and percentage. Bonferroni correction was used to adjust
p-values for multiple measures (Bonferroni-corrected α-level = 0.05/16). Statistical significance was assumed at the
p < 0.003 level.