Genetic Variability and Population Structure of Salvia lachnostachys: Implications for Breeding and Conservation Programs
Abstract
:1. Introduction
2. Results
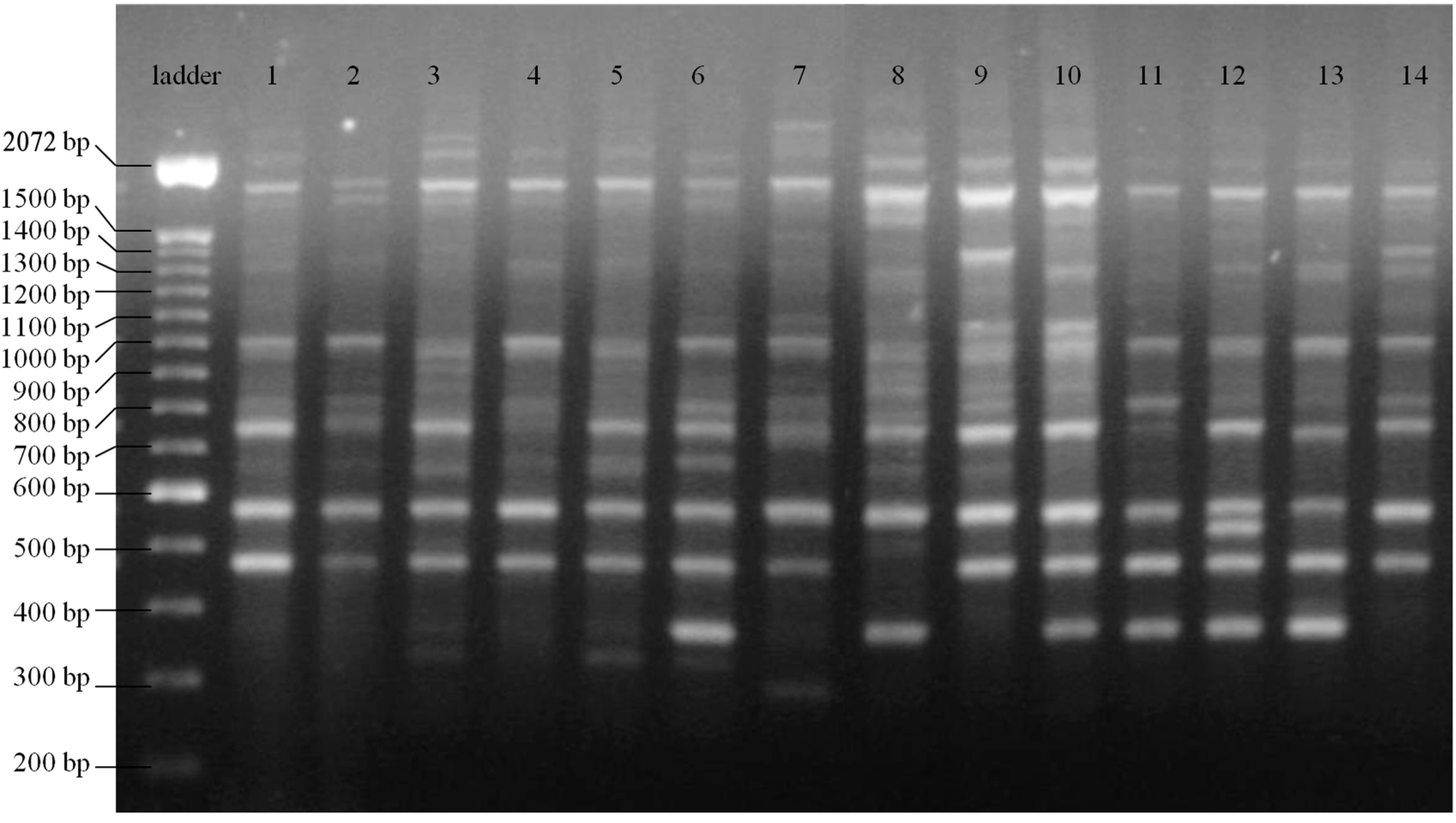
| Sequence (5'→3') | AT (°C) | NB | NPB | PPB (%) | Simpson’s Index | Reference |
|---|---|---|---|---|---|---|
| (AC)8T | 51.4 | 14 | 13 | 93 | 0.79 | [20] |
| (AG)8A | 46.7 | 12 | 11 | 92 | 0.84 | [20,21] |
| (AG)8C | 48.8 | 20 | 20 | 100 | 0.83 | [22] |
| (AG)8YC | 50.2 | 22 | 22 | 100 | 0.86 | [20] |
| (AG)8YT | 49.2 | 19 | 19 | 100 | 0.77 | [21,22] |
| (CA)8G | 51.0 | 19 | 19 | 100 | 0.78 | [20,21] |
| (CT)8A | 44.7 | 10 | 10 | 100 | 0.66 | [21] |
| (CTC)4RC | 51.7 | 22 | 19 | 86 | 0.77 | [20,21] |
| (GA)8T | 45.4 | 21 | 19 | 90 | 0.80 | [20,21,22] |
| Total | - | 159 | 152 | 96 | 0.79 | - |
| Average | - | 17.6 | - | - | - | - |
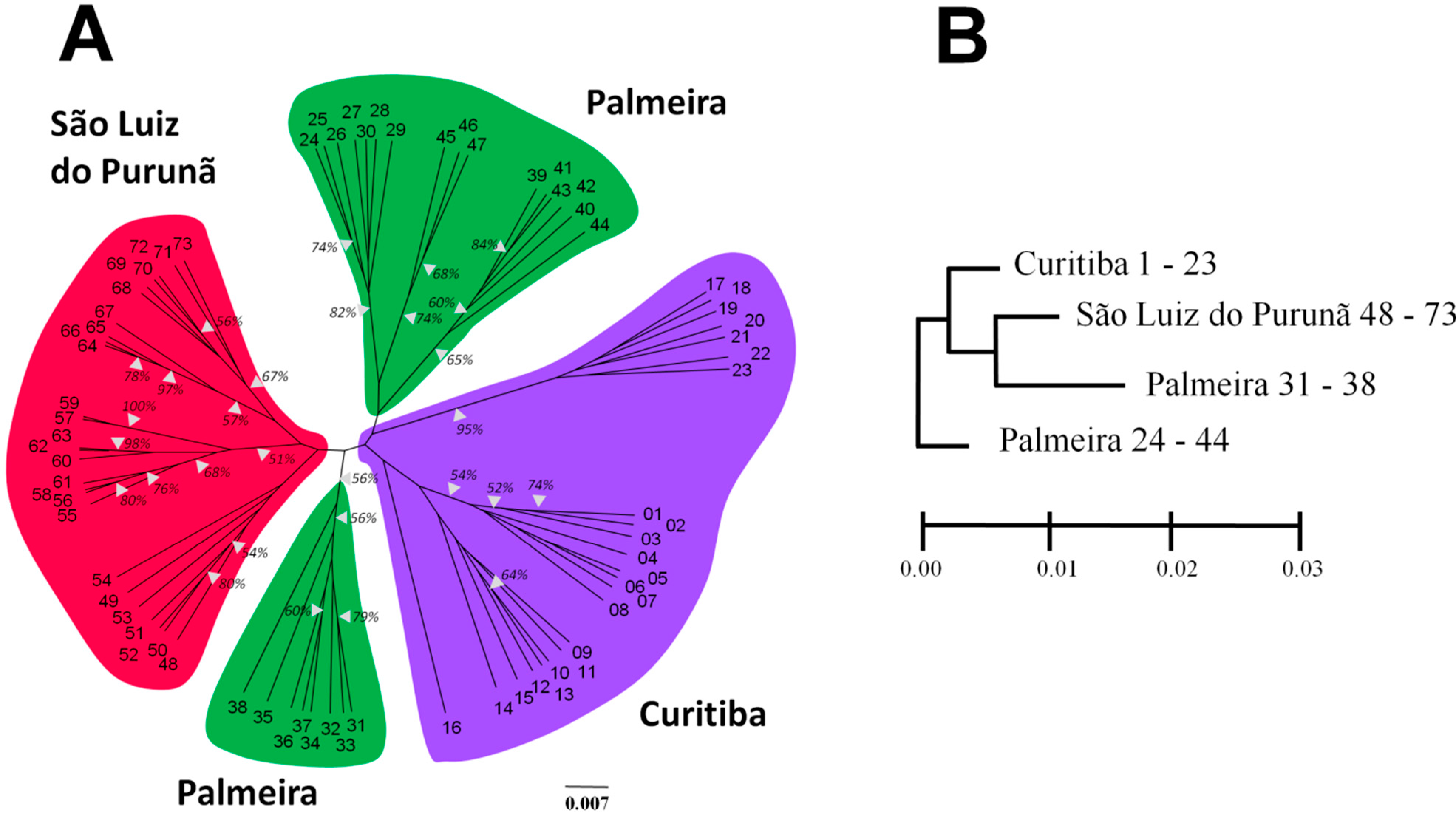
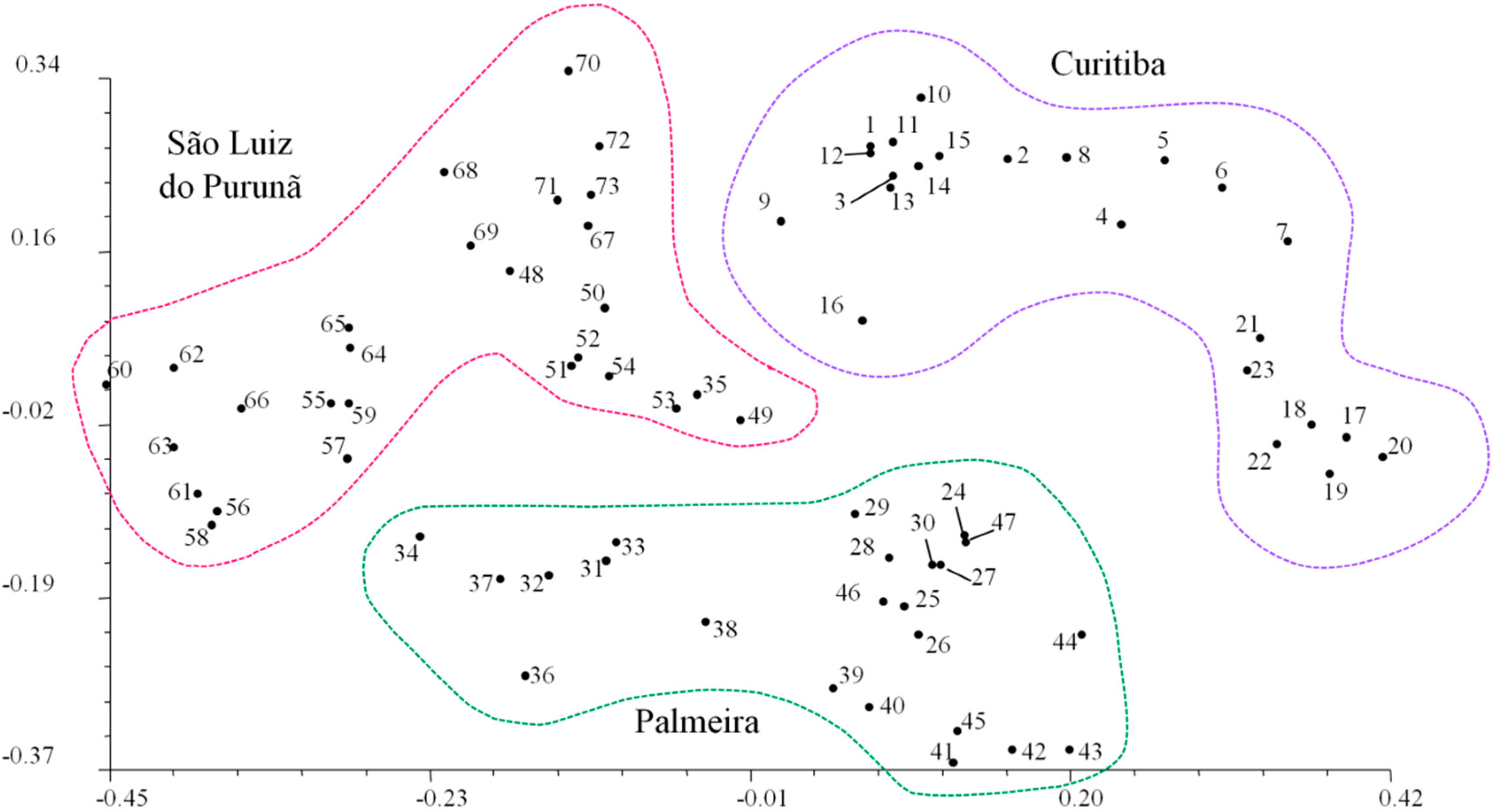
| Populations | N | M | NPB | PPB | Ao | Ae | H | I |
|---|---|---|---|---|---|---|---|---|
| Curitiba | 23 | 23 | 130 | 81.76 | 1.82 | 1.39 | 0.24 | 0.37 |
| Palmeira | 24 | 23 | 121 | 76.10 | 1.76 | 1.33 | 0.21 | 0.33 |
| São Luiz Purunã | 26 | 25 | 111 | 69.81 | 1.70 | 1.33 | 0.20 | 0.31 |
| Total | 73 | 70 | 155 | 97.48 | 1.97 | 1.39 | 0.25 | 0.40 |
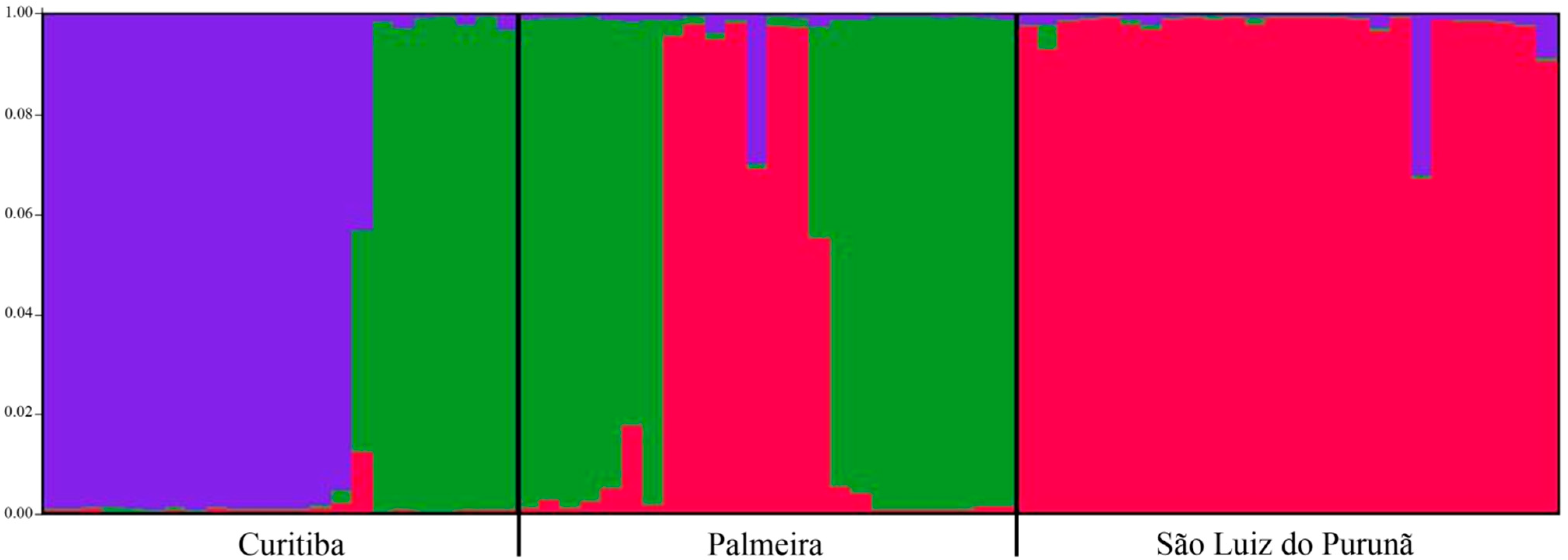
3. Discussion
4. Experimental Section
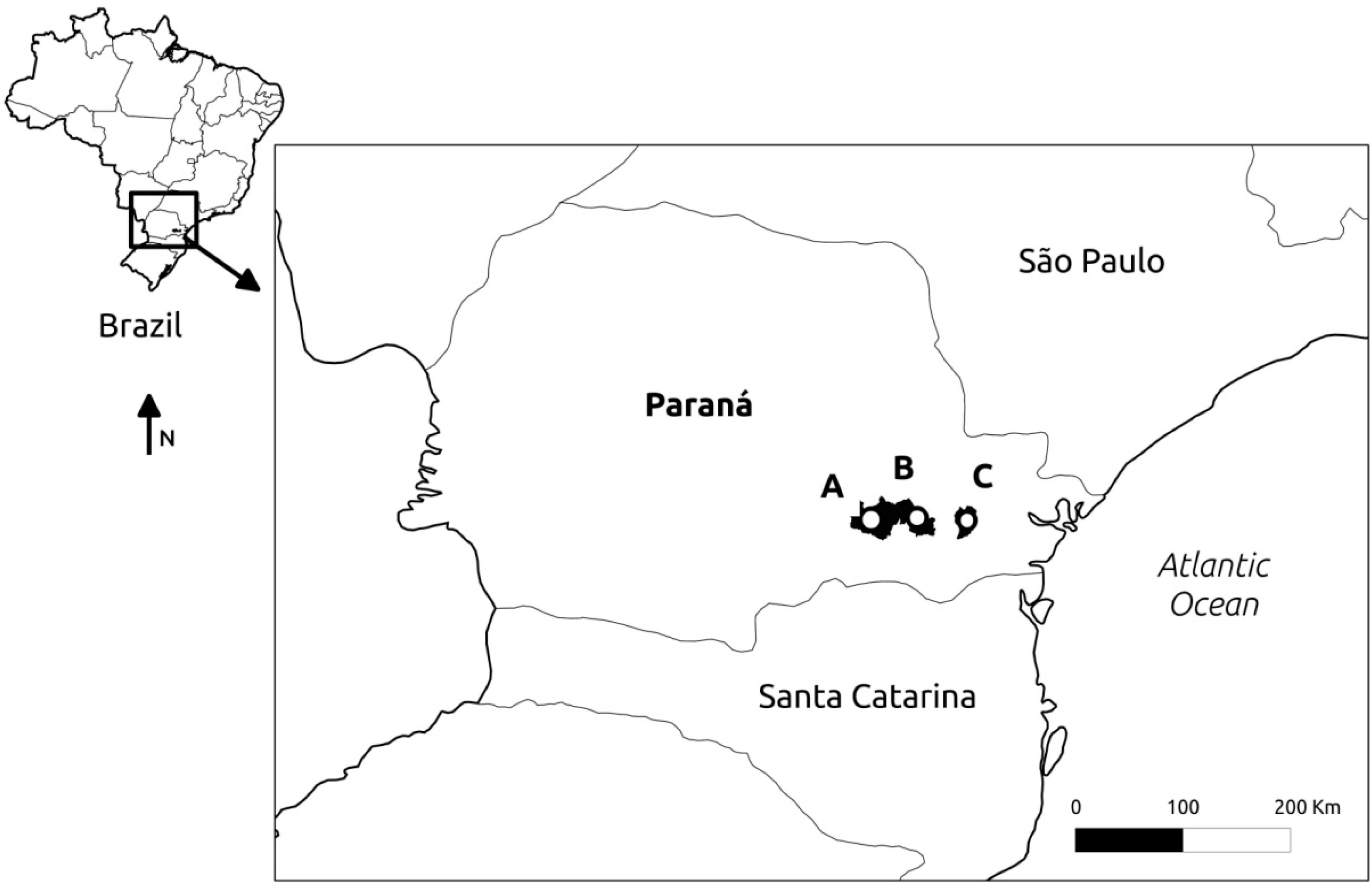
4.1. Sampling and DNA Extraction
| Location | Latitude S | Longitude W | Altitude (m) | Voucher Number | No. of Samples |
|---|---|---|---|---|---|
| Curitiba | 25°30'22" | 49°18'30" | 932 | E.P. Santos 1251 | 23 |
| Palmeira | 25°28'10" | 49°48'05" | 1069 | E.P. Santos 1264 | 24 |
| São Luiz do Purunã | 25°28'13" | 49°39'33" | 1089 | E.P. Santos 1266 | 26 |
4.2. ISSR-PCR Amplification
4.3. Data Analysis
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Harley, R.M.; Atkins, S.; Budantsev, A.L.; Cantino, P.D.; Conn, B.J.; Grayer, R.; Harley, M.M.; Kok, R.; Krestovsaja, T.; Morales, R.; et al. Labiatae. In The Families and Genera of Vascular Plant; Kadereit, J.W., Kubitzki, K., Eds.; Springer-Verlag: Berlin, Germany, 2004; Volume 7, pp. 167–236. [Google Scholar]
- Santos, E.P. Salvia L. In Lista de Espécies da Flora do Brasil. Available online: http://www.floradobrasil.jbrj.gov.br/jabot/floradobrasil/FB8296 (accessed on 19 December 2014).
- Cheng, T.O. Cardiovascular effects of Danshen. Int. J. Cardiol. 2007, 121, 9–22. [Google Scholar] [CrossRef] [PubMed]
- Hosseinzadeh, H.; Haddakhodaparast, M.H.; Arash, A.R. Antinociceptive, antiinflammatory and acute toxicity effects of Salvia leriifolia Benth. seed extract in mice and rats. Phytother. Res. 2003, 17, 422–425. [Google Scholar] [CrossRef]
- Mayer, B.; Baggio, C.H.; Freitas, C.S.; Santos, C.; Twardowschy, A.; Horst, H.; Pizzolatti, M.G.; Micke, G.H.; Heller, M.; Santos, E.P.; et al. Gastroprotective constituents of Salvia officinalis L. Fitoterapia 2009, 80, 421–426. [Google Scholar] [CrossRef]
- Fan, H.Y.; Fu, F.H.; Yang, M.Y.; Xu, H.; Zhang, A.H.; Liu, R. Antiplatelet and antithrombotic activities of salvianolic acid A. Thromb. Res. 2010, 126, 17–22. [Google Scholar] [CrossRef]
- Veitch, N.C.; Smith, M.; Barnes, J.; Anderson, L.A.; Phillipson, J.D. Herbal Medicines, 4th ed.; Pharmaceutical Press: London, UK, 2013. [Google Scholar]
- Jimena, E.S.; França, F.; Sobral, M. Plantas da Floresta Atlântica; Jardim Botânico do Rio de Janeiro: Rio de Janeiro, Brazil, 2009; pp. 297–303. [Google Scholar]
- Erbano, M.; Ehrenfried, C.A.; Stefanello, M.E.A.; Santos, E.P. Morphoanatomical and phytochemical studies of Salvia lachnostachys (Lamiaceae). Microsc. Res. Techniq. 2012, 75, 1737–1744. [Google Scholar] [CrossRef]
- Kassuya, C.A.L.; Wisniewski-Junior, A.; Simionatto, E.L.; Santos, E.P.; Stefanello, M.E.A. Composição dos óleos essenciais de Salvia lachnostachys e Salvia melissiflora (Lamiaceae). Lat. Am. J. Pharm. 2009, 28, 919–921. [Google Scholar]
- Piccinelli, A.C.; Aquino, D.F.S.; Morato, P.N.; Kuraoka-Oliveira, A.M.; Strapasson, R.L.B.; Santos, E.P.; Stefanello, M.E.A.; Oliveira, R.J.; Kassuya, C.A.L. Anti-inflammatory and antihyperalgesic activities of ethanolic extract and fruticulin A from Salvia lachnostachys leaves in mice. Evid. Based Complement. Alternat. Med. 2014, 2014. [Google Scholar] [CrossRef] [PubMed]
- Gupta, V.S.; Ramakrishna, W.; Rawat, S.R.; Ranjekar, P.K. (CAC)5 detects DNA fingerprinting and sequence homologous to gene transcripts in rice. Biochem. Genet. 1994, 32, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Barth, S.; Melchinger, A.E.; Lübberstedt, T.H. Genetic diversity in Arabidopsis thaliana L. Heynh. investigated by cleaved amplified polymorphic sequence (CAPS) and inter-simple sequence repeat (ISSR) markers. Mol. Ecol. 2002, 11, 495–505. [Google Scholar]
- Bornet, B.; Branchard, M. Nonanchored Inter Simple Sequence Repeat (ISSR) markers: Reproducible and specific tools for genome fingerprinting. Plant Mol. Biol. Rep. 2001, 9, 209–215. [Google Scholar] [CrossRef]
- Bai, C.; Wen, M.; Zhang, L.; Li, G. Genetic diversity and sampling strategy of Scutellaria baicalensis germplasm resources based on ISSR. Genet. Resour. Crop. Evol. 2013, 60, 1673–1685. [Google Scholar] [CrossRef]
- Qian, X.; Li, Q.J.; Liu, F.; Gong, M.J.; Wang, C.X.; Tian, M. Conservation genetics of an endangered Lady’s Dlipper Orchid: Cypripedium japonicum in China. Int. J. Mol. Sci. 2014, 15, 11578–11596. [Google Scholar] [CrossRef] [PubMed]
- Kurane, J.; Shinde, V.; Harsulkar, A. Application of ISSR marker in pharmacognosy: Current update. Pharmacogn. Rev. 2009, 3, 216–228. [Google Scholar]
- Lüdtke, R.; Agostini, G.; Miotto, S.T.S.; Souza-Chies, T.T. Characterizing Polygala L. (Polygalaceae) species in Southern Brazil using ISSR. Plant Mol. Biol. Rep. 2010, 28, 317–323. [Google Scholar]
- Mittermeier, R.A.; Fonseca, G.A.B.; Rylands, A.B.; Brandon, K. Uma breve história da conservação da biodiversidade do Brasil. Megadiversidade 2005, 19, 601–607. [Google Scholar]
- Agostini, G.; Echeverrigaray, S.; Souza-Chies, T.T. Genetic relationships among South American species of Cunila D. Royen ex L. based on ISSR. Plant Syst. Evol. 2008, 274, 135–141. [Google Scholar]
- Fracaro, F.; Echeverrigaray, S. Genetic variability in Hesperozygis ringens Benth. (Lamiaceae), an endangered aromatic and medicinal plant of Southern Brazil. Biochem. Genet. 2006, 44, 479–490. [Google Scholar]
- Rahimmalek, M.; Bahreininejad, B.; Khorrami, M.; Tabatabaei, B.E.S. Genetic variability and geographic differentiation in Thymus daenensis subsp. daenensis, an endangered medicinal plant, as reaveled by inter simple sequence repeat (ISSR) markers. Biochem. Genet. 2009, 47, 831–842. [Google Scholar]
- Song, Z.; Li, X.; Wang, H.; Wang, J. Genetic diversity and population structure of Salvia miltiorrhiza Bge in China revealed by ISSR and SRAP. Genetica 2010, 138, 241–249. [Google Scholar] [CrossRef] [PubMed]
- Javan, Z.S.; Rahmani, F.; Heidari, R. Assessment of genetic variation of genus Salvia by RAPD and ISSR markers. Aust. J. Crop. Sci. 2012, 6, 1068–1073. [Google Scholar]
- Zhang, Y.; Li, X.; Wang, Z. Diversity evaluations of Salvia miltiorrhiza using ISSR markers. Biochem. Genet. 2013, 51, 707–721. [Google Scholar] [CrossRef] [PubMed]
- Guo, B.L.; Lin, S.; Feng, Y.X.; Zhao, Y.J. Primary research on genetic relationship among main populations of Salvia miltiorrhiza and genuineness of herb. Zhongcaoyào 2002, 33, 1113–1116. [Google Scholar]
- Wang, Q.; Zhang, B.; Lu, Q. Conserved region amplification polymorphism (CoRAP), a novel marker technique for plant genotyping in Salvia miltiorrhiza. Plant Mol. Biol. Rep. 2009, 27, 139–143. [Google Scholar] [CrossRef]
- Ge, X.J.; Sun, M. Reproductive biology and genetic diversity of a cryptoviviparous mangrove Aegiceras corniculatum (Myrsinaceae) using allozyme and intersimple sequence repeat (ISSR) analysis. Mol. Ecol. 1999, 8, 2061–2069. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Zhang, Y.; Chen, C.B.; Li, X.L.; Chen, R.Y.; Chen, L. Analysis on genetic diversity of different Salvia miltiorrhiza geographical populations in China. Zhongguo Zhong Yao Za Zhi 2007, 32, 1988–1991. [Google Scholar] [PubMed]
- Wolfe, A.D.; Liston, A. Contributions of PCR-based methods of plant systematics and evolutionary biology. In Molecular Systematics of Plants II: DNA Sequencing; Soltis, P.S., Soltis, D.E., Doyle, J.J., Eds.; Springer Science + Business Media: New York, NY, USA, 1998; pp. 43–86. [Google Scholar]
- Camacho, F.J.; Liston, A. Population structure and genetic diversity of Botrychium pumicola (Ophioglossaceae) based on Inter-Simple Sequence Repeats (ISSR). Am. J. Bot. 2001, 88, 1065–1070. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Chen, X.Y.; Wang, X.R.; Pian, R.Q. ISSR analysis of genetic diversity among Lespedeza bicolor populations. J. Plant Genet. Resour. 2007, 8, 195–199. [Google Scholar]
- Bijlsma, R.; Ouborg, N.J.; Treuren, R. On genetic erosion and population extinction in plants: A case study in Scabiosa columbaria and Salvia pratensis. Conserv. Genet. 1994, 68, 255–271. [Google Scholar]
- Li, H.; Chen, G. Genetic diversity of mangrove plant Sonneratia caseolaris in Hainan Island based on ISSR analysis. Acta Ecol. Sin. 2004, 24, 1656–1662. [Google Scholar]
- Wang, J. Application of the one migrant per generation rule of conservation and management. Conserv. Biol. 2004, 18, 332–343. [Google Scholar] [CrossRef]
- Tivang, J.G.; Nienhuis, J.; Smith, O.S. Estimation of sampling variance of molecular marker data using the bootstrap procedure. Theor. Appl. Genet. 1994, 89, 259–264. [Google Scholar] [PubMed]
- Manly, B.F.J. Randomization, Bootstrap and Monte Carlo Methods in Biology, 2nd ed.; Chapman & Hall/CRC: London, UK, 1997. [Google Scholar]
- Cruz, C.D. Programa Genes—Biometria; Editora UFV: Viçosa, Brazil, 2006. [Google Scholar]
- Nei, M.; Li, W.H. Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc. Natl. Acad. Sci. USA 1979, 76, 5269–5273. [Google Scholar] [CrossRef] [PubMed]
- Kruskal, J.B. Nonmetric multidimensional scaling: A numerical method. Psychometrika 1964, 29, 115–129. [Google Scholar] [CrossRef]
- Yeh, F.C.; Yang, R.; Boyle, T. PopGene version 3.2: Microsoft Window-Based Freeware for Population Genetic Analysis; University of Alberta: Edmonton, AB, Canada, 1999. [Google Scholar]
- McDermott, J.M.; McDonald, B.A. Gene flow in plant pathosystems. Annu. Rev. Phytopathol. 1993, 31, 353–373. [Google Scholar] [CrossRef]
- Peakall, R.; Smouse, P.E. GENEALEX 6: Genetic analysis in Excel. Population genetic software for teaching and research. Mol. Ecol. Notes 2006, 6, 288–295. [Google Scholar]
- Excoffier, L.; Laval, G.; Schneider, S. Arlequin (version 3.0): An integrated software package for population genetics data analysis. Evol. Bioinform. Online 2005, 1, 47–50. [Google Scholar]
- Rohlf, F.J. NTSYS-pc: Numerical Taxonomy and Multivariate Analysis System; Version 2.1; Exeter Sotfware: New York, NY, USA, 2000. [Google Scholar]
- Swofford, D.L. PAUP* Phylogenetic Analysis Using Parsimony (*and Others Methods); Version 4b10; Sinauer Associates: Sunderland, MA, USA, 2003. [Google Scholar]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar] [PubMed]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Erbano, M.; Schühli, G.S.e.; Santos, É.P.d. Genetic Variability and Population Structure of Salvia lachnostachys: Implications for Breeding and Conservation Programs. Int. J. Mol. Sci. 2015, 16, 7839-7850. https://doi.org/10.3390/ijms16047839
Erbano M, Schühli GSe, Santos ÉPd. Genetic Variability and Population Structure of Salvia lachnostachys: Implications for Breeding and Conservation Programs. International Journal of Molecular Sciences. 2015; 16(4):7839-7850. https://doi.org/10.3390/ijms16047839
Chicago/Turabian StyleErbano, Marianna, Guilherme Schnell e Schühli, and Élide Pereira dos Santos. 2015. "Genetic Variability and Population Structure of Salvia lachnostachys: Implications for Breeding and Conservation Programs" International Journal of Molecular Sciences 16, no. 4: 7839-7850. https://doi.org/10.3390/ijms16047839
APA StyleErbano, M., Schühli, G. S. e., & Santos, É. P. d. (2015). Genetic Variability and Population Structure of Salvia lachnostachys: Implications for Breeding and Conservation Programs. International Journal of Molecular Sciences, 16(4), 7839-7850. https://doi.org/10.3390/ijms16047839






