High-Resolution Chromosome Ideogram Representation of Currently Recognized Genes for Autism Spectrum Disorders
Abstract
:1. Introduction
2. Results and Discussion
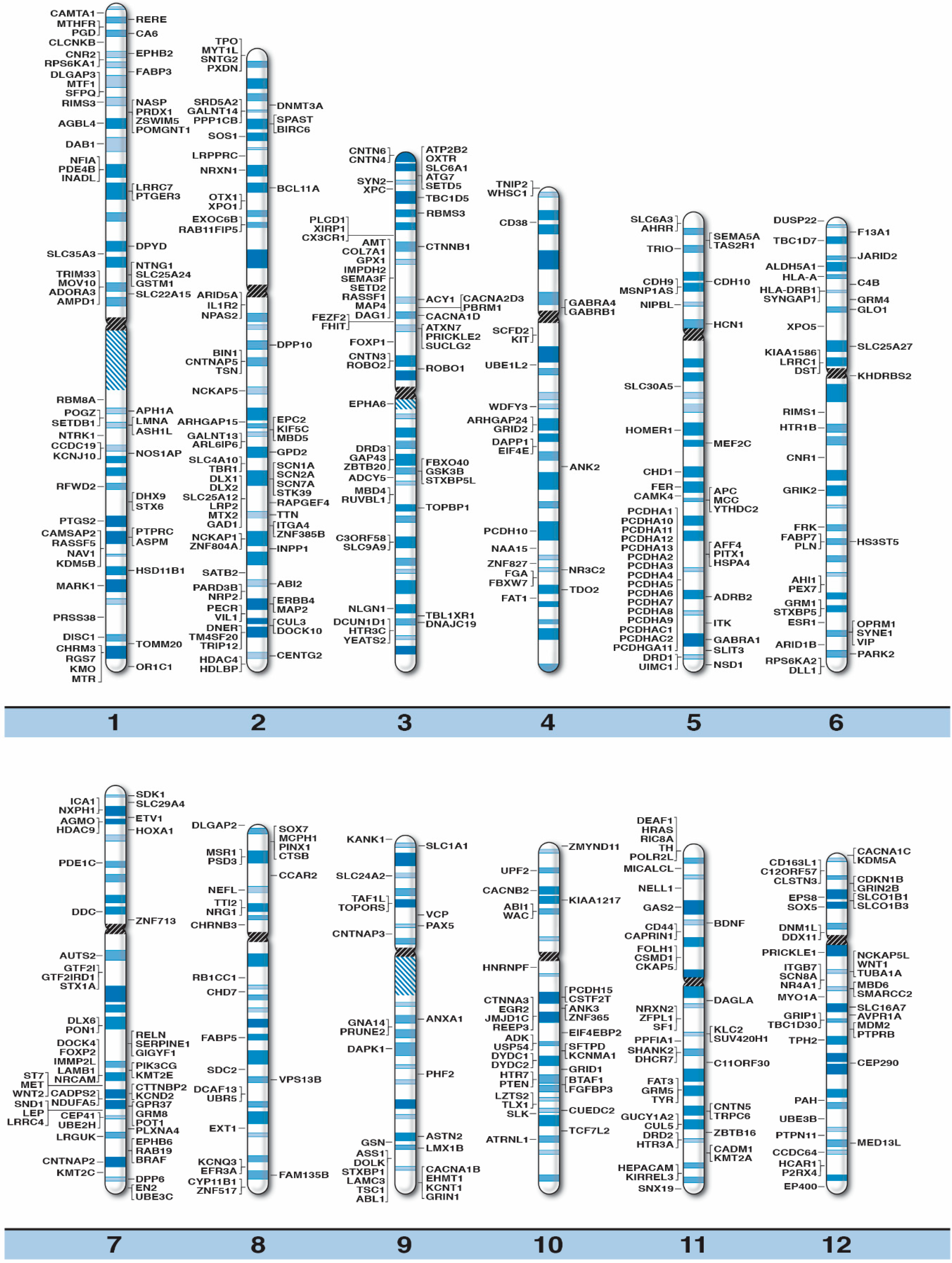
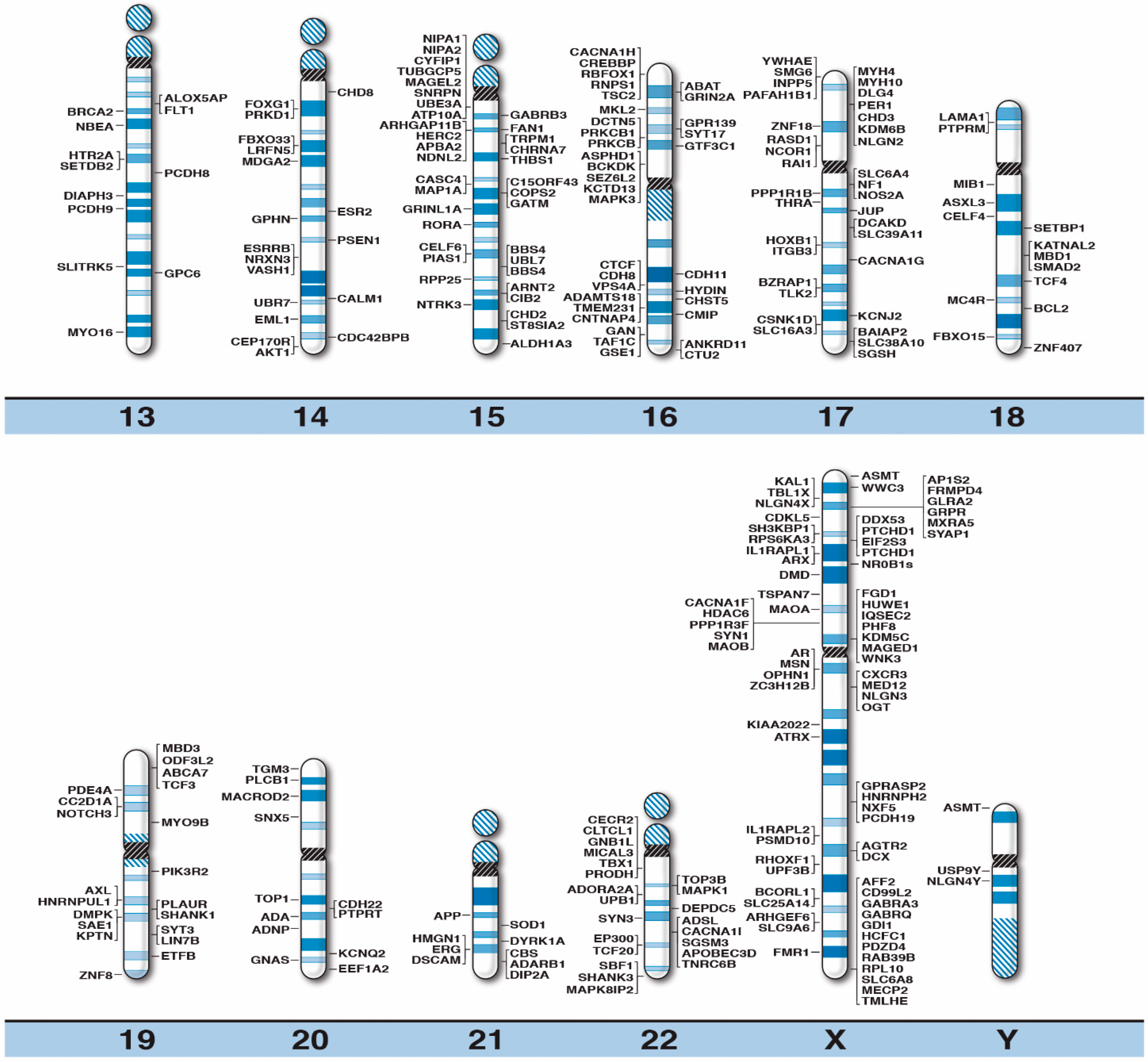
| Gene Symbol | Gene Name | Location |
|---|---|---|
| ABAT | 4-aminobutyrate aminotransferase | 16p13.2 |
| ABCA7 | ATP-binding cassette, sub-family A (ABC1), member 7 | 19p13.3 |
| ABI1 | Abl-interactor 1 | 10p12.1 |
| ABI2 | Abl-interactor 2 | 2q33.2 |
| ABL1 | C-Abl oncogene 1, non-receptor tyrosine kinase | 9q34.12 |
| ACY1 | Aminoacylase 1 | 3p21.2 |
| ADA | Adenosine deaminase | 20q13.12 |
| ADAMTS18 | A disintegrin-like and metalloproteinase with thrombospondin type 1 motif, 18 | 16q23.1 |
| ADARB1 | Adenosine deaminase, RNA-specific, B1 | 21q22.3 |
| ADCY5 | Adenylate cyclase 5 | 3q21.1 |
| ADK | Adenosine kinase | 10q22.2 |
| ADNP | Activity-dependent neuroprotector homeobox | 20q13.13 |
| ADORA2A | Adenosine A2A receptor | 22q11.23 |
| ADORA3 | Adenosine A3 receptor | 1p13.2 |
| ADRB2 | Adrenergic, β 2 receptor | 5q32 |
| ADSL | Adenylosuccinate lyase | 22q13.1 |
| AFF2 | AF4/fragile X mental retardation 2 (FMR2) family, member 2 | Xq28 |
| AFF4 | AF4/fragile X mental retardation 2 (FMR2) family, member 4 | 5q31.1 |
| AGBL4 | ATP/GTP binding protein-like 4 | 1p33 |
| AGMO | Alkylglycerol monooxygenase | 7p21.1 |
| AGTR2 | Angiotensin II receptor, type 2 | Xq23 |
| AHI1 | Abelson helper integration site 1 | 6q23.3 |
| AHRR | Aryl hydrocarbon receptor repressor | 5p15.33 |
| AKT1 | v-Akt murine thymoma viral oncogene homolog 1 | 14q32.33 |
| ALDH1A3 | Aldehyde dehydrogenase 1 family, member A3 | 15q26.3 |
| ALDH5A1 | Aldehyde dehydrogenase 5 family, member A1 | 6p22.3 |
| ALOX5AP | Arachidonate 5-lipoxygenase-activating protein | 13q12.3 |
| AMPD1 | Adenosine monophosphate deaminase 1 | 1p13.2 |
| AMT | Aminomethyltransferase | 3p21.31 |
| ANK2 | Ankyrin 2 | 4q25 |
| ANK3 | Ankyrin 3 | 10q21.2 |
| ANKRD11 | Ankyrin repeat domain 11 | 16q24.3 |
| ANXA1 | Annexin A1 | 9q21.13 |
| AP1S2 | Adaptor-related protein complex 1, sigma 2 subunit | Xp22.2 |
| APBA2 | Amyloid β precursor protein-binding, family A, member 2 | 15q13.1 |
| APC | Adenomatosis polyposis coli | 5q22.2 |
| APH1A | APH1A γ secretase subunit | 1q21.2 |
| APOBEC3D | Apolipoprotein B mRNA editing enzyme, catalytic polypeptide-like 3D | 22q13.1 |
| APP | Amyloid β precursor protein | 21q21.3 |
| AR | Androgen receptor | Xq12 |
| ARHGAP11B | Rho GTPase activating protein 11B | 15q13.2 |
| ARHGAP15 | Rho GTPase activating protein 15 | 2q22.2 |
| ARHGAP24 | Rho GTPase activating protein 24 | 4q22.1 |
| ARHGEF6 | RAC/CDC42 guanine nucleotide exchange factor (GEF) 6 | Xq26.3 |
| ARID1B | AT rich interactive domain 1B (SWI1-like) | 6q25.3 |
| ARID5A | AT rich interactive domain 5A (MRF1-like) | 2q11.2 |
| ARL6IP6 | ADP-ribosylation-like factor 6 interacting protein 6 | 2q23.3 |
| ARNT2 | Aryl-hydrocarbon receptor nuclear translocator 2 | 15q25.1 |
| ARX | Aristaless related homeobox | Xp21.3 |
| ASH1L | Ash1 (absent, small, or homeotic)-like (Drosophila) | 1q22 |
| ASMT | Acetylserotonin O-methyltransferase, X-chromosomal | Xp22.33 |
| ASMT | Acetylserotonin O-methyltransferase, Y-chromosomal | Yp11.32 |
| ASPHD1 | Aspartate β-hydroxylase domain containing 1 | 16p11.2 |
| ASPM | Asp (abnormal spindle) homolog, microcephaly associated | 1q31.3 |
| ASS1 | Argininosuccinate synthetase | 9q34.1 |
| ASTN2 | Astrotactin 2 | 9q33.1 |
| ASXL3 | Additional sex combs-like 3 | 18q12.1 |
| ATG7 | Autophagy related 7 | 3p25.3 |
| ATP10A | ATPase, Class V, type 10A | 15q11.2 |
| ATP2B2 | ATPase, Ca++ transporting, plasma membrane 2 | 3p25.3 |
| ATRNL1 | Attractin-like 1 | 10q25.3 |
| ATRX | α thalassemia/mental retardation syndrome X-linked | Xq21.1 |
| ATXN7 | Ataxin 7 | 3p14.1 |
| AUTS2 | Autism susceptibility candidate 2 | 7q11.22 |
| AVPR1A | Arginine vasopressin receptor 1A | 12q14.2 |
| AXL | AXL receptor tyrosine kinase | 19q13.2 |
| BAIAP2 | BAI1-associated protein 2 | 17q25.3 |
| BBS4 | Bardet-Biedl syndrome 4 | 15q24.1 |
| BCKDK | Branched chain ketoacid dehydrogenase kinase | 16p11.2 |
| BCL11A | B-Cell CLL/lymphoma 11A (zinc finger protein) | 2p16.1 |
| BCL2 | B-cell CLL/lymphoma 2 | 18q21.33 |
| BCORL1 | Bc16 co-repressor-like 1 | Xq26.1 |
| BDNF | Brain-derived neurotrophic factor | 11p14.1 |
| BIN1 | Bridging integrator 1 | 2q14.3 |
| BIRC6 | Baculoviral IAP repeat containing 6 | 2p22.3 |
| BRAF | v-Raf murine sarcoma viral oncogene homolog B | 7q34 |
| BRCA2 | Breast cancer 2, early onset | 13q13.1 |
| BTAF1 | RNA polymerase II, B-TFIID transcription factor-associated, 170 kDa (Mot1 homolog, S. cerevisiae) | 10q23.32 |
| BZRAP1 | Benzodiazepine receptor (peripheral) associated protein 1 | 17q23.2 |
| C11ORF30 | Chromosome 11 open reading frame 30 | 11q13.5 |
| C12ORF57 | Chromosome 12 open reading frame 57 | 12p13.31 |
| C15ORF43 | Chromosome 15 open reading frame 43 | 15q21.1 |
| C3ORF58 | Chromosome 3 open reading frame 58 | 3q24 |
| C4B | Complement component 4B | 6p21.33 |
| CA6 | Carbonic anhydrase VI | 1p36.2 |
| CACNA1B | Calcium channel, voltage-dependent, N type, α 1B subunit | 9q34.3 |
| CACNA1C | Calcium channel, voltage-dependent, L type, α 1C subunit | 12p13.33 |
| CACNA1D | Calcium channel, voltage-dependent, L type, α 1D subunit | 3p14.3 |
| CACNA1F | Calcium channel, voltage-dependent, α 1F subunit | Xp11.23 |
| CACNA1G | Calcium channel, voltage-dependent, T type, α 1G subunit | 17q21.33 |
| CACNA1H | Calcium channel, voltage-dependent, α 1H subunit | 16p13.3 |
| CACNA1I | Calcium channel, voltage-dependent, T type, α 1I subunit | 22q13.1 |
| CACNA2D3 | Calcium channel, voltage-dependent, α 2/δ subunit 3 | 3p21.1 |
| CACNB2 | Calcium channel, voltage-dependent, β 2 subunit | 10p12.33 |
| CADM1 | Cell adhesion molecule 1 | 11q23.3 |
| CADPS2 | Ca++-dependent activator protein for secretion 2 | 7q31.32 |
| CALM1 | Calmodulin 1 (phosphorylase kinase, δ) | 14q32.11 |
| CAMK4 | Calcium/calmodulin-dependent protein kinase | 5q22.1 |
| CAMSAP2 | Calmodulin regulated spectrin-associated protein family, member 2 | 1q32.1 |
| CAMTA1 | Calmodulin binding transcription activator 1 | 1p36.31 |
| CAPRIN1 | Cell cycle associated protein 1 | 11p13 |
| CASC4 | Cancer susceptibility candidate 4 | 15q15.3 |
| CBS | Cystathionine β-synthase | 21q22.3 |
| CCAR2 | Cell cycle and apoptosis regulator 2 | 8p21.3 |
| CC2D1A | Coiled-coil and C2 domain-containing 1A | 19p13.12 |
| CCDC19 | Coiled-coil domain-containing protein 19 | 1q23.2 |
| CCDC64 | Coiled-coil domain-containing 64 | 12q24.23 |
| CD38 | CD38 molecule | 4p15.32 |
| CD44 | CD44 molecule | 11p13 |
| CD163L1 | CD163 molecule-like 1 | 12p13.31 |
| CD99L2 | CD99 molecule-like 2 | Xq28 |
| CDC42BPB | CDC42 binding protein kinase β (DMPK-like) | 14q32.32 |
| CDH10 | Cadherin 10, type 2 | 5p14.2 |
| CDH22 | Cadherin-like 22 | 20q13.1 |
| CDH8 | Cadherin 8, type 2 | 16q22.1 |
| CDH9 | Cadherin 9, type 2 | 5p14.1 |
| CDH11 | Cadherin 11, type 2 | 16q21 |
| CDKL5 | Cyclin-dependent kinase-like 5 | Xp22.13 |
| CDKN1B | Cyclin-dependent kinase inhibitor 1B | 12p13.1 |
| CECR2 | Cat eye syndrome chromosome region, candidate 2 | 22q11.21 |
| CELF4 | CUGBP, Elav-like family, member 4 | 18q12.2 |
| CELF6 | CUGBP, Elav-like family, member 6 | 15q23 |
| CENTG2 | Centaurin γ-2 | 2q37.2 |
| CEP170R | Centrosomal protein 170B | 14q32.33 |
| CEP290 | Centrosomal protein 290 kDa | 12q21.32 |
| CEP41 | Centrosomal protein 41 kDa | 7q32.2 |
| CHD1 | Chromodomain helicase DNA binding protein 1 | 5q21.1 |
| CHD2 | Chromodomain helicase DNA binding protein 2 | 15q26.1 |
| CHD3 | Chromodomain helicase DNA binding protein 3 | 17p13.1 |
| CHD7 | Chromodomain helicase DNA binding protein 7 | 8q12.2 |
| CHD8 | Chromodomain helicase DNA binding protein 8 | 14q11.2 |
| CHRM3 | Cholinergic receptor, muscarinic 3 | 1q43 |
| CHRNA7 | Cholinergic receptor, neuronal nicotinic, α 7 | 15q13.3 |
| CHRNB3 | Cholinergic receptor, neuronal nicotinic, β 3 | 8p11.21 |
| CHST5 | Carbohydrate sulfotransferase 5 | 16q22.3 |
| CIB2 | Calcium and integrin binding family member 2 | 15q25.1 |
| CKAP5 | Cytoskeleton associated protein 5 | 11p11.2 |
| CLCNKB | Chloride channel voltage-sensitive kidney, B | 1p36.13 |
| CLSTN3 | Calsyntenin 3 | 12p13.31 |
| CLTCL1 | Clathrin, heavy chain-like 1 | 22q11.21 |
| CMIP | c-MAF inducing protein | 16q23.2 |
| CNR1 | Cannabinoid receptor 1 | 6q15 |
| CNR2 | Cannabinoid receptor 2 | 1p36.11 |
| CNTN3 | Contactin 3 | 3p12.3 |
| CNTN4 | Contactin 4 | 3p26.3 |
| CNTN5 | Contactin 5 | 11q22.1 |
| CNTN6 | Contactin 6 | 3p26.3 |
| CNTNAP2 | Contactin associated protein-like 2 | 7q35 |
| CNTNAP3 | Contactin associated protein-like 3 | 9p13.1 |
| CNTNAP4 | Contactin associated protein-like 4 | 16q23.1 |
| CNTNAP5 | Contactin associated protein-like 5 | 2q14.3 |
| COL7A1 | Collagen, type VII, α 1 | 3p21.31 |
| COPS2 | Thyroid hormone receptor interactor 15 | 15q21.1 |
| CREBBP | CREB binding protein | 16p13.3 |
| CSMD1 | Cytoskeleton associated protein 5 | 11p11.2 |
| CSNK1D | Casein kinase 1, δ | 17q25 |
| CSTF2T | Cleavage stimulation factor, 3' pre-RNA, subunit 2, 64 kDa, tau | 10q21.1 |
| CTCF | CCCTC-binding factor | 16q22.1 |
| CTNNA3 | Catenin (cadherin-associated protein), α 3 | 10q21.3 |
| CTNNB1 | Catenin (cadherin-associated protein), β 1, 88 kDa | 3p22.1 |
| CTSB | Cathepsin B | 8p23.1 |
| CTTNBP2 | Cortactin binding protein 2 | 7q31.31 |
| CTU2 | Cytosolic thiouridylase subunit 2 homolog (S. pombe) | 16q24.3 |
| CUEDC2 | CUE domain containing 2 | 10q24.32 |
| CUL5 | Cullin 5 | 11q22.3 |
| CUL3 | Cullin 3 | 2q36.2 |
| CX3CR1 | Chemokine (C-X3-C motif) receptor 1 | 3p22.2 |
| CXCR3 | Chemokine, CXC motif, receptor 3 | Xq13.1 |
| CYFIP1 | Cytoplasmic FMRP interacting protein 1 | 15q11.2 |
| CYP11B1 | Cytochrome P450, subfamily XIB, polypeptide 1 | 8q24.3 |
| DAB1 | Disabled homolog 1 | 1p32.2 |
| DAG1 | Dystroglycan 1 (dystrophin-associated glycoprotein 1) | 3p21.31 |
| DAGLA | Diacylglycerol lipase, α | 11q12.2 |
| DAPK1 | Death-associated protein kinase 1 | 9q21.33 |
| DAPP1 | Dual adaptor of phosphotyrosine and 3-phosphoinositides 1 | 4q23 |
| DCAF13 | DDB1 and CUL4 associated factor 13 | 8q22.3 |
| DCAKD | Dephospho-CoA kinase domain-containing protein | 17q21.31 |
| DCTN5 | Dynactin 5 | 16p12.2 |
| DCUN1D1 | DCN1, domain containing protein 1 | 3q27.1 |
| DCX | Doublecortin | Xq23 |
| DDC | DOPA decarboxylase | 7p12.1 |
| DDX11 | DEAD (Asp-Glu-Ala-Asp)/H box 11 | 12p11.21 |
| DDX53 | DEAD (Asp-Glu-Ala-Asp) box polypeptide 53 | Xp22.11 |
| DEAF1 | DEAF1 transcription factor | 11p15.5 |
| DEPDC5 | DEP domain containing 3 protein 5 | 22q12.2 |
| DHCR7 | 7-dehydrocholesterol reductase | 11q13.4 |
| DHX9 | DEAH (Asp-Glu-Ala-His) box helicase 9 | 1q25.3 |
| DIAPH3 | Diaphanous, Drosophila, homolog 3 | 13q21.2 |
| DIP2A | DIP2 disco-interacting protein 2 homolog A (Drosophila) | 21q22.3 |
| DISC1 | Disrupted in schizophrenia 1 | 1q42.2 |
| DLG4 | Discs, large, Drosophila, homolog 4 | 17p13.1 |
| DLGAP2 | Discs, large- associated protein 2 | 8p23.3 |
| DLGAP3 | Discs, large- associated protein 3 | 1p34.3 |
| DLL1 | δ-like 1 (Drosophila) | 6q27 |
| DLX1 | Distal-less homeobox 1 | 2q31.1 |
| DLX2 | Distal-less homeobox 2 | 2q31.1 |
| DLX6 | Distal-less homeobox 6 | 7q21.3 |
| DMD | Dystrophin | Xp21.1 |
| DMPK | Dystrophia myotonica-protein kinase | 19q13.32 |
| DNAJC19 | DNAJ Hsp40 homolog, subfamily C, member 19 | 3q26.33 |
| DNER | δ- and notch-like epidermal growth factor-related receptor | 2q36.3 |
| DNM1L | Dynamin 1-like | 12p11.21 |
| DNMT3A | DNA (cytosine-5)-methyltransferase 3 α | 2p23.3 |
| DOCK4 | Dedicator of cytokinesis 4 | 7q31.1 |
| DOCK10 | Dedicator of cytokinesis 10 | 2q36.2 |
| DOLK | Dolichol kinase | 9q34.1 |
| DPP10 | Dipeptidyl peptidase 10 | 2q14.1 |
| DPP6 | Dipeptidyl peptidase 6 | 7q36.2 |
| DPYD | Dihydropyrimidine dehydrogenase | 1p21.3 |
| DRD1 | Dopamine receptor D1 | 5q35.2 |
| DRD2 | Dopamine receptor D2 | 11q23.2 |
| DRD3 | Dopamine receptor D3 | 3q13.31 |
| DSCAM | Down syndrome cell adhesion molecule | 21q22.2 |
| DST | Dystonin | 6p12.1 |
| DUSP22 | Dual specificity phosphatase 22 | 6p25.3 |
| DYDC1 | DPY30 domain containing 1 | 10q23.1 |
| DYDC2 | DPY30 domain containing 2 | 10q23.1 |
| DYRK1A | Dual-specificity tyrosine-phosphorylation-regulated kinase 1A | 21q22.13 |
| EEF1A2 | Eukaryotic translation elongation factor 1 α 2 | 20q13.33 |
| EFR3A | EFR3 homolog A (S. cerevisiae) | 8q24.22 |
| EGR2 | Early growth response 2 | 10q21.3 |
| EHMT1 | Euchromatic histone methyltransferase 1 | 9q34.3 |
| EIF2S3 | Eukaryotic translation initiation factor 2, subunit 3 γ | Xp22.11 |
| EIF4E | Eukaryotic translation initiation factor 4E | 4q23 |
| EIF4EBP2 | Eukaryotic translation initiation factor 4E binding protein 2 | 10q22.1 |
| EML1 | Echinoderm microtubule associated protein like 1 | 14q32.2 |
| EN2 | Engrailed 2 | 7q36.3 |
| EP300 | E1A binding protein p300 | 22q13.2 |
| EP400 | E1A binding protein p400 | 12q24.33 |
| EPC2 | Enhancer of polycomb, Drosophila homolog of 2 | 2q23.1 |
| EPHA6 | Ephrin receptor A6 | 3q11.2 |
| EPHB2 | Ephrin receptor B2 | 1p36.12 |
| EPHB6 | Ephrin receptor B6 | 7q34 |
| EPS8 | Epidermal growth factor receptor pathway substrate 8 | 12p12.3 |
| ERBB4 | v-ERB-A avian erythroblastic leukemia viral oncogene homolog 4 | 2q34 |
| ERG | v-ETS avian erythroblastosis virus E26 oncogene homolog | 21q22.2 |
| ESR1 | Estrogen receptor 1 | 6q25.1 |
| ESR2 | Estrogen receptor 2 | 14q23.2 |
| ESRRB | Estrogen-related receptor β | 14q24.3 |
| ETFB | Electron-transfer-flavoprotein, β polypeptide | 19q13.41 |
| ETV1 | Ets variant 1 | 7p21.2 |
| EXOC6B | Exocyst complex component 6B | 2p13.2 |
| EXT1 | Exostosin 1 | 8q24.11 |
| F13A1 | Factor XIII, A1 subunit | 6p25.1 |
| FABP3 | Fatty acid binding protein 3, muscle and heart (mammary-derived growth inhibitor) | 1p35.2 |
| FABP5 | Fatty acid binding protein 5 | 8q21.13 |
| FABP7 | Fatty acid binding protein 7 | 6q22.31 |
| FAM135B | Family with sequence similarity 135, member B | 8q24.23 |
| FAN1 | FANCD2/FANCI-associated nuclease 1 | 15q13.2 |
| FAT1 | FAT tumor suppressor, Drosophila homolog of, 1 | 4q35.2 |
| FAT3 | FAT tumor suppressor, Drosophila homolog of , 3 | 11q14.3 |
| FBXO15 | F-box protein 15 | 18q22.3 |
| FBXO33 | F-box protein 33 | 14q21.1 |
| FBXO40 | F-box protein 40 | 3q13.33 |
| FBXW7 | F-box and WD repeat domain containing 7, E3 ubiquitin protein | 4q31.3 |
| FER | FPS/FES related tyrosine kinase | 5q21.3 |
| FEZF2 | FEZ family zinc finger 2 | 3p14.2 |
| FGA | Fibrinogen, A α polypeptide | 4q31.3 |
| FGD1 | FYVE, Rho GEF and PH domain containing 1 | Xp11.22 |
| FGFBP3 | Fibroblast growth factor binding protein 3 | 10q23.32 |
| FHIT | Fragile histidine triad | 3p14.2 |
| FLT1 | c-FMS-related tyrosine kinase 1 | 13q12.3 |
| FMR1 | Fragile X mental retardation 1 (FMR1) | Xq27.3 |
| FOLH1 | Folate hydrolase 1 | 11p11.2 |
| FOXG1 | Forkhead box G1 | 14q12 |
| FOXP1 | Forkhead box P1 | 3p13 |
| FOXP2 | Forkhead box P2 | 7q31.1 |
| FRK | FYN-related kinase | 6q22.1 |
| FRMPD4 | FERM and PDZ domain containing protein 4 | Xp22.2 |
| GABRA1 | γ-aminobutyric acid A receptor, α 1 | 5q34 |
| GABRA3 | γ-aminobutyric acid receptor, α 3 | Xq28 |
| GABRA4 | γ-aminobutyric acid receptor, α 4 | 4p12 |
| GABRB1 | γ-aminobutyric acid receptor, β 1 | 4p12 |
| GABRB3 | γ-aminobutyric acid receptor, β 3 | 15q12 |
| GABRQ | γ-aminobutyric acid receptor, θ | Xq28 |
| GAD1 | Glutamate decarboxylase 1 (brain, 67 kDa) | 2q31.1 |
| GALNT13 | UDP-N-acetyl-α-d-galactosamine:polypeptide N-acetylgalactosaminyl-transferase 13 | 2q23.3 |
| GALNT14 | UDP-N-acetyl-α-d-galactosamine:polypeptide N-acetylgalactosaminyl-transferase 14 | 2p23.1 |
| GAN | Gigaxonin | 16q24.1 |
| GAP43 | Growth associated protein 43 | 3q13.31 |
| GAS2 | Growth arrest-specific 2 | 11p14.3 |
| GATM | Glycine amidinotransferase (l-arginine:glycine amidinotransferase) | 15q21.1 |
| GDI1 | GDP dissociation inhibitor 1 | Xq28 |
| GIGYF1 | GRB10 interacting GYF protein 1 | 7q22.1 |
| GLO1 | Glyoxalase I | 6p21.2 |
| GLRA2 | Glycine receptor, α 2 subunit | Xp22.2 |
| GNA14 | Guanine nucleotide-binding protein, α 14 | 9q21.2 |
| GNAS | Guanine nucleotide-binding protein, α-stimulating activity polypeptide I complex locus | 20q13.32 |
| GNB1L | Guanine nucleotide-binding protein, β 1-like | 22q11.21 |
| GPC6 | Glypican 6 | 13q31.3 |
| GPD2 | Glycerol-3-phosphate dehydrogenase 2 | 2q24.1 |
| GPHN | Gephyrin | 14q23.3 |
| GPR139 | G protein-coupled receptor 139 | 16p12.3 |
| GPR37 | G protein-coupled receptor 37 | 7q31.33 |
| GPRASP2 | G protein-coupled receptor associated sorting protein 2 | Xq22.1 |
| GPX1 | Glutathione peroxidase 1 | 3p21.31 |
| GRID1 | Glutamate receptor, ionotropic, δ 1 | 10q23.2 |
| GRID2 | Glutamate receptor, ionotropic, δ 2 | 4q22.1 |
| GRIK2 | Glutamate receptor, ionotropic, kainate 2 | 6q16.3 |
| GRIN1 | Glutamate receptor, ionotropic, N-methyl d-aspartate 1 | 9q34.3 |
| GRIN2A | Glutamate receptor, ionotropic, N-methyl d-aspartate 2A | 16p13.2 |
| GRIN2B | Glutamate receptor, ionotropic, N-methyl d-aspartate 2B | 12p13.1 |
| GRINL1A | GRINL1A complex locus 1 | 15q21.3 |
| GRIP1 | Glutamate receptor interacting protein 1 | 12q14.3 |
| GRM1 | Glutamate receptor, metabotropic 1 | 6q24.3 |
| GRM4 | Glutamate receptor, metabotropic 4 | 6p21.31 |
| GRM5 | Glutamate receptor, metabotropic 5 | 11q14.3 |
| GRM8 | Glutamate receptor, metabotropic 8 | 7q31.33 |
| GRPR | Gastrin-releasing peptide receptor | Xp22.2 |
| GSE1 | Gse1 coiled-coil protein | 16q24.1 |
| GSK3B | Glycogen synthase kinase 3 β | 3q13.33 |
| GSN | Gelsolin | 9q33.2 |
| GSTM1 | Glutathione S-transferase M1 | 1p13.3 |
| GTF2I | General transcription factor III | 7q11.23 |
| GTF2IRD1 | GTF2I repeat domain containing 1 | 7q11.23 |
| GTF3C1 | General transcription factor IIIC, polypeptide 1, α | 16p12.1 |
| GUCY1A2 | Guanylate cyclase 1, soluble, α 2 | 11q22.3 |
| HCAR1 | Hydroxycarboxylic acid receptor 1/G protein-coupled receptor 81 | 12q24.31 |
| HCFC1 | Host cell factor C1 | Xq28 |
| HCN1 | Hyperpolarization activated cyclic nucleotide-gated potassium channel 1 | 5p12 |
| HDAC4 | Histone deacetylase 4 | 2q37.3 |
| HDAC6 | Histone deacetylase 6 | Xp11.23 |
| HDAC9 | Histone deacetylase 9 | 7p21.1 |
| HDLBP | High density lipoprotein binding protein | 2q37.3 |
| HEPACAM | Hepatic and glial cell adhesion molecule | 11q24.2 |
| HERC2 | HECT domain and RCC1-like domain 2 | 15q13.1 |
| HLA-A | Major histocompatibility complex, class I, A | 6p22.1 |
| HLA-DRB1 | Major histocompatibility complex, class II, DR β 1 | 6p21.32 |
| HMGN1 | High mobility group nucleosome binding domain 1 | 21q22.2 |
| HNRNPF | Heterogeneous nuclear ribonucleoprotein F | 10q11.21 |
| HNRNPH2 | Heterogeneous nuclear ribonucleoprotein H2 | Xq22.1 |
| HNRNPUL1 | Heterogeneous nuclear ribonucleoprotein U-like 1 | 19q13.2 |
| HOMER1 | Homer, Drosophila, homolog 1 of 1 | 5q14.1 |
| HOXA1 | Homeobox A1 | 7p15.3 |
| HOXB1 | Homeobox B1 | 17q21.32 |
| HRAS | v-HA-RAS Harvey rat sarcoma viral oncogene homolog | 11p15.5 |
| HS3ST5 | Heparan sulfate 3-O-sulfotransferase 5 | 6q22.31 |
| HSD11B1 | 11-β-hydroxysteroid dehydrogenase type 1 | 1q32.2 |
| HSPA4 | Heat shock 70 kDa protein 4 | 5q31.1 |
| HTR1B | 5-hydroxytryptamine receptor 1B | 6q14.1 |
| HTR2A | 5-hydroxytryptamine receptor 2A | 13q14.2 |
| HTR3A | 5-hydroxytryptamine receptor 3A | 11q23.2 |
| HTR3C | 5-hydroxytryptamine receptor 3, family member C | 3q27.1 |
| HTR7 | 5-hydroxytryptamine receptor 7 | 10q23.31 |
| HUWE1 | HECT, UBA and WWE domain containing 1, E3 ubiquitin protein ligase | Xp11.22 |
| HYDIN | Hydrocephalus-inducing, mouse, homolog of | 16q22.2 |
| ICA1 | Islet cell autoantigen 1 | 7p21.3 |
| IL1R2 | Interleukin 1 receptor, type II | 2q11.2 |
| IL1RAPL1 | Interleukin 1 receptor accessory protein-like 1 | Xp21.3 |
| IL1RAPL2 | Interleukin 1 receptor accessory protein-like 2 | Xq22.3 |
| IMMP2L | Inner mitochondrial membrane peptidase, subunit 2, S. cerevisiae, homolog of | 7q31.1 |
| IMPDH2 | Inosine-5-prime monophosphate dehydrogenase 2 | 3p21.31 |
| INADL | Inactivation no after-potential D-like | 1p31.3 |
| INPP1 | Inositol polyphosphate-1-phosphatase | 2q32.2 |
| INPP5 | Inositol polyphosphate-5-phosphatase | 17p13.3 |
| IQSEC2 | IQ motif and Sec7 domain 2 | Xp11.22 |
| ITGA4 | Integrin, α 4 | 2q31.3 |
| ITGB3 | Integrin, β 3 | 17q21.32 |
| ITGB7 | Integrin, β 7 | 12q13.13 |
| ITK | IL20 inducible t-cell kinase | 5q33.3 |
| JARID2 | Jumonji, AT rich interactive domain 2 | 6p22.3 |
| JMJD1C | Jumonji domain containing 1C | 10q21.3 |
| JUP | Junction plakoglobin | 17q21.2 |
| KAL1 | Kallmann syndrome interval 1 | Xp22.31 |
| KANK1 | KN motif and ankyrin repeat domains 1 | 9p24.3 |
| KATNAL2 | Katanin p60 subunit A-like 2 | 18q21.1 |
| KCND2 | Potassium voltage-gated channel, Shal-related subfamily, member 2 | 7q31.31 |
| KCNJ2 | Potassium inwardly-rectifying channel, subfamily J, member 2 | 17q24.3 |
| KCNJ10 | Potassium inwardly-rectifying channel, subfamily J, member 10 | 1q23.2 |
| KCNMA1 | Potassium large conductance calcium-activated channel, subfamily M, α member 1 | 10q22.3 |
| KCNQ2 | Potassium voltage-gated channel, KQT-like subfamily, member 2 | 20q13.3 |
| KCNQ3 | Potassium voltage-gated channel, KQT-like subfamily, member 3 | 8q24.22 |
| KCNT1 | Potassium channel, subfamily T, member 1 | 9q34.3 |
| KCTD13 | Potassium channel tetramerization domain containing protein 13 | 16p11.2 |
| KDM5A | Lysine (K)-specific demethylase 5A | 12p13.33 |
| KDM5B | Lysine (K)-specific demethylase 5B | 1q32.1 |
| KDM5C | Lysine (K)-specific demethylase 5C | Xp11.22 |
| KDM6B | Lysine (K)-specific demethylase 6B | 17p13.1 |
| KHDRBS2 | KH domain containing, RNA binding, signal transduction associated protein 2 | 6q11.1 |
| KIAA1217 | Sickle tail protein homolog | 10p12.31 |
| KIAA1586 | KIAA1586 | 6p12.1 |
| KIAA2022 | KIAA2022 | Xq13.3 |
| KIF5C | Kinesin family member 5C | 2q23.1 |
| KIRREL3 | Kin of IRRE like 3 | 11q24.2 |
| KIT | v-KIT Hardy-Zuckerman 4 feline sarcoma viral oncogene homolog | 4q12 |
| KLC2 | Kinesin light chain 2 | 11q13.2 |
| KMO | Kynurenine 3-monooxygenase | 1q43 |
| KMT2A | Lysine (K)-specific methyltransferase 2A | 11q23.3 |
| KMT2C | Lysine (K)-specific methyltransferase 2C | 7q36.1 |
| KMT2E | Lysine (K)-specific methyltransferase 2E | 7q22.3 |
| KPTN | Kaptin (actin binding protein) | 19q13.32 |
| LAMA1 | Laminin, α 1 | 18p11.23 |
| LAMB1 | Laminin, β 1 | 7q31.1 |
| LAMC3 | Laminin, γ 3 | 9q34.1 |
| LEP | Leptin | 7q32.1 |
| LIN7B | Lin-7 homolog B (C. elegans) | 19q13.33 |
| LMNA | Lamin A/C | 1q22 |
| LMX1B | LIM homeobox transcription factor 1, β | 9q33.3 |
| LRFN5 | Leucine-rich repeats and fibronectin type III domain containing 5 | 14q21.1 |
| LRGUK | Leucine-rich repeats and guanylate kinase domain containing | 7q33 |
| LRP2 | Low density lipoprotein receptor-related protein 2 | 2q31.1 |
| LRPPRC | Leucine-rich PPR motif containing protein | 2p21 |
| LRRC1 | Leucine-rich repeat-containing protein 1 | 6p12.1 |
| LRRC4 | Leucine-rich repeat-containing protein 4 | 7q32.1 |
| LRRC7 | Leucine-rich repeat-containing protein 7 | 1p31.1 |
| LZTS2 | Leucine zipper, putative tumor suppressor 2 | 10q24.31 |
| MACROD2 | Macro domain containing 2 | 20p12.1 |
| MAGED1 | Melanoma antigen family D, 1 | Xp11.22 |
| MAGEL2 | MAGE-like 2 | 15q11.2 |
| MAOA | Monoamine oxidase A | Xp11.3 |
| MAOB | Monoamine oxidase B | Xp11.23 |
| MAP1A | Microtubule-associated protein 1A | 15q15.3 |
| MAP2 | Microtubule-associated protein (MAP) 2 | 2q34 |
| MAP4 | Microtubule-associated protein (MAP) 4 | 3p21.31 |
| MAPK1 | Mitogen-activated protein kinase 1 | 22q11.22 |
| MAPK3 | Mitogen-activated protein kinase 3 | 16p11.2 |
| MAPK8IP2 | Mitogen-activated protein kinase 8 interacting protein 2 | 22q13.33 |
| MARK1 | MAP/microtubule affinity-regulating kinase 1 | 1q41 |
| MBD1 | Methyl-CpG binding domain protein 1 | 18q21.1 |
| MBD3 | Methyl-CpG binding domain protein 3 | 19p13.3 |
| MBD4 | Methyl-CpG binding domain protein 4 | 3q21.3 |
| MBD5 | Methyl-CpG binding domain protein 5 | 2q23.1 |
| MBD6 | Methyl-CpG binding domain protein 6 | 12q13.2 |
| MC4R | Melanocortin 4 receptor | 18q21.32 |
| MCC | Mutated in colorectal cancers | 5q22.2 |
| MCPH1 | Microcephalin 1 | 8p23.1 |
| MDGA2 | Mephrin, A5 antigen, protein tyrosine phosphatase mu (MAM) domain containing glycosylphosphatidylinositol anchor 2 | 14q21.3 |
| MDM2 | MDM2 oncogene, E3 ubiquitin protein ligase | 12q15 |
| MECP2 | Methyl CpG binding protein 2 | Xq28 |
| MED12 | Mediator complex subunit 12 | Xq13.1 |
| MED13L | Mediator complex subunit 13-like | 12q24.21 |
| MEF2C | MADS box transcription myocyte enhancer factor 2, polypeptide C | 5q14.3 |
| MET | Met proto-oncogene | 7q31.2 |
| MIB1 | Mind bomb E3 ubiquitin protein ligase 1 | 18q11.2 |
| MICAL3 | Microtubule-associated monooxygenase, calponin and lim domains-containing, 3 | 22q11.21 |
| MICALCL | MICAL C-terminus-like protein | 11p15.3 |
| MKL2 | Myocardin-like 2 | 16p13.12 |
| MOV10 | Moloney leukemia virus 10, mouse, homolog of | 1p13.2 |
| MSN | Moesin | Xq12 |
| MSNP1AS | Moesin pseudogene 1 antisense | 5p14.1 |
| MSR1 | Macrophage scavenger receptor | 8p22 |
| MTF1 | Metal-regulatory transcription factor 1 | 1p34.3 |
| MTHFR | 5-10-methylene-tetrahydrofolate reductase | 1p36.22 |
| MTR | 5-methyltetrahydrofolate-homocysteine S-methyltransferase | 1q43 |
| MTX2 | Metaxin 2 | 2q31.1 |
| MXRA5 | Matrix-remodelling associated 5 | Xp22.2 |
| MYH4 | Myosin, heavy chain 4, skeletal muscle | 17p13.1 |
| MYH10 | Myosin, heavy chain 10, non-muscle | 17p13.1 |
| MYO16 | Myosin XVI | 13q33.3 |
| MYO1A | Myosin IA | 12q13.3 |
| MYO9B | Myosin IXB | 19p13.11 |
| MYT1L | Myelin transcription factor 1-like | 2p25.3 |
| NAA15 | N(α)-acetyltransferase 15, NatA auxiliary subunit | 4q31.1 |
| NASP | Nuclear autoantigenic sperm protein (histone-binding) | 1p34.1 |
| NAV1 | Neuron navigator 1 | 1q32.1 |
| NBEA | Neurobeachin | 13q13.3 |
| NCKAP1 | NCK-associated protein 1 | 2q32.1 |
| NCKAP5 | NCK-associated protein 5 | 2q21.2 |
| NCKAP5L | NCK-associated protein 5-like | 12q13.12 |
| NCOR1 | Nuclear receptor corepressor 1 | 17p11.2 |
| NDNL2 | Necdin-like gene 2 | 15q13.1 |
| NDUFA5 | NADH-ubiquinone oxidoreductase 1 α subcomplex, 5 | 7q31.32 |
| NEFL | Neurofilament protein, light polypeptide | 8p21.2 |
| NELL1 | NEL-like 1 | 11p15.1 |
| NF1 | Neurofibromin 1 | 17q11.2 |
| NFIA | Nuclear factor I/A | 1p31.3 |
| NIPA1 | Non imprinted gene in Prader-Willi/Angelman syndrome chromosomal region 1 | 15q11.2 |
| NIPA2 | Non imprinted gene in Prader-Willi/Angelman syndrome chromosomal region 2 | 15q11.2 |
| NIPBL | Nipped-B-like | 5p13.2 |
| NLGN1 | Neuroligin 1 | 3q26.31 |
| NLGN2 | Neuroligin 2 | 17p13.1 |
| NLGN3 | Neuroligin 3 | Xq13.1 |
| NLGN4X | Neuroligin 4, X-linked | Xp22.31 |
| NLGN4Y | Neuroligin 4, Y-linked | Yq11.221 |
| NOS1AP | Nitric oxide synthase 1 (neuronal) adaptor protein | 1q23.3 |
| NOS2A | Nitric oxide synthase 2A | 17q11.2 |
| NOTCH3 | Notch 3 | 19p13.12 |
| NPAS2 | Neuronal PAS domain protein 2 | 2q11.2 |
| NR0B1 | Nuclear receptor subfamily 0, group B, member 1 | Xp21.2 |
| NR3C2 | Nuclear receptor subfamily 3, group C, member 2 | 4q31.23 |
| NR4A1 | Nuclear receptor subfamily 4, group A, member 1 | 12q13.13 |
| NRCAM | Neuronal cell adhesion molecule | 7q31.1 |
| NRG1 | Neuregulin 1 | 8p12 |
| NRP2 | Neuropilin 2 | 2q33.3 |
| NRXN1 | Neurexin I | 2p16.3 |
| NRXN2 | Neurexin II | 11q13.1 |
| NRXN3 | Neurexin III | 14q24.3 |
| NSD1 | Nuclear receptor-binding Sa-var, enhancer of zeste, and trithorax domain protein 1 | 5q35.3 |
| NTNG1 | Netrin G1 | 1p13.3 |
| NTRK1 | Neurotrophic tyrosine kinase, receptor, type 1 | 1q23.1 |
| NTRK3 | Neurotrophic tyrosine kinase, receptor, type 3 | 15q25.3 |
| NXF5 | Nuclear RNA export factor 5 | Xq22.1 |
| NXPH1 | Neurexophilin 1 | 7p21.3 |
| ODF3L2 | Outer dense fiber of sperm tails 3-like 2 | 19p13.3 |
| OGT | O-linked N-acetylglucosamine transferase | Xq13.1 |
| OPHN1 | Oligophrenin 1 | Xq12 |
| OPRM1 | Opioid receptor, mu 1 | 6q25.2 |
| OR1C1 | Olfactory receptor, family 1, subfamily C, member 1 | 1q44 |
| OTX1 | Orthodenticle Drosophila, homolog of | 2p15 |
| OXTR | Oxytocin receptor | 3p25.3 |
| P2RX4 | Purinergic receptor P2X, ligand-gated ion channel, 4 | 12q24.31 |
| PAFAH1B1 | Platelet-activating factor acetylhydrolase 1B, regulatory subunit 1 | 17p13.3 |
| PAH | Phenylalanine hydroxylase | 12q23.2 |
| PARD3B | PAR-3 family cell polarity regulator β | 2q33.3 |
| PARK2 | Parkin | 6q26 |
| PAX5 | Paired box 5 | 9p13.2 |
| PBRM1 | Polybromo 1 | 3p21.1 |
| PCDH10 | Protocadherin 10 | 4q28.3 |
| PCDH15 | Protocadherin 15 | 10q21.1 |
| PCDH19 | Protocadherin 19 | Xq22.1 |
| PCDH8 | Protocadherin 8 | 13q14.3 |
| PCDH9 | Protocadherin 9 | 13q21.32 |
| PCDHA1 | Protocadherin α 1 | 5q31.3 |
| PCDHA10 | Protocadherin α 10 | 5q31.3 |
| PCDHA11 | Protocadherin α 11 | 5q31.3 |
| PCDHA12 | Protocadherin α 12 | 5q31.3 |
| PCDHA13 | Protocadherin α 13 | 5q31.3 |
| PCDHA2 | Protocadherin α 2 | 5q31.3 |
| PCDHA3 | Protocadherin α 3 | 5q31.3 |
| PCDHA4 | Protocadherin α 4 | 5q31.3 |
| PCDHA5 | Protocadherin α 5 | 5q31.3 |
| PCDHA6 | Protocadherin α 6 | 5q31.3 |
| PCDHA7 | Protocadherin α 7 | 5q31.3 |
| PCDHA8 | Protocadherin α 8 | 5q31.3 |
| PCDHA9 | Protocadherin α 9 | 5q31.3 |
| PCDHAC1 | Protocadherin α subfamily C, member 1 | 5q31.3 |
| PCDHAC2 | Protocadherin α subfamily C, member 2 | 5q31.3 |
| PCDHGA11 | Protocadherin γ subfamily A, member 11 | 5q31.3 |
| PDE1C | Phosphodiesterase 1C | 7p14.3 |
| PDE4A | Phosphodiesterase 4A, cAMP-specific | 19p13.2 |
| PDE4B | Phosphodiesterase 4B, cAMP-specific | 1p31.3 |
| PDZD4 | PDZ domain containing 4 | Xq28 |
| PECR | Peroxisomal trans-2-enoyl-CoA reductase | 2q35 |
| PER1 | Period, Drosophila, homolog of | 17p13.1 |
| PEX7 | Peroxisomal biogenesis factor 7 | 6q23.3 |
| PGD | Phosphogluconate dehydrogenase | 1p36.22 |
| PHF2 | PHD finger protein 2 | 9q22.31 |
| PHF8 | PHD finger protein 8 | Xp11.22 |
| PIAS1 | Protein inhibitor of activated STAT, 1 | 15q23 |
| PIK3CG | Phosphatidylinositol-3-kinase, catalytic, γ | 7q22.3 |
| PIK3R2 | Phosphatidylinositol-3-kinase, regulatory subunit 2 | 19q13.11 |
| PINX1 | PIN2 interacting protein 1 | 8p23.1 |
| PITX1 | Paired-like homeodomain transcription factor 1 | 5q31.1 |
| PLAUR | Plasminogen activator receptor, urokinase-type | 19q13.31 |
| PLCB1 | Phospholipase C, β 1 | 20p12.3 |
| PLCD1 | Phospholipase C, δ 1 | 3p22.2 |
| PLN | Phospholamban | 6q22.31 |
| PLXNA4 | Plexin A4 | 7q32.3 |
| POGZ | POGO transposable element with ZNF domain | 1q21.3 |
| POLR2L | Polymerase (RNA) II (DNA directed) polypeptide L, 7.6 kDa | 11p15.5 |
| POMGNT1 | Protein O-mannose β-1, 2-N-acetylglucosaminyl-transferase | 1p34.1 |
| PON1 | Paraoxonase 1 | 7q21.3 |
| POT1 | Protection of telomeres 1 | 7q31.33 |
| PPFIA1 | Protein tyrosine phosphatase, receptor type, F polypeptide, interacting protein, α 1 | 11q13.3 |
| PPP1CB | Protein phosphatase 1, catalytic subunit, β isozyme | 2p23.2 |
| PPP1R1B | Protein phosphatase 1, regulatory (inhibitor) subunit 1B | 17q12 |
| PPP1R3F | Protein phosphatase 1, regulatory (inhibitor) subunit 3F | Xp11.23 |
| PRODH | Proline dehydrogenase (oxidase) 1 | 22q11.21 |
| PRICKLE1 | Prickle, Drosophila, homolog of, 1 | 12q12 |
| PRICKLE2 | Prickle, Drosophila, homolog of, 2 | 3p14.1 |
| PRKCB | Protein kinase C, β | 16p12.2 |
| PRKCB1 | Protein kinase C, β-1 | 16p12.2 |
| PRKD1 | Protein kinase D1 | 14q12 |
| PRDX1 | Peroxiredoxin 1 | 1p34.1 |
| PRSS38 | Protease, serine, 38 | 1q42.13 |
| PRUNE2 | Prune, Drosophila, homolog of, 2 | 9q21.2 |
| PSD3 | Pleckstrin and Sec7 domains-containing protein 3 | 8p22 |
| PSEN1 | Presenilin 1 | 14q24.2 |
| PSMD10 | Proteasome 26S subunit, non-ATPase, 10 | Xq22.3 |
| PTCHD1 | Patched domain containing protein 1 | Xp22.11 |
| PTEN | Phosphatase and tensin homolog | 10q23.31 |
| PTGER3 | Prostaglandin E receptor 3, EP3 subtype | 1p31.1 |
| PTGS2 | Prostaglandin-endoperoxide synthase 2 | 1q31.1 |
| PTPN11 | Protein tyrosine phosphatase, non-receptor type 11 | 12q24.13 |
| PTPRB | Protein tyrosine phosphatase, receptor type, B | 12q15 |
| PTPRC | Protein tyrosine phosphatase, receptor type, C | 1q31.3 |
| PTPRM | Protein tyrosine phosphatase, receptor type, M | 18p11.23 |
| PTPRT | Protein tyrosine phosphatase, receptor type, T | 20q13.11 |
| PXDN | Peroxidasin, Drosophila homolog of | 2p25.3 |
| RAB11FIP5 | RAB11 family-interacting protein 5 | 2p13.2 |
| RAB19 | RAB19, member RAS oncogene family | 7q34 |
| RAB39B | RAS-associated protein RAB39B | Xq28 |
| RAI1 | Retinoic acid induced gene 1 | 17p11.2 |
| RAPGEF4 | Rap guanine nucleotide exchange factor | 2q31.1 |
| RASD1 | RAS protein, dexamethasone-induced, 1 | 17p11.2 |
| RASSF1 | RAS association (ralGDS/AF-6) domain family member 1 | 3p21.31 |
| RASSF5 | RAS association domain family protein 5 | 1q32.1 |
| RB1CC1 | RB1-inducible coiled-coil 1 | 8q11.23 |
| RBFOX1 | RNA binding protein FOX-1, C. elegans, homolog of, 1 | 16p13.3 |
| RBM8A | RNA binding motif protein 8A | 1q21.1 |
| RBMS3 | RNA binding motif protein, single stranded interacting, 3 | 3p24.1 |
| REEP3 | Receptor expression-enhancing protein 3 | 10q21.3 |
| RELN | Reelin | 7q22.1 |
| RERE | RE-repeats encoding gene | 1p36.23 |
| RFWD2 | Ring finger and WD repeat domains-containing protein 2 | 1q25.2 |
| RGS7 | Regulator of G protein signaling 7 | 1q43 |
| RHOXF1 | RHOX homeobox family, member 1 | Xq24 |
| RIC8A | RIC8 guanine nucleotide exchange factor A | 11p15.5 |
| RIMS1 | Regulating synaptic membrane exocytosis 1 | 6q13 |
| RIMS3 | Protein regulating synaptic membrane exocytosis 3 | 1p34.2 |
| RNPS1 | RNA binding protein S1 | 16p13.3 |
| ROBO1 | Roundabout, Drosophila, homolog of, 1 | 3p12.2 |
| ROBO2 | Roundabout, Drosophila, homolog of, 2 | 3p12.3 |
| RORA | RAR-related orphan receptor A | 15q22.2 |
| RPL10 | Ribosomal protein L10 | Xq28 |
| RPP25 | Ribonuclease P/MRP 25 kDa subunit | 15q24.2 |
| RPS6KA1 | Ribosomal protein S6 kinase, 90 kDa, polypeptide 1 | 1p36.11 |
| RPS6KA2 | Ribosomal protein S6 kinase, 90 kDa, polypeptide 2 | 6q27 |
| RPS6KA3 | Ribosomal protein S6 kinase, 90 kDa, polypeptide 3 | Xp22.12 |
| RUVBL1 | RuvB-E. coli, homolog-like 1 | 3q21.3 |
| SAE1 | SUMO1 activating enzyme, subunit 1 | 19q13.32 |
| SATB2 | Special AT-rich sequence-binding protein 2 | 2q33.1 |
| SBF1 | SET binding factor 1 | 22q13.33 |
| SCFD2 | Sec1 family domain containing 2 | 4q12 |
| SCN1A | Sodium channel, neuronal, type I, α subunit | 2q24.3 |
| SCN2A | Sodium channel, voltage-gated, type II, α subunit | 2q24.3 |
| SCN7A | Sodium channel, voltage-gated, type VII, α subunit | 2q24.3 |
| SCN8A | Sodium channel, voltage-gated, type VIII, α subunit | 12q13.13 |
| SDC2 | Syndecan 2 | 8q22.1 |
| SDK1 | Sidekick cell adhesion molecule 1 | 7p22.2 |
| SEMA3F | Sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3F | 3p21.31 |
| SEMA5A | Semaphorin 5A | 5p15.31 |
| SERPINE1 | Serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 1 | 7q22.1 |
| SETBP1 | SET binding protein 1 | 18q12.3 |
| SETD2 | SET domain containing protein 2 | 3p21.31 |
| SETD5 | SET domain containing protein 5 | 3p25.3 |
| SETDB1 | SET domain, bifurcated, 1 | 1q21.3 |
| SETDB2 | SET domain, bifurcated, 2 | 13q14.2 |
| SEZ6L2 | Seizure related 6 homolog (mouse)-like 2 | 16p11.2 |
| SF1 | Splicing factor 1 | 11q13.1 |
| SFPQ | Splicing factor proline/glutamine-rich | 1p34.3 |
| SFTPD | Surfactant, pulmonary-associated protein D | 10q22.3 |
| SGSH | N-sulfoglucosamine sulfohydrolase | 17q25.3 |
| SGSM3 | Small G protein signaling modulator 3 | 22q13.1 |
| SH3KBP1 | SH3-domain kinase binding protein 1 | Xp22.12 |
| SHANK1 | SH3 and multiple ankyrin repeat domains 1 | 19q13.3 |
| SHANK2 | SH3 and multiple ankyrin repeat domains 2 | 11q13.4 |
| SHANK3 | SH3 and multiple ankyrin repeat domains 3 | 22q13.33 |
| SLC16A3 | Solute carrier family 16 (monocarboxylic acid transporter), member 3 | 17q25 |
| SLC16A7 | Solute carrier family 16 (monocarboxylic acid transporter), member 7 | 12q14.1 |
| SLC1A1 | Solute carrier family 1 (neuronal/epithelial high affinity glutamate transporter), member 1 | 9p24.2 |
| SLC22A15 | Solute carrier family 22, (organic cation transporter), member 15 | 1p13.1 |
| SLC24A2 | Solute carrier family 24 (sodium/potassium/calcium exchanger), member 2 | 9p22.1 |
| SLC25A12 | Solute carrier family 25 (mitochondrial carrier, Aralar), member 12 | 2q31.1 |
| SLC25A14 | Solute carrier family 25 (mitochondrial carrier, brain), member 14 | Xq26.1 |
| SLC25A24 | Solute carrier family 25 (mitochondrial carrier, phosphate carrier), member 24 | 1p13.3 |
| SLC25A27 | Solute carrier family 25, member 27 | 6p12.3 |
| SLC29A4 | Solute carrier family 29 (equilibrative nucleoside transporter), member 4 | 7p22.1 |
| SLC30A5 | Solute carrier family 30 (zinc transporter), member 5 | 5q13.1 |
| SLC35A3 | Solute carrier family 35 (UDP-N-acetylglucosamine transporter), member 3 | 1p21.2 |
| SLC38A10 | Solute carrier family 38, member 10 | 17q25.3 |
| SLC39A11 | Solute carrier family 39 (metal ion transporter), member 11 | 17q21.31 |
| SLC4A10 | Solute carrier family 4 (sodium bicarbonate transporter-like), member 10 | 2q24.2 |
| SLC6A1 | Solute carrier family 6 (neurotransmitter transporter), member 1 | 3p25.3 |
| SLC6A3 | Solute carrier family 6 (neurotransmitter transporter, dopamine), member 3 | 5p15.33 |
| SLC6A4 | Solute carrier family 6 (neurotransmitter transporter, serotonin), member 4 | 17q11.2 |
| SLC6A8 | Solute carrier family 6 (neurotransmitter transporter, creatine), member 8 | Xq28 |
| SLC9A6 | Solute carrier family 9 (sodium/hydrogen exchanger), member 6 | Xq26.3 |
| SLC9A9 | Solute carrier family 9 (sodium/hydrogen exchanger), member 9 | 3q24 |
| SLCO1B1 | Solute carrier organic anion transporter family, member 1B1 | 12p12.2 |
| SLCO1B3 | Solute carrier organic anion transporter family, member 1B3 | 12p12.2 |
| SLIT3 | Slit, Drosophila, homolog of, 3 | 5q35.1 |
| SLITRK5 | SLIT and NTRK-like family, member 5 | 13q31.2 |
| SLK | STE20-like kinase | 10q24.33 |
| SMAD2 | SMAD family member 2 | 18q21.1 |
| SMARCC2 | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily C, member 2 | 12q13.2 |
| SMG6 | SMG 6, C. elegans, homolog of | 17p13.3 |
| SND1 | EBNA2 coactivator p100 | 7q32.1 |
| SNRPN | Small nuclear ribonucleoprotein polypeptide N | 15q11.2 |
| SNTG2 | Syntrophin, γ 2 | 2p25.3 |
| SNX19 | Sorting nexin 19 | 11q25 |
| SNX5 | Sorting nexin 5 | 20p11.23 |
| SOD1 | Superoxide dismutase 1, soluble | 21q22.11 |
| SOS1 | Son of sevenless (SOS), Drosophila, homolog 1 | 2p22.1 |
| SOX5 | SRY (sex determining region Y)-box 5 | 12p12.1 |
| SOX7 | SRY (sex determining region Y)-box 7 | 8p23.1 |
| SPAST | Spastin | 2p22.3 |
| SRD5A2 | Steroid-5-α-reductase, 2 | 2p23.1 |
| ST7 | Suppressor of tumorigenicity 7 | 7q31.2 |
| ST8SIA2 | ST8 α-N-acetyl-neuraminide α-2,8-sialyltransferase 2 | 15q26.1 |
| STK39 | Serine/threonine protein kinase 39 | 2q24.3 |
| STX6 | Syntaxin 6 | 1q25.3 |
| STX1A | Syntaxin 1A | 7q11.23 |
| STXBP1 | Syntaxin-binding protein 1 | 9q34.1 |
| STXBP5 | Syntaxin-binding protein 5 | 6q24.3 |
| STXBP5L | Syntaxin-binding protein 5-like | 3q13.33 |
| SUCLG2 | Succinate-CoA ligase, GDP-forming, β subunit | 3p14.1 |
| SUV420H1 | Suppressor of variegation 4–20, Drosophila, homolog of, 1 | 11q13.2 |
| SYAP1 | Synapse associated protein 1 | Xp22.2 |
| SYN1 | Synapsin 1 | Xp11.23 |
| SYN2 | Synapsin II | 3p25.2 |
| SYN3 | Synapsin III | 22q12.3 |
| SYNE1 | Spectrin repeat containing nuclear envelope 1 | 6q25.2 |
| SYNGAP1 | Synaptic RAS-GTPase-activating protein 1 | 6p21.32 |
| SYT17 | Synaptotagmin XVII | 16p12.3 |
| SYT3 | Synaptotagmin III | 19q13.33 |
| TAF1C | TATA box-binding protein-associated factor 1C | 16q24.1 |
| TAF1L | TATA box-binding protein-associated factor 1-like | 9p21.1 |
| TAS2R1 | Taste receptor, type 2, member 1 | 5p15.31 |
| TBC1D30 | TBC1 domain family, member 30 | 12q14.3 |
| TBC1D5 | TBC1 domain family, member 5 | 3p24.3 |
| TBC1D7 | TBC1 domain family, member 7 | 6p24 |
| TBL1X | Transducin-β-like 1, X-linked | Xp22.31 |
| TBL1XR1 | Transducin-β-like 1 receptor 1 | 3q26.32 |
| TBR1 | T-box, brain, 1 | 2q24.2 |
| TBX1 | T-box 1 | 22q11.21 |
| TCF3 | Transcription factor 3 | 19p13.3 |
| TCF4 | Transcription factor 4 | 18q21.2 |
| TCF20 | Transcription factor 20 (AR1) | 22q13.2 |
| TCF7L2 | Transcription factor 7-like 2 (t-cell specific, HMG-box) | 10q25.2 |
| TDO2 | Tryptophan 2,3-dioxygenase | 4q32.1 |
| TGM3 | Transglutaminase 3 | 20p13 |
| TH | Tyrosine hydroxylase | 11p15.5 |
| THBS1 | Thrombospondin 1 | 15q14 |
| THRA | Thyroid hormone receptor, α-1 | 17q21.1 |
| TLK2 | Tousled-like kinase 2 | 17q23.2 |
| TLX1 | T-cell leukemia homeobox 1 | 10q24.31 |
| TM4SF20 | Transmembrane 4 L6 family, member 20 | 2q36.3 |
| TMEM231 | Transmembrane protein 231 | 16q23.1 |
| TMLHE | Epsilon-trimethyllysine hydroxylase | Xq28 |
| TNIP2 | TNFAIP3 interacting protein 2 | 4p16.3 |
| TNRC6B | Trinucleotide repeat containing 6B | 22q13.1 |
| TOMM20 | MAS20P, S. cerevisiae, homolog of | 1q42.3 |
| TOP1 | Topoisomerase, DNA, I | 20q12 |
| TOP3B | Topoisomerase, DNA, III, β | 22q11.22 |
| TOPBP1 | Topoisomerase (DNA) II-binding protein 1 | 3q22.1 |
| TOPORS | Topoisomerase I-binding, arginine/serine-rich, E3 ubiquitin protein ligase | 9p21.1 |
| TPH2 | Tryptophan hydroxylase 2 | 12q21.1 |
| TPO | Thyroid peroxidase | 2p25.3 |
| TRIM33 | Tripartite motif containing protein 33 | 1p13.2 |
| TRIO | Trio Rho guanine nucleotide exchange factor | 5p15.2 |
| TRIP12 | Thyroid hormone receptor interactor 12 | 2q36.3 |
| TRPC6 | Transient receptor potential cation channel, subfamily C, member 6 | 11q22.1 |
| TRPM1 | Transient receptor potential cation channel, subfamily M, member 1 | 15q13.3 |
| TSC1 | Tuberous sclerosis 1 | 9q34.1 |
| TSC2 | Tuberous sclerosis 2 | 16p13.3 |
| TSN | Translin | 2q14.3 |
| TSPAN7 | Tetraspanin 7 | Xp11.4 |
| TTI2 | TELO2-interacting protein 2 | 8p12 |
| TTN | Titin | 2q31.2 |
| TUBA1A | Tubulin, α-1A | 12q13.12 |
| TUBGCP5 | Tubulin-γ complex-associated protein 5 | 15q11.2 |
| TYR | Tyrosinase | 11q14.3 |
| UBE1L2 | Ubiquitin-activating enzyme, E1-like 2 | 4q13.2 |
| UBE2H | Ubiquitin-conjugating enzyme E2H | 7q32.2 |
| UBE3A | Ubiquitin protein ligase E3A | 15q11.2 |
| UBE3B | Ubiquitin protein ligase E3B | 12q24.11 |
| UBE3C | Ubiquitin protein ligase E3C | 7q36.3 |
| UBL7 | Ubiquitin-like 7 | 15q24.1 |
| UBR5 | Ubiquitin protein ligase E3 component N-recognin 5 | 8q22.3 |
| UBR7 | Ubiquitin protein ligase E3 component N-recognin 7 | 14q32.12 |
| UIMC1 | Ubiquitin interaction motif containing 1 | 5q35.2 |
| UPB1 | Ureidopropionase, β 1 | 22q11.23 |
| UPF2 | UPF2, yeast, homolog of | 10p14 |
| UPF3B | UPF3, yeast, homolog of, B | Xq24 |
| USP54 | Ubiquitin specific peptidase 54 | 10q22.2 |
| USP9Y | Ubiquitin specific protease 9, Y-chromosome | Yq11.21 |
| VASH1 | Vasohibin 1 | 14q24.3 |
| VCP | Valosin containing protein | 9p13.3 |
| VIL1 | Villin 1 | 2q35 |
| VIP | Vasoactive intestinal peptide (VIP) | 6q25.2 |
| VPS13B | Vacuolar protein sorting 13, yeast, homolog of, B | 8q22.2 |
| VPS4A | Vacuolar protein sorting 4 homolog A (S. cerevisiae) | 16q22.1 |
| WAC | WW domain containing adaptor with coiled-coil | 10p12.1 |
| WDFY3 | WD repeat and FYVE domain containing 3 | 4q21.23 |
| WHSC1 | Wolf-Hirschhorn syndrome candidate 1 | 4p16.3 |
| WNK3 | Protein kinase lysine deficient 3 | Xp11.22 |
| WNT1 | Wingless-type MMTV integration site family, member 1 | 12q13.12 |
| WNT2 | Wingless-type MMTV integration site family, member 2 | 7q31.2 |
| WWC3 | WWC family member 3 | Xp22.32 |
| XIRP1 | Cardiomyopathy-associated protein 1 | 3p22.2 |
| XPC | Xeroderma pigmentosum complementation group C | 3p25.1 |
| XPO1 | Exportin 1 | 2p15 |
| XPO5 | Exportin 5 | 6p21.1 |
| YEATS2 | YEATS domain containing 2 | 3q27.1 |
| YTHDC2 | YTH domain containing 2 | 5q22.2 |
| YWHAE | Tyrosine 3-monooxygenase, tryptophan 5-monooxygenase activation protein, epsilon isoform | 17p13.3 |
| ZBTB16 | Zinc finger- and BTB domain-containing protein 16 | 11q23.1 |
| ZBTB20 | Zinc finger- and BTB domain-containing protein 20 | 3q13.31 |
| ZC3H12B | Zinc finger CCCH domain-containing protein 12B | Xq12 |
| ZFPL1 | Zinc finger protein-like 1 | 11q13.1 |
| ZMYND11 | Zinc finger, MYND-type containing 11 | 10p15.3 |
| ZNF18 | Zinc finger protein 18 | 17p12 |
| ZNF365 | Zinc finger protein 365 | 10q21.2 |
| ZNF385B | Zinc finger protein 385B | 2q31.3 |
| ZNF407 | Zinc finger protein 407 | 18q23 |
| ZNF517 | Zinc finger protein 517 | 8q24.3 |
| ZNF8 | Zinc finger protein 8 | 19q13.43 |
| ZNF713 | Zinc finger protein 713 | 7p11.2 |
| ZNF804A | Zinc finger protein 804A | 2q32.1 |
| ZNF827 | Zinc finger protein 827 | 4q31.22 |
| ZSWIM5 | Zinc finger, SWIM-type containing 5 | 1p34.1 |
3. Experimental Section
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- American Psychiatric Association. Diagnostic and Statistical Manual of Mental Disorders, 4th ed.; American Psychiatric Association: Washington, DC, USA, 2000. [Google Scholar]
- Johnson, C.P.; Myers, S.M.; American Academy of Pediatrics Counsel on Children with Disablities. Identifiction and evaluation of children with autism spectrum disorders. Pediatrics 2007, 120, 1183–1215. [Google Scholar] [CrossRef]
- Hughes, J.R. Update on autism: A review of 1300 reports published in 2008. Epilepsy Behav. 2009, 16, 569–589. [Google Scholar] [CrossRef] [PubMed]
- Lord, C.; Risi, S.; Lambrecht, L.; Cook, E.H., Jr.; Leventhal, B.L.; DiLavore, P.C.; Pickles, A.; Rutter, M. The autism diagnostic observation schedule-generic: A standard measure of social and communication deficits associated with the spectrum of autism. J. Autism Dev. Disord. 2000, 30, 205–223. [Google Scholar] [CrossRef] [PubMed]
- Le Couteur, A.; Lord, C.; Rutter, M. Autism Diagnostic Interview-Reviewed (ADI-R); Western Psychological Services: Los Angeles, CA, USA, 2003. [Google Scholar]
- Constantino, J.N.; Davis, S.A.; Todd, R.D.; Schindler, M.K.; Gross, M.M.; Brophy, S.L.; Metzger, L.M.; Shoushtari, C.S.; Splinter, R.; Reich, W. Validation of a brief quantitative measure of autistic traits: Comparison of the social responsiveness scale with the autism diagnostic interview-revised. J. Autism Dev. Disord. 2003, 33, 427–433. [Google Scholar] [CrossRef] [PubMed]
- Rice, C. Prevalence of autism spectrum disorders-autism and developmental disabilities monitoring network, United States, 2006. MMWR Surveill. Summ. 2009, 58, 3–8. [Google Scholar]
- Fombonne, E. Epidemiology of autistic disorder and other pervasive developmental disorders. J. Clin. Psychiatry 2005, 66 (Suppl. 10), 3–8. [Google Scholar] [PubMed]
- Kanner, L. Autistic psychopathy in childhood. Nerv. Child. 1943, 2, 217–250. [Google Scholar]
- Rapin, I. Autistic regression and disintegrative disorder: How important the role of epilepsy? Semin. Pediatr. Neurol. 1995, 2, 278–285. [Google Scholar] [CrossRef] [PubMed]
- Geschwind, D.H.; Levitt, P. Autism spectrum disorders: Developmental disconnection syndromes. Curr. Opin. Neurobiol. 2007, 17, 103–111. [Google Scholar] [CrossRef] [PubMed]
- Kurtzke, J. Neuroepidemiology. In Neurology in Clinical Practice; Bradley, W., Daroff, R., Fenichel, G., Marsden, C., Eds.; Butterworth-Heinemann: Stoneham, MA, USA, 1991; pp. 545–560. [Google Scholar]
- Gadia, C.A.; Tuchman, R.; Rotta, N.T. Autism and pervasive developmental disorders. J. Pediatr. 2004, 80, S83–S94. [Google Scholar]
- Butler, M.G.; Dasouki, M.J.; Zhou, X.P.; Talebizadeh, Z.; Brown, M.; Takahashi, T.N.; Miles, J.H.; Wang, C.H.; Stratton, R.; Pilarski, R.; et al. Subset of individuals with autism spectrum disorders and extreme macrocephaly associated with germline PTEN tumour suppressor gene mutations. J. Med. Genet. 2005, 42, 318–321. [Google Scholar] [CrossRef]
- Prontera, P.; Ottaviani, V.; Toccaceli, D.; Rogaia, D.; Ardisia, C.; Romani, R.; Stangoni, G.; Pierini, A.; Donti, E. Recurrent approximately 100 kb microdeletion in the chromosomal region 14q11.2, involving CHD8 gene, is associated with autism and macrocephaly. Am. J. Med. Genet. A 2014, 164, 3137–3141. [Google Scholar] [CrossRef]
- Benvenuto, A.; Moavero, R.; Alessandrelli, R.; Manzi, B.; Curatolo, P. Syndromic autism: Causes and pathogenetic pathways. World J. Pediatr. 2009, 5, 169–176. [Google Scholar] [CrossRef] [PubMed]
- Holt, R.; Monaco, A.P. Links between genetics and pathophysiology in the autism spectrum disorders. EMBO Mol. Med. 2011, 3, 438–450. [Google Scholar] [CrossRef] [PubMed]
- Campbell, D.B.; Sutcliffe, J.S.; Ebert, P.J.; Militerni, R.; Bravaccio, C.; Trillo, S.; Elia, M.; Schneider, C.; Melmed, R.; Sacco, R.; et al. A genetic variant that disrupts MET transcription is associated with autism. Proc. Natl. Acad. Sci. USA 2006, 103, 16834–16839. [Google Scholar] [CrossRef]
- Schaefer, G.B.; Starr, L.; Pickering, D.; Skar, G.; Dehaai, K.; Sanger, W.G. Array comparative genomic hybridization findings in a cohort referred for an autism evaluation. J. Child Neurol. 2010, 25, 1498–1503. [Google Scholar] [CrossRef] [PubMed]
- Roberts, J.L.; Hovanes, K.; Dasouki, M.; Manzardo, A.M.; Butler, M.G. Chromosomal microarray analysis of consecutive individuals with autism spectrum disorders or learning disability presenting for genetic services. Gene 2014, 535, 70–78. [Google Scholar] [CrossRef] [PubMed]
- Butler, M.G.; Usrey, K.; Roberts, J.L.; Schroeder, S.R.; Manzardo, A.M. Clinical presentation and microarray analysis of Peruvian children with atypical development and/or aberrant behavior. Genet. Res. Int. 2014, 2014, 408516. [Google Scholar] [PubMed]
- Miles, J.H. Autism spectrum disorders—A genetics review. Genet. Med. 2011, 13, 278–294. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Takumi, T. Genomic and genetic aspects of autism spectrum disorder. Biochem. Biophys. Res. Commun. 2014, 452, 244–253. [Google Scholar] [CrossRef] [PubMed]
- Cox, D.M.; Butler, M.G. The 15q11.2 BP1-BP2 microdeletion syndrome: A review. Int. J. Mol. Sci. 2015, 16, 4068–4082. [Google Scholar] [CrossRef] [PubMed]
- Hempel, M.; Rivera Brugues, N.; Wagenstaller, J.; Lederer, G.; Weitensteiner, A.; Seidel, H.; Meitinger, T.; Strom, T.M. Microdeletion syndrome 16p11.2-p12.2: Clinical and molecular characterization. Am. J. Med. Genet. A 2009, 149, 2106–2112. [Google Scholar] [CrossRef]
- Fernandez, B.A.; Roberts, W.; Chung, B.; Weksberg, R.; Meyn, S.; Szatmari, P.; Joseph-George, A.M.; Mackay, S.; Whitten, K.; Noble, B.; et al. Phenotypic spectrum associated with de novo and inherited deletions and duplications at 16p11.2 in individuals ascertained for diagnosis of autism spectrum disorder. J. Med. Genet. 2010, 47, 195–203. [Google Scholar] [CrossRef]
- Miller, D.T.; Shen, Y.; Weiss, L.A.; Korn, J.; Anselm, I.; Bridgemohan, C.; Cox, G.F.; Dickinson, H.; Gentile, J.; Harris, D.J.; et al. Microdeletion/duplication at 15q13.2q13.3 among individuals with features of autism and other neuropsychiatric disorders. J. Med. Genet. 2009, 46, 242–248. [Google Scholar] [CrossRef]
- Ritvo, E.R.; Jorde, L.B.; Mason-Brothers, A.; Freeman, B.J.; Pingree, C.; Jones, M.B.; McMahon, W.M.; Petersen, P.B.; Jenson, W.R.; Mo, A. The UCLA-university of Utah epidemiologic survey of autism: Recurrence risk estimates and genetic counseling. Am. J. Psychiatry 1989, 146, 1032–1036. [Google Scholar] [CrossRef] [PubMed]
- Sandin, S.; Lichtenstein, P.; Kuja-Halkola, R.; Larsson, H.; Hultman, C.M.; Reichenberg, A. The familial risk of autism. JAMA 2014, 311, 1770–1777. [Google Scholar] [CrossRef] [PubMed]
- Sebat, J.; Lakshmi, B.; Malhotra, D.; Troge, J.; Lese-Martin, C.; Walsh, T.; Yamrom, B.; Yoon, S.; Krasnitz, A.; Kendall, J.; et al. Strong association of de novo copy number mutations with autism. Science 2007, 316, 445–449. [Google Scholar] [CrossRef] [PubMed]
- Lauritsen, M.B.; Als, T.D.; Dahl, H.A.; Flint, T.J.; Wang, A.G.; Vang, M.; Kruse, T.A.; Ewald, H.; Mors, O. A genome-wide search for alleles and haplotypes associated with autism and related pervasive developmental disorders on the Faroe islands. Mol. Psychiatry 2006, 11, 37–46. [Google Scholar] [CrossRef] [PubMed]
- Ma, D.; Salyakina, D.; Jaworski, J.M.; Konidari, I.; Whitehead, P.L.; Andersen, A.N.; Hoffman, J.D.; Slifer, S.H.; Hedges, D.J.; Cukier, H.N.; et al. A genome-wide association study of autism reveals a common novel risk locus at 5p14.1. Ann. Hum. Genet. 2009, 73, 263–273. [Google Scholar] [CrossRef]
- Weiss, L.A.; Arking, D.E.; Gene Discovery Project of Johns Hopkins & the Autism Consortium; Daly, M.J.; Chakravarti, A. A genome-wide linkage and association scan reveals novel loci for autism. Nature 2009, 461, 802–808. [Google Scholar] [CrossRef] [PubMed]
- Anney, R.; Klei, L.; Pinto, D.; Regan, R.; Conroy, J.; Magalhaes, T.R.; Correia, C.; Abrahams, B.S.; Sykes, N.; Pagnamenta, A.T.; et al. A genome-wide scan for common alleles affecting risk for autism. Hum. Mol. Genet. 2010, 19, 4072–4082. [Google Scholar] [CrossRef]
- Wang, K.; Zhang, H.; Ma, D.; Bucan, M.; Glessner, J.T.; Abrahams, B.S.; Salyakina, D.; Imielinski, M.; Bradfield, J.P.; Sleiman, P.M.; et al. Common genetic variants on 5p14.1 associate with autism spectrum disorders. Nature 2009, 459, 528–533. [Google Scholar] [CrossRef]
- Pinto, D.; Pagnamenta, A.T.; Klei, L.; Anney, R.; Merico, D.; Regan, R.; Conroy, J.; Magalhaes, T.R.; Correia, C.; Abrahams, B.S.; et al. Functional impact of global rare copy number variation in autism spectrum disorders. Nature 2010, 466, 368–372. [Google Scholar] [CrossRef]
- Pagnamenta, A.T.; Khan, H.; Walker, S.; Gerrelli, D.; Wing, K.; Bonaglia, M.C.; Giorda, R.; Berney, T.; Mani, E.; Molteni, M.; et al. Rare familial 16q21 microdeletions under a linkage peak implicate cadherin 8 (CDH8) in susceptibility to autism and learning disability. J. Med. Genet. 2011, 48, 48–54. [Google Scholar] [CrossRef]
- Klei, L.; Sanders, S.J.; Murtha, M.T.; Hus, V.; Lowe, J.K.; Willsey, A.J.; Moreno-De-Luca, D.; Yu, T.W.; Fombonne, E.; Geschwind, D.; et al. Common genetic variants, acting additively, are a major source of risk for autism. Mol. Autism 2012, 3, 9. [Google Scholar] [CrossRef]
- Campbell, M.G.; Kohane, I.S.; Kong, S.W. Pathway-based outlier method reveals heterogeneous genomic structure of autism in blood transcriptome. BMC Med. Genomics 2013, 6, 34. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.H.; Yuen, R.K.; Jin, X.; Wang, M.; Chen, N.; Wu, X.; Ju, J.; Mei, J.; Shi, Y.; He, M.; et al. Detection of clinically relevant genetic variants in autism spectrum disorder by whole-genome sequencing. Am. J. Hum. Genet. 2013, 93, 249–263. [Google Scholar] [CrossRef] [PubMed]
- Huguet, G.; Ey, E.; Bourgeron, T. The genetic landscapes of autism spectrum disorders. Ann. Rev. Genomics Hum. Genet. 2013, 14, 191–213. [Google Scholar] [CrossRef]
- Correia, C.; Oliveira, G.; Vicente, A.M. Protein interaction networks reveal novel autism risk genes within gwas statistical noise. PLoS One 2014, 9, e112399. [Google Scholar] [CrossRef] [PubMed]
- Merikangas, A.K.; Segurado, R.; Heron, E.A.; Anney, R.J.; Paterson, A.D.; Cook, E.H.; Pinto, D.; Scherer, S.W.; Szatmari, P.; Gill, M.; et al. The phenotypic manifestations of rare genic CNVs in autism spectrum disorder. Mol. Psychiatry 2014. [Google Scholar] [CrossRef]
- Lossifov, I.; O’Roak, B.J.; Sanders, S.J.; Ronemus, M.; Krumm, N.; Levy, D.; Stessman, H.A.; Witherspoon, K.T.; Vives, L.; Patterson, K.E.; et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature 2014, 515, 216–221. [Google Scholar] [CrossRef] [PubMed]
- Goldani, A.A.S.; Downs, S.R.; Widjaja, F.; Lawton, B.; Hendren, R.L. Biomarkers in autism. Front. Psychiatry 2014, 5, 100. [Google Scholar] [CrossRef] [PubMed]
- Ch’ng, C.; Kwok, W.; Rogic, S.; Pavlidis, P. Meta-analysis of gene expression in autism spectrum disorder. 2015. [Google Scholar] [CrossRef]
- Uddin, M.; Tammimies, K.; Pellecchia, G.; Alipanahi, B.; Hu, P.; Wang, Z.; Pinto, D.; Lau, L.; Nalpathamkalam, T.; Marshall, C.R.; et al. Brain-expressed exons under purifying selection are enriched for de novo mutations in autism spectrum disorder. Nat. Genet. 2014, 46, 742–747. [Google Scholar] [CrossRef] [PubMed]
- De Rubeis, S.; He, X.; Goldberg, A.P.; Poultney, C.S.; Samocha, K.; Cicek, A.E.; Kou, Y.; Liu, L.; Fromer, M.; Walker, S.; et al. Synaptic, transcriptional and chromatin genes disrupted in autism. Nature 2014, 515, 209–215. [Google Scholar] [CrossRef] [PubMed]
- Hadley, D.; Wu, Z.L.; Kao, C.; Kini, A.; Mohamed-Hadley, A.; Thomas, K.; Vazquez, L.; Qiu, H.; Mentch, F.; Pellegrino, R.; et al. The impact of the metabotropic glutamate receptor and other gene family interaction networks on autism. Nat. Commun. 2014. [Google Scholar] [CrossRef]
- Butler, M.G.; Rafi, S.K.; Hossain, W.; Stephan, D.A.; Manzardo, A.M. Whole exome sequencing in females with autism implicates novel and candidate genes. Int. J. Mol. Sci. 2015, 16, 1312–1335. [Google Scholar] [CrossRef] [PubMed]
- Yuen, R.K.; Thiruvahindrapuram, B.; Merico, D.; Walker, S.; Tammimies, K.; Hoang, N.; Chrysler, C.; Nalpathamkalam, T.; Pellecchia, G.; Liu, Y.; et al. Whole-genome sequencing of quartet families with autism spectrum disorder. Nat. Med. 2015, 21, 185–191. [Google Scholar] [CrossRef] [PubMed]
- Willsey, J.A.; State, M.W. Autism spectrum disorders: From genes to neurobiology. Curr. Opin. Neurobiol. 2015, 30, 92–99. [Google Scholar] [CrossRef] [PubMed]
- Hormozdiari, F.; Penn, O.; Borenstein, E.; Eichler, E.E. The discovery of integrated gene networks for autism and related disorders. Genome Res. 2015, 25, 142–154. [Google Scholar] [CrossRef] [PubMed]
- Butler, M.G.; Youngs, E.L.; Roberts, J.L.; Hellings, J.A. Assessment and treatment in autism spectrum disorders: A focus on genetics and psychiatry. Autism Res. Treat. 2012. [Google Scholar] [CrossRef]
- Herman, G.E.; Henninger, N.; Ratliff-Schaub, K.; Pastore, M.; Fitzgerald, S.; McBride, K.L. Genetic testing in autism: How much is enough? Genet. Med. 2007, 9, 268–274. [Google Scholar] [CrossRef] [PubMed]
- Butler, M.G. Prader-willi syndrome: Obesity due to genomic imprinting. Curr. Genomics 2011, 12, 204–215. [Google Scholar] [CrossRef] [PubMed]
- Dhillon, S.; Hellings, J.A.; Butler, M.G. Genetics and mitochondrial abnormalities in autism spectrum disorders: A review. Curr. Genomics 2011, 12, 322–332. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Butler, M.G.; Rafi, S.K.; Manzardo, A.M. High-Resolution Chromosome Ideogram Representation of Currently Recognized Genes for Autism Spectrum Disorders. Int. J. Mol. Sci. 2015, 16, 6464-6495. https://doi.org/10.3390/ijms16036464
Butler MG, Rafi SK, Manzardo AM. High-Resolution Chromosome Ideogram Representation of Currently Recognized Genes for Autism Spectrum Disorders. International Journal of Molecular Sciences. 2015; 16(3):6464-6495. https://doi.org/10.3390/ijms16036464
Chicago/Turabian StyleButler, Merlin G., Syed K. Rafi, and Ann M. Manzardo. 2015. "High-Resolution Chromosome Ideogram Representation of Currently Recognized Genes for Autism Spectrum Disorders" International Journal of Molecular Sciences 16, no. 3: 6464-6495. https://doi.org/10.3390/ijms16036464
APA StyleButler, M. G., Rafi, S. K., & Manzardo, A. M. (2015). High-Resolution Chromosome Ideogram Representation of Currently Recognized Genes for Autism Spectrum Disorders. International Journal of Molecular Sciences, 16(3), 6464-6495. https://doi.org/10.3390/ijms16036464