Silencing of the CaCP Gene Delays Salt- and Osmotic-Induced Leaf Senescence in Capsicum annuum L.
Abstract
:1. Introduction
2. Results and Discussion
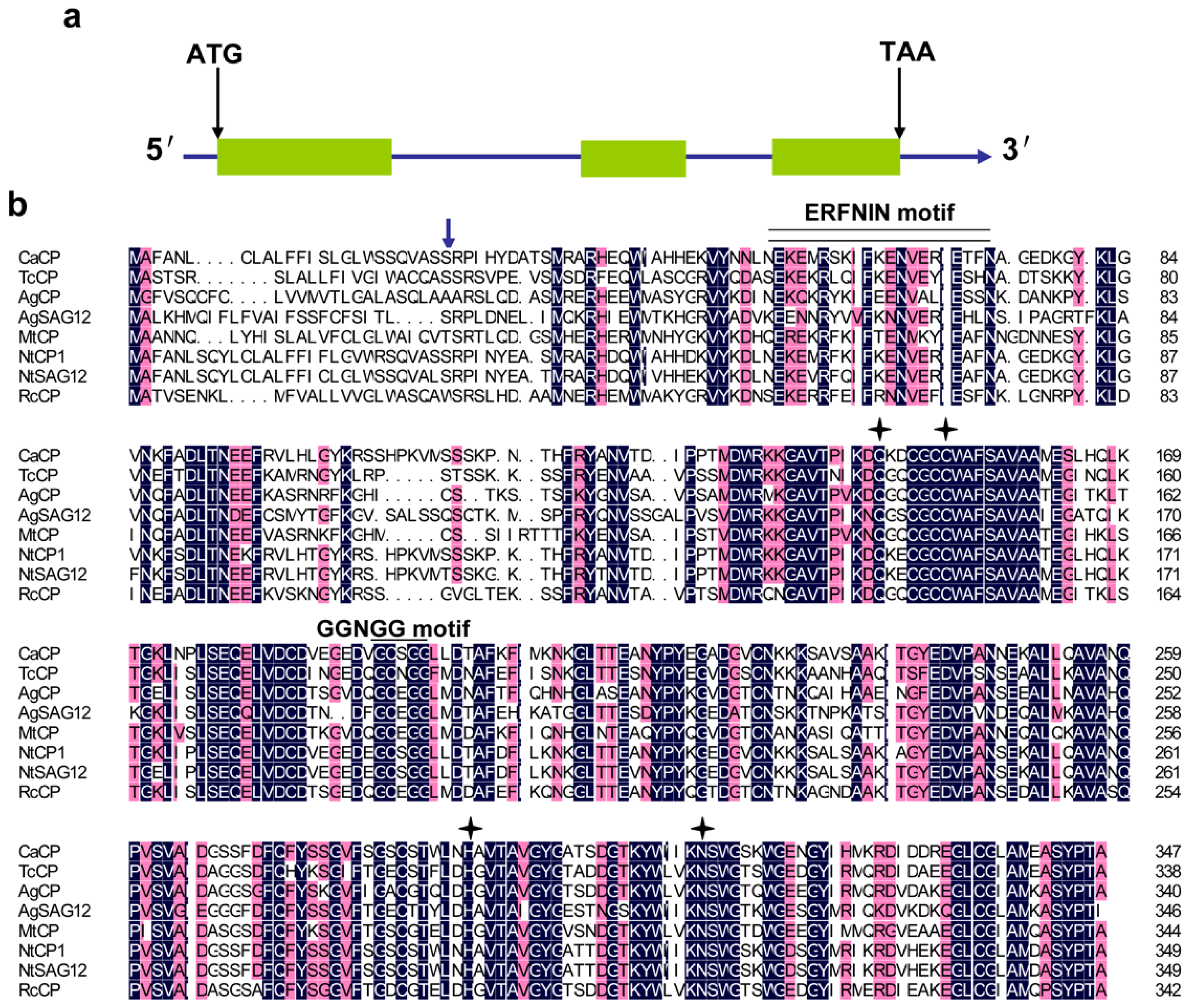
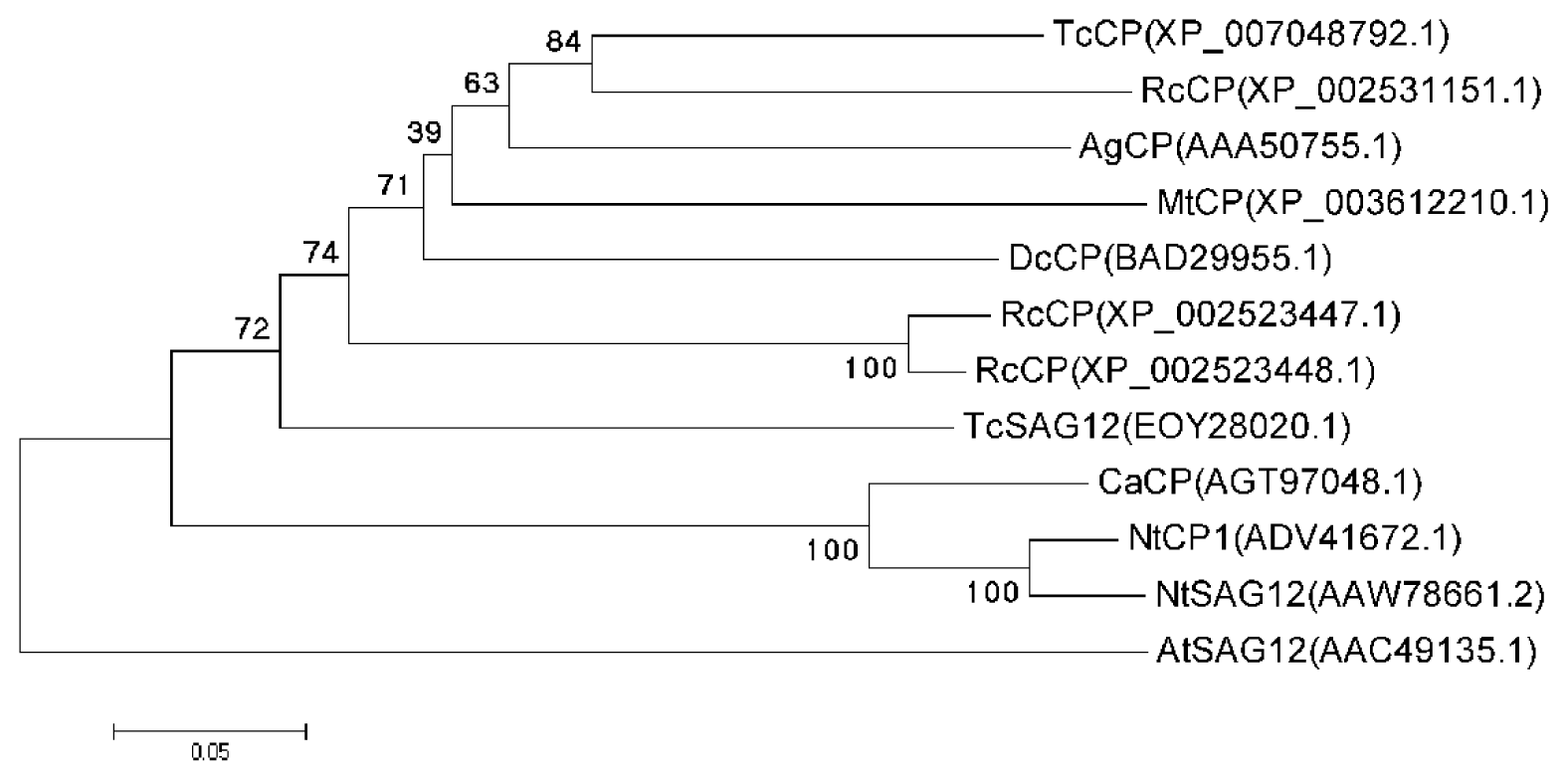
2.1. Cloning and Sequence Analysis of the CaCP Gene
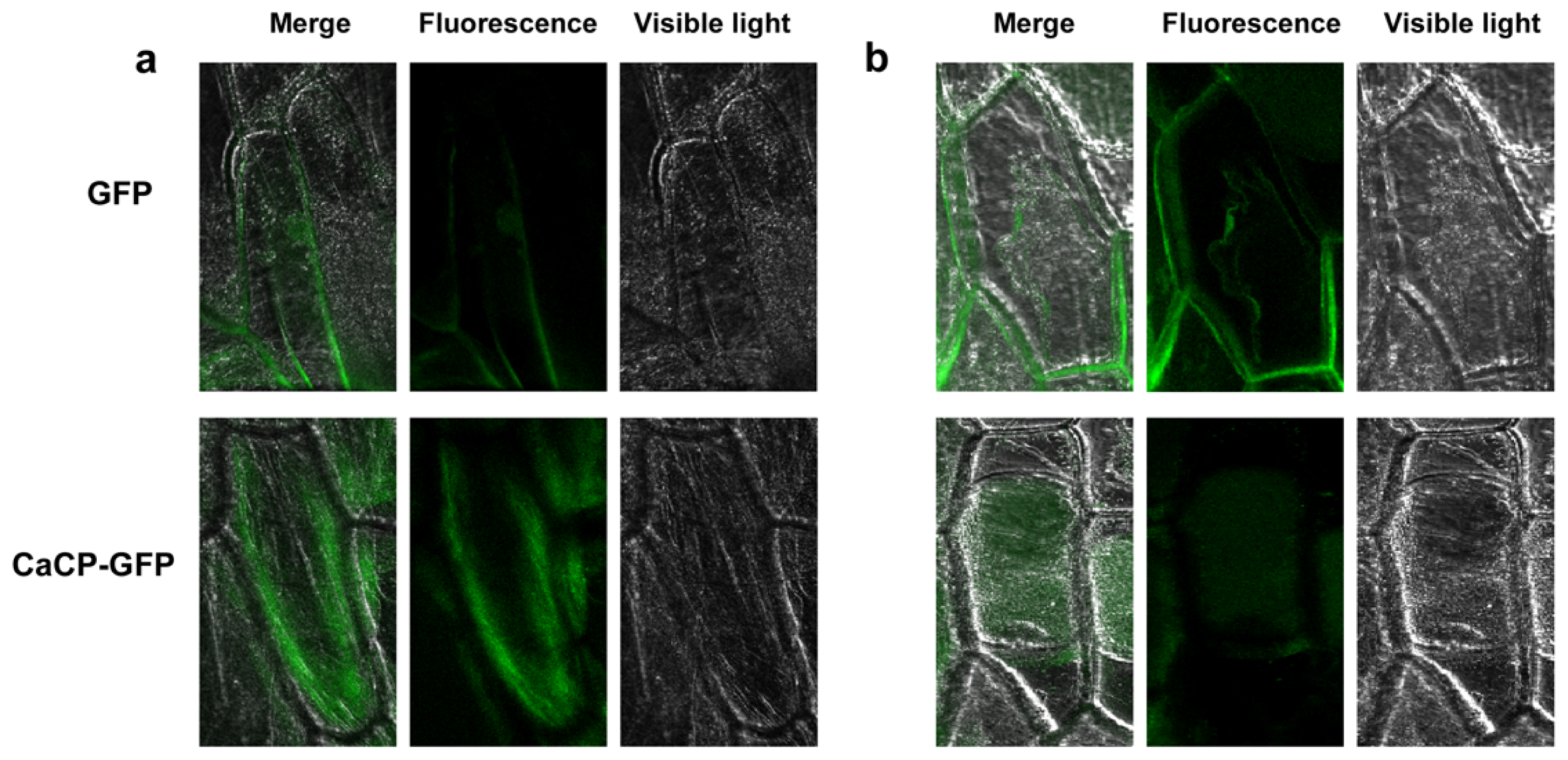
2.2. Subcellular Localization of CaCP Protein
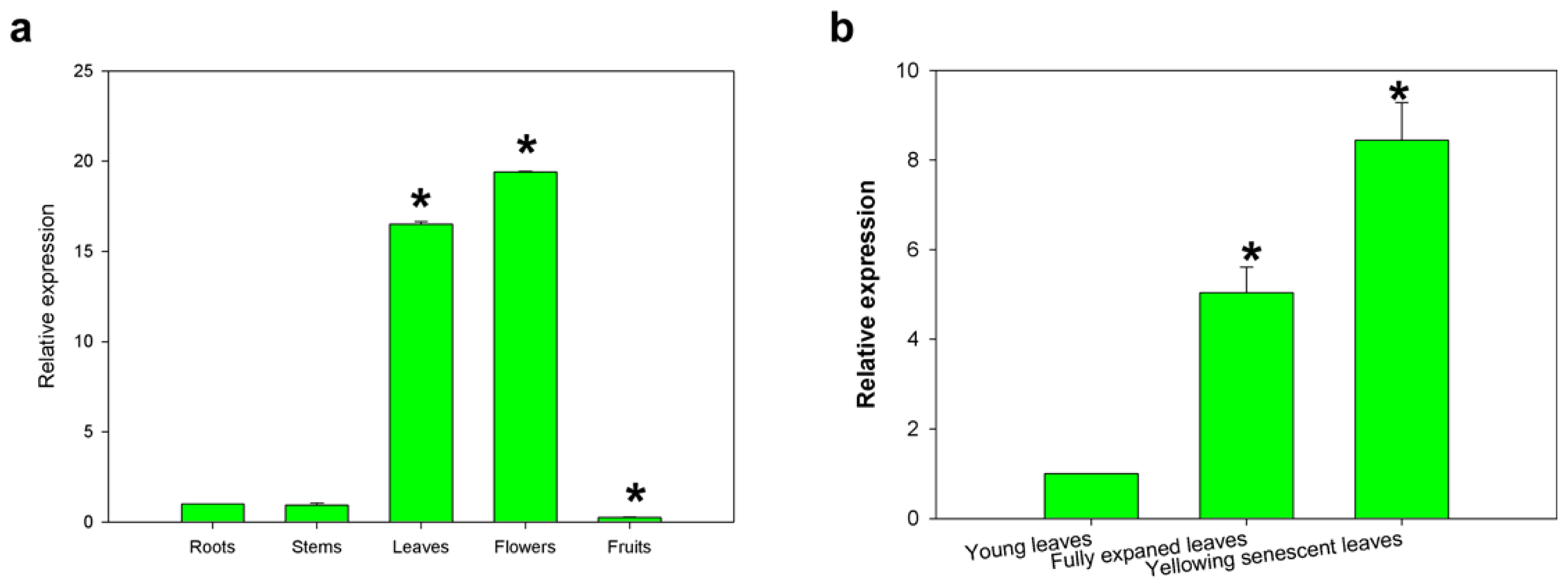
2.3. Expression of the CaCP Gene in Different Pepper Tissues and during Leaf Development
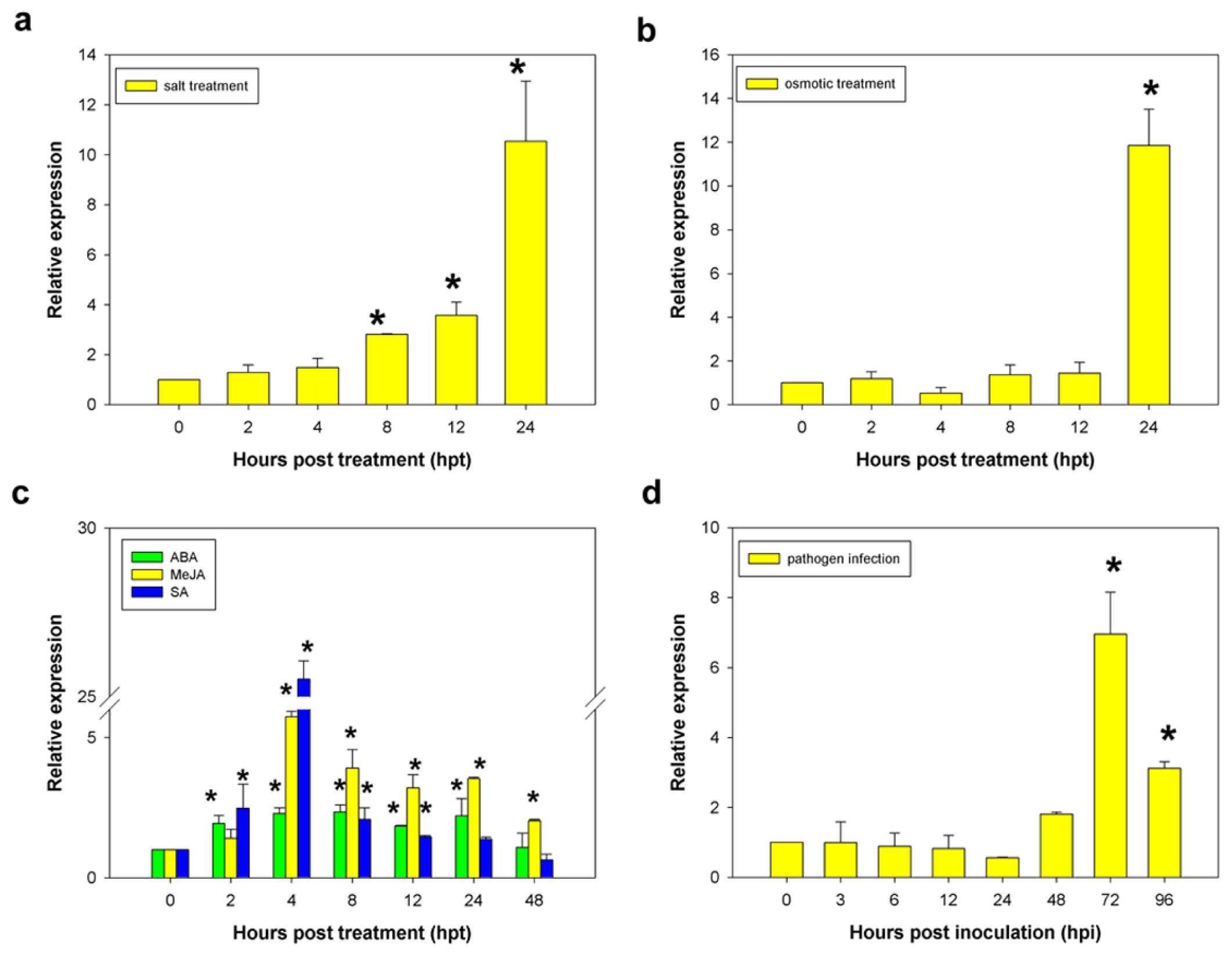
2.4. Induction of the CaCP Gene in Pepper Leaves by Signaling Molecules, Abiotic and Biotic Stresses
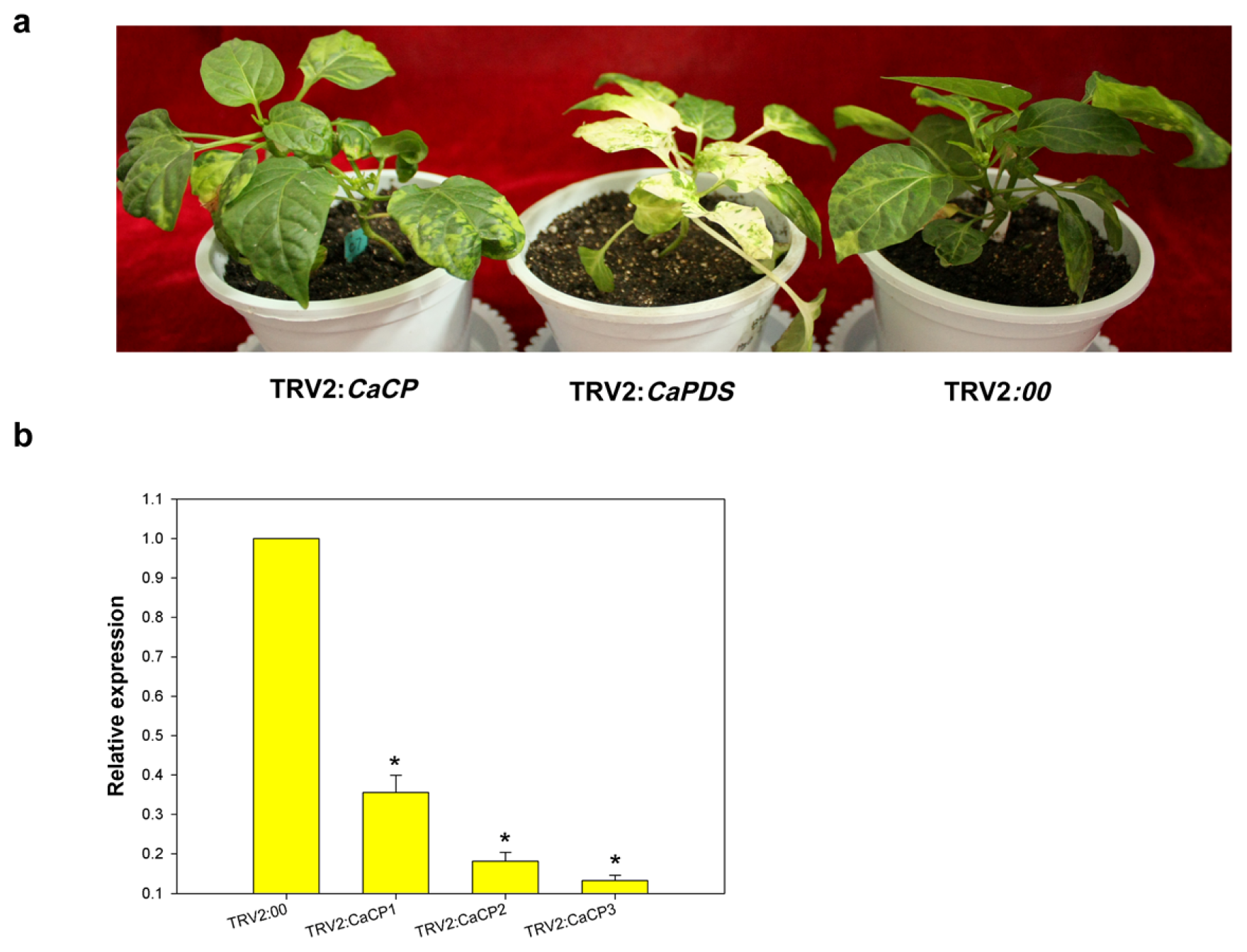
2.5. Phenotype and Silencing Efficiency of CaCP-Silenced Plants
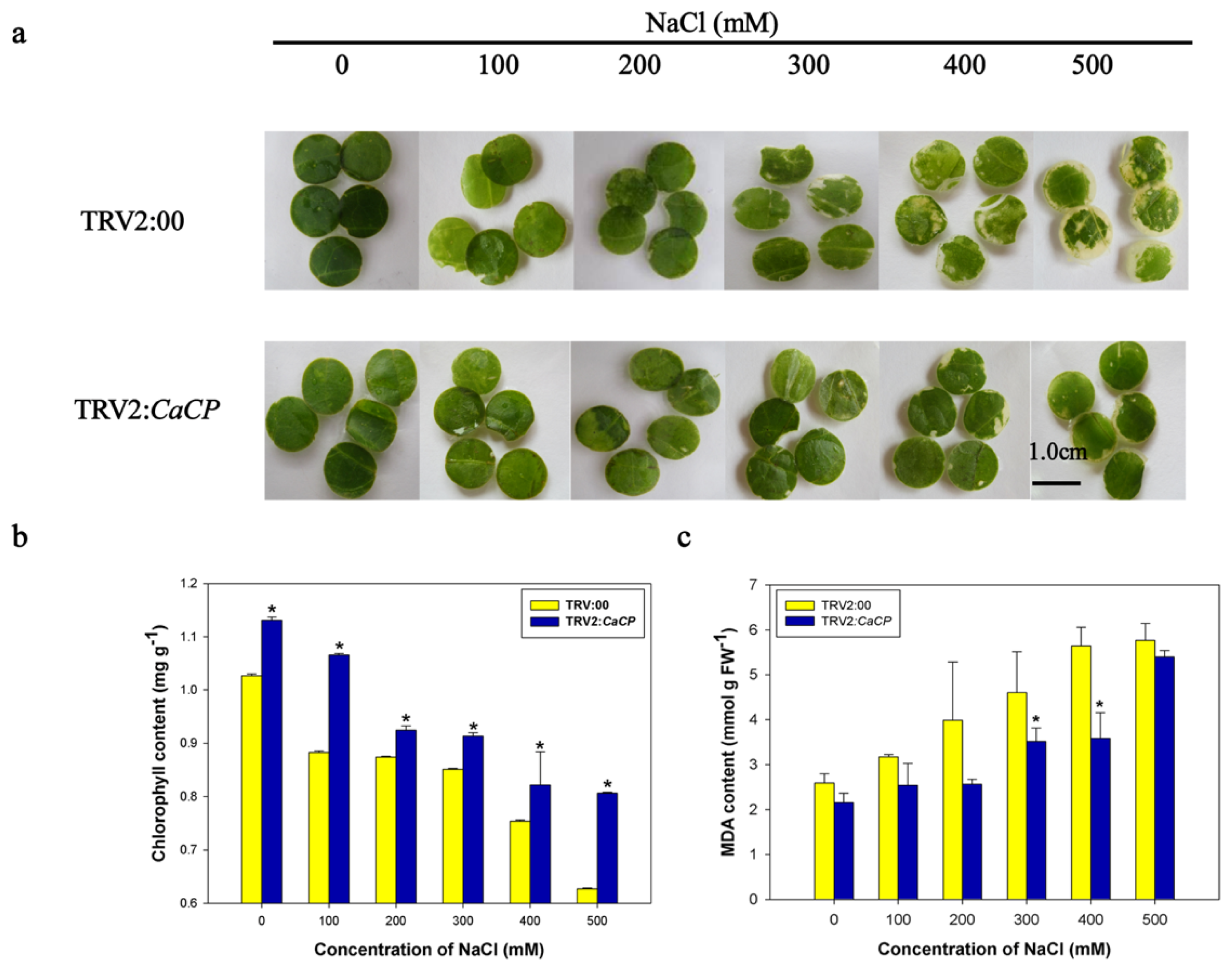
2.6. Enhanced Tolerance of CaCP-Silenced Pepper Plants to Salt Stress
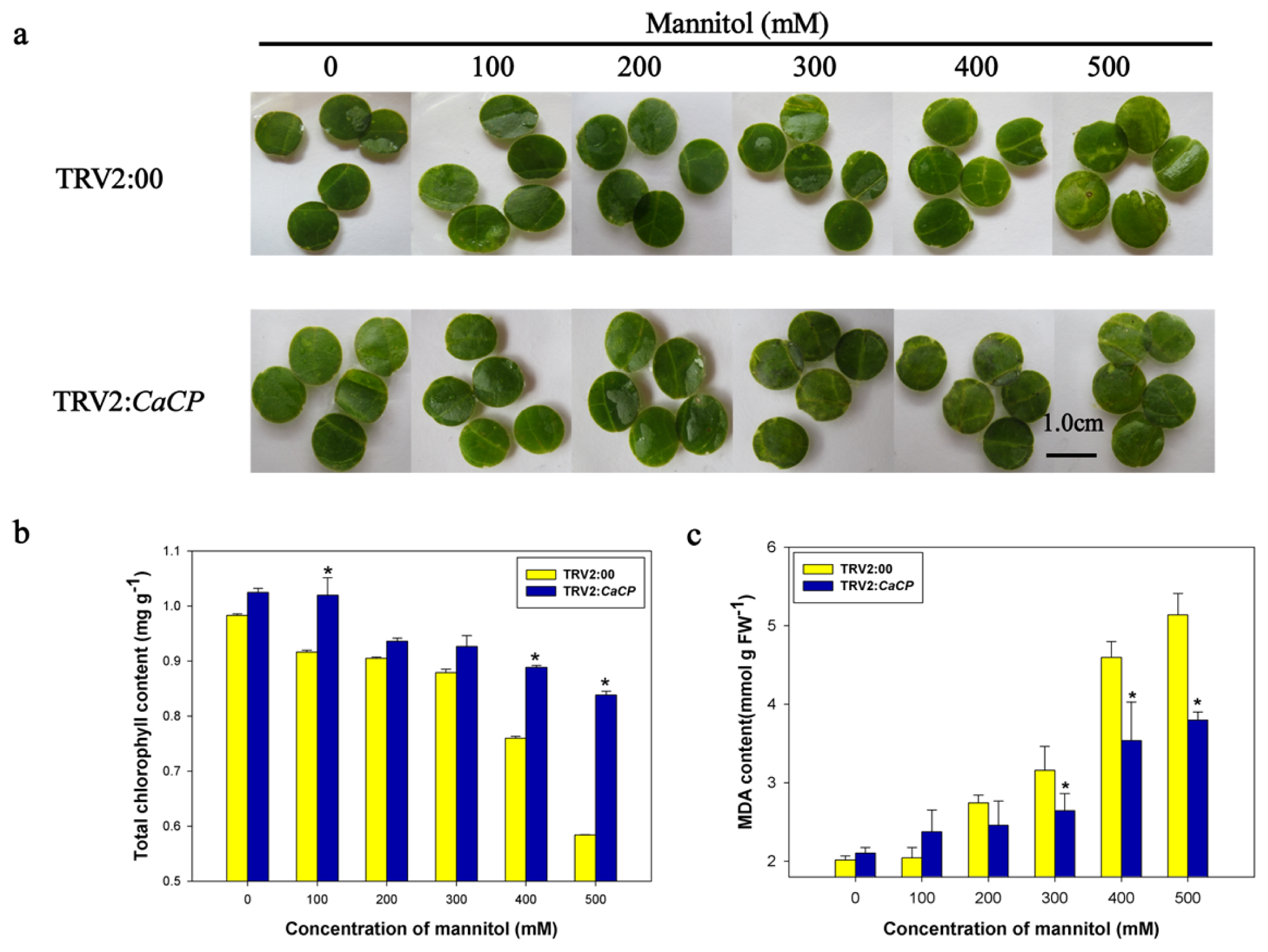
2.7. Silencing of CaCP Enhanced Tolerance to Osmotic Stress in Pepper Plants
2.8. Discussion
3. Experimental Section
3.1. Plant Material and Growth Condition
3.2. Plant Treatments
3.3. Measurement of Chlorophyll Content and MDA Content
3.4. Isolation of RNA and Genomic DNA
3.5. First-Strand cDNA Synthesis and qRT-PCR Analysis
3.6. Cloning and Sequence Analysis of the CaCP Gene
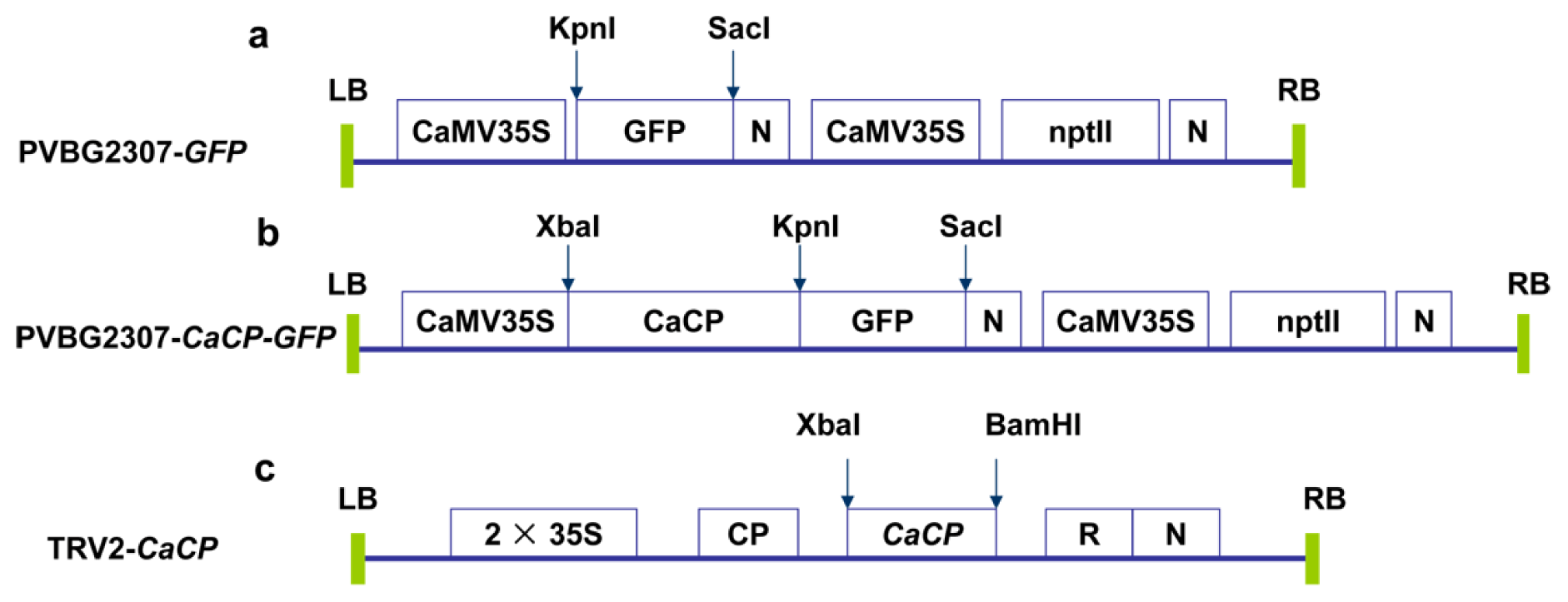
3.7. VIGS Assay of CaCP in Pepper Plant
3.8. Subcellular Localization of the CaCP Protein
3.9. Primers Used in this Study
3.10. Statistical Analysis
4. Conclusions
Acknowledgments
Conflicts of Interest
- Author ContributionsConceived and designed the experiments: H.-J.X., Z.-H.G. Performed the experiments: H.-J.X., Y.-X.Y. Analyzed the data: H.-J.X., Y.-X.Y., W.-G.C. Contributed reagents/materials/analysis tools: Z.-H.G. Wrote the paper: H.-J.X.
References
- Smart, C. Gene expression during leaf senescence. New Phytol 1994, 126, 419–448. [Google Scholar]
- Lim, P.O.; Kim, H.J.; Nam, H.G. Leaf Senescence. Annu. Rev. Plant Biol 2007, 58, 115–136. [Google Scholar]
- Guo, Y.; Cai, Z.; Gan, S. Transcriptome of Arabidopsis leaf senescence. Plant Cell Environ 2004, 27, 521–549. [Google Scholar]
- Makino, A.; Osmond, B. Effects of nitrogen nutrition on nitrogen partitioning between chloroplasts and mitochondria in pea and wheat. Plant Physiol 1991, 96, 355–362. [Google Scholar]
- Staswick, P.E. Storage proteins of vegetative plant tissues. Annu. Rev. Plant Phys 1994, 45, 303–322. [Google Scholar]
- Robertsa, I.N.; Caputo, C.; Criado, M.V.; Funk, C. Senescence-associated proteases in plants. Physiol. Plantarum 2012, 145, 130–139. [Google Scholar]
- Toyooka, K.; Okamoto, T.; Minamikawa, T. Mass transport of a proform of a KDEL-tailed cysteine proteinase (SH-EP) to protein storage vacuoles by endoplasmic reticulum-derived vesicle is involved in protein mobilization in germinating seeds. J. Cell Biol 2000, 148, 453–563. [Google Scholar]
- Jones, M.L.; Chaffin1, G.S.; Eason, J.R.; Clark, D.G. Ethylene-sensitivity regulates proteolytic activity and cysteine protease gene expression in petunia corollas. J. Exp. Bot 2005, 420, 2733–2744. [Google Scholar]
- Watanabe, Y.; Matsushima, S.; Yamaguchi, A.; Shioi, Y. Characterization and cloning of cysteine protease that is induced in green leaves of barley. Plant Sci 2009, 176, 264–271. [Google Scholar]
- Fan, J.; Yang, Y.W.; Gao, X.; Deng, W.; Falara, V.; Kanellis, A.K.; Li, Z.G. Expression of a senescence-associated cysteine protease gene related to peel pitting of navel orange (Citrus sinensis L. Osbeck). Plant Cell Tissue Org 2009, 98, 281–289. [Google Scholar]
- Zhang, X.M.; Wang, Y.; Lv, X.M.; Li, H.; Sun, P.; Lu, H.; Li, F.L. NtCP56, a new cysteine protease in Nicotiana tabacum L., involved in pollen grain development. J. Exp. Bot 2009, 6, 1569–1577. [Google Scholar]
- Naito, Y.; Fujie, M.; Usami, S.; Murooka, Y.; Yamada, T. The involvement of cysteine proteinase in the nodule development in Chinese milk vetch infected with Mesorhizobium huakuii subsp. Rengei. Plant Physiol 2000, 124, 1087–1096. [Google Scholar]
- Li, Y.X.; Zhou, L.; Li, Y.G.; Chen, D.S.; Tan, X.J.; Lei, L.; Zhou, J.C. A nodule-specific plant cysteine proteinase, AsNODF32, is involved in nodule senescence and nitrogen fixation activity of the green manure legume Astragalus sinicus. New Phytol 2008, 180, 185–192. [Google Scholar]
- Forsthoefel, N.R.; Cushman, M.A.F.; Ostrem, J.A.; Cushman, J.C. Induction of a cysteine protease cDNA from Mesembryanthemum crystallinum leaves by environmental stress and plant growth regulators. Plant Sci 1998, 136, 195–206. [Google Scholar]
- Simova, S.L.; Vaseva, I.; Grigorova, B.; Demirevska, K.; Feller, U. Proteolytic activity and cysteine protease expression in wheat leaves under severe soil drought and recovery. Plant Physiol. Biochem 2010, 48, 200–206. [Google Scholar]
- Maciel, F.M.; Salles, C.M.C.; Retamal, C.A.; Gomes, V.M.; Machado, O.L.T. Identification and partial characterization of two cysteine proteases from castor bean leaves (Ricinus communis L.) activated by wounding and methyl jasmonate stress. Acta Physiol Plant 2011, 33, 1867–1875. [Google Scholar]
- Klostermana, S.J.; Anchieta, A.; Garcia-Pedrajas, M.D.; Maruthachalam, K.; Hayes, R.J.; Subbarao, K.V. SSH reveals a linkage between a senescence-associated protease and Verticillium wilt symptom development in lettuce (Lactuca sativa). Physiol. Mol. Plant Pathol 2011, 76, 48–58. [Google Scholar]
- Shin, R.; Lee, G.J.; Park, C.J.; Kim, T.Y.; You, J.S.; Nam, Y.W.; Peak, K.H. Isolation of pepper mRNAs differentially expressed during the hypersensitive response to tobacco mosaic virus and characterization of a proteinase inhibitor gene. Plant Sci 2001, 161, 727–737. [Google Scholar]
- Karrer, K.M.; Peiffer, S.L.; DiTomas, M.E. Two distinct gene subfamilies of cysteine proteinase gene. Proc. Natl. Acad. Sci. USA 1993, 90, 3063–3067. [Google Scholar]
- Okamoto, T.; Shimada, T.; Hara-Nishimura, I.; Nishimura, M.; Minamikawa, T. C-terminal KDEL sequence of a KDEL tailed cysteine proteinase (sulfhydryl-endopeptidase) is involved in formation of KDEL vesicle and in efficient vacuolar transport of sulfhydryl-endopeptidase. Plant Physiol 2003, 132, 1892–1900. [Google Scholar]
- Ling, J.; Kojima, T.; Shiraiwa, M.; Takahara, H. Cloning of two cysteine proteinases genes: CysP1 and CysP2, from soybean cotyledons by cDNA representational difference analysis. Biochim. Biophys. Acta 2003, 1627, 129–139. [Google Scholar]
- Granell, A.; Cercos, M.; Carbonell, J. Plant cysteine proteinases in germination and senescence. In Handbook of Proteolytic Enzymes; Barrett, A.J., Rawlings, N.D., Woessner, J.F., Eds.; Academic Press: San Diego, CA, USA, 1998; pp. 578–583. [Google Scholar]
- Richau, K.H.; Kaschani, F.; Verdoes, M.; Pansuriya, T.C.; Niessen, S.; Stüber, K.; Colby, T.; Overkleeft, H.S.; Bogyo, M.; van der Hoorn, R.A. Subclassification and biochemical analysis of plant papain-like cysteine proteases displays subfamily-specific characteristics. Plant Physiol 2012, 158, 1583–1599. [Google Scholar]
- Guerrero, C.; Calle, M.; Reid, M.S.; Valpuesta, V. Analysis of the expression of two thiolprotease genes from daylily (Hemerocallis spp.) during flower senescence. Plant Mol. Biol 1998, 36, 565–571. [Google Scholar]
- Beyene, G.; Foyer, C.H.; Kunert, K.J. Two new cysteine proteinases with specific expression patterns in mature and senescent tobacco (Nicotiana tabacum L.) leaves. J. Exp. Bot 2006, 57, 1431–1443. [Google Scholar]
- Drake, R.; John, I.; Farrell, A.; Cooper, W.; Schuch, W.; Grierson, D. Isolation and analysis of cDNAs encoding tomato cysteine proteases expressed during leaf senescence. Plant Mol. Biol 1996, 30, 755–767. [Google Scholar]
- Lohman, K.N.; Gan, S.; John, M.C.; Amasino, R.M. Molecular analysis of natural leaf senescence in Arabidopsis thaliana. Physiol. Plantarum 1994, 92, 322–328. [Google Scholar]
- Ueda, T.; Seo, S.; Ohashi, Y.; Hashimoto, J. Circadian and senescence-enhanced expression of a toabcco cysteine protease gene. Plant Mol. Biol 2000, 44, 649–657. [Google Scholar]
- Chen, G.H.; Huang, L.T.; Yap, M.N.; Lee, R.H.; Huang, Y.J.; Cheng, M.C.; Chen, S.C.G. Molecular characterization of a senescence-associated gene encoding cysteine proteinase and its gene expression during leaf senescence in sweet potato. Plant Cell Physiol 2002, 43, 984–991. [Google Scholar]
- Liu, Y.; Schiff, M.; Dinesh-Kumar, S.P. Virus-induced gene silencing in tomato. Plant J 2002, 31, 777–786. [Google Scholar]
- Brigneti, G.; Martin-Hernandez, A.; Jin, H.; Chen, J.; Baulcombe, D.C.; Baker, B.; Jones, J.D. Virus-induced gene silencing in Solanum species. Plant J 2004, 39, 264–272. [Google Scholar]
- CM334 Genome Assembly, Pseudomolecules, Annotations and C. chinense Genome Assembly. Available online: http://peppergenome.snu.ac.kr/download.php access on 19 January 2014.
- Callis, J. Regulation of protein degradation. Plant Cell 1995, 7, 845–857. [Google Scholar]
- Srivalli, B.; Bharti, S.; Khanna-Chopra, R. Vacuole cysteine proteases and ribulose-1,5-bisphosphate carboxylase/oxygenase degradation during monocarpic senescence in cowpea leaves. Photosynthetica 2001, 39, 87–93. [Google Scholar]
- Martínez, D.E.; Bartoli, C.G.; Grbic, V.; Guiamet, J.J. Vacuolar cysteine proteases of wheat (Triticum aestivum L.) are common to leaf senescence induced by different factors. J. Exp. Bot 2007, 58, 1099–1107. [Google Scholar]
- Prins, A.; van Heerden, P.D.R.; Olmos, E.; Kunert, K.J.; Foyer, C.H. Cysteine proteinases regulate chloroplast protein content and composition in tobacco leaves: A model for dynamic interactions with ribulose-1,5-bisphosphate carboxylase/oxygenase (Rubisco)vesicular bodies. J. Exp. Bot 2008, 59, 1935–1950. [Google Scholar]
- Kong, Z.; Li, M.; Yang, W.; Xu, W.; Xue, Y. A novel nuclear-localized CCCH-type zinc finger protein, OsDOS, is involved in delaying leaf senescence in rice (Oryza sativa L.). Plant Physiol 2006, 141, 1376–1388. [Google Scholar]
- Morris, K.; Mackerness, S.A.; Page, T.; John, C.F.; Murphy, A.M.; Carr, J.P.; Buchanan-Wollaston, V. Salicylic acid has a role in regulating gene expression during senescence. Plant J 2000, 23, 677–685. [Google Scholar]
- Gepstein, S.; Thimann, K.V. Changes in the abscisic acid content of oat leaves during senescence. Proc. Natl. Acad. Sci. USA 1980, 77, 2050–2053. [Google Scholar]
- Rosenvasser, S.; Mayak, S.; Friedman, H. Increase in reactive oxygen species (ROS) and in senescence-associated gene transcript (SAG) levels during dark-induced senescence of Pelargonium cuttings, and the effect of gibberellic acid. Plant Sci 2006, 170, 873–879. [Google Scholar]
- Vanacker, H.; Sandalio, L.M.; Jimenez, A.; Palma, J.M.; Corpas, F.J.; Meseguer, V.; Gómez, M.; Sevilla, F.; Leterrier, M.; Foyer, C.H.; et al. Role of redox regulation in leaf senescence of pea plants grown in different sources of nitrogen nutrition. J. Exp. Bot 2006, 57, 1735–1745. [Google Scholar]
- Chopra, K.R. Leaf senescence and abiotic stresses share reactive oxygen species-mediated chloroplast degradation. Protoplasma 2012, 249, 469–481. [Google Scholar]
- Solomon, M.; Belenghi, B.; Delledonne, M.; Menachem, E.; Levine, A. The involvement of cysteine proteases and protease inhibitor genes in the regulation of programmed cell death in plants. Plant Cell 1999, 11, 431–444. [Google Scholar]
- Palma, M.J.; Sandalio, M.L.; Javier, C.F. Plant proteases, protein degradation, and oxidative stress: role of peroxisomes. Plant Physiol. Biochem 2002, 40, 521–530. [Google Scholar]
- De, D.N. Functions of vacuoles. In Plant Cell Vacuoles: An Introduction; CSIRO Publishing: Collingwood, Australia, 2000; Volume 5, pp. 163–249. [Google Scholar]
- Canut, H.; Alibert, G.; Carrasco, A.; Boudet, A.M. Rapid degradation of abnormal proteins in vacuoles from Acer pseudoplatanus L. cells. Plant Physiol 1986, 81, 460–463. [Google Scholar]
- Zhang, X.; Zhang, Z.; Li, J.; Wu, L.; Guo, J.; Ouyang, L.; Xia, Y.; Huang, X.; Pang, X. Correlation of leaf senescence and gene expression/activities of chlorophyll degradation enzymes in harvested Chinese flowering cabbage (Brassica rapa var. parachinensis). J. Plant Physiol 2011, 168, 2081–2087. [Google Scholar]
- Abreu, M.E.; Munné-Bosch, S. Salicylic acid may be involved in the regulation of drought-induced leaf senescence in perennials: A case study in field-grown Salvia officinalis L. plants. Environ. Exp. Bot 2008, 64, 105–112. [Google Scholar]
- Wang, P.; Yin, L.H.; Liang, D.; Li, C.; Ma, F.W.; Yue, Z.Y. Delayed senescence of apple leaves by exogenous melatonin treatment: Toward regulating the ascorbate–glutathione cycle. J. Pineal Res 2012, 53, 11–20. [Google Scholar]
- Otegui, M.S.; Noh, Y.S.; Martínez, D.E.; Petroff, M.G.V.; Staehelin, L.A.; Amasino, R.M.; Guiamet, J.J. Senescence-associated vacuoles with intense proteolytic activity develop in leaves of Arabidopsis and soybean. Plant J 2005, 41, 831–844. [Google Scholar]
- Wang, J.E.; Li, D.W.; Zhang, Y.L.; Zhao, Q.; He, Y.M.; Gong, Z.H. Defence responses of pepper (Capsicum annuum L.) infected with incompatible and compatible strains of Phytophthora capsici. Eur. J. Plant Pathol 2013, 136, 625–638. [Google Scholar]
- Arkus, K.A.J.; Cahoon, E.B.; Jez, J.M. Mechanistic analysis of wheat chlorophyllase. Arch. Biochem. Biophys 2005, 438, 146–155. [Google Scholar]
- Buege, J.A.; Aust, S.D. Microsomal lipid peroxidation. Method Enzymol 1978, 52, 302–310. [Google Scholar]
- Wang, J.E.; Liu, K.K.; Li, D.W.; Zhang, Y.L.; Zhao, Q.; He, Y.M.; Gong, Z.H. A novel peroxidase CanPOD gene of pepper is involved in defense responses to Phytophtora capsici infection as well as Aabiotic stress tolerance. Int. J. Mol. Sci 2013, 14, 3158–3177. [Google Scholar]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCt Method. Methods 2001, 25, 402–408. [Google Scholar]
- Li, Z.; Wang, S.; Tao, Q.Y.; Pan, J.S.; Si, L.T.; Gong, Z.H.; Cai, R. A putative positive feedback regulation mechanism in CsACS2 expression suggests a modified model for sex determination in cucumber (Cucumis sativus L.). J. Exp. Bot 2012, 63, 4475–4484. [Google Scholar]
- Genovesi, V.; Fornalé, S.; Fry, S.C.; Ruel, K.; Ferrer, P.; Encina, A.; Sonbol, F.M.; Bosch, J.; Puigdomènech, P.; Rigau, J.; et al. ZmXTH1, a new xyloglucan endotransglucosylase/hydrolase in maize, affects cell wall structure and composition in Arabidopsis thaliana. J. Exp. Bot 2008, 59, 875–889. [Google Scholar]
| Primers | Sequence(5′–3′) |
|---|---|
| Cloning of CaCP cDNA sequence | |
| EST-CaCPF | TACAGGATATGAAGATGTCCCAGC |
| EST-CaCPR | AAGCCTCCATGGCAAGTCC |
| 5′RACE-CaCPGSP | GTTTTCACCCCATTTACTGCCCC |
| 5′RACE-CaCPNGSP | CCACTGACACTGGTTGATTTGCC |
| 3′RACE-CaCPGSP | GATATGAAGATGTCCCAGCCAAC |
| 3′RACE-CaCPNGSP | GTGGTGTATTCAGTGGATCATGC |
| Cloning of CaCP DNA sequence | |
| DNA-CaCPF | TAGTTGTTCTATAATGGCCTTTGCA |
| DNA-CaCPR | ACAATTTAGAAGCTGGCACCATT |
| quantitative real-time PCR | |
| RT-CaCPF | TTGTTCTATAATGGCCTTTGCA |
| RT-CaCPR | TCATAGTGAATTGGACGTGGTG |
| CaUBI3F | TGTCCATCTGCTCTCTGTTG |
| CaUBI3R | CACCCCAAGCACAATAAGAC |
| Virus-induced gene-silencing (VIGS) vector construction | |
| TRV-CaCPF | GAGTAGCTTCGATTTCCAGTTC |
| TRV-CaCPR | CAAATCCACAACGTATTCACAT |
| CaPDSF | TGTTGTCAAAACTCCAAGGTCTGTA |
| CaPDSR | TTTCTCCCACTTGGTTCACTCTTGT |
| PVBG2307-CaCP-GFP vector construction | |
| GFP-CaCPF | TTCTATAATGGCCTTTGCAAACC |
| GFP-CaCPR | GGCAGTGGGATAAGAAGCCTC |
© 2014 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Xiao, H.-J.; Yin, Y.-X.; Chai, W.-G.; Gong, Z.-H. Silencing of the CaCP Gene Delays Salt- and Osmotic-Induced Leaf Senescence in Capsicum annuum L. Int. J. Mol. Sci. 2014, 15, 8316-8334. https://doi.org/10.3390/ijms15058316
Xiao H-J, Yin Y-X, Chai W-G, Gong Z-H. Silencing of the CaCP Gene Delays Salt- and Osmotic-Induced Leaf Senescence in Capsicum annuum L. International Journal of Molecular Sciences. 2014; 15(5):8316-8334. https://doi.org/10.3390/ijms15058316
Chicago/Turabian StyleXiao, Huai-Juan, Yan-Xu Yin, Wei-Guo Chai, and Zhen-Hui Gong. 2014. "Silencing of the CaCP Gene Delays Salt- and Osmotic-Induced Leaf Senescence in Capsicum annuum L." International Journal of Molecular Sciences 15, no. 5: 8316-8334. https://doi.org/10.3390/ijms15058316
APA StyleXiao, H.-J., Yin, Y.-X., Chai, W.-G., & Gong, Z.-H. (2014). Silencing of the CaCP Gene Delays Salt- and Osmotic-Induced Leaf Senescence in Capsicum annuum L. International Journal of Molecular Sciences, 15(5), 8316-8334. https://doi.org/10.3390/ijms15058316




