Genetic Diversity and Population Structure of Siberian apricot (Prunus sibirica L.) in China
Abstract
:1. Introduction
2. Results and Discussion
2.1. Genetic Diversity Analysis
2.2. Genetic Differentiation Analysis
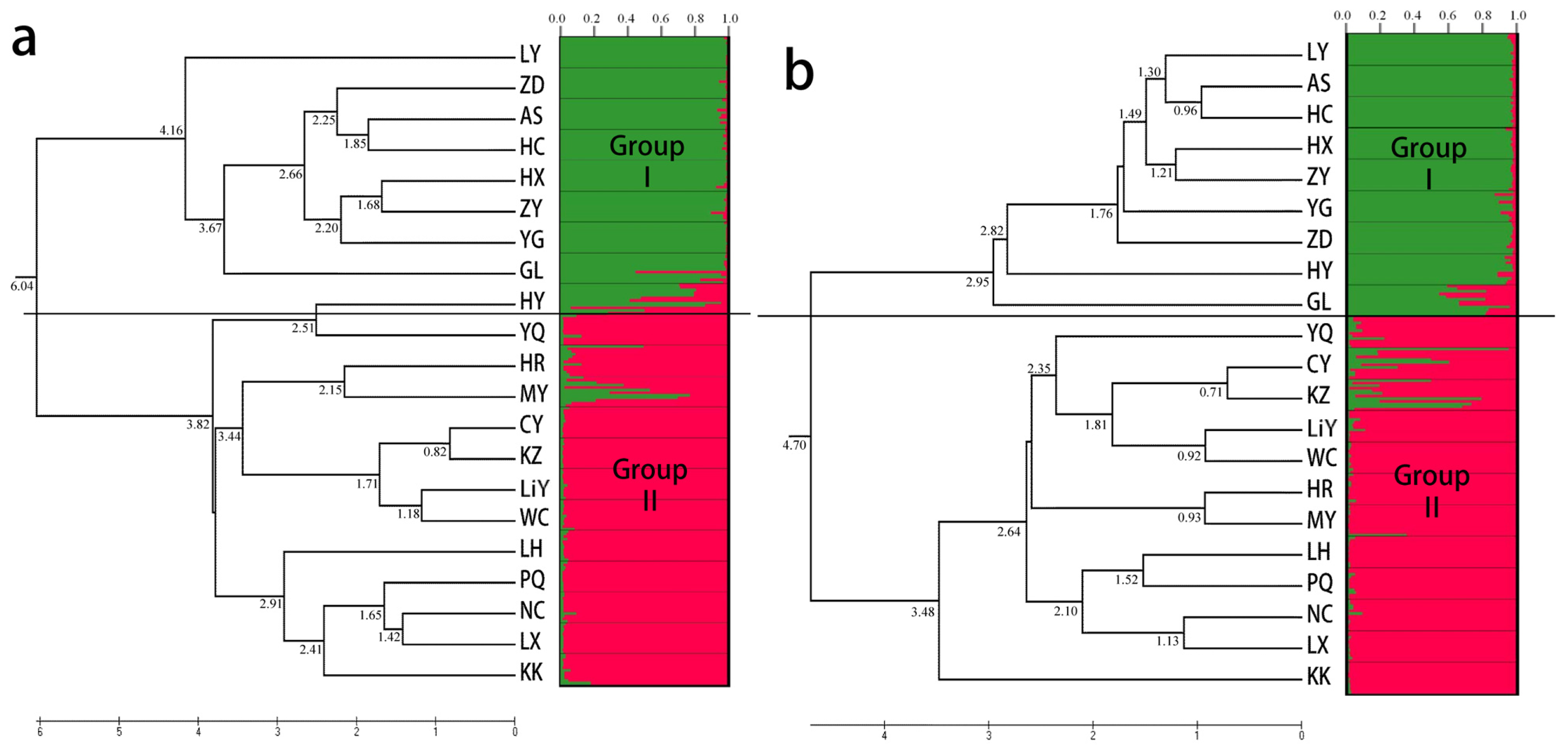
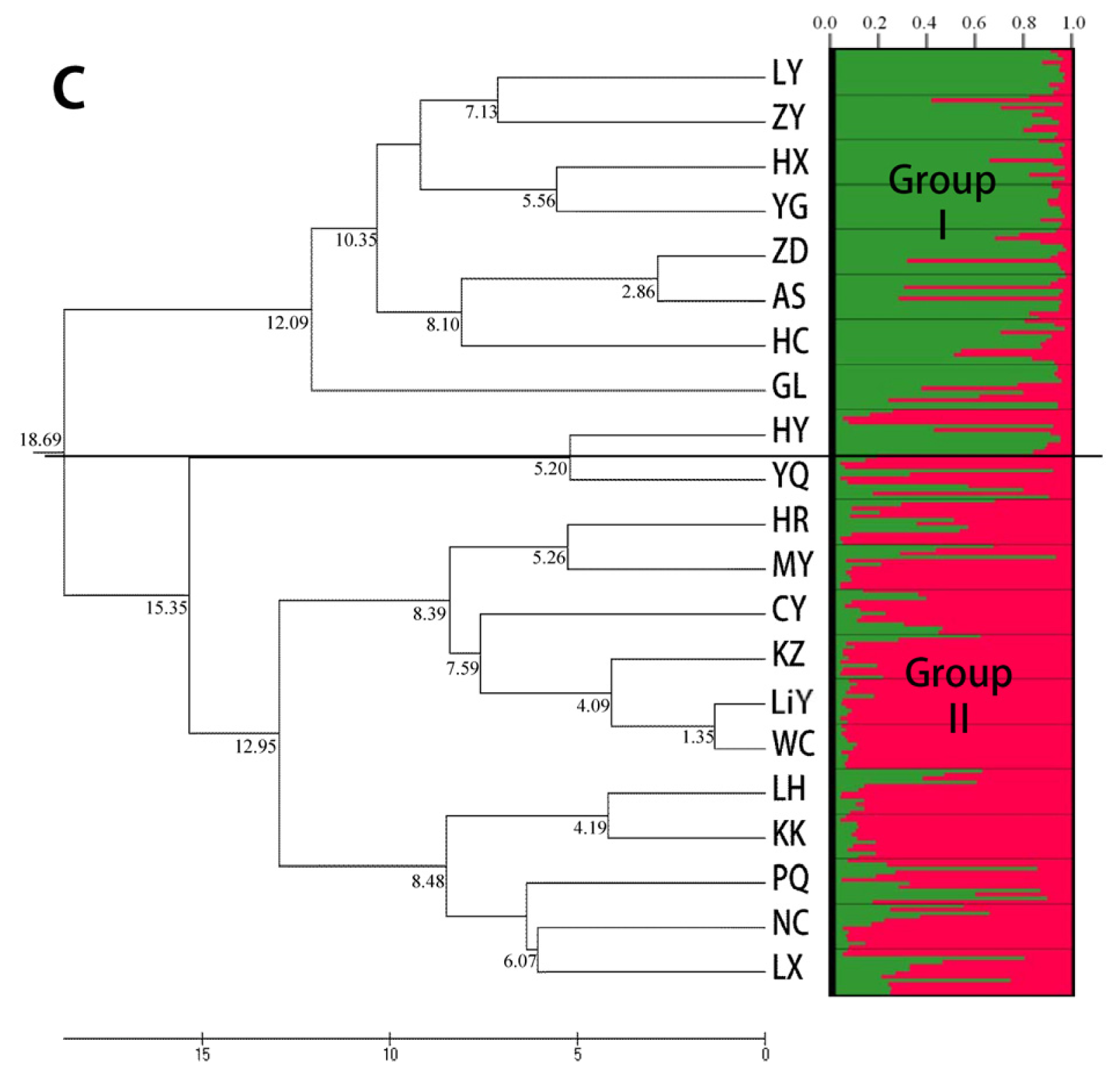
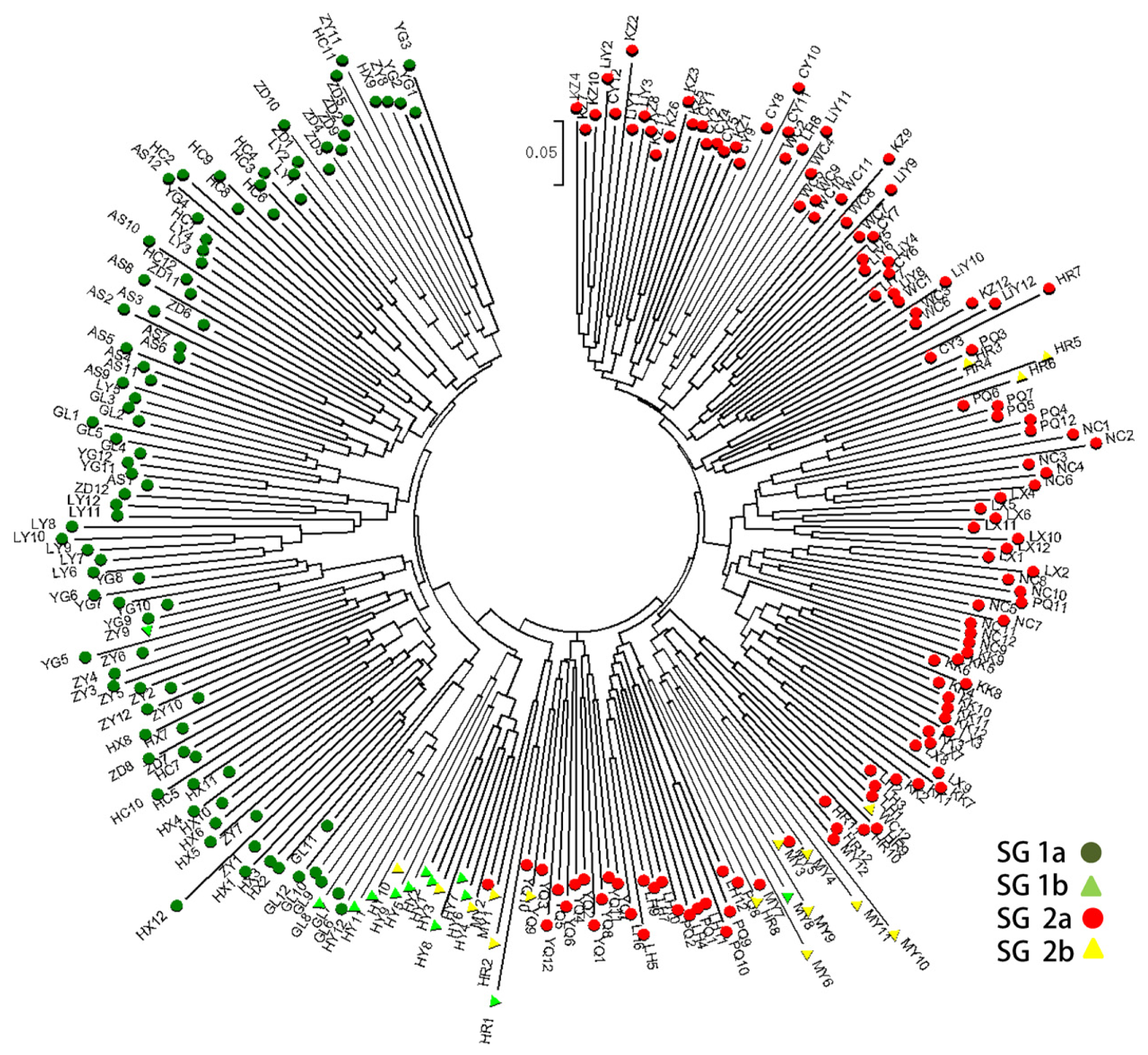
2.3. Population Structure and Cluster Analysis
2.4. Comparison of ISSR, SSR, and SRAP Markers
2.5. Combined Analysis
2.6. Conservation Considerations
3. Experimental Section
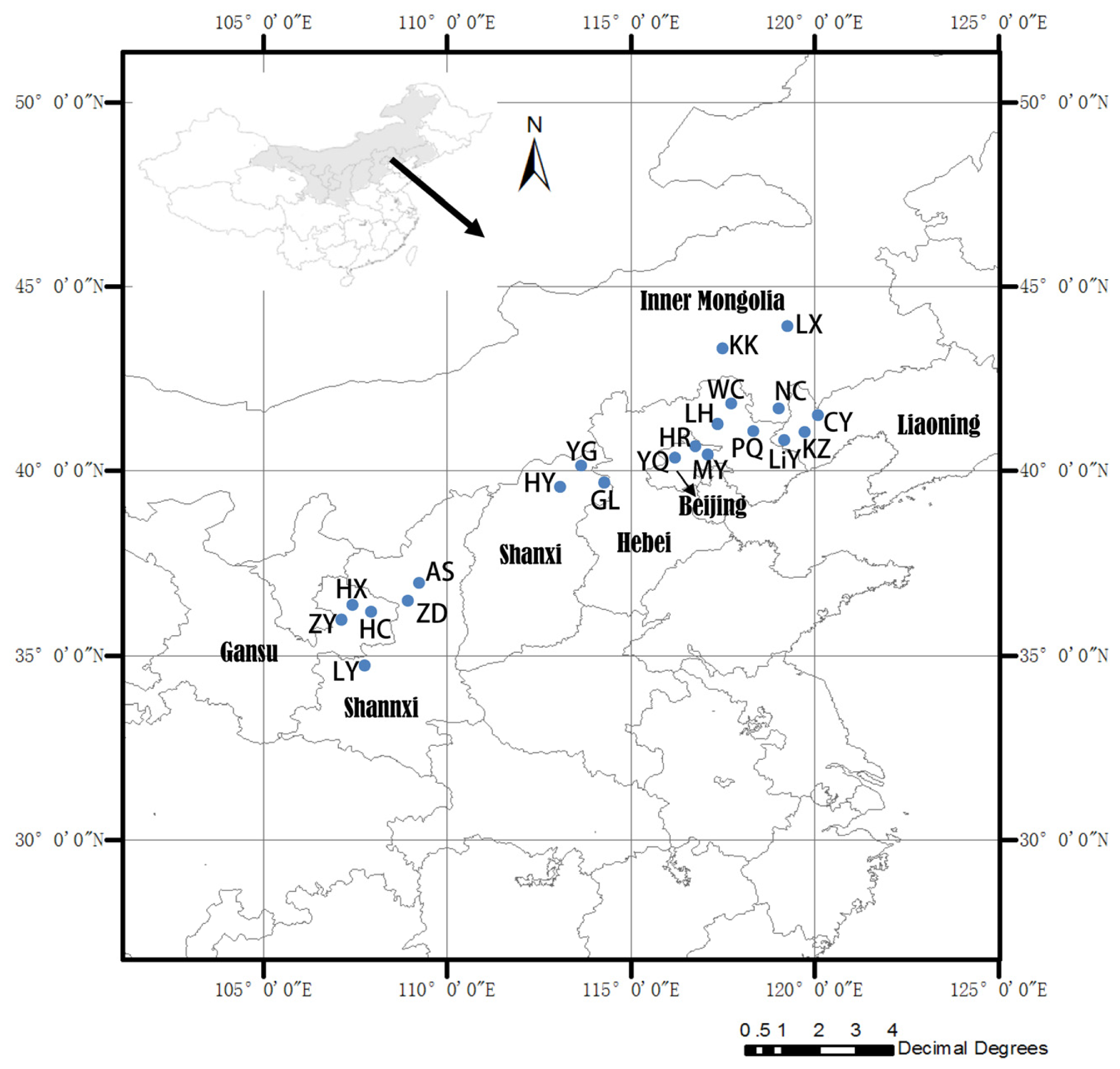
3.1. Plant Materials
3.2. DNA Extraction
3.3. DNA Amplification
3.4. Data Analysis
4. Conclusions
Supplementary Information
| Primer | Annealing temp (°C) | Total bands | Polymorphism bands | Sequence (5′–3′) | Reference |
|---|---|---|---|---|---|
| BC807 | 50 | 10 | 10 | (AG)8T | UBC Primer Set #9 [61] |
| BC818 | 52 | 9 | 9 | (CA)8G | UBC Primer Set #9 [61] |
| BC827 | 51 | 10 | 10 | (AC)8G | UBC Primer Set #9 [61] |
| BC835 | 54 | 15 | 15 | (AG)8YC | UBC Primer Set #9 [61] |
| BC843 | 50 | 8 | 7 | (CT)8GA | UBC Primer Set #9 [61] |
| BC847 | 54 | 13 | 13 | (CA)8RC | UBC Primer Set #9 [61] |
| BC868 | 48 | 8 | 6 | (GAA)6 | UBC Primer Set #9 [61] |
| BC873 | 49 | 11 | 9 | (GACA)4 | UBC Primer Set #9 [61] |
| BC880 | 48 | 8 | 7 | (GGAGA)3 | UBC Primer Set #9 [61] |
| BC888 | 50 | 11 | 8 | BDB(CA)7 | UBC Primer Set #9 [61] |
| Me1/Em1 | 50 | 15 | 14 | TGAGTCCAAACCGGAGC/GACTGCGTACGAATTTGC | G. Li et al. [16] |
| Me1/Em4 | 50 | 9 | 6 | TGAGTCCAAACCGGAGC/GACTGCGTACGAATTGAG | G. Li et al. [16] |
| Me2/Em1 | 50 | 9 | 6 | TGAGTCCAAACCGGACC/GACTGCGTACGAATTTGC | G. Li et al. [16] |
| Me2/Em3 | 50 | 12 | 11 | TGAGTCCAAACCGGACC/GACTGCGTACGAATTAAC | G. Li et al. [16] |
| Me2/Em9 | 50 | 10 | 6 | TGAGTCCAAACCGGACC/GACTGCGTACGAATTATT | PF. Ai et al. [24] |
| Me4/Em7 | 50 | 11 | 8 | TGAGTCCAAACCGGTCC/GACTGCGTACGAATTGCA | PF. Ai et al. [24] |
| Me5/Em2 | 50 | 9 | 8 | TGAGTCCAAACCGGTGC/GACTGCGTACGAATTTGA | G. Li et al. [16] |
| Me8/Em9 | 50 | 9 | 8 | TGAGTCCAAACCGGAAG/GACTGCGTACGAATTATT | PF. Ai et al. [24] |
| Me1/Em6 | 50 | 19 | 17 | TGAGTCCAAACCGGAGC/GACTGCGTACGAATTCTT | PF. Ai et al. [24] |
| Me8/Em8 | 50 | 17 | 15 | TGAGTCCAAACCGGAAG/GACTGCGTACGAATTGCC | PF. Ai et al. [24] |
| Primer | Reference | SSR motive | Annealing temp (°C) | Observed alleles | Effective alleles | Shannon’s index | Observed heterozygosity | Expected heterozygosity | Genetic differentiation coefficient | Gene flow |
|---|---|---|---|---|---|---|---|---|---|---|
| AMPA101 | Hagen et al. [51] | (TC)11(AC)12 | 56 | 5 | 4.7825 | 1.5882 | 0.5119 | 0.7925 | 0.1268 | 1.7221 |
| AMPA119 | Hagen et al. [51] | (TA)9 | 57 | 7 | 5.6433 | 1.8253 | 0.5714 | 0.8244 | 0.1964 | 1.0230 |
| BPPCT039 | Dirlewanger et al. [50] | (GA)20 | 55 | 5 | 4.4628 | 1.5488 | 0.4444 | 0.7775 | 0.1223 | 1.7946 |
| pchgms3 | Sosinski et al. [54] | (CT)19 | 57 | 5 | 3.5636 | 1.3522 | 0.4921 | 0.7208 | 0.1726 | 1.1986 |
| pchgms5 | Sosinski et al. [54] | (CA)9(TA)8 | 51 | 2 | 1.9027 | 0.6673 | 0.5357 | 0.4754 | 0.0872 | 2.6160 |
| ssrPaCITA23 | Lopes et al. [52] | (AC)2(AG)18 | 51 | 6 | 4.5883 | 1.6390 | 0.8651 | 0.7836 | 0.0357 | 6.7527 |
| UDAp-414 | Messina et al. [53] | (AG)21 | 56 | 4 | 3.3321 | 1.2652 | 0.1944 | 0.7013 | 0.3336 | 0.4995 |
| UDAp-415 | Messina et al. [53] | (GA)21 | 56 | 4 | 3.6956 | 1.3480 | 0.4683 | 0.7309 | 0.1658 | 1.2577 |
| UDAp-420 | Messina et al. [53] | (CT)20 | 56 | 5 | 3.9018 | 1.3908 | 0.6429 | 0.7452 | 0.1196 | 1.8403 |
| UDP96-001 | Cipriani et al. [49] | (CA)17 | 57 | 2 | 1.8529 | 0.6529 | 0.2500 | 0.4612 | 0.1871 | 1.0860 |
| Mean | - | - | - | 4.5 | 3.7726 | 1.3278 | 0.4976 | 0.7013 | 0.1543 | 1.3706 |
| POP | LY | ZD | AS | HC | HX | ZY | YG | GL | HY | YQ | HR | MY | CY | KZ | LiY | WC | LH | PQ | NC | LX | KK |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| LY | - | 0.9246 | 0.9248 | 0.9363 | 0.9112 | 0.9125 | 0.9192 | 0.9124 | 0.8673 | 0.8438 | 0.8695 | 0.8911 | 0.8425 | 0.8553 | 0.8775 | 0.8641 | 0.8645 | 0.8543 | 0.8437 | 0.8394 | 0.8399 |
| ZD | 0.0784 | - | 0.9505 | 0.9616 | 0.9528 | 0.9509 | 0.9466 | 0.9309 | 0.9161 | 0.8839 | 0.8925 | 0.9205 | 0.8779 | 0.8758 | 0.8888 | 0.8807 | 0.8861 | 0.8807 | 0.8697 | 0.8707 | 0.8688 |
| AS | 0.0782 | 0.0507 | - | 0.9637 | 0.9426 | 0.9406 | 0.9429 | 0.9239 | 0.9017 | 0.8797 | 0.8999 | 0.9282 | 0.8969 | 0.8903 | 0.9067 | 0.8948 | 0.8865 | 0.8949 | 0.8735 | 0.8799 | 0.8708 |
| HC | 0.0658 | 0.0391 | 0.037 | - | 0.9519 | 0.9467 | 0.9589 | 0.9306 | 0.9089 | 0.8837 | 0.8942 | 0.9317 | 0.8784 | 0.879 | 0.9068 | 0.8986 | 0.8997 | 0.8962 | 0.8914 | 0.8769 | 0.8717 |
| HX | 0.093 | 0.0483 | 0.0591 | 0.0492 | - | 0.967 | 0.9592 | 0.9324 | 0.939 | 0.8795 | 0.8992 | 0.9092 | 0.8613 | 0.8689 | 0.8749 | 0.8798 | 0.8786 | 0.8722 | 0.8815 | 0.878 | 0.8831 |
| ZY | 0.0916 | 0.0503 | 0.0612 | 0.0548 | 0.0336 | - | 0.9549 | 0.9139 | 0.9186 | 0.8872 | 0.8951 | 0.9167 | 0.8921 | 0.8922 | 0.9032 | 0.9093 | 0.8928 | 0.8806 | 0.8733 | 0.8812 | 0.884 |
| YG | 0.0843 | 0.0549 | 0.0588 | 0.042 | 0.0417 | 0.0461 | - | 0.9438 | 0.9293 | 0.8793 | 0.896 | 0.9114 | 0.8726 | 0.8659 | 0.8879 | 0.9 | 0.8942 | 0.885 | 0.8782 | 0.8762 | 0.878 |
| GL | 0.0917 | 0.0716 | 0.0792 | 0.0719 | 0.07 | 0.0901 | 0.0578 | - | 0.9406 | 0.8989 | 0.8978 | 0.911 | 0.8732 | 0.8875 | 0.8876 | 0.8898 | 0.8924 | 0.8868 | 0.8661 | 0.8805 | 0.8994 |
| HY | 0.1424 | 0.0876 | 0.1035 | 0.0955 | 0.063 | 0.0849 | 0.0733 | 0.0612 | - | 0.9511 | 0.9133 | 0.9248 | 0.8989 | 0.9054 | 0.9022 | 0.9092 | 0.9108 | 0.9178 | 0.9036 | 0.9123 | 0.9232 |
| YQ | 0.1698 | 0.1234 | 0.1282 | 0.1237 | 0.1284 | 0.1197 | 0.1286 | 0.1066 | 0.0502 | - | 0.9397 | 0.9441 | 0.9452 | 0.956 | 0.9424 | 0.9461 | 0.9341 | 0.9477 | 0.9244 | 0.9386 | 0.9471 |
| HR | 0.1398 | 0.1137 | 0.1055 | 0.1119 | 0.1063 | 0.1108 | 0.1098 | 0.1078 | 0.0906 | 0.0622 | - | 0.9578 | 0.9216 | 0.9251 | 0.9376 | 0.934 | 0.9204 | 0.933 | 0.9169 | 0.9339 | 0.9183 |
| MY | 0.1153 | 0.0829 | 0.0745 | 0.0708 | 0.0952 | 0.087 | 0.0928 | 0.0932 | 0.0781 | 0.0575 | 0.0431 | - | 0.939 | 0.9334 | 0.9461 | 0.9318 | 0.92 | 0.9411 | 0.929 | 0.9283 | 0.9145 |
| CY | 0.1714 | 0.1302 | 0.1088 | 0.1296 | 0.1493 | 0.1141 | 0.1363 | 0.1356 | 0.1066 | 0.0564 | 0.0817 | 0.0629 | - | 0.9837 | 0.9677 | 0.9665 | 0.9272 | 0.9262 | 0.895 | 0.926 | 0.9187 |
| KZ | 0.1563 | 0.1326 | 0.1161 | 0.1289 | 0.1406 | 0.1141 | 0.144 | 0.1194 | 0.0994 | 0.045 | 0.0779 | 0.0689 | 0.0164 | - | 0.968 | 0.9635 | 0.9335 | 0.9338 | 0.9112 | 0.9296 | 0.935 |
| LiY | 0.1307 | 0.1178 | 0.0979 | 0.0978 | 0.1336 | 0.1018 | 0.1189 | 0.1192 | 0.1029 | 0.0593 | 0.0645 | 0.0554 | 0.0329 | 0.0325 | - | 0.9768 | 0.9427 | 0.9425 | 0.9128 | 0.9342 | 0.9258 |
| WC | 0.146 | 0.127 | 0.1111 | 0.1069 | 0.128 | 0.0951 | 0.1053 | 0.1168 | 0.0952 | 0.0554 | 0.0682 | 0.0706 | 0.0341 | 0.0372 | 0.0235 | - | 0.9584 | 0.9422 | 0.901 | 0.9286 | 0.9377 |
| LH | 0.1457 | 0.1209 | 0.1205 | 0.1057 | 0.1295 | 0.1134 | 0.1118 | 0.1139 | 0.0935 | 0.0682 | 0.083 | 0.0834 | 0.0755 | 0.0688 | 0.059 | 0.0425 | - | 0.9587 | 0.9282 | 0.9422 | 0.9448 |
| PQ | 0.1575 | 0.127 | 0.111 | 0.1096 | 0.1368 | 0.1272 | 0.1221 | 0.1201 | 0.0858 | 0.0537 | 0.0693 | 0.0607 | 0.0767 | 0.0685 | 0.0592 | 0.0595 | 0.0421 | - | 0.9688 | 0.9662 | 0.9508 |
| NC | 0.1699 | 0.1396 | 0.1352 | 0.1149 | 0.1261 | 0.1354 | 0.1299 | 0.1438 | 0.1013 | 0.0786 | 0.0867 | 0.0737 | 0.1109 | 0.0929 | 0.0912 | 0.1043 | 0.0745 | 0.0317 | - | 0.9721 | 0.9388 |
| LX | 0.1751 | 0.1385 | 0.128 | 0.1313 | 0.1301 | 0.1264 | 0.1322 | 0.1272 | 0.0918 | 0.0633 | 0.0684 | 0.0744 | 0.0769 | 0.073 | 0.068 | 0.0741 | 0.0595 | 0.0343 | 0.0283 | - | 0.9693 |
| KK | 0.1744 | 0.1406 | 0.1383 | 0.1373 | 0.1243 | 0.1233 | 0.1301 | 0.106 | 0.0799 | 0.0543 | 0.0852 | 0.0894 | 0.0848 | 0.0673 | 0.0771 | 0.0643 | 0.0568 | 0.0504 | 0.0632 | 0.0311 | - |
| POP | LY | ZD | AS | HC | HX | ZY | YG | GL | HY | YQ | HR | MY | CY | KZ | LiY | WC | LH | PQ | NC | LX | KK |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| LY | - | 0.9636 | 0.9757 | 0.9729 | 0.9744 | 0.9718 | 0.961 | 0.9439 | 0.947 | 0.9011 | 0.9272 | 0.9497 | 0.9358 | 0.9238 | 0.9258 | 0.9127 | 0.8983 | 0.9065 | 0.9186 | 0.9033 | 0.873 |
| ZD | 0.0371 | - | 0.974 | 0.9716 | 0.9536 | 0.9688 | 0.9606 | 0.9294 | 0.9374 | 0.9001 | 0.9046 | 0.9227 | 0.9168 | 0.8995 | 0.9044 | 0.8971 | 0.8838 | 0.8919 | 0.9044 | 0.8901 | 0.8613 |
| AS | 0.0246 | 0.0263 | - | 0.981 | 0.9692 | 0.9699 | 0.9622 | 0.937 | 0.9487 | 0.9118 | 0.9274 | 0.9419 | 0.9337 | 0.9088 | 0.9123 | 0.907 | 0.8865 | 0.8976 | 0.9079 | 0.892 | 0.8553 |
| HC | 0.0275 | 0.0288 | 0.0192 | - | 0.9684 | 0.9701 | 0.9726 | 0.9428 | 0.9477 | 0.9111 | 0.9242 | 0.9388 | 0.9303 | 0.9103 | 0.9132 | 0.9032 | 0.8955 | 0.9043 | 0.9188 | 0.8893 | 0.8664 |
| HX | 0.0259 | 0.0475 | 0.0312 | 0.0321 | - | 0.9762 | 0.9653 | 0.9527 | 0.9422 | 0.9153 | 0.9363 | 0.9445 | 0.932 | 0.9168 | 0.9083 | 0.9024 | 0.892 | 0.8908 | 0.9122 | 0.8978 | 0.8708 |
| ZY | 0.0286 | 0.0317 | 0.0306 | 0.0304 | 0.0241 | - | 0.9714 | 0.951 | 0.9439 | 0.9213 | 0.9295 | 0.9415 | 0.9264 | 0.9212 | 0.9119 | 0.9012 | 0.9008 | 0.8978 | 0.911 | 0.901 | 0.8791 |
| YG | 0.0398 | 0.0402 | 0.0385 | 0.0278 | 0.0354 | 0.029 | - | 0.9482 | 0.9491 | 0.906 | 0.9295 | 0.9427 | 0.9201 | 0.9063 | 0.8994 | 0.8897 | 0.8883 | 0.8931 | 0.9006 | 0.893 | 0.8594 |
| GL | 0.0577 | 0.0732 | 0.065 | 0.059 | 0.0485 | 0.0502 | 0.0531 | - | 0.9361 | 0.9197 | 0.9323 | 0.9406 | 0.9416 | 0.9234 | 0.905 | 0.9046 | 0.9134 | 0.8904 | 0.8958 | 0.8842 | 0.8719 |
| HY | 0.0545 | 0.0646 | 0.0526 | 0.0537 | 0.0595 | 0.0578 | 0.0522 | 0.066 | - | 0.946 | 0.9369 | 0.9547 | 0.953 | 0.9323 | 0.9196 | 0.928 | 0.9407 | 0.9431 | 0.9226 | 0.912 | 0.8855 |
| YQ | 0.1042 | 0.1052 | 0.0923 | 0.0931 | 0.0885 | 0.082 | 0.0988 | 0.0837 | 0.0555 | - | 0.9365 | 0.9345 | 0.9648 | 0.9639 | 0.9406 | 0.9472 | 0.9437 | 0.9275 | 0.9229 | 0.931 | 0.9281 |
| HR | 0.0755 | 0.1003 | 0.0754 | 0.0789 | 0.0658 | 0.0731 | 0.0731 | 0.0701 | 0.0652 | 0.0656 | - | 0.9817 | 0.9609 | 0.9564 | 0.9469 | 0.941 | 0.938 | 0.9396 | 0.9498 | 0.9395 | 0.9112 |
| MY | 0.0517 | 0.0804 | 0.0599 | 0.0632 | 0.0572 | 0.0602 | 0.059 | 0.0612 | 0.0464 | 0.0678 | 0.0185 | - | 0.9653 | 0.9584 | 0.9541 | 0.9423 | 0.9449 | 0.9627 | 0.9616 | 0.9423 | 0.9099 |
| CY | 0.0664 | 0.0869 | 0.0686 | 0.0722 | 0.0705 | 0.0764 | 0.0833 | 0.0602 | 0.0481 | 0.0359 | 0.0398 | 0.0353 | - | 0.9859 | 0.9649 | 0.9625 | 0.958 | 0.9551 | 0.9521 | 0.9462 | 0.9235 |
| KZ | 0.0792 | 0.1059 | 0.0957 | 0.094 | 0.0868 | 0.0821 | 0.0984 | 0.0797 | 0.0701 | 0.0368 | 0.0446 | 0.0425 | 0.0142 | - | 0.9702 | 0.9599 | 0.9498 | 0.9515 | 0.9462 | 0.9471 | 0.9325 |
| LiY | 0.0771 | 0.1005 | 0.0918 | 0.0908 | 0.0962 | 0.0922 | 0.1061 | 0.0998 | 0.0838 | 0.0613 | 0.0545 | 0.047 | 0.0357 | 0.0302 | - | 0.9817 | 0.9482 | 0.9568 | 0.9528 | 0.9547 | 0.9272 |
| WC | 0.0914 | 0.1086 | 0.0976 | 0.1018 | 0.1027 | 0.104 | 0.1168 | 0.1002 | 0.0747 | 0.0543 | 0.0608 | 0.0595 | 0.0382 | 0.0409 | 0.0184 | - | 0.9611 | 0.967 | 0.9573 | 0.9559 | 0.943 |
| LH | 0.1072 | 0.1235 | 0.1205 | 0.1103 | 0.1143 | 0.1045 | 0.1185 | 0.0906 | 0.0611 | 0.058 | 0.064 | 0.0567 | 0.0429 | 0.0515 | 0.0532 | 0.0397 | - | 0.9701 | 0.9469 | 0.9547 | 0.9466 |
| PQ | 0.0982 | 0.1144 | 0.108 | 0.1006 | 0.1156 | 0.1079 | 0.113 | 0.1161 | 0.0585 | 0.0752 | 0.0623 | 0.038 | 0.0459 | 0.0497 | 0.0441 | 0.0336 | 0.0304 | - | 0.9718 | 0.9621 | 0.9281 |
| NC | 0.0849 | 0.1005 | 0.0966 | 0.0847 | 0.0919 | 0.0932 | 0.1047 | 0.11 | 0.0806 | 0.0803 | 0.0515 | 0.0392 | 0.0491 | 0.0553 | 0.0484 | 0.0436 | 0.0545 | 0.0286 | - | 0.9777 | 0.9422 |
| LX | 0.1017 | 0.1164 | 0.1143 | 0.1173 | 0.1078 | 0.1043 | 0.1132 | 0.1231 | 0.0921 | 0.0715 | 0.0624 | 0.0594 | 0.0553 | 0.0544 | 0.0464 | 0.0451 | 0.0463 | 0.0387 | 0.0225 | - | 0.9706 |
| KK | 0.1359 | 0.1493 | 0.1563 | 0.1434 | 0.1383 | 0.1289 | 0.1515 | 0.1371 | 0.1216 | 0.0746 | 0.093 | 0.0944 | 0.0796 | 0.0699 | 0.0756 | 0.0587 | 0.0549 | 0.0746 | 0.0595 | 0.0298 | - |
| POP | LY | ZD | AS | HC | HX | ZY | YG | GL | HY | YQ | HR | MY | CY | KZ | LiY | WC | LH | PQ | NC | LX | KK |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| LY | - | 0.8003 | 0.8311 | 0.7872 | 0.8099 | 0.8672 | 0.809 | 0.729 | 0.6243 | 0.6804 | 0.6218 | 0.6774 | 0.677 | 0.6046 | 0.546 | 0.545 | 0.6504 | 0.7714 | 0.7101 | 0.7559 | 0.6019 |
| ZD | 0.2228 | - | 0.9443 | 0.8546 | 0.8016 | 0.8244 | 0.8604 | 0.8024 | 0.7017 | 0.6287 | 0.6696 | 0.6399 | 0.7268 | 0.6782 | 0.6801 | 0.6508 | 0.656 | 0.6962 | 0.5933 | 0.6535 | 0.617 |
| AS | 0.185 | 0.0573 | - | 0.8464 | 0.8379 | 0.8475 | 0.9035 | 0.83 | 0.7434 | 0.6438 | 0.6488 | 0.7216 | 0.8105 | 0.6975 | 0.6991 | 0.6934 | 0.7128 | 0.7846 | 0.702 | 0.6994 | 0.6562 |
| HC | 0.2392 | 0.1571 | 0.1668 | - | 0.7722 | 0.7341 | 0.7711 | 0.6664 | 0.6516 | 0.6127 | 0.7047 | 0.7677 | 0.7633 | 0.7042 | 0.7333 | 0.6802 | 0.6064 | 0.7278 | 0.6794 | 0.5961 | 0.5919 |
| HX | 0.2108 | 0.2212 | 0.1769 | 0.2585 | - | 0.827 | 0.8947 | 0.7989 | 0.747 | 0.682 | 0.7273 | 0.6751 | 0.7476 | 0.6368 | 0.5905 | 0.577 | 0.762 | 0.7316 | 0.6233 | 0.7329 | 0.6726 |
| ZY | 0.1425 | 0.1931 | 0.1655 | 0.3091 | 0.1899 | - | 0.8851 | 0.8579 | 0.7143 | 0.7104 | 0.7548 | 0.7583 | 0.7573 | 0.6788 | 0.6487 | 0.6829 | 0.8021 | 0.7641 | 0.7384 | 0.808 | 0.7918 |
| YG | 0.2119 | 0.1504 | 0.1015 | 0.26 | 0.1112 | 0.122 | - | 0.8299 | 0.8099 | 0.6677 | 0.7073 | 0.7239 | 0.7122 | 0.5829 | 0.6231 | 0.5759 | 0.7519 | 0.7415 | 0.7034 | 0.6997 | 0.696 |
| GL | 0.3161 | 0.2201 | 0.1863 | 0.4059 | 0.2245 | 0.1533 | 0.1864 | - | 0.6919 | 0.7172 | 0.7896 | 0.7092 | 0.7029 | 0.6491 | 0.667 | 0.6641 | 0.8048 | 0.7257 | 0.6277 | 0.7291 | 0.7736 |
| HY | 0.4712 | 0.3543 | 0.2965 | 0.4284 | 0.2917 | 0.3365 | 0.2109 | 0.3683 | - | 0.9012 | 0.7513 | 0.7189 | 0.766 | 0.7634 | 0.6967 | 0.6836 | 0.7802 | 0.6988 | 0.6088 | 0.5735 | 0.7292 |
| YQ | 0.3851 | 0.4641 | 0.4404 | 0.4898 | 0.3827 | 0.3419 | 0.404 | 0.3324 | 0.1041 | - | 0.8894 | 0.7893 | 0.807 | 0.8517 | 0.7448 | 0.7605 | 0.8324 | 0.7818 | 0.6413 | 0.6261 | 0.7811 |
| HR | 0.4752 | 0.4011 | 0.4326 | 0.35 | 0.3184 | 0.2813 | 0.3463 | 0.2362 | 0.2859 | 0.1172 | - | 0.9001 | 0.823 | 0.8558 | 0.8292 | 0.8372 | 0.8457 | 0.7507 | 0.6371 | 0.6412 | 0.7849 |
| MY | 0.3895 | 0.4465 | 0.3263 | 0.2644 | 0.393 | 0.2766 | 0.3231 | 0.3436 | 0.33 | 0.2366 | 0.1053 | - | 0.8536 | 0.8227 | 0.8858 | 0.8572 | 0.8454 | 0.8471 | 0.8346 | 0.7321 | 0.7997 |
| CY | 0.3901 | 0.3191 | 0.2102 | 0.2702 | 0.2909 | 0.2781 | 0.3394 | 0.3525 | 0.2666 | 0.2144 | 0.1948 | 0.1583 | - | 0.8832 | 0.8515 | 0.8431 | 0.8401 | 0.8129 | 0.7762 | 0.7455 | 0.8575 |
| KZ | 0.5033 | 0.3883 | 0.3603 | 0.3507 | 0.4514 | 0.3875 | 0.5397 | 0.4322 | 0.27 | 0.1605 | 0.1557 | 0.1952 | 0.1242 | - | 0.9143 | 0.9286 | 0.8044 | 0.773 | 0.6944 | 0.695 | 0.7668 |
| LiY | 0.6051 | 0.3855 | 0.358 | 0.3102 | 0.5268 | 0.4327 | 0.4731 | 0.405 | 0.3615 | 0.2946 | 0.1873 | 0.1212 | 0.1607 | 0.0896 | - | 0.9734 | 0.8342 | 0.828 | 0.7771 | 0.7336 | 0.7883 |
| WC | 0.6069 | 0.4296 | 0.3661 | 0.3853 | 0.5498 | 0.3814 | 0.5519 | 0.4094 | 0.3804 | 0.2738 | 0.1777 | 0.1541 | 0.1706 | 0.0741 | 0.027 | - | 0.8601 | 0.7937 | 0.6997 | 0.6804 | 0.7524 |
| LH | 0.4301 | 0.4215 | 0.3385 | 0.5001 | 0.2718 | 0.2205 | 0.2852 | 0.2172 | 0.2482 | 0.1835 | 0.1676 | 0.1679 | 0.1742 | 0.2176 | 0.1813 | 0.1507 | - | 0.8712 | 0.8101 | 0.8705 | 0.9197 |
| PQ | 0.2595 | 0.3621 | 0.2426 | 0.3177 | 0.3125 | 0.269 | 0.2991 | 0.3206 | 0.3585 | 0.2462 | 0.2868 | 0.166 | 0.2071 | 0.2575 | 0.1888 | 0.2311 | 0.1378 | - | 0.8791 | 0.882 | 0.8061 |
| NC | 0.3423 | 0.522 | 0.3539 | 0.3865 | 0.4727 | 0.3032 | 0.3519 | 0.4657 | 0.4963 | 0.4443 | 0.4508 | 0.1808 | 0.2534 | 0.3647 | 0.2522 | 0.3571 | 0.2106 | 0.1288 | - | 0.8857 | 0.8414 |
| LX | 0.2799 | 0.4254 | 0.3575 | 0.5173 | 0.3108 | 0.2132 | 0.3571 | 0.316 | 0.556 | 0.4683 | 0.4444 | 0.3119 | 0.2937 | 0.3638 | 0.3097 | 0.3851 | 0.1387 | 0.1256 | 0.1213 | - | 0.8669 |
| KK | 0.5076 | 0.4829 | 0.4214 | 0.5245 | 0.3967 | 0.2334 | 0.3624 | 0.2568 | 0.3158 | 0.2471 | 0.2422 | 0.2235 | 0.1538 | 0.2656 | 0.2379 | 0.2845 | 0.0837 | 0.2156 | 0.1726 | 0.1428 | - |
ΔK values for different numbers of populations assumed (K) in the STRUCURE analysis.
Principal component analysis on combined three markers data sets of the entire population.
Acknowledgments
Conflicts of Interest
References
- Cohen, J.I.; Williams, J.T.; Plucknett, D.L.; Shands, H. Ex situ conservation of plant genetic resources: Global development and environmental concerns. Science 1991, 253, 866–872. [Google Scholar]
- Laidò, G.; Mangini, G.; Taranto, F.; Gadaleta, A.; Blanco, A.; Cattivelli, L.; Marone, D.; Mastrangelo, A.M.; Papa, R.; de Vita, P. Genetic diversity and population structure of tetraploid Wheats (Triticum turgidum L.) estimated by SSR, DArT and pedigree data. PLoS One 2013, 8, e67280. [Google Scholar]
- Evans, L.T. Crop Evolution, Adaptation and Yield; Cambridge University Press: London, UK, 1996. [Google Scholar]
- Sreekanth, P.M.; Balasundaran, M.; Nazeem, P.A.; Suma, T.B. Genetic diversity of nine natural Tectona grandis L.f. populations of the Western Ghats in Southern India. Conserv. Genet 2012, 13, 1409–1419. [Google Scholar]
- Wang, L.; Yu, H. Biodiesel from Siberian apricot (Prunus. sibirica L.) seed kernel oil. Bioresour. Technol 2012, 112, 355–358. [Google Scholar]
- Zhang, J.Y.; Fu, D.; Wei, Z.Z.; Zhao, H.L.; Zhang, T.H. Determination of the ability of several tree and shrub species to endure and survive extreme aridity with methods of limited areas under field condition in Horqin Sandy Land. Acta Ecol. Sin 2006. [Google Scholar] [CrossRef]
- Wei, Q.; Ma, X.; Dong, J. Preparation, chemical constituents and antimicrobial activity of pyroligneous acids from walnut tree branches. J. Anal. Appl. Pyrolysis 2010, 87, 24–28. [Google Scholar]
- Sze-Tao, K.; Sathe, S. Functional properties and in vitro digestibility of almond (Prunus. dulcis L.) protein isolate. Food Chem 2000, 69, 153–160. [Google Scholar]
- Kris-Etherton, P.M.; Zhao, G.; Binkoski, A.E.; Coval, S.M.; Etherton, T.D. The effects of nuts on coronary heart disease risk. Nutr. Rev 2001, 59, 103–111. [Google Scholar]
- Wang, L. Evaluation of Siberian apricot (Prunus. sibirica L.) germplasm variability for biodiesel properties. J. Am. Oil Chem. Soc 2012, 89, 1743–1747. [Google Scholar]
- Zhebentyayeva, T.; Ledbetter, C.; Burgos, L.; Llácer, G. Apricot. In Fruit Breeding; Springer: Berlin, Germany, 2012; pp. 415–458. [Google Scholar]
- Zhebentyayeva, T.N.; Reighard, G.L.; Lalli, D.; Gorina, V.M.; Krška, B.; Abbott, A.G. Origin of resistance to plum pox virus in apricot: What new AFLP and targeted SSR data analyses tell. Tree Genet. Genomes 2008, 4, 403–417. [Google Scholar]
- Michurin, I.V. Production of New Cultivated Varieties of Fruit Trees and Shrubs from Seed; Foreign Languages Publishing House: Moscow, Soviet Union, 1952. [Google Scholar]
- Arcade, A.; Anselin, F.; Rampant, P.F.; Lesage, M.C.; Paques, L.E.; Prat, D. Application of AFLP, RAPD and ISSR markers to genetic mapping of European and Japanese larch. Theor. Appl. Genet 2000, 100, 299–307. [Google Scholar]
- McGregor, C.; Lambert, C.; Greyling, M.; Louw, J.; Warnich, L. A comparative assessment of DNA fingerprinting techniques (RAPD, ISSR, AFLP and SSR) in tetraploid potato (Solanum tuberosum L.) germplasm. Euphytica 2000, 113, 135–144. [Google Scholar]
- Li, G.; Quiros, C.F. Sequence-related amplified polymorphism (SRAP), a new marker system based on a simple PCR reaction: Its application to mapping and gene tagging in Brassica. Theor. Appl. Genet 2001, 103, 455–461. [Google Scholar]
- Dondini, L.; Lain, O.; Geuna, F.; Banfi, R.; Gaiotti, F.; Tartarini, S.; Bassi, D.; Testolin, R. Development of a new SSR-based linkage map in apricot and analysis of synteny with existing Prunus maps. Tree Genet. Genomes 2006, 3, 239–249. [Google Scholar]
- Pedryc, A.; Ruthner, S.; Hermán, R.; Krska, B.; Hegedűs, A.; Halász, J. Genetic diversity of apricot revealed by a set of SSR markers from linkage group G1. Sci. Horticult 2009, 121, 19–26. [Google Scholar]
- Soriano, J.M.; Domingo, M.L.; Zuriaga, E.; Romero, C.; Zhebentyayeva, T.; Abbott, A.G.; Badenes, M.L. Identification of simple sequence repeat markers tightly linked to plum pox virus resistance in apricot. Mol. Breed 2012, 30, 1017–1026. [Google Scholar]
- Zhebentyayeva, T.; Reighard, G.; Gorina, V.; Abbott, A. Simple sequence repeat (SSR) analysis for assessment of genetic variability in apricot germplasm. Theor. Appl. Genet 2003, 106, 435–444. [Google Scholar]
- Ganopoulos, I.V.; Kazantzis, K.; Chatzicharisis, I.; Karayiannis, I.; Tsaftaris, A.S. Genetic diversity, structure and fruit trait associations in Greek sweet cherry cultivars using microsatellite based (SSR/ISSR) and morpho-physiological markers. Euphytica 2011, 181, 237–251. [Google Scholar]
- Li, M.-M.; Cai, Y.-L.; Qian, Z.-Q.; Zhao, G.-F. Genetic diversity and differentiation in Chinese sour cherry Prunus pseudocerasus Lindl., and its implications for conservation. Genet. Resour. Crop Evol 2009, 56, 455–464. [Google Scholar]
- Yilmaz, K.U.; Paydas-Kargi, S.; Dogan, Y.; Kafkas, S. Genetic diversity analysis based on ISSR, RAPD and SSR among Turkish Apricot Germplasms in Iran Caucasian eco-geographical group. Sci. Horticult 2012, 138, 138–143. [Google Scholar]
- Ai, P.-F.; Zhen, Z.-J.; Jin, Z.-Z. Genetic diversity and relationships within sweet kernel apricot and related Armeniaca species based on sequence-related amplified polymorphism markers. Biochem. Syst. Ecol 2011, 39, 694–699. [Google Scholar]
- Uzun, A.; Gulsen, O.; Seday, U.; Bircan, M.; Yilmaz, K.U. SRAP based genetic analysis of some apricot cultivars. Romanian Biotechnol. Lett 2010, 15, 5396–5404. [Google Scholar]
- Li, M.; Zhao, Z.; Miao, X.J. Genetic variability of wild apricot (Prunus. armeniac a L.) populations in the Ili Valley as revealed by ISSR markers. Genet. Resour. Crop Evol 2013. [Google Scholar] [CrossRef]
- Bourguiba, H.; Audergon, J.-M.; Krichen, L.; Trifi-Farah, N.; Mamouni, A.; Trabelsi, S.; Khadari, B. Genetic diversity and differentiation of grafted and seed propagated apricot (Prunus. armeniaca L.) in the Maghreb region. Sci. Horticult 2012, 142, 7–13. [Google Scholar]
- Wang, Y.; Zhang, J.; Sun, H.; Ning, N.; Yang, L. Construction and evaluation of a primary core collection of apricot germplasm in China. Sci. Horticult 2011, 128, 311–319. [Google Scholar]
- Tian-Ming, H.; Xue-Sen, C.; Zheng, X.; Jiang-Sheng, G.; Pei-Jun, L.; Wen, L.; Qing, L.; Yan, W. Using SSR markers to determine the population genetic structure of wild apricot (Prunus. armeniaca L.) in the Ily Valley of West China. Genet. Resour. Crop Evol 2006, 54, 563–572. [Google Scholar]
- Wang, L.; Chu, J. Optimization of biodiesel production from Siberian apricot (Prunus. sibirica L.) oil using response surface methodology. Asian J. Chem 2013, 25, 2577–2582. [Google Scholar]
- Liu, W.; Liu, N.; Zhang, Y.; Yu, X.; Sun, M.; Xu, M.; Zhang, Q.; Liu, S. Kernel-Using Apricot Resources and its Utilization. In XV International Symposium on Apricot Breeding and Culture; Avagyan, A., Ed.; Int. Soc. Horticultural Science: Leuven, Belgium, 2012; Volume 966, pp. 189–191. [Google Scholar]
- Slatkin, M. Gene flow in natural populations. Ann. Rev. Ecol. Syst 1985, 16, 393–430. [Google Scholar]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol 2005, 14, 2611–2620. [Google Scholar]
- Slatkin, M. Gene flow and the geographic structure of natural. Science 1987, 236, 787–792. [Google Scholar]
- Schaal, B.; Hayworth, D.; Olsen, K.; Rauscher, J.; Smith, W. Phylogeographic studies in plants: Problems and prospects. Mol. Ecol 1998, 7, 465–474. [Google Scholar]
- Zhai, P.; Sun, A.; Ren, F.; Liu, X.; Gao, B.; Zhang, Q. Changes of Climate Extremes in China. Climatic Change 1999, 42, 203–218. [Google Scholar]
- Olukolu, B.A.; Trainin, T.; Fan, S.; Kole, C.; Bielenberg, D.G.; Reighard, G.L.; Abbott, A.G.; Holland, D. Genetic linkage mapping for molecular dissection of chilling requirement and budbreak in apricot (Prunus. armeniaca L.). Genome 2009, 52, 819–828. [Google Scholar]
- Walther, G.-R.; Post, E.; Convey, P.; Menzel, A.; Parmesan, C.; Beebee, T.J.; Fromentin, J.-M.; Hoegh-Guldberg, O.; Bairlein, F. Ecological responses to recent climate change. Nature 2002, 416, 389–395. [Google Scholar]
- Nei, M. Analysis of gene diversity in subdivided populations. Proc. Natl. Acad. Sci. USA 1973, 70, 3321–3323. [Google Scholar]
- Budak, H.; Shearman, R.C.; Parmaksiz, I.; Dweikat, I. Comparative analysis of seeded and vegetative biotype buffalograsses based on phylogenetic relationship using ISSRs, SSRs, RAPDs, and SRAPs. Theor. Appl. Genet 2004, 109, 280–288. [Google Scholar]
- Zietkiewicz, E.; Rafalski, A.; Labuda, D. Genome fingerprinting by simple sequence repeat (SSR)-anchored polymerase chain reaction amplification. Genomics 1994, 20, 176–183. [Google Scholar]
- Russell, J.; Fuller, J.; Macaulay, M.; Hatz, B.; Jahoor, A.; Powell, W.; Waugh, R. Direct comparison of levels of genetic variation among barley accessions detected by RFLPs, AFLPs, SSRs and RAPDs. Theor. Appl. Genet 1997, 95, 714–722. [Google Scholar]
- Rohlf, F. NTSYS-PC, Numerical Taxonomy System for the PC Exeter Software, Version 2.1; Applied Biostatistics Inc Setauket: E. Setauket, NY, USA, 2000. [Google Scholar]
- Zhang, P.; Li, J.; Li, X.; Liu, X.; Zhao, X.; Lu, Y. Population structure and genetic diversity in a rice core collection (Oryza sativa L.) investigated with SSR markers. PLoS One 2011, 6, e27565. [Google Scholar]
- Bourguiba, H.; Khadari, B.; Krichen, L.; Trifi-Farah, N.; Mamouni, A.; Trabelsi, S.; Audergon, J.-M. Genetic relationships between local north african apricot (Prunus armeniaca L.) germplasm and recently introduced varieties. Sci. Horticult 2013, 152, 61–69. [Google Scholar]
- Martín, C.; Herrero, M.; Hormaza, J.I. Molecular characterization of apricot germplasm from an old stone collection. PLoS One 2011, 6, e23979. [Google Scholar]
- Karp, A. Molecular Tools in Plant Genetic Resources Conservation: A Guide to the Technologies; Bioversity International: Rome, Italy, 1997. [Google Scholar]
- Cipriani, G.; Lot, G.; Huang, W.-G.; Marrazzo, M.; Peterlunger, E.; Testolin, R. AC/GT and AG/CT microsatellite repeats in peach [Prunus persica (L) Batsch]: Isolation, characterisation and cross-species amplification in. Prunus. Theor. Appl. Genet 1999, 99, 65–72. [Google Scholar]
- Dirlewanger, E.; Cosson, P.; Tavaud, M.; Aranzana, M.; Poizat, C.; Zanetto, A.; Arús, P.; Laigret, F. Development of microsatellite markers in peach [Prunus persica (L.) Batsch] and their use in genetic diversity analysis in peach and sweet cherry (Prunus avium L.). Theor. Appl. Genet 2002, 105, 127–138. [Google Scholar]
- Hagen, L.S.; Chaib, J.; Fady, B.; Decroocq, V.; Bouchet, J.P.; Lambert, P.; Audergon, J.M. Genomic and cDNA microsatellites from apricot (Prunus armeniaca L.). Mol. Ecol. Notes 2004, 4, 742–745. [Google Scholar]
- Lopes, M.; Sefc, K.; Laimer, M.; da Câmara Machado, A. Identification of microsatellite loci in apricot. Mol. Ecol. Notes 2002, 2, 24–26. [Google Scholar]
- Messina, R.; Lain, O.; Marrazzo, M.; Cipriani, G.; Testolin, R. New set of microsatellite loci isolated in Apricot. Mol. Ecol. Notes 2004, 4, 432–434. [Google Scholar]
- Sosinski, B.; Gannavarapu, M.; Hager, L.; Beck, L.; King, G.J.; Ryder, C.; Rajapakse, S.; Baird, W.; Ballard, R.; Abbott, A. Characterization of microsatellite markers in peach [Prunus persica (L.) Batsch]. Theor. Appl. Genet 2000, 101, 421–428. [Google Scholar]
- Yeh, F.; Yang, R.-C.; Boyle, T.; Ye, Z.; Mao, J.X. POPGENE, the User-Friendly Shareware for Population Genetic Analysis. In Molecular Biology and Biotechnology Centre; University of Alberta: Edmonton, AB, Canada, 1997. [Google Scholar]
- Shannon, C.E.; Weaver, W. The Mathematical Theory of Communication; Urbana, I.L., Ed.; University of Illinois Press: Champaign, IL, USA, 1949; Volume 19, p. 1. [Google Scholar]
- Excoffier, L.; Laval, G.; Schneider, S. Arlequin (version 3.0): An integrated software package for population genetics data analysis. Evol. Bioinforma. Online 2005, 1, 47–50. [Google Scholar]
- Liedloff, A. Mantel (Version 2.0): Mantel Nonparametric Test Calculator; Queensland University of Technology: Brisbane, Australia, 1999. [Google Scholar]
- Cao, Q.; Lu, B.R.; Xia, H.; Rong, J.; Sala, F.; Spada, A.; Grassi, F. Genetic diversity and origin of weedy rice (Oryza sativa f. spontanea) populations found in North-eastern China revealed by simple sequence repeat (SSR) markers. Ann. Bot 2006, 98, 1241–1252. [Google Scholar]
- Tamura, K.; Dudley, J.; Nei, M.; Kumar, S. MEGA4: Molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol. Biol. Evol 2007, 24, 1596–1599. [Google Scholar]
- Isshiki, S.; Iwata, N.; Khan, M.M.R. Issr variations in eggplant (Solanum melongena L.) and related Solanum species. Scientia Hortic 2008, 117, 186–190. [Google Scholar]
- Nei, M. The theory of genetic distance and evolution of human races. J. Hum. Genet 1978, 23, 341–369. [Google Scholar]
| Population | ISSR | SRAP | SSR | |||||
|---|---|---|---|---|---|---|---|---|
| h | I | PPB (%) | h | I | PPB (%) | h | I | |
| LY | 0.191 | 0.285 | 54.4 | 0.145 | 0.220 | 43.3 | 1.099 | 0.612 |
| ZD | 0.156 | 0.238 | 48.5 | 0.141 | 0.216 | 45.0 | 1.120 | 0.632 |
| AS | 0.200 | 0.301 | 59.2 | 0.165 | 0.254 | 55.8 | 1.231 | 0.662 |
| HC | 0.208 | 0.310 | 59.2 | 0.176 | 0.271 | 57.5 | 1.157 | 0.631 |
| HX | 0.197 | 0.294 | 56.3 | 0.178 | 0.271 | 55.0 | 1.097 | 0.591 |
| ZY | 0.198 | 0.293 | 53.4 | 0.190 | 0.288 | 59.2 | 1.095 | 0.606 |
| YG | 0.156 | 0.230 | 42.7 | 0.158 | 0.241 | 49.2 | 1.134 | 0.629 |
| GL | 0.140 | 0.212 | 45.6 | 0.148 | 0.228 | 48.3 | 1.097 | 0.596 |
| HY | 0.161 | 0.239 | 44.7 | 0.138 | 0.209 | 42.5 | 1.008 | 0.577 |
| YQ | 0.166 | 0.248 | 47.6 | 0.165 | 0.248 | 49.2 | 1.027 | 0.579 |
| HR | 0.204 | 0.304 | 58.3 | 0.213 | 0.326 | 69.2 | 1.106 | 0.602 |
| MY | 0.197 | 0.291 | 54.4 | 0.186 | 0.283 | 60.0 | 1.141 | 0.633 |
| CY | 0.154 | 0.235 | 48.5 | 0.138 | 0.213 | 48.3 | 1.100 | 0.612 |
| KZ | 0.184 | 0.275 | 50.5 | 0.172 | 0.262 | 54.2 | 0.977 | 0.544 |
| LiY | 0.160 | 0.241 | 48.6 | 0.166 | 0.254 | 55.0 | 1.025 | 0.573 |
| WC | 0.161 | 0.242 | 47.6 | 0.149 | 0.230 | 50.0 | 0.945 | 0.541 |
| LH | 0.174 | 0.259 | 50.5 | 0.137 | 0.210 | 45.8 | 1.081 | 0.592 |
| PQ | 0.144 | 0.221 | 47.6 | 0.134 | 0.213 | 50.0 | 1.133 | 0.621 |
| NC | 0.161 | 0.243 | 48.5 | 0.146 | 0.226 | 52.5 | 0.983 | 0.554 |
| LX | 0.161 | 0.238 | 43.7 | 0.132 | 0.200 | 40.8 | 0.891 | 0.512 |
| KK | 0.139 | 0.211 | 41.8 | 0.120 | 0.183 | 36.7 | 0.939 | 0.534 |
| Mean | 0.172 | 0.258 | 50.1 | 0.157 | 0.240 | 50.8 | 1.066 | 0.592 |
| At the species level | 0.248 | 0.387 | 91.3 | 0.218 | 0.344 | 82.5 | 1.639 | 0.782 |
| ISSR | SRAP | SSR | ||||
|---|---|---|---|---|---|---|
| Source of variance | Variance component | Ratio (%) | Variance component | Ratio (%) | Variance component | Ratio (%) |
| Among populations | 3.098 | 25.01 | 3.561 | 23.84% | 1.437 | 16.65 |
| Within populations | 9.289 | 74.99 | 11.378 | 76.16% | 7.192 | 83.35 |
| Population code | Seed collection sites | Latitude (°N) | Longitude (°E) | Elevation (m) |
|---|---|---|---|---|
| LY | Lingyou, Shannxi Province | 34°35′ | 107°46′ | 1,292 |
| ZD | Zhidan, Shannxi Province | 36°48′ | 108°45′ | 1,238 |
| AS | Ansai, Shannxi Province | 37°04′ | 109°09′ | 1,252 |
| HC | Huachi, Gansu Province | 36°12′ | 107°56′ | 1,244 |
| HX | Huanxian, Gansu Province | 36°31′ | 107°17′ | 1,190 |
| ZY | Zhenyuan, Gansu Province | 35°37′ | 107°02′ | 1,281 |
| YG | Yanggao, Shanxi Province | 40°07′ | 113°54′ | 1,097 |
| GL | Guangling, Shanxi Province | 39°49′ | 114°34′ | 1,284 |
| HY | Hunyuan, Shanxi Province | 39°32′ | 113°28′ | 1,442 |
| YQ | Yanqing, Beijing Province | 40°26′ | 116°14′ | 641 |
| HR | Huairou, Beijing Province | 40°36′ | 116°44′ | 382 |
| MY | Miyun, Beijing Province | 40°31′ | 117°13′ | 228 |
| CY | Chaoyang, Liaoning Province | 41°42′ | 120°03′ | 627 |
| KZ | Kazuo, Liaoning Province | 41°01′ | 119°44′ | 405 |
| LiY | Lingyuan, Liaoning Province | 40°53′ | 119°12′ | 560 |
| WC | Weichang, Hebei Province | 41°56′ | 117°44′ | 1,127 |
| LH | Longhua, Hebei Province | 41°15′ | 117°21′ | 675 |
| PQ | Pingquan, Hebei Province | 41°02′ | 118°32′ | 583 |
| NC | Ningcheng, Inner Mongolia | 41°45′ | 119°01′ | 1,134 |
| LX | Linxi, Inner Mongolia | 44°01′ | 118°20′ | 1,207 |
| KK | Keshiketeng, Inner Mongolia | 43°16′ | 117°33′ | 1,221 |
© 2014 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Li, M.; Zhao, Z.; Miao, X.; Zhou, J. Genetic Diversity and Population Structure of Siberian apricot (Prunus sibirica L.) in China. Int. J. Mol. Sci. 2014, 15, 377-400. https://doi.org/10.3390/ijms15010377
Li M, Zhao Z, Miao X, Zhou J. Genetic Diversity and Population Structure of Siberian apricot (Prunus sibirica L.) in China. International Journal of Molecular Sciences. 2014; 15(1):377-400. https://doi.org/10.3390/ijms15010377
Chicago/Turabian StyleLi, Ming, Zhong Zhao, Xingjun Miao, and Jingjing Zhou. 2014. "Genetic Diversity and Population Structure of Siberian apricot (Prunus sibirica L.) in China" International Journal of Molecular Sciences 15, no. 1: 377-400. https://doi.org/10.3390/ijms15010377
APA StyleLi, M., Zhao, Z., Miao, X., & Zhou, J. (2014). Genetic Diversity and Population Structure of Siberian apricot (Prunus sibirica L.) in China. International Journal of Molecular Sciences, 15(1), 377-400. https://doi.org/10.3390/ijms15010377




