Development of Quorum-Based Anti-Virulence Therapeutics Targeting Gram-Negative Bacterial Pathogens
Abstract
:1. Introduction
2. Quorum Sensing in Gram-Negative Bacteria
2.1. Classical AHL- and AI-2-Mediated Quorum Sensing
2.2. Alternative Quorum Sensing Pathways in Gram-Negative Bacteria
2.3. Quorum Sensing Mechanisms Used by Clinically Relevant Gram-Negative Pathogens
2.3.1. Pseudomonas aeruginosa
2.3.2. Acinetobacter baumannii and Klebsiella pneumoniae
2.3.3. Enterobacter spp. and Escherichia coli
3. Quorum Sensing Disruption Strategies in Gram-Negative Bacteria
3.1. Quorum Sensing Disruption in Pseudomonas aeruginosa
3.1.1. Inhibition of AHL Detection
3.1.2. Inhibition of AHL Accumulation
3.1.3. Inhibition of the pqs System
3.1.4. Inhibition by Natural Compounds
3.2. Quorum Sensing Disruption in Escherichia coli
3.3. Quorum Sensing Disruption in Klebsiella pneumoniae, Enterobacter spp. and Acinetobacter baumannii
4. Quorum-Quenching Considerations and Implications
5. Conclusions
Acknowledgments
Conflict of Interest
References
- Hastings, J.W.; Nealson, K.H. Bacterial bioluminescence. Annu. Rev. Microbiol 1977, 31, 549–595. [Google Scholar]
- Winzer, K.; Williams, P. Quorum sensing and the regulation of virulence gene expression in pathogenic bacteria. Int. J. Med. Microbiol 2001, 291, 131–143. [Google Scholar]
- Waters, C.M.; Bassler, B.L. Quorum sensing: Cell-to-cell communication in bacteria. Annu. Rev. Cell Dev. Biol 2005, 21, 319–346. [Google Scholar]
- LaSarre, B.; Federle, M.J. Exploiting quorum sensing to confuse bacterial pathogens. Microbiol. Mol. Biol. Rev 2013, 77, 73–111. [Google Scholar]
- Cegelski, L.; Marshall, G.R.; Eldridge, G.R.; Hultgren, S.J. The biology and future prospects of antivirulence therapies. Nat. Rev. Microbiol 2008, 6, 17–27. [Google Scholar]
- Alanis, A.J. Resistance to antibiotics: Are we in the post-antibiotic era? Arch. Med. Res 2005, 36, 697–705. [Google Scholar]
- Walsh, C. Antibiotics: Actions, Origins, Resistance; Asm Press: Washington, DC, USA, 2003. [Google Scholar]
- Werner, G.; Strommenger, B.; Witte, W. Acquired vancomycin resistance in clinically relevant pathogens. Future Microbiol 2008, 3, 547–562. [Google Scholar]
- Sengupta, S.; Chattopadhyay, M.K.; Grossart, H.P. The multifaceted roles of antibiotics and antibiotic resistance in nature. Front. Microbiol 2013, 4, 47. [Google Scholar]
- Dellit, T.H.; Owens, R.C.; McGowan, J.E.; Gerding, D.N.; Weinstein, R.A.; Burke, J.P.; Huskins, W.C.; Paterson, D.L.; Fishman, N.O.; Carpenter, C.F.; et al. Infectious diseases society of america and the society for healthcare epidemiology of america guidelines for developing an institutional program to enhance antimicrobial stewardship. Clin. Infect. Dis 2007, 44, 159–177. [Google Scholar]
- Fernandes, P. Antibacterial discovery and development[mdash]the failure of success? Nat. Biotech 2006, 24, 1497–1503. [Google Scholar]
- Rasko, D.A.; Sperandio, V. Anti-virulence strategies to combat bacteria-mediated disease. Nature reviews. Drug Discov 2010, 9, 117–128. [Google Scholar]
- Kiran, S. Enzymatic quorum quenching increases antibiotic susceptibility of multidrug resistant Pseudomonas aeruginosa. Irani. J. Microbiol 2011, 3, 1–12. [Google Scholar]
- McLean, R.J.C.; Whiteley, M.; Stickler, D.J.; Fuqua, W.C. Evidence of autoinducer activity in naturally occurring biofilms. FEMS Microbiol. Lett 1997, 154, 259–263. [Google Scholar]
- Weinstein, R.A.; Gaynes, R.; Edwards, J.R. National Nosocomial Infections Surveillance System Overview of nosocomial infections caused by gram-negative bacilli. Clin. Infect. Dis. 2005, 41, 848–854. [Google Scholar]
- Vincent, J.L.; Rello, J.; Marshall, J.; Silva, E.; Anzueto, A.; Martin, C.D.; Moreno, R.; Lipman, J.; Gomersall, C.; Sakr, Y.; et al. International study of the prevalence and outcomes of infection in intensive care units. JAMA 2009, 302, 2323–2329. [Google Scholar]
- Rice, L.B. Federal funding for the study of antimicrobial resistance in nosocomial pathogens: No eskape. J. Infect. Dis 2008, 197, 1079–1081. [Google Scholar]
- Boucher, H.W.; Talbot, G.H.; Bradley, J.S.; Edwards, J.E.; Gilbert, D.; Rice, L.B.; Scheld, M.; Spellberg, B.; Bartlett, J. Bad bugs, no drugs: No eskape! An update from the infectious diseases society of america. Clin. Infect. Dis 2009, 48, 1–12. [Google Scholar]
- Ramphal, R.; Ambrose, P.G. Extended-spectrum β-lactamases and clinical outcomes: Current data. Clin. Infect. Dis 2006, 42, S164–S172. [Google Scholar]
- Tschudin-Sutter, S.; Frei, R.; Dangel, M.; Stranden, A.; Widmer, A.F. Rate of transmission of extended-spectrum beta-lactamase-producing enterobacteriaceae without contact isolation. Clin. Infect. Dis 2012, 55, 1505–1511. [Google Scholar]
- Kaplan, H.B.; Greenberg, E.P. Diffusion of autoinducer is involved in regulation of the Vibrio fischeri luminescence system. J. Bacteriol 1985, 163, 1210–1214. [Google Scholar]
- Fuqua, C.; Winans, S.C.; Greenberg, E.P. Census and consensus in bacterial ecosystems: The luxr-luxi family of quorum-sensing transcriptional regulators. Annu. Rev. Microbiol 1996, 50, 727–751. [Google Scholar]
- Re, S.D.; Bertagnoli, S.; Fourment, J.; Reyrat, J.-M.; Kahn, D. Intramolecular signal transduction within the fixj transcriptional activator: In vitro evidence for the inhibitory effect of the phosphorylatable regulatory domain. Nucleic Acids Res 1994, 22, 1555–1561. [Google Scholar]
- Freeman, J.A.; Lilley, B.N.; Bassler, B.L. A genetic analysis of the functions of luxn: A two-component hybrid sensor kinase that regulates quorum sensing in Vibrio harveyi. Mol. Microbiol 2000, 35, 139–149. [Google Scholar]
- Lenz, D.H.; Mok, K.C.; Lilley, B.N.; Kulkarni, R.V.; Wingreen, N.S.; Bassler, B.L. The small rna chaperone hfq and multiple small rnas control quorum sensing in Vibrio harveyi and Vibrio cholerae. Cell 2004, 118, 69–82. [Google Scholar]
- Ni, N.; Li, M.; Wang, J.; Wang, B. Inhibitors and antagonists of bacterial quorum sensing. Med. Res. Rev 2009, 29, 65–124. [Google Scholar]
- Xavier, K.B.; Bassler, B.L. Interference with ai-2-mediated bacterial cell-cell communication. Nature 2005, 437, 750–753. [Google Scholar]
- Rajamani, S.; Sayre, R.T. A sensitive fluorescence reporter for monitoring quorum sensing regulated protease production in Vibrio harveyi. J. Microbiol. Methods 2011, 84, 189–193. [Google Scholar]
- Schauder, S.; Shokat, K.; Surette, M.G.; Bassler, B.L. The luxs family of bacterial autoinducers: Biosynthesis of a novel quorum-sensing signal molecule. Mol. Microbiol 2001, 41, 463–476. [Google Scholar]
- Chen, X.; Schauder, S.; Potier, N.; Van Dorsselaer, A.; Pelczer, I.; Bassler, B.L.; Hughson, F.M. Structural identification of a bacterial quorum-sensing signal containing boron. Nature 2002, 415, 545–549. [Google Scholar]
- De Keersmaecker, S.C.J.; Varszegi, C.; van Boxel, N.; Habel, L.W.; Metzger, K.; Daniels, R.; Marchal, K.; de Vos, D.; Vanderleyden, J. Chemical synthesis of (S)-4,5-dihydroxy-2,3-pentanedione, a bacterial signal molecule precursor, and validation of its activity in Salmonella typhimurium. J. Biol. Chem 2005, 280, 19563–19568. [Google Scholar]
- Pereira, C.S.; Thompson, J.A.; Xavier, K.B. Ai-2-mediated signaling in bacteria. FEMS Microbiol. Rev 2013, 37, 156–181. [Google Scholar]
- Galloway, W.R.J.D.; Hodgkinson, J.T.; Bowden, S.D.; Welch, M.; Spring, D.R. Quorum sensing in gram-negative bacteria: Small-molecule modulation of ahl and ai-2 quorum sensing pathways. Chem. Rev 2010, 111, 28–67. [Google Scholar]
- Henke, J.M.; Bassler, B.L. Three parallel quorum-sensing systems regulate gene expression in vibrio harveyi. J. Bacteriol 2004, 186, 6902–6914. [Google Scholar]
- Neiditch, M.B.; Federle, M.J.; Miller, S.T.; Bassler, B.L.; Hughson, F.M. Regulation of luxpq receptor activity by the quorum-sensing signal autoinducer-2. Mol. Cell 2005, 18, 507–518. [Google Scholar]
- Pereira, C.S.; de Regt, A.K.; Brito, P.H.; Miller, S.T.; Xavier, K.B. Identification of functional lsrb-like autoinducer-2 receptors. J. Bacteriol 2009, 191, 6975–6987. [Google Scholar]
- Xavier, K.B.; Bassler, B.L. Luxs quorum sensing: More than just a numbers game. Curr. Opin. Microbiol 2003, 6, 191–197. [Google Scholar]
- Vendeville, A.; Winzer, K.; Heurlier, K.; Tang, C.M.; Hardie, K.R. Making ‘sense’ of metabolism: Autoinducer-2, luxs and pathogenic bacteria. Nat. Rev. Microbiol 2005, 3, 383–396. [Google Scholar]
- Déziel, E.; Lépine, F.; Milot, S.; He, J.; Mindrinos, M.N.; Tompkins, R.G.; Rahme, L.G. Analysis of Pseudomonas aeruginosa 4-hydroxy-2-alkylquinolines (haqs) reveals a role for 4-hydroxy-2-heptylquinoline in cell-to-cell communication. Proc. Nat. Acad. Sci. USA 2004, 101, 1339–1344. [Google Scholar]
- Pesci, E.C.; Milbank, J.B.J.; Pearson, J.P.; McKnight, S.; Kende, A.S.; Greenberg, E.P.; Iglewski, B.H. Quinolone signaling in the cell-to-cell communication system of Pseudomonas aeruginosa. Proc. Nat. Acad. Sci 1999, 96, 11229–11234. [Google Scholar]
- Dubern, J.-F.; Diggle, S.P. Quorum sensing by 2-alkyl-4-quinolones in Pseudomonas aeruginosa and other bacterial species. Mol. BioSyst 2008, 4, 882–888. [Google Scholar]
- Ng, W.L.; Perez, L.J.; Wei, Y.; Kraml, C.; Semmelhack, M.F.; Bassler, B.L. Signal production and detection specificity in vibrio cqsa/cqss quorum-sensing systems. Mol. Microbiol 2011, 79, 1407–1417. [Google Scholar]
- Gilbert, K.B.; Kim, T.H.; Gupta, R.; Greenberg, E.P.; Schuster, M. Global position analysis of the Pseudomonas aeruginosa quorum-sensing transcription factor lasr. Mol. Microbiol 2009, 73, 1072–1085. [Google Scholar]
- Chugani, S.A.; Whiteley, M.; Lee, K.M.; D’Argenio, D.; Manoil, C.; Greenberg, E.P. Qscr, a modulator of quorum-sensing signal synthesis and virulence in Pseudomonas aeruginosa. Proc. Nat. Acad. Sci. USA 2001, 98, 2752–2757. [Google Scholar]
- Lee, J.-H.; Lequette, Y.; Greenberg, E.P. Activity of purified qscr, a Pseudomonas aeruginosa orphan quorum-sensing transcription factor. Mol. Microbiol 2006, 59, 602–609. [Google Scholar]
- Wang, X.D.; de Boer, P.A.; Rothfield, L.I. A factor that positively regulates cell division by activating transcription of the major cluster of essential cell division genes of Escherichia coli. EMBO J 1991, 10, 3363–3372. [Google Scholar]
- Rezzonico, F.; Smits, T.H.M.; Duffy, B. Detection of ai-2 receptors in genomes of enterobacteriaceae suggests a role of type-2 quorum sensing in closed ecosystems. Sensors 2012, 12, 6645–6665. [Google Scholar]
- Wang, L.; Li, J.; March, J.C.; Valdes, J.J.; Bentley, W.E. Luxs-dependent gene regulation in Escherichia coli k-12 revealed by genomic expression profiling. J. Bacteriol 2005, 187, 8350–8360. [Google Scholar]
- Walters, M.; Sperandio, V. Quorum sensing in Escherichia coli and Salmonella. Int. J. Med. Microbiol 2006, 296, 125–131. [Google Scholar]
- Sperandio, V.; Torres, A.G.; Kaper, J.B. Quorum sensing Escherichia coli regulators b and c (qsebc): A novel two-component regulatory system involved in the regulation of flagella and motility by quorum sensing in e. Coli. Mol. Microbiol 2002, 43, 809–821. [Google Scholar]
- Bhargava, N.; Sharma, P.; Capalash, N. Quorum sensing in acinetobacter: An emerging pathogen. Crit. Rev. Microbiol 2010, 36, 349–360. [Google Scholar]
- Yin, W.-F.; Purmal, K.; Chin, S.; Chan, X.-Y.; Chan, K.-G. Long chain N-acyl homoserine lactone production by Enterobacter sp. isolated from human tongue surfaces. Sensors 2012, 12, 14307–14314. [Google Scholar]
- Shankar, M.; Ponraj, P.; Illakkiam, D.; Rajendhran, J.; Gunasekaran, P. Inactivation of the transcriptional regulator-encoding gene sdia enhances rice root colonization and biofilm formation in Enterobacter cloacae gs1. J. Bacteriol 2013, 195, 39–45. [Google Scholar]
- Yin, W.-F.; Purmal, K.; Chin, S.; Chan, X.-Y.; Koh, C.-L.; Sam, C.-K.; Chan, K.-G. N-acyl homoserine lactone production by Klebsiella pneumoniae isolated from human tongue surface. Sensors 2012, 12, 3472–3483. [Google Scholar]
- Balestrino, D.; Haagensen, J.A.J.; Rich, C.; Forestier, C. Characterization of type 2 quorum sensing in Klebsiella pneumoniae and relationship with biofilm formation. J. Bacteriol 2005, 187, 2870–2880. [Google Scholar]
- Higgins, D.A.; Pomianek, M.E.; Kraml, C.M.; Taylor, R.K.; Semmelhack, M.F.; Bassler, B.L. The major vibrio cholerae autoinducer and its role in virulence factor production. Nature 2007, 450, 883–886. [Google Scholar]
- Barber, C.E.; Tang, J.L.; Feng, J.X.; Pan, M.Q.; Wilson, T.J.G.; Slater, H.; Dow, J.M.; Williams, P.; Daniels, M.J. A novel regulatory system required for pathogenicity of Xanthomonas campestris is mediated by a small diffusible signal molecule. Mol. Microbiol 1997, 24, 555–566. [Google Scholar]
- Fouhy, Y.; Lucey, J.F.; Ryan, R.P.; Dow, J.M. Cell-cell signaling, cyclic di-gmp turnover and regulation of virulence in Xanthomonas campestris. Res. Microbiol 2006, 157, 899–904. [Google Scholar]
- Newman, K.L.; Almeida, R.P.P.; Purcell, A.H.; Lindow, S.E. Cell-cell signaling controls Xylella fastidiosa interactions with both insects and plants. Proc. Nat. Acad. Sci. USA 2004, 101, 1737–1742. [Google Scholar]
- Fouhy, Y.; Scanlon, K.; Schouest, K.; Spillane, C.; Crossman, L.; Avison, M.B.; Ryan, R.P.; Dow, J.M. Diffusible signal factor-dependent cell-cell signaling and virulence in the nosocomial pathogen Stenotrophomonas maltophilia. J. Bacteriol 2007, 189, 4964–4968. [Google Scholar]
- Deng, Y.; Wu, J.E.; Eberl, L.; Zhang, L.-H. Structural and functional characterization of diffusible signal factor family quorum-sensing signals produced by members of the Burkholderia cepacia complex. Appl. Environ. Microbiol 2010, 76, 4675–4683. [Google Scholar]
- Ryan, R.P.; Dow, J.M. Communication with a growing family: Diffusible signal factor (dsf) signaling in bacteria. Trends Microbiol 2011, 19, 145–152. [Google Scholar]
- Ryan, R.P.; Fouhy, Y.; Garcia, B.F.; Watt, S.A.; Niehaus, K.; Yang, L.; Tolker-Nielsen, T.; Dow, J.M. Interspecies signaling via the stenotrophomonas maltophilia diffusible signal factor influences biofilm formation and polymyxin tolerance in Pseudomonas aeruginosa. Mol. Microbiol 2008, 68, 75–86. [Google Scholar]
- Sperandio, V.; Torres, A.G.; Jarvis, B.; Nataro, J.P.; Kaper, J.B. Bacteria-host communication: The language of hormones. Proc. Nat. Acad. Sci 2003, 100, 8951–8956. [Google Scholar]
- Clarke, M.B.; Hughes, D.T.; Zhu, C.; Boedeker, E.C.; Sperandio, V. The qsec sensor kinase: A bacterial adrenergic receptor. Proc. Nat. Acad. Sci 2006, 103, 10420–10425. [Google Scholar]
- Juhas, M.; Eberl, L.; Tümmler, B. Quorum sensing: The power of cooperation in the world of Pseudomonas. Environ. Microbiol 2005, 7, 459–471. [Google Scholar]
- Wilder, C.N.; Diggle, S.P.; Schuster, M. Cooperation and cheating in Pseudomonas aeruginosa: The roles of the las, rhl and pqs quorum-sensing systems. ISME J 2011, 5, 1332–1343. [Google Scholar]
- Schuster, M.; Lostroh, C.P.; Ogi, T.; Greenberg, E.P. Identification, timing, and signal specificity of Pseudomonas aeruginosa quorum-controlled genes: A transcriptome analysis. J. Bacteriol 2003, 185, 2066–2079. [Google Scholar]
- Deziel, E.; Gopalan, S.; Tampakaki, A.P.; Lepine, F.; Padfield, K.E.; Saucier, M.; Xiao, G.; Rahme, L.G. The contribution of mvfr to Pseudomonas aeruginosa pathogenesis and quorum sensing circuitry regulation: Multiple quorum sensing-regulated genes are modulated without affecting lasri, rhlri or the production of N-acyl-l-homoserine lactones. Mol. Microbiol 2005, 55, 998–1014. [Google Scholar]
- McGrath, S.; Wade, D.S.; Pesci, E.C. Dueling quorum sensing systems in Pseudomonas aeruginosa control the production of the Pseudomonas quinolone signal (pqs). FEMS Microbiol. Lett 2004, 230, 27–34. [Google Scholar]
- McKnight, S.L.; Iglewski, B.H.; Pesci, E.C. The Pseudomonas quinolone signal regulates rhl quorum sensing in Pseudomonas aeruginosa. J. Bacteriol 2000, 182, 2702–2708. [Google Scholar]
- Wade, D.S.; Calfee, M.W.; Rocha, E.R.; Ling, E.A.; Engstrom, E.; Coleman, J.P.; Pesci, E.C. Regulation of Pseudomonas quinolone signal synthesis in Pseudomonas aeruginosa. J. Bacteriol 2005, 187, 4372–4380. [Google Scholar]
- Pesci, E.C.; Pearson, J.P.; Seed, P.C.; Iglewski, B.H. Regulation of las and rhl quorum sensing in Pseudomonas aeruginosa. J. Bacteriol 1997, 179, 3127–3132. [Google Scholar]
- Pearson, J.P.; Pesci, E.C.; Iglewski, B.H. Roles of Pseudomonas aeruginosa las and rhl quorum-sensing systems in control of elastase and rhamnolipid biosynthesis genes. J. Bacteriol 1997, 179, 5756–5767. [Google Scholar]
- Ha, C.; Park, S.J.; Im, S.J.; Lee, J.H. Interspecies signaling through qscr, a quorum receptor of Pseudomonas aeruginosa. Mol. Cells 2012, 33, 53–59. [Google Scholar]
- Ledgham, F.; Ventre, I.; Soscia, C.; Foglino, M.; Sturgis, J.N.; Lazdunski, A. Interactions of the quorum sensing regulator qscr: Interaction with itself and the other regulators of Pseudomonas aeruginosa lasr and rhlr. Mol. Microbiol 2003, 48, 199–210. [Google Scholar]
- Lequette, Y.; Lee, J.H.; Ledgham, F.; Lazdunski, A.; Greenberg, E.P. A distinct qscr regulon in the Pseudomonas aeruginosa quorum-sensing circuit. J. Bacteriol 2006, 188, 3365–3370. [Google Scholar]
- Niu, C.; Clemmer, K.M.; Bonomo, R.A.; Rather, P.N. Isolation and characterization of an autoinducer synthase from Acinetobacter baumannii. J. Bacteriol 2008, 190, 3386–3392. [Google Scholar]
- Wand, M.E. Acinetobacter baumannii virulence is enhanced in Galleria mellonella following biofilm adaptation. J. Med. Microbiol 2012, 61, 470–477. [Google Scholar]
- Bhargava, N.; Sharma, P.; Capalash, N. N-acyl homoserine lactone mediated interspecies interactions between A. baumannii and P. aeruginosa. Biofouling 2012, 28, 813–822. [Google Scholar]
- Zhu, H.; Liu, H.-J.; Ning, S.-J.; Gao, Y.-L. A luxs-dependent transcript profile of cell-to-cell communication in Klebsiella pneumoniae. Mol. BioSyst 2011, 7, 3164–3168. [Google Scholar]
- Zhu, H.; Liu, H.J.; Ning, S.J.; Gao, Y.L. The response of type 2 quorum sensing in Klebsiella pneumoniae to a fluctuating culture environment. DNA Cell Biol 2012, 31, 455–459. [Google Scholar]
- Russo, T.A.; Shon, A.S.; Beanan, J.M.; Olson, R.; MacDonald, U.; Pomakov, A.O.; Visitacion, M.P. Hypervirulent k. Pneumoniae secretes more and more active iron-acquisition molecules than “classical” k. Pneumoniae thereby enhancing its virulence. PLoS One 2011, 6, e26734. [Google Scholar]
- Sanders, W.E.; Sanders, C.C. Enterobacter spp.: Pathogens poised to flourish at the turn of the century. Clin. Microbiol. Rev 1997, 10, 220–241. [Google Scholar]
- Pinto, U.M.; de Souza Viana, E.; Martins, M.L.; Vanetti, M.C.D. Detection of acylated homoserine lactones in gram-negative proteolytic psychrotrophic bacteria isolated from cooled raw milk. Food Control 2007, 18, 1322–1327. [Google Scholar]
- Gram, L.; Christensen, A.B.; Ravn, L.; Molin, S.; Givskov, M. Production of acylated homoserine lactones by psychrotrophic members of the Enterobacteriaceae isolated from foods. Appl. Environ. Microbiol 1999, 65, 3458–3463. [Google Scholar]
- Reis Ponce, A.; Martins, M.; Araujo, E.; Mantovani, H.; Vanetti, M. Aiia quorum-sensing quenching controls proteolytic activity and biofilm formation by Enterobacter cloacae. Curr. Microbiol 2012, 65, 758–763. [Google Scholar]
- Rahmati, S.; Yang, S.; Davidson, A.L.; Zechiedrich, E.L. Control of the acrab multidrug efflux pump by quorum-sensing regulator sdia. Mol. Microbiol 2002, 43, 677–685. [Google Scholar]
- Tavío, M.M.; Aquili, V.D.; Poveda, J.B.; Antunes, N.T.; Sánchez-Céspedes, J.; Vila, J. Quorum-sensing regulator sdia and mara overexpression is involved in in vitro-selected multidrug resistance of Escherichia coli. J. Antimicrob. Chemother 2010, 65, 1178–1186. [Google Scholar]
- Kanamaru, K.; Kanamaru, K.; Tatsuno, I.; Tobe, T.; Sasakawa, C. Sdia, an Escherichia coli homologue of quorum-sensing regulators, controls the expression of virulence factors in enterohaemorrhagic Escherichia coli o157:H7. Mol. Microbiol 2000, 38, 805–816. [Google Scholar]
- Lee, J.; Maeda, T.; Hong, S.H.; Wood, T.K. Reconfiguring the quorum-sensing regulator sdia of Escherichia coli to control biofilm formation via indole and n-acylhomoserine lactones. Appl. Environ. Microbiol 2009, 75, 1703–1716. [Google Scholar]
- Xue, T.; Zhao, L.; Sun, H.; Zhou, X.; Sun, B. Lsrr-binding site recognition and regulatory characteristics in Escherichia coli ai-2 quorum sensing. Cell Res 2009, 19, 1258–1268. [Google Scholar]
- Xavier, K.B.; Miller, S.T.; Lu, W.; Kim, J.H.; Rabinowitz, J.; Pelczer, I.; Semmelhack, M.F.; Bassler, B.L. Phosphorylation and processing of the quorum-sensing molecule autoinducer-2 in enteric bacteria. ACS Chem. Biol 2007, 2, 128–136. [Google Scholar]
- Wang, L.; Hashimoto, Y.; Tsao, C.-Y.; Valdes, J.J.; Bentley, W.E. Cyclic amp (camp) and camp receptor protein influence both synthesis and uptake of extracellular autoinducer 2 in Escherichia coli. J. Bacteriol 2005, 187, 2066–2076. [Google Scholar]
- Zhou, X.; Meng, X.; Sun, B. An eal domain protein and cyclic amp contribute to the interaction between the two quorum sensing systems in Escherichia coli. Cell Res 2008, 18, 937–948. [Google Scholar]
- González Barrios, A.F.; Zuo, R.; Hashimoto, Y.; Yang, L.; Bentley, W.E.; Wood, T.K. Autoinducer 2 controls biofilm formation in Escherichia coli through a novel motility quorum-sensing regulator (mqsr, b3022). J. Bacteriol 2006, 188, 305–316. [Google Scholar]
- Yang, Y.X.; Xu, Z.H.; Zhang, Y.Q.; Tian, J.; Weng, L.X.; Wang, L.H. A new quorum-sensing inhibitor attenuates virulence and decreases antibiotic resistance in Pseudomonas aeruginosa. J. Microbiol 2012, 50, 987–993. [Google Scholar]
- Geske, G.D.; O’Neill, J.C.; Miller, D.M.; Mattmann, M.E.; Blackwell, H.E. Modulation of bacterial quorum sensing with synthetic ligands: Systematic evaluation of n-acylated homoserine lactones in multiple species and new insights into their mechanisms of action. J. Am. Chem. Soc 2007, 129, 13613–13625. [Google Scholar]
- Mattmann, M.E.; Shipway, P.M.; Heth, N.J.; Blackwell, H.E. Potent and selective synthetic modulators of a quorum sensing repressor in Pseudomonas aeruginosa identified from second-generation libraries of N-acylated l-homoserine lactones. ChemBioChem 2011, 12, 942–949. [Google Scholar]
- Amara, N.; Mashiach, R.; Amar, D.; Krief, P.; Spieser, S.A.H.; Bottomley, M.J.; Aharoni, A.; Meijler, M.M. Covalent inhibition of bacterial quorum sensing. J. Am. Chem. Soc 2009, 131, 10610–10619. [Google Scholar]
- Ganguly, K.; Wu, R.; Ollivault-Shiflett, M.; Goodwin, P.M.; Silks, L.A.; Iyer, R. Design, synthesis, and a novel application of quorum-sensing agonists as potential drug-delivery vehicles. J. Drug Target 2010, 19, 528–539. [Google Scholar]
- Lu, C.; Kirsch, B.; Zimmer, C.; de Jong, J.C.; Henn, C.; Maurer, C.K.; Müsken, M.; Häussler, S.; Steinbach, A.; Hartmann, R.W. Discovery of antagonists of pqsr, a key player in 2-alkyl-4-quinolone-dependent quorum sensing in Pseudomonas aeruginosa. Chem. Biol 2012, 19, 381–390. [Google Scholar]
- Rasko, D.A.; Moreira, C.G.; Li de, R.; Reading, N.C.; Ritchie, J.M.; Waldor, M.K.; Williams, N.; Taussig, R.; Wei, S.; Roth, M.; et al. Targeting qsec signaling and virulence for antibiotic development. Science 2008, 321, 1078–1080. [Google Scholar]
- Stacy, D.M.; Welsh, M.A.; Rather, P.N.; Blackwell, H.E. Attenuation of quorum sensing in the pathogen Acinetobacter baumannii using non-native N-acyl homoserine lactones. ACS Chem. Biol 2012, 7, 1719–1728. [Google Scholar]
- Passador, L.; Tucker, K.D.; Guertin, K.R.; Journet, M.P.; Kende, A.S.; Iglewski, B.H. Functional analysis of the Pseudomonas aeruginosa autoinducer pai. J. Bacteriol 1996, 178, 5995–6000. [Google Scholar]
- Smith, K.M.; Bu, Y.; Suga, H. Induction and inhibition of Pseudomonas aeruginosa quorum sensing by synthetic autoinducer analogs. Chem. Biol 2003, 10, 81–89. [Google Scholar]
- Morkunas, B.; Galloway, W.R.J.D.; Wright, M.; Ibbeson, B.M.; Hodgkinson, J.T.; O’Connell, K.M.G.; Bartolucci, N.; Valle, M.D.; Welch, M.; Spring, D.R. Inhibition of the production of the Pseudomonas aeruginosa virulence factor pyocyanin in wild-type cells by quorum sensing autoinducer-mimics. Org. Biomol. Chem 2012, 10, 8452–8464. [Google Scholar]
- Annapoorani, A.; Umamageswaran, V.; Parameswari, R.; Pandian, S.; Ravi, A. Computational discovery of putative quorum sensing inhibitors against lasr and rhlr receptor proteins of Pseudomonas aeruginosa. J. Comput. Aided Mol. Des 2012, 26, 1067–1077. [Google Scholar]
- Lin, Y.-H.; Xu, J.-L.; Hu, J.; Wang, L.-H.; Ong, S.L.; Leadbetter, J.R.; Zhang, L.-H. Acyl-homoserine lactone acylase from ralstonia strain xj12b represents a novel and potent class of quorum-quenching enzymes. Mol. Microbiol 2003, 47, 849–860. [Google Scholar]
- Dong, Y.-H.; Xu, J.-L.; Li, X.-Z.; Zhang, L.-H. Aiia, an enzyme that inactivates the acylhomoserine lactone quorum-sensing signal and attenuates the virulence of erwinia carotovora. Proc. Nat. Acad. Sci 2000, 97, 3526–3531. [Google Scholar]
- Uroz, S.; Chhabra, S.R.; Cámara, M.; Williams, P.; Oger, P.; Dessaux, Y. N-acylhomoserine lactone quorum-sensing molecules are modified and degraded by rhodococcus erythropolis w2 by both amidolytic and novel oxidoreductase activities. Microbiology 2005, 151, 3313–3322. [Google Scholar]
- Reimmann, C.; Ginet, N.; Michel, L.; Keel, C.; Michaux, P.; Krishnapillai, V.; Zala, M.; Heurlier, K.; Triandafillu, K.; Harms, H.; et al. Genetically programmed autoinducer destruction reduces virulence gene expression and swarming motility in Pseudomonas aeruginosa pao1. Microbiology 2002, 148, 923–932. [Google Scholar]
- Bijtenhoorn, P.; Mayerhofer, H.; Müller-Dieckmann, J.; Utpatel, C.; Schipper, C.; Hornung, C.; Szesny, M.; Grond, S.; Thürmer, A.; Brzuszkiewicz, E.; et al. A novel metagenomic short-chain dehydrogenase/reductase attenuates Pseudomonas aeruginosa biofilm formation and virulence on caenorhabditis elegans. PLoS One 2011, 6, e26278. [Google Scholar]
- Wahjudi, M.; Papaioannou, E.; Hendrawati, O.; van Assen, A.H.G.; van Merkerk, R.; Cool, R.H.; Poelarends, G.J.; Quax, W.J. Pa0305 of Pseudomonas aeruginosa is a quorum quenching acylhomoserine lactone acylase belonging to the ntn hydrolase superfamily. Microbiology 2011, 157, 2042–2055. [Google Scholar]
- Hong, K.-W.; Koh, C.-L.; Sam, C.-K.; Yin, W.-F.; Chan, K.-G. Quorum quenching revisited—from signal decays to signaling confusion. Sensors 2012, 12, 4661–4696. [Google Scholar]
- De Lamo Marin, S.; Xu, Y.; Meijler, M.M.; Janda, K.D. Antibody catalyzed hydrolysis of a quorum sensing signal found in gram-negative bacteria. BioOrg. Med. Chem. Lett 2007, 17, 1549–1552. [Google Scholar]
- Kaufmann, G.F.; Park, J.; Mee, J.M.; Ulevitch, R.J.; Janda, K.D. The quorum quenching antibody rs2-1g9 protects macrophages from the cytotoxic effects of the Pseudomonas aeruginosa quorum sensing signaling molecule N-3-oxo-dodecanoyl-homoserine lactone. Mol. Immunol 2008, 45, 2710–2714. [Google Scholar]
- Xu, N.; Yu, S.; Moniot, S.; Weyand, M.; Blankenfeldt, W. Crystallization and preliminary crystal structure analysis of the ligand-binding domain of pqsr (mvfr), the Pseudomonas quinolone signal (pqs) responsive quorum-sensing transcription factor of Pseudomonas aeruginosa. Acta Crystallogr. Section F 2012, 68, 1034–1039. [Google Scholar]
- Pustelny, C.; Albers, A.; Büldt-Karentzopoulos, K.; Parschat, K.; Chhabra, S.R.; Cámara, M.; Williams, P.; Fetzner, S. Dioxygenase-mediated quenching of quinolone-dependent quorum sensing in Pseudomonas aeruginosa. Chem. Biol 2009, 16, 1259–1267. [Google Scholar]
- Calfee, M.W.; Coleman, J.P.; Pesci, E.C. Interference with pseudomonas quinolone signal synthesis inhibits virulence factor expression by Pseudomonas aeruginosa. Proc. Natl. Acad. Sci 2001, 98, 11633–11637. [Google Scholar]
- Lesic, B.; Lépine, F.; Déziel, E.; Zhang, J.; Zhang, Q.; Padfield, K.; Castonguay, M.-H.; Milot, S.; Stachel, S.; Tzika, A.A.; et al. Inhibitors of pathogen intercellular signals as selective anti-infective compounds. PLoS Pathog 2007, 3, e126. [Google Scholar]
- Plyuta, V.; Zaitseva, J.; Lobakova, E.; Zagoskina, N.; Kuznetsov, A.; Khmel, I. Effect of plant phenolic compounds on biofilm formation by Pseudomonas aeruginosa. APMIS 2013. [Google Scholar] [CrossRef]
- Adonizio, A.; Leal, S.M.; Ausubel, F.M.; Mathee, K. Attenuation of Pseudomonas aeruginosa virulence by medicinal plants in a Caenorhabditis elegans model system. J. Med. Microbiol 2008, 57, 809–813. [Google Scholar]
- Sarabhai, S.; Sharma, P.; Capalash, N. Ellagic acid derivatives from Terminalia chebula retz. Downregulate the expression of quorum sensing genes to attenuate Pseudomonas aeruginosa pao1 virulence. PLoS One 2013, 8, e53441. [Google Scholar]
- Zimmer, K.R.; Macedo, A.J.; Nicastro, G.G.; Baldini, R.L.; Termignoni, C. Egg wax from the cattle tick Rhipicephalus(Boophilus) microplus inhibits Pseudomonas aeruginosa biofilm. Ticks Tick-Borne Dis. 2013. [Google Scholar] [CrossRef]
- Skindersoe, M.; Ettinger-Epstein, P.; Rasmussen, T.; Bjarnsholt, T.; Nys, R.; Givskov, M. Quorum sensing antagonism from marine organisms. Mar. Biotechnol 2008, 10, 56–63. [Google Scholar]
- Gutierrez, J.A.; Crowder, T.; Rinaldo-Matthis, A.; Ho, M.-C.; Almo, S.C.; Schramm, V.L. Transition state analogs of 5[prime]-methylthioadenosine nucleosidase disrupt quorum sensing. Nat. Chem. Biol 2009, 5, 251–257. [Google Scholar]
- Guo, M.; Gamby, S.; Nakayama, S.; Smith, J.; Sintim, H.O. A pro-drug approach for selective modulation of ai-2-mediated bacterial cell-to-cell communication. Sensors 2012, 12, 3762–3772. [Google Scholar]
- Roy, V.; Fernandes, R.; Tsao, C.-Y.; Bentley, W.E. Cross species quorum quenching using a native ai-2 processing enzyme. ACS Chem. Biol 2009, 5, 223–232. [Google Scholar]
- Vikram, A.; Jesudhasan, P.R.; Pillai, S.D.; Patil, B.S. Isolimonic acid interferes with Escherichia coli o157:H7 biofilm and ttss in qsebc and qsea dependent fashion. BMC Microbiol 2012. [Google Scholar] [CrossRef]
- Ravichandiran, V.; Karthi, S.; Princy, S.A. Screening of sdia inhibitors from melia dubia seed extracts towards the hold back of uropathogenic E. coli quorum sensing-regulated factors. Med. Chem 2012, 8, 1190–1197. [Google Scholar]
- Allison, K.R.; Brynildsen, M.P.; Collins, J.J. Metabolite-enabled eradication of bacterial persisters by aminoglycosides. Nature 2011, 473, 216–220. [Google Scholar]
- Lee, J.H.; Park, J.H.; Kim, J.A.; Neupane, G.P.; Cho, M.H.; Lee, C.S.; Lee, J. Low concentrations of honey reduce biofilm formation, quorum sensing, and virulence in Escherichia coli o157:H7. Biofouling 2011, 27, 1095–1104. [Google Scholar]
- Givskov, M.; de Nys, R.; Manefield, M.; Gram, L.; Maximilien, R.; Eberl, L.; Molin, S.; Steinberg, P.D.; Kjelleberg, S. Eukaryotic interference with homoserine lactone-mediated prokaryotic signaling. J. Bacteriol 1996, 178, 6618–6622. [Google Scholar]
- Ren, D.; Bedzyk, L.A.; Ye, R.W.; Thomas, S.M.; Wood, T.K. Differential gene expression shows natural brominated furanones interfere with the autoinducer-2 bacterial signaling system of Escherichia coli. Biotechnol. Bioeng 2004, 88, 630–642. [Google Scholar]
- Sievert, D.M.; Ricks, P.P.; Edwards, J.R.; Schneider, A.; Patel, J.; Srinivasan, A.; Kallen, A.; Limbago, B.; Fridkin, S.; et al. National Healthcare Safety Network (NHSN) Team. Antimicrobial-resistant pathogens associated with healthcare-associated infections: Summary of data reported to the national healthcare safety network at the centers for disease control and prevention 2009–2010. Infect. Control Hosp. Epidemiol. 2013, 34, 1–14. [Google Scholar]
- Cai, Y.; Chai, D.; Wang, R.; Liang, B.; Bai, N. Colistin resistance of Acinetobacter baumannii: Clinical reports, mechanisms and antimicrobial strategies. J. Antimicrob. Chemother 2012, 67, 1607–1615. [Google Scholar]
- Falagas, M.E.; Kasiakou, S.K.; Saravolatz, L.D. Colistin: The revival of polymyxins for the management of multidrug-resistant gram-negative bacterial infections. Clin. Infect. Dis 2005, 40, 1333–1341. [Google Scholar]
- Derakhshan, S.; Sattari, M.; Bigdeli, M. Effect of cumin (cuminum cyminum) seed essential oil on biofilm formation and plasmid integrity of Klebsiella pneumoniae. Pharmacogn. Mag 2010, 6, 57–61. [Google Scholar]
- Kim, H.-W.; Oh, H.-S.; Kim, S.-R.; Lee, K.-B.; Yeon, K.-M.; Lee, C.-H.; Kim, S.; Lee, J.-K. Microbial population dynamics and proteomics in membrane bioreactors with enzymatic quorum quenching. Appl. Microbiol. Biotechnol. 2013, 97, 4665–4675. [Google Scholar]
- Saroj, S.D.; Rather, P.N. Streptomycin inhibits quorum sensing in Acinetobacter baumannii. Antimicrob. Agents Chemother 2013, 57, 1926–1929. [Google Scholar]
- Hoffman, L.R.; D’Argenio, D.A.; MacCoss, M.J.; Zhang, Z.; Jones, R.A.; Miller, S.I. Aminoglycoside antibiotics induce bacterial biofilm formation. Nature 2005, 436, 1171–1175. [Google Scholar]
- Linares, J.F.; Gustafsson, I.; Baquero, F.; Martinez, J.L. Antibiotics as intermicrobial signaling agents instead of weapons. Proc. Nat. Acad. Sci 2006, 103, 19484–19489. [Google Scholar]
- Singh, V.; Evans, G.B.; Lenz, D.H.; Mason, J.M.; Clinch, K.; Mee, S.; Painter, G.F.; Tyler, P.C.; Furneaux, R.H.; Lee, J.E.; et al. Femtomolar transition state analogue inhibitors of 5′-methylthioadenosine/s-adenosylhomocysteine nucleosidase from Escherichia coli. J. Biol. Chem 2005, 280, 18265–18273. [Google Scholar]
- Schramm, V.L.; Gutierrez, J.A.; Cordovano, G.; Basu, I.; Guha, C.; Belbin, T.J.; Evans, G.B.; Tyler, P.C.; Furneaux, R.H. Transition state analogues in quorum sensing and sam recycling. Nucleic Acids Symp. Ser 2008, 52, 75–76. [Google Scholar]
- Hoang, T.T.; Schweizer, H.P. Characterization of Pseudomonas aeruginosa enoyl-acyl carrier protein reductase (fabi): A target for the antimicrobial triclosan and its role in acylated homoserine lactone synthesis. J. Bacteriol 1999, 181, 5489–5497. [Google Scholar]
- Czajkowski, R.; Jafra, S. Quenching of acyl-homoserine lactone-dependent quorum sensing by enzymatic disruption of signal molecules. Acta Biochim. Pol 2009, 56, 1–16. [Google Scholar]
- Romero, M.; Diggle, S.P.; Heeb, S.; Cámara, M.; Otero, A. Quorum quenching activity in anabaena sp. Pcc 7120: Identification of aiic, a novel ahl-acylase. FEMS Microbiol. Lett 2008, 280, 73–80. [Google Scholar]
- Uroz, S.; Oger, P.M.; Chapelle, E.; Adeline, M.-T.; Faure, D.; Dessaux, Y. A rhodococcus qsda-encoded enzyme defines a novel class of large-spectrum quorum-quenching lactonases. Appl. Environ. Microbiol 2008, 74, 1357–1366. [Google Scholar]
- Chow, J.Y.; Xue, B.; Lee, K.H.; Tung, A.; Wu, L.; Robinson, R.C.; Yew, W.S. Directed evolution of a thermostable quorum-quenching lactonase from the amidohydrolase superfamily. J. Biol. Chem 2010, 285, 40911–40920. [Google Scholar]
- Chow, J.Y.; Yang, Y.; Tay, S.B.; Chua, K.L.; Yew, W.S. Disruption of biofilm formation by the human pathogen Acinetobacter baumannii using engineered quorum-quenching lactonases; National University of Singapore: Singapore, Unpublished work; 2013. [Google Scholar]
- Dekimpe, V.; Déziel, E. Revisiting the quorum-sensing hierarchy in Pseudomonas aeruginosa: The transcriptional regulator rhlr regulates lasr-specific factors. Microbiology 2009, 155, 712–723. [Google Scholar]
- Defoirdt, T.; Boon, N.; Bossier, P. Can bacteria evolve resistance to quorum sensing disruption? PLoS Pathog 2010, 6, e1000989. [Google Scholar]
- Martinez, J.L.; Baquero, F.; Andersson, D.I. Predicting antibiotic resistance. Nat. Rev. Micro 2007, 5, 958–965. [Google Scholar]
- Diggle, S.P.; Griffin, A.S.; Campbell, G.S.; West, S.A. Cooperation and conflict in quorum-sensing bacterial populations. Nature 2007, 450, 411–414. [Google Scholar]
- Sandoz, K.M.; Mitzimberg, S.M.; Schuster, M. Social cheating in Pseudomonas aeruginosa quorum sensing. Proc. Nat. Acad. Sci 2007, 104, 15876–15881. [Google Scholar]
- Mellbye, B.; Schuster, M. The sociomicrobiology of antivirulence drug resistance: A proof of concept. MBio 2011, 2. [Google Scholar] [CrossRef]


| Bacteria | Receptor(s) | Synthase(s) | Quorum molecule(s) | References |
|---|---|---|---|---|
| V. harveyi | LuxN | LuxM | 3-hydroxy-C4-HSL | [24] |
| LuxP | LuxS | AI-2 | [33] | |
| CqsS | CqsA | CAI-1 | [42] | |
| P. aeruginosa | RhlR | RhlI | C4-HSL | [43] |
| LasR | LasI | 3-oxo-C12-HSL | [43] | |
| QscR | NA | 3-oxo-C12-HSL | [44,45] | |
| PqsR | PqsABCD, PqsH | PQS, HHQ | [40,41] | |
| E. coli | SdiA | N.A. | 3-oxo-C8-HSL | [4,46] |
| LsrB | LuxS | AI-2 | [47,48] | |
| QseC | - - | AI-3/ Epinephrine/Norepinephrine | [49,50] | |
| A. baumannii | AbaR | AbaI | 3-hydroxy-C12-HSL | [51] |
| Enterobacter spp. | - - | - - | C12-HSL | [52] |
| SdiA | NA | - - | [53] | |
| LsrB | LuxS | AI-2 | [47] | |
| K. pneumonia | - - | - - | C8-HSL | [54] |
| - - | - - | C12-HSL | [54] | |
| LsrB | LuxS | AI-2 | [47,55] |
| Inhibitor | Structure | Target bacteria | Target | Effect/value | References |
|---|---|---|---|---|---|
| N-Decanoyl-L-homoserine benzylester (C2) | 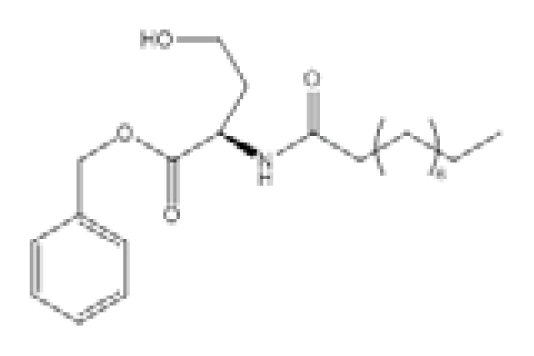 | P. aeruginosa | LasR, RhlR expression | LasR - at 100 μM C2: 29.67% reduction RhlR - at 100 μM C2: 28.20% reduction | [97] |
| Compound D15 | 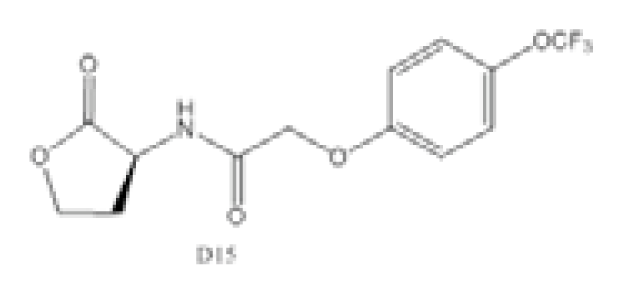 | P. aeruginosa | IC50 | 4.67 μM | [98] |
| Compound Q9 | 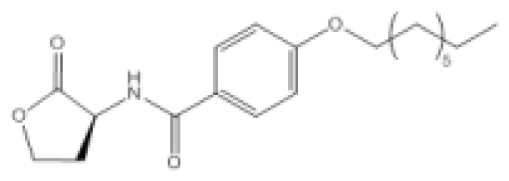 | P. aeruginosa | IC50 | 11 nM | [99] |
| Isothiocyanate-13 (itc-13) |  | P. aeruginosa | IC50 | 45.2 μM | [100] |
| QS0108-ciprofloxacin conjugate | 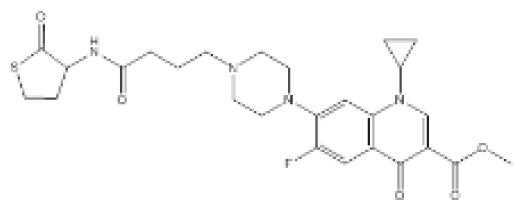 | P. aeruginosa | MIC | 50 μM | [101] |
| Compound 18 |  | P. aeruginosa | IC50 | 259 nM | [102] |
| Compound 19 | 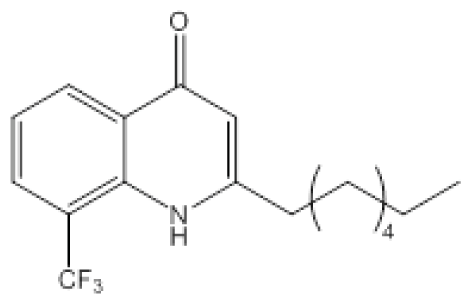 | P. aeruginosa | IC50 | 54 nM | [102] |
| N-Phenyl-4-{[(phenylamino)thioxomethyl]amino}-benzenesulfonamide (LED209) |  | E. coli | IC50 | <10 μM | [103] |
| Compound C8 | 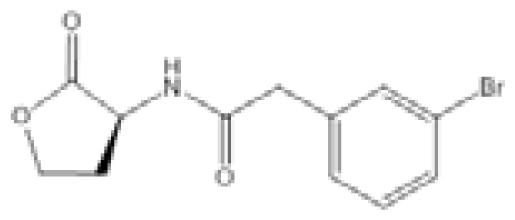 | A. baumannii | IC50 | 5.06 μM | [104] |
| Compound C11 | 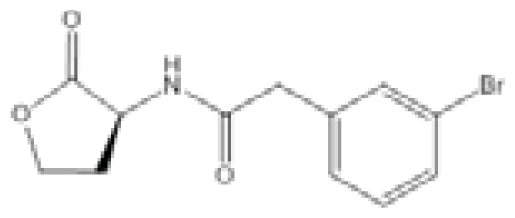 | A. baumannii | IC50 | 2.32 μM | [104] |
© 2013 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Tay, S.B.; Yew, W.S. Development of Quorum-Based Anti-Virulence Therapeutics Targeting Gram-Negative Bacterial Pathogens. Int. J. Mol. Sci. 2013, 14, 16570-16599. https://doi.org/10.3390/ijms140816570
Tay SB, Yew WS. Development of Quorum-Based Anti-Virulence Therapeutics Targeting Gram-Negative Bacterial Pathogens. International Journal of Molecular Sciences. 2013; 14(8):16570-16599. https://doi.org/10.3390/ijms140816570
Chicago/Turabian StyleTay, Song Buck, and Wen Shan Yew. 2013. "Development of Quorum-Based Anti-Virulence Therapeutics Targeting Gram-Negative Bacterial Pathogens" International Journal of Molecular Sciences 14, no. 8: 16570-16599. https://doi.org/10.3390/ijms140816570
APA StyleTay, S. B., & Yew, W. S. (2013). Development of Quorum-Based Anti-Virulence Therapeutics Targeting Gram-Negative Bacterial Pathogens. International Journal of Molecular Sciences, 14(8), 16570-16599. https://doi.org/10.3390/ijms140816570





