Proteome Analysis of Rice (Oryza sativa L.) Mutants Reveals Differentially Induced Proteins during Brown Planthopper (Nilaparvata lugens) Infestation
Abstract
:1. Introduction
2. Results
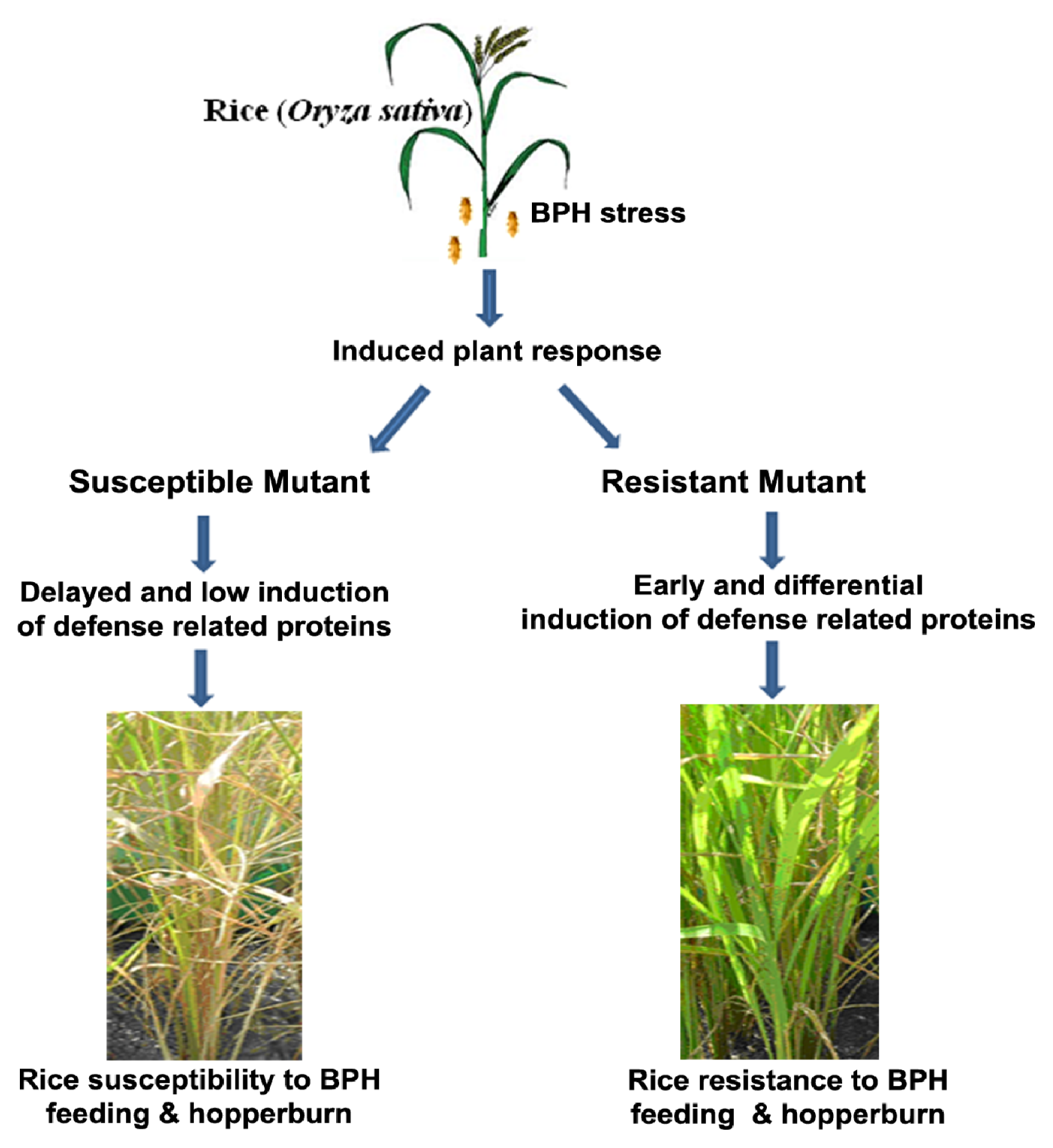
2.1. Rice Phenotype during BPH Stress
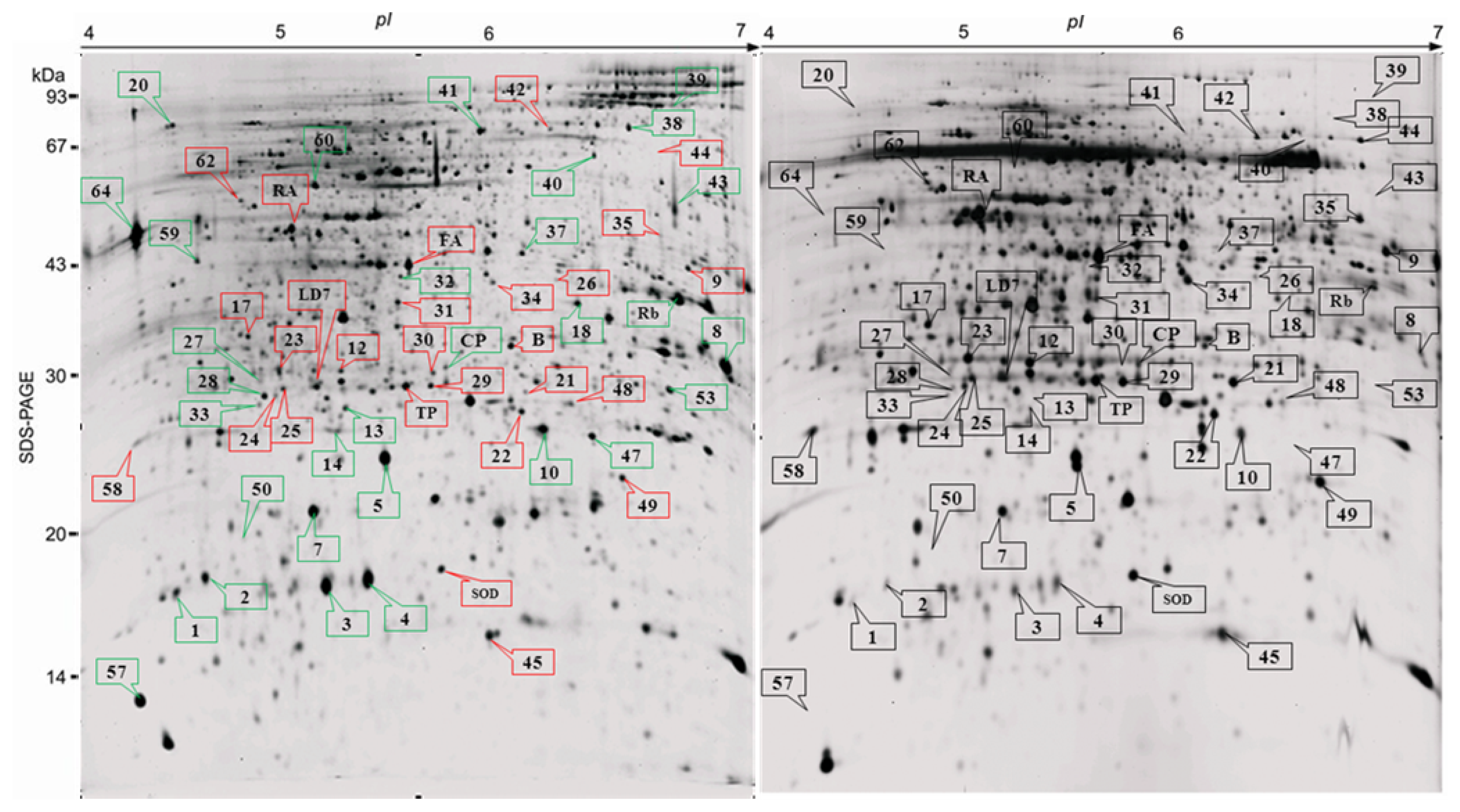
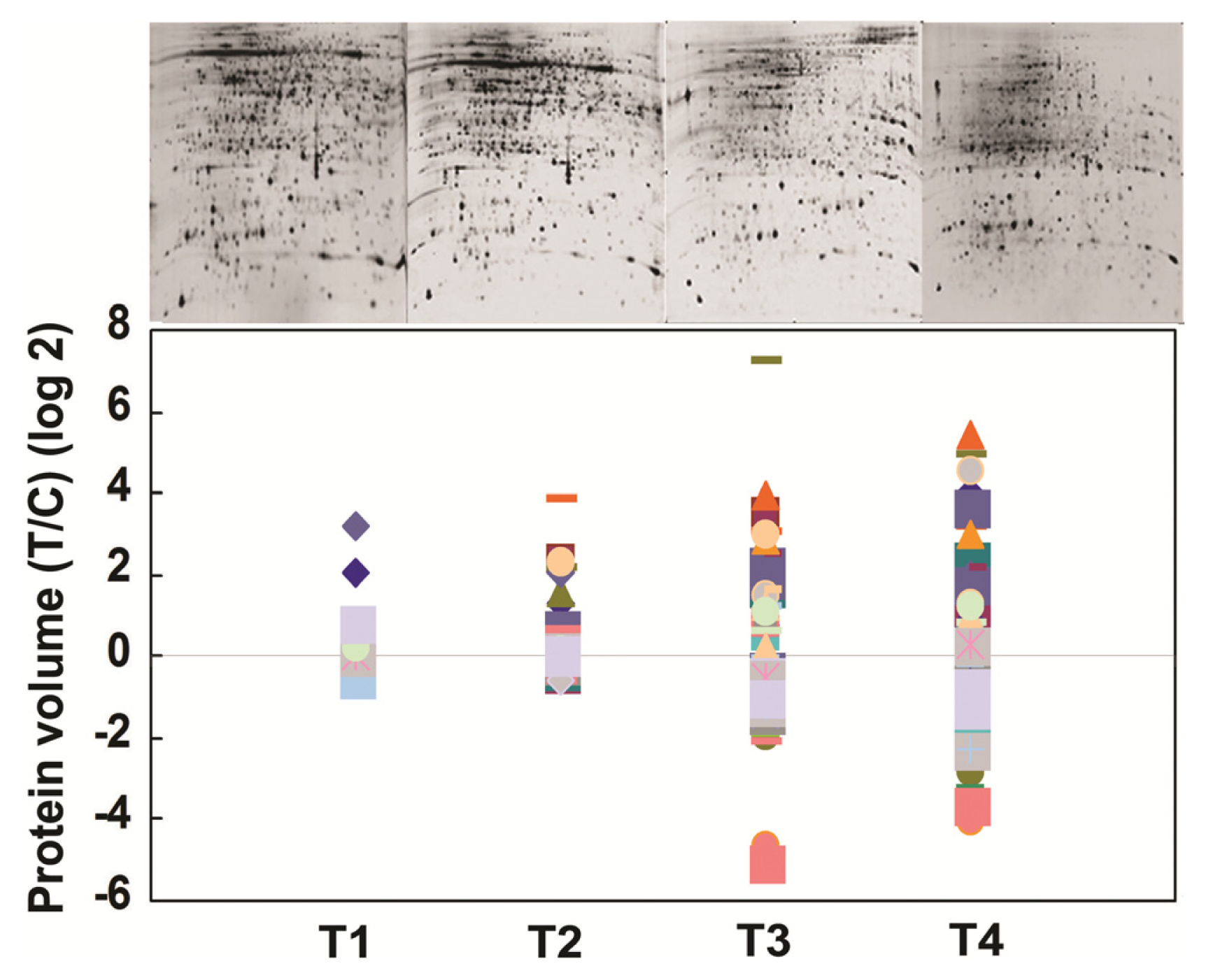
2.2. Proteome Analysis of BPH Induced Proteins in IR64
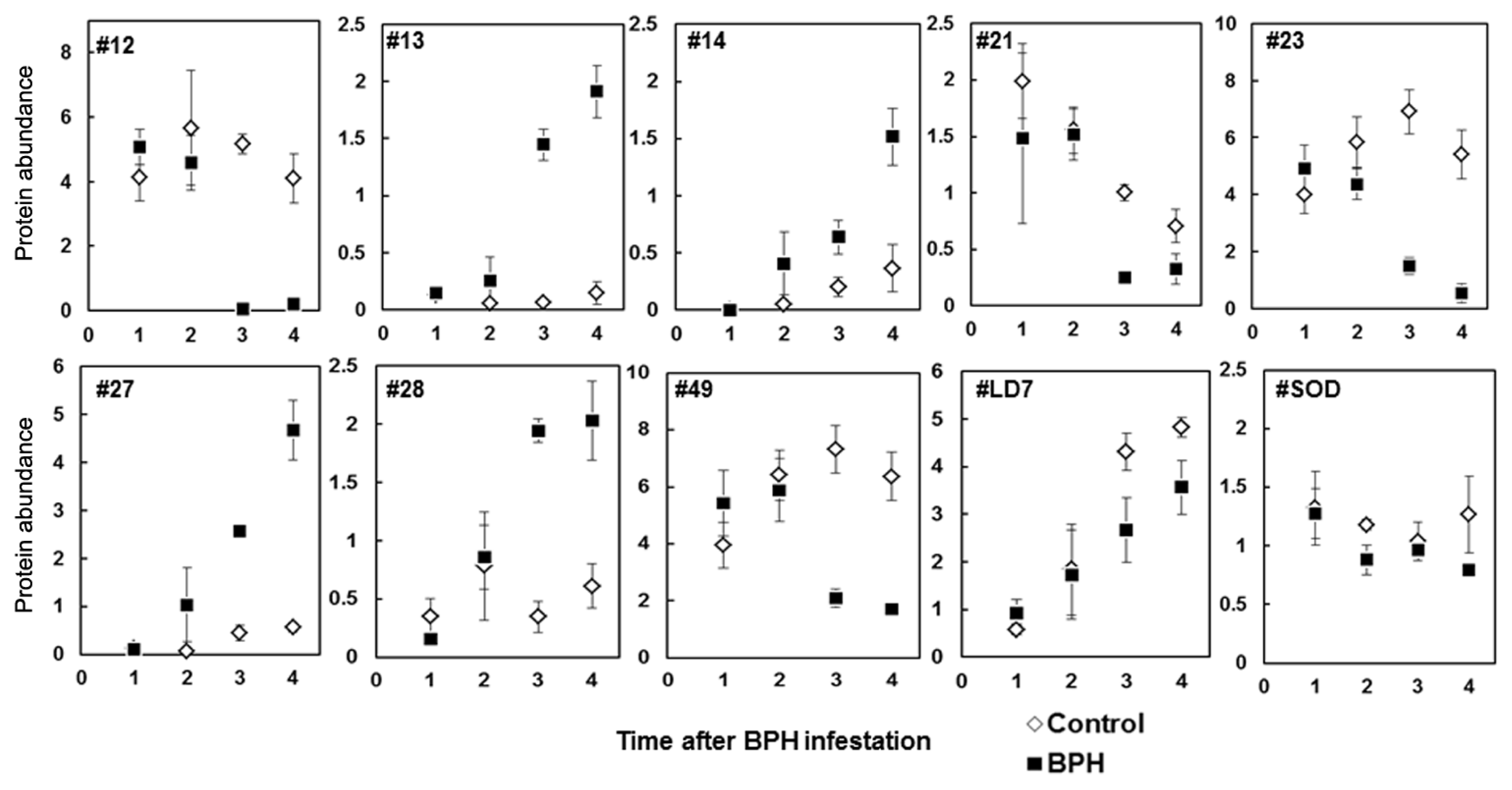
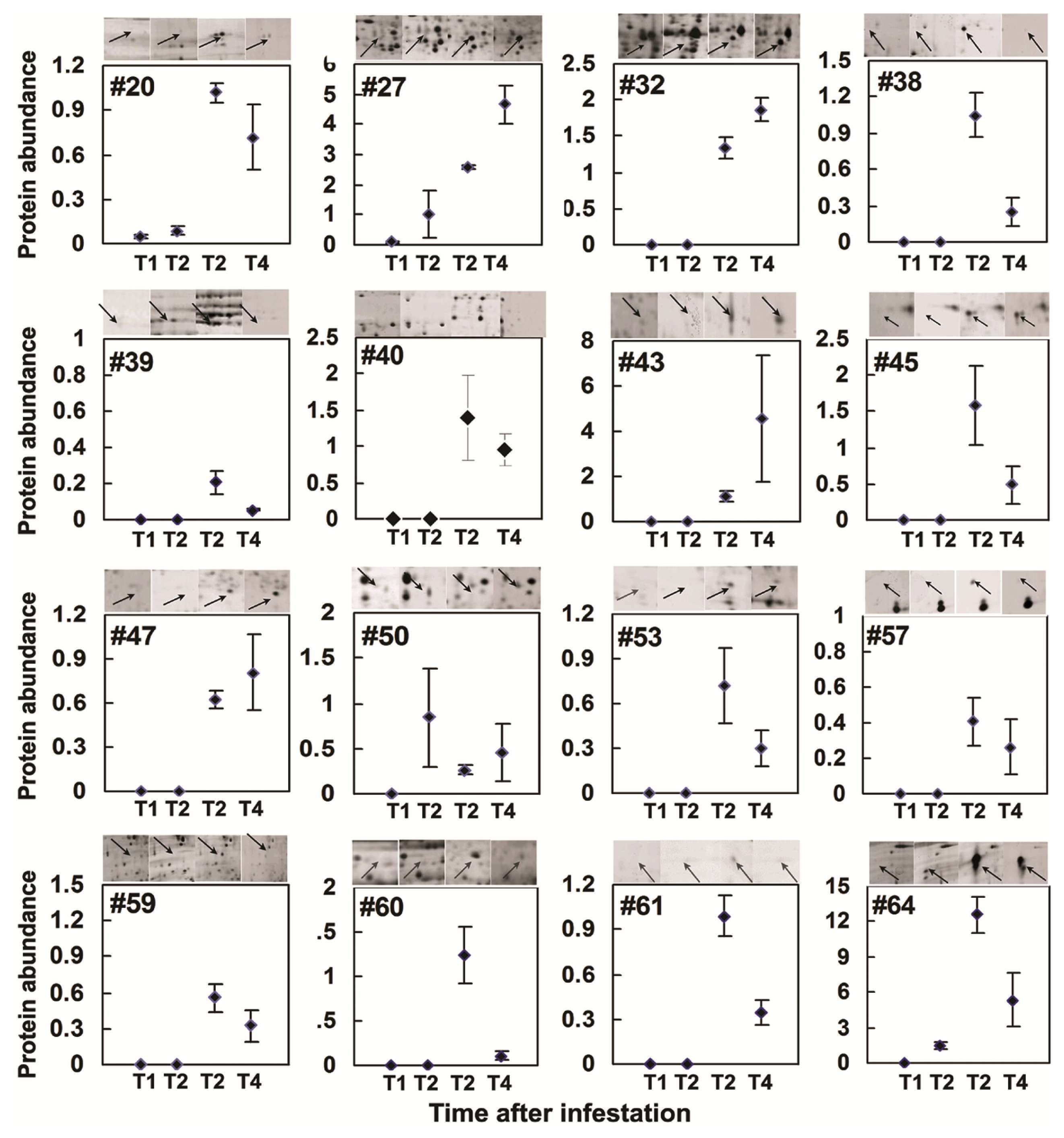
2.3. Rice Proteins Induced in BPH Infested Plants
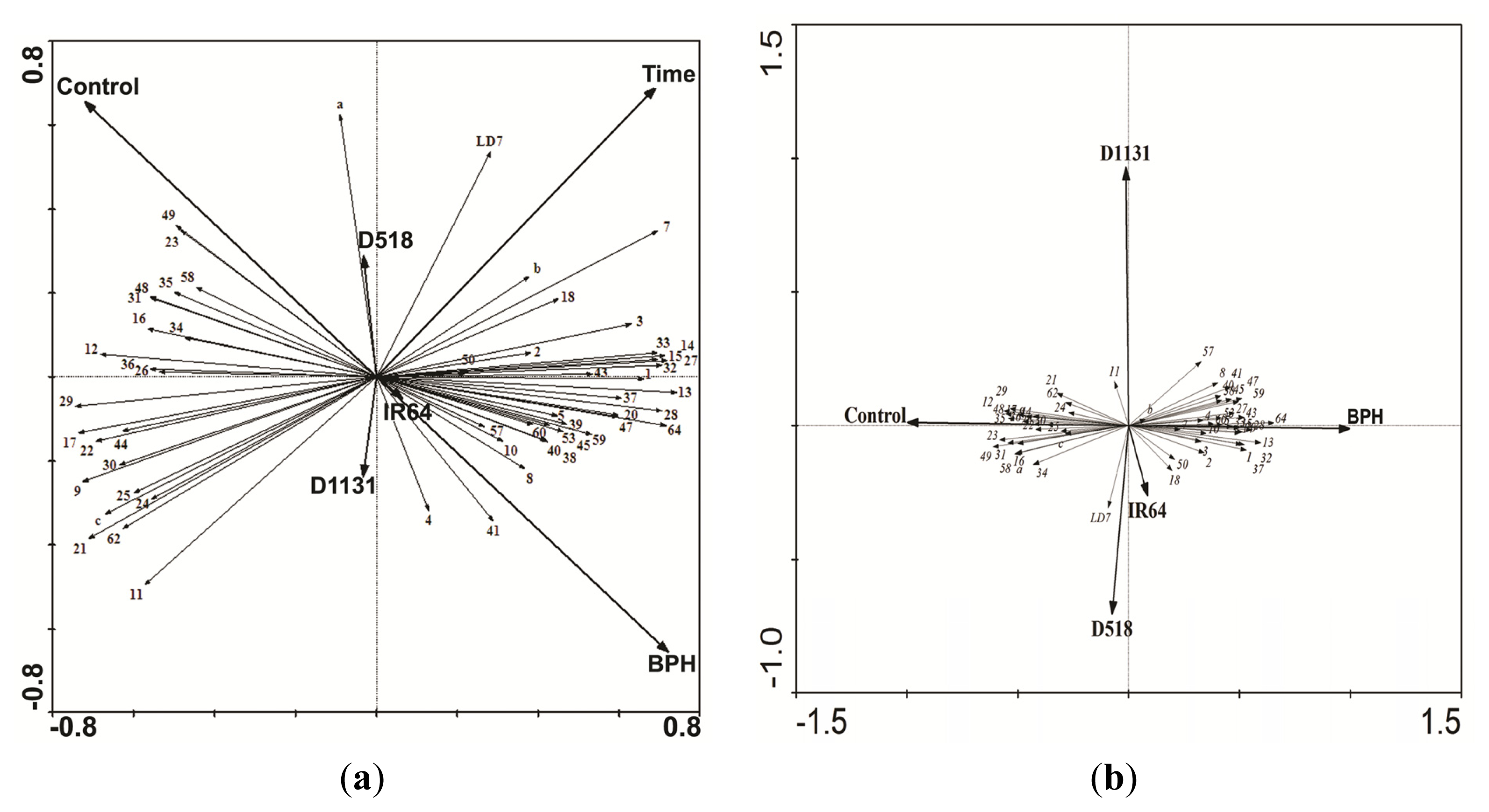
2.4. Comparative Proteomics of IR64 and Mutants
3. Discussion
4. Experimental Section
4.1. Insect Culture and Plant Material
4.2. Plant Phenotype to BPH Infestation
4.3. Proteomics Response after BPH Infestation
4.4. Protein Analysis
4.5. Protein Identification
4.6. Statistical Analysis
5. Conclusions
Acknowledgments
References
- Walling, L.L. The myriad plant responses to herbivores. J. Plant Growth Regul 2000, 19, 195–216. [Google Scholar]
- Wu, J.; Baldwin, I.T. Herbivory-induced signalling in plants: Perception and action. Plant Cell Environ 2009, 32, 1161–1174. [Google Scholar]
- Sarmento, R. A.; Lemos, F.; Dias, C. R.; Kikuchi, W. T.; Rodrigues, J.C. P.; Pallini, A.; Sabelis, M. W.; Janssen, A. A herbivorous mite down-regulates plant defence and produces web to exclude competitors. PLoS One 2011, 6. [Google Scholar] [CrossRef] [Green Version]
- Rakwal, R.; Komatsu, S. Role of jasmonate in the rice (Oryza sativa L.) self defense mechanism using proteome analysis. Electrophoresis 2000, 21, 2492–2500. [Google Scholar]
- Wu, J.; Baldwin, I.T. New insights into plant responses to the attack from insect herbivores. Ann. Rev. Genet 2010, 44, 1–24. [Google Scholar]
- Brar, D. S.; Khush, G.S. Utilization of Wild Species of Genus Oryza in Rice Improvement. In Monograph on Genus Oryza. Plymouth; Nanda, J.S., Sharma, S.D., Eds.; Science Publishers: Enfield, UK, 2003; pp. 283–309. [Google Scholar]
- Zhang, F.; Zhu, L.; He, G. Differential gene expression in response to brown planthopper feeding in rice. J. Plant Physiol 2004, 161, 53–62. [Google Scholar]
- Wang, X.- L.; He, R. F.; He, G.C. Construction of suppression subtractive hybridization libraries and identification of brown planthopper-induced genes. J. Plant Physiol. 2005, 162, 1254–1262. [Google Scholar]
- Wei, Z.; Hu, W.; Lin, Q.; Cheng, X.; Tong, M.; Zhu, L.; Chen, R; He, G. Understanding rice plant resistance to the Brown Planthopper (Nilaparvata lugens): A proteomic approach. Proteomics 2009, 9, 2798–2808. [Google Scholar]
- Park, D. S.; Lee, S. K.; Lee, J. H.; Song, M. Y.; Song, S. Y.; Kwak, D. Y.; Yeo, U. S.; Jeon, N. S.; Park, S. K.; Yi, G.; et al. The identification of candidate rice genes that confer resistance to the brown planthopper (Nilaparvata lugens) through representational difference analysis. Theor. Appl. Genet 2007, 115, 537–547. [Google Scholar]
- Liu, C.; Hao, F.; Hu, J.; Zhang, W.; Wan, L.; Zhu, L.; Tang, H.; He, G. Revealing different systems responses to brown planthopper infestation for pest susceptible and resistant rice plants with the combined metabonomic and gene-expression analysis. J. Proteome Res 2010, 9, 6774–6785. [Google Scholar]
- Zhang, Q. Strategies for developing green super rice. Proc. Natl. Acad. Sci. USA 2007, 104, 16402–16409. [Google Scholar]
- Alam, S. N.; Cohen, M.B. Detection and analysis of QTLs for resistance to the brown planthopper, Nilaparvata lugens in a double-haploid rice production. Theor. Appl. Genet 1998, 97, 1370–1379. [Google Scholar]
- Yuan, H.; Chen, X.; Zhu, L.; He, G. Identification of genes responsive to brown planthopper Nilaparvata lugens Stål (Homoptera: Delphacidae) feeding in rice. Planta 2005, 221, 105–112. [Google Scholar]
- Wang, Y. Y.; Wang, X. L.; Yuan, H.; Chen, R.; Zhu, L.; He, R.; He, G. Responses of two contrasting genotypes of rice to brown planthopper. Mol. Plant Microbe Interact 2008, 21, 122–132. [Google Scholar]
- Hao, P.; Liu, C.; Wang, Y.; Chen, R.; Tang, M.; Du, B.; Zhu, L.; He, G. Herbivore-induced callose deposition on the sieve plates of rice: an important mechanism for host resistance. Plant Physiol 2008, 146, 1810–1820. [Google Scholar]
- Agrawal, G. K.; Rakwal, R. Rice proteomics: A move toward expanded proteome coverage to comparative and functional proteomics uncovers the mysteries of rice and plant biology. Proteomics 2011, 11, 1630–1649. [Google Scholar]
- Kim, S. T.; Kim, S. G.; Kang, Y. H.; Wang, Y.; Kim, J. Y.; Yi, N.; Kim, J. K.; Rakwal, R.; Koh, H. J.; Kang, K.Y. Proteomics analysis of rice lesion mimic mutant (spl1) reveals tightly localized probenazole-induced protein (PBZ1) in cells undergoing programmed cell death. J. Proteome Res 2008, 7, 1750–1760. [Google Scholar]
- Tsunezuka, H.; Fujiwara, M.; Kawasaki, T.; Shimamoto, K. Proteome analysis of programmed cell death and defense signaling using the rice lesion mimic mutant cdr2. Mol. Plant Microbe Interact 2005, 18, 52–59. [Google Scholar]
- Jung, Y. H.; Rakwal, R.; Agrawal, G. K.; Shibato, J.; Kim, J. A.; Lee, M. O.; Choi, P. K.; Jung, S. H.; Kim, S. H.; Koh, H.J.; et al. Differential expression of defense/stress-related marker proteins in leaves of a unique rice blast lesion mimic mutant (blm). J. Proteome Res 2006, 5, 2586–2598. [Google Scholar]
- Kang, S. G.; Matin, M. N.; Bae, H.; Natarajan, S. Proteome analysis and characterization of phenotypes of lesion mimic mutant spotted leaf 6 in rice. Proteomics 2007, 7, 2447–2458. [Google Scholar]
- Lee, J.; Bricker, T. M.; Lefevre, M.; Pinson, S. R.; Oard, J.H. Proteomic and genetic approaches to identifying defence-related proteins in rice challenged with the fungal pathogen Rhizoctonia solani. Mol. Plant Pathol 2006, 7, 405–416. [Google Scholar]
- Sangha, J. S.; Chen, Y. H.; Palchamy, K.; Jahn, G. C.; Maheswaran, M.; Adalla, C. B.; Leung, H. Categories and inheritance of resistance to Nilaparvata lugens (Hemiptera: Delphacidae) in mutants of indica rice ‘IR64’. J. Econ. Entomol 2008, 101, 575–583. [Google Scholar]
- Schiltz, S.; Gallardo, K.; Huart, M.; Negroni, L.; Sommerer, N.; Burstin, J. Proteome reference maps of vegetative tissues in pea. An investigation of nitrogen mobilization from leaves during seed filling. Plant Physiol 2004, 135, 2241–2260. [Google Scholar]
- Salekdeh, H. G.; Siopongco, J.; Wade, L. J.; Ghareyazie, B.; Bennett, J. Proteomic analysis of rice leaves during drought stress and recovery. Proteomics 2002, 2, 1131–1145. [Google Scholar]
- Kraepiel, Y.; Pédron, J.; Patrit, O.; Simond-Côte, E.; Hermand, V.; van Gijsegem, F. Analysis of the Plant bos1 Mutant Highlights Necrosis as an Efficient Defence Mechanism during D. dadantii/Arabidospis thaliana Interaction. PLoS One 2011, 6. [Google Scholar] [CrossRef]
- Watanabe, T.; Kitagawa, H. Photosynthesis and translocation of assimilates in rice plants following phloem feeding by the planthopper Nilaparvata lugens (Homoptera: Delphacidae). J. Econ. Entomol 2000, 93, 1192–1198. [Google Scholar]
- Zhu-Salzman, K.; Salzman, R. A.; Ahn, J.- E.; Koiwa, H. Transcriptional regulation of sorghum defense determinants against a phloem-feeding aphid. Plant Physiol. 2004, 134, 420–431. [Google Scholar]
- Scheideler, M.; Schlaich, N. L.; Fellenberg, K.; Beissbarth, T.; Hauser, N. C.; Vingron, M.; Slusarenko, A. J.; Hoheisel, J.D. Monitoring the switch from housekeeping to pathogen defense metabolism in Arabidopsis thaliana using cDNA arrays. J. Biol. Chem 2002, 277, 10555–10561. [Google Scholar]
- Bilgin, D. D.; Zavala, J. A.; Zhu, J.; Clough, S. J.; Ort, D. R.; DeLucia, E.H. Biotic stress globally downregulates photosynthesis genes. Plant Cell Environ 2010, 33, 1597–1613. [Google Scholar]
- Nakano, R.; Ishida, H.; Makino, A.; Mae, T. In vivo fragmentation of the large subunit of ribulose-1,5-bisphosphate carboxylase by reactive oxygen species in an intact leaf of cucumber under chilling-light conditions. Plant Cell Physiol 2006, 47, 270–276. [Google Scholar]
- Navarre, D.; Wolpert, T.J. Victorin induction of an apoptotic/senescence-like response in oats. Plant Cell 1999, 11, 237–250. [Google Scholar]
- Matile, P. Chloroplast Senescence. In Crop Photosynthesis: Spatial and Temporal Determinant; Baker, N.R., Thomas, H., Eds.; Elsevier: Amsterdam, The Netherlands, 1992; pp. 413–440. [Google Scholar]
- Evans, J. R.; Seeman, J.R. The Allocation of Protein Nitrogen in the Photosynthetic Apparatus: Cost, Consequence and Control. In Photosynthesis; Briggs, W.R., Ed.; Alan R. Liss Inc: New York, NY, USA, 1989; pp. 183–205. [Google Scholar]
- Asada, K. Ascorbate peroxidase: A hydrogen peroxidase scavenging system in plants. Physiol. Plant 1992, 85, 235–241. [Google Scholar]
- Kocsy, G.; von Ballmoos, P.; Ruegsegger, A.; Szalai, G.; Galiba, G.; Brunold, C. Increasing the glutathione content in a chilling-sensitive maize genotype using safeners increased protection against chilling-induced injury. Plant Physiol 2001, 127, 1147–1156. [Google Scholar]
- Zhu-Salzman, K.; Luthe, D. S.; Felton, G.W. Arthropod-inducible proteins: Broad spectrum defenses against multiple herbivores. Plant Physiol 2008, 146, 852–858. [Google Scholar]
- Barbehenn, R.; Dukatz, C.; Holt, C.; Reese, A.; Martiskainen, O.; Salminen, J. P.; Yip, L.; Tran, L.; Constabel, C.P. Feeding on poplar leaves by caterpillars potentiates foliar peroxidase action in their guts and increases plant resistance. Oecologia 2010, 164, 993–1004. [Google Scholar]
- Kerchev, P. I.; Fenton, B.; Foyer, C. H.; Hancock, R.D. Plant responses to insect herbivory: Interactions between photosynthesis, reactive oxygen species and hormonal signalling pathways. Plant Cell Environ 2012, 35, 441–453. [Google Scholar]
- Yoshimura, K.; Yabuta, Y.; Ishikawa, T.; Shigeoka, S. Expression of spinach ascorbate peroxidase isoenzymes in response to oxidative stresses. Plant Physiol 2000, 123, 223–234. [Google Scholar]
- Kim, J. K.; Bamba, T.; Harada, K.; Fukusaki, E.; Kobayashi, A. Time-course metabolic profiling in Arabidopsis thaliana cell cultures after salt stress treatment. J. Exp. Bot 2007, 58, 415–424. [Google Scholar]
- Shulaev, V.; Cortes, D.; Miller, G.; Mittler, R. Metabolomics for plant stress response. Physiol. Plant 2008, 132, 199–208. [Google Scholar]
- Baxter, C. J.; Redestig, H.; Schauer, N.; Repsilber, D.; Patil, K. R.; Nielsen, J.; Selbig, J.; Liu, J.; Fernie, A. R.; Sweetlove, L.J. The metabolic response of heterotrophic arabidopsis cells to oxidative stress. Plant Physiol 2007, 143, 312–325. [Google Scholar]
- Cabrera, H. M.; Argandona, V. H.; Corcuera, L.J. Metabolic changes in barley seedlings at different aphid infestation levels. Phytochemistry 1994, 35, 317–319. [Google Scholar]
- Divol, F.; Vilaine, F.; Thibivilliers, S.; Amselem, J.; Palauqui, J. C.; Kusiak, C.; Dinant, S. Systemic response to aphid infestation by Myzus persicae in the phloem of Apium graveolens. Plant Mol. Biol 2005, 57, 517–540. [Google Scholar]
- Veena-Reddy, V. S.; Sopory, S.K. Glyoxalase I from Brassica juncea: Molecular cloning, regulation and its over-expression confer tolerance in transgenic tobacco under stress. Plant J 1999, 17, 385–395. [Google Scholar]
- Caldas, T. D.; El Yaagoubi, A.; Richarme, G. Chaperone properties of bacterial elongation factor EF-Tu. J. Biol. Chem 1998, 273, 11478–11482. [Google Scholar]
- Bhadula, S. K.; Elthon, T. E.; Habben, J. E.; Helentjaris, T. G.; Jiao, S.; Ristic, Z. Heat stress induced synthesis of chloroplast protein synthesis elongation factor (EF-Tu) in a heat-tolerant maize line. Planta 2001, 212, 359–366. [Google Scholar]
- Bariola, P. A.; Howard, C. J.; Taylor, C. B.; Verburg, M. T.; Jaglan, V. D.; Green, P.J. The Arabidopsis ribonuclease gene RNS1 is tightly controlled in response to phosphate limitation. Plant J 1994, 6, 673–685. [Google Scholar]
- Galiana, E.; Bonnet, P.; Conrod, S.; Keller, H.; Panabières, F.; Ponchet, M.; Poupet, A.; Ricci, P. RNase activity prevents the growth of a fungal pathogen in tobacco leaves and increases upon induction of systemic acquired resistance with elicitin. Plant Physiol 1997, 115, 1557–1567. [Google Scholar]
- Lers, A.; Sonego, L.; Green, P. J.; Burd, S. Suppression of LX ribonuclease in tomato results in a delay of leaf senescence and abscission. Plant Physiol 2006, 142, 710–721. [Google Scholar]
- Okabe, T.; Yoshimoto, I.; Hitoshi, M.; Ogawa, T.; Ohyama, T. An S-like ribonuclease gene is used to generate a trap-leaf enzyme in the carnivorous plant Drosera adelae. FEBS Lett 2005, 579, 5729–5733. [Google Scholar]
- Hugot, K.; Ponchet, M.; Marais, A.; Ricci, P.; Galiana, E. A tobacco S-like RNase inhibits hyphal elongation of plant pathogens. Mol. Plant-Microbe Interact 2002, 15, 243–250. [Google Scholar]
- Damerval, C.; de Vienne, D.; Zivy, M.; Thiellement, H. Technical improvements in two-dimensional electrophoresis increase the level of genetic variation detected in wheat-seedling proteins. Electrophoresis 1986, 7, 52–54. [Google Scholar]
- Australian Proteome Analysis Facility. Available online: http://www.proteome.org.au/Services/Mass-Spectrometry/default.aspx accessed on 4 February 2013.
- Mascot Search Engine. Available online: http://www.matrixscience.com accesses on 4 February 2013.
- Ter Braak, C.J. F.; Smilauer, P. CANOCO Reference Manual and CanoDraw for Windows User’s Guide: Software for Canonical Community Ordination (version 4.5); Microcomputer Power: Ithaca, NY, USA, 2002; p. 500. [Google Scholar]
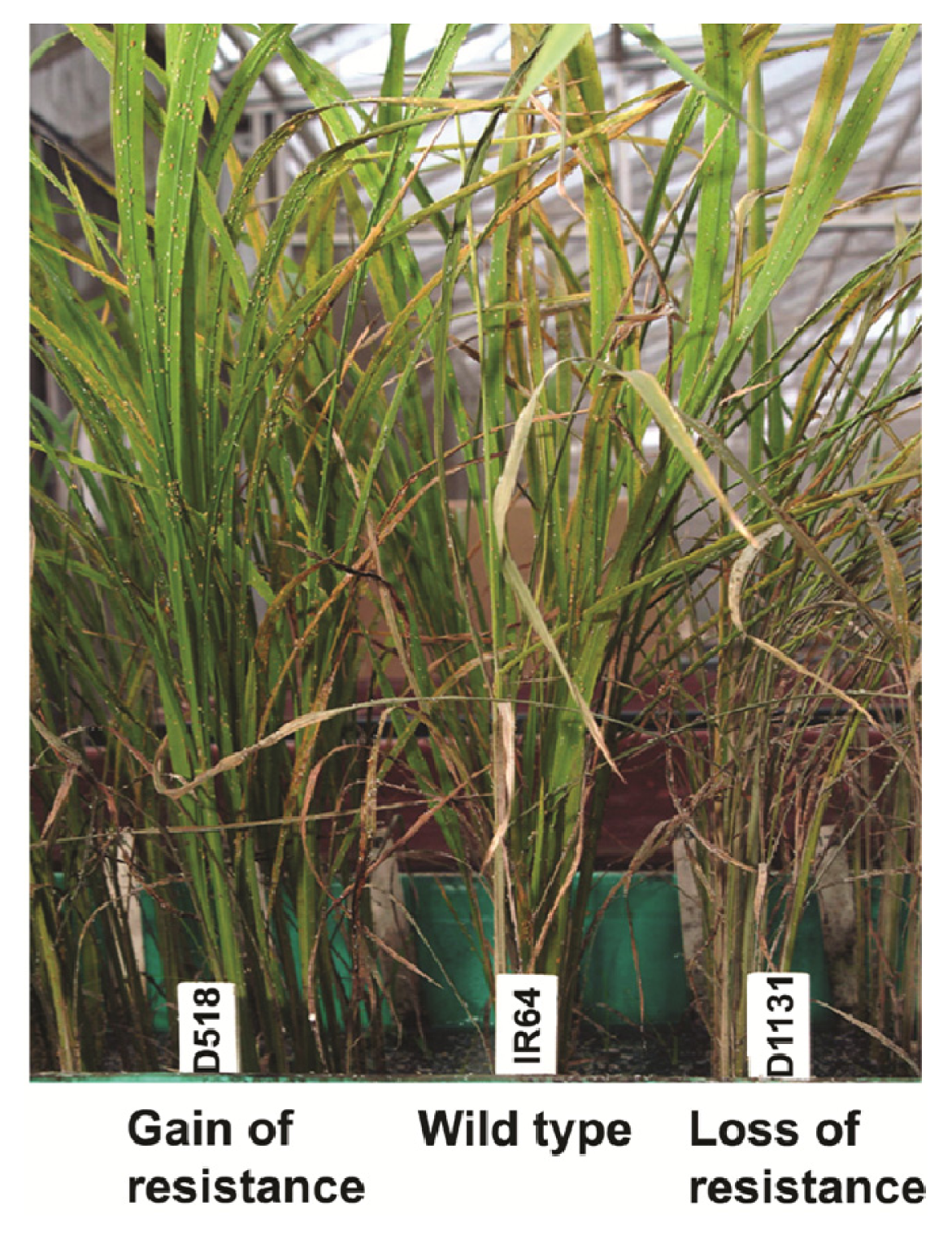


| Rice line | BPH damage (1–9 scale) | |||
|---|---|---|---|---|
| T1 | T2 | T3 | T4 | |
| IR64 | 1.0 ± 0.0 | 1.6 ± 0.55 | 3.6 ± 0.55 | 5.2 ± 0.85 |
| D518 | 1.0 ± 0.0 | 1.4 ± 0.48 | 3.0 ± 0.76 | 3.6 ± 0.56 |
| D1131 | 1.0 ± 0.0 | 1.8 ± 0.59 | 4.8 ± 0.65 | 6.8 ± 0.66 |
| Spot | PM (%C) | Identity/source | Accession | Exp. (Theo.) Mr | Exp. (Theo.) pI | Mascot score | Fold change | P-value |
|---|---|---|---|---|---|---|---|---|
| Energy/pentose phosphate | ||||||||
| 1 | 2(4) | Rubisco large subunit | gi11955 | 17.2(52.8) | 4.5(6.13) | 64 | >10 ↑ | 0.047 |
| 2 | 1(4) | Rubisco large subunit | gi476752 | 17.3(45.1) | 4.6(8.4) | 104 | >10 ↑ | 0.006 |
| 3 | 2(5) | Ribulose bisphosphate carboxylase/oxygenase large chain | gi11466795 | 17.2(52.8) | 5.1(6.2) | 98 | 4.56 ↑ | 0.008 |
| 5 | 3(9) | Rubisco large subunit from chromosome 10 chloroplast insertion | gi37533338 | 23.7(52.8) | 5.4(6.4) | 128 | 1.53 ↑ | 0.130 |
| 10 | 5(13) | Rubisco large subunit | gi476752 | 24.9(45.1) | 6.1(8.4) | 174 | >10 ↑ | 0.005 |
| RA* | – | Ribulose bis phosphate carboxylase/oxygenase activase | P93431 | 47(42.07) | 5.0(5.0) | – | 11.45 ↓ | 0.0019 |
| Rb | 3(9) | Rubisco large subunit | gi2734976 | 34.1(43.7) | 6.3 | 332 | 3.35 ↑ | 0.006 |
| Energy/photosynthesis | ||||||||
| 61 | 1(33) | Putative oxygen evolving enhancer protein 3-1 chloroplast precursor | gi50938199 | 18.7(22.9) | 9.8(9.8) | 114 | Ind ↑ | 0.0053 |
| 34 | 13 (38) | Ferredoxin-NADP (H) oxidoreductase | Q6ZFJ3_ORYSA | 36.0(40.8) | 5.9(7.9) | 90 | 4.21 ↓ | 0.0114 |
| 63 | 3(44) | Putative oxygen evolving enhancer protein 3-1 chloroplast precursor | gi50938199 | 14.5(22.9) | 9.9(9.8) | 400 | 2.57 ↓ | 0.0053 |
| Energy/glycolysis | ||||||||
| 32 | 2(8) | Enolase | gi33113259 | 37.8(47.9) | 5.5(5.4) | 77 | Ind ↑ | 0.0009 |
| 37 | 14 (49) | Enolase | gi780372 | 39.9(47.9) | 6.7(5.4) | 104 | 7.95 ↑ | 0.0522 |
| TP | 8(25) | Triose phosphate isomerase, cytosolic | P48494 | 27.5(27.1) | 5.6(5.4) | 70 | <10 ↓ | <0.0001 |
| 9 | 6(33) | Glyceraldehyde-3- phosphate dehydrogenase, cytosolic | G3PC_HORV | 37.4(33.2) | 6.7(6.2) | 258 | 9.6 ↓ | 0.0090 |
| 44 | 6(27) | Putative dihydrolipoamide dehydrogenase precursor | gi34894800 | 57.0(52.6) | 6.6(7.2) | 111 | <10 ↓ | 0.0011 |
| 35 | 3(17) | Formate dehydrogenase | gi51536124 | 41.2(41.3) | 6.6(6.7) | 100 | <10 ↓ | 0.0005 |
| FB* | – | Fructose bisphosphate aldolase | Q40677 | 37.7(36.4) | 5.7(5.8) | – | 1.31 ↓ | 0.1620 |
| Energy/electron transport | ||||||||
| 58 | 3(34) | Putative H(+)− transporting ATP synthase | gi50912809 | 25.5(26.2) | 4.3(4.9) | 305 | 3.57 ↓ | 0.202 |
| 30 | 8(50) | Probable ATP synthase 24 kDa subunit | gi50905037 | 28.1(27.2) | 5.5(6.5) | 300 | 4.89 ↓ | 0.0007 |
| Plant defense | ||||||||
| 13 | 6(32) | L-Ascorbate peroxidase 2, cytosolic Oryza sativa subsp. japonica (Rice) | APX2_ORYSJ | 26.3(27.1) | 5.3(5.2) | 65 | >10 ↑ | 0.0005 |
| 14 | 5(36) | Putative ascorbate peroxidase | gi50920595 | 26.2(27.1) | 5.2(5.4) | 94 | 3.16 ↑ | 0.0604 |
| 28 | 5(39) | Ascorbate peroxidase | gi50940199 | 28.0(27.1) | 5.5(5.2) | 239 | 5.55 ↑ | 0.0007 |
| 49 | 5(36) | Putative ascorbate peroxidase | gi50920595 | 22.6(27.1) | 6.5(5.4) | 71 | 3.49 ↓ | 0.0041 |
| 12 | 5(50) | Ascorbate peroxidase | gi50940199 | 29.1(27.1) | 5.2(5.2) | 419 | <10 ↓ | <0.0001 |
| SOD * | – | Superoxide dismutase | P93407 | 17.7(15.7) | 5.8(5.3) | – | 1.47 ↓ | 0.0652 |
| Stress induced | ||||||||
| LD7 | 3(23) | Drought induced S-like RNase protein | gi17105171 | 28.2(28.4) | 5.1(5.2) | 187 | 1.6 ↓ | 0.1033 |
| 27 | 4(38) | Unnamed protein product (Salt stress induced protein) | gi34904362 | 29.1(21.8) | 4.9(4.9) | 153 | Ind ↑ | 0.0003 |
| 21 * | – | GSH-dependent dehydro ascorbate reductase | BAA90672 | 27.0(27.1) | 6.1(5.4) | – | 4.03 ↓ | 0.0009 |
| 23 | 3(40) | Unnamed protein product (Salt stress induced protein) | gi34904362 | 30.3(21.8) | 4.9(4.9) | 179 | 4.6 ↓ | 0.0029 |
| Protein synthesis | ||||||||
| 64 | 3(23) | Chloroplast translation elongation factor Tu1 | gi50910077 | 43.7(50.4) | 4.3(6.19) | 306 | Ind ↑ | 0.0012 |
| 4 | 3(45) | Putative ribosomal protein s12 | gi50934241 | 17.2(14.8) | 5.3(5.3) | 290 | 2.64 ↑ | 0.0390 |
| 22 | 8(34) | Putative ribosome recycling factor, chloroplast precursor | XP_478772.1 | 26.4(29.7) | 6.0(9.3) | 66 | 2.77 ↓ | 0.0007 |
| Protein destination and storage | ||||||||
| CP | 6(23) | Putative clp protease | OS02g42290 | 30.2(31.9) | 5.7(6.7) | 70 | 1.77--↑ | 0.0051 |
| 24 | 3(20) | Putative chaperonin 21 precursor | gi51091339 | 27.2(25.4) | 4.9(5.9) | 60 | 4.58 ↓ | 0.0148 |
| Growth and division | ||||||||
| 41 | 4(20) | (O65316) Actin (Mesostigma viride) | ACT_MESVI | 68.3(41.5) | 5.8(5.3) | 181 | 7.27 ↑ | 0.0097 |
| Secondary metabolism | ||||||||
| 47 | 3(21) | Putative 1,4-benzoquinone reductase | gi34910128 | 24.7(21.7) | 6.3(6.0) | 79 | Ind ↑ | 0.0004 |
| 26 | 4(20) | Putative NADPH-dependent mannose 6-phosphate reductase | gi50904895 | 36.3(35.4) | 6.2(5.9) | 142 | >10 ↓ | 0.0106 |
| 31 | 4(23) | Glyoxalase I | gi16580747 | 34.0(32.5) | 5.5(5.5) | 173 | 9.12 ↓ | 0.0004 |
| Miscellaneous | ||||||||
| 20 | 9(15) | Putative proteophosphoglycan | gi50918953 | 74.0(96.8) | 4.4 (10.5) | 104 | Ind ↑ | 0.0001 |
| 53 | 8(41) | Putative defective chloroplasts and leaves (DCL) protein Oryza sativa | Q6UUF7_ORYSA | 30.8(21.3) | 6.9(9.0) | 69 | Ind ↑ | 0.04 |
| 59 | 18 (27) | Putative FH protein NFH2.-Oryza sativa (japonica cultivar-group) | Q8S0F0_ORYSA | 13.3(10.2) | 4.7(8.9) | 66 | Ind ↑ | 0.0099 |
| 60 | 4(53) | Hypothetical protein P0677B10.12 | Q67VJ8_ORYSA | 52.5(12.5) | 5.1(9.3) | 68 | Ind ↑ | 0.0185 |
| 69 | 12 (44) | Putative glyceraldehyde-3- phosphate dehydrogenase (Phosphorylating) Oryza sativa | gi115459078 | 37.2(36.5) | 7.8(7.68) | 94 | 1.75 ↑ | 0.0341 |
| 40 | 9(40) | hypothetical protein OsJ_015102 [Oryza sativa] | gi125591269 | 70.0(25.3) | 6.4(11.0) | 66 | Ind ↑ | 0.0185 |
| B | 6(24) | hypothetical protein OsJ_012934 | gi125589101 | 32.0(35.2) | 6.1(5.3) | 76 | 1.37 ↓ | 0.0277 |
| 39 | 5(3) | Vitellogenin [Nilaparvata lugens] | gi342318865 | 72.5(22.7) | 7.9(8.5) | 64 | Ind ↑ | 0.0372 |
| 39a | 9(12) | Chain E, Leech-Derived Tryptase Inhibitor TRYPSIN COMPLEX | gi3318722 | 97.5(23.4) | 6.7 | 240 | Ind ↑ | 0.0139 |
| 17 | 2(18) | Putative DREPP2 protein | gi50906969 | 32.0(24.0) | 4.8(4.7) | 105 | 8.47 ↓ | 0.0003 |
| 62 | 11 (36) | hypothetical protein OsI_021661 | Q5Z6P9_ORYSA | 47.0(43.0) | 4.7(4.7) | 109 | 3.55 ↓ | 0.0095 |
| 38 | 19 (26) | ATP-dependent DNA helicase UvrD Shewanella denitrificans OS217 | Q3P3H8_9GAMM | 70.0(81.6) | 6.6(5.9) | 78 | Ind ↑ | 0.0050 |
| 42 | 12 (38) | Os12g0420200 [Oryza sativa (japonica cultivar-group)] | gi115488340 | 68.7(41.5) | 6.3(8.5) | 135 | 1.87 ↓ | 0.0476 |
| 68 | 8(22) | Succinyl-CoA ligase [ADP-forming] subunit beta OS = Mesorhizobium sp. (strain BNC1) | SUCC_MESSB | 35.5(42.2) | 7.5(5.0) | 74 | 1.59 ↓ | 0.0327 |
| Protein | Spot | D518 | D1131 | IR64 | Prob. > F |
|---|---|---|---|---|---|
| Unknown | 7 | 1.06 ± 0.07 b,* | 2.18 ± 0.30 a,▲ | 1.17 ± 0.14 b,* | 0.014 |
| GSH-dependent dehydro ascorbate reductase | 21 | 0.81 ± 0.11 a,▼ | 0.47 ± 0.12 b,▼▼ | 0.73 ± 0.01 a,▼ | 0.059 |
| Enolase | 32 | 0.77 ± 0.19 a,▼ | 0.31 ± 0.12 b,▼▼ | 1.24 ± 0.15 a,* | 0.011 |
| Unknown | 43 | 1.04 ± 0.20 b,* | 1.81 ± 0.11 a,▲ | 1.10 ± 0.21 b,* | 0.042 |
| Unknown | 45 | 1.68 ± 0.34 a,b,▲ | 3.46 ± 0.67 a,▲▲ | 1.58 ± 0.54 b,▲ | 0.085 |
| Putative 1,4-benzoquinone reductase | 47 | 0.89 ± 0.19 b,* | 1.85 ± 0.20 a,▲ | 0.62 ± 0.05 b,▼ | 0.004 |
| Putative defective chloroplasts and leaves (DCL) protein Oryza sativa | 53 | 0.67 ± 0.19 b,▼ | 1.76 ± 0.17 a,▲ | 0.72 ± 0.25 b,▼ | 0.018 |
| Unknown | 57 | 0.51 ± 0.06 b,▼ | 3.10 ± 1.42 a,▲ | 0.58 ± 0.13 b▼ | 0.008 |
| Putative FH protein NFH2 Oryza sativa (japonica cultivar-group) | 59 | 0.61 ± 0.06 a,▼ | 1.14 ± 0.10 b | 0.55 ± 0.12 b,▼ | 0.011 |
| Hypothetical protein OsJ_012934 | B | 1.18 ± 0.09 b,* | 1.67 ± 0.10 a,▲ | 0.74 ± 0.07 c,▼ | 0.002 |
| S-like Rnase | LD7 | 2.07 ± 0.37 a,▲ | 0.37 ± 0.07 b,▼ | 0.62 ± 0.15 b,▼ | 0.005 |
| Unknown | 8 | 2.58 ± 0.42 b,▲ | 9.15 ± 1.08 a,▲▲▲ | 9.71 ± 2.44 a,▲▲▲ | 0.030 |
| Glyceraldehyde-3-phosphate dehydrogenase, cytosolic | 9 | 0.27–0.05 a,▼▼ | 0.11 ± 0.01b▼▼▼ | 0.10 ± 0.02 b,▼▼▼ | 0.024 |
| Salt stress root protein “RS1” | 27 | 4.09 ± 0.24 a,▲▲ | 1.95 ± 1.36 b,▲ | 2.75 ± 0.17 a,b,▲ | 0.018 |
| Unknown | 29 | 0.37 ± 0.05 a,▼ | 0.19 ± 0.03 b,▼▼▼ | 0.08 ± 0.02 b,▼▼▼ | 0.008 |
| Probable ATP synthase 24kDa subunit | 30 | 0.43 ± 0.01 a,▼ | 0.18 ± 0.02 b,▼▼▼ | 0.20 ± 0.02 b,▼▼▼ | 0.061 |
| Glyoxalase I | 31 | 0.94 ± 0.09 a,* | 0.47 ± 0.07 a,b,▼▼ | 0.33 ± 0.01 b,▼▼ | 0.087 |
| Formate dehydrogenase | 35 | 0.46 ± 0.09 b,▼ | 1.28 ± 0.17 a,* | 1.16 ± 0.21 a,* | 0.015 |
| ATP-dependent DNA helicase UvrD Shewanella denitrificans OS217 | 38 | 0.49 ± 0.15b,▼ | 1.39 ± 0.21a,▲ | 1.04 ± 0.32 a,* | 0.011 |
| Hypothetical protein OsJ_015102 | 40 | 0.33 ± 0.07 b,▼ | 1.20 ± 0.12 a,* | 0.99 ± 0.29 a,* | 0.041 |
| (O65316) Actin (Mesostigma viride) | 41 | 1.81 ± 0.36 b,▲ | 5.94 ± 1.08 a,▲▲ | 7.27 ± 1.33 a,▲▲ | 0.021 |
| EFTu1 | 64 | 14.05 ± 1.49 a,▲▲▲ | 7.78 ± 0.83 b,▲▲ | 12.58 ± 1.52 a,▲▲▲ | 0.065 |
Supplementary Files
© 2013 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Sangha, J.S.; Chen, Y.H.; Kaur, J.; Khan, W.; Abduljaleel, Z.; Alanazi, M.S.; Mills, A.; Adalla, C.B.; Bennett, J.; Prithiviraj, B.; et al. Proteome Analysis of Rice (Oryza sativa L.) Mutants Reveals Differentially Induced Proteins during Brown Planthopper (Nilaparvata lugens) Infestation. Int. J. Mol. Sci. 2013, 14, 3921-3945. https://doi.org/10.3390/ijms14023921
Sangha JS, Chen YH, Kaur J, Khan W, Abduljaleel Z, Alanazi MS, Mills A, Adalla CB, Bennett J, Prithiviraj B, et al. Proteome Analysis of Rice (Oryza sativa L.) Mutants Reveals Differentially Induced Proteins during Brown Planthopper (Nilaparvata lugens) Infestation. International Journal of Molecular Sciences. 2013; 14(2):3921-3945. https://doi.org/10.3390/ijms14023921
Chicago/Turabian StyleSangha, Jatinder Singh, Yolanda H. Chen, Jatinder Kaur, Wajahatullah Khan, Zainularifeen Abduljaleel, Mohammed S. Alanazi, Aaron Mills, Candida B. Adalla, John Bennett, Balakrishnan Prithiviraj, and et al. 2013. "Proteome Analysis of Rice (Oryza sativa L.) Mutants Reveals Differentially Induced Proteins during Brown Planthopper (Nilaparvata lugens) Infestation" International Journal of Molecular Sciences 14, no. 2: 3921-3945. https://doi.org/10.3390/ijms14023921
APA StyleSangha, J. S., Chen, Y. H., Kaur, J., Khan, W., Abduljaleel, Z., Alanazi, M. S., Mills, A., Adalla, C. B., Bennett, J., Prithiviraj, B., Jahn, G. C., & Leung, H. (2013). Proteome Analysis of Rice (Oryza sativa L.) Mutants Reveals Differentially Induced Proteins during Brown Planthopper (Nilaparvata lugens) Infestation. International Journal of Molecular Sciences, 14(2), 3921-3945. https://doi.org/10.3390/ijms14023921




