Wild and Hatchery Populations of Korean Starry Flounder (Platichthys stellatus) Compared Using Microsatellite DNA Markers
Abstract
:1. Introduction
2. Results and Discussion
2.1. Microsatellite Marker Isolation
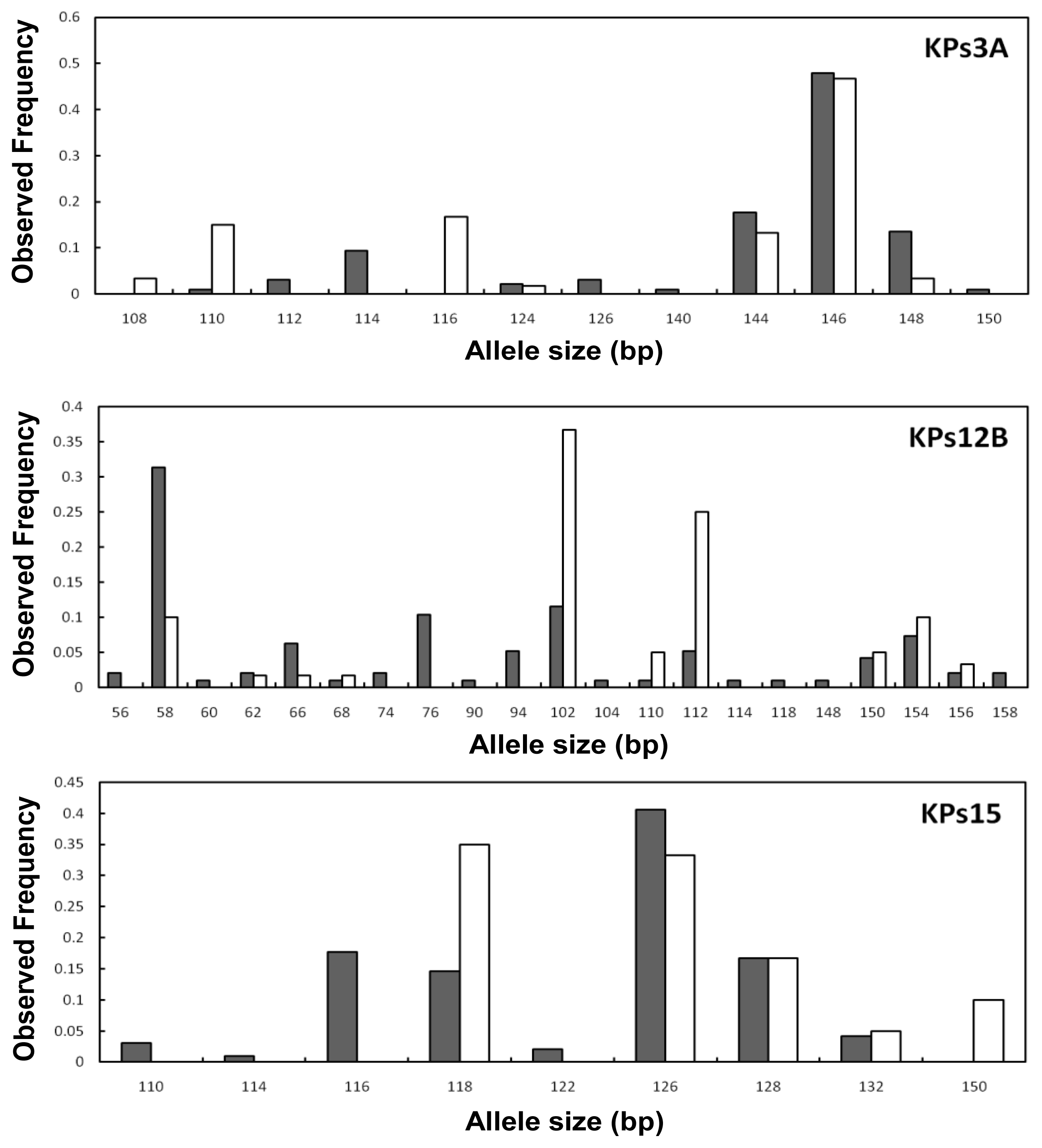
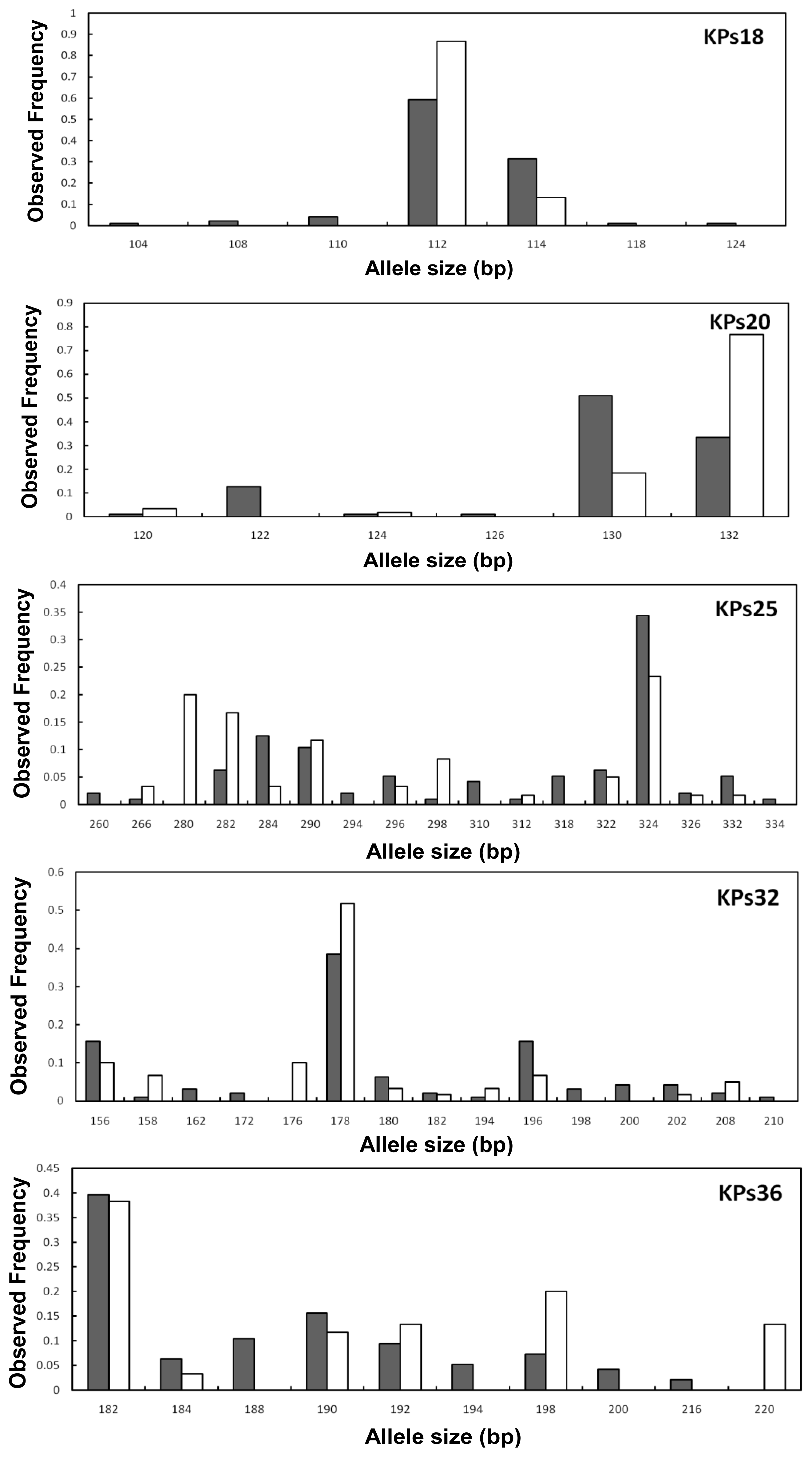
2.2. Genetic Variation within Populations
2.3. Genetic Differentiation between the Wild and Hatchery Populations
3. Experimental Section
3.1. Sample Collection and DNA Extraction
3.2. Microsatellite-Enriched Genomic Library Construction and Microsatellite Sequencing
3.3. Isolation of Microsatellite-Containing DNA Fragments and Microsatellite Sequencing
3.4. Primer Design and Allele Scoring
3.5. Sample Comparisons
4. Conclusions
Acknowledgments
References
- Hart, J.L. Pacific fishes of Canada. Bull. Fish. Res. Boars Can 1973, 180, 740. [Google Scholar]
- Alverson, D.L.; Pruter, A.T.; Ronholt, L.L. A Study of Demersal Fishes and Fisheries of the Northeastern Pacific Ocean; Institute for Fisheries, University of British Columbia: Vancouver, Canada, 1964; p. 190. [Google Scholar]
- Manual of Starry Flounder Culture; National Fisheries Research and Development Institute: Busan, Korean, 2010.
- Evaluation of a Stock Enhancement Programme Effectiveness; Korean Fisheries Resources Agency: Seoul, Korean, 2010.
- Allendorf, F.W.; Phelps, S.R. Loss of genetic variation in a hatchery stock of cutthroat trout. Trans. Am. Fish. Soc 1980, 109, 537–543. [Google Scholar]
- Lacy, R.C. Loss of genetic diversity from managed populations: Interacting effects of drift, mutation, selection, and population subdivision. Conserv. Biol 1987, 2, 143–158. [Google Scholar]
- Report of the Expert Consultation on Utilization and Conservation of Aquatic Genetic Resources; FAO Fisheries Report No. 491; FAO: Rome, Italy, 1993.
- Liu, Z.J.; Cordes, F.J. DNA marker technology and their applications in aquaculture genetics. Aquaculture 2004, 238, 1–37. [Google Scholar]
- Feral, J.P. How useful are the genetic markers in attempts to understand and manage marine biodiversity? J. Exp. Mar. Biol. Ecol 2002, 268, 121–145. [Google Scholar]
- Chistiakov, D.A.; Hellemans, B.; Volckaert, F.A.M. Microsatellites and their genomic distribution, evolution, function and applications: A review with special reference to fish genetics. Aquaculture 2006, 255, 1–29. [Google Scholar]
- Kohlmann, K.; Kerten, P.; Flajshans, M. Microsatellite-based genetic variability and differentiation of domesticated, wild and feral common carp (Cyprinus carpio L.) populations. Aquaculture 2005, 247, 253–266. [Google Scholar]
- An, H.S.; Kim, E.-M.; Lee, J.-H.; Noh, J.K.; An, C.M.; Yoon, S.J.; Park, K.D.; Myeong, J.-I. Population genetic structure of wild and hatchery black rockfish Sebastes inermis in Korea, assessed using cross-species microsatellite markers. Genet. Mol. Res 2011, 10, 2492–2504. [Google Scholar]
- Wang, L.; meng, Z.; Liu, X.; Lin, H. Genetic diversity and differentiation of the orange-spotted grouper (Epinephelus coiodes) between and within cultured stocks and wild populations from microsatellite DNA analysis. Int. J. Mol. Sci 2011, 12, 4378–4394. [Google Scholar]
- Kang, J.-H.; Kim, Y.-K.; Park, J.-Y.; An, C.M.; Nam, M.-M.; Byun, S.G.; Lee, B.I.; Lee, J.H.; Choi, T.-J. Microsatellite analysis as a tool for discriminating an interfamily hybrid between olive flounder and starry flounder. Genet. Mol. Res 2011, 10, 2786–2794. [Google Scholar]
- Hamilton, M.B.; Pincus, E.L.; DiFiore, A.; Fleischer, R.C. Universal linker and ligation procedures for construction of genomic DNA libraries enriched for microsatellites. BioTechniques 1999, 27, 500–507. [Google Scholar]
- Gardner, M.G.; Cooper, S.J.B.; Bull, C.M.; Grant, W.N. Isolation of microsatellite loci from a social lizard, Egernia stokesii, using a modified enrichment procedure. J. Hered 1999, 90, 301–304. [Google Scholar]
- Carleton, K.L.; Streelman, J.T.; Lee, B.Y.; Garnhart, N.; Kidd, M.; Kocher, T.D. Rapid isolation of CA microsatellites from the tilapia genome. Anim. Genet 2002, 33, 140–144. [Google Scholar]
- Yue, G.H.; Zhu, Z.Y.; Wang, C.M.; Xia, J.H. A simple and efficient method for isolating polymorphic microsatellites from cDNA. BMC Genomics 2009, 10, 125:1–125:8. [Google Scholar]
- Sekino, M.; Hara, M. Isolation and characterization of microsatellite DNA loci in Japanese flounder Paralichthys olivaceus (Pleuronectiformes, Pleuronectoidei, Paralichthyidae). Mol. Ecol 2000, 9, 2200–2203. [Google Scholar]
- Cruz, F.; Pérez, M.; Presa, P. Distribution and abundance of microsatellites in the genome of bivalves. Gene 2005, 241–247. [Google Scholar]
- DeWoody, J.A.; Avise, J.C. Microsatellite variation in marine, freshwater and anadromous fishes compared with other animals. J. Fish Biol 2000, 56, 461–473. [Google Scholar]
- Li, Q.; Park, C.; Kobayashi, T.; Kijima, A. Inheritance of microsatellite DNA markers in the Pacific abalone Haliotis discus hannai. Mar. Biotech 2003, 5, 331–338. [Google Scholar]
- Callen, D.F.; Thompson, A.D.; Shen, Y.; Phillips, H.A.; Mulley, J.C.; Sutherland, G.R. Incidence and origin of “null” alleles in the (AC)n microsatellite markers. Am. J. Hum. Genet 1993, 52, 922–927. [Google Scholar]
- De Sousa, S.N.; Finkeldey, R.; Gailing, O. Experimental verification of microsatellite null alleles in Norway spruce (Picea abies [L.] Karst.): Implications for population genetic studies. Plant Mol. Biol. Rep 2005, 23, 113–119. [Google Scholar]
- Kim, M.-J.; An, H.S.; Choi, K.H. Genetic characteristics of Pacific cod populations in Korea based on microsatellite markers. Fish. Sci 2010, 76, 595–603. [Google Scholar]
- Comps, B.; Gomory, D.; Letouzey, J.; Thiebaut, B.; Petit, R.J. Diverging trends between heterozygosity and allelic richness during postglacial colonization in the European beech. Genetics 2001, 157, 389–397. [Google Scholar]
- Norris, A.T.; Bradley, D.G.; Cunningham, E.P. Microsatellite genetic variation between and within farmed and wild Atlantic salmon (Salmo salar) populations. Aquaculture 1999, 180, 247–264. [Google Scholar]
- Mjolnerod, I.B.; Refseth, U.H.; Karlsen, E.; Balstad, T.; Jakobsen, K.S.; Hindar, K. Genetic differences between two wild and one farmed population of Atlantic salmon (Salmo salar) revealed by three classes of genetic markers. Hereditas 1997, 127, 239–248. [Google Scholar]
- Vuorinen, J. Reduction of genetic variability in a hatchery stock of brown trout, Salmo trutta. J. Fish Biol 1984, 24, 339–348. [Google Scholar]
- Asahida, T.; Kobayashi, T.; Saitoh, K.; Nakayama, I. Tissue preservation and total DNA extraction from fish stored at ambient temperature using buffers containing high concentrations of urea. Fish. Sci. Tokyo 1996, 62, 727–730. [Google Scholar]
- Li, Q.; Park, C.; Kijima, A. Isolation and characterization of microsatellite loci in the Pacific abalone, Haliotis discus hannai. J. Shell. Res 2002, 212, 811–815. [Google Scholar]
- Van Oosterhout, C.; Hutchinson, W.F.; Wills, D.P.M.; Shipley, P. MICRO-CHECKER: Software for identifying and correcting genotyping errors in microsatellite data. Mol. Ecol. Notes 2004, 4, 535–538. [Google Scholar]
- Raymond, M.; Rousset, F. GENEPOP (version 1.2): Population genetics software for exact tests and ecumenicism. J. Hered 1995, 86, 248–249. [Google Scholar]
- Goudet, J. FSTAT (version 1.2): A computer program to calculate F-statistics. J. Hered 1995, 86, 485–486. [Google Scholar]
- Weir, B.S.; Cockerham, C.C. Estimating F-statistics for the analysis of population structure. Evolution 1984, 38, 1358–1370. [Google Scholar]
- El Mousadik, A.; Petit, R.J. High level of genetic differentiation for allelic richness among populations of the argan tree (Argania spinosa (L.) Skeels) endemic to Morocco. Theor. Appl. Genet 1996, 92, 832–839. [Google Scholar]
- Slatkin, M.; Excoffier, L. Testing for linkage disequilibrium in genotypic data using the EM algorithm. Heredity 1996, 76, 377–383. [Google Scholar]
- Rice, W.R. Analyzing tables of statistical tests. Evolution 1989, 43, 223–225. [Google Scholar]
| Locus | Repeat motif | Primer sequence (5′ → 3′) | Ta (°C) | Genbank accession no. |
|---|---|---|---|---|
| KPs1 | (CA)11 | CAGCAGTAAGAGTGTGTCCTG Hex TTCAGCCTGTTTTCTGTCAT | 55 | EF157643 |
| KPs2 | (TG)12 | TTAGGGGTGGGACAGACT Hex GTCATCAGATGGGAGAAAGAT | 55 | EF157644 |
| KPs3A | (TG)12 | AGGTTATCGCATCTGTGTGAT Hex GAACTCTTGTTTCGCTTCAG | 55 | EF157645 |
| KPs5A | (CA)11 | ACAGATAGCAAGGTCATAGAA Ned TACTAATTCCAAGGTGTTGAT | 60 | EF157646 |
| KPs12B | (TG)15TC(TG)12GT(TG)3 | TGTTTACTGCTTTCCTGTGTG Fam TGTATTTCAGCCTGCTTTATG | 58 | EF157647 |
| KPs15 | (TG)5TA(TG)8TC(TG)4 | GAGCCAGACCTCTCATGTTAC Fam CGCTTCCATGTGAACCAG | 60 | EF157648 |
| KPs17A | (TG)5TC(TG)9 | CAACCACGTTATCCTCTGTG Fam CCAGAATAAATCTCATGCTCA | 60 | EF157649 |
| KPs18 | (TG)8CG(TG)4CG(TG)3 | TCTTGGATGTAGTGTATGTGC Fam GAAAACACGAAATTTGACAG | 60 | EF157650 |
| KPs20 | (CA)14 | TGGGCAACTACGTACACACTA Fam GCCGACATTACAAAAACAAA | 58 | EF157651 |
| KPs23 | (CA)9CG(CA)5CG(CA)20TA(CA)5 | ACAAACACTTGCATGGGTAT Ned ATCCTCTAGCAGAAGCATTTC | 55 | EF157652 |
| KPs25 | (TG)6TA(TG)8TA(TG)10 | TGTTATCGGGTGTTGATTGTA Ned GTTGATTGTGAAACGCTGTT | 60 | EF157653 |
| KPs27 | (CA)27 | TGTTGAAATCTAATGGGCTAT Fam GTGCGCAGCAAAAACTA | 51 | EF157654 |
| KPs29 | (CA)6AA(CA)14 | TCCTACTCTACACACCCACAT Hex GGATCGGAAAAGACAGACTA | 55 | EF157655 |
| KPs31 | (CA)10 | TGATTTCCAATTACTCACATA Fam CTAGCAGAAGCCTTTCAG | 58 | EF157656 |
| KPs32 | (TG)8TT(TG)7TA(TG)22 | TTAAATAAGTGTCTGGGGATT Hex GCCACACTTCTGCTTCTG | 55 | EF157657 |
| KPs33 | (CA)9-(CA)11 | TTTCACTTCTCTTTGGGTTAC Ned GGCAGACTGATTCCTCATT | 60 | EF157658 |
| KPs36 | (TG)4TT(TG)13 | ATGTGCCCAATAAAACAAAC Hex CTAAGCCCTAGAGCAAACAGT | 58 | EF157659 |
| Population (No) | Microsatellite Loci
| |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| KPs1 | KPs3A | KPs12B | KPs15 | KPs17A | KPs18 | KPs20 | KPs25 | KPs27 | KPs32 | KPs33 | KPs36 | Mean | ||
| FST | 0.012 | 0.065 | 0.038 | 0.028 | 0.102 | 0.135 | 0.035 | 0.007 | 0.015 | 0.038 | 0.008 | 0.035 | 0.043 | |
| Uijin Captured (48) | NA | 10 | 9 | 18 | 7 | 11 | 7 | 6 | 16 | 14 | 14 | 8 | 9 | 10.75 |
| AR | 9.06 | 7.92 | 14.81 | 6.57 | 10.10 | 5.72 | 4.88 | 14.05 | 12.64 | 12.32 | 7.09 | 8.84 | 9.50 | |
| S | 140–174 | 110–150 | 56–158 | 110–132 | 62–104 | 104–124 | 120–132 | 260–334 | 82–130 | 156–210 | 274–294 | 182–216 | ||
| F | 0.385 | 0.531 | 0.313 | 0.563 | 0.208 | 0.594 | 0.479 | 0.344 | 0.208 | 0.385 | 0.510 | 0.396 | 0.41 | |
| U | 5 | 5 | 8 | 2 | 2 | 5 | 4 | 5 | 3 | 5 | 1 | 4 | 4.08 | |
| He | 0.819 | 0.667 | 0.868 | 0.630 | 0.873 | 0.583 | 0.528 | 0.845 | 0.888 | 0.832 | 0.661 | 0.793 | 0.749 | |
| Ho | 0.483 | 0.698 | 0.833 | 0.625 | 0.633 | 0.553 | 0.475 | 0.792 | 0.938 | 0.763 | 0.604 | 0.604 | 0.667 | |
| FIS | 0.297 (0.000) | −0.176 (0.147) | 0.100 (0.486) | 0.078 (0.661) | 0.355 (0.000) | 0.088 (0.453) | 0.151 (0.014) | 0.041 (0.399) | −0.117 (0.580) | 0.097 (0.028) | 0.050 (0.208) | 0.211 (0.009) | 0.061 (0.120) | |
| P | 0.000 | 0.069 | 0.376 | 0.784 | 0.000 | 0.628 | 0.009 | 0.475 | 0.661 | 0.010 | 0.148 | 0.000 | 0.074 | |
| Uljin Cultured (30) | NA | 5 | 7 | 10 | 5 | 9 | 2 | 2 | 12 | 12 | 10 | 7 | 6 | 7.25 |
| AR | 5.00 | 7.00 | 10.00 | 5.00 | 9.00 | 2.00 | 2.00 | 12.00 | 12.00 | 10.00 | 7.00 | 6.00 | 7.25 | |
| S | 142–170 | 108–148 | 58–156 | 118–150 | 68–104 | 112–114 | 120–132 | 266–332 | 82–140 | 156–208 | 274–294 | 182–220 | ||
| F | 0.517 | 0.683 | 0.367 | 0.367 | 0.283 | 0.867 | 0.767 | 0.233 | 0.283 | 0.600 | 0.500 | 0.317 | 0.48 | |
| U | 0 | 3 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 1 | 0 | 1 | 0.58 | |
| He | 0.683 | 0.512 | 0.867 | 0.800 | 0.868 | 0.200 | 0.364 | 0.871 | 0.867 | 0.627 | 0.666 | 0.800 | 0.677 | |
| Ho | 0.533 | 0.578 | 0.789 | 0.743 | 0.667 | 0.235 | 0.300 | 0.865 | 0.897 | 0.467 | 0.633 | 0.733 | 0.620 | |
| FIS | 0.364 (0.016) | −0.031 (0.848) | 0.080 (0.290) | 0.007 (0.094) | 0.405 (0.044) | −0.055 (0.414) | 0.046 (0.025) | 0.063 (0.312) | −0.056 (0.243) | 0.299 (0.053) | 0.087 (0.349) | 0.100 (0.013) | 0.109 (0.215) | |
| P | 0.000 | 1.000 | 0.021 | 0.016 | 0.000 | 0.428 | 0.012 | 0.156 | 0.189 | 0.000 | 0.352 | 0.023 | 0.011 | |
| Locus | P-value | Locus | P-value |
|---|---|---|---|
| KPs1 | 0.016 | KPs20 | 0.000* |
| KPs3A | 0.000* | KPs25 | 0.000* |
| KPs12B | 0.000* | KPs27 | 0.577 |
| KPs15 | 0.000* | KPs32 | 0.003* |
| KPs17A | 0.461 | KPs33 | 0.352 |
| KPs18 | 0.002* | KPs36 | 0.000* |
© 2011 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
An, H.S.; Byun, S.G.; Kim, Y.C.; Lee, J.W.; Myeong, J.-I. Wild and Hatchery Populations of Korean Starry Flounder (Platichthys stellatus) Compared Using Microsatellite DNA Markers. Int. J. Mol. Sci. 2011, 12, 9189-9202. https://doi.org/10.3390/ijms12129189
An HS, Byun SG, Kim YC, Lee JW, Myeong J-I. Wild and Hatchery Populations of Korean Starry Flounder (Platichthys stellatus) Compared Using Microsatellite DNA Markers. International Journal of Molecular Sciences. 2011; 12(12):9189-9202. https://doi.org/10.3390/ijms12129189
Chicago/Turabian StyleAn, Hye Suck, Soon Gyu Byun, Yi Cheong Kim, Jang Wook Lee, and Jeong-In Myeong. 2011. "Wild and Hatchery Populations of Korean Starry Flounder (Platichthys stellatus) Compared Using Microsatellite DNA Markers" International Journal of Molecular Sciences 12, no. 12: 9189-9202. https://doi.org/10.3390/ijms12129189




