Molecular Modeling of Peroxidase and Polyphenol Oxidase: Substrate Specificity and Active Site Comparison
Abstract
:1. Introduction
2. Results and Discussion
2.1. Three-Dimensional Model of Grape Ascorbate Peroxidase and Polyphenol Oxidase
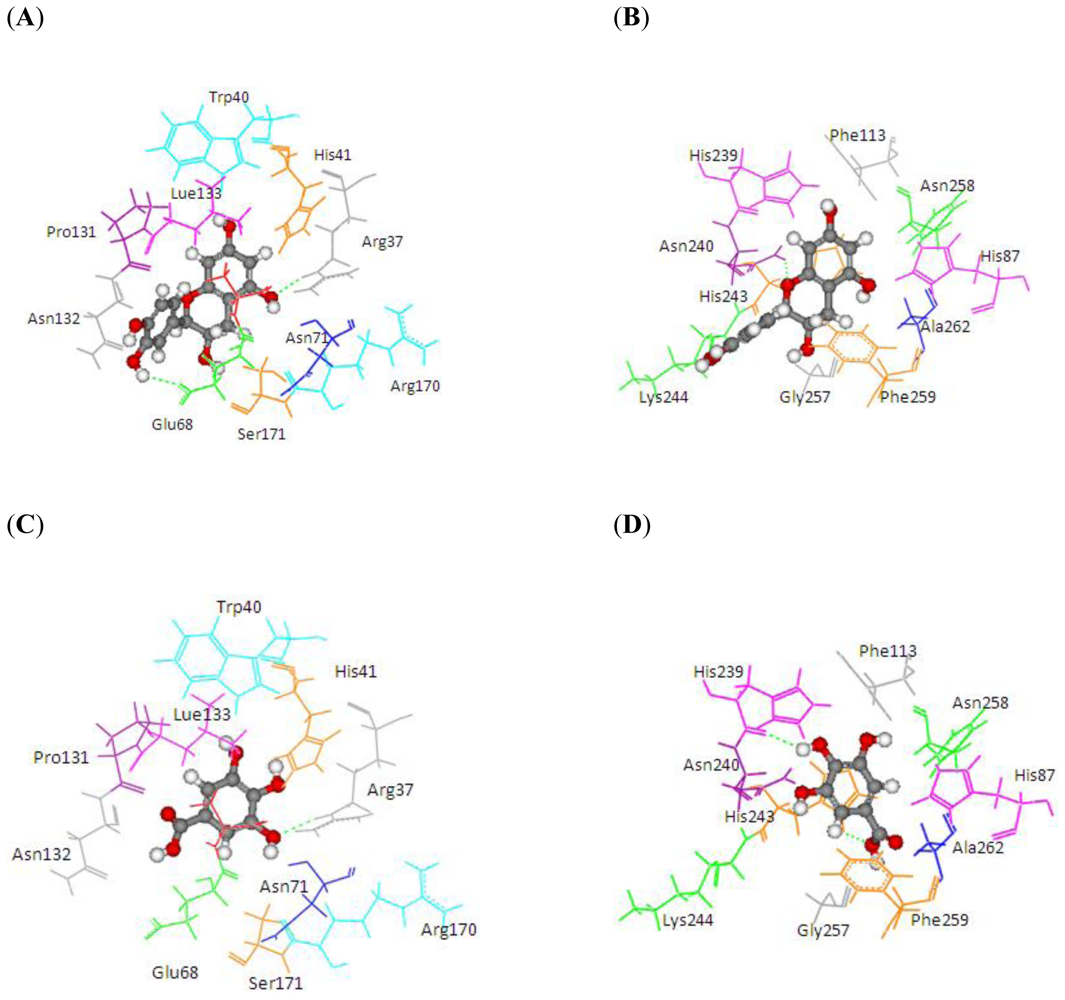
2.2. Comparison of Substrate Binding Site for PPO and POD from Molecular Docking
2.3. Specificity of Inhibitors for PPO and POD: Theoretical and Experimental Comparison
3. Experimental Section
3.1. Three Dimensional Structure Modeling
3.2. Docking Study
4. Conclusions
Acknowledgements
References
- Chen, L; Mehta, A; Berenbaum, M; Zangerl, AR; Engeseth, NJ. Honeys from different floral sources as inhibitors of enzymatic browning in fruit and vegetable homogenates. J. Agric. Food Chem 2000, 48, 4997–5000. [Google Scholar]
- Sapers, GM. Browning of foods: Control by sulfites, antioxidants, and other means. Food Technol 1993, 68, 5–84. [Google Scholar]
- Francisco, AT-B; Juan Carlos, E. Phenolic compounds and related enzymes as determinants of quality in fruits and vegetables. J. Sci. Food Agric 2001, 81, 853–876. [Google Scholar]
- Yemenicioglu, A; Özkan, M; Cemeroglu, B. Some characteristics of polyphenol oxidase and peroxidase from taro (Colocasia antiquorum). Tr. J. Agric. Forestry 1999, 23, 425–430. [Google Scholar]
- Banci, L. Structural properties of peroxidases. J. Biotechnol 1997, 53, 253–263. [Google Scholar]
- Onsa, GH; Saari, NB; Selamat, J; Bakar, J. Latent polyphenol oxidase from Sago Log(Metroxylon sagu): Partial purification, activation, and some properties. J. Agric. Food Chem 2000, 48, 5041–5045. [Google Scholar]
- Lengauer, T; Rarey, M. Computational methods for biomolecular docking. Curr. Opin. Struct. Biol 1996, 6, 402–406. [Google Scholar]
- Khan, AA; Robinson, DS. Hydrogen donor specificity of mango isoperoxidases. Food Chem 1994, 49, 407–410. [Google Scholar]
- Sugai, AY; Tadini, CC. Thermal inactivation of mango (Mangifera indica variety Palmer) puree peroxidase. Proceedings of 2006 CIGR Section VI International Symposium on Future of Food Engineering, Warsaw, Poland; 2006. [Google Scholar]
- Onsa, GH; Saari, NB; Selamat, J; Bakar, J. Purification and characterization of membrane-bound peroxidases from Metroxylon sagu. Food Chem 2004, 85, 365–376. [Google Scholar]
- Furumo, NC; Furutani, S. A simple method for assaying total protein, polyphenol oxidase and peroxidase activity from ‘Kaimana’ Litchi chinensis Sonn. J. Haw. Pac. Agric 2008, 15. [Google Scholar]
- Altschul, SF; Madden, TL; Schaffer, AA; Zhang, J; Zhang, Z; Miller, W; Lipman, DJ. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucl. Acids Res 1997, 25, 3389–3402. [Google Scholar]
- Larkin, MA; Blackshields, G; Brown, NP; Chenna, R; McGettigan, PA; McWilliam, H; Valentin, F; Wallace, IM; Wilm, A; Lopez, R; Thompson, JD; Gibson, TJ; Higgins, DG. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar]
- Tatoli, S; Zazza, C; Sanna, N; Palma, A; Aschi, M. The role of arginine 38 in horseradish peroxidase enzyme revisited: A computational investigation. Biophys. Chem 2009, 141, 87–93. [Google Scholar]
- Poulos, TL; Kraut, J. The stereochemistry of peroxidase catalyst. J. Biol. Chem 1980, 255, 8199–8205. [Google Scholar]
- Rapeanu, G; Loey, AV; Smout, C; Hendrickx, M. Biochemical characterization and process stability of polyphenoloxidase extracted from Victoria grape (Vitis vinifera ssp. Sativa). Food Chem 2006, 94, 253–261. [Google Scholar]
- Guerrero-Beltrán, JA; Swanson, BG; Barbosa-Cánovas, GV. Inhibition of polyphenoloxidase in mango puree with 4-hexylresorcinol, cysteine and ascorbic acid. LWT Food Sci. Technol 2005, 38, 625–630. [Google Scholar]
- Özođlu, H; BayIndIrlI, A. Inhibition of enzymic browning in cloudy apple juice with selected antibrowning agents. Food Control 2002, 13, 213–221. [Google Scholar]
- Ýyidođan, NF; BayIndIrlI, A. Effect of l-cysteine, kojic acid and 4-hexylresorcinol combination on inhibition of enzymatic browning in Amasya apple juice. J. Food Eng 2004, 62, 299–304. [Google Scholar]
- Vámos-Vigyázó, L; Haard, NF. Polyphenol oxidases and peroxidases in fruits and vegetables. CRC Crit. Rev. Food Sci. Nutr 1981, 15, 49–127. [Google Scholar]
- Peter, HF; John, RLW. Inhibition of diphenol oxidases: A comparative study. J. Food Biochem 1996, 20, 15–30. [Google Scholar]
- Patterson, WR; Poulos, TL. Crystal structure of recombinant pea cytosolic ascorbate peroxidase. Biochemistry 1995, 34, 4331–4341. [Google Scholar]
- Sali, A; Blundell, TL. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol 1993, 234, 779–815. [Google Scholar]
- Wu, G; Robertson, DH; Brooks, CL, III; Vieth, M. Detailed analysis of grid-based molecular docking: A case study of CDOCKER-A CHARMm-based MD docking algorithm. J. Comput. Chem 2003, 24, 1549–1562. [Google Scholar]

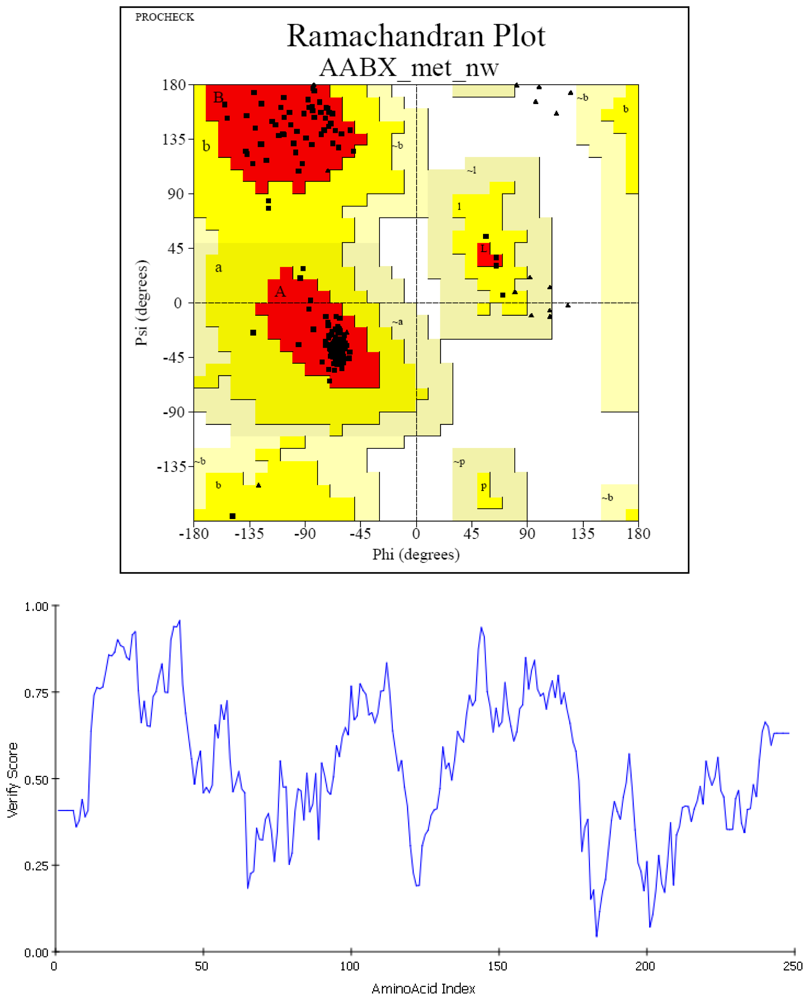
| PROCHECK | Ramachandran Plot Quality (%) | |||
|---|---|---|---|---|
| Core | Allowed | General | Disallowed | |
| Model | 95.6 | 4.40 | 0 | 0 |
| Template | 93.7 | 5.8 | 0 | 0.5 |
| Substrate | Structure | ABX (POD) | 2P3X (PPO) | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Experimental Value [10] Km(×10−3 M) | Interaction Energy (kcal/mol) | No. of Hydrogen Bonding | Residue in hydrogen Bonding | Relative Activity [6] | Interaction Energy (kcal/mol) | No. of Hydrogen Bonding | Residue in Hydrogen Bonding | ||
| Substrates | |||||||||
| 4MC | 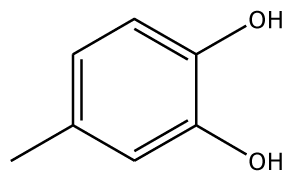 | 22.0 | −28.23 | 1 | Arg37 | 100 | −41.85 | 1 | His239 |
| GAC | 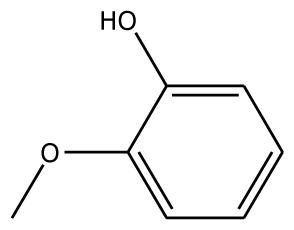 | 32.2 | −28.49 | 2 | Arg37 | −23.93 | 0 | ||
| PGL | 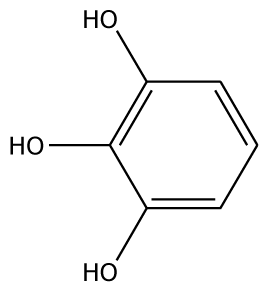 | 32.2 | −30.45 | 2 | Arg37 | 78.1 | −28.78 | 0 | |
| 3,4-DHPA | 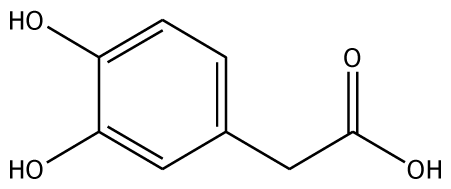 | na | −35.46 | 2 | Trp40 Arg170 | na | −53.55 | 2 | His239 Gly257 |
| CN | 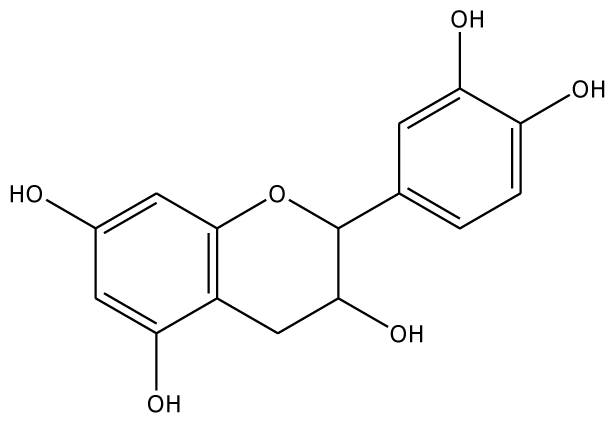 | 5.2 | 44.75 | 2 | Arg37 Glu68 | na | −45.55 | 2 | Asn240 Gly257 |
| EPC |  | 5.2 | −45.63 | 2 | Arg37 Glu68 | 93.1 | −42.99 | 1 | Asn240 |
| Inhibitors | |||||||||
| 2,3-DHBA |  | na | −32.15 | 1 | Pro131 | na | 37.37 | 1 | Gly257 |
| 3,4-DHBA | 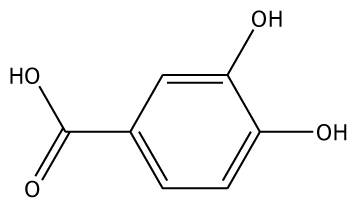 | na | −31.38 | 1 | Arg170 | na | −44.71 | 1 | His239 |
| 3,4,5-THBA | 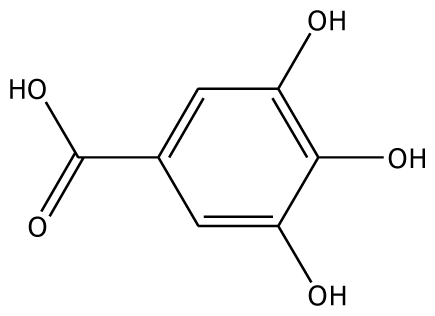 | na | 34.76 | 1 | Arg37 | na | −43.01 | 4 | His239His243 Gly257 Asn258 |
| o-HBA | 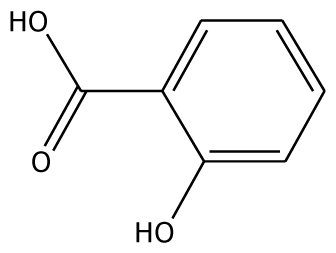 | na | −29.14 | 1 | Arg37 | na | −33.99 | 1 | His239 |
| m-HBA | 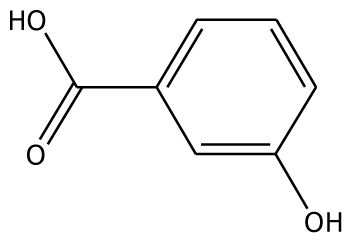 | na | −29.17 | 0 | na | −39.04 | 1 | Gly257 | |
| p-HBA |  | na | −26.23 | 1 | Trp40 | na | −36.68 | 2 | Glu235 Gly257 |
© 2010 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Nokthai, P.; Lee, V.S.; Shank, L. Molecular Modeling of Peroxidase and Polyphenol Oxidase: Substrate Specificity and Active Site Comparison. Int. J. Mol. Sci. 2010, 11, 3266-3276. https://doi.org/10.3390/ijms11093266
Nokthai P, Lee VS, Shank L. Molecular Modeling of Peroxidase and Polyphenol Oxidase: Substrate Specificity and Active Site Comparison. International Journal of Molecular Sciences. 2010; 11(9):3266-3276. https://doi.org/10.3390/ijms11093266
Chicago/Turabian StyleNokthai, Prontipa, Vannajan Sanghiran Lee, and Lalida Shank. 2010. "Molecular Modeling of Peroxidase and Polyphenol Oxidase: Substrate Specificity and Active Site Comparison" International Journal of Molecular Sciences 11, no. 9: 3266-3276. https://doi.org/10.3390/ijms11093266
APA StyleNokthai, P., Lee, V. S., & Shank, L. (2010). Molecular Modeling of Peroxidase and Polyphenol Oxidase: Substrate Specificity and Active Site Comparison. International Journal of Molecular Sciences, 11(9), 3266-3276. https://doi.org/10.3390/ijms11093266




