Identification of Tetranectin as a Potential Biomarker for Metastatic Oral Cancer
Abstract
:1. Introduction
2. Materials and methods
2.1. Patients and Samples
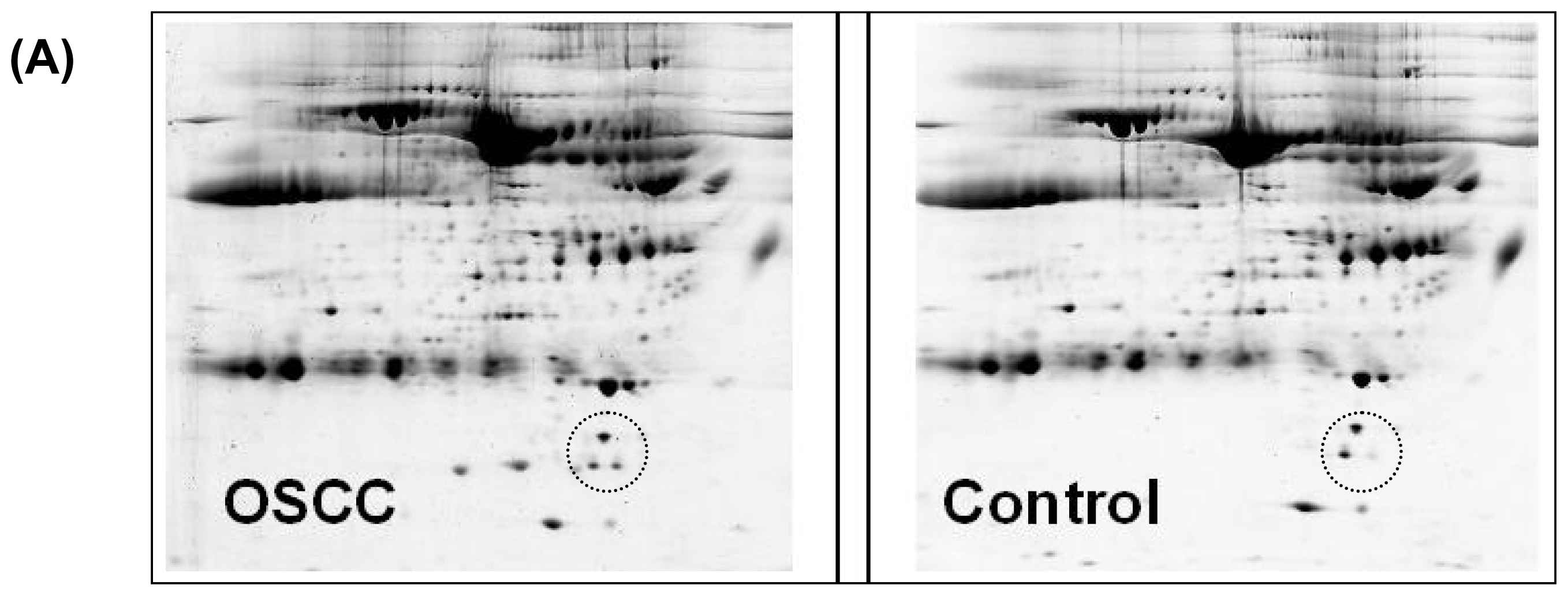
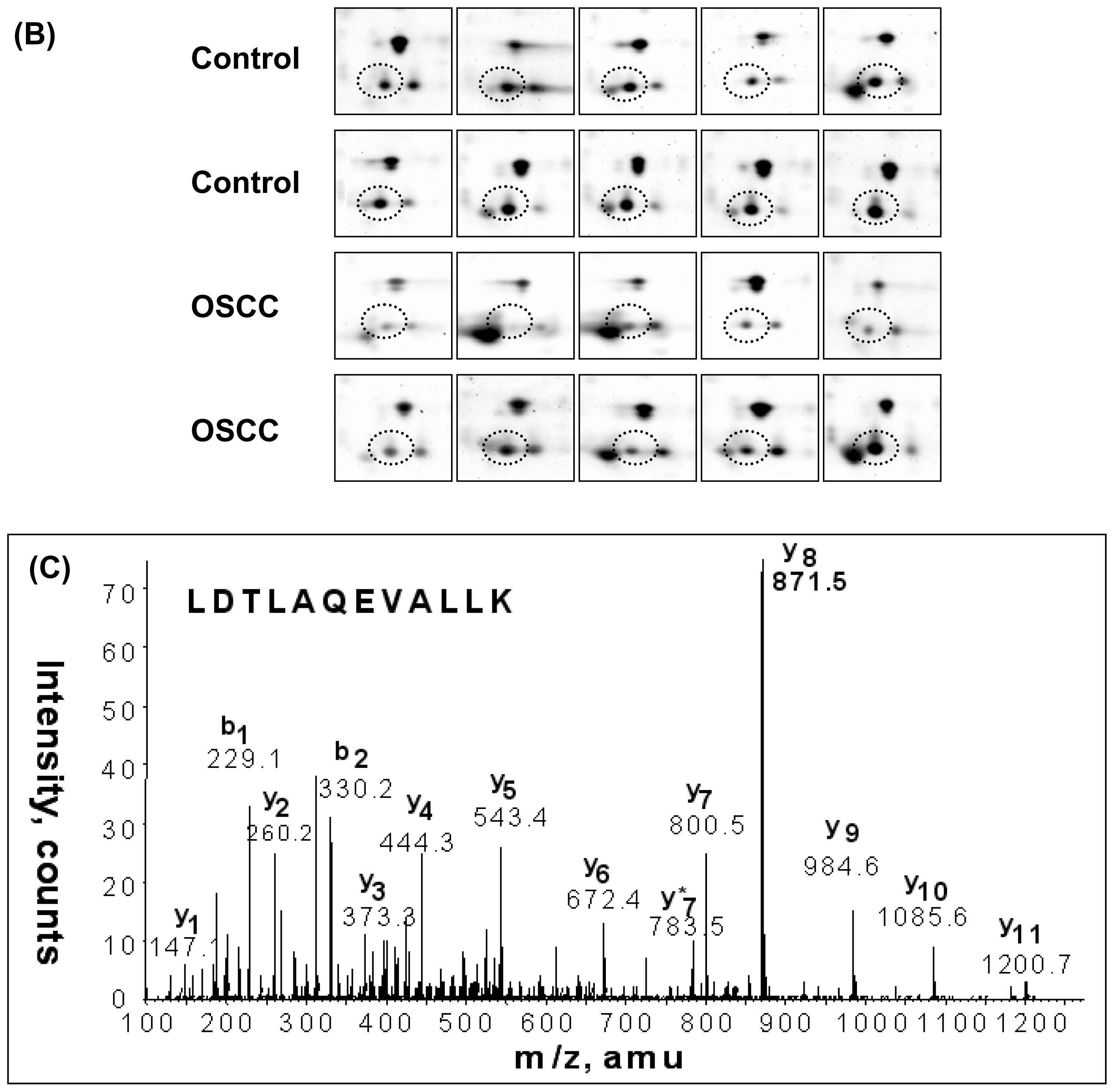
2.2. 2-D Gel Electrophoresis
2.3. LC-MS/MS and Database Searching
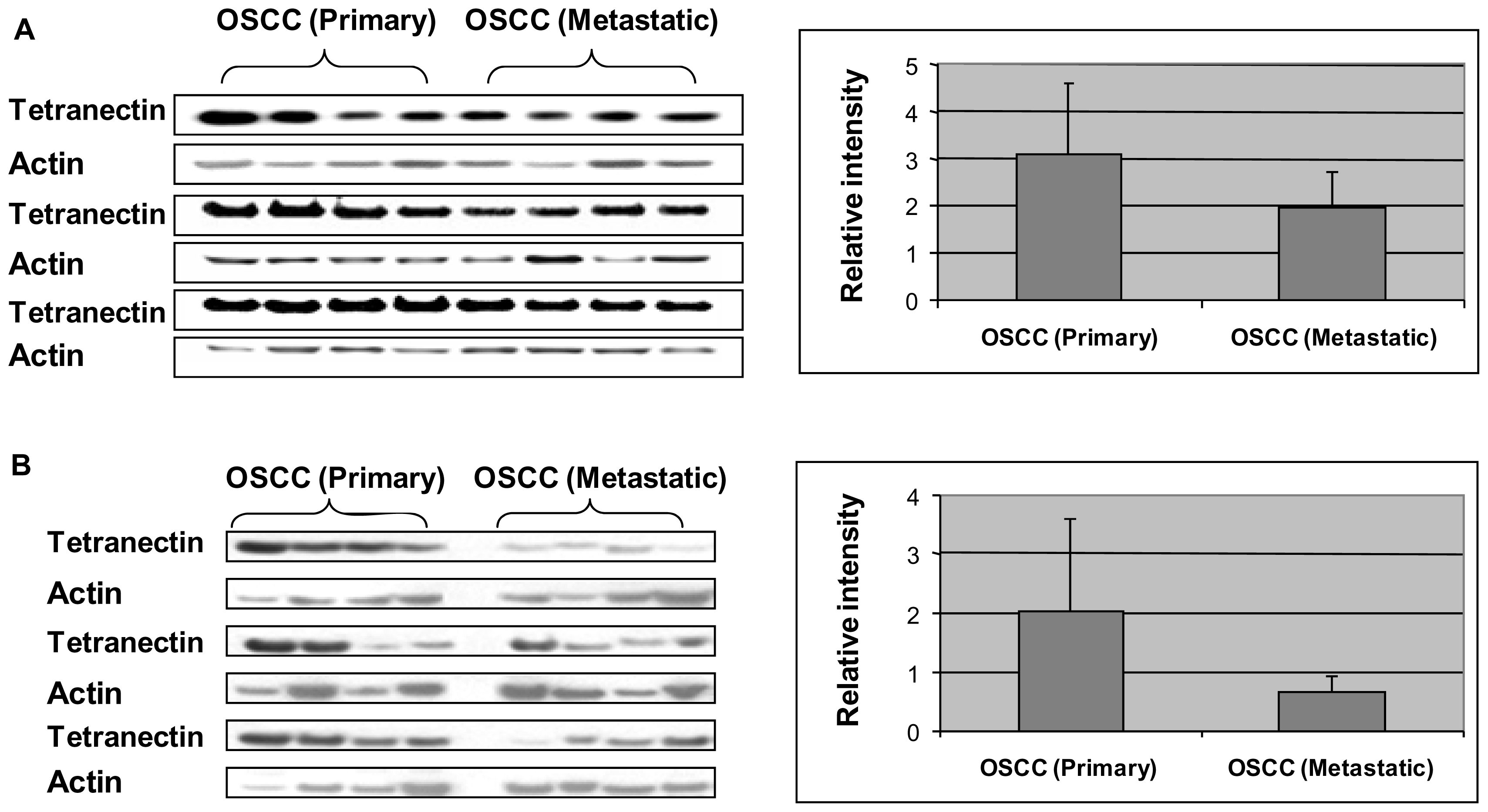
2.4. Western Blot Analysis
3. Results
4. Discussion
5. Conclusion
Acknowledgement
References
- Argiris, A; Karamouzis, MV; Raben, D; Ferris, RL. Head and neck cancer. Lancet 2008, 371, 1695–1709. [Google Scholar]
- Parkin, DM; Bray, F; Ferlay, J; Pisani, P. Global Cancer Statistics, 2002. CA: Cancer J. Clin 2005, 55, 74–108. [Google Scholar]
- Grandi, C; Alloisio, M; Moglia, D; Podrecca, S; Sala, L; Salvatori, P; Molinari, R. Prognostic significance of lymphatic spread in head and neck carcinomas: Therapeutic implications. Head Neck Surg 1985, 8, 67–73. [Google Scholar]
- Greenberg, JS; Fowler, R; Gomez, J; Mo, V; Roberts, D; Naggar, AKE; Myers, JN. Extent of extracapsular spread. Cancer 2003, 97, 1464–1470. [Google Scholar]
- Sano, D; Myers, J. Metastasis of squamous cell carcinoma of the oral tongue. Cancer Metastasis Rev 2007, 26, 645–662. [Google Scholar]
- Woolgar, JA. Pathology of the N0 neck. Br. J. Oral Maxillofac. Surg 1999, 37, 205–209. [Google Scholar]
- Jones, AS; Phillips, DE; Helliwell, TR; Roland, NJ. Occult node metastases in head and neck squamous carcinoma. Eur. Arch. Oto-Rhino-Laryngol 1993, 250, 446–449. [Google Scholar]
- Ferris, RL; Xi, L; Raja, S; Hunt, JL; Wang, J; Gooding, WE; Kelly, L; Ching, J; Luketich, JD; Godfrey, TE. Molecular staging of cervical lymph nodes in squamous cell carcinoma of the head and neck. Cancer Res 2005, 65, 2147–2156. [Google Scholar]
- Wilgen, CPv; Dijkstra, PU; Laan, BFAMvd; Plukker, JT; Roodenburg, JLN. Shoulder and neck morbidity in quality of life after surgery for head and neck cancer. Head Neck 2004, 26, 839–844. [Google Scholar]
- Ferlito, A; Partridge, M; Brennan, J; Hamakawa, H. Lymph node micrometastases in head and neck cancer: A review. Acta Oto-laryngol 2009, 121, 660–665. [Google Scholar]
- Wang, J; Xi, LQ; Hunt, JL; Gooding, W; Whiteside, TL; Chen, Z; Godfrey, TE; Ferris, RL. Expression pattern of chemokine receptor 6 (CCR6) and CCR7 in squamous cell carcinoma of the head and neck identifies a novel metastatic phenotype. Cancer Res 2004, 64, 1861–1866. [Google Scholar]
- Nikitakis, NG; Rivera, H; Lopes, MA; Siavash, H; Reynolds, MA; Ord, RA; Sauk, JJ. Immunohistochemical expression of angiogenesis-related markers in oral squamous cell carcinomas with multiple metastatic lymph nodes. Am. J. Clin. Pathol 2003, 119, 574–586. [Google Scholar]
- Stoll, C; Baretton, G; Soost, F; Terpe, HJ; Domide, P; Lohrs, U. Prognostic importance of the expression of CD44 splice variants in oral squamous cell carcinomas. Oral Oncol 1999, 35, 484–489. [Google Scholar]
- Agnes, B; Melanie, KO; Igor, BB; Andras, V; Endre, F; Jozsef, P. Gains and losses of adhesion molecules (CD44, E-cadherin, and beta-catenin) during oral carcinogenesis and tumour progression. J. Pathol 2002, 198, 343–351. [Google Scholar]
- Onda, T; Uzawa, K; Nakashima, D; Saito, K; Iwadate, Y; Seki, N; Shibahara, T; Tanzawa, H. Lin-7C/VELI3/MALS-3: An essential component in metastasis of human squamous cell carcinoma. Cancer Res 2007, 67, 9643–9648. [Google Scholar]
- Xia, W; Lau, Y; Hu, M; Li, L; Johnston, D; Sheng, S; El-Naggar, A; Hung, M. High tumoral maspin expression is associated with improved survival of patients with oral squamous cell carcinoma. Oncogene 2000, 19, 2398–2403. [Google Scholar]
- Lo Muzio, L; Mignogna, M; Pannone, G; Staibano, S; Procaccini, M; Serpico, R; de Rosa, G; Scully, C. The NM23 gene and its expression in oral squamous cell carcinoma. Oncol. Rep 1999, 6, 747–751. [Google Scholar]
- Ohtsuki, K; Shintani, S; Kimura, N; Matsumura, T. Immunohistochemical study on the nm23 gene produce (NDP kinase) in oral squamous cell carcinoma. Oral Oncol 1997, 33, 237–239. [Google Scholar]
- Sartini, DSA; Rossi, V; Goteri, G; Rubini, C; Ciavarella, D; Lo Muzio, L; Emanuelli, M. Nicotinamide N-methyltransferase upregulation inversely correlates with lymph node metastasis in oral squamous cell carcinoma. Mol. Med 2007, 13, 415–421. [Google Scholar]
- Hasina, R; Hulett, K; Bicciato, S; Di Bello, C; Petruzzelli, GJ; Lingen, MW. Plasminogen activator inhibitor-2: A molecular biomarker for head and neck cancer progression. Cancer Res 2003, 63, 555–559. [Google Scholar]
- Ferris, RL; Xi, LQ; Raja, S; Hunt, JL; Wang, J; Gooding, WE; Kelly, L; Ching, J; Luketich, JD; Godfrey, TE. Molecular staging of cervical lymph nodes in squamous cell carcinoma of the head and neck. Cancer Res 2005, 65, 2147–2156. [Google Scholar]
- Kurahara, S; Shinohara, M; Ikebe, T; Nakamura, S; Beppu, M; Hiraki, A; Takeuchi, H; Shirasuna, K. Expression of MMPs, MT-MMP, and TIMPs in squamous cell carcinoma of the oral cavity: Correlations with tumor invasion and metastasis. Head Neck-J. Sci. Spec 1999, 21, 627–638. [Google Scholar]
- Wang, J; Zhang, X; Thomas, SM; Grandis, JR; Wells, A; Chen, Z; Ferris, RL. Chemokine receptor 7 activates phosphoinositide-3 kinase-mediated invasive and prosurvival pathways in head and neck cancer cells independent of EGFR. Oncogene 2005, 24, 5897–5904. [Google Scholar]
- Masuda, M; Suzui, M; Yasumatu, R; Nakashima, T; Kuratomi, Y; Azuma, K; Tomita, K; Komiyama, S; Weinstein, IB. Constitutive Activation of Signal Transducers and Activators of Transcription 3 Correlates with Cyclin D1 Overexpression and May Provide a Novel Prognostic Marker in Head and Neck Squamous Cell Carcinoma. Cancer Res 2002, 62, 3351–3355. [Google Scholar]
- Yu, ZW; Weinberger, PM; Haffty, BG; Sasaki, C; Zerillo, C; Joe, J; Kowalski, D; Dziura, J; Camp, RL; Rimm, DL; Psyrri, A. Cyclin D1 is a valuable prognostic marker in oropharyngeal squamous cell carcinoma. Clin. Cancer Res 2005, 11, 1160–1166. [Google Scholar]
- Strojan, P; Budihna, M; Smid, L; Svetic, B; Vrhovec, I; Kos, J; Skrk, J. Prognostic significance of cysteine proteinases cathepsins B and L and their endogenous inhibitors stefins A and B in patients with squamous cell carcinoma of the head and neck. Clin. Cancer Res 2000, 6, 1052–1062. [Google Scholar]
- Yang, M-H; Wu, M-Z; Chiou, S-H; Chen, P-M; Chang, S-Y; Liu, C-J; Teng, S-C; Wu, K-J. Direct regulation of TWIST by HIF-1 alpha promotes metastasis. Nat. Cell Biol 2008, 10, 295–305. [Google Scholar]
- Nozawa, H; Howell, G; Suzuki, S; Zhang, Q; Qi, Y; Klein-Seetharaman, J; Wells, A; Grandis, JR; Thomas, SM. Combined Inhibition of PLC gamma-1 and c-Src Abrogates Epidermal Growth Factor Receptor-Mediated Head and Neck Squamous Cell Carcinoma Invasion. Clin. Cancer Res 2008, 14, 4336–4344. [Google Scholar]
- Thomas, SM; Coppelli, FM; Wells, A; Gooding, WE; Song, J; Kassis, J; Drenning, SD; Grandis, JR. Epidermal growth factor receptor-stimulated activation of phospholipase C gamma-1 promotes invasion of head and neck squamous cell carcinoma. Cancer Res 2003, 63, 5629–5635. [Google Scholar]
- Lu, Z; Jiang, G; Blume-Jensen, P; Hunter, T. Epidermal Growth Factor-Induced Tumor Cell Invasion and Metastasis Initiated by Dephosphorylation and Downregulation of Focal Adhesion Kinase. Mol. Cell. Biol 2001, 21, 4016–4031. [Google Scholar]
- Seethala, RR; Gooding, WE; Handler, PN; Collins, B; Zhang, Q; Siegfried, JM; Grandis, JR. Immunohistochemical Analysis of Phosphotyrosine Signal Transducer and Activator of Transcription 3 and Epidermal Growth Factor Receptor Autocrine Signaling Pathways in Head and Neck Cancers and Metastatic Lymph Nodes. Clin. Cancer Res 2008, 14, 1303–1309. [Google Scholar]
- Xia, W; Lau, Y-K; Zhang, H-Z; Xiao, F-Y; Johnston, DA; Liu, A-R; Li, L; Katz, RL; Hung, M-C. Combination of EGFR, HER-2/neu, and HER-3 Is a Stronger Predictor for the Outcome of Oral Squamous Cell Carcinoma Than Any Individual Family Members. Clin. Cancer Res 1999, 5, 4164–4174. [Google Scholar]
- Rehman, AO; Wang, C-Y. SDF-1 alpha promotes invasion of head and neck squamous cell carcinoma by activating NF-kappa B. J. Biol. Chem 2008, 283, 19888–19894. [Google Scholar]
- Zhou, HY; Wan, KF; Ip, CKM; Wong, CKC; Mak, NK; Lo, KW; Wong, AST. Hepatocyte growth factor enhances proteolysis and invasiveness of human nasopharyngeal cancer cells through activation of PI3K and JNK. FEBS Lett 2008, 582, 3415–3422. [Google Scholar]
- Yang, MH; Chang, SY; Chiou, SH; Liu, CJ; Chi, CW; Chen, PM; Teng, SC; Wu, KJ. Overexpression of NBS1 induces epithelial-mesenchymal transition and co-expression of NBS1 and Snail predicts metastasis of head and neck cancer. Oncogene 2006, 26, 1459–1467. [Google Scholar]
- Li, X; Yang, Y; Hu, Y; Dang, D; Regezi, J; Schmidt, BL; Atakilit, A; Chen, B; Ellis, D; Ramos, DM. αvβ6-Fyn Signaling Promotes Oral Cancer Progression. J. Biol. Chem 2003, 278, 41646–41653. [Google Scholar]
- Hassan, KA; Ang, KK; El-Naggar, AK; Story, MD; Lee, JI; Liu, D; Hong, WK; Mao, L. Cyclin B1 Overexpression and Resistance to Radiotherapy in Head and Neck Squamous Cell Carcinoma. Cancer Res 2002, 62, 6414–6417. [Google Scholar]
- Lai, SY; Childs, EE; Xi, S; Coppelli, FM; Gooding, WE; Wells, A; Ferris, RL; Grandis, JR. Erythropoietin-mediated activation of JAK-STAT signaling contributes to cellular invasion in head and neck squamous cell carcinoma. Oncogene 2005, 24, 4442–4449. [Google Scholar]
- Aronsohn, M; Brown, H; Hauptman, G; Kornberg, LJ. Expression of focal adhesion kinase and phosphorylated focal adhesion kinase in squamous cell carcinoma of the larynx. Laryngoscope 2003, 113, 1944–1948. [Google Scholar]
- Xia, W; Lau, YK; Zhang, HZ; Liu, AR; Li, L; Kiyokawa, N; Clayman, GL; Katz, RL; Hung, MC. Strong correlation between c-erbB-2 overexpression and overall survival of patients with oral squamous cell carcinoma. Clin. Cancer Res 1997, 3, 3–9. [Google Scholar]
- Ramos, DM; But, M; Regezi, J; Schmidt, BL; Atakilit, A; Dang, D; Ellis, D; Jordan, R; Li, X. Expression of integrin beta 6 enhances invasive behavior in oral squamous cell carcinoma. Matrix Biol 2002, 21, 297–307. [Google Scholar]
- Bancroft, CC; Chen, Z; Dong, G; Sunwoo, JB; Yeh, N; Park, C; van Waes, C. Coexpression of Proangiogenic Factors IL-8 and VEGF by Human Head and Neck Squamous Cell Carcinoma Involves Coactivation by MEK-MAPK and IKK-NF-κB Signal Pathways. Clin. Cancer Res 2001, 7, 435–442. [Google Scholar]
- Dong, G; Chen, Z; Li, Z-Y; Yeh, NT; Bancroft, CC; van Waes, C. Hepatocyte Growth Factor/Scatter Factor-induced Activation of MEK and PI3K Signal Pathways Contributes to Expression of Proangiogenic Cytokines Interleukin-8 and Vascular Endothelial Growth Factor in Head and Neck Squamous Cell Carcinoma. Cancer Res 2001, 61, 5911–5918. [Google Scholar]
- Brunner, A; Ensinger, C; Christiansen, M; Heiss, S; Verdorfer, I; Mikuz, G; Tzankov, A. Expression and prognostic significance of Tetranectin in invasive and non-invasive bladder cancer. Virchows Arch 2007, 450, 659–664. [Google Scholar]
- Hogdall, CK; Soletormos, G; Nielsen, D; Norgaardpedersen, B; Dombernowsky, P; Clemmensen, I. Prognostic value of serum tetranectin in patients with metastatic breast cancer. Acta Oncol 1993, 32, 631–636. [Google Scholar]
- Jensen, BA; Clemmensen, I. Plasma tetranectin is reduced in cancer and related to metastasis. Cancer 1988, 62, 869–872. [Google Scholar]
- Li, Y; Elashoff, D; Oh, M; Sinha, U; St John, MAR; Zhou, X; Abemayor, E; Wong, DT. Serum Circulating Human mRNA Profiling and Its Utility for Oral Cancer Detection. J. Clin. Oncol 2006, 24, 1754–1760. [Google Scholar]
- Hu, S; Zhou, M; Jiang, J; Wang, J; Elashoff, D; Gorr, S; Michie, SA; Spijkervet, FKL; Bootsma, H; Kallenberg, CGM; Vissink, A; Horvath, S; Wong, DT. Systems biology analysis of sjoren’s syndrome and mucosa-associated lymphoid tissue lymphoma in parotid glands. Arthritis Rheum 2009, 60, 81–92. [Google Scholar]
- Chen, Y; Lin, S; Yeh, Y; Hsiao, H; Wu, C; Chen, S; Wang, A. A modified protein precipitation procedure for efficient removal of albumin from serum. Electrophoresis 2005, 26, 2117–2127. [Google Scholar]
- Nielsen, BB; Kastrup, JS; Rasmussen, H; Holtet, TL; Graversen, JH; Etzerodt, M; Th⊘gersen, HC; Larsen, IK. Crystal structure of tetranectin, a trimeric plasminogen-binding protein with an alpha-helical coiled coil. FEBS Lett 1997, 412, 388–396. [Google Scholar]
- Clemmensen, I; Petersen, LC; Kluft, C. Purification and characterization of a novel, oligomeric, plasminogen kringle 4 binding protein from human plasma: Tetranectin. Eur. J. Biochem 1986, 156, 327–333. [Google Scholar]
- Andreasen, PA; Egelund, R; Petersen, HH. The plasminogen activation system in tumor growth, invasion, and metastasis. Cell. Mol. Life Sci 2000, 57, 25–40. [Google Scholar]
- Westergaard, UB; Andersen, MH; Heegaard, CW; Fedosov, SN; Petersen, TE. Tetranectin binds hepatocyte growth factor and tissue-type plasminogen activator. Eur. J. Biochem 2003, 270, 1850–1854. [Google Scholar]
- H⊘gdall, C; Christiansen, M; N⊘rgaard-Pedersen, B; Bentzen, S; Kronborg, O; Clemmensen, I. Plasma tetranectin and colorectal cancer. Eur. J. Cancer 1995, 31A, 888–894. [Google Scholar]
- H⊘gdall, C; H⊘gdall, E; H⊘rding, U; Clemmensen, I; N⊘rgaard-Pedersen, B; Toftager-Larsen, K. Pre-operative plasma tetranectin as a prognostic marker in ovarian cancer patients. Scand. J. Clin. Lab. Invest 1993, 53, 741–746. [Google Scholar]
- Christensen, L; Clemmensen, I. Differences in tetranectin immunoreactivity between benign and malignant breast tissue. Histochemistry 1991, 95, 427–433. [Google Scholar]
- Wewer, U; Albrechtsen, R. Tetranectin, a plasminogen kringle 4-binding protein. Cloning and gene expression pattern in human colon cancer. Lab. Invest 1992, 67, 253–262. [Google Scholar]
- de Vries, TJ; de Wit, PEJ; Clemmensen, I; Verspaget, HW; Weidle, UH; Brocker, EB; Ruiter, DJ; van Muijen, GNP. Tetranectin and plasmin/plasminogen are similarly distributed at the invasive front of cutaneous melanoma lesions. J. Pathol 1996, 179, 260–265. [Google Scholar]
- Mogues, T; Etzerodt, M; Hall, C; Engelich, G; Graversen, J; Hartshorn, K. Tetranectin binds to the kringle 1–4 form of angiostatin and modifies Its functional activity. J. Biomed. Biotechnol 2004, 2, 73–78. [Google Scholar]
- Hu, S; Arellano, M; Boontheung, P; Wang, J; Zhou, H; Jiang, J; Elashoff, D; Wei, R; Loo, JA; Wong, DT. Salivary proteomics for oral cancer biomarker discovery. Clin. Cancer Res 2008, 14, 6246–6252. [Google Scholar]
- Hu, S; Wang, J; Meijer, J; Ieong, S; Xie, Y; Yu, T; Zhou, H; Henry, S; Vissink, A; Pijpe, J; Kallenberg, C; Elashoff, D; Loo, J; Wong, D. Salivary proteomic and genomic biomarkers for primary Sjögren’s syndrome. Arthritis Rheum 2007, 56, 3588–3600. [Google Scholar]
- Zhang, L; Farrell, JJ; Zhou, H; Elashoff, D; Akin, D; Park, N-H; Chia, D; Wong, DT. Salivary transcriptomic biomarkers for detection of resectable pancreatic cancer. Gastroenterology 2010, 138, 949–957. [Google Scholar]
- Denny, P; Hagen, FK; Hardt, M; Liao, L; Yan, W; Arellanno, M; Bassilian, S; Bedi, GS; Boontheung, P; Cociorva, D; et al. The proteomes of human parotid and submandibular/sublingual gland salivas collected as the ductal secretions. J. Proteome Res 2008, 7, 1994–2006. [Google Scholar]
- Loo, J; Yan, W; Ramachandran, P; Wong, D. Comparative human salivary and plasma proteomes. J. Dent. Res 2010, in press. [Google Scholar]
- Yan, W; Apweiler, R; Balgley, B; Boontheung, P; Bundy, J; Cargile, B; Cole, S; Fang, X; Gonzalez-Begne, M; Griffin, T; et al. Systematic comparison of the human saliva and plasma proteomes. Proteomics Clin. Appl 2009, 3, 116–134. [Google Scholar]
- Findeisen, P; Zapatka, M; Peccerella, T; Matzk, H; Neumaier, M; Schadendorf, D; Ugurel, S. Serum amyloid A as a prognostic marker in melanoma identified by proteomic profiling. J. Clin. Oncol 2009, 27, 2199–2208. [Google Scholar]

| Protein | Up/Down | Role in metastasis | Reference |
|---|---|---|---|
| Chemokine receptor 6 | Down-regulation | Controls immune cell trafficking in response to inflammatory stimuli | [11] |
| Endostatin & Collagen XVII | Down-regulation | Angiogenesis inhibitor | [12] |
| CD44 | Down-regulation | Cell adhesion molecule | [13,14] |
| Lin-7C/VELI3/MALS-3 | Down-regulation | Signals through β-catenin | [15] |
| E-cadherin, β-catenin | Down-regulation | Adhesion molecule, reduction in cell-cell adhesion | [14] |
| Maspin | Down-regulation | Member of the serpin family of protease inhibitors | [16] |
| NM23-H1/NDPK | Down-regulation | Metastasis suppressor factor | [17,18] |
| Nicotinamide N-Methyltransferase (NNMT) | Down-regulation | Enzyme participating in nicotinate and nicotinamide metabolism | [19] |
| PAI-2 | Down-regulation | Inhibits conversion of plasminogen to plasmin and inhibit fibrolysis | [20] |
| Pemphigus vulgaris antigen | Up-regulation | Cell adhesion molecule | [21] |
| MMP-1, 2, 3, 9, MT1-MMP, TIMP-1 | Up-regulation | Proteolytic activity against the components of ECM, MT1-MMP activates MMP-2, TIMP-1 inhibits MMP-2 | [22] |
| Chemokine receptor 7 | Up-regulation | Mediator of immune cell survival and migration to lymph nodes | [11,23] |
| Cyclin D1 | Up-regulation | Cell cycle regulator | [24,25] |
| Cathepsin B & L | Up-regulation | Lysosomal proteolytic enzymes | [26] |
| Stefin A and B | Up-regulation | Inhibitors of cathepsin B and L | [26] |
| HIF-1 alpha | Up-regulation | Promotes tumor progression and metastasis | [27] |
| TWIST | Up-regulation | Essential mediator of cancer metastasis | [27] |
| PLC γ-1 | Up-regulation | Promotes tumor cell invasion when activated by EGFR | [28,29] |
| c-Src | Up-regulation | Proto-oncogene | [28] |
| EGFR & TGF-alpha | Up-regulation | Cell proliferation activator | [30–32] |
| pSTAT3 | Up-regulation | Transcription factor, regulates MMP-2 | [24,31] |
| SDF-1 alpha | Up-regulation | Promotes metastasis through activation of NF-kappa B signaling | [33] |
| Met | Up-regulation | Increases MMP-9 | [34] |
| NBS1 | Up-regulation | Up-regulates Snail and MMP-2 | [35] |
| Integrin beta 6 | Up-regulation | Activates Fyn and promotes oral cancer progression | [36] |
| Cyclin B1 | Up-regulation | Cell cycle regulator | [37] |
| Erythropoietin (EPO) & EPO receptor | Up-regulation | Activates JAK-STAT signaling and cell invasion | [38] |
| FAK | Up-regulation | Cell differentiation and cell invasion | [39] |
| HER-2 & HER-3 | Up-regulation | Cell growth and differentiation | [32,40] |
| Integrin alpha v beta 6 | Up-regulation | Promotes tumor growth and invasion, activates MMP-3 | [41] |
| VEGF & Interleukin 8 | Up-regulation | Promotes tumor angiogenesis, growth and metastasis | [42] |
| Hepatocyte growth factor (HGF) | Up-regulation | Induces the expression of Interleukin 8 and VEGF | [43] |
| Accession | Protein Name | Isoelectric Point | Molecular Weight | Peptides Identified | P Value | Ratio (ctrl/cancer) |
|---|---|---|---|---|---|---|
| IPI00332128 | 48 kD Protein | 5 | 48396 | 2 | 0.021 | 0.28 |
| IPI00384697 | ALB protein | 5.97 | 47330 | 8 | 0.048 | 1.72 |
| IPI00022429 | Alpha-1-acid glycoprotein 1 | 5.11 | 24770 | 13 | 0.030 | 1.99 |
| IPI00020091 | Alpha-1-acid glycoprotein 2 | 5.03 | 23588 | 2 | 0.029 | 1.59 |
| IPI00305457 | Alpha-1-antitrypsin | 5.19 | 48208 | 12 | 0.018 | 0.47 |
| IPI00021842 | Apolipoprotein E | 5.65 | 36132 | 6 | 0.040 | 1.85 |
| IPI00177869 | Apolipoprotein L1 Isoform A | 5.6 | 43947 | 2 | 0.015 | 1.68 |
| IPI00025204 | CD-5 Antigen like protein | 5.28 | 38063 | 7 | 0.029 | 1.59 |
| IPI00164623 | Complement C3 fragment | 6.02 | 187046 | 4 | 0.022 | 0.00 |
| IPI00027827 | Extracellular superoxide dismutase | 6.14 | 25865 | 2 | 0.031 | 0.27 |
| IPI00017530 | Ficolin 2 | 6.09 | 33998 | 2 | 0.019 | 0.65 |
| IPI00478493 | Haptoglobin | 6.13 | 45177 | 10 | 0.041 | 0.35 |
| IPI00296170 | Haptoglobin-related protein | 6.42 | 43049 | 2 | 0.003 | 2.00 |
| IPI00477597 | Haptoglobin-related protein | 6.42 | 38983 | 5 | 0.012 | 5.59 |
| IPI00431645 | HP protein | 8.48 | 31362 | 4 | 0.006 | 0.55 |
| IPI00472610 | Hypothetical protein | 7.5 | 52633 | 10 | 0.013 | 2.32 |
| IPI00332161 | Ig gamma 1 Chain C region | 8.46 | 36083 | 8 | 0.020 | 0.41 |
| IPI00385058 | Ig kappa chain C region | 7.51 | 26283 | 4 | 0.031 | 1.81 |
| IPI00335356 | Ig mu chain C region | 6.35 | 49526 | 8 | 0.029 | 0.31 |
| IPI00022420 | Plasma retinol-binding protein | 5.76 | 23029 | 5 | 0.024 | 2.55 |
| IPI00009544 | RalA binding protein 1 | 5.68 | 76016 | 2 | 0.008 | 1.84 |
| IPI00009544 | RalA binding protein 1 | 5.68 | 76016 | 2 | 0.034 | 1.73 |
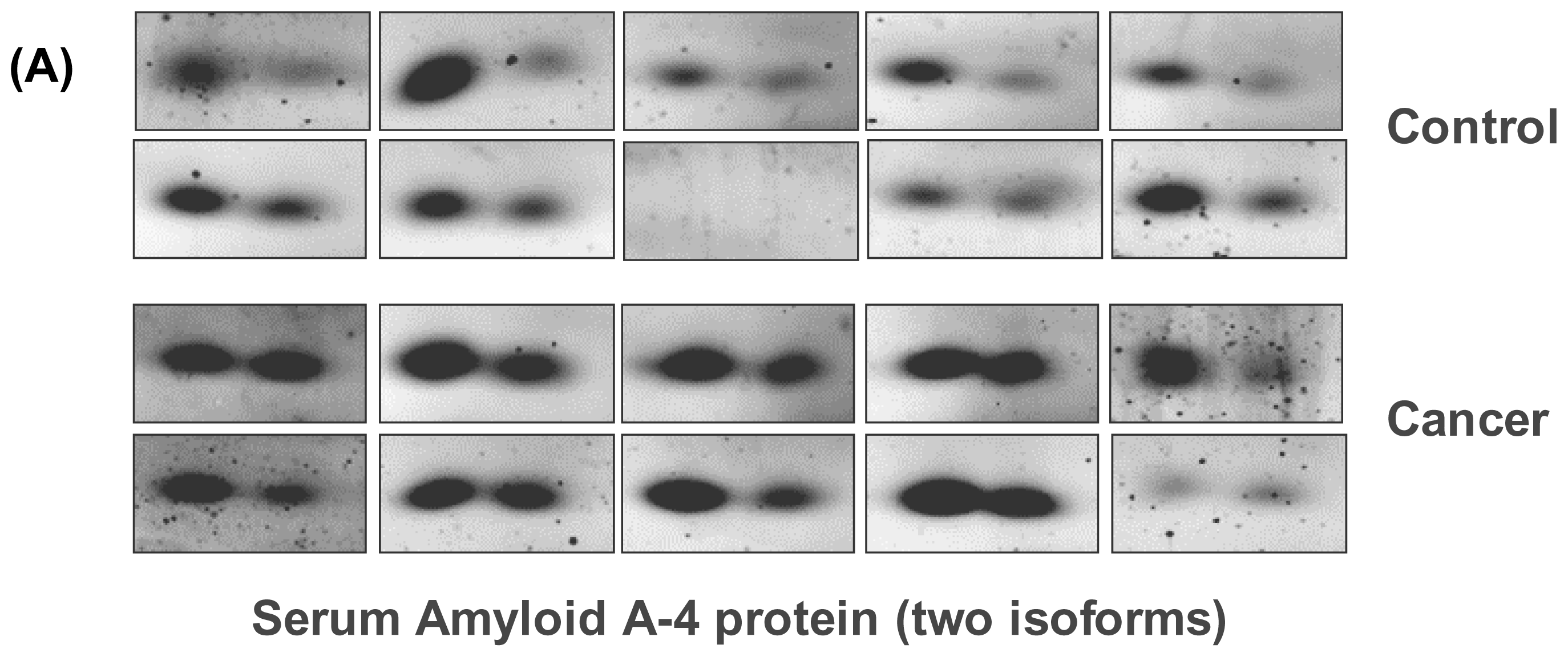
| IPI00019399 | Serum Amyloid A-4 protein isoform 1 | 9.27 | 14797 | 2 | 0.044 | 0.37 |
| IPI00019399 | Serum amyloid A-4 protein isoform 2 | 9.27 | 14797 | 3 | 0.038 | 0.58 |
| IPI00550061 | Similar to Ig gamma 3 Chain C region | 7.47 | 56823 | 6 | 0.032 | 1.67 |
| IPI00221125 | Voltage-gated potassium channel beta-1 subunit | 9.23 | 45462 | 4 | 0.006 | 1.79 |
| IPI00009028 | Tetranectin | 5.52 | 22552 | 7 | 0.045 | 1.84 |
| IPI00009028 | Tetranectin | 5.52 | 22552 | 8 | 0.030 | 1.68 |
| IPI00022432 | Transthyretin | 5.52 | 15877 | 16 | 0.022 | 1.45 |
| IPI00022432 | Transthyretin | 5.52 | 15877 | 7 | 0.040 | 3.76 |
© 2010 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Arellano-Garcia, M.E.; Li, R.; Liu, X.; Xie, Y.; Yan, X.; Loo, J.A.; Hu, S. Identification of Tetranectin as a Potential Biomarker for Metastatic Oral Cancer. Int. J. Mol. Sci. 2010, 11, 3106-3121. https://doi.org/10.3390/ijms11093106
Arellano-Garcia ME, Li R, Liu X, Xie Y, Yan X, Loo JA, Hu S. Identification of Tetranectin as a Potential Biomarker for Metastatic Oral Cancer. International Journal of Molecular Sciences. 2010; 11(9):3106-3121. https://doi.org/10.3390/ijms11093106
Chicago/Turabian StyleArellano-Garcia, Martha E., Roger Li, Xiaojun Liu, Yongming Xie, Xiaofei Yan, Joseph A. Loo, and Shen Hu. 2010. "Identification of Tetranectin as a Potential Biomarker for Metastatic Oral Cancer" International Journal of Molecular Sciences 11, no. 9: 3106-3121. https://doi.org/10.3390/ijms11093106
APA StyleArellano-Garcia, M. E., Li, R., Liu, X., Xie, Y., Yan, X., Loo, J. A., & Hu, S. (2010). Identification of Tetranectin as a Potential Biomarker for Metastatic Oral Cancer. International Journal of Molecular Sciences, 11(9), 3106-3121. https://doi.org/10.3390/ijms11093106





