The Research Progress of Bioactive Peptides Derived from Traditional Natural Products in China
Abstract
1. Introduction
2. The Preparation Method of Bioactive Peptides
2.1. Enzymatic Hydrolysis
2.2. Solvent Extraction
3. The Separation Method of Bioactive Peptides
3.1. Membrane Separation
3.2. Gel Filtration Chromatography
3.3. Ion Exchange Chromatography
3.4. Reversed-Phase High-Performance Liquid Chromatography
3.5. Multidimensional Chromatographic Separation
4. The Identification of Bioactive Peptides
4.1. Identification Based on Database Search
4.2. Peptide Identification Based on De Novo
4.3. Methods Combination
5. Functional Classification of Bioactive Peptides
5.1. Antioxidant Peptides
5.2. Anti-Hypertensive Peptides
5.3. Anti-Inflammatory Peptides
5.4. Anti-Cancer Peptides
5.5. Other Functional Peptides
6. Discussion
7. Conclusions and Prospect
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Sample Availability
References
- Wang, Z.-G.; Ren, J. Current Status and Future Direction of Chinese Herbal Medicine. Trends Pharmacol. Sci. 2002, 23, 347–348. [Google Scholar] [CrossRef]
- Jiao, Q.; Wang, R.; Jiang, Y.; Liu, B. Study on the Interaction between Active Components from Traditional Chinese Medicine and Plasma Proteins. Chem. Cent. J. 2018, 12, 48. [Google Scholar] [CrossRef]
- Zhu, J. Through a Century: Traditional Chinese Medicine Since 1912. Chin. Med. Cult. 2018, 1, 5–10. [Google Scholar] [CrossRef]
- Li, Q.-Q.; Liu, Z.-Z.; Zhang, J.; Xu, D.-L. Research progress in the biosynthesis of medicinal active ingredients of traditional. Chin. Tradit. Pat. Med. 2022, 44, 3603–3608. [Google Scholar]
- Zhang, W.-J.; Wang, S.; Kang, C.; Lv, C.; Zhou, L.; Huang, L.-Q.; Guo, L.-P. Pharmacodynamic Material Basis of Traditional Chinese Medicine Based on Biomacromolecules: A Review. Plant Methods 2020, 16, 26. [Google Scholar] [CrossRef] [PubMed]
- Lafarga, T.; Hayes, M. Bioactive Peptides from Meat Muscle and By-Products: Generation, Functionality and Application as Functional Ingredients. Meat Sci. 2014, 98, 227–239. [Google Scholar] [CrossRef] [PubMed]
- Rutherfurd-Markwick, K.J. Food Proteins as a Source of Bioactive Peptides with Diverse Functions. Br. J. Nutr. 2012, 108, S149–S157. [Google Scholar] [CrossRef] [PubMed]
- Manzanares, P.; Salom, J.B.; García-Tejedor, A.; Fernández-Musoles, R.; Ruiz-Giménez, P.; Gimeno-Alcañíz, J.V. Unraveling the Mechanisms of Action of Lactoferrin-Derived Antihypertensive Peptides: ACE Inhibition and Beyond. Food Funct. 2015, 6, 2440–2452. [Google Scholar] [CrossRef]
- Daliri, E.B.-M.; Lee, B.H.; Oh, D.H. Current Trends and Perspectives of Bioactive Peptides. Crit. Rev. Food Sci. Nutr. 2018, 58, 2273–2284. [Google Scholar] [CrossRef]
- Korhonen, H.; Pihlanto, A. Bioactive Peptides: Production and Functionality. Int. Dairy J. 2006, 16, 945–960. [Google Scholar] [CrossRef]
- Hartmann, R.; Meisel, H. Food-Derived Peptides with Biological Activity: From Research to Food Applications. Curr. Opin. Biotechnol. 2007, 18, 163–169. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Wang, Y.; Liu, Y.; Ruan, R. Bioactive Peptides Derived from Traditional Chinese Medicine and Traditional Chinese Food: A Review. Food Res. Int. 2016, 89, 63–73. [Google Scholar] [CrossRef]
- Williams, A.P. Enzymic Hydrolysis of Food Proteins. Food Chem. 1987, 26, 81–82. [Google Scholar] [CrossRef]
- Udenigwe, C.; Rouvinen-Watt, K. The Role of Food Peptides in Lipid Metabolism during Dyslipidemia and Associated Health Conditions. Int. J. Mol. Sci. 2015, 16, 9303–9313. [Google Scholar] [CrossRef] [PubMed]
- Marciniak, A.; Suwal, S.; Naderi, N.; Pouliot, Y.; Doyen, A. Enhancing Enzymatic Hydrolysis of Food Proteins and Production of Bioactive Peptides Using High Hydrostatic Pressure Technology. Trends Food Sci. Technol. 2018, 80, 187–198. [Google Scholar] [CrossRef]
- de Castro, R.J.S.; Sato, H.H. Biologically Active Peptides: Processes for Their Generation, Purification and Identification and Applications as Natural Additives in the Food and Pharmaceutical Industries. Food Res. Int. 2015, 74, 185–198. [Google Scholar] [CrossRef]
- Wang, X.; Deng, Y.; Xie, P.; Liu, L.; Zhang, C.; Cheng, J.; Zhang, Y.; Liu, Y.; Huang, L.; Jiang, J. Novel Bioactive Peptides from Ginkgo Biloba Seed Protein and Evaluation of Their α-Glucosidase Inhibition Activity. Food Chem. 2023, 404, 134481. [Google Scholar] [CrossRef]
- Cao, X.; Yang, J.; Ma, H.; Guo, P.; Cai, Y.; Xu, H.; Ding, G.; Gao, D. Angiotensin I Converting Enzyme (ACE) Inhibitory Peptides Derived from Alfalfa (Medicago Sativa L.) Leaf Protein and Its Membrane Fractions. J. Food Process. Preserv. 2021, 45, e15834. [Google Scholar] [CrossRef]
- Guo, Z.; Lin, D.; Guo, J.; Zhang, Y.; Zheng, B. In Vitro Antioxidant Activity and In Vivo Anti-Fatigue Effect of Sea Horse (Hippocampus) Peptides. Molecules 2017, 22, 482. [Google Scholar] [CrossRef]
- Ghanbari, R.; Zarei, M.; Ebrahimpour, A.; Abdul-Hamid, A.; Ismail, A.; Saari, N. Angiotensin-I Converting Enzyme (ACE) Inhibitory and Anti-Oxidant Activities of Sea Cucumber (Actinopyga Lecanora) Hydrolysates. Int. J. Mol. Sci. 2015, 16, 28870–28885. [Google Scholar] [CrossRef]
- Memarpoor-Yazdi, M.; Zare-Zardini, H.; Mogharrab, N.; Navapour, L. Purification, Characterization and Mechanistic Evaluation of Angiotensin Converting Enzyme Inhibitory Peptides Derived from Zizyphus Jujuba Fruit. Sci. Rep. 2020, 10, 3976. [Google Scholar] [CrossRef]
- Li, S.-L.; Lin, J.; Jiang, C.-X.; Chen, X.-M.; Wang, X.-H.; Ye, H.; Wang, H. Study on preparation and antioxidant activity of the Lycium barbarum polypeptide. J. Light Ind. 2016, 31, 31–37. [Google Scholar]
- Bao, M.-L. Study on the Preparation and Properties of the Hypoglycemic Peptide from Velvet Antler. Master’s Thesis, Northeast Forestry University, Harbin, China, 2017. [Google Scholar]
- Liu, E.-Q.; Wu, Y.-H.; Zhang, J.-P.; Chen, Z.-J.; Li, Y.-E. Separation and Purification of Black Soybean Peptide and Its Hypolipidemic Effect in Mice. Food Sci. 2011, 32, 248–252. [Google Scholar]
- Wei, P.; You, X.-R.; Zhang, Y.-Y.; Xie, X.-Q.; Sun, J.; Li, M.-J. Preparation of Cannabis sativa L. Peptides by Enzyme Method and Their Activities of Inhibiting Angiotensin Converting Enzyme. Sci. Technol. Food Ind. 2019, 40, 127–132. [Google Scholar] [CrossRef]
- Fan, Y.-M.; Ru, W.-W.; Shi, C.-C.; Qian, L.-Y.; Liao, F. Composition and Its Antioxidant Activity of Colla Corii Asini Oligopeptides. Sci. Technol. Food Ind. 2020, 41, 314–318+323. [Google Scholar] [CrossRef]
- Liu, H.-J.; Le, C.-Y.; Shao, W.; Ye, J.-L.; Pan, H.; Li, J.-J. Researching advancement of bioactive peptides. China Brew. 2010, 5–8. [Google Scholar] [CrossRef]
- Sridhar, K.; Inbaraj, B.S.; Chen, B.-H. Recent Developments on Production, Purification and Biological Activity of Marine Peptides. Food Res. Int. 2021, 147, 110468. [Google Scholar] [CrossRef] [PubMed]
- Chen, R.; Wang, X.-J.; Zhang, Y.-Y.; Xing, Y.; Yang, L.; Ni, H.; Li, H.-H. Simultaneous Extraction and Separation of Oil, Proteins, and Glucosinolates from Moringa Oleifera Seeds. Food Chem. 2019, 300, 125162. [Google Scholar] [CrossRef]
- Liu, X.-J.; Lin, S.-J. Study on antiviral effect of isatis root polypeptide on mice infected with influenza virus. China Pharm. 2014, 25, 590–592. [Google Scholar]
- Kumar, M.; Tomar, M.; Potkule, J.; Verma, R.; Punia, S.; Mahapatra, A.; Belwal, T.; Dahuja, A.; Joshi, S.; Berwal, M.K.; et al. Advances in the Plant Protein Extraction: Mechanism and Recommendations. Food Hydrocoll. 2021, 115, 106595. [Google Scholar] [CrossRef]
- Cui, Q.; Ni, X.; Zeng, L.; Tu, Z.; Li, J.; Sun, K.; Chen, X.; Li, X. Optimization of Protein Extraction and Decoloration Conditions for Tea Residues. Hortic. Plant J. 2017, 3, 172–176. [Google Scholar] [CrossRef]
- Liu, J.; Wang, H.; Luo, Q.; Qiu, S.; He, Z.; Liu, Z.; Yang, L.; Liu, X.; Sun, X. LingZhi Oligopeptides Amino Acid Sequence Analysis and Anticancer Potency Evaluation. RSC Adv. 2020, 10, 8377–8384. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Xu, P.; Wang, Y.; Gao, T.-T.; Pan, H.-N. The effect of gecko crude polypeptide on the proliferation and expression of autophagy-related protein of HepG2 cells. Chin. J. Clin. Pharmacol. 2019, 35, 1791–1793. [Google Scholar] [CrossRef]
- Qian, G.; Pan, G.-F.; Guo, J.-Y. Anti-Inflammatory and Antinociceptive Effects of Cordymin, a Peptide Purified from the Medicinal Mushroom. Nat. Prod. Res. 2012, 26, 2358–2362. [Google Scholar] [CrossRef]
- Tie, R.; Ji, L.; Nan, Y.; Wang, W.; Liang, X.; Tian, F.; Xing, W.; Zhu, M.; Li, R.; Zhang, H. Achyranthes Bidentata Polypeptides Reduces Oxidative Stress and Exerts Protective Effects against Myocardial Ischemic/Reperfusion Injury in Rats. Int. J. Mol. Sci. 2013, 14, 19792–19804. [Google Scholar] [CrossRef]
- Zhang, C.; Sanders, J.P.M.; Xiao, T.T.; Bruins, M.E. How Does Alkali Aid Protein Extraction in Green Tea Leaf Residue: A Basis for Integrated Biorefinery of Leaves. PLoS ONE 2015, 10, e0133046. [Google Scholar] [CrossRef]
- Acquah, C.; Chan, Y.W.; Pan, S.; Agyei, D.; Udenigwe, C.C. Structure-Informed Separation of Bioactive Peptides. J. Food Biochem. 2019, 43, e12765. [Google Scholar] [CrossRef]
- Wu, S.-H.; Liu, G.-M. Research progress in protein separation and purification methods. China Pharm. 2012, 21, 1–3. [Google Scholar]
- Capriotti, A.L.; Cavaliere, C.; Foglia, P.; Piovesana, S.; Samperi, R.; Zenezini Chiozzi, R.; Laganà, A. Development of an Analytical Strategy for the Identification of Potential Bioactive Peptides Generated by In Vitro Tryptic Digestion of Fish Muscle Proteins. Anal. Bioanal. Chem. 2015, 407, 845–854. [Google Scholar] [CrossRef]
- Ye, J.; Shen, C.; Huang, Y.; Zhang, X.; Xiao, M. Anti-Fatigue Activity of Sea Cucumber Peptides Prepared from Stichopus Japonicus in an Endurance Swimming Rat Model: Anti-Fatigue Activity of Sea Cucumber Peptides. J. Sci. Food Agric. 2017, 97, 4548–4556. [Google Scholar] [CrossRef]
- Ding, Y.; Ko, S.-C.; Moon, S.-H.; Lee, S.-H. Protective Effects of Novel Antioxidant Peptide Purified from Alcalase Hydrolysate of Velvet Antler Against Oxidative Stress in Chang Liver Cells In Vitro and in a Zebrafish Model In Vivo. Int. J. Mol. Sci. 2019, 20, 5187. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.-K. Study on Preparation of Immunoactive Soybean Peptides by Membrane Separation Method. Master’s Thesis, Jiangnan University, Wuxi, China, 2007. [Google Scholar]
- Yuan, X.; Gu, X.; Tang, J. Purification and Characterisation of a Hypoglycemic Peptide from Momordica Charantia L. Var. Abbreviata Ser. Food Chem. 2008, 111, 415–420. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Tan, S.; Luo, J.; Pinelo, M. Nanofiltration for Separation and Purification of Saccharides from Biomass. Front. Chem. Sci. Eng. 2021, 15, 837–853. [Google Scholar] [CrossRef]
- Muro Urista, C.; Álvarez Fernández, R.; Riera Rodriguez, F.; Arana Cuenca, A.; Téllez Jurado, A. Review: Production and Functionality of Active Peptides from Milk. Food Sci. Technol. Int. 2011, 17, 293–317. [Google Scholar] [CrossRef]
- Georgiou, H.M.; Rice, G.E.; Baker, M.S. Proteomic Analysis of Human Plasma: Failure of Centrifugal Ultrafiltration to Remove Albumin and Other High Molecular Weight Proteins. Proteomics 2001, 1, 1503–1506. [Google Scholar] [CrossRef]
- Capriotti, A.L.; Caruso, G.; Cavaliere, C.; Piovesana, S.; Samperi, R.; Laganà, A. Comparison of Three Different Enrichment Strategies for Serum Low Molecular Weight Protein Identification Using Shotgun Proteomics Approach. Anal. Chim. Acta. 2012, 740, 58–65. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Du, M.-X.; Wang, F.-L.; Affirmative, M.; Xie, J.-H.; Xie, M.-Y. Research progress in preparation and purification of bioactive peptides. Sci. Technol. Food Ind. 2017, 38, 336–340+346. [Google Scholar] [CrossRef]
- Mora, L.; Escudero, E.; Fraser, P.D.; Aristoy, M.-C.; Toldrá, F. Proteomic Identification of Antioxidant Peptides from 400 to 2500Da Generated in Spanish Dry-Cured Ham Contained in a Size-Exclusion Chromatography Fraction. Food Res. Int. 2014, 56, 68–76. [Google Scholar] [CrossRef]
- Wang, L.; Dong, C.; Li, X.; Han, W.; Su, X. Anticancer Potential of Bioactive Peptides from Animal Sources. Oncol. Rep. 2017, 38, 637–651. [Google Scholar] [CrossRef]
- Fekete, S.; Beck, A.; Veuthey, J.-L.; Guillarme, D. Theory and Practice of Size Exclusion Chromatography for the Analysis of Protein Aggregates. J. Pharm. Biomed. Anal. 2014, 101, 161–173. [Google Scholar] [CrossRef]
- Lee, S.H.; Song, K.B. Isolation of an Angiotensin Converting Enzyme Inhibitory Peptide from Irradiated Bovine Blood Plasma Protein Hydrolysates. J. Food Sci. 2003, 68, 2469–2472. [Google Scholar] [CrossRef]
- Hong, P.; Koza, S.; Bouvier, E.S.P. A Review Size-Exclusion Chromatography for the Analysis of Protein Biotherapeutics and Their Aggregates. J. Liq. Chromatogr. Relat. Technol. 2012, 35, 2923–2950. [Google Scholar] [CrossRef]
- Wang, Q.; Huang, Y.; Qin, C.; Liang, M.; Mao, X.; Li, S.; Zou, Y.; Jia, W.; Li, H.; Ma, C.W.; et al. Bioactive Peptides from Angelica sinensis Protein Hydrolyzate Delay Senescence in Caenorhabditis elegans through Antioxidant Activities. Oxidative Med. Cell. Longev. 2016, 2016, 8956981. [Google Scholar] [CrossRef]
- Mishra, J.; Rajput, R.; Singh, K.; Puri, S.; Goyal, M.; Bansal, A.; Misra, K. Antibacterial Natural Peptide Fractions from Indian Ganoderma Lucidum. Int. J. Pept. Res. Ther. 2018, 24, 543–554. [Google Scholar] [CrossRef]
- Zhou, H.-J.; Zhu, Z.B.; Yi, J.-H. Purification of ACE inhibitory peptides from enzymatic hydrolysate of walnut protein. Cereals Oils 2013, 26, 16–18. [Google Scholar]
- Gilar, M.; Olivova, P.; Daly, A.E.; Gebler, J.C. Orthogonality of Separation in Two-Dimensional Liquid Chromatography. Anal. Chem. 2005, 77, 6426–6434. [Google Scholar] [CrossRef] [PubMed]
- Otte, J.; Shalaby, S.M.; Zakora, M.; Nielsen, M.S. Fractionation and Identification of ACE-Inhibitory Peptides from α-Lactalbumin and β-Casein Produced by Thermolysin-Catalysed Hydrolysis. Int. Dairy J. 2007, 17, 1460–1472. [Google Scholar] [CrossRef]
- Wang, Q.; Li, W.; He, Y.; Ren, D.; Kow, F.; Song, L.; Yu, X. Novel Antioxidative Peptides from the Protein Hydrolysate of Oysters (Crassostrea talienwhanensis). Food Chem. 2014, 145, 991–996. [Google Scholar] [CrossRef]
- Levison, P.R. Large-Scale Ion-Exchange Column Chromatography of Proteins. Comparison of Different Formats. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 2003, 790, 17–33. [Google Scholar] [CrossRef]
- Schmidt, M.; Hafner, M.; Frech, C. Modeling of Salt and PH Gradient Elution in Ion-Exchange Chromatography. J. Sep. Sci. 2014, 37, 5–13. [Google Scholar] [CrossRef]
- Fekete, S.; Beck, A.; Veuthey, J.-L.; Guillarme, D. Ion-Exchange Chromatography for the Characterization of Biopharmaceuticals. J. Pharm. Anal. 2015, 113, 43–55. [Google Scholar] [CrossRef]
- Kammerer, J.; Carle, R.; Kammerer, D.R. Adsorption and Ion Exchange: Basic Principles and Their Application in Food Processing. J. Agric. Food Chem. 2011, 59, 22–42. [Google Scholar] [CrossRef] [PubMed]
- Coskun, O. Separation Tecniques: Chromatography. N. Clin. Istanb. 2016, 3, 156–160. [Google Scholar] [CrossRef]
- Yigzaw, Y.; Hinckley, P.; Hewig, A.; Vedantham, G. Ion Exchange Chromatography of Proteins and Clearance of Aggregates. Curr. Pharm. Biotechnol. 2009, 10, 421–426. [Google Scholar] [CrossRef]
- Zhao, C.-X.; Li, Y.-T.; Bu, F.-L.; Zheng, M. Lycium barbarum Cyclic Peptide Regulates NLRP3 and NF-κB Signaling Pathways to Reduce BaP-Induced Inflammatory Airway Epithelial Cell Injury. Mod. Food Sci. Technol. 2022, 38, 92–99. [Google Scholar] [CrossRef]
- Wang, Y.-X.; Jiang, Z.-Q.; Li, P.; Ding, T.-L.; Yang, X.-C. Research progress in separation and purification of bioactive peptides. J. Shandong Inst. Build. Mater. 2014, 28, 321–325. [Google Scholar] [CrossRef]
- Heemskerk, A.A.M.; Deelder, A.M.; Mayboroda, O.A. CE-ESI-MS for Bottom-up Proteomics: Advances in Separation, Interfacing and Applications: CE-Esi-Ms For Bottom-Up Proteomics. Mass Spectrom. Rev. 2016, 35, 259–271. [Google Scholar] [CrossRef]
- Gruber, K.A.; Stein, S.; Brink, L.; Radhakrishnan, A.; Udenfriend, S. Fluorometric Assay of Vasopressin and Oxytocin: A General Approach to the Assay of Peptides in Tissues. Proc. Natl. Acad. Sci. USA 1976, 73, 1314–1318. [Google Scholar] [CrossRef]
- Galli, V.; Mazzoli, L.; Luti, S.; Venturi, M.; Guerrini, S.; Paoli, P.; Vincenzini, M.; Granchi, L.; Pazzagli, L. Effect of Selected Strains of Lactobacilli on the Antioxidant and Anti-Inflammatory Properties of Sourdough. Int. J. Food Microbiol. 2018, 286, 55–65. [Google Scholar] [CrossRef]
- Qiu, H.; Liang, X.; Sun, M.; Jiang, S. Development of Silica-Based Stationary Phases for High-Performance Liquid Chromatography. Anal. Bioanal. Chem. 2011, 399, 3307–3322. [Google Scholar] [CrossRef]
- He, L.; Zhang, J.; Sun, Y.; Liu, J.; Jiang, X.; Qu, L. A Multiple-Function Stationary Phase Based on Perhydro-26-Membered Hexaazamacrocycle for High-Performance Liquid Chromatography. J. Chromatogr. A. 2010, 1217, 5971–5977. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Huang, Y.-B.; Gao, K.-X.; Sun, H.; Gao, Z.-L. Preparation, purification and antioxidant activity of enzymolysis antler peptide. Chem. J. Chin. Univ. 2010, 31, 2390–2395. [Google Scholar]
- Yang, L.-J.; You, Y.-H.; Lin, Z.-B.; Lin, Y.-F. Protective effect of Ganoderma lucidum polysaccharide peptide on oxidative damage of human umbilical vein endothelial cells. Chin. Pharmacol. Bull. 2010, 26, 657–660. [Google Scholar]
- Josic, D.; Kovac, S. Reversed-Phase High Performance Liquid Chromatography of Proteins. Curr. Protoc. Protein Sci. 2010, 61, 8.7.1–8.7.22. [Google Scholar] [CrossRef]
- Zare-Zardini, H.; Tolueinia, B.; Hashemi, A.; Ebrahimi, L.; Fesahat, F. Antioxidant and Cholinesterase Inhibitory Activity of a New Peptide from Ziziphus jujuba Fruits. Am. J. Alzheimer’s Dis. Other Dementiasr. 2013, 28, 702–709. [Google Scholar] [CrossRef]
- Lillehoj, E.P.; Malik, V.S. Protein purification. In Bioprocesses and Engineering; Advances in Biochemical Engineering/Biotechnology; Springer: Berlin/Heidelberg, Germany, 1989; Volume 40, pp. 19–71. [Google Scholar]
- Cacciola, F.; Mangraviti, D.; Rigano, F.; Donato, P.; Dugo, P.; Mondello, L.; Cortes, H.J. Novel Comprehensive Multidimensional Liquid Chromatography Approach for Elucidation of the Microbosphere of Shikimate-Producing Escherichia coli SP1.1/PKD15.071 Strain. Anal. Bioanal. Chem. 2018, 410, 3473–3482. [Google Scholar] [CrossRef]
- Everley, R.A.; Croley, T.R. Ultra-Performance Liquid Chromatography/Mass Spectrometry of Intact Proteins. J. Chromatogr. A. 2008, 1192, 239–247. [Google Scholar] [CrossRef] [PubMed]
- Di Palma, S.; Hennrich, M.L.; Heck, A.J.R.; Mohammed, S. Recent Advances in Peptide Separation by Multidimensional Liquid Chromatography for Proteome Analysis. J. Proteom. 2012, 75, 3791–3813. [Google Scholar] [CrossRef]
- Blacken, G.R.; Gelb, M.H.; Turecek, F. Metal Affinity Capture Tandem Mass Spectrometry for the Selective Detection of Phosphopeptides. Anal. Chem. 2006, 78, 6065–6073. [Google Scholar] [CrossRef][Green Version]
- Giddings, J.C. Two-Dimensional Separations: Concept and Promise. Anal. Chem. 1984, 56, 1258A–1270A. [Google Scholar] [CrossRef]
- Yu, T.-Y.; Morton, J.D.; Clerens, S.; Dyer, J.M. Cooking-Induced Protein Modifications in Meat. Compr. Rev. Food Sci. Food Saf. 2017, 16, 141–159. [Google Scholar] [CrossRef]
- Giddings, J.C. Concepts and Comparisons in Multidimensional Separation. J. High Resolut. Chromatogr. 1987, 10, 319–323. [Google Scholar] [CrossRef]
- Lan, Y.-R.; Huang, S.; Zhao, F.; Wu, H.-Y.; Guo, Y.-X. Optimization of enzymatic hydrolysis conditions for antioxidant peptide preparation from velvet antler collagen by response surface methodology. J. Process Eng. 2020, 20, 91–98. [Google Scholar]
- Zhang, J.; Xu, X.; Gao, M.; Yang, P.; Zhang, X. Comparison of 2-D LC and 3-D LC with Post- and Pre-Tryptic-Digestion SEC Fractionation for Proteome Analysis of Normal Human Liver Tissue. Proteomics 2007, 7, 500–512. [Google Scholar] [CrossRef]
- Gao, Y.; Qin, H.; Wu, D.; Liu, C.; Fang, L.; Wang, J.; Liu, X.; Min, W. Walnut Peptide WEKPPVSH in Alleviating Oxidative Stress and Inflammation in Lipopolysaccharide-Activated BV-2 Microglia via the Nrf2/HO-1 and NF-ΚB/P38 MAPK Pathways. J. Biosci. Bioeng. 2021, 132, 496–504. [Google Scholar] [CrossRef]
- Tanaka, K. The Origin of Macromolecule Ionization by Laser Irradiation (Nobel Lecture). Angew. Chem. Int. Ed. Engl. 2003, 42, 3860–3870. [Google Scholar] [CrossRef]
- Wilm, M.; Mann, M. Analytical Properties of the Nanoelectrospray Ion Source. Anal. Chem. 1996, 68, 1–8. [Google Scholar] [CrossRef]
- Rizzello, C.G.; Tagliazucchi, D.; Babini, E.; Rutella, G.S.; Saa, D.L.T.; Gianotti, A. Bioactive Peptides from Vegetable Food Matrices: Research Trends and Novel Biotechnologies for Synthesis and Recovery. J. Funct. Foods. 2016, 27, 549–569. [Google Scholar] [CrossRef]
- Zhang, X.; Fang, A.; Riley, C.P.; Wang, M.; Regnier, F.E.; Buck, C. Multi-Dimensional Liquid Chromatography in Proteomics—A Review. Anal. Chim. Acta. 2010, 664, 101–113. [Google Scholar] [CrossRef]
- Piovesana, S.; Capriotti, A.L.; Cavaliere, C.; La Barbera, G.; Montone, C.M.; Zenezini Chiozzi, R.; Laganà, A. Recent Trends and Analytical Challenges in Plant Bioactive Peptide Separation, Identification and Validation. Anal. Bioanal. Chem. 2018, 410, 3425–3444. [Google Scholar] [CrossRef]
- Fields, K.; Falla, T.J.; Rodan, K.; Bush, L. Bioactive Peptides: Signaling the Future. J. Cosmet. Dermatol. 2009, 8, 8–13. [Google Scholar] [CrossRef]
- Craig, R.; Beavis, R.C. TANDEM: Matching Proteins with Tandem Mass Spectra. Bioinformatics 2004, 20, 1466–1467. [Google Scholar] [CrossRef]
- Eng, J.K.; Searle, B.C.; Clauser, K.R.; Tabb, D.L. A Face in the Crowd: Recognizing Peptides through Database Search. Mol. Cell. Proteom. 2011, 10, R111.009522. [Google Scholar] [CrossRef] [PubMed]
- Nesvizhskii, A.I. Protein Identification by Tandem Mass Spectrometry and Sequence Database Searching. In Mass Spectrometry Data Analysis in Proteomics; Humana Press: Totowa, NJ, USA, 2006; Volume 367, pp. 87–120. ISBN 978-1-59745-275-5. [Google Scholar]
- Wang, Q.; Shi, J.; Zhong, H.; Abdullah; Zhuang, J.; Zhang, J.; Wang, J.; Zhang, X.; Feng, F. High-Degree Hydrolysis Sea Cucumber Peptides Improve Exercise Performance and Exert Antifatigue Effect via Activating the NRF2 and AMPK Signaling Pathways in Mice. J. Funct. Foods 2021, 86, 104677. [Google Scholar] [CrossRef]
- Wang, Y.-X.; Fan, Y.-M.; Liao, F.; Yu, Z.-P. Structure Identification and Activity Screening of Colla Corii Asini (E’jiao) Active Peptides. Food Sci. 2022, 43, 207–213. [Google Scholar]
- Kumar, D.; Yadav, A.K.; Dash, D. Choosing an Optimal Data. In Proteome Bioinformatics; Keerthikumar, S., Mathivanan, S., Eds.; Methods in Molecular Biology; Springer: New York, NY, USA, 2017; Volume 1549, pp. 17–29. ISBN 978-1-4939-6738-4. [Google Scholar]
- Silva do Nascimento, E.; Anaya, K.; de Oliveira, J.M.C.; de Lacerda, J.T.J.G.; Miller, M.E.; Dias, M.; Mendes, M.A.; de Azevedo Lima Pallone, J.; Weis Arns, C.; Juliano, M.A.; et al. Identification of Bioactive Peptides Released from In Vitro Gastrointestinal Digestion of Yam Proteins (Dioscorea cayennensis). Food Res. Int. 2021, 143, 110286. [Google Scholar] [CrossRef]
- Chai, T.T.; Soo, Z.-Y.; Hsu, K.-C.; Li, J.-C.; Abd, M.-F.; Wong, F.-C. Antioxidant Activity of Semen Cassiae Protein Hydrolysate: Thermal and Gastrointestinal Stability, Peptide Identification, and In Silico Analysis. Mod. Food Sci. Technol. 2019, 35, 38–48. [Google Scholar] [CrossRef]
- Taylor, J.A.; Johnson, R.S. Implementation and Uses of Automated de Novo Peptide Sequencing by Tandem Mass Spectrometry. Anal. Chem. 2001, 73, 2594–2604. [Google Scholar] [CrossRef]
- Doellinger, J.; Schaade, L.; Nitsche, A. Comparison of the Cowpox Virus and Vaccinia Virus Mature Virion Proteome: Analysis of the Species- and Strain-Specific Proteome. PLoS ONE 2015, 10, e0141527. [Google Scholar] [CrossRef]
- Käll, L.; Storey, J.D.; MacCoss, M.J.; Noble, W.S. Assigning Significance to Peptides Identified by Tandem Mass Spectrometry Using Decoy Databases. J. Proteome Res. 2008, 7, 29–34. [Google Scholar] [CrossRef]
- Liska, A.J.; Shevchenko, A. Expanding the Organismal Scope of Proteomics: Cross-Species Protein Identification by Mass Spectrometry and Its Implications. Proteomics 2003, 3, 19–28. [Google Scholar] [CrossRef] [PubMed]
- O’Bryon, I.; Jenson, S.C.; Merkley, E.D. Flying Blind, or Just Flying under the Radar? The Underappreciated Power of de Novo Methods of Mass Spectrometric Peptide Identification. Protein Sci. 2020, 29, 1864–1878. [Google Scholar] [CrossRef]
- Yan, Y.; Kusalik, A.J.; Wu, F.-X. NovoHCD: De Novo Peptide Sequencing from HCD Spectra. IEEE Trans. NanoBioscience 2014, 13, 65–72. [Google Scholar] [CrossRef]
- Kim, H.J.; Bae, I.Y.; Ahn, C.-W.; Lee, S.; Lee, H.G. Purification and Identification of Adipogenesis Inhibitory Peptide from Black Soybean Protein Hydrolysate. Peptides 2007, 28, 2098–2103. [Google Scholar] [CrossRef]
- Degenkolb, T.; Brückner, H. Peptaibiomics: Towards a Myriad of Bioactive Peptides Containing Cα-Dialkylamino Acids? Chem. Biodivers. 2008, 5, 1817–1843. [Google Scholar] [CrossRef] [PubMed]
- Yingchutrakul, Y.; Krobthong, S.; Choowongkomon, K.; Papan, P.; Samutrtai, P.; Mahatnirunkul, T.; Chomtong, T.; Srimongkolpithak, N.; Jaroenchuensiri, T.; Aonbangkhen, C. Discovery of a Multifunctional Octapeptide from Lingzhi with Antioxidant and Tyrosinase Inhibitory Activity. Pharmaceuticals 2022, 15, 684. [Google Scholar] [CrossRef]
- Shih, Y.-H.; Chen, F.-A.; Wang, L.-F.; Hsu, J.-L. Discovery and Study of Novel Antihypertensive Peptides Derived from Cassia Obtusifolia Seeds. J. Agric. Food Chem. 2019, 67, 7810–7820. [Google Scholar] [CrossRef] [PubMed]
- Jiang, S.; Dong, P.-P.; Li, H.-R.; Xu, J.; Li, H.-J.; Yu, Y.-Y. Study on lipid-lowering mechanism of active peptide DP17 from Eupolyphaga steleophaga in hyperlipidemia rats. China J. Chin. Mater. Med. 2020, 45, 5265–5272. [Google Scholar] [CrossRef]
- Johnson, R.S.; Davis, M.T.; Taylor, J.A.; Patterson, S.D. Informatics for Protein Identification by Mass Spectrometry. Methods 2005, 35, 223–236. [Google Scholar] [CrossRef]
- Babini, E.; Tagliazucchi, D.; Martini, S.; Dei Più, L.; Gianotti, A. LC-ESI-QTOF-MS Identification of Novel Antioxidant Peptides Obtained by Enzymatic and Microbial Hydrolysis of Vegetable Proteins. Food Chem. 2017, 228, 186–196. [Google Scholar] [CrossRef]
- Frank, A.; Tanner, S.; Bafna, V.; Pevzner, P. Peptide Sequence Tags for Fast Database Search in Mass-Spectrometry. J. Proteome Res. 2005, 4, 1287–1295. [Google Scholar] [CrossRef] [PubMed]
- Muth, T.; Renard, B.Y. Evaluating de Novo Sequencing in Proteomics: Already an Accurate Alternative to Database-Driven Peptide Identification? Brief. Bioinform. 2018, 19, 954–970. [Google Scholar] [CrossRef] [PubMed]
- Devabhaktuni, A.; Lin, S.; Zhang, L.; Swaminathan, K.; Gonzalez, C.G.; Olsson, N.; Pearlman, S.M.; Rawson, K.; Elias, J.E. TagGraph Reveals Vast Protein Modification Landscapes from Large Tandem Mass Spectrometry Datasets. Nat. Biotechnol. 2019, 37, 469–479. [Google Scholar] [CrossRef]
- Li, X.-C.; Yan, L.-H.; Wang, Z.-M.; Zhang, Q.-W.; Gao, J.-P.; Chen, L.-M. Separation and mass spectrometric analysis of antioxidant peptides from enzymatic hydrolysates of donkey-hide gelatin. China J. Chin. Mater. Med. 2013, 38, 1021–1025. [Google Scholar]
- Lu, Y. Purification, Structure Identification and Properties of ACE Inhibitory Peptide from Walnut Protein. Ph.D. Thesis, Shaanxi University of Science and Technology, Xi’an, China, 2015. [Google Scholar]
- Ma, B.; Johnson, R. De Novo Sequencing and Homology Searching. Mol. Cell Proteomics. 2012, 11, O111.014902. [Google Scholar] [CrossRef]
- He, H.; Sun, J.; Xie, B.-J.; Shi, Y.-L. Studied on the Ganoderan Lucidum peptides’antioxidation capacities in lipid system. Food Res. Dev. 2006, 27, 8–10. [Google Scholar] [CrossRef]
- Zhu, N.; Xu, M.-H.; Li, Y. Bioactive Oligopeptides from Ginseng (Panax Ginseng Meyer) Suppress Oxidative Stress-Induced Senescence in Fibroblasts via NAD+/SIRT1/PGC-1α Signaling Pathway. Nutrients 2022, 14, 5289. [Google Scholar] [CrossRef]
- Ren, J.-W.; Li, D.; Chen, Q.-H.; Liu, R.; Sun, J.-Q.; Li, H. Antioxidant Effect of Oligopeptide Extracted from Panax ginseng Grown in Jilin Province. Food Sci. 2018, 24, 54–57. [Google Scholar]
- Jiang, H.; Le, G.-W.; Yin, X.-P.; Gao, X.-L.; Shi, H. Effect of Peptides Hydrolyzed from Cervus elaphus Velvet Antler Blood on the Immunity and Antioxidation of Mice. Nat. Prod. Res. Dev. 2009, 21, 757–760. [Google Scholar] [CrossRef]
- Ma, F.-F.; Wang, H.; Wei, C.-K.; Thakur, K.; Wei, Z.-J.; Jiang, L. Three Novel ACE Inhibitory Peptides Isolated From Ginkgo Biloba Seeds: Purification, Inhibitory Kinetic and Mechanism. Front. Pharmacol. 2019, 9, 1579. [Google Scholar] [CrossRef]
- González-Montoya, M.; Hernández-Ledesma, B.; Silván, J.M.; Mora-Escobedo, R.; Martínez-Villaluenga, C. Peptides Derived from In Vitro Gastrointestinal Digestion of Germinated Soybean Proteins Inhibit Human Colon Cancer Cells Proliferation and Inflammation. Food Chem. 2018, 242, 75–82. [Google Scholar] [CrossRef]
- Gao, C.; Liu, J.; Zhao, C.-X. Ginseng Oligopeptides against Benzopyrene-Induced Inflammatory Injury of Lung Epithelial Cells. Sci. Technol. Food Ind. 2021, 42, 336–343. [Google Scholar] [CrossRef]
- Zhao, L.; Zhang, Y.-L.; Zhang, X.-L.; Xie, Q.-F.; Wang, C.-T. Anti-inflammatory and Intestinal Absorption Mechanism of Peptides Hydrolyzed from Velvet Antler Protein. J. Chin. Inst. Food Sci. Technol. 2018, 18, 45–51. [Google Scholar] [CrossRef]
- Itokawa, H.; Takeya, K.; Mori, N.; Sonobe, T.; Mihashi, S.; Hamanaka, T. Studies on Antitumor Cyclic Hexapeptides RA Obtained from Rubiae Radix, Rubiaceae. VI Minor Antitumor Constituents. Chem. Pharm. Bull. 1986, 34, 3762–3768. [Google Scholar] [CrossRef]
- Zhang, X.; Bi, Q.; Wu, X.; Wang, Z.; Miao, Y.; Tan, N. Systematic characterization and quantification of Rubiaceae-type cyclopeptides in 20 Rubia species by ultra performance liquid chromatography tandem mass spectrometry combined with chemometrics. J. Chromatogr. A 2018, 1581–1582, 43–54. [Google Scholar] [CrossRef]
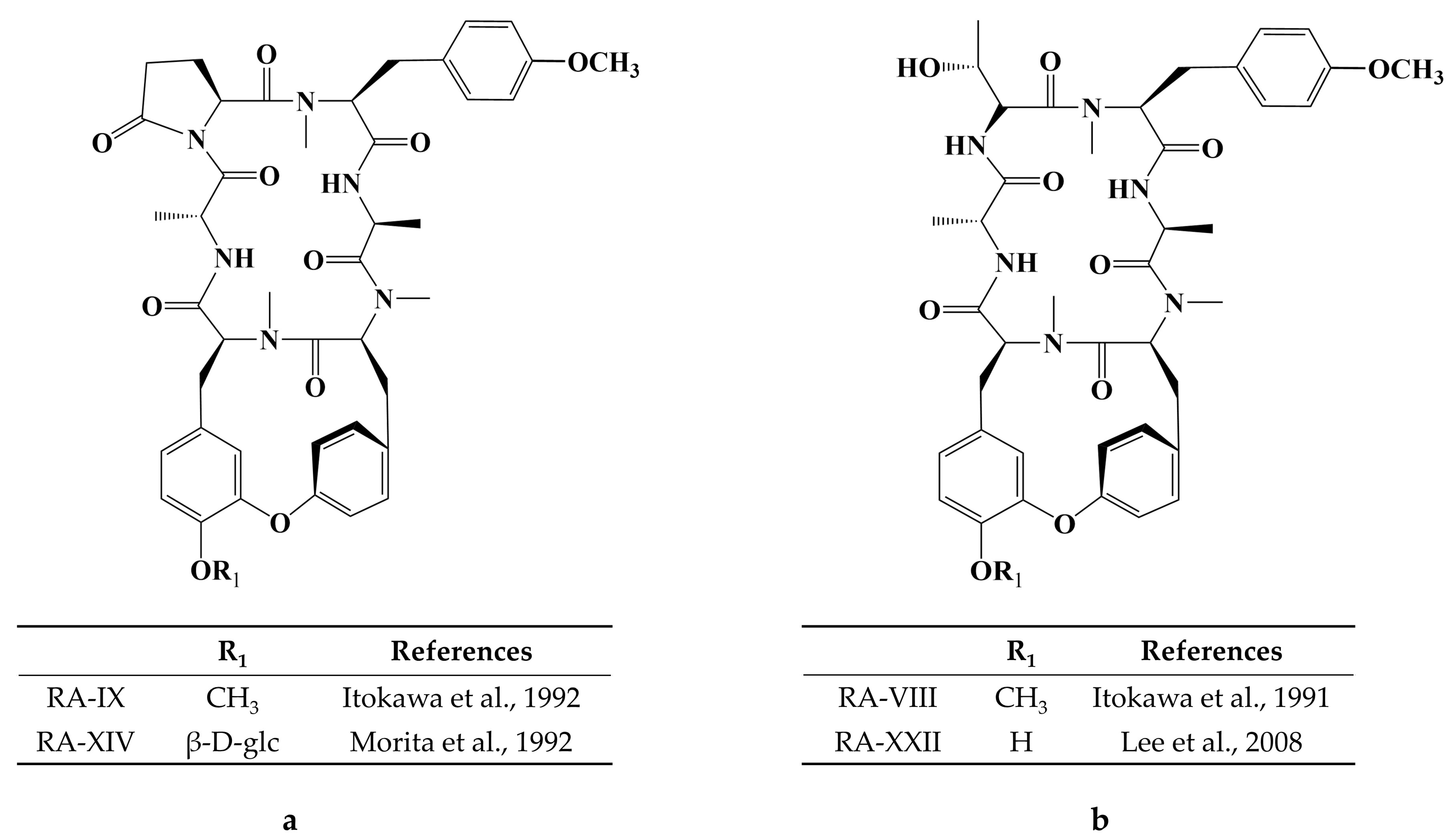
- Itokawa, H.; Morita, H.; Takeya, K.; Tomioka, N.; Itai, A.; Iitaka, Y. New antitumor bicyclic hexapeptides, RA-VI and -VIII from rubia cordifolia; Conformation-activity relationship II. Tetrahedron. 1991, 47, 7007–7020. [Google Scholar] [CrossRef]
- Itokawa, H.; Yamamiya, T.; Morita, H.; Takeya, K. New Antitumour Bicyclic Hexapeptides, RA-IX and -X from Rubia cordifolia. Part 3. Conformation-Antitumour Activity Relationship. J. Chem. Soc. Perkin Trans. 1992, 1, 455–459. [Google Scholar] [CrossRef]
- Morita, H.; Yamamiya, T.; Takeya, K.; Itokawa, H. New Antitumor Bicyclic Hexapeptides, RA-XI, -XII, -XIII and -XIV from Rubia Cordifolia. Chem. Pharm. Bull. 1992, 40, 1352–1354. [Google Scholar] [CrossRef]
- Takeya, K.; Yamamiya, T.; Morita, H.; Itokawa, H. Two Antitumour Bicyclic Hexapeptides from Rubia cordifolia. Phytochemistry 1993, 33, 613–615. [Google Scholar] [CrossRef]
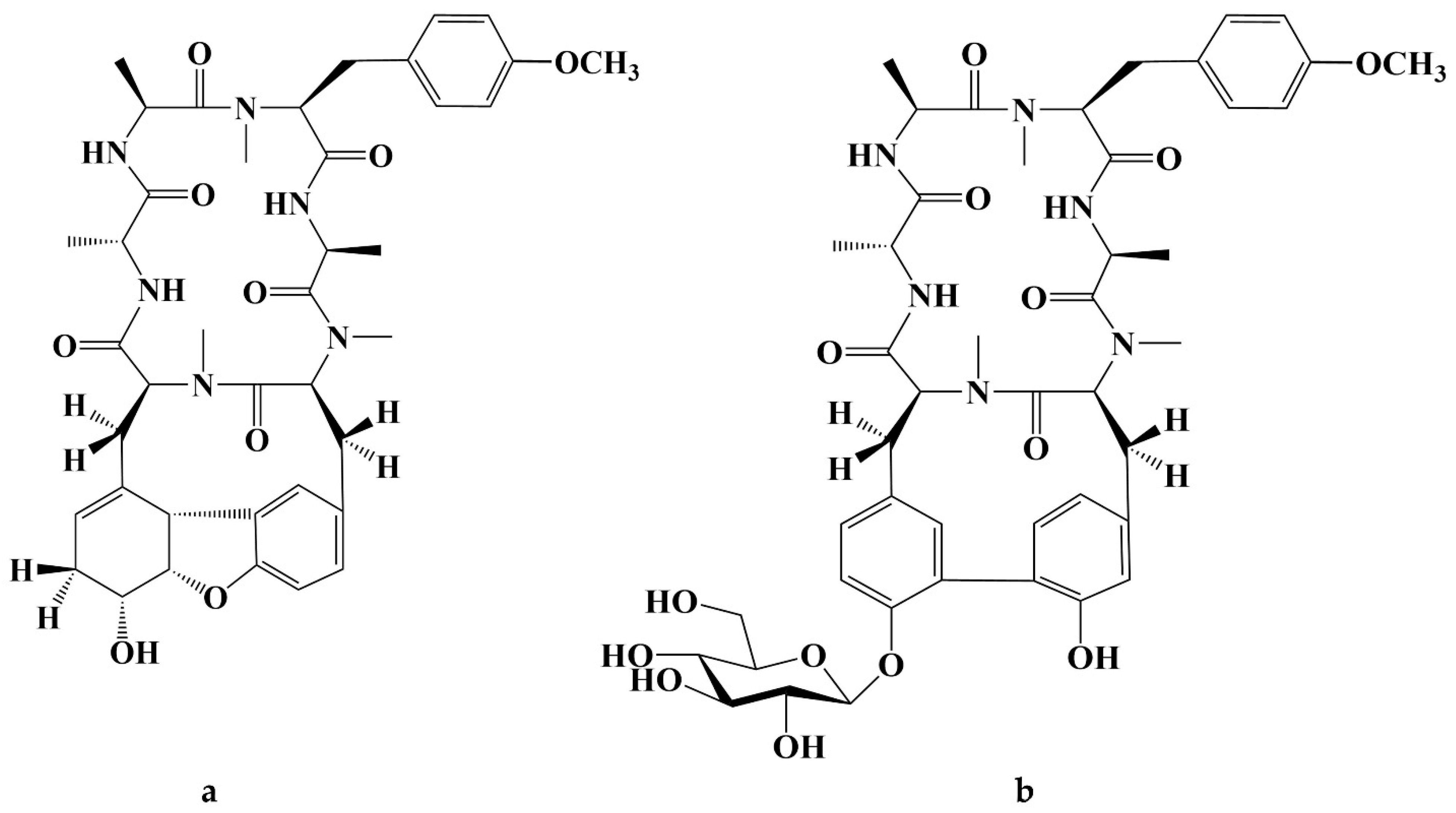
- Hitotsuyanagi, Y.; Ishikawa, H.; Hasuda, T.; Takeya, K. Isolation, structural elucidation, and synthesis of RA-XVII, a novel bicyclic hexapeptide from Rubia cordifolia, and the effect of side chain at residue 1 upon the conformation and cytotoxic activity. Tetrahedron Lett. 2004, 45, 935–938. [Google Scholar] [CrossRef]
- Lee, J.-E.; Hitotsuyanagi, Y.; Kim, I.-H.; Hasuda, T.; Takeya, K. A novel bicyclic hexapeptide, RA-XVIII, from Rubia cordifolia: Structure, semi-synthesis, and cytotoxicity. Bioorganic Med. Chem. Lett. 2008, 18, 808–811. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.-E.; Hitotsuyanagi, Y.; Takeya, K. Structures of cytotoxic bicyclic hexapeptides, RA-XIX, -XX, -XXI, and -XXII, from Rubia cordifolia L. Tetrahedron 2008, 64, 4117–4125. [Google Scholar] [CrossRef]
- Lee, J.-E.; Hitotsuyanagi, Y.; Fukaya, H.; Kondo, K.; Takeya, K. New Cytotoxic Bicyclic Hexapeptides, RA-XXIII and RA-XXIV, from Rubia cordifolia L. Chem. Pharm. Bull. 2008, 56, 730–733. [Google Scholar] [CrossRef]
- Hitotsuyanagi, Y.; Hirai, M.; Odagiri, M.; Komine, M.; Hasuda, T.; Fukaya, H.; Takeya, K. RA-XXV and RA-XXVI, Bicyclic Hexapeptides from Rubia cordifolia L. Structure, Synthesis, and Conformation. Chem. Asian J. 2019, 14, 205–215. [Google Scholar] [CrossRef] [PubMed]
- Fan, J.T.; Chen, Y.S.; Xu, W.Y.; Du, L.C.; Tan, N.H. Rubiyunnanins A and B, Two Novel Cyclic Hexapeptides from Rubia yunnanensis. Tetrahedron Lett. 2010, 51, 6810–6813. [Google Scholar] [CrossRef]
- Fan, J.-T.; Su, J.; Peng, Y.-M.; Li, Y.; Li, J.; Zhou, Y.-B.; Zeng, G.-Z.; Yan, H.; Tan, N.-H. Rubiyunnanins C–H, cytotoxic cyclic hexapeptides from Rubia yunnanensis inhibiting nitric oxide production and NF-κB activation. Bioorganic Med. Chem. Lett. 2010, 18, 8226–8234. [Google Scholar] [CrossRef]
- Xue, J.-W.; Wang, P.; Luo, X.-H.; Wu, S.-P.; Zhu, Q.-N.; He, Z.-P. Optimized Preparation of Hypoglycemic Peptides from Torreya grandis using Response Surface Methodology. J. Chin. Cereals Oils Assoc. 2022, 1–13. [Google Scholar] [CrossRef]
- Wang, B.-X.; Yang, M.; Jin, Y.-L.; Liu, P. Study on the hypoglycemic mechanism of ginseng peptides. Acta Pharm. Sin. B 1990, 25, 401–405. [Google Scholar] [CrossRef]
- Jiang, N.; Zhang, S.; Zhu, J.; Shang, J.; Gao, X. Hypoglycemic, Hypolipidemic and Antioxidant Effects of Peptides from Red Deer Antlers in Streptozotocin-Induced Diabetic Mice. Tohoku J. Exp. Med. 2015, 236, 71–79. [Google Scholar] [CrossRef]
- Zanoni, C.; Aiello, G.; Arnoldi, A.; Lammi, C. Hempseed Peptides Exert Hypocholesterolemic Effects with a Statin-Like Mechanism. J. Agric. Food Chem. 2017, 65, 8829–8838. [Google Scholar] [CrossRef]
- Wang, N.; Tong, Z.; Wang, D.; Zhang, Y.; Liu, T. Effects of Hericium Erinaceus Polypeptide on Lowering Blood Lipids of Mice with Hyperlipidemia Induced by a High-Fat Diet. J. Future Foods 2022, 2, 346–357. [Google Scholar] [CrossRef]
- Li, D.; Liu, R.; Li, H.; Chen, Q.-H.; Zhang, Z.-F.; Wang, J.-B. Hypolipidemic Effect of Oligopeptide Derived from Panax ginseng Grown in Jilin Province on Rats with Hyperlipidemia. Food Sci. 2017, 38, 227–232. [Google Scholar]
- Zhao, H.-L.; Deng, S.-H.; Hao, T.; Yin, M.-M.; Yin, Z.-F.; Wang, L.-Y. In vitro Antibacterial activity of different enzymolysis extracts from eupolyphaga sinesis walker. Chem. Bioeng. 2017, 34, 59–62. [Google Scholar]
- Bao, L.; Cai, X.; Wang, J.; Zhang, Y.; Sun, B.; Li, Y. Anti-Fatigue Effects of Small Molecule Oligopeptides Isolated from Panax ginseng C. A. Meyer in Mice. Nutrients 2016, 8, 807. [Google Scholar] [CrossRef] [PubMed]
- He, L.-X.; Ren, J.-W.; Liu, R.; Chen, Q.-H.; Zhao, J.; Wu, X.; Zhang, Z.-F.; Wang, J.-B.; Pettinato, G.; Li, Y. Ginseng (Panax ginseng Meyer) Oligopeptides Regulate Innate and Adaptive Immune Responses in Mice via Increased Macrophage Phagocytosis Capacity, NK Cell Activity and Th Cells Secretion. Food Funct. 2017, 8, 3523–3532. [Google Scholar] [CrossRef] [PubMed]
- Liang, R.; Fan, C.; Li, Y.; Zeng, Q.-H.; Liu, R.; Guo, X.-F. Immunomodulatory Effects of Low Molecular Weight Peptide of Colla Corii Asini. Sci. Technol. Food Ind. 2019, 40, 306–310+315. [Google Scholar] [CrossRef]
- Fan, N.-J.; Wang, D.-M.; Gao, Y.; Guo, Y.-N.; Cheng, T.-C.; Fang, Q.-X. Immunomodulatory effects of the iron yam peptides on mice with immunocompromised. Food Ferment. Ind. 2020, 46, 101–107. [Google Scholar] [CrossRef]
- Fan, R.; Hao, Y.-T.; Liu, X.-R.; Li, Y. Effects of Ginseng Oligopeptide on Blood Glucose and Immune regulation in the Aging db/db Mice. Sci. Technol. Food Ind. 2020, 41, 297–301+310. [Google Scholar] [CrossRef]
- He, L.-X.; Liu, R.; Ren, J.-W.; Chen, Q.-H.; Wang, J.-B.; Zhang, Z.-F. Immunomodulatory effects of Panax ginseng oligopeptide of Jilin Province. Sci. Technol. Rev. 2015, 33, 62–67. [Google Scholar]
- Liu, Y.; Li, D.; Wei, Y.; Ma, Y.; Wang, Y.; Huang, L.; Wang, Y. Hydrolyzed Peptides from Purple Perilla (Perilla Frutescens L. Britt.) Seeds Improve Muscle Synthesis and Exercise Performance in Mice. J. Food Biochem. 2020, 44, 13461. [Google Scholar] [CrossRef]
- Astafieva, A.A.; Rogozhin, E.A.; Andreev, Y.A.; Odintsova, T.I.; Kozlov, S.A.; Grishin, E.V.; Egorov, T.A. A Novel Cysteine-Rich Antifungal Peptide ToAMP4 from Taraxacum Officinale Wigg. Flowers. Plant Physiol. Biochem. 2013, 70, 93–99. [Google Scholar] [CrossRef]
- Mandal, S.M.; Migliolo, L.; Franco, O.L.; Ghosh, A.K. Identification of an Antifungal Peptide from Trapa Natans Fruits with Inhibitory Effects on Candida Tropicalis Biofilm Formation. Peptides 2011, 32, 1741–1747. [Google Scholar] [CrossRef] [PubMed]
- Khani, S.; Seyedjavadi, S.S.; Zare-Zardini, H.; Hosseini, H.M.; Goudarzi, M.; Khatami, S.; Amani, J.; Imani Fooladi, A.A.; Razzaghi-Abyaneh, M. Isolation and Functional Characterization of an Antifungal Hydrophilic Peptide, Skh-AMP1, Derived from Satureja khuzistanica Leaves. Phytochemistry 2019, 164, 136–143. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.-M.; Yang, M.; Zhou, S.-W.; Zhang, H.; Feng, Y.; Shi, L.; Li, D.-S.; Lu, Q.-M.; Zhang, Z.-H.; Zhao, M. Blapstin, a Diapause-Specific Peptide-Like Peptide from the Chinese Medicinal Beetle Blaps rhynchopetera, Has Antifungal Function. Microbiol. Spectr. 2023, 11, e03089-22. [Google Scholar] [CrossRef] [PubMed]
- Mirzaei, M.; Mirdamadi, S.; Ehsani, M.R.; Aminlari, M.; Hosseini, E. Purification and Identification of Antioxidant and ACE-Inhibitory Peptide from Saccharomyces cerevisiae Protein Hydrolysate. J. Funct. Foods 2015, 19, 259–268. [Google Scholar] [CrossRef]
- Elias, R.J.; Kellerby, S.S.; Decker, E.A. Antioxidant Activity of Proteins and Peptides. Crit. Rev. Food Sci. Nutr. 2008, 48, 430–441. [Google Scholar] [CrossRef]
- Halliwell, B. Free Radicals, Antioxidants, and Human Disease: Curiosity, Cause, or Consequence? Lancet 1994, 344, 721–724. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Zheng, Z.; Ai, Z.; Zhang, Y.; Tan, C.P.; Liu, Y. Exploring the Antioxidant and Structural Properties of Black Bean Protein Hydrolysate and Its Peptide Fractions. Front. Nutr. 2022, 9, 884537. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhou, F.-Y.; Pan, L.-M.; He, P.; Zhang, Y.-Z. Identification and structure-activity relationship of antioxidant peptides derived from earthworm protein. Mod. Food Sci. Technol. 2023, 39, 164–173. [Google Scholar] [CrossRef]
- He, L.-S.; Lin, M.-N.; Jiang, B.-B.; Peng, M.-S.; Zhang, W.-Y.; Han, J.-Q. Study on four antioxidant activities of endogenous peptides from Isatis indigotica in vitro. J. Shaanxi Univ. Chin. Med. 2022, 45, 55–58. [Google Scholar] [CrossRef]
- Kumar, N.S.S.; Nazeer, R.A.; Jaiganesh, R. Purification and Identification of Antioxidant Peptides from the Skin Protein Hydrolysate of Two Marine Fishes, Horse Mackerel (Magalaspis cordyla) and Croaker (Otolithes ruber). Amino Acids 2012, 42, 1641–1649. [Google Scholar] [CrossRef]
- Qian, Z.-J.; Jung, W.-K.; Byun, H.-G.; Kim, S.-K. Protective Effect of an Antioxidative Peptide Purified from Gastrointestinal Digests of Oyster, Crassostrea Gigas against Free Radical Induced DNA Damage. Bioresour. Technol. 2008, 99, 3365–3371. [Google Scholar] [CrossRef]
- Rajapakse, N.; Mendis, E.; Byun, H.-G.; Kim, S.-K. Purification and In Vitro Antioxidative Effects of Giant Squid Muscle Peptides on Free Radical-Mediated Oxidative Systems. J. Nutr. Biochem. 2005, 16, 562–569. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.J. Study on antioxidant activity of antler polypeptides with different molecular weights. Sci. Technol. Food Ind. 2015, 36, 53–56. [Google Scholar] [CrossRef]
- Li, Y.; Yu, J. Research Progress in Structure-Activity Relationship of Bioactive Peptides. J. Med. Food. 2015, 18, 147–156. [Google Scholar] [CrossRef] [PubMed]
- Cheison, S.C.; Wang, Z.; Xu, S.-Y. Preparation of Whey Protein Hydrolysates Using a Single- and Two-Stage Enzymatic Membrane Reactor and Their Immunological and Antioxidant Properties: Characterization by Multivariate Data Analysis. J. Agric. Food Chem. 2007, 55, 3896–3904. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.-C.; Xu, X.; Wang, L. Research Progress in Enzymatic Hydrolysis of Protein to Prepare Antioxidant Peptides. Cereals Oils 2008, 7, 1–2. [Google Scholar]
- Boschin, G.; Scigliuolo, G.M.; Resta, D.; Arnoldi, A. ACE-Inhibitory Activity of Enzymatic Protein Hydrolysates from Lupin and Other Legumes. Food Chem. 2014, 145, 34–40. [Google Scholar] [CrossRef]
- Shobako, N. Hypotensive Peptides Derived from Plant Proteins. Peptides 2021, 142, 170573. [Google Scholar] [CrossRef]
- Lee, S.Y.; Hur, S.J. Antihypertensive Peptides from Animal Products, Marine Organisms, and Plants. Food Chem. 2017, 228, 506–517. [Google Scholar] [CrossRef]
- Shi, A.; Liu, H.; Liu, L.; Hu, H.; Wang, Q.; Adhikari, B. Isolation, Purification and Molecular Mechanism of a Peanut Protein-Derived ACE-Inhibitory Peptide. PLoS ONE 2014, 9, e111188. [Google Scholar] [CrossRef]
- Natesh, R.; Schwager, S.L.U.; Sturrock, E.D.; Acharya, K.R. Crystal Structure of the Human Angiotensin-Converting Enzyme-Lisinopril Complex. Nature 2003, 421, 551–554. [Google Scholar] [CrossRef]
- Jia, J.Q.; Ma, H.L.; Wang, Z.B.; Luo, L.; He, R.H. Structure-activity relationship of antihypertensive peptides. J. Chin. Cereals Oils Assoc. 2009, 24, 110–114. [Google Scholar]
- Guha, S.; Majumder, K. Structural-Features of Food-Derived Bioactive Peptides with Anti-Inflammatory Activity: A Brief Review. J. Food Biochem. 2019, 43, e12531. [Google Scholar] [CrossRef]
- Chakrabarti, S.; Jahandideh, F.; Wu, J. Food-Derived Bioactive Peptides on Inflammation and Oxidative Stress. Biomed. Res. Int. 2014, 2014, 608979. [Google Scholar] [CrossRef]
- Lau, J.L.; Dunn, M.K. Therapeutic Peptides: Historical Perspectives, Current Development Trends, and Future Directions. Bioorganic Med. Chem. 2018, 26, 2700–2707. [Google Scholar] [CrossRef]
- Anderson, P.; Delgado, M. Endogenous Anti-Inflammatory Neuropeptides and pro-Resolving Lipid Mediators: A New Therapeutic Approach for Immune Disorders. J. Cell. Mol. Med. 2008, 12, 1830–1847. [Google Scholar] [CrossRef]
- Harrison, D.A. The Jak/STAT Pathway. Cold Spring Harb. Perspect. Biol. 2012, 4, a011205. [Google Scholar] [CrossRef]
- Haddad, J.J. The Role of Inflammatory Cytokines and NF-κB/MAPK Signaling Pathways in the Evolution of Familial Mediterranean Fever: Current Clinical Perspectives and Potential Therapeutic Approaches. Cell. Immunol. 2009, 260, 6–13. [Google Scholar] [CrossRef]
- Roberts, P.R.; Burney, J.D.; Black, K.W.; Zaloga, G.P. Effect of Chain Length on Absorption of Biologically Active Peptides from the Gastrointestinal Tract. Digestion 1999, 60, 332–337. [Google Scholar] [CrossRef]
- Vogel, H.J.; Schibli, D.J.; Jing, W.; Lohmeier-Vogel, E.M.; Epand, R.F.; Epand, R.M. Towards a Structure-Function Analysis of Bovine Lactoferricin and Related Tryptophan- and Arginine-Containing Peptides. Biochem. Cell Biol. 2002, 80, 49–63. [Google Scholar] [CrossRef] [PubMed]
- Ichikawa, S.; Morifuji, M.; Ohara, H.; Matsumoto, H.; Takeuchi, Y.; Sato, K. Hydroxyproline-Containing Dipeptides and Tripeptides Quantified at High Concentration in Human Blood after Oral Administration of Gelatin Hydrolysate. Int. J. Food Sci. Nutr. 2010, 61, 52–60. [Google Scholar] [CrossRef]
- Peer, D.; Karp, J.M.; Hong, S.; Farokhzad, O.C.; Margalit, R.; Langer, R. Nanocarriers as an Emerging Platform for Cancer Therapy. Nat. Nanotechnol. 2007, 2, 751–760. [Google Scholar] [CrossRef]
- Kang, T.H.; Mao, C.-P.; He, L.; Tsai, Y.-C.; Liu, K.; La, V.; Wu, T.-C.; Hung, C.-F. Tumor-Targeted Delivery of IL-2 by NKG2D Leads to Accumulation of Antigen-Specific CD8+ T Cells in the Tumor Loci and Enhanced Anti-Tumor Effects. PLoS ONE 2012, 7, e35141. [Google Scholar] [CrossRef]
- Mulder, K.C.L.; Lima, L.A.; Miranda, V.J.; Dias, S.C.; Franco, O.L. Current Scenario of Peptide-Based Drugs: The Key Roles of Cationic Antitumor and Antiviral Peptides. Front. Microbiol. 2013, 4, 321. [Google Scholar] [CrossRef]
- Call, J.A.; Eckhardt, S.G.; Camidge, D.R. Targeted Manipulation of Apoptosis in Cancer Treatment. Lancet Oncol. 2008, 9, 1002–1011. [Google Scholar] [CrossRef]
- Scott, A.M.; Wolchok, J.D.; Old, L.J. Antibody Therapy of Cancer. Nat. Rev. Cancer 2012, 12, 278–287. [Google Scholar] [CrossRef] [PubMed]
- He, C.; Tang, Z.; Tian, H.; Chen, X. Co-Delivery of Chemotherapeutics and Proteins for Synergistic Therapy. Adv. Drug Deliv. Rev. 2016, 98, 64–76. [Google Scholar] [CrossRef]
- Cicero, A.F.G.; Fogacci, F.; Colletti, A. Potential Role of Bioactive Peptides in Prevention and Treatment of Chronic Diseases: A Narrative Review. Br. J. Pharmacol. 2017, 174, 1378–1394. [Google Scholar] [CrossRef] [PubMed]
- Chiangjong, W.; Chutipongtanate, S.; Hongeng, S. Anticancer Peptide: Physicochemical Property, Functional Aspect and Trend in Clinical Application (Review). Int. J. Oncol. 2020, 57, 678–696. [Google Scholar] [CrossRef] [PubMed]
- Schweizer, F. Cationic Amphiphilic Peptides with Cancer-Selective Toxicity. Eur. J. Pharmacol. 2009, 625, 190–194. [Google Scholar] [CrossRef] [PubMed]
- Lohner, K. New Strategies for Novel Antibiotics: Peptides Targeting Bacterial Cell Membranes. Gen. Physiol. Biophys. 2009, 28, 105–116. [Google Scholar] [CrossRef]
- Huang, Y.-B.; Wang, X.-F.; Wang, H.-Y.; Liu, Y.; Chen, Y. Studies on Mechanism of Action of Anticancer Peptides by Modulation of Hydrophobicity within a Defined Structural Framework. Mol. Cancer Ther. 2011, 10, 416–426. [Google Scholar] [CrossRef] [PubMed]
- Pace, C.N.; Scholtz, J.M. A Helix Propensity Scale Based on Experimental Studies of Peptides and Proteins. Biophys. J. 1998, 75, 422–427. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Liu, Z.; Liu, Y.; Su, Z.; Liu, Y. Plant Polypeptides: A Review on Extraction, Isolation, Bioactivities and Prospects. Int. J. Biol. Macromol. 2022, 207, 169–178. [Google Scholar] [CrossRef]
- Wang, L.; Gagey-Eilstein, N.; Broussy, S.; Reille-Seroussi, M.; Huguenot, F.; Vidal, M.; Liu, W.-Q. Design and Synthesis of C-Terminal Modified Cyclic Peptides as VEGFR1 Antagonists. Molecules 2014, 19, 15391–15407. [Google Scholar] [CrossRef]
- Gang, D.; Kim, D.W.; Park, H.-S. Cyclic Peptides: Promising Scaffolds for Biopharmaceuticals. Genes 2018, 9, 557. [Google Scholar] [CrossRef] [PubMed]
- Evans, B.J.; King, A.T.; Katsifis, A.; Matesic, L.; Jamie, J.F. Methods to Enhance the Metabolic Stability of Peptide-Based PET Radiopharmaceuticals. Molecules 2020, 25, 2314. [Google Scholar] [CrossRef]
- Itokawa, H.; Takeya, K.; Mori, N.; Takanashi, M.; Yamamoto, H.; Sonobe, T.; Kidokoro, S. Cell Growth-Inhibitory Effects of Derivatives of Antitumor Cyclic Hexapeptide RA-V Obtained from Rubiae radix (V). Gan 1984, 75, 929–936. [Google Scholar]
- Yue, G.G.; Fan, J.-T.; Lee, J.K.; Zeng, G.-Z.; Ho, T.W.; Fung, K.-P.; Leung, P.-C.; Tan, N.-H.; Lau, C.B. Cyclopeptide RA-V Inhibits Angiogenesis by down-Regulating ERK1/2 Phosphorylation in HUVEC and HMEC-1 Endothelial Cells: RA-V Inhibits Angiogenesis in Endothelial Cells. Br. J. Pharmacol. 2011, 164, 1883–1898. [Google Scholar] [CrossRef] [PubMed]
- Chow, H.Y.; Zhang, Y.; Matheson, E.; Li, X. Ligation Technologies for the Synthesis of Cyclic Peptides. Chem. Rev. 2019, 119, 9971–10001. [Google Scholar] [CrossRef] [PubMed]
- Craik, D.J.; Conibear, A.C. The Chemistry of Cyclotides. J. Org. Chem. 2011, 76, 4805–4817. [Google Scholar] [CrossRef] [PubMed]
- Sun, B.; Li, D.; Mao, R.-X.; Li, H.; Wang, J.-B.; Li, Y. Anti-diabetic Effects of Panax ginseng Oligopeptides in Jilin Province on Diabetes Mice. Biotechnol. Bull. 2016, 22, 62–65. [Google Scholar]
- Liu, R.; Chen, Q.-H.; Ren, J.-W.; Sun, B.; Cai, X.-X.; Li, D.; Mao, R.-X.; Wu, X.; Li, Y. Ginseng (Panax ginseng Meyer) Oligopeptides Protect against Binge Drinking-Induced Liver Damage through Inhibiting Oxidative Stress and Inflammation in Rats. Nutrients 2018, 10, 1665. [Google Scholar] [CrossRef]
- Xu, M.; Chen, Q.; Fan, R.; Wang, J.; Li, Y. Anti-Inflammation Effect of Small Molecule Oligopeptides Prepared from Panax ginseng C. A. Meyer in Rats. Molecules 2019, 24, 858. [Google Scholar] [CrossRef] [PubMed]
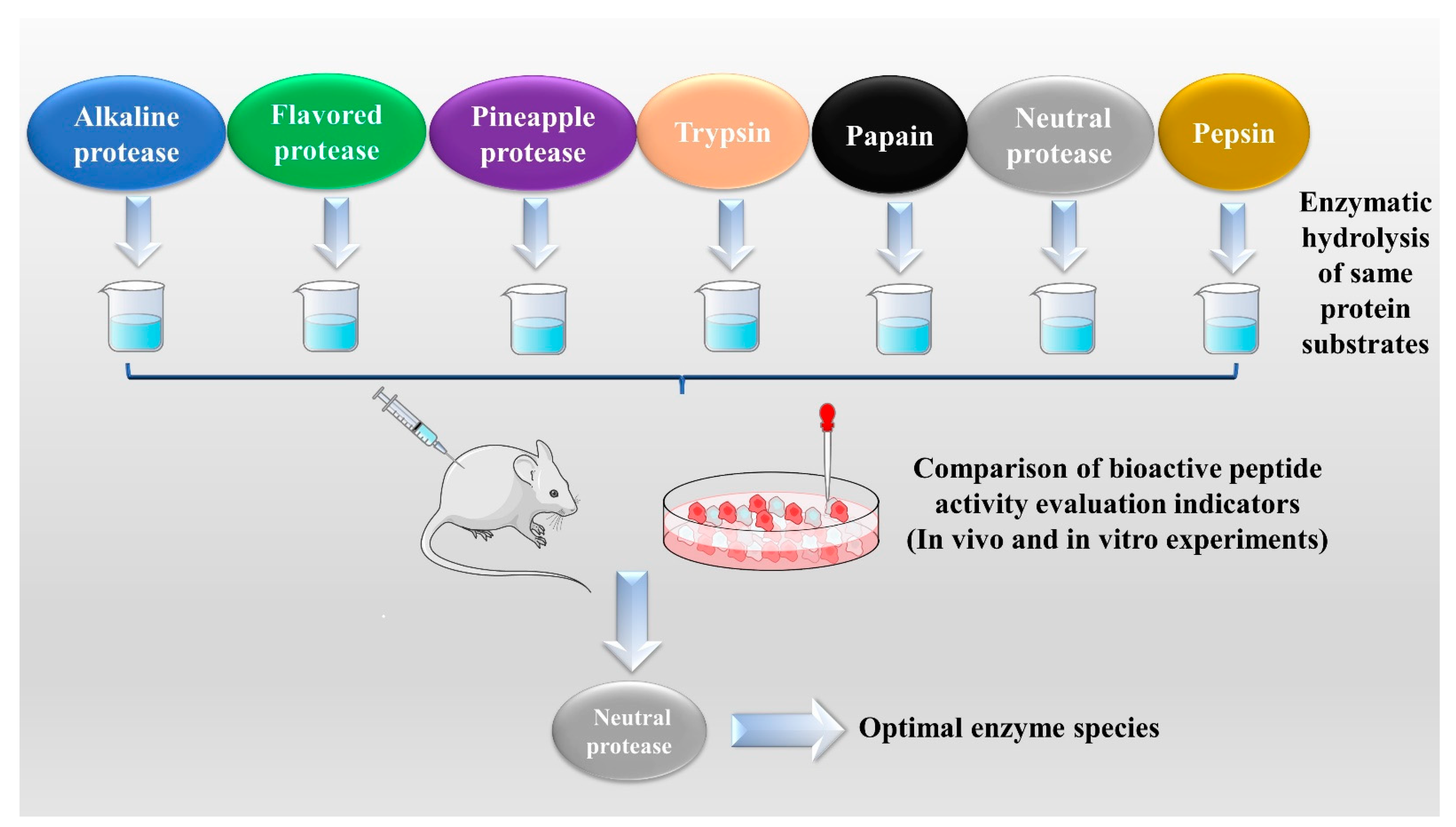



| Enzymatic Methods | Raw Materials | Types of Enzymes Used | Enzymatic Hydrolysis Conditions | Reference |
|---|---|---|---|---|
| Single EH | Ginkgo biloba seed | Alcalase | pH value: 8.0, temperature: 50 °C, time: 4.5 h, enzyme addition: 3500 U/g. | [17] |
| Alfalfa (Medicago sativa L.) Leaf | Papain | pH value: 7.5, temperature: 55 °C, time: 4 h, enzyme addition: 3:100 (enzyme/substrate w/w). | [18] | |
| Seahorse (Hippocampus) | Papain | pH value: 6.0, temperature: 60 °C, time: 40 min, enzyme addition: 2000 U/g. | [19] | |
| Sea cucumber (Actinopyga lecanora) | Alcalase | pH value: 8.0, temperature: 37 °C, time: 8 h, enzyme addition: 1:100 (enzyme/substrate w/w). | [20] | |
| Combinatorial EH | Zizyphus jujuba fruits | Trypsin, papain | pH value: 7.5, temperature: 37 °C, time: 4 h, enzyme addition: 1:50 (enzyme/substrate w/w). | [21] |
| Lycium barbarum | Neutrase, papain | pH value: 7.0, temperature: 51 °C, time: 4.3 h, neutral protease-papain addition rate: 1: 2.65. | [22] | |
| Successive EH | Red deer (Cervus elaphus) antler | Alcalase- Flavourzyme | Alcalase pH value: 8.0, temperature: 60 °C, time: 3 h, enzyme addition: 5000 U/g, substrate concentration: 12%. Flavourzyme pH value: 6.5, temperature: 45 °C, time: 1 h, enzyme addition: 6000 U/g, substrate concentration: 5%. | [23] |
| Black Soybean | Alcalase- Neutrase- Flavourzyme | Alcalase pH value: 8.5, temperature: 60 °C, time: 30 min. Neutrase pH value: 7.0, temperature: 60 °C, time: 60 min. Flavourzyme pH value: 6.0, temperature: 60 °C, time: 30 min. | [24] | |
| Hempseed (Cannabis sativa L.) | Alcalase- Neutrase | Alcalase pH value: 10.0, temperature: 50 °C, time: 4h, enzyme addition: 8000 U/g. Neutrase pH value: 7.0, temperature: 45 °C, time: 4h, enzyme addition: 8000 U/g. | [25] |
| Methods | Mechanism | Advantage | Disadvantage | Reference |
|---|---|---|---|---|
| Aqueous extraction | Polar similarity solubility (Polar peptides) | Mild reaction conditions (ensure peptide stability); Simple and convenient operation | Time-consuming; Low extraction efficiency; Introduction of water-soluble impurities | [31] |
| Organic solvent extraction | Polar similarity solubility (Peptides containing aromatic amino acids and peptides with many nonpolar side chains) | Higher extraction rate (compared to aqueous extraction); Reaction conditions can be controlled to obtain different peptides (e.g., solvent polarity, pH, etc.) | Destruction of essential amino acids such as serine, threonine, and tryptophan; Large solvent usage; Degradation or denaturation of peptides; Peptide toxicity due to organic solvent residues | [31,32,35,37] |
| Acid or alkali extraction | Disruption of disulfide, hydrogen, and peptide bonds increases peptide solubilization; Acidic and neutral amino acids undergo ionization to increase solubility in a high pH environment |
| Bioactive Peptides’s Functions | Bioactive Peptides Source | References |
|---|---|---|
| Antioxidant | Colla Corii Asini | [26,119] |
| Velvet Antlers | [74] | |
| Ganoderma lucidum | [75,111,122] | |
| Ginseng (Panax ginseng Meyer) | [123,124] | |
| Cervus elaphus velvet antlers | [125] | |
| Anti-hypertensive | Zizyphus Jujuba Fruit | [21] |
| Cassia Obtusifolia Seeds | [112] | |
| Ginkgo biloba seeds | [126] | |
| Anti-inflammatory | walnut | [88] |
| Soybean | [127] | |
| Ginseng | [128] | |
| Velvet Antler | [129] | |
| Lycium barbarum L. | [67] | |
| Anti-cancer | Colla Corii Asini | [99] |
| genus Rubia | [130,131,132,133,134,135,136,137,138,139,140,141,142] | |
| Hypoglycemic | Torreya grandis Merrillii | [143] |
| Ginseng | [144] | |
| Hypoglycemic Hypolipidemic | Red Deer Antlers | [145] |
| Hypolipidemic | Hempseed | [146] |
| Hericium erinaceus | [147] | |
| Eupolyphaga steleophaga | [113] | |
| Panax ginseng | [148] | |
| Antibacterial | Indian Ganoderma lucidum | [56] |
| eupolyphaga sinesis walker | [149] | |
| Antifatigue | Sea Horse (Hippocampus) | [19] |
| Panax ginseng C. A. Meyer | [150] | |
| Immunomodulatory | Ginseng (Panax ginseng Meyer) | [151] |
| Colla Corii Asini | [152] | |
| Iron yam | [153] | |
| Ginseng | [154] | |
| Panax ginseng | [155] | |
| Isatis indigotica | [30] | |
| Improve muscle synthesis exercise performance | Purple perilla (Perilla frutescens L. Britt.) seeds | [156] |
| Antifungal | Taraxacum officinale Wigg. flowers | [157] |
| Trapa natans fruits | [158] | |
| Satureja khuzistanica leaves | [159] | |
| Beetle Blaps rhynchopetera | [160] |
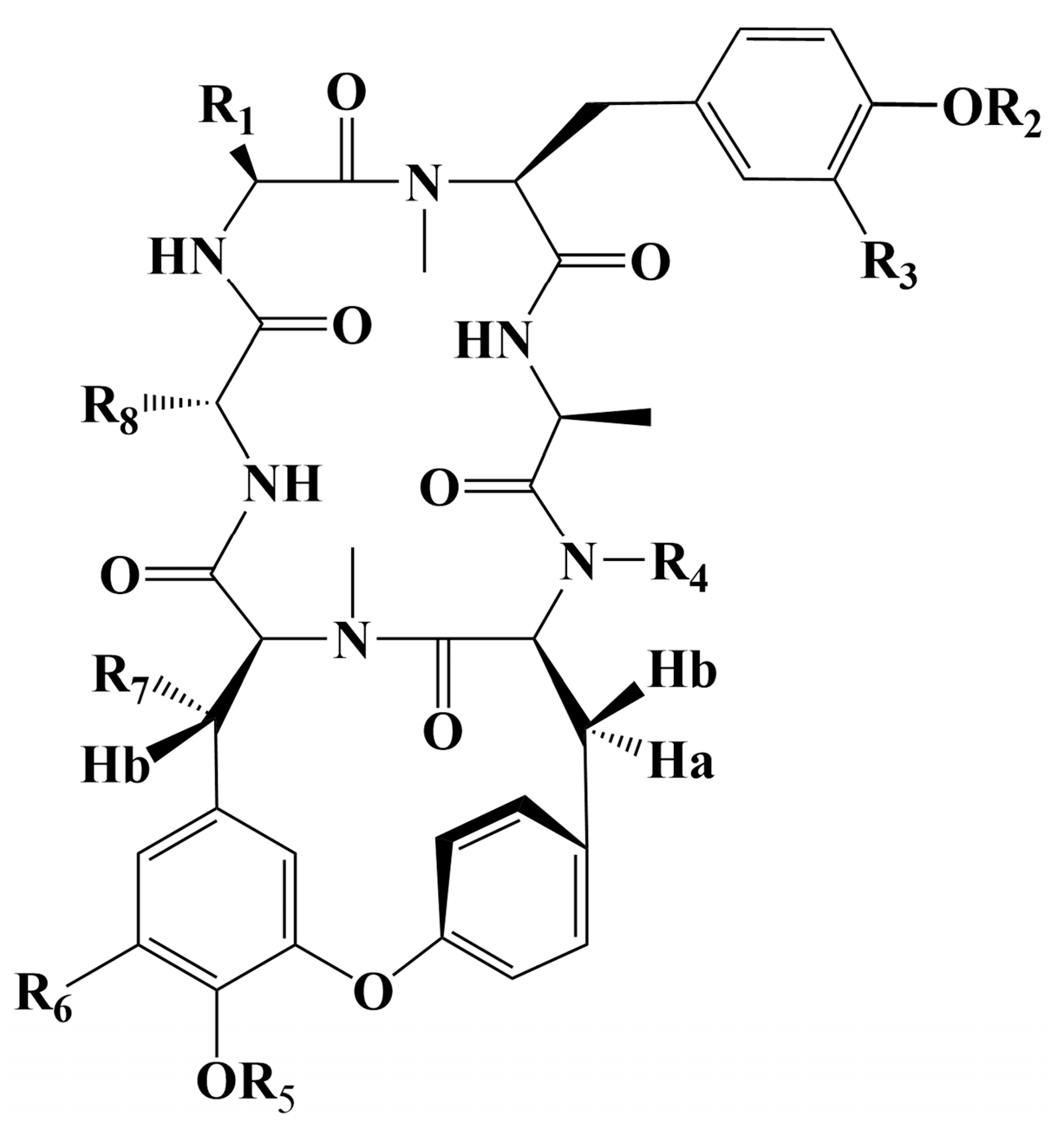
| R1 | R2 | R3 | R4 | R5 | R6 | R7 | R8 | References | |
|---|---|---|---|---|---|---|---|---|---|
| RA-I | CH2OH | CH3 | H | CH3 | H | H | H | CH3 | [130] |
| RA-II | CH3 | H | H | CH3 | CH3 | H | H | CH3 | [130] |
| RA-III | CH2OH | CH3 | H | CH3 | CH3 | H | H | CH3 | [130] |
| RA-IV | CH3 | CH3 | H | CH3 | CH3 | H | OH | CH3 | [130] |
| RA-V | CH3 | CH3 | H | CH3 | H | H | H | CH3 | [130] |
| RA-VII | CH3 | CH3 | H | CH3 | CH3 | H | H | CH3 | [131] |
| RA-X | CH2CH2COOH | CH3 | H | CH3 | CH3 | H | H | CH3 | [133] |
| RA-XI | CH2CH2COOH | CH3 | H | CH3 | H | H | H | CH3 | [134] |
| RA-XII | CH3 | CH3 | H | CH3 | β-d-glc | H | H | CH3 | [134] |
| RA-XIII | CH2CH2COOH | CH3 | H | CH3 | β-d-glc | H | H | CH3 | [134] |
| RA-XV | CH3 | CH3 | H | CH3 | β-d-glc-Ac | H | H | CH3 | [135] |
| RA-XVI | CH3 | CH3 | H | CH3 | β-d-glc | H | AcO | CH3 | [135] |
| RA-XVII | CH3 | CH3 | H | CH3 | H | H | H | CH2CH3 | [136] |
| RA-XVIII | CH3 | CH3 | H | CH3 | CH3 | OH | H | CH3 | [137] |
| RA-XIX | i-Pr | CH3 | H | CH3 | CH3 | H | H | CH3 | [138] |
| RA-XX | CH2CH3 | CH3 | H | CH3 | CH3 | H | H | CH3 | [138] |
| RA-XXI | CH2CH3 | CH3 | H | CH3 | H | H | H | CH3 | [138] |
| RA-XXIII | CH2CH2CONH2 | CH3 | H | CH3 | CH3 | H | H | CH3 | [139] |
| RA-XXVI | CH2CH2CONH2 | CH3 | H | CH3 | H | H | H | CH3 | [139] |
| RA-XXV | CH3 | CH3 | H | H | CH3 | H | H | CH3 | [140] |
| RA-XXVI | CH3 | CH3 | H | H | H | H | H | CH3 | [140] |
| Rubiyunnanin C | CH2CH2COOCH3 | CH3 | H | CH3 | H | H | H | CH3 | [142] |
| Rubiyunnanin D | CH2CH2COOH | H | H | CH3 | H | H | H | CH3 | [142] |
| Rubiyunnanin E | CH2CH2COOH | H | OH | H | H | H | H | CH3 | [142] |
| Rubiyunnanin F | CH2CH2CONH2 | CH3 | H | CH3 | β-d-glc | H | H | CH3 | [142] |
| Rubiyunnanin G | CH3 | H | H | CH3 | β-d-glc | H | H | CH3 | [142] |
| Rubiyunnanin H | CH3 | CH3 | H | CH3 | β-d-glc | H | H | CH3 | [142] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, Y.; Liu, L.; Zhang, M.; Li, S.; Wu, J.; Sun, Q.; Ma, S.; Cai, W. The Research Progress of Bioactive Peptides Derived from Traditional Natural Products in China. Molecules 2023, 28, 6421. https://doi.org/10.3390/molecules28176421
Zhang Y, Liu L, Zhang M, Li S, Wu J, Sun Q, Ma S, Cai W. The Research Progress of Bioactive Peptides Derived from Traditional Natural Products in China. Molecules. 2023; 28(17):6421. https://doi.org/10.3390/molecules28176421
Chicago/Turabian StyleZhang, Yanyan, Lianghong Liu, Min Zhang, Shani Li, Jini Wu, Qiuju Sun, Shengjun Ma, and Wei Cai. 2023. "The Research Progress of Bioactive Peptides Derived from Traditional Natural Products in China" Molecules 28, no. 17: 6421. https://doi.org/10.3390/molecules28176421
APA StyleZhang, Y., Liu, L., Zhang, M., Li, S., Wu, J., Sun, Q., Ma, S., & Cai, W. (2023). The Research Progress of Bioactive Peptides Derived from Traditional Natural Products in China. Molecules, 28(17), 6421. https://doi.org/10.3390/molecules28176421







