Identification of Novel Cannabinoid CB2 Receptor Agonists from Botanical Compounds and Preliminary Evaluation of Their Anti-Osteoporotic Effects
Abstract
:1. Introduction
2. Results
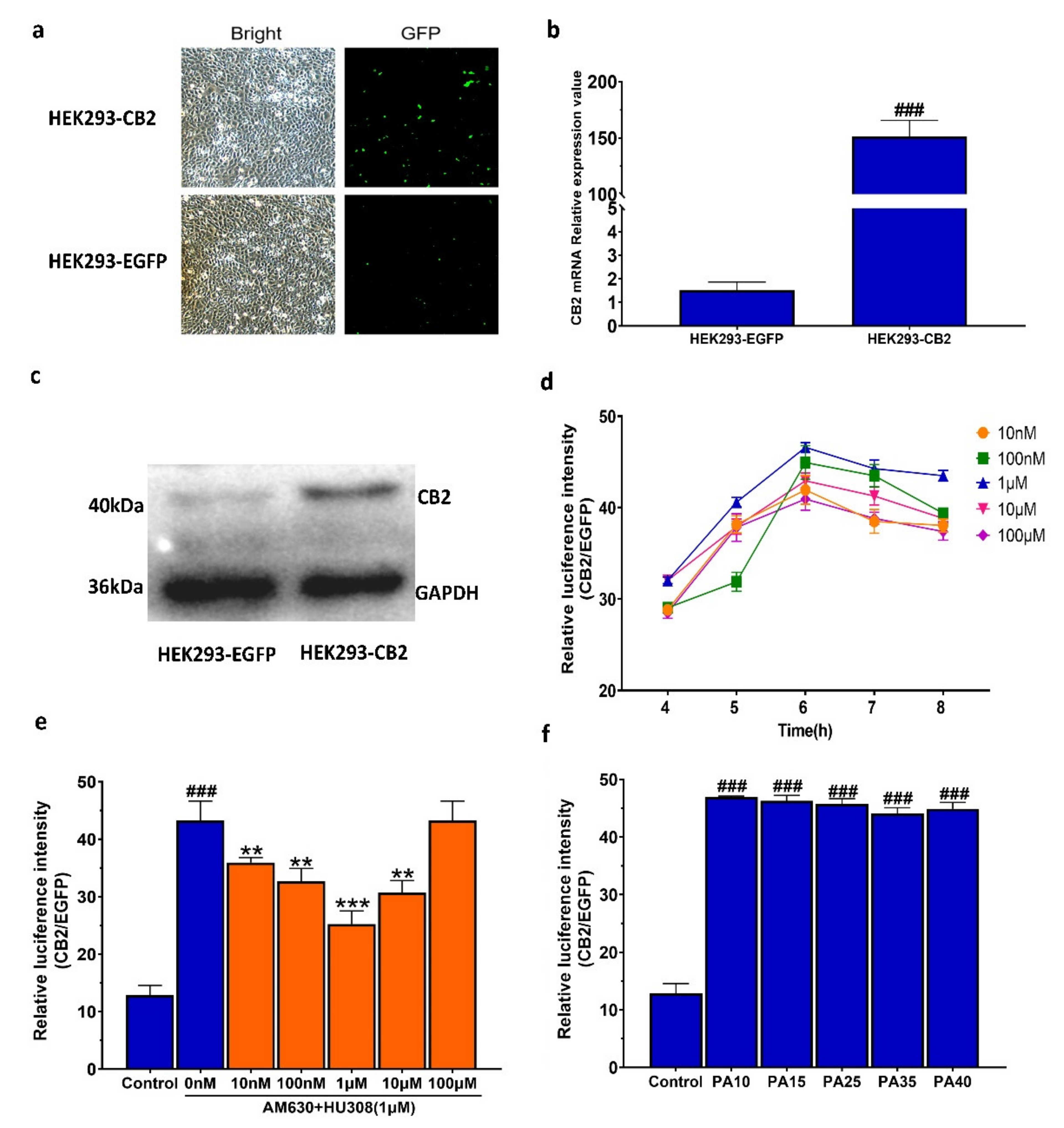
2.1. Establishment of the Double Luciferase Screening System for Screening CB2R Modulators
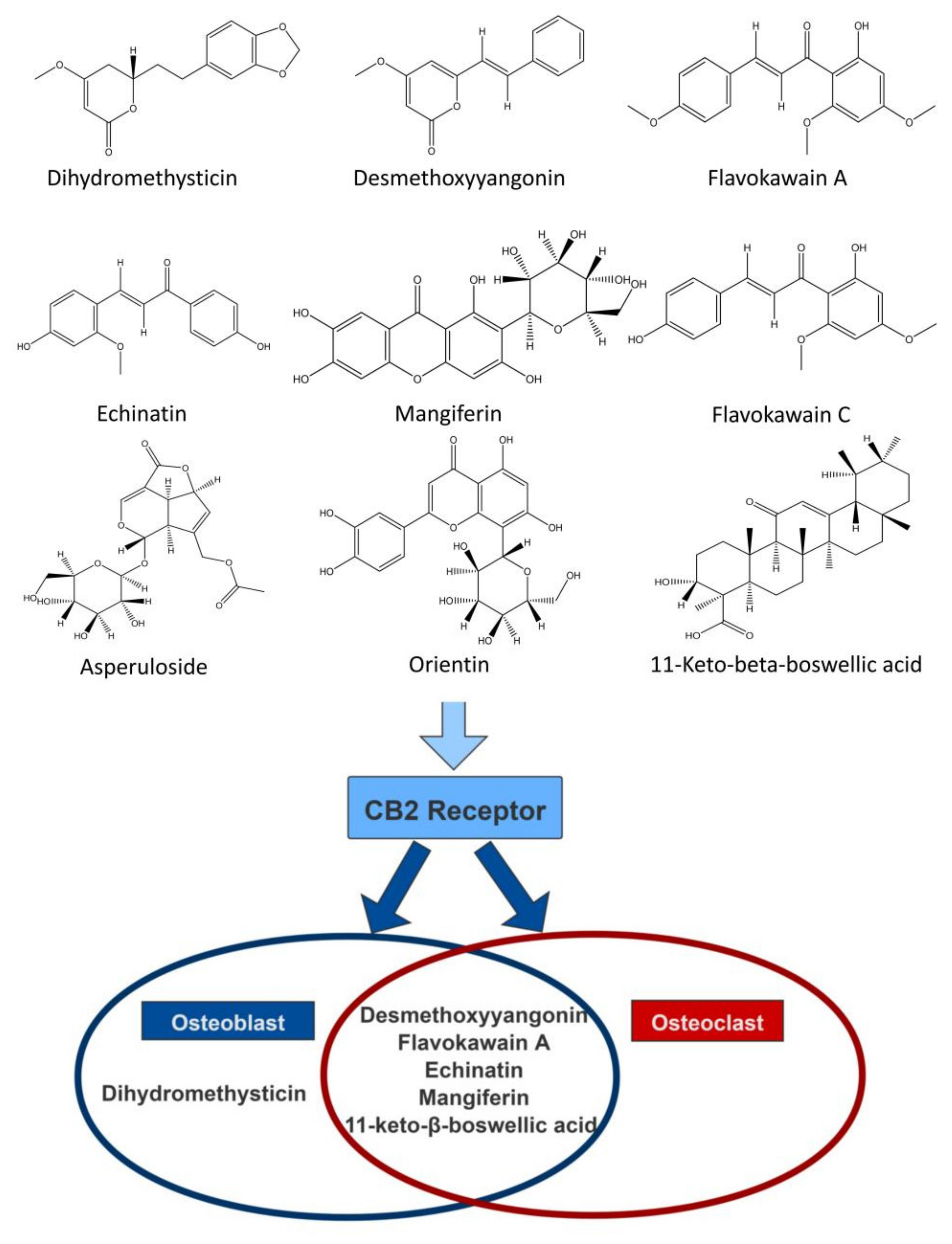
2.2. Screening of CB2R Agonists from Botanical Compounds
2.3. Effects of the Screened CB2R Agonists on the Expression Level of Intracellular cAMP in HEK293-CB2 Cells
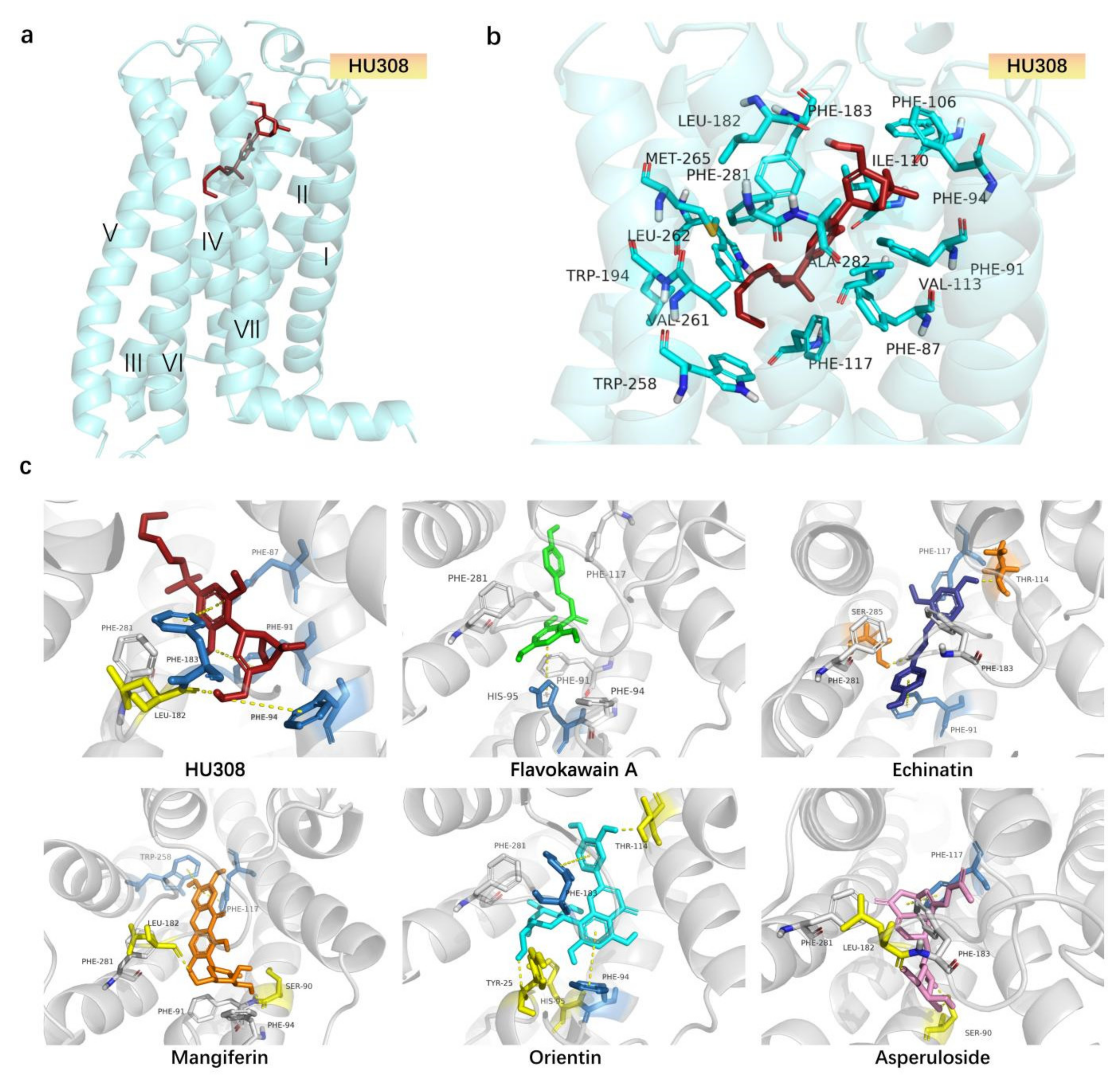
2.4. Molecular Docking Analysis of CB2R Agonists from the Botanical Compounds by Comparison with HU308
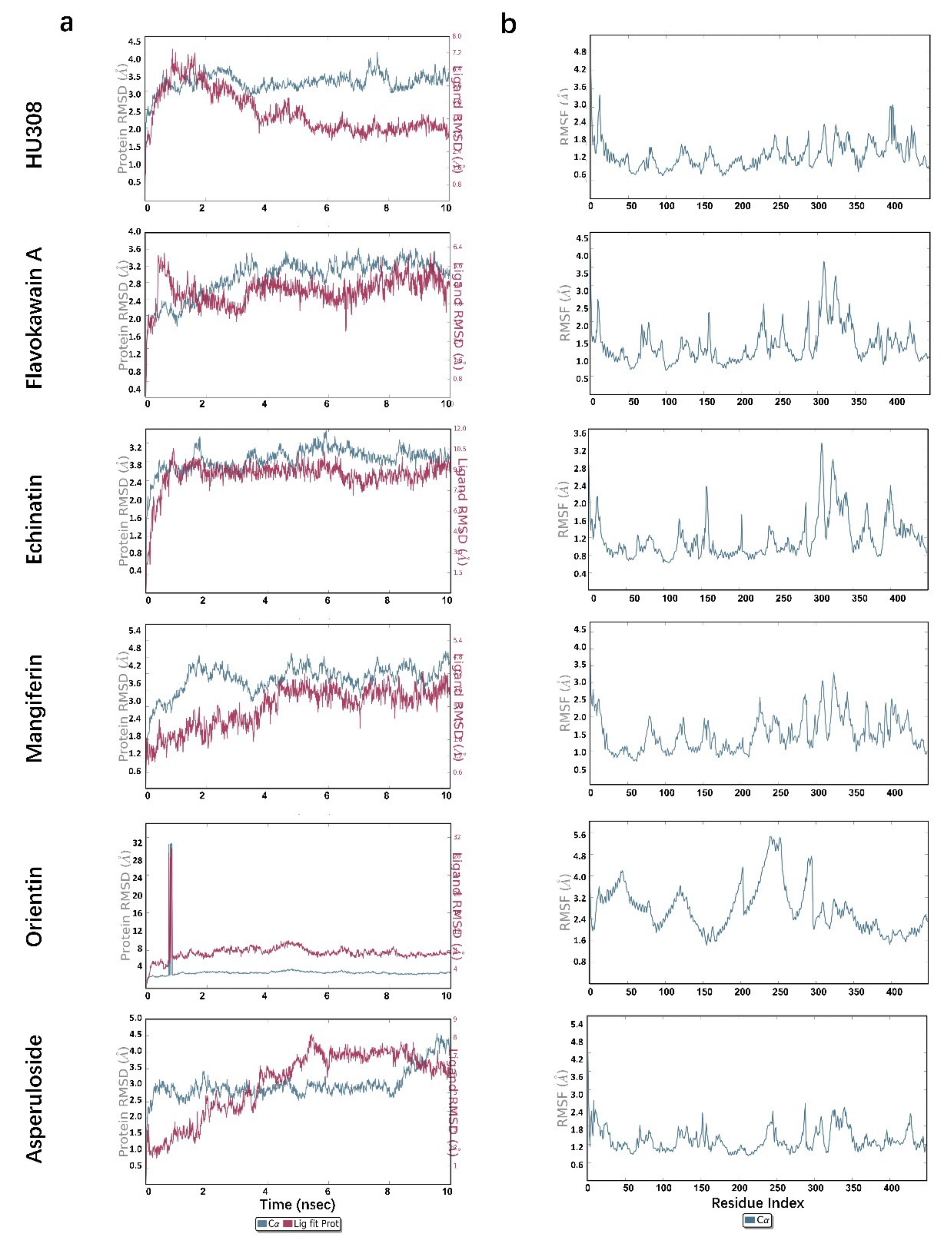
2.5. Molecular Dynamics Simulations
2.6. CB2R Agonists Improve the Function of Osteoblasts Derived from Bone Marrow Mesenchymal Stem Cells (BMSCs) in a CB2-Dependent Manner
2.7. CB2R Agonists Inhibitthe Activity of Osteoclastsin a CB2-Dependent Manner
3. Discussion
4. Materials and Methods
4.1. Reagents
4.2. Cell Culture
4.3. Establishment of CB2R Double Luciferase Screening System
4.3.1. Construction of CB2-Specific Reporter Plasmid
4.3.2. Establishment of HEK293-CB2R Cells
4.3.3. Fluorescence Quantitative Reverse Transcriptase-Polymerase Chain Reaction (FQ-RT-PCR)
4.3.4. Western Blot Analysis
4.3.5. Assay Optimization
4.3.6. Stability Evaluation of HEK293–CB2 Cells
4.4. Screening of Botanical Compounds
4.5. Determination of Levels of Cyclic Adenosine Monophosphate (cAMP) in Cells
4.6. Molecular Docking
4.7. Molecular Dynamics Simulation
4.8. Determination of Proliferation, ALP Activities, and Bone Mineralized Nodules of Osteoblasts
4.9. Determination of Viability and TRAP Activityin Osteoclasts
4.10. F-Actin Ring Formation Assay
4.11. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Di Marzo, V.; Bifulco, M.; De Petrocellis, L. The Endocannabinoid System and Its Therapeutic Exploitation. Nat. Rev. Drug. Discov. 2004, 3, 771–784. [Google Scholar] [CrossRef] [PubMed]
- Pollastro, F.; Minassi, A.; Fresu, L.G. Cannabis Phenolics and their Bioactivities. Curr. Med. Chem. 2018, 25, 1160–1185. [Google Scholar] [CrossRef]
- Bukke, V.N.; Archana, M.; Villani, R.; Serviddio, G.; Cassano, T. Pharmacological and Toxicological Effects of Phytocannabinoids and Recreational Synthetic Cannabinoids: Increasing Risk of Public Health. Pharmaceuticals 2021, 14, 965. [Google Scholar] [CrossRef] [PubMed]
- Mlost, J.; Kac, P.; Kędziora, M.; Starowicz, K. Antinociceptive and chondroprotective effects of prolonged β-caryophyllene treatment in the animal model of osteoarthritis: Focus on tolerance development. Neuropharmacology 2021, 204, 108908. [Google Scholar] [CrossRef]
- Meyer, H.C.; Lee, F.S.; Gee, D.G. The Role of the Endocannabinoid System and Genetic Variation in Adolescent Brain Development. Neuropsychopharmacology 2018, 43, 21–33. [Google Scholar] [CrossRef] [Green Version]
- Morales, P.; Hernandez-Folgado, L.; Goya, P.; Jagerovic, N. Cannabinoid receptor 2 (CB 2) agonists and antagonists: A patent update. Expert Opin. Ther. Patents 2016, 26, 843–856. [Google Scholar] [CrossRef]
- Apostu, D.; Lucaciu, O.; Mester, A.; Benea, H.; Oltean-Dan, D.; Onisor, F.; Baciut, M.; Bran, S. Cannabinoids and bone regeneration. Drug Metab. Rev. 2019, 51, 65–75. [Google Scholar] [CrossRef] [PubMed]
- Galiegue, S.; Mary, S.; Marchand, J.; Dussossoy, D.; Carriere, D.; Carayon, P.; Bouaboula, M.; Shire, D.; Le Fur, G.; Casellas, P. Expression of Central and Peripheral Cannabinoid Receptors in Human Immune Tissues and Leukocyte Subpopulations. Eur. J. Biochem. 1995, 232, 54–61. [Google Scholar] [CrossRef]
- Schatz, A.R.; Lee, M.; Condie, R.B.; Pulaski, J.T.; Kaminski, N.E. Cannabinoid Receptors CB1 and CB2: A Characterization of Expression and Adenylate Cyclase Modulation within the Immune System. Toxicol. Appl. Pharmacol. 1997, 142, 278–287. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, G.C.; Pai, M.; Chandran, P.; Hooker, B.A.; Zhu, C.Z.; Salyers, A.K.; Wensink, E.J.; Zhan, C.; Carroll, W.A.; Dart, M.J.; et al. Central and peripheral sites of action for CB2 receptor mediated analgesic activity in chronic inflammatory and neuropathic pain models in rats. Br. J. Pharmacol. 2011, 162, 428–440. [Google Scholar] [CrossRef] [Green Version]
- Yang, L.; Li, Y.; Ren, J.; Zhu, C.; Fu, J.; Lin, D.; Qiu, Y. Celastrol Attenuates Inflammatory and Neuropathic Pain Mediated by Cannabinoid Receptor Type 2. Int. J. Mol. Sci. 2014, 15, 13637–13648. [Google Scholar] [CrossRef] [Green Version]
- Cui, J.H.; Kim, W.M.; Lee, H.G.; Kim, Y.O.; Kim, C.M.; Yoon, M.H. Antinociceptive effect of intrathecal cannabinoid receptor agonist WIN 55,212-2 in a rat bone tumor pain model. Neurosci. Lett. 2011, 493, 67–71. [Google Scholar] [CrossRef] [PubMed]
- Wilkinson, S.T.; D’souza, D.C. Problems with the Medicalization of Marijuana. JAMA 2014, 311, 2377–2378. [Google Scholar] [CrossRef]
- Zhu, M.; Yu, B.; Bai, J.; Wang, X.; Guo, X.; Liu, Y.; Lin, J.; Hu, S.; Zhang, W.; Tao, Y.; et al. Cannabinoid Receptor 2 Agonist Prevents Local and Systemic Inflammatory Bone Destruction in Rheumatoid Arthritis. J. Bone Miner. Res. 2019, 34, 739–751. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y. The Investigation of Energy Metabolism in Osteoblasts and Osteoclasts. Hua Xi Kou Qiang Yi Xue Za Zhi 2021, 39, 501–509. [Google Scholar]
- Xu, A.; Yang, Y.; Shao, Y.; Wu, M.; Sun, Y. Activation of cannabinoid receptor type 2-induced osteogenic differentiation involves autophagy induction and p62-mediated Nrf2 deactivation. Cell Commun. Signal. 2020, 18, 1–11. [Google Scholar] [CrossRef] [Green Version]
- Ofek, O.; Karsak, M.; Leclerc, N.; Fogel, M.; Frenkel, B.; Wright, K.; Tam, J.; Attar-Namdar, M.; Kram, V.; Shohami, E.; et al. Peripheral cannabinoid receptor, CB2, regulates bone mass. Proc. Natl. Acad. Sci. USA 2006, 103, 696–701. [Google Scholar] [CrossRef] [Green Version]
- Sun, Y.-X.; Xu, A.-H.; Yang, Y.; Zhang, J.-X.; Yu, A.-W. Activation of Cannabinoid Receptor 2 Enhances Osteogenic Differentiation of Bone Marrow Derived Mesenchymal Stem Cells. BioMed Res. Int. 2015, 2015, 1–8. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tian, F.; Yang, H.; Huang, T.; Chen, F.; Xiong, F. Involvement of CB2 signalling pathway in the development of osteoporosis by regulating the proliferation and differentiation of hBMSCs. J. Cell. Mol. Med. 2021, 25, 2426–2435. [Google Scholar] [CrossRef]
- Geng, D.-C.; Xu, Y.-Z.; Yang, H.-L.; Zhu, G.-M.; Wang, X.-B.; Zhu, X.-S. Cannabinoid receptor-2 selective antagonist negatively regulates receptor activator of nuclear factor kappa B ligand mediated osteoclastogenesis. Chin. Med. J. 2011, 124, 586–590. [Google Scholar]
- Sophocleous, A.; Marino, S.; Logan, J.G.; Mollat, P.; Ralston, S.H.; Idris, A.I. Bone Cell-autonomous Contribution of Type 2 Cannabinoid Receptor to Breast Cancer-induced Osteolysis. J. Biol. Chem. 2015, 290, 22049–22060. [Google Scholar] [CrossRef] [Green Version]
- Lozano-Ondoua, A.N.; Hanlon, K.E.; Symons-Liguori, A.M.; Largent-Milnes, T.M.; Havelin, J.J.; Ferland, H.L., 3rd; Chandramouli, A.; Owusu-Ankomah, M.; Nikolich-Zugich, T.; Bloom, A.P.; et al. Disease Modification of Breast Cancer-Induced Bone Remodeling by Cannabinoid 2 Receptor Agonists. J. Bone Miner. Res. 2013, 28, 92–107. [Google Scholar] [CrossRef] [PubMed]
- Rossi, F.; Tortora, C.; Punzo, F.; Bellini, G.; Argenziano, M.; Di Paola, A.; Torella, M.; Perrotta, S. The Endocannabinoid/Endovanilloid System in Bone: From Osteoporosis to Osteosarcoma. Int. J. Mol. Sci. 2019, 20, 1919. [Google Scholar] [CrossRef] [Green Version]
- Kuchta, K.; Cameron, S. Tradition to Pathogenesis: A Novel Hypothesis for Elucidating the Pathogenesis of Diseases Based on the Traditional Use of Medicinal Plants. Front. Pharmacol. 2021, 12, 705077. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.-H.; Chung, T.D.Y.; Oldenburg, K.R. A Simple Statistical Parameter for Use in Evaluation and Validation of High Throughput Screening Assays. J. Biomol. Screen. 1999, 4, 67–73. [Google Scholar] [CrossRef]
- Mammana, S.; Cavalli, E.; Gugliandolo, A.; Silvestro, S.; Pollastro, F.; Bramanti, P.; Mazzon, E. Could the combination of two non-psychotropic cannabinoids counteract neuroinflammation? Effectiveness of cannabidiol associated with cannabigerol. Medicina 2019, 55, 747. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Munro, S.; Thomas, K.L.; Abu-Shaar, M. Molecular characterization of a peripheral receptor for cannabinoids. Nature 1993, 365, 61–65. [Google Scholar] [CrossRef] [PubMed]
- Bouaboula, M.; Desnoyer, N.; Carayon, P.; Combes, T.; Casellas, P. Gi protein modulation induced by a selective inverse agonist for the peripheral cannabinoid receptor CB2: Implication for intracellular signalization cross-regulation. Mol. Pharmacol. 1999, 55, 473–480. [Google Scholar]
- Zoete, V.; Grosdidier, A.; Michielin, O. Docking, virtual high throughput screening and in silico fragment-based drug design. J. Cell. Mol. Med. 2009, 13, 238–248. [Google Scholar] [CrossRef] [Green Version]
- Brandman, R.; Brandman, Y.; Pande, V.S. A-Site Residues Move Independently from P-Site Residues in all-Atom Molecular Dynamics Simulations of the 70S Bacterial Ribosome. PLoS ONE 2012, 7, e29377. [Google Scholar] [CrossRef]
- Bab, I.; Ofek, O.; Tam, J.; Rehnelt, J.; Zimmer, A. Endocannabinoids and the Regulation of Bone Metabolism. J. Neuroendocr. 2008, 20, 69–74. [Google Scholar] [CrossRef]
- Takada, Y.; Ichikawa, H.; Badmaev, V.; Aggarwal, B.B. Acetyl-11-Keto-Beta-Boswellic Acid Potentiates Apoptosis, Inhibits Invasion, and Abolishes Osteoclastogenesis by Suppressing Nf-Kappa B and Nf-Kappa B-Regulated Gene Expression. J. Immunol. 2006, 176, 3127–3140. [Google Scholar] [CrossRef] [Green Version]
- Sekiguchi, Y.; Mano, H.; Nakatani, S.; Shimizu, J.; Kataoka, A.; Ogura, K.; Kimira, Y.; Ebata, M.; Wada, M. Mangiferin positively regulates osteoblast differentiation and suppresses osteoclast differentiation. Mol. Med. Rep. 2017, 16, 1328–1332. [Google Scholar] [CrossRef] [Green Version]
- Ding, L.Z.; Teng, X.; Zhang, Z.B.; Zheng, C.J.; Chen, S.H. Mangiferin Inhibits Apoptosis and Oxidative Stress Via Bmp2/Smad-1 Signaling in Dexamethasone-Induced Mc3t3-E1 Cells. Int. J. Mol. Med. 2018, 41, 2517–2526. [Google Scholar] [CrossRef] [Green Version]
- Ang, E.; Liu, Q.; Qi, M.; Liu, H.G.; Yang, X.; Chen, H.; Zheng, M.H.; Xu, J. Mangiferin Attenuates Osteoclastogenesis, Bone Resorption, and Rankl-Induced Activation of Nf-Κb and Erk. J. Cell Biochem. 2011, 112, 89–97. [Google Scholar] [CrossRef] [PubMed]
- Xu, G.; Fu, S.; Zhan, X.; Wang, Z.; Zhang, P.; Shi, W.; Qin, N.; Chen, Y.; Wang, C.; Niu, M.; et al. Echinatin effectively protects against NLRP3 inflammasome–driven diseases by targeting HSP90. JCI Insight 2021, 6, e134601. [Google Scholar] [CrossRef]
- Shi, J.; Gu, Y.; Wang, Y.; Bai, J.; Xiong, L.; Tao, Y.; Xue, Y.; Xu, Y.; Yang, H.; Ye, H.; et al. Inhibitory Effect of Acetyl-11-Keto-Beta-Boswellic Acid on Titanium Particle-Induced Bone Loss by Abrogating Osteoclast Formation and Downregulating the Erk Signaling Pathway. Int. Immunopharmacol. 2021, 94, 107459. [Google Scholar] [CrossRef]
- Xiong, L.; Liu, Y.; Zhu, F.; Lin, J.; Wen, D.; Wang, Z.; Bai, J.; Ge, G.; Xu, C.; Gu, Y.; et al. Acetyl-11-Keto-Beta-Boswellic Acid Attenuates Titanium Particle-Induced Osteogenic Inhibition Via Activation of the Gsk-3beta/Beta-Catenin Signaling Pathway. Theranostics 2019, 9, 7140–7155. [Google Scholar] [CrossRef]
- Al-Dhubiab, B.E.; Patel, S.S.; Morsy, M.A.; Duvva, H.; Nair, A.B.; Deb, P.K.; Shah, J. The Beneficial Effect of Boswellic Acid on Bone Metabolism and Possible Mechanisms of Action in Experimental Osteoporosis. Nutrients 2020, 12, 3186. [Google Scholar] [CrossRef]
- Tucci, M.A.; Pride, Y.; Strickland, S.; Marocho, S.M.S.; Jackson, R.J.; Jefferson, J.R.; Chade, A.R.; Grill, R.J.; Grayson, B.E. Delayed Systemic Treatment with Cannabinoid Receptor 2 Agonist Mitigates Spinal Cord Injury-Induced Osteoporosis More Than Acute Treatment Directly after Injury. Neurotrauma Rep. 2021, 2, 270–284. [Google Scholar] [CrossRef]
- Sophocleous, A.; Yiallourides, M.; Zeng, F.; Pantelas, P.; Stylianou, E.; Li, B.; Carrasco, G.; Idris, A.I. Association of cannabinoid receptor modulation with normal and abnormal skeletal remodelling: A systematic review and meta-analysis of in vitro, in vivo and human studies. Pharmacol. Res. 2021, 175, 105928. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, J.V.; Pierotte, I.C.; Rodrigues, F.F.; Souza, L.C.R.; Bastos, R.W.; Carmo, P.H.F.; Cassali, G.D.; Tagliati, C.A.; Machado, R.R.; Santos, D.A.; et al. Acute Oral Toxicity, Antinociceptive and Antimicrobial Activities of Kava Dried Extracts and Synthetic Kavain. Nat. Prod. Res. 2021, 1–6. [Google Scholar] [CrossRef]
- Li, X.; Song, L.; Xu, S.; Tippin, M.; Meng, S.; Xie, J.; Uchio, E.; Zi, X. Kava root extracts hinder prostate cancer development and tumorigenesis by involvement of dual inhibition of MAO-A and LSD1. J. Transl. Genet. Genom. 2021, 5, 163–172. [Google Scholar] [CrossRef] [PubMed]
- Chou, T.-W.; Feng, J.-H.; Huang, C.-C.; Cheng, Y.-W.; Chien, S.-C.; Wang, S.-Y.; Shyur, L.-F. A Plant Kavalactone Desmethoxyyangonin Prevents Inflammation and Fulminant Hepatitis in Mice. PLoS ONE 2013, 8, e77626. [Google Scholar] [CrossRef] [Green Version]
- Hu, Q.; Corral, P.; Narayanapillai, S.C.; Leitzman, P.; Upadhyaya, P.; O’Sullivan, M.G.; Hecht, S.S.; Lu, J.; Xing, C. Oral Dosing of Dihydromethysticin Ahead of Tobacco Carcinogen NNK Effectively Prevents Lung Tumorigenesis in A/J Mice. Chem. Res. Toxicol. 2020, 33, 1980–1988. [Google Scholar] [CrossRef] [PubMed]
- Narayanapillai, S.C.; Lin, S.-H.; Leitzman, P.; Upadhyaya, P.; Baglole, C.J.; Xing, C. Dihydromethysticin (DHM) Blocks Tobacco Carcinogen 4-(Methylnitrosamino)-1-(3-pyridyl)-1-butanone (NNK)-Induced O6-Methylguanine in a Manner Independent of the Aryl Hydrocarbon Receptor (AhR) Pathway in C57BL/6 Female Mice. Chem. Res. Toxicol. 2016, 29, 1828–1834. [Google Scholar] [CrossRef] [PubMed]
- Hseu, Y.C.; Yang, T.Y.; Li, M.L.; Rajendran, P.; Mathew, D.C.; Tsai, C.H.; Lin, R.W.; Lee, C.C.; Yang, H.L. Chalcone Flavokawain a Attenuates Tgf-Beta1-Induced Fibrotic Pathology Via Inhibition of Ros/Smad3 Signaling Pathways and Induction of Nrf2/Are-Mediated Antioxidant Genes in Vascular Smooth Muscle Cells. J. Cell Mol. Med. 2019, 23, 775–788. [Google Scholar] [CrossRef]
- Yang, H.L.; Yang, T.Y.; Gowrisankar, Y.V.; Liao, C.H.; Liao, J.W.; Huang, P.J.; Hseu, Y.C. Suppression of Lps-Induced Inflammation by Chalcone Flavokawain a through Activation of Nrf2/Are-Mediated Antioxidant Genes and Inhibition of Ros/Nfκb Signaling Pathways in Primary Splenocytes. Oxid. Med. Cell Longev. 2020, 2020, 3476212. [Google Scholar] [CrossRef]
- Rajendran, P.; Alzahrani, A.M.; Veeraraghavan, V.P.; Ahmed, E.A. Anti-Apoptotic Effect of Flavokawain A on Ochratoxin-A-Induced Endothelial Cell Injury by Attenuation of Oxidative Stress via PI3K/AKT-Mediated Nrf2 Signaling Cascade. Toxins 2021, 13, 745. [Google Scholar] [CrossRef]
- Malek, S.N.A.; Phang, C.-W.; Karsani, S.A. Induction of apoptosis and cell cycle arrest by flavokawain C on HT-29 human colon adenocarcinoma via enhancement of reactive oxygen species generation, upregulation of p21, p27, and Gadd153, and inactivation of inhibitor of apoptosis proteins. Pharmacogn. Mag. 2017, 13, S321–S328. [Google Scholar] [CrossRef] [Green Version]
- Phang, C.-W.; Gandah, N.A.; Malek, S.N.A.; Karsani, S.A. Proteomic analysis of flavokawain C-induced cell death in HCT 116 colon carcinoma cell line. Eur. J. Pharmacol. 2019, 853, 388–399. [Google Scholar] [CrossRef]
- Boswellia Serrata. In Livertox: Clinical and Research Information on Drug-Induced Liver Injury; National Institute of Diabetes and Digestive and Kidney Diseases: Bethesda, MD, USA, 2012.
- Ammon, H. Boswellic extracts and 11-keto-ß-boswellic acids prevent type 1 and type 2 diabetes mellitus by suppressing the expression of proinflammatory cytokines. Phytomedicine 2019, 63, 153002. [Google Scholar] [CrossRef]
- Jiang, X.; Wang, Y.; Zhang, B.; Fei, X.; Guo, X.; Jia, Y.; Yu, W. Acetyl-11-Keto-Beta-Boswellic Acid Regulates the Repair of Rat Sciatic Nerve Injury by Promoting the Proliferation of Schwann Cells. Life Sci. 2020, 254, 116887. [Google Scholar] [CrossRef]
- Marefati, N.; Beheshti, F.; Memarpour, S.; Bayat, R.; Shafei, M.N.; Sadeghnia, H.R.; Ghazavi, H.; Hosseini, M. The Effects of Acetyl-11-Keto-Beta-Boswellic Acid on Brain Cytokines and Memory Impairment Induced by Lipopolysaccharide in Rats. Cytokine 2020, 131, 155107. [Google Scholar] [CrossRef] [PubMed]
- Schmiech, M.; Ulrich, J.; Lang, S.J.; Büchele, B.; Paetz, C.; St-Gelais, A.; Syrovets, T.; Simmet, T. 11-Keto-α-Boswellic Acid, a Novel Triterpenoid from Boswellia spp. with Chemotaxonomic Potential and Antitumor Activity against Triple-Negative Breast Cancer Cells. Molecules 2021, 26, 366. [Google Scholar] [CrossRef] [PubMed]
- Lu, C.W.; Lin, T.Y.; Wang, S.J. 11-Keto-Beta-Boswellic Acid Attenuates Glutamate Release and Kainic Acid-Induced Excitotoxicity in the Rat Hippocampus. Planta Med. 2020, 86, 434–441. [Google Scholar]
- Siddiqui, A.; Shah, Z.; Jahan, R.N.; Othman, I.; Kumari, Y. Mechanistic role of boswellic acids in Alzheimer’s disease: Emphasis on anti-inflammatory properties. Biomed. Pharmacother. 2021, 144, 112250. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.A.; Ali, R.; Parveen, R.; Najmi, A.K.; Ahmad, S. Pharmacological evidences for cytotoxic and antitumor properties of Boswellic acids from Boswellia serrata. J. Ethnopharmacol. 2016, 191, 315–323. [Google Scholar] [CrossRef]
- Ammon, H.P.T. Modulation of the immune system by Boswellia serrata extracts and boswellic acids. Phytomedicine 2010, 17, 862–867. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, A.; Yokoyama, Y.; Tanaka, K.; Benegiamo, G.; Hirayama, A.; Zhu, Q.; Kitamura, N.; Sugizaki, T.; Morimoto, K.; Itoh, H.; et al. Asperuloside Improves Obesity and Type 2 Diabetes through Modulation of Gut Microbiota and Metabolic Signaling. iScience 2020, 23, 101522. [Google Scholar] [CrossRef]
- Manzione, M.G.; Martorell, M.; Sharopov, F.; Bhat, N.G.; Kumar, N.V.A.; Fokou, P.V.T.; Pezzani, R. Phytochemical and pharmacological properties of asperuloside, a systematic review. Eur. J. Pharmacol. 2020, 883, 173344. [Google Scholar] [CrossRef] [PubMed]
- Chan, Y.; Ng, S.W.; Tan, J.Z.X.; Gupta, G.; Tambuwala, M.M.; Bakshi, H.A.; Dureja, H.; Dua, K.; Ishaq, M.; Caruso, V.; et al. Emerging therapeutic potential of the iridoid molecule, asperuloside: A snapshot of its underlying molecular mechanisms. Chem. Interact. 2020, 315, 108911. [Google Scholar] [CrossRef] [PubMed]
- He, J.; Lu, X.; Wei, T.; Dong, Y.; Cai, Z.; Tang, L.; Liu, M. Asperuloside and Asperulosidic Acid Exert an Anti-Inflammatory Effect Via Suppression of the Nf-Κb and Mapk Signaling Pathways in Lps-Induced Raw 264.7 Macrophages. Int. J. Mol. Sci. 2018, 19, 2027. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, D.; Yu, J.; Huang, P.; Qader, M.; Manavalan, A.; Wu, X.; Kim, J.-C.; Pang, C.; Cao, S.; Kang, K.; et al. Identification of Anti-Inflammatory Compounds from Hawaiian Noni (Morindacitrifolia L.) Fruit Juice. Molecules 2020, 25, 4968. [Google Scholar] [CrossRef]
- Akkol, E.K.; Ilhan, M.; Karpuz, B.; Taştan, H.; Sobarzo-Sánchez, E.; Khan, H. Beneficial effects of Ajuga chamaepitys (L.) Schreber subsp. chia (Schreber) and its iridoids on the colitis model: Histopathological and biochemical evidence. Food Chem. Toxicol. 2020, 144, 111589. [Google Scholar] [CrossRef]
- Ran, H.; Liu, H.; Wu, P. Echinatin Mitigates H2O2-Induced Oxidative Damage and Apoptosis in Lens Epithelial Cells Via the Nrf2/Ho-1 Pathway. Adv. Clin. Exp. Med. 2021, 30, 1195–1203. [Google Scholar] [CrossRef] [PubMed]
- Hong, P.; Liu, Q.W.; Xie, Y.; Zhang, Q.H.; Liao, L.; He, Q.Y.; Li, B.; Xu, W.W. Echinatin Suppresses Esophageal Cancer Tumor Growth and Invasion through Inducing Akt/Mtor-Dependent Autophagy and Apoptosis. Cell Death Dis. 2020, 11, 524. [Google Scholar] [CrossRef] [PubMed]
- Mittal, S.; Iqubal, M.K.; Iqbal, B.; Gupta, M.M.; Ali, J.; Baboota, S. A pervasive scientific overview on mangiferin in the prevention and treatment of various diseases with preclinical and clinical updates. J. Complement. Integr. Med. 2020, 18, 9–21. [Google Scholar] [CrossRef]
- Khare, P.; Shanker, K. Mangiferin: A review of sources and interventions for biological activities. BioFactors 2016, 42, 504–514. [Google Scholar] [CrossRef]
- Wu, T.; Wang, X.; Xiong, H.; Deng, Z.; Peng, X.; Xiao, L.; Jiang, L.; Sun, Y. Bioactives and their metabolites from Tetrastigmahemsleyanum leaves ameliorate DSS-induced colitis via protecting the intestinal barrier, mitigating oxidative stress and regulating the gut microbiota. Food Funct. 2021, 12, 11760–11776. [Google Scholar] [CrossRef]
- Wang, L.; Jing, S.; Qu, H.; Wang, K.; Jin, Y.; Ding, Y.; Yang, L.; Yu, H.; Shi, Y.; Li, Q.; et al. Orientin mediates protection against MRSA-induced pneumonia by inhibiting Sortase A. Virulence 2021, 12, 2149–2161. [Google Scholar] [CrossRef] [PubMed]
- Mthembu, S.X.H.; Muller, C.J.F.; Dludla, P.V.; Madoroba, E.; Kappo, A.P.; Mazibuko-Mbeje, S.E. Rooibos Flavonoids, Aspalathin, Isoorientin, and Orientin Ameliorate Antimycin A-Induced Mitochondrial Dysfunction by Improving Mitochondrial Bioenergetics in Cultured Skeletal Muscle Cells. Molecules 2021, 26, 6289. [Google Scholar] [CrossRef]
- Mazibuko-Mbeje, S.E.; Mthembu, S.X.H.; Tshiitamune, A.; Muvhulawa, N.; Mthiyane, F.T.; Ziqubu, K.; Muller, C.J.F.; Dludla, P.V. Orientin Improves Substrate Utilization and the Expression of Major Genes Involved in Insulin Signaling and Energy Regulation in Cultured Insulin-Resistant Liver Cells. Molecules 2021, 26, 6154. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Hua, T.; Vemuri, K.; Ho, J.-H.; Wu, Y.; Wu, L.; Popov, P.; Benchama, O.; Zvonok, N.; Locke, K.; et al. Crystal Structure of the Human Cannabinoid Receptor CB2. Cell 2019, 176, 459–467.e13. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Case, D.A.; Cheatham, T.E., 3rd; Darden, T.; Gohlke, H.; Luo, R.; Merz, K.M., Jr.; Onufriev, A.; Simmerling, C.; Wang, B.; Woods, R.J. The Amber Biomolecular Simulation Programs. J.Comput. Chem. 2005, 26, 1668–1688. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, J.; Wolf, R.M.; Caldwell, J.W.; Kollman, P.A.; Case, D.A. Development and testing of a general amber force field. J. Comput. Chem. 2004, 25, 1157–1174. [Google Scholar] [CrossRef]
- Dickson, C.J.; Madej, B.D.; Skjevik, Å.A.; Betz, R.M.; Teigen, K.; Gould, I.R.; Walker, R.C. Lipid14: The Amber Lipid Force Field. J. Chem. Theory Comput. 2014, 10, 865–879. [Google Scholar] [CrossRef]
- Maier, J.A.; Martinez, C.; Kasavajhala, K.; Wickstrom, L.; Hauser, K.E.; Simmerling, C. ff14SB: Improving the accuracy of protein side chain and backbone parameters from ff99SB. J. Chem. Theory Comput. 2015, 11, 3696–3713. [Google Scholar] [CrossRef] [Green Version]
- Emperador, A.; Crehuet, R.; Guàrdia, E. Effect of the Water Model in Simulations of Protein–Protein Recognition and Association. Polymers 2021, 13, 176. [Google Scholar] [CrossRef]




| S. No | Compound | Docking Score | Amino Acid Residues within 4 Å | No. of Hydrogen Bonds | Hydrogen Bond Interactions | π-π Interactions |
|---|---|---|---|---|---|---|
| 1 | HU308 | −11.604 | TYR25, PHE87, SER90, PHE91, PHE94, HIS95, PHE106, ILE110, VAL113, THR114, PHE117, LEU182, PHE183, PRO184, TRP194, TRP258, VAL261, LEU262, MET265, PHE281, ALA282, SER285, CYS288 | 1 | LEU182 | PHE87, PHE183 |
| 2 | Dihydromethysticin-1 | −8.053 | PHE87, PHE91, HIS95, ILE110, VAL113, THR114, PHE117, PHE183, TRP194, TRP258, VAL261, LEU262, MET265, PHE281, SER285, CYS288 | 0 | - | PHE117 |
| 3 | Desmethoxyyangonin | −7.985 | ILE110, VAL113, THR114, PHE117, PHE183, ILE186, TYR190, LEU191, TRP194, LEU195, TRP258, VAL261, LEU262, MET265, CYS288 | 0 | - | PHE117, PHE183, TRP194 |
| 4 | Flavokawain A | −8.508 | TYR25, ILE27, PHE87, PHE91, PHE94, HIS95, PHE106, ILE110, VAL113, PHE117, LEU182, PHE183, PRO184, TRP194, TRP258, VAL261, MET265, LYS278, PHE281, ALA282, SER285, CYS288 | 1 | SER285 | HIS95 |
| 5 | Echinatin | −8.926 | TYR25, ILE27, PHE87, PHE91, HIS95, ILE110, VAL113, THR114, PHE117, LEU182, PHE183, PRO184, TRP194, VAL261, LYS278, PHE281, ALA282, SER285, CYS288 | 2 | THR114, SER285 | HIS95, PHE183 |
| 6 | Mangiferin | −13.968 | TYR25, PHE87, SER90, PHE91, PHE94, HIS95, PHE106, ILE110, VAL113, THR114, PHE117, THR118, LEU182, PHE183, PRO184, TRP194, TRP258, VAL261, PHE281, ALA282, SER285, CYS288 | 3 | SER90, LEU182, SER285 | PHE87, PHE183 |
| 7 | 11-Keto-beta-boswellic acid | −6.477 | TYR25, PHE87, SER90, PHE91, PHE94, HIS95, PHE106, ILE110, VAL113, THR114, PHE117, LEU182, PHE183, PRO184, TRP194, VAL261, LEU262, MET265, PHE281, SER285, CYS288 | 1 | SER285 | - |
| 8 | Flavokawain C | −7.769 | TYR25, ILE27, PHE87, SER90, PHE91, PHE94, HIS95, PHE106, LYS109, ILE110, VAL113, THR114, PHE117, LEU182, PHE183, TRP194, VAL261, MET265, LYS278, PHE281, ALA282, SER285 | 1 | LYS278 | PHE94, HIE95 |
| 9 | Orientin | −14.469 | TYR25, ILE27, PHE87, SER90, PHE91, PHE94, HIS95, PHE106, LYS109, ILE110, VAL113, THR114, PHE117, LEU182, PHE183, TRP194, VAL261, MET265, LYS278, PHE281, ALA282, SER285 | 3 | TYR25, PHE95, TPR114 | PHE94, PHE183 |
| 10 | Asperuloside | −12.473 | PHE87, SER90, PHE91, PHE94, HIS95, PHE106, ILE110, VAL113, THR114, PHE117, LEU182, PHE183, PRO184, ILE186, TYR190, LEU191, TRP194, VAL261, MET265, PHE281, SER285, CYS288 | 2 | SER90, LEU182 | - |
| S. No | Compound | T stable of the Protein (ns) | RMSD Value of the Protein stable (Å) | T stable of the ligand (ns) | RMSD Value of the Ligand stable (Å) |
|---|---|---|---|---|---|
| 1 | HU308 | 1~10 | 3.0 | 4~10 | 1.7 |
| 2 | Dihydromethysticin | 8~10 | 3.0 | 2~10 | 3.5 |
| 3 | Desmethoxyyangonin | 4~9 | 3.2 | 4~10 | 3.0 |
| 4 | Flavokawain A | 4~10 | 3.0 | 4~10 | 2.6 |
| 5 | Echinatin | 2~10 | 2.8 | 1~10 | 2.6 |
| 6 | Mangiferin | 4~10 | 3.9 | 4~10 | 3.3 |
| 7 | 11-Keto-beta-boswellic acid | 6~9 | 3.2 | 4~9 | 2.2 |
| 8 | Flavokawain C | 7~10 | 3.2 | 4~8 | 2.5 |
| 9 | Orientin | 1~10 | 3.0 | 1~10 | 7.0 |
| 10 | Asperuloside | 0.2~8 | 2.8 | 6~8.5 | 3.8 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hu, S.-J.; Cheng, G.; Zhou, H.; Zhang, Q.; Zhang, Q.-L.; Wang, Y.; Shen, Y.; Lian, C.-X.; Ma, X.-Q.; Zhang, Q.-Y.; et al. Identification of Novel Cannabinoid CB2 Receptor Agonists from Botanical Compounds and Preliminary Evaluation of Their Anti-Osteoporotic Effects. Molecules 2022, 27, 702. https://doi.org/10.3390/molecules27030702
Hu S-J, Cheng G, Zhou H, Zhang Q, Zhang Q-L, Wang Y, Shen Y, Lian C-X, Ma X-Q, Zhang Q-Y, et al. Identification of Novel Cannabinoid CB2 Receptor Agonists from Botanical Compounds and Preliminary Evaluation of Their Anti-Osteoporotic Effects. Molecules. 2022; 27(3):702. https://doi.org/10.3390/molecules27030702
Chicago/Turabian StyleHu, Si-Jing, Gang Cheng, Hao Zhou, Qi Zhang, Quan-Long Zhang, Yang Wang, Yi Shen, Chen-Xia Lian, Xue-Qin Ma, Qiao-Yan Zhang, and et al. 2022. "Identification of Novel Cannabinoid CB2 Receptor Agonists from Botanical Compounds and Preliminary Evaluation of Their Anti-Osteoporotic Effects" Molecules 27, no. 3: 702. https://doi.org/10.3390/molecules27030702
APA StyleHu, S.-J., Cheng, G., Zhou, H., Zhang, Q., Zhang, Q.-L., Wang, Y., Shen, Y., Lian, C.-X., Ma, X.-Q., Zhang, Q.-Y., & Qin, L.-P. (2022). Identification of Novel Cannabinoid CB2 Receptor Agonists from Botanical Compounds and Preliminary Evaluation of Their Anti-Osteoporotic Effects. Molecules, 27(3), 702. https://doi.org/10.3390/molecules27030702






