Label-Free Quantitative Proteomics to Explore the Action Mechanism of the Pharmaceutical-Grade Triticum vulgare Extract in Speeding Up Keratinocyte Healing
Abstract
:1. Introduction
2. Results
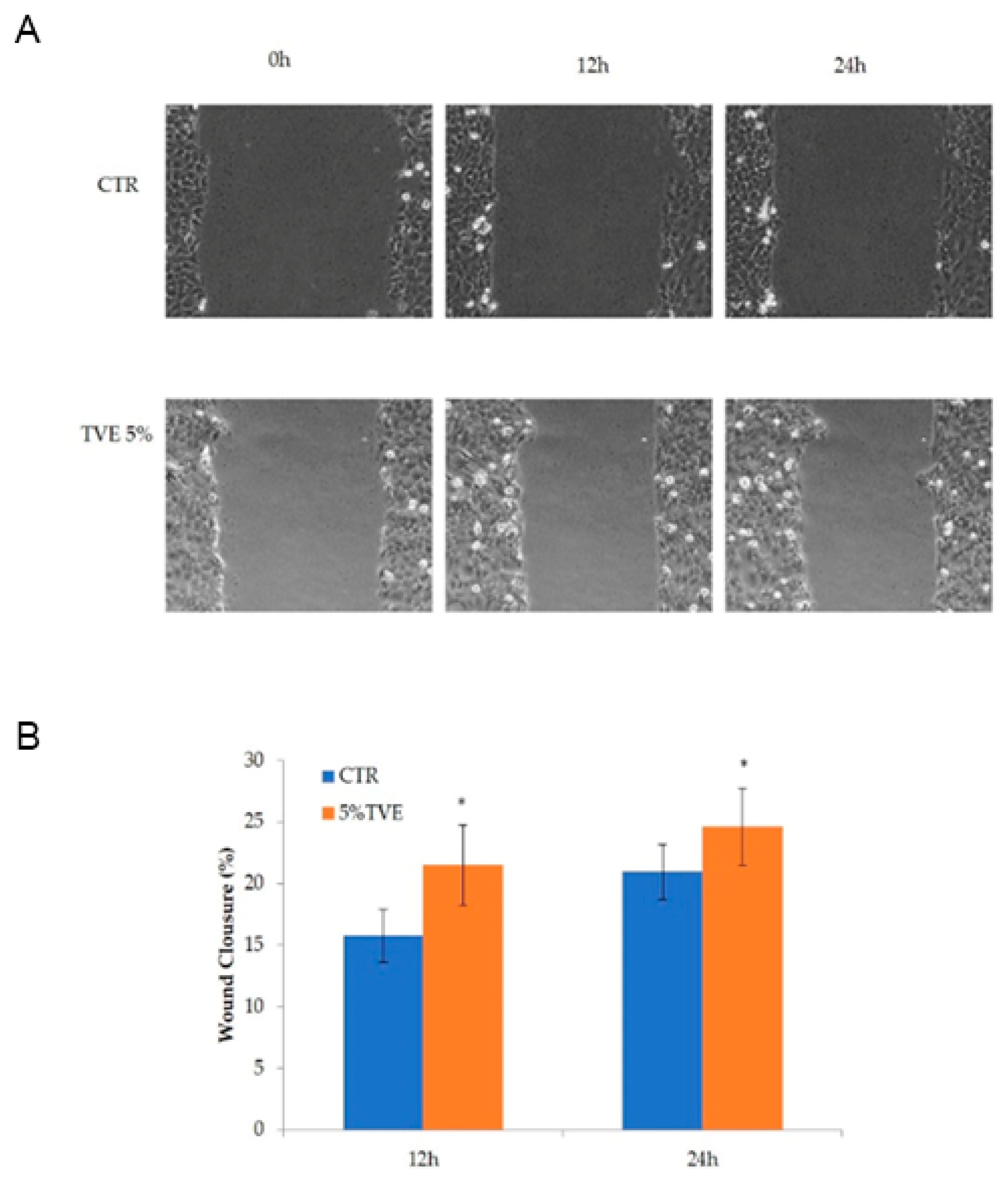
2.1. Evaluation of the TVE Effect in the Keratinocytes Monolayer Scratch Assay
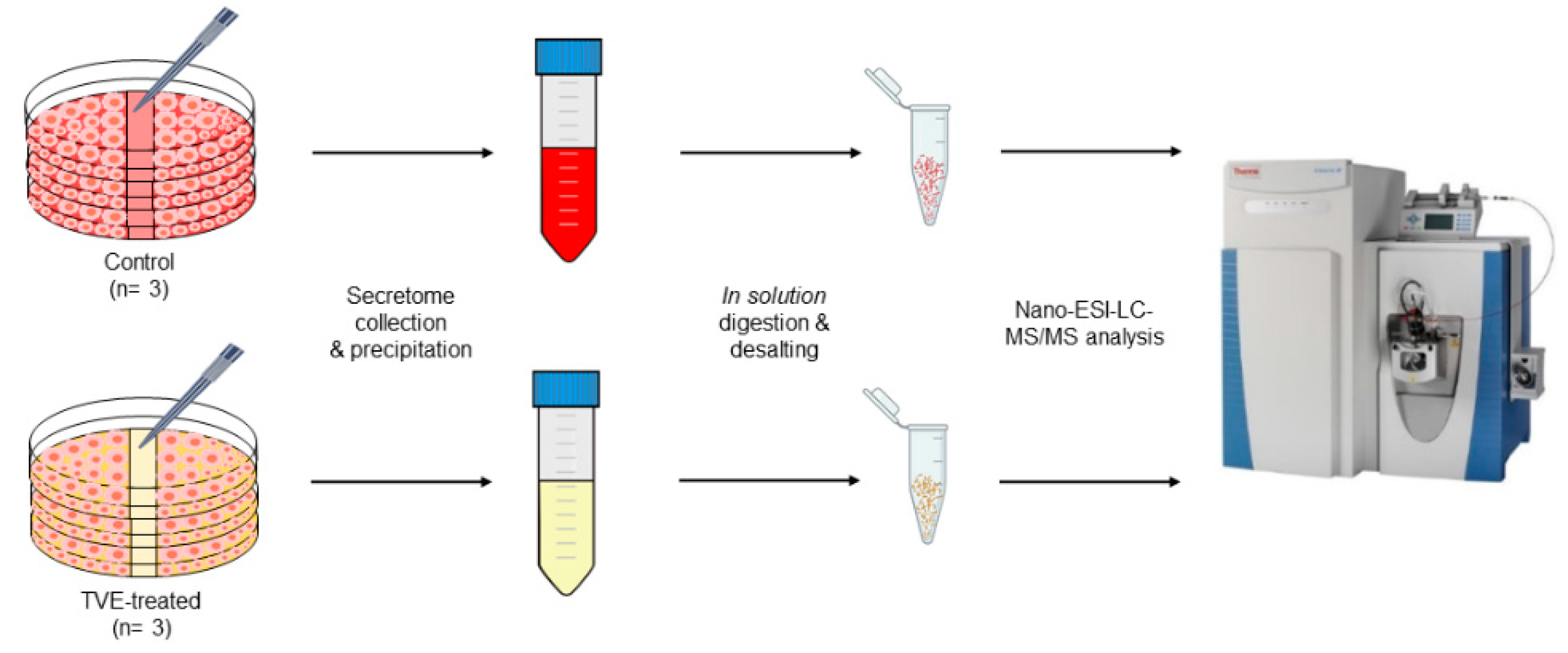
2.2. Proteome Profiling of HaCat Secretomes
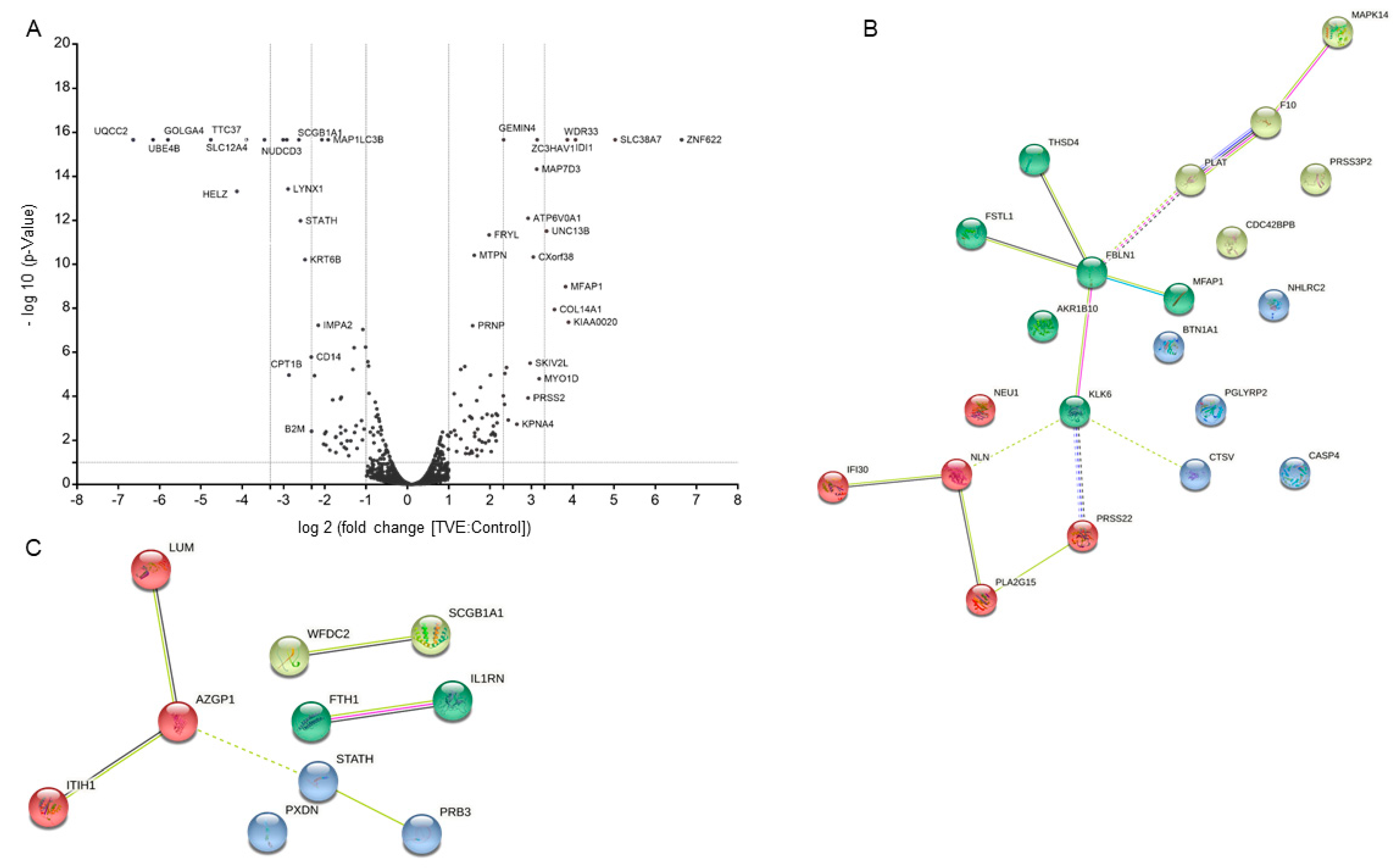
2.3. MS-Based Quantitative Proteomics to Disclose Secretome Content Alteration by TVE
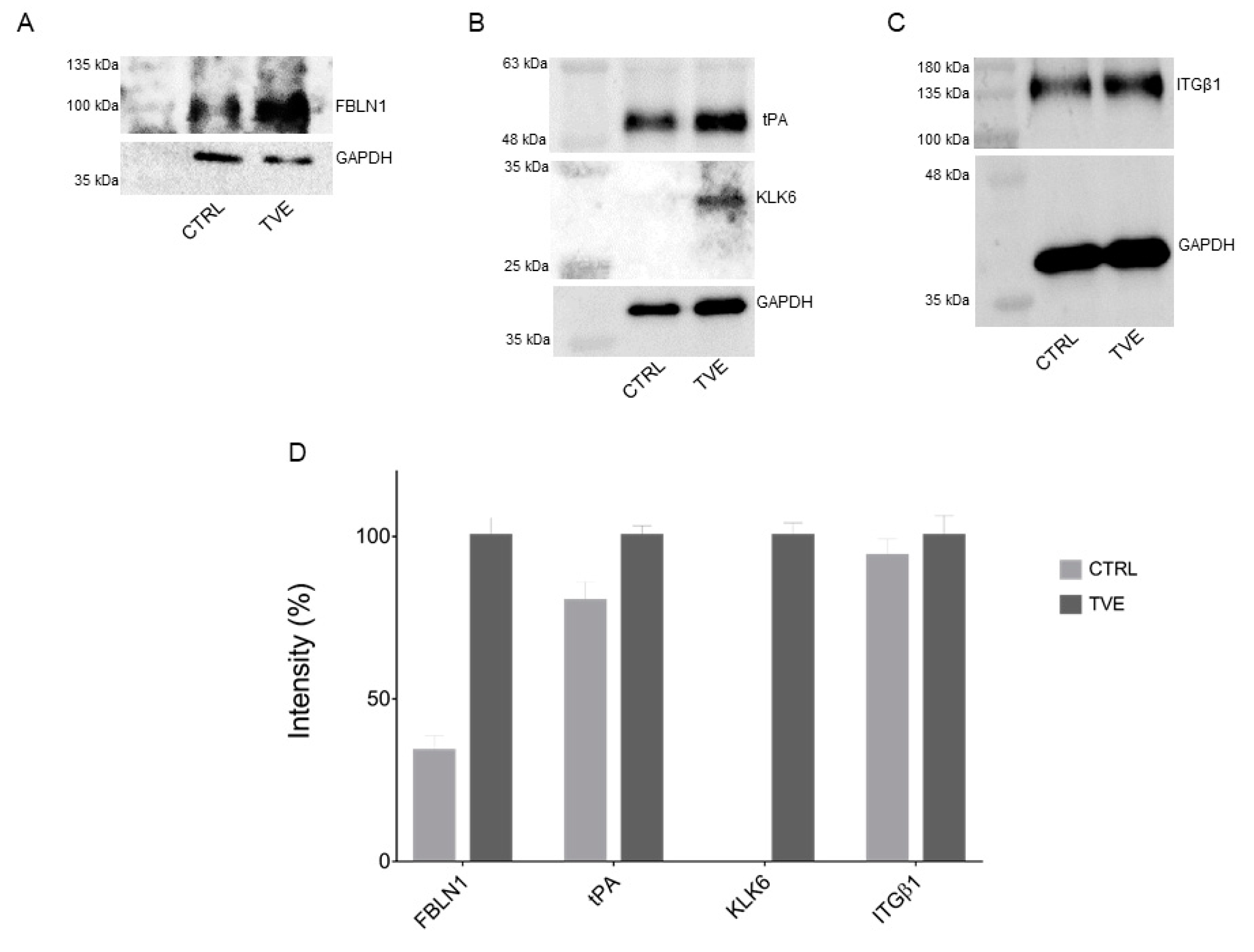
2.4. Validation through Immunoblotting Analysis
3. Discussion
4. Materials and Methods
4.1. Cell Viability Assay
4.2. Evaluation of TVE Effect on the Keratinocytes Monolayer Scratch Assay
4.3. HaCat Secretome Sample Preparation for Proteomics Analysis
4.4. Proteomics Analysis: Nano-UPLC-MS/MS
4.5. Data Processing through Proteome Discoverer
4.6. Data Visualization
4.7. Immunoblotting Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Sample Availability
References
- Kleinpenning, F.; Steigenberger, B.; Wu, W.; Heck, A.J.R. Fishing for newly synthesized proteins with phosphonate-handles. Nat. Commun. 2020, 11, 3244. [Google Scholar] [CrossRef] [PubMed]
- Tjalsma, H.; Bolhuis, A.; Jongbloed, J.D.H.; Bron, S.; van Dijl, J.M. Signal Peptide-Dependent Protein Transport in Bacillus subtilis: A Genome-Based Survey of the Secretome. Microbiol. Mol. Biol. Rev. 2000, 64, 515–547. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McCotter, S.W.; Horianopoulos, L.C.; Kronstad, J.W. Regulation of the fungal secretome. Curr. Genet. 2016, 62, 533–545. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, P.; Mani, S. Methodologies to decipher the cell secretome. Biochim. Biophys. Acta (BBA) Proteins Proteom. 2013, 1834, 2226–2232. [Google Scholar] [CrossRef] [Green Version]
- Ibrahim, N.; Wong, S.K.; Mohamed, I.N.; Mohamed, N.; Chin, K.-Y.; Ima-Nirwana, S.; Shuid, A.N. Wound Healing Properties of Selected Natural Products. Int. J. Environ. Res. Public Health 2018, 15, 2360. [Google Scholar] [CrossRef] [Green Version]
- Harper, D.; Young, A.; McNaught, C.-E. The physiology of wound healing. Surgery (Oxford) 2014, 32, 445–450. [Google Scholar] [CrossRef]
- Funel, N.; Dini, V.; Janowska, A.; Loggini, B.; Minale, M.; Grieco, F.; Riccio, S.; Romanelli, M. Triticum vulgare Extract Modulates Protein-Kinase B and Matrix Metalloproteinases 9 Protein Expression in BV-2 Cells: Bioactivity on Inflammatory Pathway Associated with Molecular Mechanism Wound Healing. Mediat. Inflamm. 2020, 2020, 2851949. [Google Scholar] [CrossRef]
- Romanelli, M.; Macchia, M.; Panduri, S.; Paggi, B.; Saponati, G.; Dini, V. Clinical evaluation of the efficacy and safety of a medical device in various forms containing Triticum vulgare for the treatment of venous leg ulcers—A randomized pilot study. Drug Des. Dev. Ther. 2015, 9, 2787–2792. [Google Scholar] [CrossRef] [Green Version]
- Sanguigno, L.; Minale, M.; Vannini, E.; Arato, G.; Riccio, R.; Casapullo, A.; Monti, M.C.; Riccio, R.; Formisano, S.; Di Renzo, G.; et al. Oligosaccharidic fractions derived from Triticum vulgare extract accelerate tissutal repairing processes in in vitro and in vivo models of skin lesions. J. Ethnopharmacol. 2015, 159, 198–208. [Google Scholar] [CrossRef] [Green Version]
- D’Agostino, A.; Pirozzi, A.V.A.; Finamore, R.; Grieco, F.; Minale, M.; Schiraldi, C. Molecular Mechanisms at the Basis of Pharmaceutical Grade Triticum vulgare Extract Efficacy in Prompting Keratinocytes Healing. Molecules 2020, 25, 431. [Google Scholar] [CrossRef] [Green Version]
- Dennisjr, G.; Sherman, B.T.; Hosack, D.A.; Yang, J.; Gao, W.; Lane, H.C.; Lempicki, R.A. DAVID: Database for Annotation, Visualization, and Integrated Discovery. Genome Biol. 2003, 4, R60. [Google Scholar] [CrossRef] [Green Version]
- Huang, D.W.; Sherman, B.T.; Tan, Q.; Kir, J.; Liu, D.; Bryant, D.; Guo, Y.; Stephens, R.; Baseler, M.W.; Lane, H.C.; et al. DAVID Bioinformatics Resources: Expanded annotation database and novel algorithms to better extract biology from large gene lists. Nucleic Acids Res. 2007, 35, W169–W175. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Morris, J.H.; Cook, H.; Kuhn, M.; Wyder, S.; Simonovic, M.; Santos, A.; Doncheva, N.T.; Roth, A.; Bork, P.; et al. The STRING database in 2017: Quality-controlled protein–protein association networks, made broadly accessible. Nucleic Acids Res. 2017, 45, D362–D368. [Google Scholar] [CrossRef]
- Belvedere, R.; Pessolano, E.; Novizio, N.; Tosco, A.; Eletto, D.; Porta, A.; Filippelli, A.; Petrella, F.; Petrella, A. The promising pro-healing role of the association of mesoglycan and lactoferrin on skin lesions. Eur. J. Pharm. Sci. 2021, 163, 105886. [Google Scholar] [CrossRef] [PubMed]
- Tan, I.; Lai, J.; Yong, J.; Li, S.F.; Leung, T. Chelerythrine perturbs lamellar actomyosin filaments by selective inhibition of myotonic dystrophy kinase-related Cdc42-binding kinase. FEBS Lett. 2011, 585, 1260–1268. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Heikkila, T.; Wheatley, E.; Crighton, D.; Schroder, E.; Boakes, A.; Kaye, S.J.; Mezna, M.; Pang, L.; Rushbrooke, M.; Turnbull, A.; et al. Co-Crystal Structures of Inhibitors with MRCKβ, a Key Regulator of Tumor Cell Invasion. PLoS ONE 2011, 6, e24825. [Google Scholar] [CrossRef] [Green Version]
- Mecham, R.P.; Gibson, M.A. The microfibril-associated glycoproteins (MAGPs) and the microfibrillar niche. Matrix Biol. 2015, 47, 13–33. [Google Scholar] [CrossRef]
- Rosenbloom, J.; Abrams, W.R.; Mecham, R. Extracellular matrix 4: The elastic fiber. FASEB J. 1993, 7, 1208–1218. [Google Scholar] [CrossRef] [Green Version]
- Wagenseil, J.E.; Mecham, R.P. New insights into elastic fiber assembly. Birth Defects Res. Part C Embryo Today Rev. 2007, 81, 229–240. [Google Scholar] [CrossRef]
- Ge, Q.; Chen, L.; Jaffar, J.; Argraves, W.S.; Twal, W.O.; Hansbro, P.; Black, J.L.; Burgess, J.; Oliver, B. Fibulin1C peptide induces cell attachment and extracellular matrix deposition in lung fibroblasts. Sci. Rep. 2015, 5, 9496. [Google Scholar] [CrossRef] [Green Version]
- Zhu, S.; Ye, L.; Bennett, S.; Xu, H.; He, D.; Xu, J. Molecular structure and function of microfibrillar-associated proteins in skeletal and metabolic disorders and cancers. J. Cell. Physiol. 2020, 236, 41–48. [Google Scholar] [CrossRef] [PubMed]
- Parfenova, O.K.; Kukes, V.G.; Grishin, D.V. Follistatin-Like Proteins: Structure, Functions and Biomedical Importance. Biomedicines 2021, 9, 999. [Google Scholar] [CrossRef]
- Nauroy, P.; Nyström, A. Kallikreins: Essential epidermal messengers for regulation of the skin microenvironment during homeostasis, repair and disease. Matrix Biol. Plus 2019, 6–7, 100019. [Google Scholar] [CrossRef] [PubMed]
- Klucky, B.; Mueller, R.; Vogt, I.; Teurich, S.; Hartenstein, B.; Breuhahn, K.; Flechtenmacher, C.; Angel, P.; Hess, J. Kallikrein 6 Induces E-Cadherin Shedding and Promotes Cell Proliferation, Migration, and Invasion. Cancer Res. 2007, 67, 8198–8206. [Google Scholar] [CrossRef] [Green Version]
- Bizzarro, V.; Belvedere, R.; Pessolano, E.; Parente, L.; Petrella, F.; Perretti, M.; Petrella, A. Mesoglycan induces keratinocyte activation by triggering syndecan-4 pathway and the formation of the annexin A1/S100A11 complex. J. Cell. Physiol. 2019, 234, 20174–20192. [Google Scholar] [CrossRef] [PubMed]
- Berdowska, I. Cysteine proteases as disease markers. Clin. Chim. Acta 2004, 342, 41–69. [Google Scholar] [CrossRef]
- Martin, P. Wound Healing—Aiming for Perfect Skin Regeneration. Science 1997, 276, 75–81. [Google Scholar] [CrossRef]
- Singer, A.J.; Clark, R.A. Cutaneous Wound Healing. N. Engl. J. Med. 1999, 341, 738–746. [Google Scholar] [CrossRef]
- Friedl, P. Prespecification and plasticity: Shifting mechanisms of cell migration. Curr. Opin. Cell Biol. 2004, 16, 14–23. [Google Scholar] [CrossRef]
- Everts, V.; Van Der Zee, E.; Creemers, L.; Beertsen, W. Phagocytosis and intracellular digestion of collagen, its role in turnover and remodelling. Histochem. J. 1996, 28, 229–245. [Google Scholar] [CrossRef]
- Murphy, G.; Gavrilovic, J. Proteolysis and cell migration: Creating a path? Curr. Opin. Cell Biol. 1999, 11, 614–621. [Google Scholar] [CrossRef]
- Wolf, K.; Friedl, P. Functional imaging of pericellular proteolysis in cancer cell invasion. Biochimie 2005, 87, 315–320. [Google Scholar] [CrossRef] [PubMed]
- Eeckhout, Y.; Vaes, G. Further studies on the activation of procollagenase, the latent precursor of bone collagenase. Effects of lysosomal cathepsin B, plasmin and kallikrein, and spontaneous activation. Biochem. J. 1977, 166, 21–31. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guinec, N.; Dalet-Fumeron, V.; Pagano, M. “In vitro” Study of Basement Membrane Degradation by the Cysteine Proteinases, Cathepsins B, B-Like and L. Digestion of Collagen IV, Laminin, Fibronectin, and Release of Gelatinase Activities front Basement Membrane Fibronectin. Biol. Chem. Hoppe-Seyler 1993, 374, 1135–1146. [Google Scholar] [CrossRef] [PubMed]
- Yadati, T.; Houben, T.; Bitorina, A.; Shiri-Sverdlov, R. The Ins and Outs of Cathepsins: Physiological Function and Role in Disease Management. Cells 2020, 9, 1679. [Google Scholar] [CrossRef]
- Rünger, T.M.; Quintanilla-Dieck, M.J.; Bhawan, J. Role of Cathepsin K in the Turnover of the Dermal Extracellular Matrix during Scar Formation. J. Investig. Dermatol. 2007, 127, 293–297. [Google Scholar] [CrossRef] [Green Version]
- Balce, D.R.; Allan, E.R.O.; McKenna, N.; Yates, R.M. γ-Interferon-inducible Lysosomal Thiol Reductase (GILT) Maintains Phagosomal Proteolysis in Alternatively Activated Macrophages. J. Biol. Chem. 2014, 289, 31891–31904. [Google Scholar] [CrossRef] [Green Version]
- Ghosh, A.K.; Vaughan, D.E. PAI-1 in tissue fibrosis. J. Cell. Physiol. 2011, 227, 493–507. [Google Scholar] [CrossRef] [Green Version]
- Simone, T.M.; Higgins, C.E.; Czekay, R.-P.; Law, B.K.; Higgins, S.P.; Archambeault, J.; Kutz, S.M.; Higgins, P.J. SERPINE1: A Molecular Switch in the Proliferation-Migration Dichotomy in Wound-“Activated” Keratinocytes. Adv. Wound Care 2014, 3, 281–290. [Google Scholar] [CrossRef] [Green Version]
- Fuchs, Y.; Steller, H. Programmed Cell Death in Animal Development and Disease. Cell 2011, 147, 742–758. [Google Scholar] [CrossRef] [Green Version]
- Liu, X.-J.; Kong, F.-Z.; Wang, Y.-H.; Zheng, J.-H.; Wan, W.-D.; Deng, C.-L.; Mao, G.-Y.; Li, J.; Yang, X.-M.; Zhang, Y.-L.; et al. Lumican Accelerates Wound Healing by Enhancing α2β1 Integrin-Mediated Fibroblast Contractility. PLoS ONE 2013, 8, e67124. [Google Scholar] [CrossRef] [PubMed]
- Corsuto, L.; Rother, S.; Koehler, L.; Bedini, E.; Moeller, S.; Schnabelrauch, M.; Hintze, V.; Schiraldi, C.; Scharnweber, D. Sulfation degree not origin of chondroitin sulfate derivatives modulates keratinocyte response. Carbohydr. Polym. 2018, 191, 53–64. [Google Scholar] [CrossRef] [PubMed]
- LoBello, N.; Biamonte, F.; Pisanu, M.E.; Faniello, M.C.; Jakopin, Ž.; Chiarella, E.; Giovannone, E.D.; Mancini, R.; Ciliberto, G.; Cuda, G.; et al. Ferritin heavy chain is a negative regulator of ovarian cancer stem cell expansion and epithelial to mesenchymal transition. Oncotarget 2016, 7, 62019–62033. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Loughner, C.L.; Bruford, E.; McAndrews, M.S.; Delp, E.E.; Swamynathan, S.; Swamynathan, S. Organization, evolution and functions of the human and mouse Ly6/uPAR family genes. Hum. Genom. 2016, 10, 10. [Google Scholar] [CrossRef] [PubMed] [Green Version]

| Accession | Gene Symbol | Description | Abundance Ratio | p-Value |
|---|---|---|---|---|
| O60218 | AKR1B10 | Aldo-keto reductase family 1-member B10 | 100.0000 | 0.0000 |
| Q9GZN4 | PRSS22 | Brain-specific serine protease 4 | 100.0000 | 0.0000 |
| P49662 | CASP4 | Caspase-4 | 100.0000 | 0.0000 |
| O60911 | CTSV | Cathepsin L2 | 100.0000 | 0.0000 |
| Q8NBF2 | NHLRC2 | NHL repeat-containing protein 2 | 100.0000 | 0.0000 |
| Q9Y5S2 | CDC42BPB | Serine/threonine-protein kinase MRCK beta | 100.0000 | 0.0000 |
| Q6ZMP0 | THSD4 | Thrombospondin-type-1 domain-containing protein 4 | 100.0000 | 0.0000 |
| P55081 | MFAP1 | Microfibrillar-associated protein 1 | 14.2020 | 0.0000 |
| A6XMV8 | PRSS2 | Protease serine 2 preproprotein | 7.5430 | 0.0001 |
| Q92876 | KLK6 | Kallikrein-6 | 5.4410 | 0.0012 |
| Q96PD5 | PGLYRP2 | N-acetylmuramoyl-l-alanine amidase | 4.9880 | 0.0001 |
| P23142 | FBLN1 | Fibulin-1 | 4.5230 | 0.0007 |
| Q99519 | NEU1 | Sialidase-1 | 4.5030 | 0.0016 |
| Q13410 | BTN1A1 | Butyrophilin subfamily 1 member A1 | 4.1290 | 0.0007 |
| Q12841 | FSTL1 | Follistatin-related protein 1 | 4.0930 | 0.0056 |
| B4E0K5 | MAPK14 | Mitogen-activated protein kinase | 3.1000 | 0.0629 |
| Q9BYT8 | NLN | Neurolysin, mitochondrial | 3.0870 | 0.0066 |
| Q8NCC3 | PLA2G15 | Phospholipase A2 group XV | 2.7950 | 0.0683 |
| P00742 | F10 | Coagulation factor X | 2.6100 | 0.0013 |
| P13284 | IFI30 | Gamma-interferon-inducible lysosomal thiol reductase | 2.3500 | 0.0778 |
| P00750 | PLAT | Tissue-type plasminogen activator | 2.2870 | 0.0317 |
| P19827 | ITIH1 | Inter-alpha-trypsin inhibitor heavy chain H1 | 0.4820 | 0.0030 |
| P02794 | FTH1 | Ferritin heavy chain | 0.4730 | 0.0000 |
| P25311 | AZGP1 | Zinc-alpha-2-glycoprotein | 0.4430 | 0.0637 |
| P51884 | LUM | Lumican | 0.4100 | 0.0000 |
| Q92626 | PXDN | Peroxidasin homolog | 0.4000 | 0.0000 |
| Q14508 | WFDC2 | WAP four-disulfide core domain protein 2 | 0.3870 | 0.0104 |
| P18510 | IL1RN | Interleukin-1 receptor antagonist protein | 0.3000 | 0.0073 |
| H0YLF3 | B2M | Beta-2-microglobulin (Fragment) | 0.2000 | 0.0038 |
| P02808 | STATH | Statherin | 0.1660 | 0.0000 |
| P11684 | SCGB1A1 | Uteroglobin | 0.1610 | 0.0000 |
| P0DP57 | SLURP2 | Secreted Ly-6/uPAR domain-containing protein 2 | 0.1350 | 0.0000 |
| Q04118 | PRB3 | Basic salivary proline-rich protein 3 | 0.0100 | 0.0000 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Morretta, E.; D’Agostino, A.; Cassese, E.; Maglione, B.; Petrella, A.; Schiraldi, C.; Monti, M.C. Label-Free Quantitative Proteomics to Explore the Action Mechanism of the Pharmaceutical-Grade Triticum vulgare Extract in Speeding Up Keratinocyte Healing. Molecules 2022, 27, 1108. https://doi.org/10.3390/molecules27031108
Morretta E, D’Agostino A, Cassese E, Maglione B, Petrella A, Schiraldi C, Monti MC. Label-Free Quantitative Proteomics to Explore the Action Mechanism of the Pharmaceutical-Grade Triticum vulgare Extract in Speeding Up Keratinocyte Healing. Molecules. 2022; 27(3):1108. https://doi.org/10.3390/molecules27031108
Chicago/Turabian StyleMorretta, Elva, Antonella D’Agostino, Elisabetta Cassese, Barbara Maglione, Antonello Petrella, Chiara Schiraldi, and Maria Chiara Monti. 2022. "Label-Free Quantitative Proteomics to Explore the Action Mechanism of the Pharmaceutical-Grade Triticum vulgare Extract in Speeding Up Keratinocyte Healing" Molecules 27, no. 3: 1108. https://doi.org/10.3390/molecules27031108
APA StyleMorretta, E., D’Agostino, A., Cassese, E., Maglione, B., Petrella, A., Schiraldi, C., & Monti, M. C. (2022). Label-Free Quantitative Proteomics to Explore the Action Mechanism of the Pharmaceutical-Grade Triticum vulgare Extract in Speeding Up Keratinocyte Healing. Molecules, 27(3), 1108. https://doi.org/10.3390/molecules27031108









