Gadolinium-Based Paramagnetic Relaxation Enhancement Agent Enhances Sensitivity for NUS Multidimensional NMR-Based Metabolomics
Abstract
1. Introduction
2. Results
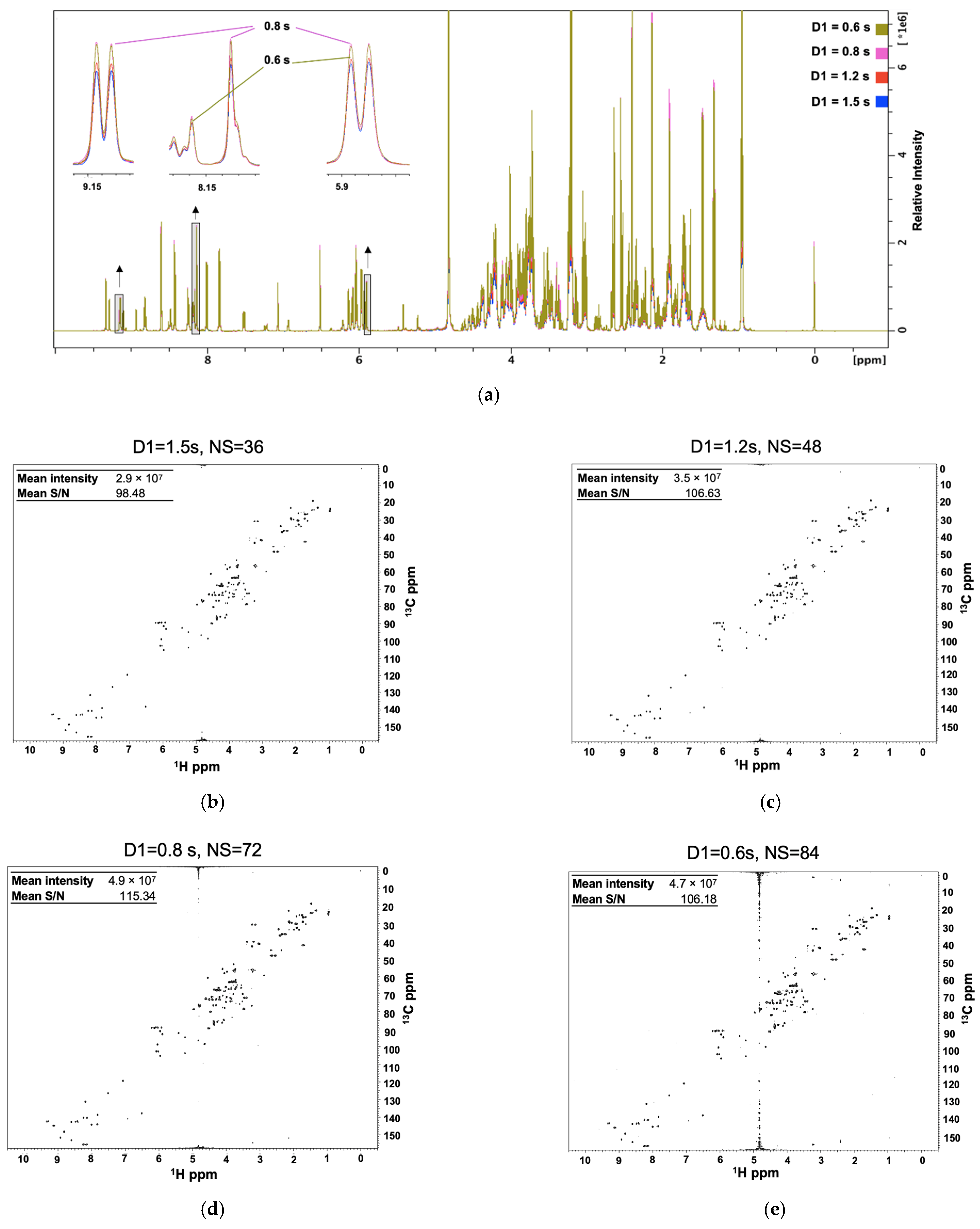
2.1. Optimizing the Relaxation Delay for Semi-Quantitative Metabolomics
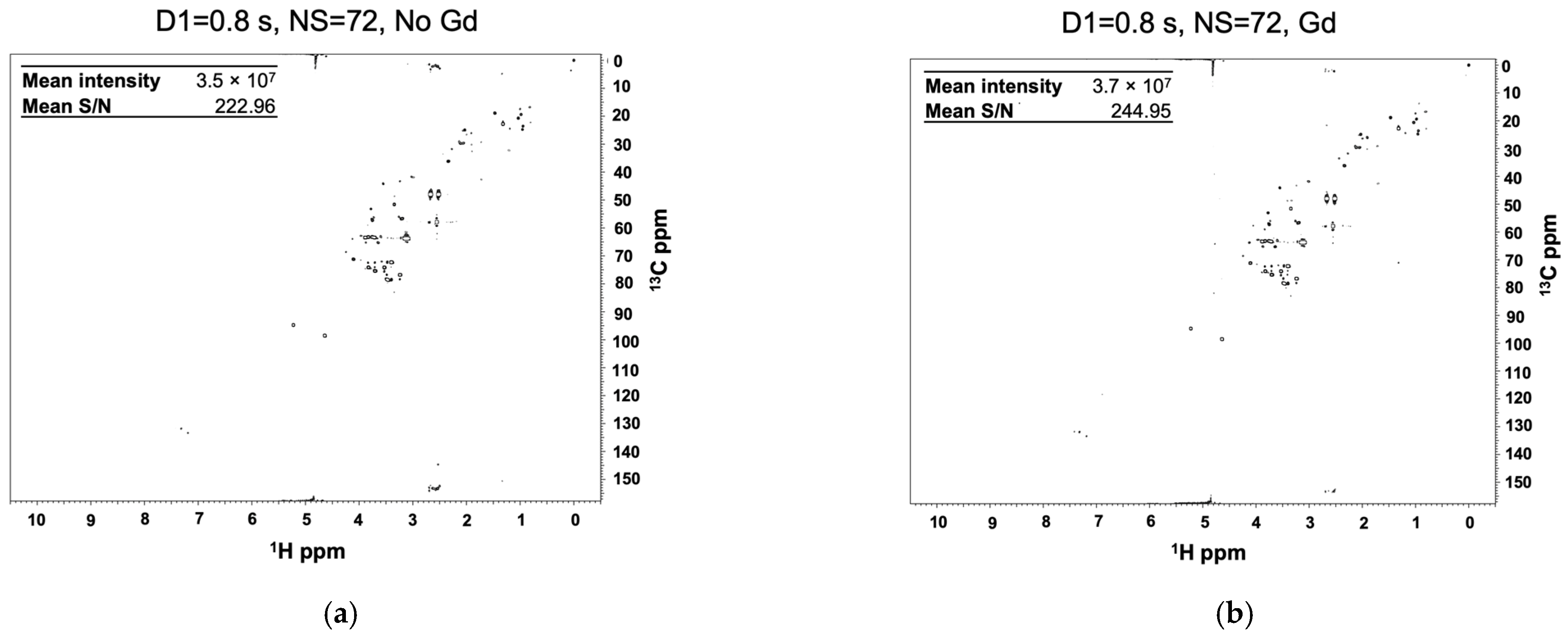
2.2. Gadolinium Provides Enhanced Sensitivity
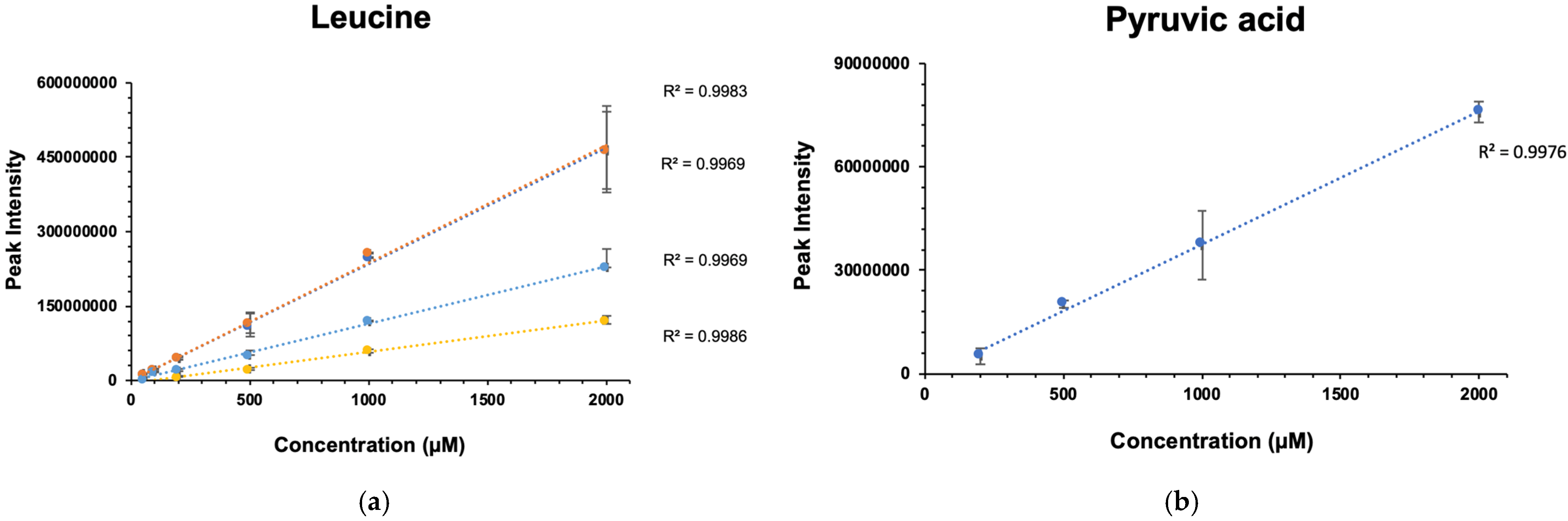
2.3. Gadolinium Maintains Linearity
2.4. Gadolinium Improves the Lower Limit of Detection and Quantification
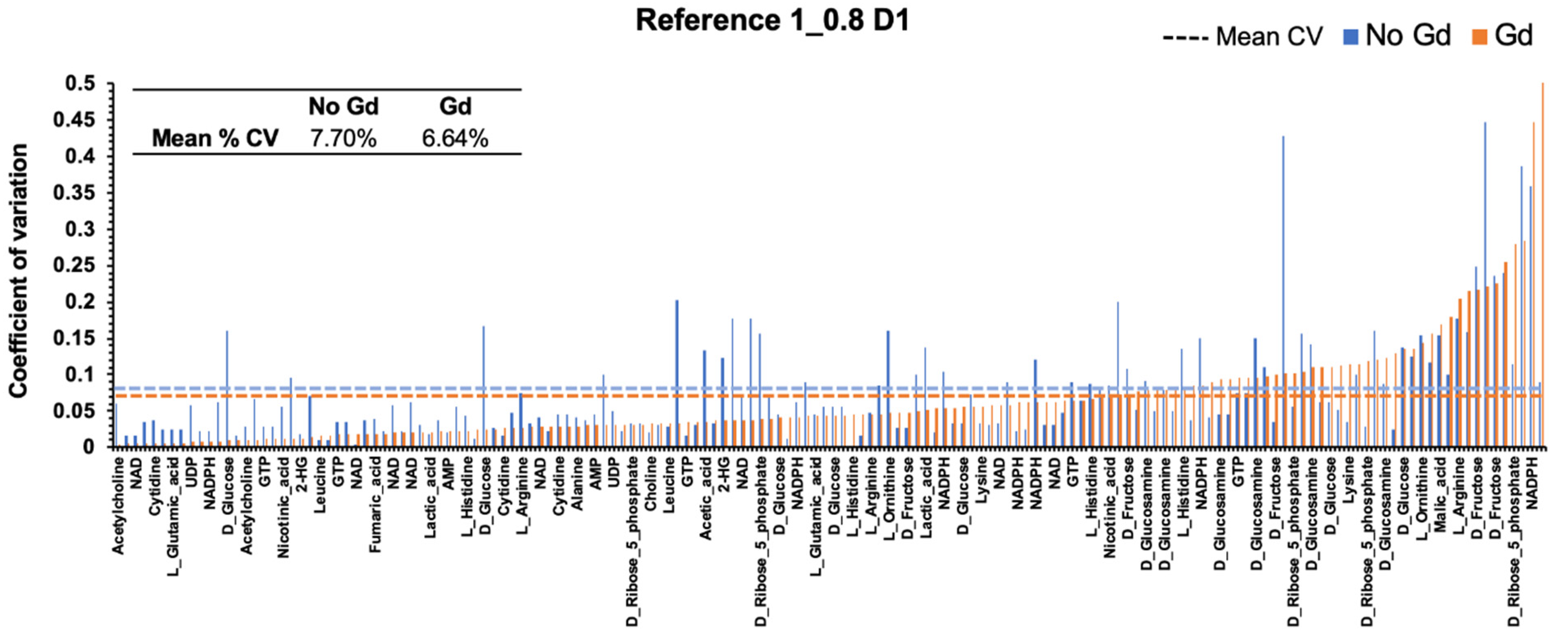
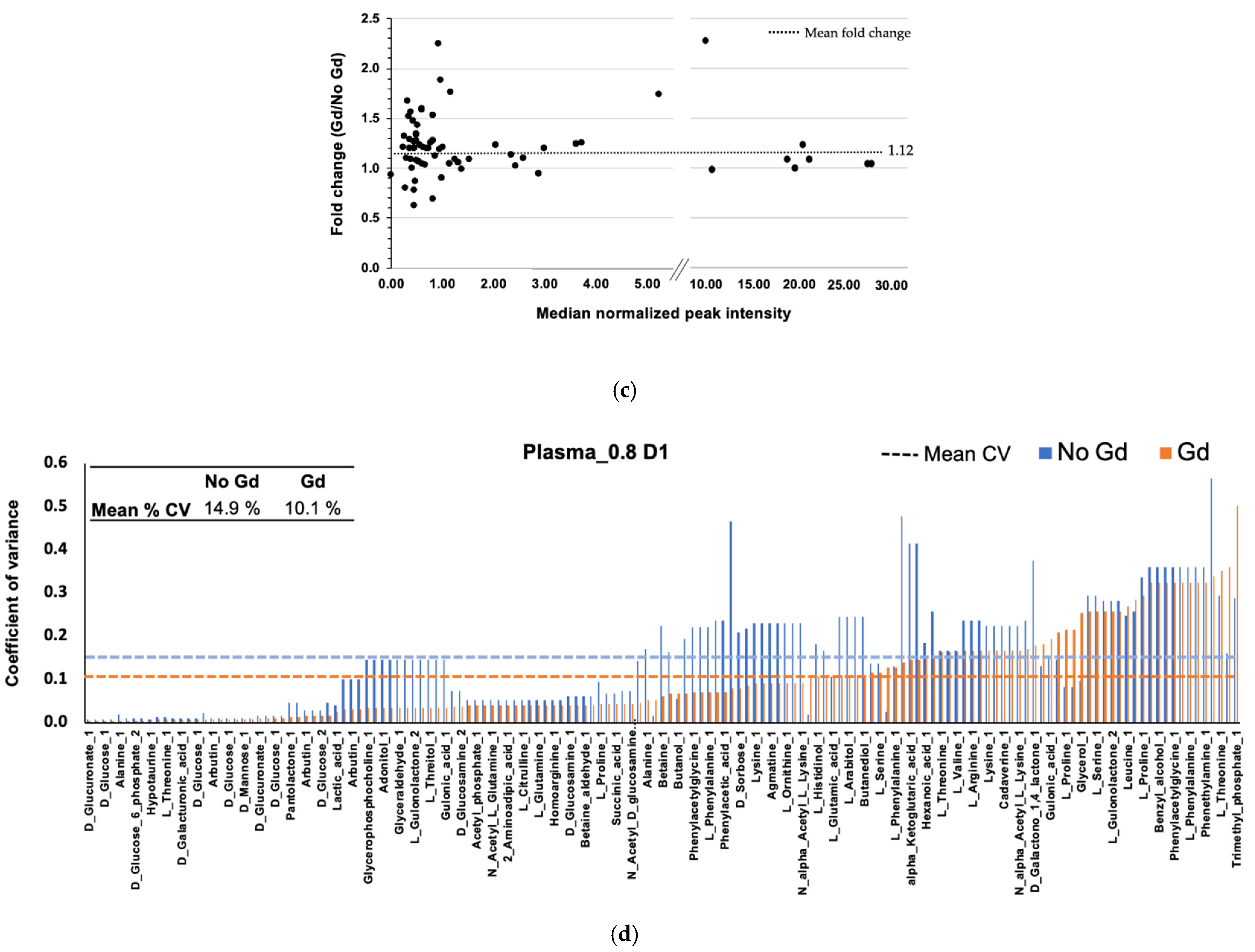
2.5. Gadolinium Maintains Reproducibility
2.6. Gadolinium Effect on Plasma Metabolites
3. Discussion
4. Materials and Methods
4.1. NMR Sample Preparation
4.2. NMR Experiments and Data Processing
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Sample Availability
References
- Riekeberg, E.; Powers, R. New frontiers in metabolomics: From measurement to insight. F1000Research 2017, 6, 1148. [Google Scholar] [CrossRef]
- Li, L.; Krznar, P.; Erban, A.; Agazzi, A.; Martin-Levilain, J.; Supale, S.; Kopka, J.; Zamboni, N.; Maechler, P. Metabolomics Identifies a Biomarker Revealing In Vivo Loss of Functional β-Cell Mass Before Diabetes Onset. Diabetes 2019, 68, 2272–2286. [Google Scholar] [CrossRef]
- Shao, Y.; Le, W. Recent advances and perspectives of metabolomics-based investigations in Parkinson’s disease. Mol. Neurodegener. 2019, 14, 1–12. [Google Scholar] [CrossRef]
- Puchades-Carrasco, L. Metabolomics Applications in Precision Medicine: An Oncological Perspective. Curr. Top. Med. Chem. 2017, 17, 2740–2751. [Google Scholar] [CrossRef]
- Cirulli, E.T.; Guo, L.; Swisher, C.L.; Shah, N.; Huang, L.; Napier, L.A.; Kirkness, E.F.; Spector, T.D.; Caskey, C.T.; Thorens, B.; et al. Profound Perturbation of the Metabolome in Obesity Is Associated with Health Risk. Cell Metab. 2019, 29, 488–500. [Google Scholar] [CrossRef]
- Ussher, J.R.; Elmariah, S.; Gerszten, R.E.; Dyck, J.R. The Emerging Role of Metabolomics in the Diagnosis and Prognosis of Cardiovascular Disease. J. Am. Coll. Cardiol. 2016, 68, 2850–2870. [Google Scholar] [CrossRef] [PubMed]
- Darst, B.F.; Koscik, R.L.; Hogan, K.J.; Johnson, S.C.; Engelman, C.D. Longitudinal plasma metabolomics of aging and sex. Aging 2019, 11, 1262–1282. [Google Scholar] [CrossRef] [PubMed]
- Bhinderwala, F.; Powers, R. NMR Metabolomics Protocols for Drug Discovery. In Cardiovascular Development; Springer Science and Business Media LLC: Berlin, Germany, 2019; Volume 2037, pp. 265–311. [Google Scholar]
- Markley, J.L.; Brüschweiler, R.; Edison, A.; Eghbalnia, H.R.; Powers, R.; Raftery, D.; Wishart, D.S. The future of NMR-based metabolomics. Curr. Opin. Biotechnol. 2017, 43, 34–40. [Google Scholar] [CrossRef] [PubMed]
- McAlpine, J.B.; Chen, S.-N.; Kutateladze, A.; MacMillan, J.B.; Appendino, G.; Barison, A.; Beniddir, M.A.; Biavatti, M.W.; Bluml, S.; Boufridi, A.; et al. The value of universally available raw NMR data for transparency, reproducibility, and integrity in natural product research. Nat. Prod. Rep. 2019, 36, 35–107. [Google Scholar] [CrossRef]
- Dona, A.; Kyriakides, M.; Scott, F.; Shephard, E.; Varshavi, D.; Veselkov, K.; Everett, J.R. A guide to the identification of metabolites in NMR-based metabonomics/metabolomics experiments. Comput. Struct. Biotechnol. J. 2016, 14, 135–153. [Google Scholar] [CrossRef] [PubMed]
- Bingol, K.; Bruschweiler-Li, L.; Li, D.-W.; Zhang, B.; Xie, M.; Brüschweiler, R. Emerging new strategies for successful metabolite identification in metabolomics. Bioanalysis 2016, 8, 557–573. [Google Scholar] [CrossRef]
- Rai, R.; Sinha, N. Fast and Accurate Quantitative Metabolic Profiling of Body Fluids by Nonlinear Sampling of1H–13C Two-Dimensional Nuclear Magnetic Resonance Spectroscopy. Anal. Chem. 2012, 84, 10005–10011. [Google Scholar] [CrossRef]
- Von Schlippenbach, T.; Oefner, P.J.; Gronwald, W. Systematic Evaluation of Non-Uniform Sampling Parameters in the Targeted Analysis of Urine Metabolites by 1H,1H 2D NMR Spectroscopy. Sci. Rep. 2018, 8, 1–10. [Google Scholar] [CrossRef]
- Zhang, B.; Powers, R.; O’Day, E.M. Evaluation of Non-Uniform Sampling 2D 1H–13C HSQC Spectra for Semi-Quantitative Metabolomics. Metabolites 2020, 10, 203. [Google Scholar] [CrossRef] [PubMed]
- Sørensen, O.W. James Keeler. Understanding NMR Spectroscopy. Magn. Reson. Chem. 2006, 44, 820. [Google Scholar] [CrossRef]
- Mulder, F.A.A.; Tenori, L.; Luchinat, C. Fast and Quantitative NMR Metabolite Analysis Afforded by a Paramagnetic Co-Solute. Angew. Chem. Int. Ed. 2019, 58, 15283–15286. [Google Scholar] [CrossRef]
- Cai, X.; Li, R. Concurrent profiling of polar metabolites and lipids in human plasma using HILIC-FTMS. Sci. Rep. 2016, 6, 36490. [Google Scholar] [CrossRef]
- Simón-Manso, Y.; Lowenthal, M.S.; Kilpatrick, L.E.; Sampson, M.L.; Telu, K.H.; Rudnick, P.A.; Mallard, W.G.; Bearden, D.W.; Schock, T.; Tchekhovskoi, D.V.; et al. Metabolite Profiling of a NIST Standard Reference Material for Human Plasma (SRM 1950): GC-MS, LC-MS, NMR, and Clinical Laboratory Analyses, Libraries, and Web-Based Resources. Anal. Chem. 2013, 85, 11725–11731. [Google Scholar] [CrossRef]
- Dunn, W.B.; Broadhurst, D.; Begley, P.; Zelena, E.; Francis-McIntyre, S.; Anderson, N.; Brown, M.; Knowles, J.D.; Halsall, A.; Haselden, J.N.; et al. Procedures for large-scale metabolic profiling of serum and plasma using gas chromatography and liquid chromatography coupled to mass spectrometry. Nat. Protoc. 2011, 6, 1060–1083. [Google Scholar] [CrossRef] [PubMed]
- Telu, K.H.; Yan, X.; Wallace, W.E.; Stein, S.E.; Simón-Manso, Y. Analysis of human plasma metabolites across different liquid chromatography/mass spectrometry platforms: Cross-platform transferable chemical signatures. Rapid Commun. Mass Spectrom. 2016, 30, 581–593. [Google Scholar] [CrossRef] [PubMed]
- Murphy, P.S.; Leach, M.; Rowland, I.J. Signal modulation in1H magnetic resonance spectroscopy using contrast agents: Proton relaxivities of choline, creatine, andN-acetylaspartate. Magn. Reson. Med. 1999, 42, 1155–1158. [Google Scholar] [CrossRef]
- Valkovič, L.; Lau, J.Y.C.; Abdesselam, I.; Rider, O.J.; Frollo, I.; Tyler, D.J.; Rodgers, C.T.; Miller, J.J.J. Effects of contrast agents on relaxation properties of 31 P metabolites. Magn. Reson. Med. 2021, 85, 1805–1813. [Google Scholar] [CrossRef]
- Botta, M.; Aime, S.; Barge, A.; Bobba, G.; Dickins, R.S.; Parker, D.; Terreno, E. Ternary Complexes between Cationic GdIII Chelates and Anionic Metabolites in Aqueous Solution: An NMR Relaxometric Study. Chem. A Eur. J. 2003, 9, 2102–2109. [Google Scholar] [CrossRef] [PubMed]
- Scott, L.J. Gadobutrol: A Review in Contrast-Enhanced MRI and MRA. Clin. Drug Investig. 2018, 38, 773–784. [Google Scholar] [CrossRef]
- Clore, G.M.; Iwahara, J. Theory, Practice, and Applications of Paramagnetic Relaxation Enhancement for the Characterization of Transient Low-Population States of Biological Macromolecules and Their Complexes. Chem. Rev. 2009, 109, 4108–4139. [Google Scholar] [CrossRef] [PubMed]
- Giannoulis, A.; Ben-Ishay, Y.; Goldfarb, D. Characteristics of Gd(III) Spin Labels for the Study of Protein Conformations; Elsevier BV: Amsterdam, The Netherlands, 2021; Volume 651, pp. 235–290. [Google Scholar]
- Sakol, N.; Egawa, A.; Fujiwara, T. Gadolinium Complexes as Contrast Agent for Cellular NMR Spectroscopy. Int. J. Mol. Sci. 2020, 21, 4042. [Google Scholar] [CrossRef]
- Tang, H.; Wang, Y.; Nicholson, J.; Lindon, J. Use of relaxation-edited one-dimensional and two dimensional nuclear magnetic resonance spectroscopy to improve detection of small metabolites in blood plasma. Anal. Biochem. 2004, 325, 260–272. [Google Scholar] [CrossRef] [PubMed]
- Shahbazi-Gahrouei, D.; Williams, M.; Allen, B.J. In vitro study of relationship between signal intensity and gadolinium-DTPA concentration at high magnetic field strength. Australas. Radiol. 2001, 45, 298–304. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.-J.; Kim, M.-J.; Yoon, C.-S.; Song, S.Y.; Park, K.; Kim, W.S. The T2-Shortening Effect of Gadolinium and the Optimal Conditions for Maximizing the CNR for Evaluating the Biliary System: A Phantom Study. Korean J. Radiol. 2011, 12, 358–364. [Google Scholar] [CrossRef]
- Rovnyak, D.; Hoch, J.; Stern, A.; Wagner, G. Resolution and sensitivity of high field nuclear magnetic resonance spectroscopy. J. Biomol. NMR 2004, 30, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Patterson, R.E.; Ducrocq, A.J.; McDougall, D.J.; Garrett, T.J.; Yost, R.A. Comparison of blood plasma sample preparation methods for combined LC–MS lipidomics and metabolomics. J. Chromatogr. B 2015, 1002, 260–266. [Google Scholar] [CrossRef] [PubMed]
- Krug, S.; Kastenmüller, G.; Stückler, F.; Rist, M.J.; Skurk, T.; Sailer, M.; Raffler, J.; Römisch-Margl, W.; Adamski, J.; Prehn, C.; et al. The dynamic range of the human metabolome revealed by challenges. FASEB J. 2012, 26, 2607–2619. [Google Scholar] [CrossRef] [PubMed]
- Bingol, K.; Li, D.-W.; Zhang, B.; Brüschweiler, R. Comprehensive Metabolite Identification Strategy Using Multiple Two-Dimensional NMR Spectra of a Complex Mixture Implemented in the COLMARm Web Server. Anal. Chem. 2016, 88, 12411–12418. [Google Scholar] [CrossRef]
- Johnson, C.; Ivanisevic, J.; Siuzdak, G. Metabolomics: Beyond biomarkers and towards mechanisms. Nat. Rev. Mol. Cell Biol. 2016, 17, 451–459. [Google Scholar] [CrossRef] [PubMed]
- Clish, C.B. Metabolomics: An emerging but powerful tool for precision medicine. Mol. Case Stud. 2015, 1, a000588. [Google Scholar] [CrossRef]
- Fiehn, O. Metabolomics—The link between genotypes and phenotypes. Plant Mol. Biol. 2002, 48, 155–171. [Google Scholar] [CrossRef] [PubMed]
- Hameed, I.; Masoodi, S.R.; Mir, S.A.; Nabi, M.; Ghazanfar, K.; A Ganai, B. Type 2 diabetes mellitus: From a metabolic disorder to an inflammatory condition. World J. Diabetes 2015, 6, 598–612. [Google Scholar] [CrossRef]
- Procaccini, C.; Santopaolo, M.; Faicchia, D.; Colamatteo, A.; Formisano, L.; de Candia, P.; Galgani, M.; De Rosa, V.; Matarese, G. Role of metabolism in neurodegenerative disorders. Metab. Clin. Exp. 2016, 65, 1376–1390. [Google Scholar] [CrossRef]
- Zhang, B.; Warner, J.; Pinto, C.; Juric, D.; ODay, E. NMR-metabolite-resonance signature to predict HR+ breast cancer patient response to CDK4/6 inhibitors. J. Clin. Oncol. 2019, 37. [Google Scholar] [CrossRef]
- O’Day, E.; Leitzel, K.; Ali, S.M.; Zhang, B.; Dong, C.; Gu, H.; Shi, X.; Drabick, J.J.; Cream, L.; Vasekar, M.; et al. Abstract P4-10-25: Pretreatment serum metabolome predicts PFS in first-line trastuzumab-treated metastatic breast cancer. Poster Sess. Abstr. 2020, 80, 4–10. [Google Scholar]
- Honrao, C.; Rao, S.R.; Teissier, N.; Call, S.G.; Oday, E.M.; Janku, F. Abstract LB031: Plasma based metabolic profiling in metastatic gastrointestinal stromal tumors (GIST). Clin. Res. 2021, 81, LB031. [Google Scholar] [CrossRef]
- Gong, L.-L.; Yang, S.; Zhang, W.; Han, F.-F.; Xuan, L.-L.; Lv, Y.-L.; Liu, H.; Liu, L.-H. Targeted Metabolomics for Plasma Amino Acids and Carnitines in Patients with Metabolic Syndrome Using HPLC-MS/MS. Dis. Markers 2020, 2020, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Tomita, R.; Todoroki, K.; Maruoka, H.; Yoshida, H.; Fujioka, T.; Nakashima, M.; Yamaguchi, M.; Nohta, H. Amino Acid Metabolomics Using LC-MS/MS: Assessment of Cancer-Cell Resistance in a Simulated Tumor Microenvironment. Anal. Sci. 2016, 32, 893–900. [Google Scholar] [CrossRef][Green Version]
- Ni, J.; Xu, L.; Li, W.; Wu, L. Simultaneous determination of thirteen kinds of amino acid and eight kinds of acylcarnitine in human serum by LC-MS/MS and its application to measure the serum concentration of lung cancer patients. Biomed. Chromatogr. 2016, 30, 1796–1806. [Google Scholar] [CrossRef] [PubMed]
- Schätzlein, M.P.; Becker, J.; Schulze-Sünninghausen, D.; Pineda-Lucena, A.; Herance, J.R.; Luy, B. Rapid two-dimensional ALSOFAST-HSQC experiment for metabolomics and fluxomics studies: Application to a 13C-enriched cancer cell model treated with gold nanoparticles. Anal. Bioanal. Chem. 2018, 410, 2793–2804. [Google Scholar] [CrossRef]
- Clore, G.M.; Tang, C.; Iwahara, J. Elucidating transient macromolecular interactions using paramagnetic relaxation enhancement. Curr. Opin. Struct. Biol. 2007, 17, 603–616. [Google Scholar] [CrossRef] [PubMed]
- Clore, G.M. Practical Aspects of Paramagnetic Relaxation Enhancement in Biological Macromolecules. In Biofilms; Elsevier BV: Amsterdam, The Netherlands, 2015; Volume 564, pp. 485–497. [Google Scholar]
- Xiao, Y.-D.; Paudel, R.; Liu, J.; Ma, C.; Zhang, Z.-S.; Zhou, S.-K. MRI contrast agents: Classification and application (Review). Int. J. Mol. Med. 2016, 38, 1319–1326. [Google Scholar] [CrossRef]
- Hyberts, S.G.; Milbradt, A.G.; Wagner, A.B.; Arthanari, H.; Wagner, G. Application of iterative soft thresholding for fast reconstruction of NMR data non-uniformly sampled with multidimensional Poisson Gap scheduling. J. Biomol. NMR 2012, 52, 315–327. [Google Scholar] [CrossRef]
- Delaglio, F.; Grzesiek, S.; Vuister, G.W.; Zhu, G.; Pfeifer, J.; Bax, A. NMRPipe: A multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 1995, 6, 277–293. [Google Scholar] [CrossRef]

| Metabolites/R2 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NAD | 0.996 | 0.999 | 0.999 | 0.997 | 1.000 | 0.999 | 0.998 | 0.999 | 1.000 | 0.999 | 1.000 | 0.999 | 0.999 |
| NADPH | 1.000 | 1.000 | 0.886 | 0.929 | 0.995 | 0.966 | 0.967 | 0.966 | 0.972 | 0.968 | 0.950 | ||
| Cytidine | 0.912 | 0.999 | 0.997 | 0.998 | 0.997 | 0.998 | 0.998 | ||||||
| UDP | 0.999 | 0.999 | 0.999 | 0.999 | 0.999 | 0.999 | 1.000 | ||||||
| Fructose | 1.000 | 1.000 | 1.000 | 1.000 | 0.984 | 0.996 | |||||||
| AMP | 0.951 | 0.973 | 0.970 | 0.992 | 0.901 | 0.994 | |||||||
| Lysine | 0.998 | 0.999 | 0.999 | 0.998 | 0.942 | ||||||||
| Histidine | 0.998 | 1.000 | 0.998 | 0.999 | 0.998 | ||||||||
| Glucose | 0.995 | 0.997 | 0.993 | 0.996 | 0.974 | ||||||||
| Ribose 5P | 0.983 | 0.990 | 0.989 | 0.996 | 0.994 | ||||||||
| Glucosamine | 0.997 | 0.959 | 0.992 | 0.951 | |||||||||
| 2-HG | 0.994 | 0.998 | 0.999 | 0.984 | |||||||||
| Leucine | 0.998 | 0.997 | 0.999 | 0.997 | |||||||||
| Nicotinic acid | 0.998 | 0.996 | 0.994 | 0.997 | |||||||||
| Acetylcholine | 0.999 | 0.989 | 0.998 | 0.998 | |||||||||
| Glutamic acid | 0.998 | 0.998 | 0.952 | ||||||||||
| Malic acid | 0.993 | 0.989 | 0.996 | ||||||||||
| Arginine | 0.991 | 0.997 | 0.999 | ||||||||||
| Ornithine | 0.998 | 0.996 | 1.000 | ||||||||||
| Choline | 0.999 | 0.997 | 0.996 | ||||||||||
| Glutamine | 0.998 | 0.998 | 0.998 | ||||||||||
| GTP | 0.998 | 0.998 | 0.967 | ||||||||||
| Citrate | 0.990 | 0.998 | |||||||||||
| Alanine | 0.998 | 0.995 | |||||||||||
| Lactic acid | 0.999 | 0.997 | |||||||||||
| Pyruvic acid | 0.998 | ||||||||||||
| Acetic acid | 0.990 | ||||||||||||
| Fumaric acid | 0.989 | ||||||||||||
| Succinic acid | 0.998 |
| LOD (µM) per NMR Resonance | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Metabolites | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | Minimal Conc. (µM) |
| NAD | 12.7 ± 0.49 | 12.58 ± 0.26 | 26.05 ± 0.32 | 12.2 ± 0.04 | 10.61 ± 0.19 | 15.54 ± 0.97 | 7.9 ± 0.08 | 13.48 ± 0.15 | 16.5 ± 0.04 | 14.41 ± 0.11 | 12.98 ± 0.16 | 9.2 ± 0.09 | 12.9 ± 0.02 | 7.90 ± 0.08 |
| NADPH | 26.85 ± 0.21 | 21.76 ± 0.99 | 7.46 ± 0.17 | 19.01 ± 0.38 | 13.4 ± 0.48 | 10.66 ± 0.42 | 14.4 ± 0.14 | 25.37 ± 0.5 | 19.46 ± 0.96 | 12.98 ± 0.27 | 24.4 ± 0.62 | 7.46 ± 0.17 | ||
| Cytidine | 18.1 ± 0.66 | 8.44 ± 0.02 | 9.04 ± 0.08 | 7.4 ± 0.00 | 6.62 ± 0.05 | 7.86 ± 0.06 | 7.89 ± 0.05 | 6.62 ± 0.05 | ||||||
| UDP | 6.78 ± 0.09 | 9.47 ± 0.15 | 9.82 ± 0.2 | 9.85 ± 0.01 | 8.69 ± 0.11 | 10.65 ± 0.15 | 10.69 ± 0.05 | 6.78 ± 0.09 | ||||||
| Fructose | 19.58 ± 0.93 | 13.82 ± 0.45 | 13.85 ± 0.44 | 18.89 ± 1.17 | 18.71 ± 0.33 | 9.49 ± 0.09 | 9.49 ± 0.09 | |||||||
| AMP | 8.35 ± 1.68 | 12.14 ± 1.54 | 3.94 ± 0.16 | 13.17 ± 2.97 | 10.75 ± 1.7 | 14.96 ± 2.96 | 3.94 ± 0.16 | |||||||
| Lysine | 23.24 ± 0.19 | 10.07 ± 0.24 | 9.28 ± 0.13 | 6.59 ± 0.03 | 3.6 ± 0.1 | 3.60 ± 0.10 | ||||||||
| Histidine | 19.15 ± 2.19 | 18.79 ± 1.91 | 10.85 ± 1.66 | 7.9 ± 0.38 | 11.3 ± 2.09 | 7.90 ± 0.38 | ||||||||
| Glucose | 21.62 ± 3.18 | 18.61 ± 2.01 | 16.35 ± 1.1 | 14.32 ± 0.11 | 15.79 ± 2.56 | 14.32 ± 0.11 | ||||||||
| Ribose 5P | 41.21 ± 2.58 | 30.31 ± 1.82 | 21.22 ± 0.59 | 8.98 ± 0.26 | 19.94 ± 0.71 | 8.98 ± 0.26 | ||||||||
| Glucosamine | 30.31 ± 1.11 | 19.2 ± 0.570.57 | 33.83 ± 1.51 | 22.32 ± 1.71 | 19.20 ± 0.57 | |||||||||
| 2-HG | 33.83 ± 1.76 | 33.3 ± 1.89 | 25.95 ± 0.75 | 12.7 ± 0.3 | 12.70 ± 0.3 | |||||||||
| Leucine | 5.64 ± 0.98 | 5.58 ± 1.07 | 20.77 ± 1.54 | 11.49 ± 1.85 | 5.58 ± 1.07 | |||||||||
| Nicotinic acid | 10.78 ± 0.03 | 9.71 ± 0.19 | 8.04 ± 0.16 | 7.26 ± 0.01 | 7.26 ± 0.01 | |||||||||
| Acetylcholine | 8.56 ± 0.12 | 0.71 ± 0.02 | 8.42 ± 0.1 | 8.06 ± 0.07 | 0.71 ± 0.02 | |||||||||
| Glutamic acid | 22.1± 1.1 | 8.87 ± 0.13 | 4.09 ± 0.08 | 4.09 ± 0.08 | ||||||||||
| Malic acid | 30.52 ± 2.05 | 30.12 ± 0.41 | 8.98 ± 0.26 | 8.98 ± 0.26 | ||||||||||
| Arginine | 25.49 ± 2.75 | 9.06 ± 0.09 | 6.09 ± 0.06 | 6.09 ± 0.06 | ||||||||||
| Ornithine | 28.27 ± 1.77 | 4.59 ± 0.05 | 6.71 ± 0.05 | 4.59 ± 0.05 | ||||||||||
| Choline | 1.41 ± 0.05 | 8.09 ± 0.3 | 7.76 ± 0.29 | 1.41 ± 0.05 | ||||||||||
| Glutamine | 8.32 ± 0.13 | 9.46 ± 0.22 | 5.06 ± 0.07 | 5.06 ± 0.07 | ||||||||||
| GTP | 11.66 ± 0.18 | 15.58 ± 0.51 | 7.54 ± 1.44 | 7.54 ± 1.44 | ||||||||||
| Citrate | 8.2 ± 1.11 | 8.15 ± 1.21 | 5.14 ± 0.01 | |||||||||||
| Alanine | 5.14 ± 0.01 | 13.4 ± 0.08 | 8.15 ± 1.21 | |||||||||||
| Lactic acid | 5.4 ± 0.02 | 13.89 ± 0.22 | 5.4 ± 0.02 | |||||||||||
| Pyruvic acid | 30.79± 2.83 | 30.79 ± 2.83 | ||||||||||||
| Acetic acid | 7.21 ± 0.08 | 7.21 ± 0.08 | ||||||||||||
| Fumaric acid | 5.6 ± 0.14 | 5.67 ± 0.12 | ||||||||||||
| Succinic acid | 3.21 ± 0.03 | 3.21 ± 0.03 | ||||||||||||
| LOQ (μM) per NMR Resonance | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Metabolites | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | Minimal Conc. (µM) |
| NAD | 42.34 ± 1.62 | 41.92 ± 0.88 | 86.83 ± 1.06 | 40.66 ± 0.13 | 35.37 ± 0.62 | 51.78 ± 3.22 | 26.35 ± 0.27 | 44.92 ± 0.51 | 55.01 ± 0.14 | 48.03 ± 0.38 | 43.26 ± 0.55 | 30.66 ± 0.3 | 43.01 ± 0.06 | 26.35 ± 0.27 |
| NADPH | 89.49 ± 0.7 | 72.53 ± 3.29 | 24.87 ± 0.55 | 63.38 ± 1.25 | 44.66 ± 1.61 | 35.52 ± 1.39 | 48.01 ± 0.47 | 84.56 ± 1.67 | 64.88 ± 3.19 | 43.27 ± 0.91 | 81.33 ± 2.07 | 35.52 ± 1.39 | ||
| Cytidine | 60.34 ± 2.2 | 28.12 ± 0.08 | 30.15 ± 0.28 | 24.68 ± 0.01 | 22.05 ± 0.18 | 26.2 ± 0.19 | 26.29 ± 0.15 | 22.05 ± 0.18 | ||||||
| UDP | 22.59 ± 0.29 | 31.55 ±0.51 | 32.74 ± 0.68 | 32.85 ± 0.03 | 28.95 ± 0.36 | 35.49 ± 0.49 | 35.64 ± 0.15 | 22.59 ± 0.29 | ||||||
| Fructose | 65.27 ± 3.1 | 46.06 ±1.5 | 46.18 ± 1.47 | 62.98 ± 3.89 | 62.38 ± 1.09 | 31.63 ± 0.29 | 31.63 ± 0.29 | |||||||
| AMP | 27.83 ± 5.61 | 40.46 ± 5.12 | 13.13 ± 0.52 | 43.92 ± 9.89 | 35.84 ± 5.68 | 49.86 ± 9.88 | 13.13 ± 0.52 | |||||||
| Lysine | 77.47 ± 0.64 | 33.57 ± 0.8 | 30.92 ± 0.44 | 21.95 ± 0.08 | 12.01 ± 0.34 | 12.01 ± 0.34 | ||||||||
| Histidine | 63.83 ± 7.29 | 62.63 ± 6.38 | 36.18 ± 5.53 | 26.32 ± 1.25 | 37.67 ± 6.97 | 26.32 ± 1.25 | ||||||||
| Glucose | 72.07 ±10.59 | 62.02 ± 6.69 | 54.5 ± 3.66 | 47.74 ± 0.36 | 52.63 ± 8.54 | 47.74 ± 0.36 | ||||||||
| Ribose 5P | 137.37 ± 8.59 | 101.02 ± 6.08 | 70.73 ± 1.97 | 30.11 ± 0.89 | 66.45 ± 2.38 | 30.11 ± 0.89 | ||||||||
| Glucosamine | 101.02 ± 3.71 | 42.87 ± 2.83 | 112.76 ± 5.02 | 74.4 ± 5.69 | 42.87 ± 2.83 | |||||||||
| 2-HG | 112.76 ± 5.86 | 110.99 ± 6.3 | 86.49 ± 2.51 | 42.34 ±1.00 | 42.34 ± 1.00 | |||||||||
| Leucine | 18.78 ± 3.27 | 18.61 ± 3.58 | 69.24 ± 5.13 | 38.28 ± 6.16 | 18.61 ± 3.58 | |||||||||
| Nicotinic acid | 35.93 ± 0.1 | 32.36 ± 0.62 | 26.8 ± 0.54 | 24.19 ± 0.04 | 24.19 ± 0.04 | |||||||||
| Acetylcholine | 28.54 ± 0.4 | 2.38 ± 0.08 | 28.08 + BL86 ± 0.33 | 26.88 ± 0.24 | 2.38 ± 0.08 | |||||||||
| Glutamic acid | 73.67 ± 3.65 | 29.55 ± 0.44 | 13.64 ± 0.28 | 13.64 ± 0.28 | ||||||||||
| Malic acid | 101.74 ± 6.84 | 100.41 ± 1.36 | 29.92 ± 0.88 | 29.92 ± 0.88 | ||||||||||
| Arginine | 84.96 ± 9.16 | 30.2 ± 0.3 | 20.32 ± 0.19 | 20.32 ± 0.19 | ||||||||||
| Ornithine | 94.24 ± 5.91 | 15.3 ± 0.16 | 22.35 ± 0.18 | 15.30 ± 0.16 | ||||||||||
| Choline | 4.69 ± 0.15 | 26.98 ± 0.99 | 25.85 ± 0.97 | 4.69 ± 0.15 | ||||||||||
| Glutamine | 27.72 ± 0.44 | 31.54 ± 0.73 | 16.85 ± 0.24 | 16.85 ± 0.24 | ||||||||||
| GTP | 38.87 ± 0.6 | 51.94 ± 1.7 | 25.13 ± 4.8 | 25.13 ± 4.80 | ||||||||||
| Citrate | 27.33 ± 3.71 | 27.18 ± 4.03 | 27.18 ± 4.03 | |||||||||||
| Alanine | 17.15 ± 0.02 | 44.67 ± 0.26 | 17.15 ± 0.02 | |||||||||||
| Lactic acid | 18.08 ± 0.19 | 46.29 ± 0.74 | 18.08 ± 0.19 | |||||||||||
| Pyruvic acid | 102.65 ± 9.42 | 102.65 ± 9.42 | ||||||||||||
| Acetic acid | 24.61 ± 0.35 | 24.61 ± 0.35 | ||||||||||||
| Fumaric acid | 18.89 ± 0.39 | 18.89 ± 0.39 | ||||||||||||
| Succinic acid | 10.69 ± 0.11 | 10.69 ± 0.11 | ||||||||||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Honrao, C.; Teissier, N.; Zhang, B.; Powers, R.; O’Day, E.M. Gadolinium-Based Paramagnetic Relaxation Enhancement Agent Enhances Sensitivity for NUS Multidimensional NMR-Based Metabolomics. Molecules 2021, 26, 5115. https://doi.org/10.3390/molecules26175115
Honrao C, Teissier N, Zhang B, Powers R, O’Day EM. Gadolinium-Based Paramagnetic Relaxation Enhancement Agent Enhances Sensitivity for NUS Multidimensional NMR-Based Metabolomics. Molecules. 2021; 26(17):5115. https://doi.org/10.3390/molecules26175115
Chicago/Turabian StyleHonrao, Chandrashekhar, Nathalie Teissier, Bo Zhang, Robert Powers, and Elizabeth M. O’Day. 2021. "Gadolinium-Based Paramagnetic Relaxation Enhancement Agent Enhances Sensitivity for NUS Multidimensional NMR-Based Metabolomics" Molecules 26, no. 17: 5115. https://doi.org/10.3390/molecules26175115
APA StyleHonrao, C., Teissier, N., Zhang, B., Powers, R., & O’Day, E. M. (2021). Gadolinium-Based Paramagnetic Relaxation Enhancement Agent Enhances Sensitivity for NUS Multidimensional NMR-Based Metabolomics. Molecules, 26(17), 5115. https://doi.org/10.3390/molecules26175115








