Recent Research Progress in Taxol Biosynthetic Pathway and Acylation Reactions Mediated by Taxus Acyltransferases
Abstract
1. Introduction
2. Taxol Biosynthetic Pathway
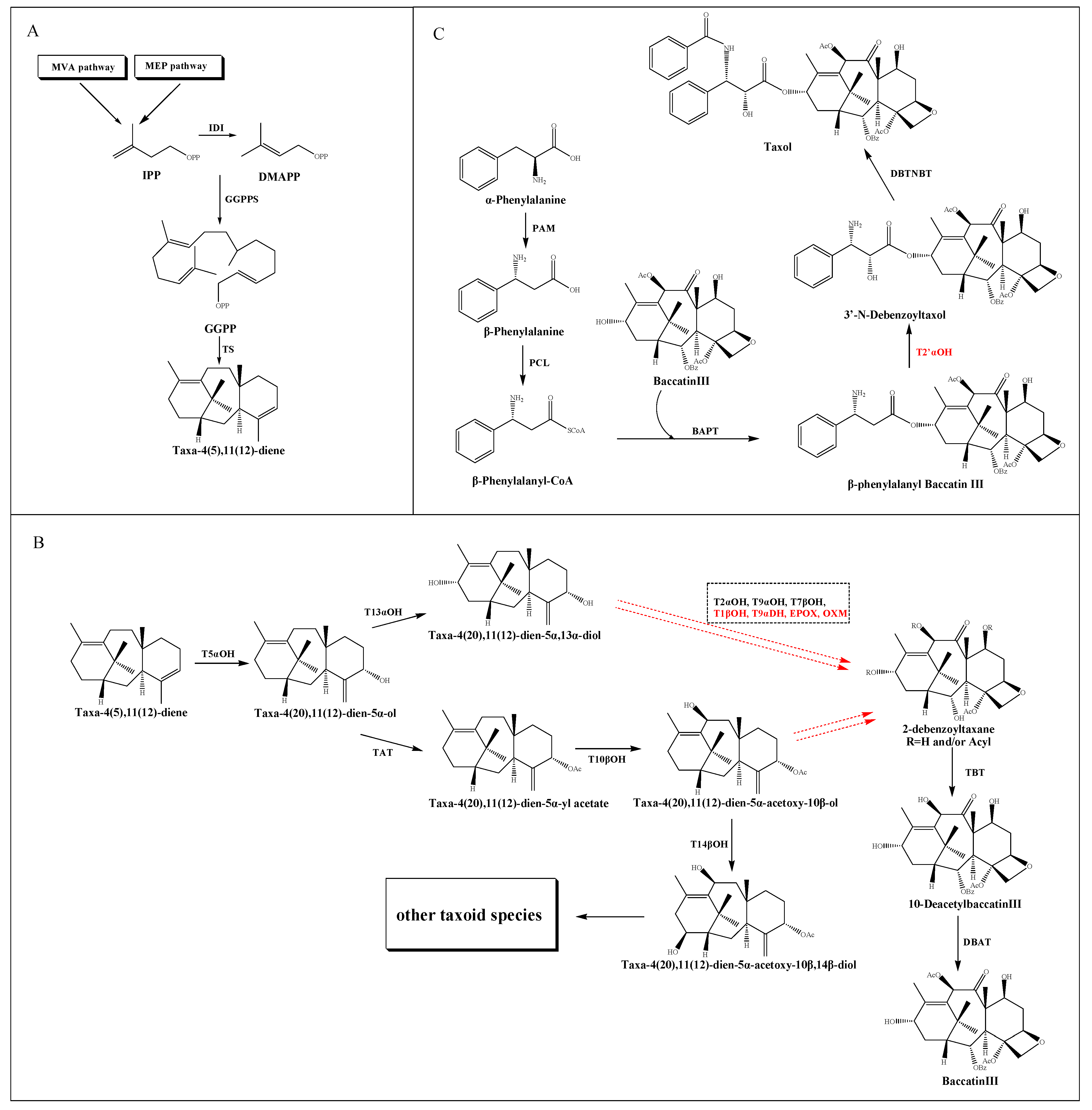
2.1. The Source and Formation of Taxane Core
2.2. The Modification of the Core Taxane Skeleton
2.3. The Synthesis of β-Phenylalanyl-CoA Side Chain and the Assembly of Taxol
2.4. Key Enzymes Involved in the Taxol Pathway
3. Acylation Reactions Mediated by Taxus ACTs
3.1. Characterized Taxus ACTs Involved in the Taxol Pathway
3.1.1. Taxadiene-5α-ol-O-acetyl Transferase (TAT)
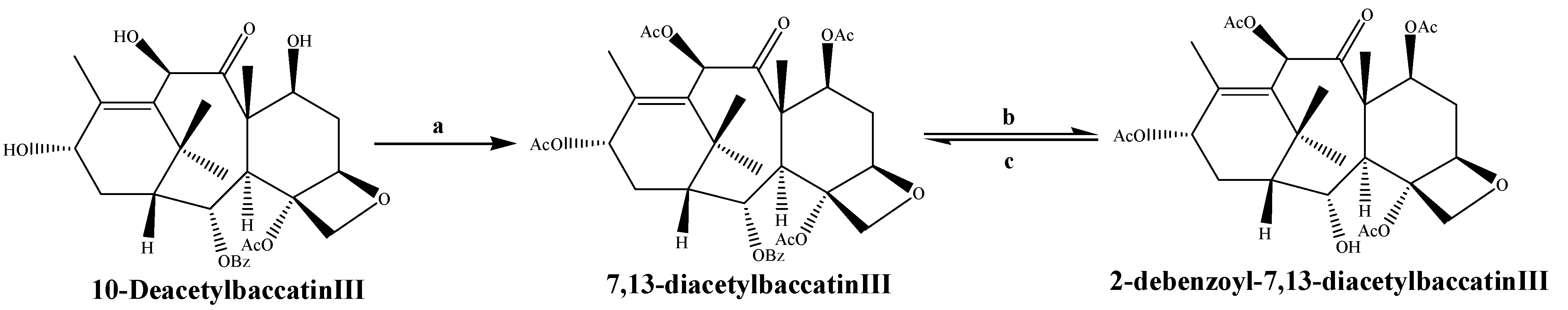
3.1.2. Taxane-2α-O-benzoyl Transferase (TBT)
3.1.3. 10-Deacetylbaccatin III-10-O-acetyl Transferase (DBAT)
3.1.4. Baccatin III: 3-amino, 3-phenylpropanoyl Transferase (BAPT)
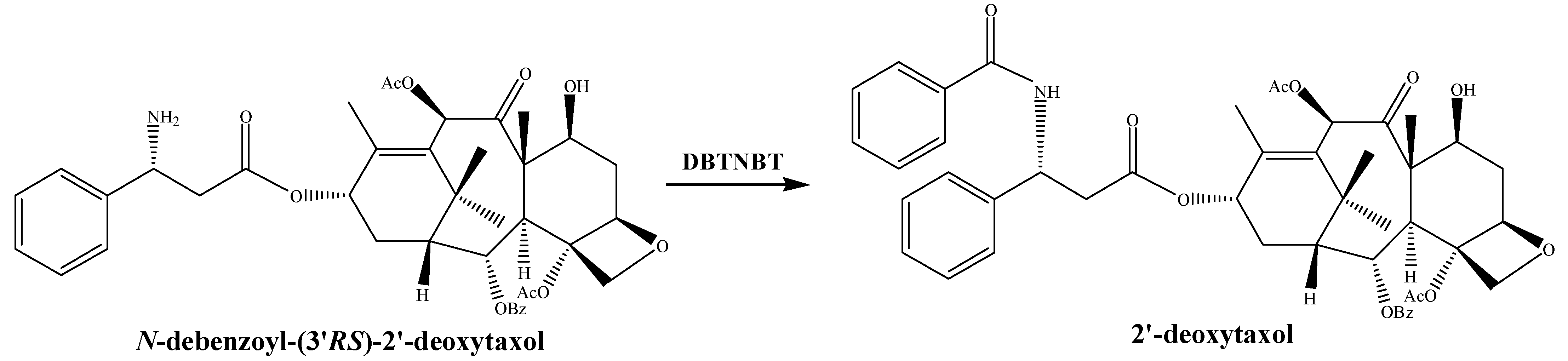
3.1.5. N-benzoyl Transferase (DBTNBT)
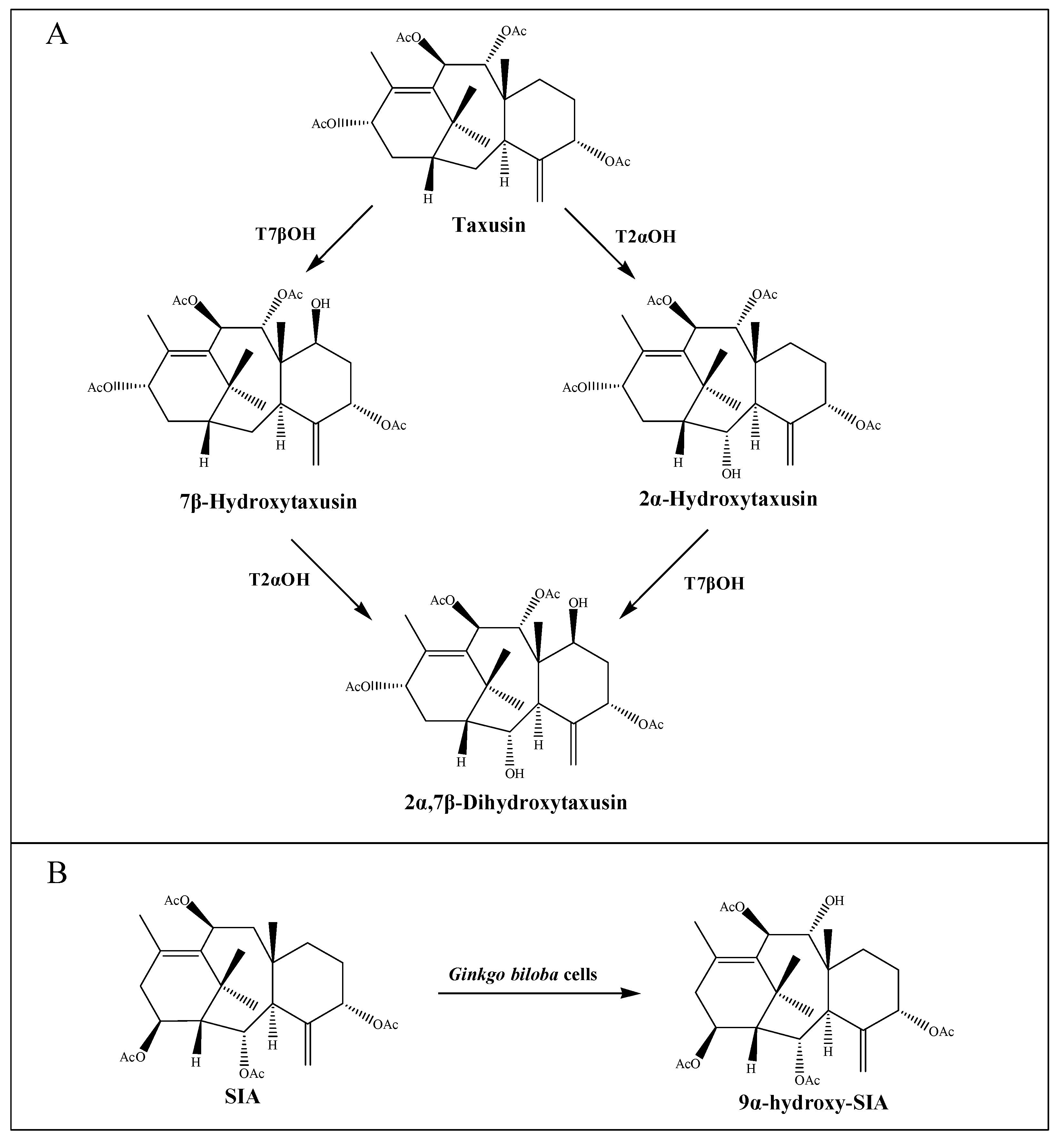
3.2. Other Taxus ACT Genes Involved in the Taxane Metabolism
3.3. Formation of Taxane Core Mediated by Taxus ACTs
3.4. Acylation Mechanism Mediated by Taxus ACTs
4. Conclusions and Perspectives
Author Contributions
Funding
Conflicts of Interest
References
- Wani, M.C.; Taylor, H.L.; Wall, M.E.; Coggon, P.; McPhail, A.T. Plant antitumor agents. VI. Isolation and structure of taxol, a novel antileukemic and antitumor agent from Taxus brevifolia. J. Am. Chem. Soc. 1971, 93, 2325–2327. [Google Scholar] [CrossRef]
- Wani, M.C.; Horwitz, S.B. Nature as a remarkable chemist: A personal story of the discovery and development of taxol. Anti-Cancer Drug. 2014, 25, 482–487. [Google Scholar] [CrossRef] [PubMed]
- Howat, S.; Park, B.; Oh, I.S.; Jin, Y.; Lee, E.; Loake, G.J. Paclitaxel: Biosynthesis, production and future prospects. New Biotechnol. 2014, 31, 242–245. [Google Scholar] [CrossRef] [PubMed]
- Priyadarshini, K.; Keerthi, A.U. Paclitaxel against cancer: A short review. Med. Chem. 2012, 2, 139–141. [Google Scholar]
- Holton, R.A.; Somoza, C.; Kim, H.B.; Liang, F.; Biediger, R.J.; Boatman, P.D.; Shindo, M.; Smith, C.C.; Kim, S. First total synthesis of taxol. 1. Functionalization of the B ring. J. Am. Chem. Soc. 1994, 116, 1597–1598. [Google Scholar] [CrossRef]
- Nicolaou, K.C.; Yang, Z.; Liu, J.J.; Ueno, H.; Nantermet, P.G.; Guy, R.K.; Claiborne, C.F.; Renaud, J.; Couladouros, E.A.; Paulvannan, K.; et al. Total synthesis of taxol. Nature 1994, 367, 630–634. [Google Scholar] [CrossRef]
- Danishefsky, S.J.; Masters, J.J.; Young, W.B.; Link, J.T.; Snyder, L.B.; Magee, T.V.; Jung, D.K.; Isaacs, R.C.A.; Bornmann, W.G.; Alaimo, C.A.; et al. Total synthesis of baccatin III and Taxol. J. Am. Chem. Soc. 1996, 118, 2843–2859. [Google Scholar] [CrossRef]
- Fu, Y.; Li, S.; Zu, Y.; Yang, G.; Yang, Z.; Luo, M.; Jiang, S.; Wink, M.; Efferth, T. Medicinal chemistry of paclitaxel and its analogues. Curr. Med. Chem. 2009, 16, 3966–3985. [Google Scholar] [CrossRef] [PubMed]
- Shao, F.; Wilson, I.; Qiu, D. The research progress of taxol in Taxus. Curr. Pharm. Biotechnol. 2021, 22, 360–366. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.C.; Gong, T.; Zhu, P. Advances in exploring alternative Taxol sources. RSC Adv. 2016, 6, 48800–48809. [Google Scholar] [CrossRef]
- Wang, T.; Chen, Y.; Zhuang, W.; Zhang, F.; Shu, X.; Wang, Z.; Yang, Q. Transcriptome sequencing reveals regulatory mechanisms of taxol synthesis in Taxus wallichiana var. mairei. Int. J. Genom. 2019, 2019, 1596895. [Google Scholar] [CrossRef]
- Gond, S.K.; Kharwar, R.N.; White, J.F., Jr. Will fungi be the new source of the blockbuster drug taxol? Fungal Biol. Rev. 2014, 28, 77–84. [Google Scholar] [CrossRef]
- Ajikumar, P.K.; Xiao, W.H.; Tyo, K.E.J.; Wang, Y.; Simeon, F.; Leonard, E.; Mucha, O.; Phon, T.H.; Pfeifer, B.; Stephanopoulos, G. Isoprenoid pathway optimization for Taxol precursor overproduction in Escherichia coli. Science 2010, 330, 70–74. [Google Scholar] [CrossRef]
- Zhou, K.; Qiao, K.; Edgar, S.; Stephanopoulos, G. Distributing a metabolic pathway among a microbial consortium enhances production of natural products. Nat. Biotechnol. 2015, 33, 377–383. [Google Scholar] [CrossRef] [PubMed]
- Besumbes, Ó.; Sauret-Güeto, S.; Phillips, M.A.; Imperial, S.; Rodríguez-Concepción, M.; Boronat, A. Metabolic engineering of isoprenoid biosynthesis in Arabidopsis for the production of taxadiene, the first committed precursor of Taxol. Biotechnol. Bioeng. 2004, 88, 168–175. [Google Scholar] [CrossRef]
- Kovacs, K.; Zhang, L.; Linforth, R.T.; Whittaker, B.; Hayes, C.; Fray, R. Redirection of carotenoid metabolism for the efficient production of taxadiene [taxa-4(5),11(12)-diene] in transgenic tomato fruit. Transgen. Res. 2007, 16, 121–126. [Google Scholar] [CrossRef]
- Anterola, A.; Shanle, E.; Perroud, P.; Quatrano, R. Production of taxa-4(5),11(12)-diene by transgenic Physcomitrella patens. Transgen. Res. 2009, 18, 655–660. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Mutanda, I.; Wang, K.; Yang, L.; Wang, J.; Wang, Y. Chloroplastic metabolic engineering coupled with isoprenoid pool enhancement for committed taxanes biosynthesis in Nicotiana benthamiana. Nat. Commun. 2019, 10, 1–12. [Google Scholar] [CrossRef]
- Liu, Y.Y.; Mo, T.; Wang, X.H.; Shi, S.P.; Liu, X.; Tu, P.F. Research progress of plant BAHD acyltransferase family. China J. Chin. Mater. Med. 2016, 41, 2175–2182. [Google Scholar]
- Li, Y.; Zhang, G.; Pfeifer, B.A. Current and emerging options for taxol production. Adv. Biochem. Eng. Biot. 2015, 148, 405–425. [Google Scholar]
- Sah, B.; Subban, K.; Jayabaskaran, C. Biochemical insights into the recombinant 10-deacetylbaccatin III-10-β-O-acetyltransferase enzyme from the Taxol-producing endophytic fungus Lasiodiplodia theobromae. Fems Microbiol. Lett. 2019, 366, fnz072. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Li, S.T.; Liu, T.T.; Fu, C.H.; Zhou, P.P.; Zhao, C.F.; Yu, L.J. Overexpression of a 10-deacetylbaccatin III-10 β-O-acetyltransferase gene leads to increased taxol yield in cells of Taxus chinensis. Plant Cell Tiss. Org. 2011, 106, 63–70. [Google Scholar] [CrossRef]
- Croteau, R.; Ketchum, R.E.; Long, R.M.; Kaspera, R.; Wildung, M.R. Taxol biosynthesis and molecular genetics. Phytochem. Rev. 2006, 5, 75–97. [Google Scholar] [CrossRef]
- Wildung, M.R.; Croteau, R. A cDNA clone for taxadiene synthase, the diterpene cyclase that catalyzes the committed step of taxol biosynthesis. J. Biol. Chem. 1996, 271, 9201–9204. [Google Scholar] [CrossRef] [PubMed]
- Hefner, J.; Ketchum, R.E.; Croteau, R. Cloning and functional expression of a cDNA encoding geranylgeranyl diphosphate synthase from Taxus canadensis and assessment of the role of this prenyltransferase in cells induced for taxol production. Arch. Biochem. Biophys. 1998, 360, 62–74. [Google Scholar] [CrossRef]
- Jarchow-Choy, S.K.; Koppisch, A.T.; Fox, D.T. Synthetic routes to methylerythritol phosphate pathway intermediates and downstream isoprenoids. Curr. Org. Chem. 2014, 18, 1050–1072. [Google Scholar] [CrossRef][Green Version]
- Roberts, S.C. Production and engineering of terpenoids in plant cell culture. Nat. Chem. Biol. 2007, 3, 387. [Google Scholar] [CrossRef] [PubMed]
- Malik, S.; Cusidó, R.M.; Mirjalili, M.H.; Moyano, E.; Palazón, J.; Bonfill, M. Production of the anticancer drug taxol in Taxus baccata suspension cultures: A review. Process. Biochem. 2011, 46, 23. [Google Scholar] [CrossRef]
- Kuang, X.J.; Sun, S.J.; Wei, J.H.; Li, Y.; Sun, C. Iso-Seq analysis of the Taxus cuspidata transcriptome reveals the complexity of Taxol biosynthesis. BMC Plant Biol. 2019, 19, 210. [Google Scholar] [CrossRef] [PubMed]
- Jennewein, S.; Wildung, M.R.; Chau, M.; Walker, K.; Croteau, R. Random sequencing of an induced Taxus cell cDNA library for identification of clones involved in taxol biosynthesis. Proc. Natl. Acad. Sci. USA 2004, 101, 9149–9154. [Google Scholar] [CrossRef] [PubMed]
- Jennewein, S.; Rithner, C.D.; Williams, R.M.; Croteau, R.B. Taxol biosynthesis: Taxane 13 alpha-hydroxylase is a cytochrome p450-dependent monooxygenase. Proc. Natl. Acad. Sci. USA 2001, 98, 13595–13600. [Google Scholar] [CrossRef]
- Walker, K.; Schoendorf, A.; Croteau, R. Molecular cloning of a taxa-4 (20), 11 (12)-dien-5α-ol-O-acetyl transferase cDNA from Taxus and functional expression in Escherichia coli. Arch. Biochem. Biophys. 2000, 374, 371–380. [Google Scholar] [CrossRef]
- Schoendorf, A.; Rithner, C.D.; Williams, R.M.; Croteau, R.B. Molecular cloning of a cytochrome P450 taxane 10 beta-hydroxylase cDNA from Taxus and functional expression in yeast. Proc. Natl. Acad. Sci. USA 2001, 98, 1501–1506. [Google Scholar] [CrossRef] [PubMed]
- Jennewein, S.; Rithner, C.D.; Williams, R.M.; Croteau, R. Taxoid metabolism: Taxoid 14beta-hydroxylase is a cytochrome P450-dependent monooxygenase. Arch. Biochem. Biophys. 2003, 413, 262. [Google Scholar] [CrossRef]
- Holton, R.A.; Kim, H.B.; Somoza, C.; Liang, F.; Biediger, R.J.; Boatman, P.D.; Shindo, M.; Smith, C.C.; Kim, S.C.; Nadizadeh, H.; et al. First total synthesis of taxol. 2. Completion of the C-ring and D-ring. J. Am. Chem. Soc. 1994, 116, 1599–1600. [Google Scholar] [CrossRef]
- Vongpaseuth, K.; Roberts, S.C. Advancements in the understanding of paclitaxel metabolism in tissue culture. Curr. Pharm. Biotechnol. 2007, 8, 219–236. [Google Scholar] [CrossRef] [PubMed]
- Walker, K.; Croteau, R. Taxol biosynthesis: Molecular cloning of a benzoyl-CoA: Taxane 2α-O-benzoyltransferase cDNA from Taxus and functional expression in Escherichia coli. Proc. Natl. Acad. Sci. USA 2000, 97, 13591–13596. [Google Scholar] [CrossRef] [PubMed]
- Walker, K.; Croteau, R.M. Molecular cloning of a 10-deacetylbaccatin Ⅲ-10-O-acetyl transferase cDNA from Taxus and functional expression in Escherichia coli. Proc. Natl. Acad. Sci. USA 2000, 97, 583–587. [Google Scholar] [CrossRef] [PubMed]
- Walker, K.D.; Klettke, K.; Akiyama, T.; Croteau, R. Cloning, heterologous expression, and characterization of a phenylalanine aminomutase involved in Taxol biosynthesis. J. Biol. Chem. 2004, 279, 53947–53954. [Google Scholar] [CrossRef]
- Ramírez-Estrada, K.; Altabella, T.; Onrubia, M.; Moyano, E.; Notredame, C.; Osuna, L.; VandenBossche, R.; Goossens, A.; Cusido, R.; Palazon, J. Transcript profiling of jasmonate-elicited Taxus cells reveals a β-phenylalanine-CoA ligase. Plant Biotechnol. J. 2016, 14, 85–96. [Google Scholar] [CrossRef]
- Walker, K.; Fujisaki, S.; Long, R.; Croteau, R. Molecular cloning and heterologous expression of the C-13 phenylpropanoid side chain-CoA acyltransferase that functions in taxol biosynthesis. Proc. Natl. Acad. Sci. USA 2002, 99, 12715–12720. [Google Scholar] [CrossRef] [PubMed]
- Walker, K.; Long, R.; Croteau, R. The final acylation step in taxol biosynthesis: Cloning of the taxoid C13-side-chain N-benzoyltransferase from Taxus. Proc. Natl. Acad. Sci. USA 2002, 99, 9166. [Google Scholar] [CrossRef] [PubMed]
- Chau, M.; Croteau, R. Molecular cloning and characterization of a cytochrome P450 taxoid 2alpha-hydroxylase involved in Taxol biosynthesis. Arch. Biochem. Biophys. 2004, 427, 48–57. [Google Scholar] [CrossRef] [PubMed]
- Chau, M.; Jennewein, S.; Walker, K.; Croteau, R. Taxol biosynthesis: Molecular cloning and characterization of a cytochrome P450 taxoid 7 β-hydroxylase. Chem. Biol. 2004, 11, 663–672. [Google Scholar]
- Zhang, N.; Han, Z.T.; Sun, G.L.; Hoffman, A.; Wilson, I.W.; Yang, Y.F.; Gao, Q.; Wu, J.Q.; Xie, D.; Dai, J.G.; et al. Molecular cloning and characterization of a cytochrome P450 taxoid 9α-hydroxylase in Ginkgo biloba cells. Biochem. Bioph. Res. Commun. 2014, 443, 938–943. [Google Scholar] [CrossRef]
- McElroy, C.; Jennewein, S. Taxol biosynthesis and production: From forests to fermenters. In Biotechnology of Natural Products; Schwab, W., Lange, B., Wüst, M., Eds.; Springer: Cham, Switzerland, 2018; pp. 145–185. [Google Scholar]
- Hao, D.C.; Ge, G.B.; Xiao, P.G.; Zhang, Y.Y.; Ling, Y.; Hans, E. The first insight into the tissue specific Taxus transcriptome via Illumina second generation sequencing. PLoS ONE 2011, 6, e21220. [Google Scholar]
- He, C.T.; Li, Z.L.; Zhou, Q.; Shen, C.; Huang, Y.Y.; Mubeen, S.; Yang, J.Z.; Yuan, J.G.; Yang, Z.Y. Transcriptome profiling reveals specific patterns of paclitaxel synthesis in a new Taxus yunnanensis cultivar. Plant Physiol. Biochem. 2018, 122, 10–18. [Google Scholar] [CrossRef]
- Mubeen, S.; Li, Z.L.; Huang, Q.M.; He, C.T.; Yang, Z.Y. Comparative transcriptome analysis revealed the tissue-specific accumulations of taxanes among three experimental lines of Taxus yunnanensis. J. Agric. Food. Chem. 2018, 66, 10410–10420. [Google Scholar] [CrossRef]
- Yu, C.N.; Guo, H.; Zhang, Y.Y.; Song, Y.B.; Pi, E.; Yu, C.L.; Zhang, L.; Dong, M.; Zheng, B.S.; Wang, H.Z.; et al. Identification of potential genes that contributed to the variation in the taxoid contents between two Taxus species (Taxus media and Taxus mairei). Tree Physiol. 2017, 37, 1659–1671. [Google Scholar] [CrossRef] [PubMed]
- Zhou, T.; Luo, X.J.; Yu, C.N.; Zhang, C.C.; Zhang, L.; Song, Y.B.; Dong, M.; Shen, C.J. Transcriptome analyses provide insights into the expression pattern and sequence similarity of several taxol biosynthesis related genes in three Taxus species. BMC Plant Biol. 2019, 19, 33. [Google Scholar] [CrossRef]
- Hao, J.; Guo, H.; Shi, X.A.; Wang, Y.; Wan, Q.H.; Song, Y.B.; Zhang, L.; Dong, M.; Shen, C.J. Comparative proteomic analyses of two Taxus species (Taxus × media and Taxus mairei) reveals variations in the metabolisms associated with paclitaxel and other metabolites. Plant Cell Physiol. 2017, 58, 1878–1890. [Google Scholar] [CrossRef]
- Yu, C.N.; Luo, X.J.; Zhan, X.R.; Hao, J.; Zhang, L.; Song, Y.B.; Shen, C.J.; Dong, M. Comparative metabolomics reveals the metabolic variations between two endangered Taxus species (T. fuana and T. yunnanensis) in the Himalayas. BMC Plant Biol. 2018, 18, 197. [Google Scholar] [CrossRef]
- Yu, C.N.; Zhang, C.C.; Xu, X.Y.; Huang, J.F.; Chen, Y.Y.; Luo, X.J.; Wang, H.Z.; Shen, C.J. Omic analysis of the endangered Taxaceae species Pseudotaxus chienii revealed the differences in taxol biosynthesis pathway between Pseudotaxus and Taxus yunnanensis trees. BMC Plant Biol. 2021, 21, 104. [Google Scholar] [CrossRef]
- Li, S.T.; Zhang, P.; Zhang, M.; Fu, C.H.; Zhao, C.F.; Dong, Y.S.; Guo, A.Y.; Yu, L.J. Transcriptional profile of Taxus chinensis cells in response to methyl jasmonate. BMC Genom. 2012, 13, 295. [Google Scholar] [CrossRef]
- Sun, G.; Yang, Y.; Xie, F.; Wen, J.-F.; Wu, J.; Wilson, I.W.; Liu, H.; Qiu, D. Deep sequencing reveals transcriptome re-programming of Taxus× media cells to the elicitation with methyl jasmonate. PLoS ONE 2013, 8, e62865. [Google Scholar]
- Jiang, L.Y.; Zhang, K.K.; Lü, X.; Yang, L.Y.; Wang, S.; Chen, D.F.; Yang, Y.F.; Qiu, D.Y. Characterization and expression analysis of genes encoding Taxol biosynthetic enzymes in Taxus spp. J. For. Res. 2021. [Google Scholar] [CrossRef]
- Zhang, M.; Li, S.; Nie, L.; Chen, Q.; Xu, X.; Yu, L.; Fu, C. Two jasmonate-responsive factors, TcERF12 and TcERF15, respectively act as repressor and activator of tasy gene of taxol biosynthesis in Taxus chinensis. Plant Mol. Biol. 2015, 89, 463–473. [Google Scholar] [CrossRef]
- Zhang, M.; Jin, X.; Chen, Y.; Wei, M.; Liao, W.; Zhao, S.; Fu, C.; Yu, L. TcMYC2a, a basic helix–loop–helix transcription factor, transduces JA-signals and regulates taxol biosynthesis in Taxus chinensis. Front. Plant Sci. 2018, 9, 863. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Zhang, P.; Zhang, M.; Fu, C.; Yu, L. Functional analysis of a WRKY transcription factor involved in transcriptional activation of the DBAT gene in Taxus chinensis. Plant Biol. 2012, 15, 19–26. [Google Scholar] [CrossRef] [PubMed]
- Yu, C.N.; Luo, X.J.; Zhang, C.C.; Xu, X.Y.; Huang, J.F.; Chen, Y.Y.; Feng, S.G.; Zhan, X.R.; Zhang, L.; Yuan, H.W.; et al. Tissue-specific study across the stem of Taxus media identifies a phloem-specific TmMYB3 involved in the transcriptional regulation of paclitaxel biosynthesis. Plant J. 2020, 103, 95–110. [Google Scholar] [CrossRef] [PubMed]
- Hu, X.L.; Zhang, L.S.; Wilson, I.; Shao, F.J.; Qiu, D.Y. The R2R3-MYB transcription factor family in Taxus chinensis: Identification, characterization, expression profiling and posttranscriptional regulation analysis. PeerJ 2020, 8, e8473. [Google Scholar] [CrossRef]
- Sun, Y.M.; Li, Y.H.; Cheng, X.; Li, G.H.; Sheng, L.L.; Jin, Q.; Cai, Y.P.; Lin, Y. Identification, sequence characterization and expression analysis of BAHD family in pear. J. Nucl. Agric. Sci. 2018, 32, 1917–1930. [Google Scholar]
- D’Auria, J.C. Acyltransferases in plants: A good time to be BAHD. Curr. Opin. Plant Biol. 2006, 9, 331–340. [Google Scholar] [CrossRef]
- Zhao, D.L.; Chi, Y.; Yu, F.; Lu, Y.; Li, Y.J. Cloning and Characterization of a Taxane 2α-O-benzoyl Transferase (TBT) cDNA from Taxus yunnanensis. Acta Bot. Boreal. Occident. Sin. 2007, 27, 1713–1719. [Google Scholar]
- Long, R.M.; Lagisetti, C.; Coates, R.M.; Croteau, R.B. Specificity of the N-benzoyl transferase responsible for the last step of taxol biosynthesis. Arch. Biochem. Biophys. 2008, 477, 384–389. [Google Scholar] [CrossRef] [PubMed]
- Chau, M.D.; Walker, K.; Long, R.; Croteau, R. Regioselectivity of taxoid-O-acetyltransferases: Heterologous expression and characterization of a new taxadien-5α-ol-O-acetyltransferase. Arch. Biochem. Biophy. 2004, 430, 237–246. [Google Scholar] [CrossRef]
- Hampel, D.; Mau, C.J.; Croteau, R. Taxol biosynthesis: Identification and characterization of two acetyl CoA: Taxoid-O-acetyl transferases that divert pathway flux away from Taxol production. Arch. Biochem. Biophy. 2009, 487, 91–97. [Google Scholar] [CrossRef][Green Version]
- Nevarez, D.M.; Mengistu, Y.A.; Nawarathne, I.N.; Walker, K.D. An N-aroyltransferase of the BAHD superfamily has broad aroyl CoA specificity in vitro with analogues of N-dearoylpaclitaxel. J. Am. Chem. Soc. 2009, 131, 5994–6002. [Google Scholar] [CrossRef] [PubMed]
- Kusano, H.; Li, H.; Minami, H.; Kato, Y.; Tabata, H.; Yazaki, K. Evolutionary developments in plant specialized metabolism, exemplified by two transferase families. Front. Plant Sci. 2019, 10, 794. [Google Scholar] [CrossRef]
- Fani, R.; Fondi, M. Origin and evolution of metabolic pathways. Phys. Life Rev. 2009, 6, 23–52. [Google Scholar] [CrossRef]
- Li, B.J.; Wang, H.; Gong, T.; Chen, J.J.; Chen, T.J.; Yang, J.L.; Zhu, P. Improving 10-deacetylbaccatin III-10-β-O-acetyltransferase catalytic fitness for Taxol production. Nat. Commun. 2017, 8, 15544. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Koepke, J.; Panjikar, S.; Fritzsch, G.; Stöckigt, J. Crystal structure of vinorine synthase, the first representative of the BAHD superfamily. J. Biol. Chem. 2005, 280, 13576. [Google Scholar] [CrossRef] [PubMed]
- You, L.F.; Wei, T.; Zheng, Q.W.; Lin, J.F.; Guo, L.Q.; Jiang, B.H.; Huang, J.J. Activity essential residue analysis of Taxoid 10β-O-acetyl transferase for enzymatic synthesis of Baccatin. Appl. Biochem. Biotechnol. 2018, 186, 949–959. [Google Scholar] [CrossRef] [PubMed]
- You, L.F.; Huang, J.J.; Wei, T.; Lin, S.H.; Jiang, B.H.; Guo, L.Q.; Lin, J.F. Enhanced catalytic activities and modified substrate preferences for taxoid 10β-O-acetyl transferase mutants by engineering catalytic histidine residues. Biotechnol. Lett. 2018, 40, 1245–1251. [Google Scholar] [CrossRef] [PubMed]
| Enzyme Abbreviation | Size (kDa) | Probable Localization | GenBank Number |
|---|---|---|---|
| Geranylgeranyl diphosphate synthase GGPPS | 42 | Plastids | AF081514 |
| Taxadiene synthase TS | 98 | Plastids | AY364469 |
| Taxadiene-5α-hydroxylase T5αOH | 56 | Endoplasmic reticulum | AY289209 |
| Taxane-10β-hydroxylase T10βOH | 56 | Endoplasmic reticulum | AF318211 |
| Taxane-10β-hydroxylase a T10βOH | 55 | Endoplasmic reticulum | AY563635 |
| Taxadiene-13α-hydroxylase T13αOH | 54 | Endoplasmic reticulum | AY056019 |
| Taxane-2α-hydroxylase T2αOH | 55 | Endoplasmic reticulum | AY518383 |
| Taxane-9α-hydroxylase T9αOH | 55 | Endoplasmic reticulum | KF773141 |
| Taxane-7β-hydroxylase T7βOH | 56 | Endoplasmic reticulum | AY307951 |
| Taxadiene-5α-ol-O-acetyl transferase TAT | 49 | Cytosol | AF190130 |
| Taxane-2α-O-benzoyl transferase TBT | 50 | Cytosol | AF297618 |
| 10-deacetylbaccatin III-10-O acetyl transferase DBAT | 49 | Cytosol | AF193765 |
| Baccatin III: 3-amino, 13-phenylpropanoyltransferase BAPT | 50 | Cytosol | AY082804 |
| N-benzoyl transferase DBTNBT | 49 | Cytosol | AF466397 |
| Phenylalanine aminomutase PAM | 76 | Cytosol | AY582743 |
| β-phenylalanyl-CoA ligase b PCL | 59 | Cytosol | KM593667 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, T.; Li, L.; Zhuang, W.; Zhang, F.; Shu, X.; Wang, N.; Wang, Z. Recent Research Progress in Taxol Biosynthetic Pathway and Acylation Reactions Mediated by Taxus Acyltransferases. Molecules 2021, 26, 2855. https://doi.org/10.3390/molecules26102855
Wang T, Li L, Zhuang W, Zhang F, Shu X, Wang N, Wang Z. Recent Research Progress in Taxol Biosynthetic Pathway and Acylation Reactions Mediated by Taxus Acyltransferases. Molecules. 2021; 26(10):2855. https://doi.org/10.3390/molecules26102855
Chicago/Turabian StyleWang, Tao, Lingyu Li, Weibing Zhuang, Fengjiao Zhang, Xiaochun Shu, Ning Wang, and Zhong Wang. 2021. "Recent Research Progress in Taxol Biosynthetic Pathway and Acylation Reactions Mediated by Taxus Acyltransferases" Molecules 26, no. 10: 2855. https://doi.org/10.3390/molecules26102855
APA StyleWang, T., Li, L., Zhuang, W., Zhang, F., Shu, X., Wang, N., & Wang, Z. (2021). Recent Research Progress in Taxol Biosynthetic Pathway and Acylation Reactions Mediated by Taxus Acyltransferases. Molecules, 26(10), 2855. https://doi.org/10.3390/molecules26102855





