A Half-Century History of Applications of Antisense Oligonucleotides in Medicine, Agriculture and Forestry: We Should Continue the Journey
Abstract
1. Introduction
2. ASO Diversity and Their Functional Activities
3. Four Recent Successful Applications of ASO to Medicine
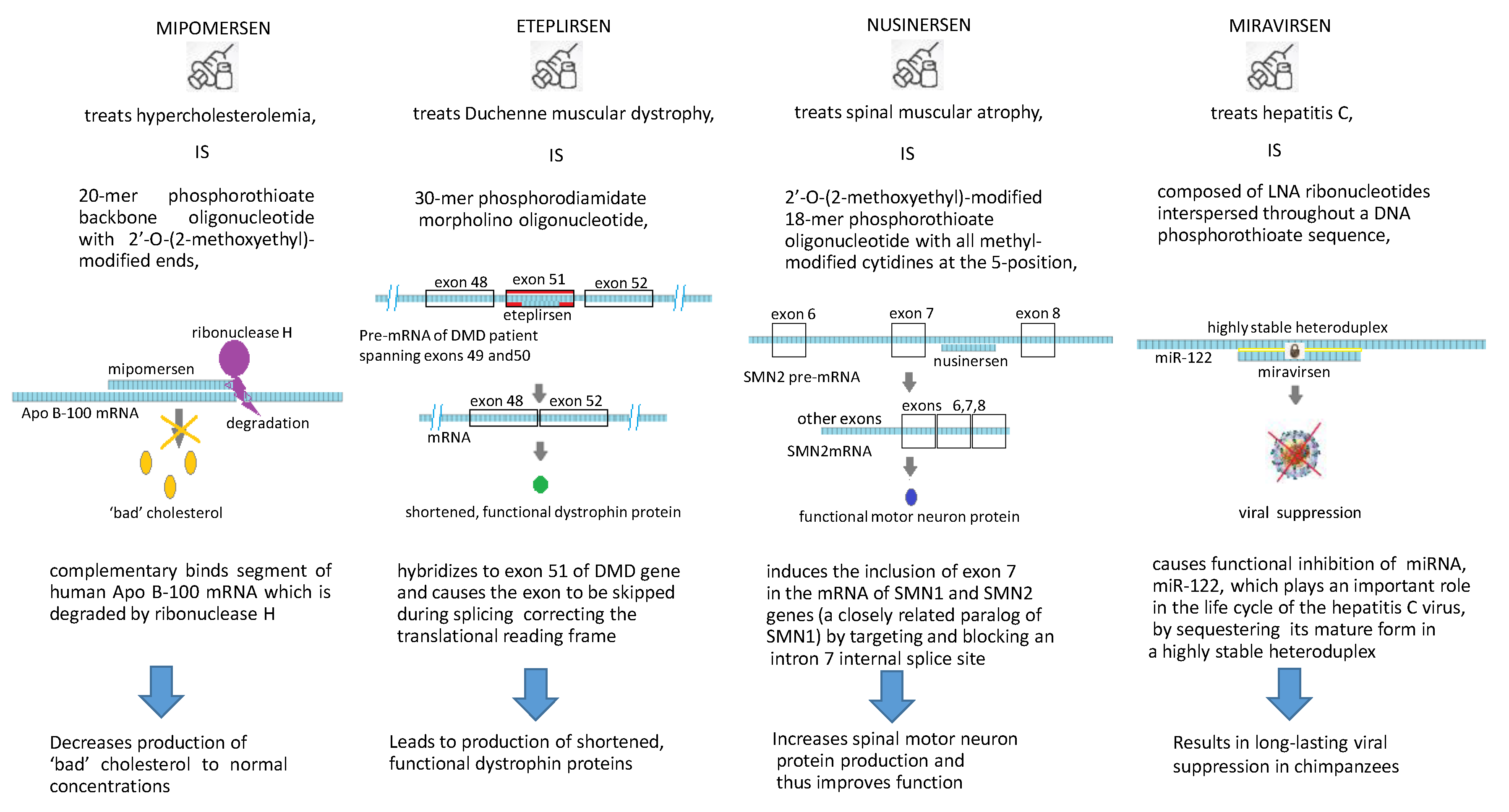
3.1. Mipomersen (Kynamro®)
3.2. Nusinersen (Spinraza®)
3.3. Eteplirsen (EXONDYS 51TM)
3.4. Miravirsen (SPC3649)
4. ASO-Based Insect Pest Management in Agriculture and Forestry
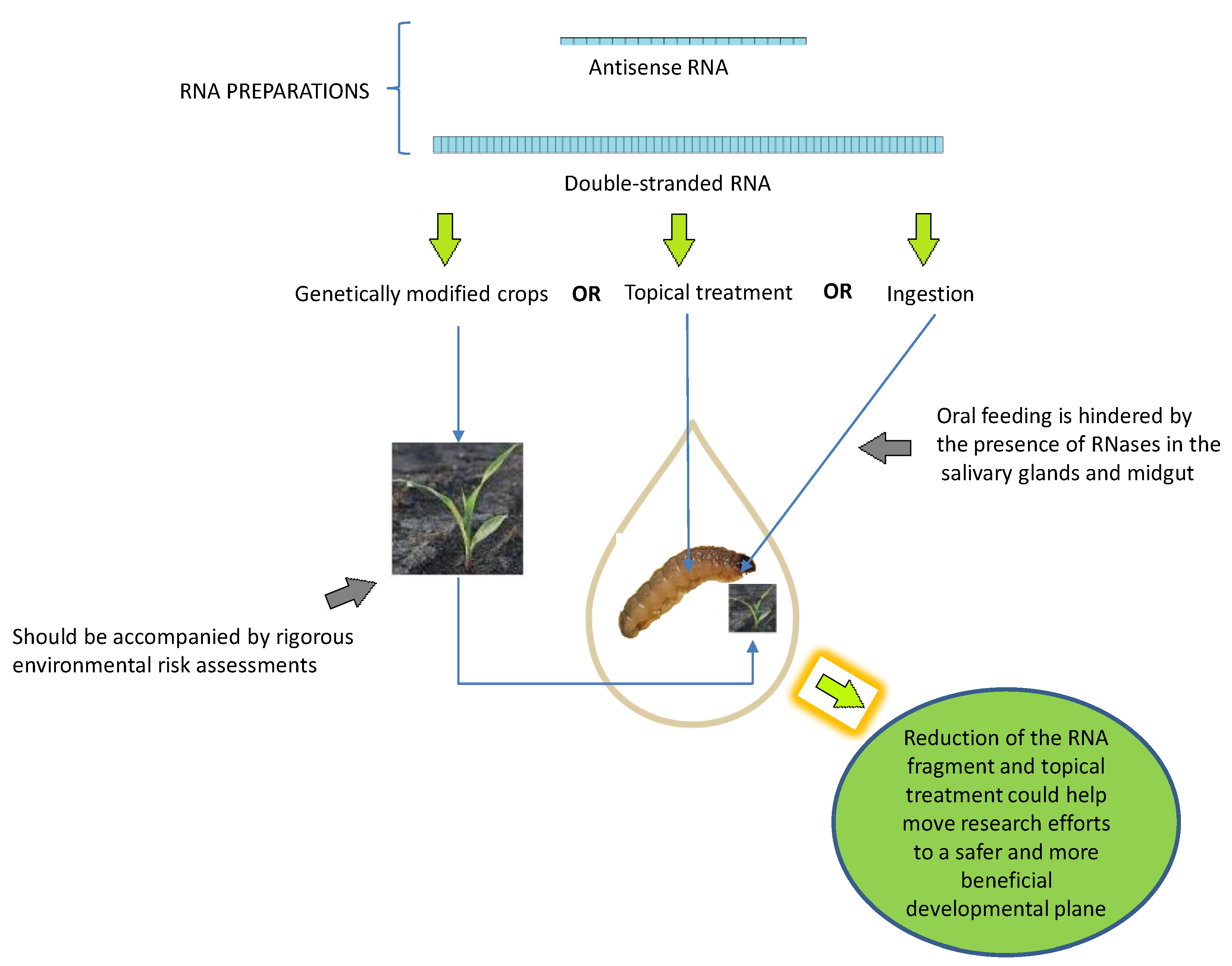
4.1. RNA Preparations
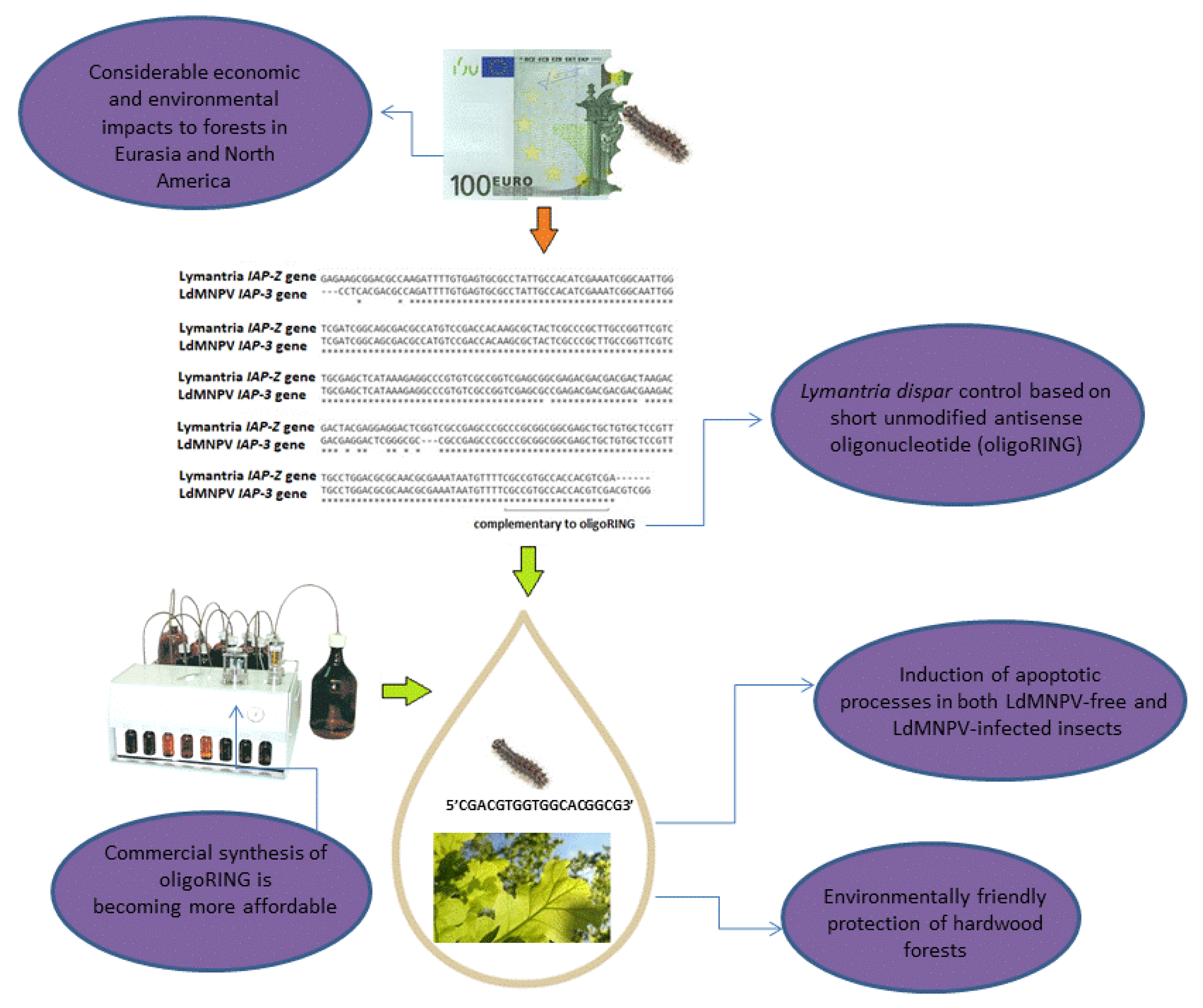
4.2. DNA Insecticides
5. Experiments and Ready-to-Use Preparations Have always Been Stronger than Words of Skepticism
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Belikova, A.; Zarytova, V.; Grineva, N. Synthesis of ribonucleosides and diribonucleoside phosphates containing 2-chloroethylamine and nitrogen mustard residues. Tetrahedron Lett. 1967, 37, 3557–3562. [Google Scholar] [CrossRef]
- Knorre, D.G. Chemical instruments in modern biology (on the example of antisense effects on genetic structures). Soros Educ. J. 1998, 12, 25–31. [Google Scholar]
- Khorana, H.G.; Agarwal, K.L.; Buchi, H.; Caruthers, M.H.; Gupta, N.K.; Kleppe, K.; Kumar, A.; Otsuka, E.; RajBhandary, U.L.; Van de Sande, J.H.; et al. Studies on polynucleotides. 103. Total synthesis of the structural gene for an alanine transfer ribonucleic acid from yeast. J. Mol. Biol. 1972, 72, 209–217. [Google Scholar] [CrossRef]
- Letsinger, R.L.; Finnan, J.L.; Heavner, G.A.; Lunsford, W.B. Letter: Phosphite coupling procedure for generating internucleotide links. Am. J. Chem. Soc. 1975, 97, 3278–3279. [Google Scholar] [CrossRef]
- Caruthers, M.H.; Beaucage, S.L.; Efcavitch, J.W.; Fisher, E.F.; Matteucci, M.D.; Stabinsky, Y. New chemical methods for synthesizing polynucleotides. Nucleic Acids Res. 1980, 7, 215–223. [Google Scholar]
- Ogilvie, K.K.; Nemer, M.J. Silica gel as solid support in the synthesis of oligoribonucleotides. Tetrahedron Lett. 1980, 21, 4159–4162. [Google Scholar] [CrossRef]
- Burgers, P.M.J.; Eckstein, F. Synthesis of dinucleoside mono-phosphorothioates via addition of sulphur to phosphite tries-ter. Tetrahedron Lett. 1978, 40, 3835–3838. [Google Scholar] [CrossRef]
- Paterson, B.M.; Roberts, B.E.; Kuff, E.L. Structural gene identification and mapping by DNA-mRNA hybrid-arrested cell-free translation. Proc. Natl. Acad. Sci. USA 1977, 74, 4370–4374. [Google Scholar] [CrossRef] [PubMed]
- Zamecnik, P.C.; Stephenson, M.L. Inhibition of Rous sarcoma virus replication and cell transformation by a specific oligodeoxynucleotide. Proc. Natl Acad. Sci. USA 1978, 75, 280–284. [Google Scholar] [CrossRef] [PubMed]
- Donis-Keller, H. Site specific enzymatic cleavage of RNA. Nucleic Acids Res. 1979, 7, 179–192. [Google Scholar] [CrossRef] [PubMed]
- Sinha, N.D.; Biernat, J.; McManus, J.; Koster, H. Polymer support oligonucleotide synthesis XVIII: Use of beta-cyanoethyl-N,N-dialkylamino-/N-morpholino phosphoramidite of deoxynucleosides for the synthesis of DNA fragments simplifying deprotection and isolation of the final product. Nucleic Acids Res. 1984, 12, 4539–4557. [Google Scholar] [CrossRef] [PubMed]
- Usman, N.; Ogilvie, K.K.; Jiang, M.Y.; Cedergren, R.J. Automated chemical synthesis of long oligoribonucleotides using 2′-O-silylated ribonucleoside 3′-O-phosphoramidites on a controlled-pore glass support: Synthesis of a 43-nucleotide sequence similar to the 3′-half molecule of an Escherichia coli formylmethionine tRNA. J. Am. Chem. Soc. 1987, 109, 7845–7854. [Google Scholar] [CrossRef]
- Tamm, I.B.; Dorken, B.; Hartmann, G. Antisense therapy in oncology: New hope for an old idea? Lancet 2001, 358, 489–497. [Google Scholar] [CrossRef]
- Simons, R.W.; Kleckner, N. Translational control of IS10 transposition. Cell 1983, 34, 683–691. [Google Scholar] [CrossRef]
- Harland, R.; Weintraub, H. Translation of mRNA injected into Xenopus oocytes is specifically inhibited by antisense RNA. J. Cell Biol. 1985, 101, 1094–1099. [Google Scholar] [CrossRef] [PubMed]
- Melton, D.A. Injected anti-sense RNAs specifically block messenger RNA translation in vivo. Proc. Natl. Acad. Sci. USA 1985, 82, 144–148. [Google Scholar] [CrossRef] [PubMed]
- Izant, J.G.; Weintraub, H. Constitutive and conditional suppression of exogenous and endogenous genes by anti-sense RNA. Science 1985, 229, 345–352. [Google Scholar] [CrossRef] [PubMed]
- Gewirtz, A.M. Oligonucleotide therapeutics: A step forward. J. Clin. Oncol. 2000, 18, 1809–1811. [Google Scholar] [CrossRef] [PubMed]
- De Smet, M.D.; Meenken, C.J.; van den Horn, G.J. Fomivirsen: A phosphorothioate oligonucleotide for the treatment of CMV retinitis. Ocul. Immunol. Inflamm. 1999, 7, 189–198. [Google Scholar] [CrossRef] [PubMed]
- Stein, A.; Castanotto, D. FDA-Approved Oligonucleotide Therapies in 2017. Mol. Ther. 2017, 25, 1069–1075. [Google Scholar] [CrossRef] [PubMed]
- Fire, A.; Xu, S.; Montgomery, M.K.; Kostas, S.A.; Driver, S.E.; Mello, C.C. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 1998, 391, 806–811. [Google Scholar] [CrossRef] [PubMed]
- Guo, S.; Kemphues, K.J. Par-1, a gene required for establishing polarity in C. elegans embryos, encodes a putative Ser/Thr kinase that is asymmetrically distributed. Cell 1995, 81, 611–620. [Google Scholar] [CrossRef]
- Tiemann, K.; Rossi, J.J. RNAi-based therapeutics-current status, challenges and prospects. EMBO Mol. Med. 2009, 1, 142–151. [Google Scholar] [CrossRef] [PubMed]
- Titze-de-Almeida, R.; David, C.; Titze-de-Almeida, S.S. The Race of 10 Synthetic RNAi-Based Drugs to the Pharmaceutical Market. Pharm. Res. 2017, 34, 1339–1363. [Google Scholar] [CrossRef] [PubMed]
- Belles, X. Beyond Drosophila: RNAi in vivo and functional genomics in insects. Annu. Rev. Entomol. 2010, 55, 111–128. [Google Scholar] [CrossRef] [PubMed]
- Baum, J.A.; Bogaert, T.; Clinton, W.; Heck, G.R.; Feldmann, P.; Ilagan, O.; Johnson, S.; Plaetinck, G.; Munyikwa, T.; Pleau, M.; et al. Control of coleopteran insect pests through RNA interference. Nat. Biotechnol. 2007, 25, 1322–1326. [Google Scholar] [CrossRef] [PubMed]
- Sturm, A.C. The role of genetic counselors for patients with familial hypercholesterolemia. Curr. Genet. Med. Rep. 2014, 2, 68–74. [Google Scholar] [CrossRef]
- Gu, L.; Knipple, D.C. Recent advances in RNA interference research in insects: Implications for future insect pest management strategies. Crop Protect. 2013, 45, 36–40. [Google Scholar] [CrossRef]
- Zotti, M.J.; Smagghe, G. RNAi technology for insect management and protection of beneficial insects from diseases: Lessons, challenges and risk assessments. Neotrop. Entomol. 2015, 44, 197–213. [Google Scholar] [CrossRef] [PubMed]
- Terenius, O.; Papanicolaou, A.; Garbutt, J.S.; Eleftherianos, I.; Huvenne, H.; Kanginakudru, S.; Albrechtsen, M.; An, C.; Aymeric, J.L.; Barthel, A.; et al. RNA interference in Lepidoptera: An overview of successful and unsuccessful studies and implications for experimental design. J. Insect Physiol. 2011, 57, 231–245. [Google Scholar] [CrossRef] [PubMed]
- Lundgren, J.G.; Duan, J.J. RNAi-based insecticidal crops: Potential effects on non-target species. Bioscience 2013, 63, 657–665. [Google Scholar] [CrossRef]
- Oberemok, V.V. Method of Elimination of Phyllophagous Insects from Order Lepidoptera. Ukraine Patent UA 36,445, 27 October 2008. [Google Scholar]
- Oberemok, V.V.; Laikova, K.V.; Zaitsev, A.S.; Gushchin, V.A.; Skorokhod, O.A. The RING for gypsy moth control: Topical application of fragment of its nuclear polyhedrosis virus anti-apoptosis gene as insecticide. Pest. Biochem. Physiol. 2016, 131, 32–39. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhang, H.; Li, H.; Miao, X. Second-generation sequencing supply an effective way to screen RNAi targets in large scale for potential application in pest insect control. PLoS ONE 2011, 6, e18644. [Google Scholar] [CrossRef] [PubMed]
- Tkachev, A.V. Pyrethroid insecticides as analogs of the defensive chemicals of plants. Soros Obraz. Zh. 2004, 8, 56–63. [Google Scholar]
- Dias, N.; Stein, C.A. Antisense oligonucleotides: Basic concepts and mechanisms. Mol. Cancer Ther. 2002, 1, 347–355. [Google Scholar] [PubMed]
- Vaerman, J.L.; Moureau, P.; Deldime, F.; Lewalle, P.; Lammineur, C.; Morschhauser, F.; Martiat, P. Antisense oligodeoxyribonucleotides suppress hematologic cell growth through stepwise release of deoxyribonucleotides. Blood 1997, 90, 331–339. [Google Scholar] [PubMed]
- Koziolkiewicz, M.; Gendaszewska, E.; Maszewska, M.; Stein, C.A.; Stec, W.J. The mononucleotide-dependent, nonantisense mechanism of action of phosphodiester and phosphorothioate oligonucleotides depends upon the activity of an ecto-5′-nucleotidase. Blood 2001, 98, 995–1002. [Google Scholar] [CrossRef] [PubMed]
- Eckstein, F. Nucleoside phosphorothioates. J. Am. Chem. Soc. 1966, 88, 4292–4294. [Google Scholar] [CrossRef]
- Stein, C.A.; Cheng, Y.C. Antisense oligonucleotides as therapeutic agents–is the bullet really magical? Science 1993, 261, 1004–1012. [Google Scholar] [CrossRef] [PubMed]
- Stein, C.A. Controversies in the cellular pharmacology of oligodeoxynucleotides. Ciba Found. Symp. 1997, 209, 79–89. [Google Scholar] [CrossRef] [PubMed]
- Dean, N.M.; Bennet, C.F. Antisense oligonucleotide-based therapeutics for cancer. Oncogene 2003, 22, 9087–9096. [Google Scholar] [CrossRef] [PubMed]
- Crooke, S.T.; Vickers, T.; Lima, W.; Wu, H.J.; Crooke, S.T. Mechanisms of antisense drug action, an introduction. In Antisense Drug Technology—Principles, Strategies, and Application; CRC Press: Boca Raton, FL, USA, 2006; pp. 3–46. [Google Scholar]
- Frazier, K.S. Antisense oligonucleotide therapies: The promise and the challenges from a toxicologic pathologist’s perspective. Toxicol. Pathol. 2015, 43, 78–89. [Google Scholar] [CrossRef] [PubMed]
- Corey, D.R.; Abrams, J.M. Morpholino antisense oligonucleotides: Tools for investigating vertebrate development. Genome Biol. 2001, 2. [Google Scholar] [CrossRef]
- Summerton, J. Morpholino antisense oligomers: The case for an RNase H-independent structural type. Biochim. Biophys. Acta 1999, 1489, 141–158. [Google Scholar] [CrossRef]
- Draper, B.W.; Morcos, P.A.; Kimmel, C.B. Inhibition of zebrafish fgf8 pre-mRNA splicing with morpholino oligos: A quantifiable method for gene knockdown. Genesis 2001, 30, 154–156. [Google Scholar] [CrossRef] [PubMed]
- Summerton, J. Morpholinos and PNAs compared. Lett. Pept. Sci. 2003, 10, 215–236. [Google Scholar] [CrossRef]
- Eisen, J.S.; Smith, J.C. Controlling morpholino experiments: Don’t stop making antisense. Development 2008, 135, 1735–1743. [Google Scholar] [CrossRef] [PubMed]
- Larsen, H.J.; Bentin, T.; Nielsen, P.E. Antisense properties of peptide nucleic acid. Biochim. Biophys. Acta 1999, 1489, 159–166. [Google Scholar] [CrossRef]
- Choi, Y.; Metcalf, G.; Sleiman, M.H.; Vair-Turnbull, D.; Ladame, S. Oligonucleotide-templated reactions based on Peptide Nucleic Acid (PNA) probes: Concept and biomedical applications. Bioorg. Med. Chem. 2014, 22, 4395–4398. [Google Scholar] [CrossRef] [PubMed]
- Grunweller, A.; Hartmann, R.K. Locked nucleic acid oligonucleotides: The next generation of antisense agents? BioDrugs 2007, 21, 235–243. [Google Scholar] [CrossRef] [PubMed]
- Wahlestedt, C.; Salmi, P.; Good, L.; Kela, J.; Johnsson, T.; Hokfelt, T.; Broberger, C.; Porreca, F.; Lai, J.; Ren, K.; et al. Potent and nontoxic antisense oligonucleotides containing locked nucleic acids. Proc. Natl. Acad. Sci. USA 2000, 97, 5633–5638. [Google Scholar] [CrossRef] [PubMed]
- Gupta, N.; Fisker, N.; Asselin, M.C.; Lindholm, M.; Rosenbohm, C.; Ørum, H.; Elmén, J.; Seidah, N.G.; Straarup, E.M. A locked nucleic acid antisense oligonucleotide (LNA) silences PCSK9 and enhances LDLR expression in vitro and in vivo. PLoS ONE 2010, 5, e10682. [Google Scholar] [CrossRef] [PubMed]
- Braasch, D.A.; Corey, D.R. Locked nucleic acid (LNA): Fine-tuning the recognition of DNA and RNA. Chem. Biol. 2001, 8, 1–7. [Google Scholar] [CrossRef]
- Kurreck, J.; Wyszko, E.; Gillen, C.; Erdmann, V.A. Design of antisense oligonucleotides stabilized by locked nucleic acids. Nucleic Acids Res. 2002, 30, 1911–1918. [Google Scholar] [CrossRef] [PubMed]
- Greenberger, L.M.; Horak, I.D.; Filpula, D.; Sapra, P.; Westergaard, M.; Frydenlund, H.F.; Albæk, C.; Schrøder, H.; Ørum, H. A RNA antagonist of hypoxia-inducible factor-1alpha, EZN-2968, inhibits tumor cell growth. Mol. Cancer Ther. 2008, 7, 3598–3608. [Google Scholar] [CrossRef] [PubMed]
- Hansen, J.B.; Fisker, N.; Westergaard, M.; Kjaerulff, L.S.; Hansen, H.F.; Thrue, C.A.; Rosenbohm, C.; Wissenbach, M.; Ørum, H.; Koch, T. SPC3042: A proapoptotic survivin inhibitor. Mol. Cancer Ther. 2008, 7, 2736–2745. [Google Scholar] [CrossRef] [PubMed]
- Swayze, E.E.; Siwkowski, A.M.; Wancewicz, E.V.; Migawa, M.T.; Wyrzykiewicz, T.K.; Hung, G.; Monia, B.P.; Bennett, C.F. Antisense oligonucleotides containing locked nucleic acid improve potency but cause significant hepatotoxicity in animals. Nucleic Acids Res. 2007, 35, 687–700. [Google Scholar] [CrossRef] [PubMed]
- Stanton, R.; Sciabola, S.; Salatto, C.; Weng, Y.; Moshinsky, D.; Little, J.; Walters, E.; Kreeger, J.; DiMattia, D.; Chen, T.; et al. Chemical modification study of antisense gapmers. Nucleic Acid Ther. 2012, 22, 344–359. [Google Scholar] [CrossRef] [PubMed]
- Burnett, J.C.; Rossi, J.J. RNA-based therapeutics: Current progress and future prospects. Chem. Biol. 2012, 19, 60–71. [Google Scholar] [CrossRef] [PubMed]
- Hammond, S.M.; Bernstein, E.; Beach, D.; Hannon, G.J. An RNA-directed nuclease mediates post-transcriptional gene silencing in Drosophila cells. Nature 2000, 404, 293–296. [Google Scholar] [CrossRef] [PubMed]
- Hutvagner, G.; Zamore, P.D. A microRNA in a multiple-turnover RNAi enzyme complex. Science 2002, 297, 2056–2060. [Google Scholar] [CrossRef] [PubMed]
- Garzon, R.; Marcucci, G.; Croce, C.M. Targeting microRNAs in cancer: Rationale, strategies and challenges. Nat. Rev. Drug Discov. 2010, 9, 775–789. [Google Scholar] [CrossRef] [PubMed]
- Wong, E.; Goldberg, T. Mipomersen (kynamro): A novel antisense oligonucleotide inhibitor for the management of homozygous familial hypercholesterolemia. Pharm. Ther. 2014, 39, 119–122. [Google Scholar]
- Toth, P.P. Emerging LDL therapies: Mipomersen-antisense oligonucleotide therapy in the management of hypercholesterolemia. J. Clin. Lipidol. 2013, 7, S6–S10. [Google Scholar] [CrossRef] [PubMed]
- Brautbar, A.; Leary, E.; Rasmussen, K.; Wilson, D.P.; Steiner, R.D.; Virani, S. Genetics of familial hypercholesterolemia. Curr. Atheroscler. Rep. 2015, 17, 491. [Google Scholar] [CrossRef] [PubMed]
- Toth, P.P. Antisense therapy and emerging applications for the management of dyslipidemia. J. Clin. Lipidol. 2011, 5, 441–449. [Google Scholar] [CrossRef] [PubMed]
- Visser, M.E.; Witztum, J.L.; Stroes, E.S.; Kastelein, J.J. Antisense oligonucleotides for the treatment of dyslipidaemia. Eur. Heart J. 2012, 33, 1451–1458. [Google Scholar] [CrossRef] [PubMed]
- Geary, R.S.; Baker, B.F.; Crooke, S.T. Clinical and preclinical pharmacokinetics and pharmacodynamics of mipomersen (Kynamro): A second-generation antisense oligonucleotide inhibitor of apolipoprotein B. Clin. Pharmacokinet. 2015, 54, 133–146. [Google Scholar] [CrossRef] [PubMed]
- Hua, Y.; Sahaski, K.; Hung, G.; Rigo, F.; Passini, M.; Bennett, C.; Krainer, A. Antisense correction of SMN2 splicing in the CNS rescues necrosis in a type III SMA mouse model. Genes Dev. 2010, 24, 1634–1644. [Google Scholar] [CrossRef] [PubMed]
- Lefebvre, S.; Burlet, P.; Liu, Q.; Bertrandy, S.; Clermont, O.; Munnich, A.; Dreyfuss, G.; Melki, J. Correlation between severity and SMN protein level in spinal muscular atrophy. Nat. Genet. 1997, 16, 265–269. [Google Scholar] [CrossRef] [PubMed]
- Hua, Y.; Liu, Y.H.; Sahashi, K.; Rigo, F.; Bennett, C.F.; Krainer, A.R. Motor neuron cell-nonautonomous rescue of spinal muscular atrophy phenotypes in mild and severe transgenic mouse models. Genes Dev. 2015, 29, 288–297. [Google Scholar] [CrossRef] [PubMed]
- Hua, Y.; Vickers, T.A.; Okunola, H.L.; Bennett, C.F.; Krainer, A.R. Antisense masking of an hnRNP A1/A2 intronic splicing silencer corrects SMN2 splicing in transgenic mice. Am. J. Hum. Genet. 2008, 82, 834–848. [Google Scholar] [CrossRef] [PubMed]
- Meijboom, K.E.; Wood, M.J.A.; McClorey, G. Splice-switching therapy for spinal muscular atrophy. Genes (Basel) 2017, 8, 161. [Google Scholar] [CrossRef] [PubMed]
- Chiriboga, C.A.; Swoboda, K.J.; Darras, B.T.; Iannaccone, S.T.; Montes, J.; Darryl, C.; Norris, D.A.; Bennett, C.F.; Bishop, K.M. Results from a phase 1 study of nusinersen (ISIS-SMNRx) in children with spinal muscular atrophy. Neurology 2016, 86, 890–897. [Google Scholar] [CrossRef] [PubMed]
- Mendell, J.R.; Shilling, C.; Leslie, N.D.; Flanigan, K.M.; al-Dahhak, R.; Gastier-Foster, J.; Kneile, K.; Dunn, D.M.; Duval, B.; Aoyagi, A.; et al. Evidence-based path to newborn screening for Duchenne muscular dystrophy. Ann. Neurol. 2012, 71, 304–313. [Google Scholar] [CrossRef] [PubMed]
- Emery, A.E. Population frequencies of inherited neuromuscular diseases: A world survey. Neuromuscul. Disord. 1991, 1, 19–29. [Google Scholar] [CrossRef]
- Lim, K.R.; Maruyama, R.; Yokota, T. Eteplirsen in the treatment of Duchenne muscular dystrophy. Drug Des. Dev. Ther. 2017, 11, 533–545. [Google Scholar] [CrossRef] [PubMed]
- Koenig, M.; Monaco, A.P.; Kunkel, L.M. The complete sequence of dystrophin predicts a rod-shaped cytoskeletal protein. Cell 1988, 53, 219–228. [Google Scholar] [CrossRef]
- Ervasti, J.M. Dystrophin, its interactions with other proteins, and implications for muscular dystrophy. Biochim. Biophys. Acta 2007, 1772, 108–117. [Google Scholar] [CrossRef] [PubMed]
- Arechavala-Gomeza, V.; Graham, I.R.; Popplewell, L.J.; Adams, A.M.; Aartsma-Rus, A.; Kinali, M.; Morgan, J.E.; Van Deutekom, J.C.; Wilton, S.D.; Dickson, G.; et al. Comparative analysis of antisense oligonucleotide sequences for targeted skipping of exon 51 during dystrophin pre-mRNA splicing in human muscle. Hum. Gene Ther. 2007, 18, 798–810. [Google Scholar] [CrossRef] [PubMed]
- Kole, R.; Krieg, A.M. Exon skipping therapy for Duchenne muscular dystrophy. Adv. Drug Deliv. Rev. 2015, 87, 104–107. [Google Scholar] [CrossRef] [PubMed]
- Simonson, B.; Das, S. MicroRNA therapeutics: The next magic bullet? Mini Rev. Med. Chem. 2011, 15, 467–474. [Google Scholar] [CrossRef]
- Gebert, L.F.; Rebhan, M.A.; Crivelli, S.E.; Denzler, R.; Stoffel, M.; Hall, J. Miravirsen (SPC3649) can inhibit the biogenesis of miR-122. Nucleic Acids Res. 2014, 42, 609–621. [Google Scholar] [CrossRef] [PubMed]
- Hajarizadeh, B.; Grebely, J.; Dore, G.J. Epidemiology and natural history of HCV infection. Nat. Rev. Gastroenterol. Hepatol. 2013, 10, 553–562. [Google Scholar] [CrossRef] [PubMed]
- Lauer, G.M.; Walker, B.D. Hepatitis C virus infection. N. Engl. J. Med. 2001, 345, 41–52. [Google Scholar] [CrossRef] [PubMed]
- Janssen, H.L.; Reesink, H.W.; Lawitz, E.J.; Zeuzem, S.; Rodriguez-Torres, M.; Patel, K.; van der Meer, A.J.; Patick, A.K.; Chen, A.; Zhou, Y.; et al. Treatment of HCV infection by targeting microRNA. N. Engl. J. Med. 2013, 368, 1685–1694. [Google Scholar] [CrossRef] [PubMed]
- Hildebrandt-Eriksen, E.S.; Bagger, Y.Z.; Knudsen, T.B.; Petri, A.; Persson, R.; Boergesen, H.M.; McHulchison, J.G.; Levin, A.A. A unique therapy for HCV inhibits microRNA-122 in humans and results in HCV RNA suppression in chronically infected chimpanzees: Results from primate and first-in-human studies. Hepatology 2009, 50, 12A. [Google Scholar]
- Lanford, R.E.; Hildebrandt-Eriksen, E.S.; Petri, A.; Persson, R.; Lindow, M.; Munk, M.E.; Kauppinen, S.; Ørum, H. Therapeutic silencing of microRNA-122 in primates with chronic hepatitis C virus infection. Science 2010, 327, 198–201. [Google Scholar] [CrossRef] [PubMed]
- Sanchis, V. From microbial sprays to insect-resistant transgenic plants: History of the biopesticide Bacillus thuringiensis. A review. Agron. Sustain. Dev. 2011, 31, 217–231. [Google Scholar] [CrossRef]
- Oerke, E.C.; Dehne, H.W. Safeguarding production-losses in major crops and the role of crop protection. Crop Protect. 2004, 23, 275–285. [Google Scholar] [CrossRef]
- Daly, H.; Doyen, J.T.; Purcell, A.H. Introduction to Insect Biology and Diversity; Oxford University Press: New York, NY, USA, 1998; pp. 1–9. [Google Scholar]
- Weston, D.P.; Poynton, H.C.; Wellborn, G.A.; Lydy, M.J.; Blalock, B.J.; Sepulveda, M.S.; Colbourne, J.K. Multiple origins of pyrethroid insecticide resistance across the species complex of a nontarget aquatic crustacean, Hyalella azteca. Proc. Natl. Acad. Sci. USA 2013, 110, 16532–16537. [Google Scholar] [CrossRef] [PubMed]
- Joga, M.R.; Zotti, M.J.; Smagghe, G.; Christiaens, O. RNAi efficiency, systemic properties, and novel delivery methods for pest insect control: What we know so far. Front. Physiol. 2016, 7, 553. [Google Scholar] [CrossRef] [PubMed]
- Roberts, A.F.; Devos, Y.; Lemgo, G. Biosafety research for non-target organism risk assessment of RNAi-based GE plants. Front. Plant Sci. 2015, 6, 958. [Google Scholar] [CrossRef] [PubMed]
- Mao, Y.B.; Cai, W.J.; Wang, J.W.; Hong, G.J.; Tao, X.Y.; Wang, L.J.; Huang, Y.P.; Chen, X.Y. Silencing a cotton bollworm P450 monooxygenase gene by plant-mediated RNAi impairs larval tolerance of gossypol. Nat. Biotechnol. 2017, 25, 1307–1313. [Google Scholar] [CrossRef] [PubMed]
- Ramesh, S.V. Non-coding RNAs in crop genetic modification: Considerations and predictable environmental risk assessments (ERA). Mol. Biotechnol. 2013, 55, 87–100. [Google Scholar] [CrossRef] [PubMed]
- Ramon, M.; Devos, Y.; Lanzoni, A.; Liu, Y.; Gomes, A.; Gennaro, A.; Waigmann, E. RNAi-based GM plants: food for thought for risk assessors. Plant Biotechnol. J. 2014, 12, 1271–1273. [Google Scholar] [CrossRef] [PubMed]
- Huvenne, H.; Smagghe, G. Mechanisms of dsRNA uptake in insects and potential of RNAi for pest control: A review. J. Insect Physiol. 2010, 56, 227–235. [Google Scholar] [CrossRef] [PubMed]
- Siomi, H.; Siomi, M.C. Posttranscriptional regulation of microRNA biogenesis in animals. Mol. Cell 2011, 38, 323–332. [Google Scholar] [CrossRef] [PubMed]
- Agrawal, N.; Dasaradhi, P.V.N.; Mohmmed, A.; Malhotra, P.; Bhatnagar, R.K.; Mukherjee, S.K. RNA interference: Biology, mechanism, and applications. Microbiol. Mol. Biol. Rev. 2003, 67, 657–685. [Google Scholar] [CrossRef] [PubMed]
- Dillin, A. The specifics of small interfering RNA specificity. Proc. Natl. Acad. Sci. USA 2003, 100, 6289–6291. [Google Scholar] [CrossRef] [PubMed]
- Bolognesi, R.; Ramaseshadri, P.; Anderson, J.; Bachman, P.; Clinton, W.; Flannagan, R.; Ilagan, O.; Lawrence, C.; Levine, S.; Moar, W.; et al. Characterizing the mechanism of action of double-stranded RNA activity against western corn rootworm (Diabrotica virgifera Le Conte). PLoS ONE 2012, 7, e47534. [Google Scholar] [CrossRef] [PubMed]
- Ivashuta, S.; Zhang, Y.; Wiggins, B.E.; Ramaseshadri, P.; Segers, G.C.; Johnson, S.; Meyer, S.E.; Kerstetter, R.A.; McNulty, B.C.; Bolognesi, R.; et al. Environmental RNAi in herbivorous insects. RNA 2015, 21, 840–850. [Google Scholar] [CrossRef] [PubMed]
- Palli, S.R. RNA interference in Colorado potato beetle: Steps toward development of dsRNA as a commercial insecticide. Curr. Opin. Insect Sci. 2004, 6, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Turner, C.T.; Davy, M.W.; MacDiarmid, R.M.; Plummer, K.M.; Birch, N.P.; Newcomb, R.D. RNA interference in the light brown apple moth, Epiphyas postvittana (Walker) induced by double-stranded RNA feeding. Insect Mol. Biol. 2006, 15, 383–391. [Google Scholar] [CrossRef] [PubMed]
- Araujo, R.N.; Santos, A.; Pinto, F.S.; Gontijo, N.F.; Lehane, M.J.; Pereira, M.H. RNA interference of the salivary gland Nitrophorin 2 in the triatomine bug Rhodnius prolixus (Hemiptera: Reduviidae) by dsRNA ingestion or injection. Insect Biochem. Mol. Biol. 2006, 36, 683–693. [Google Scholar] [CrossRef] [PubMed]
- Yu, N.; Christiaens, O.; Liu, J.; Niu, J.; Cappelle, K.; Caccia, S.; Huvenne, H.; Smagghe, G. Delivery of dsRNA for RNAi in insects: An overview and future directions. Insect Sci. 2013, 20, 4–14. [Google Scholar] [CrossRef] [PubMed]
- Oberemok, V.V.; Laikova, K.V.; Zaitsev, A.S.; Nyadar, P.M.; Gninenko, Y.I.; Gushchin, V.A.; Makarov, V.V.; Agranovsky, A.A. Topical treatment of LdMNPV-infected gypsy moth larvae with 18 nucleotides long antisense fragment from LdMNPV IAP3 gene triggers higher levels of apoptosis in infected cells and mortality of the pest. J. Plant Protect. Res. 2017, 57, 18–24. [Google Scholar] [CrossRef]
- Oberemok, V.V.; Nyadar, P.M. Investigation of mode of action of DNA insecticides on the basis of LdMNPV IAP-3 gene. Turk. J. Biol. 2015, 39, 258–264. [Google Scholar] [CrossRef]
- Oberemok, V.V.; Skorokhod, O.A. Single-stranded DNA fragments of insect-specific nuclear polyhedrosis virus act as selective DNA insecticides for gypsy moth control. Pest. Biochem. Physiol. 2014, 113, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Grayson, K.L.; Parry, D.; Faske, T.M.; Hamilton, A.; Tobin, P.C.; Agosta, S.J.; Johnson, D.M. Performance of wild and laboratory-reared gypsy moth (Lepidoptera: Erebidae): A comparison between foliage and artificial diet. Environ. Entomol. 2015, 44, 864–873. [Google Scholar] [CrossRef] [PubMed]
- Chen, F.; Shi, J.; Luo, Y.; Sun, S.; Pu, M. Genetic characterization of the gypsy moth from China (Lepidoptera, Lymantriidae) using inter simple sequence repeats markers. PLoS ONE 2013, 8, e73017. [Google Scholar] [CrossRef] [PubMed]
- Alalouni, U.; Schädler, M.; Brandl, R. Natural enemies and environmental factors affecting the population dynamics of the gypsy moth. J. Appl. Entomol. 2013, 137, 721–738. [Google Scholar] [CrossRef]
- Pitt, J.P.; Regniere, J.; Worner, S. Risk assessment of gypsy moth, Lymantria dispar (L), in New Zealand based on phenology modeling. Int. J. Biometeorol. 2004, 51, 295–305. [Google Scholar] [CrossRef] [PubMed]
- Cerio, R.J.; Vandergaast, R.; Friesen, P.D. Host insect inhibitor-of-apoptosis SfIAP functionally replaces baculovirus IAP but is differentially regulated by its N-terminal leader. J. Virol. 2010, 84, 11448–11460. [Google Scholar] [CrossRef] [PubMed]
- Rumble, J.M.; Duckett, C. Diverse functions within the IAP family. J. Cell Sci. 2008, 121, 3505–3507. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, M.; Yamada, H.; Ito, H.; Kobayashi, M. Baculovirus IAP1 induces caspase-dependent apoptosis in insect cells. J. Gen. Virol. 2011, 92, 2654–2663. [Google Scholar] [CrossRef] [PubMed]
- Oberemok, V.V.; Nyadar, P.; Zaytsev, O.; Levchenko, N.; Shiyntum, H.; Omelchenko, O. Pioneer evaluation of the possible side effects of the DNA insecticides on wheat (Triticum aestivum L.). Int. J. Biochem. Biophys. 2013, 1, 57–63. [Google Scholar] [CrossRef]
- Zaitsev, A.S.; Omel’chenko, O.V.; Nyadar, P.M.; Oberemok, V.V. Influence of DNA oligonucleotides used as insecticides on biochemical parameters of Quercus robur and Malus domestica. Bull. Transilv. Univ. Brasov 2015, 8, 37–46. [Google Scholar]
- Thorp, H.H. The importance of being r: Greater oxidative stability of RNA compared with DNA. Chem. Biol. 2000, 7, 33–36. [Google Scholar] [CrossRef]
- Perry, W.B.; Christiansen, T.A.; Perry, S.A. Response of soil and leaf litter microarthropods to forest application of diflubenzuron. Ecotoxicology 1997, 6, 87–99. [Google Scholar] [CrossRef]
- Msangi, S.; Lyatuu, E.; Kweka, E. Field and laboratory evaluation of bioefficacy of an insect growth regulator (Dimilin) as a larvicide against mosquito and housefly larvae. J. Trop. Med. 2011, 2011, 394541. [Google Scholar] [CrossRef] [PubMed]

| ASOs | Ability to Penetrate Cells | Stability in Vivo | Binding to Target mRNA | Level of Efficiency | Non-Target Effects |
|---|---|---|---|---|---|
| Phosphophodiesters (antisense RNA and DNA, dsRNA) | Moderate | Moderate | Moderate | Moderate | Moderate |
| First generation (phosphorothioates, methylphosphonates) | Moderate | Higher nuclease stability | Lower binding capacity | Higher effectiveness | More pronounced non-specific effects |
| Second generation (2′-O-methyl and 2′-O-methyloxyethyl oligonucleotides) | Moderate | Much higher nuclease stability | Higher binding capacity | Higher effectiveness | More pronounced non-specific effects |
| Third generation (locked nucleic acids, peptide nucleic acids, morpholino oligomers) | Higher penetration ability | Nuclease resistant | Much higher binding capacity | Much higher effectiveness | Moderate |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oberemok, V.V.; Laikova, K.V.; Repetskaya, A.I.; Kenyo, I.M.; Gorlov, M.V.; Kasich, I.N.; Krasnodubets, A.M.; Gal’chinsky, N.V.; Fomochkina, I.I.; Zaitsev, A.S.; et al. A Half-Century History of Applications of Antisense Oligonucleotides in Medicine, Agriculture and Forestry: We Should Continue the Journey. Molecules 2018, 23, 1302. https://doi.org/10.3390/molecules23061302
Oberemok VV, Laikova KV, Repetskaya AI, Kenyo IM, Gorlov MV, Kasich IN, Krasnodubets AM, Gal’chinsky NV, Fomochkina II, Zaitsev AS, et al. A Half-Century History of Applications of Antisense Oligonucleotides in Medicine, Agriculture and Forestry: We Should Continue the Journey. Molecules. 2018; 23(6):1302. https://doi.org/10.3390/molecules23061302
Chicago/Turabian StyleOberemok, Volodymyr V., Kateryna V. Laikova, Anna I. Repetskaya, Igor M. Kenyo, Mikhail V. Gorlov, Igor N. Kasich, Alisa M. Krasnodubets, Nikita V. Gal’chinsky, Iryna I. Fomochkina, Aleksei S. Zaitsev, and et al. 2018. "A Half-Century History of Applications of Antisense Oligonucleotides in Medicine, Agriculture and Forestry: We Should Continue the Journey" Molecules 23, no. 6: 1302. https://doi.org/10.3390/molecules23061302
APA StyleOberemok, V. V., Laikova, K. V., Repetskaya, A. I., Kenyo, I. M., Gorlov, M. V., Kasich, I. N., Krasnodubets, A. M., Gal’chinsky, N. V., Fomochkina, I. I., Zaitsev, A. S., Bekirova, V. V., Seidosmanova, E. E., Dydik, K. I., Meshcheryakova, A. O., Nazarov, S. A., Smagliy, N. N., Chelengerova, E. L., Kulanova, A. A., Deri, K., ... Kubyshkin, A. V. (2018). A Half-Century History of Applications of Antisense Oligonucleotides in Medicine, Agriculture and Forestry: We Should Continue the Journey. Molecules, 23(6), 1302. https://doi.org/10.3390/molecules23061302









