In vivo, haemostasis and blood circulation for removing blood stasis are two contradictory activities. However, both activities objectively exist at the same time. In gynaecological diseases or after surgery, wounds require stopping bleeding, but the medicines used cannot increase patient blood viscosity, which will form thrombosis; at the same time, the medicines are needed to slowly dissipate the congestion in blood circulation and promote the restoration of the patient’s body.
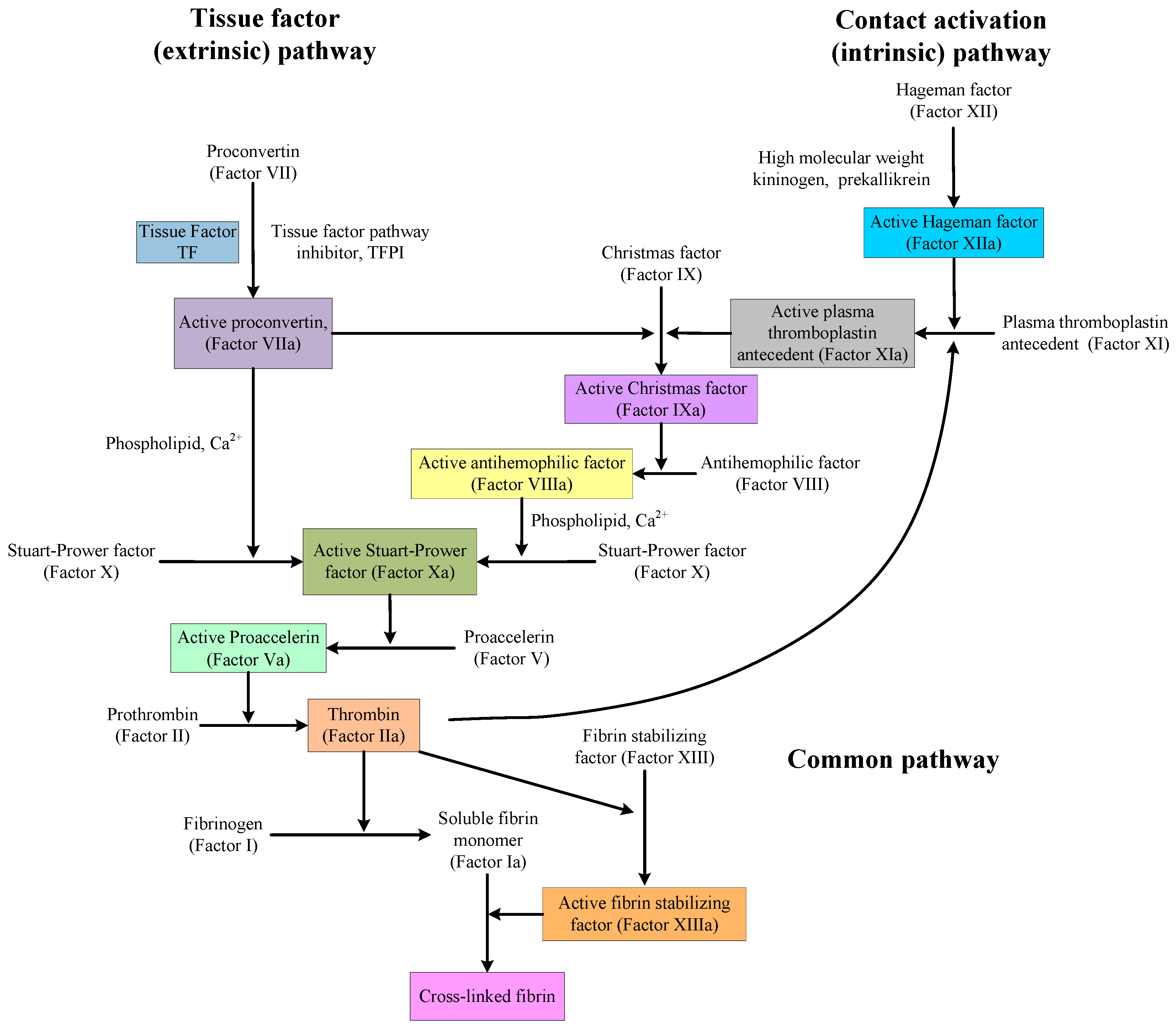
There is also a paradox in discovering and designing medicines that affect the mechanisms of anticoagulation and procoagulation. As shown in
Figure 1, the coagulation pathway is very complex, but the situation is different near thrombin. Indeed, in that area, there is single factor control. Therefore, in the design of medicines or the software simulation of medicine activity, thrombin or other proteins and factors in that area are, in many cases, used as the targets. However, no matter how to design or evaluate medicines, all obtained compounds and precursors will be used in actual blood systems to evaluate their activities, and the activities of these compounds are a combination of their performance on every protein and factor in the coagulation pathway. The compounds, showing good anticoagulation activity with thrombin, may not have the same anticoagulation activity or procoagulant effect in blood.
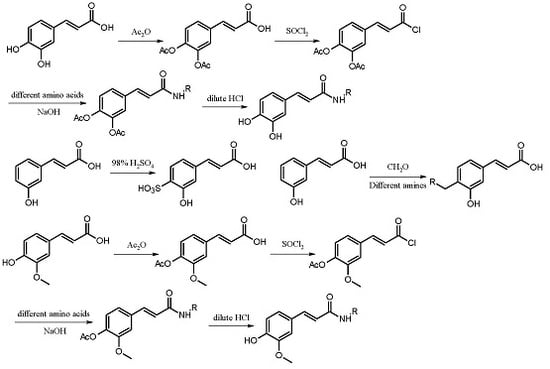
In the present study, the anticoagulant and procoagulant activities of three phenolic acid derivative series, 18 PADs, were investigated. Comparing the results of molecular docking and SPR with those of five blood assays, we found that using molecular docking and SPR to assist the molecule design had some drawbacks (one of the reasons was the paradox mentioned above). However, the experimental data also showed that using molecular docking and SPR could generate some important information for molecule design.
3.1. Virtual Screening
Virtual screening of chemical libraries, as an alternative approach for the identification of new lead compounds in drug discovery, has been widely used [
47]. As the structures of biomolecules deposited in the Protein Data Bank (PDB) have substantially increased in the past decades, searching for the targets of a given drug or small compound (also known as inverse screening, target fishing, off-target prediction, etc.) has become a useful approach [
48]. In PharmMapper tool, 18 newly-synthesized PADs were aligned to 2241 human protein targets, among which the top 300 targets were analysed. From the results, three types, a total of seven protein targets, were screened out. The first type was targets associated with coagulation, including thrombin, prothrombin, coagulation factor X, and fibrinogen. The second type was targets associated with anticoagulant, including antithrombin-III and plasminogen. The third type was the two-way regulatory target, vitamin K-dependent protein Z.
3.2. AutoDock Docking
The results of the docking study for all 18 synthesized PADs with seven protein targets, including thrombin (3RLW), prothrombin (1YPK), coagulation factor X (2J4I), fibrinogen (3GHG), antithrombin-III (2ANT), plasminogen (1QRZ), and vitamin K-dependent protein Z (3H5C), are listed in
Table 7.
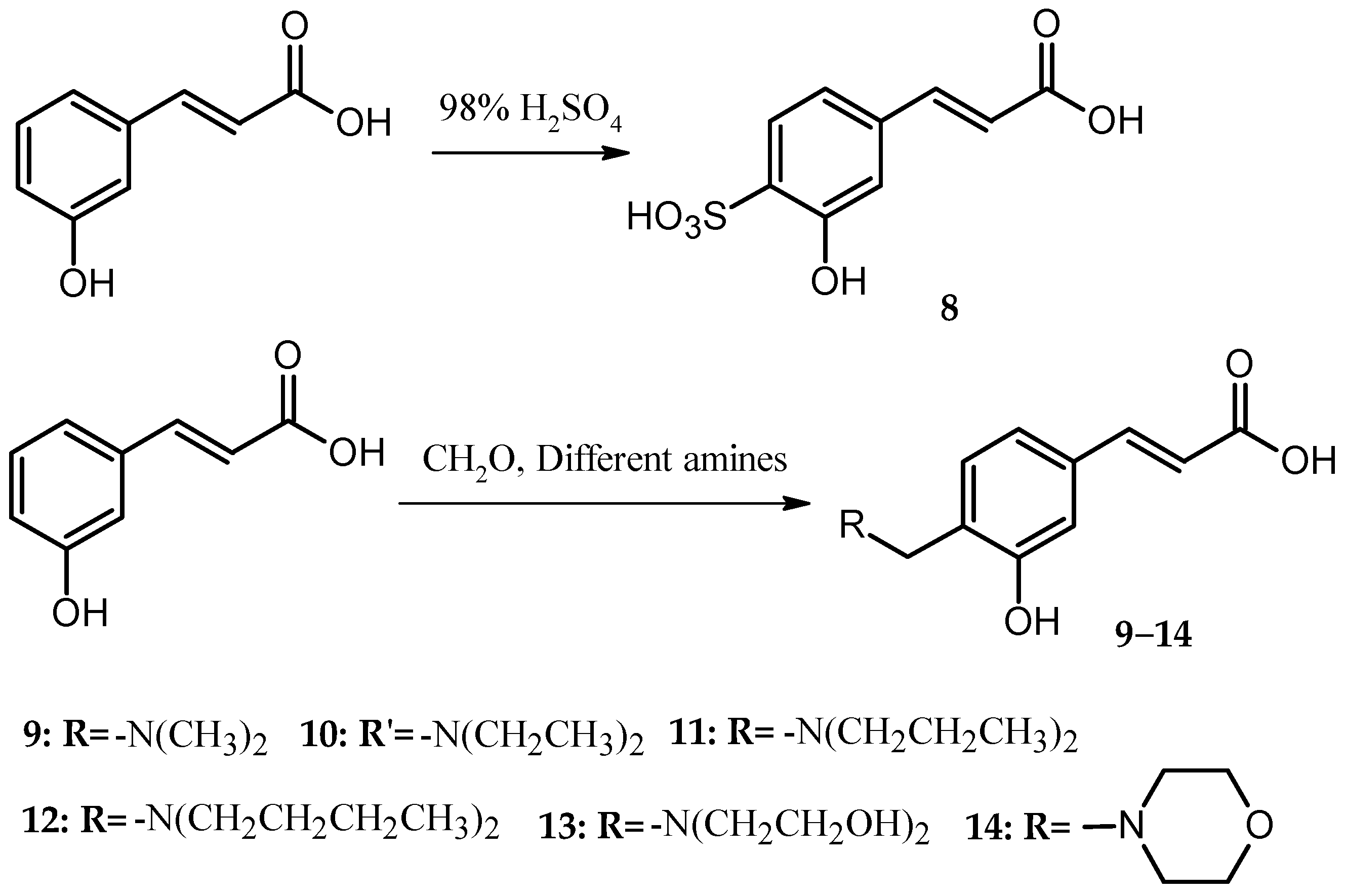
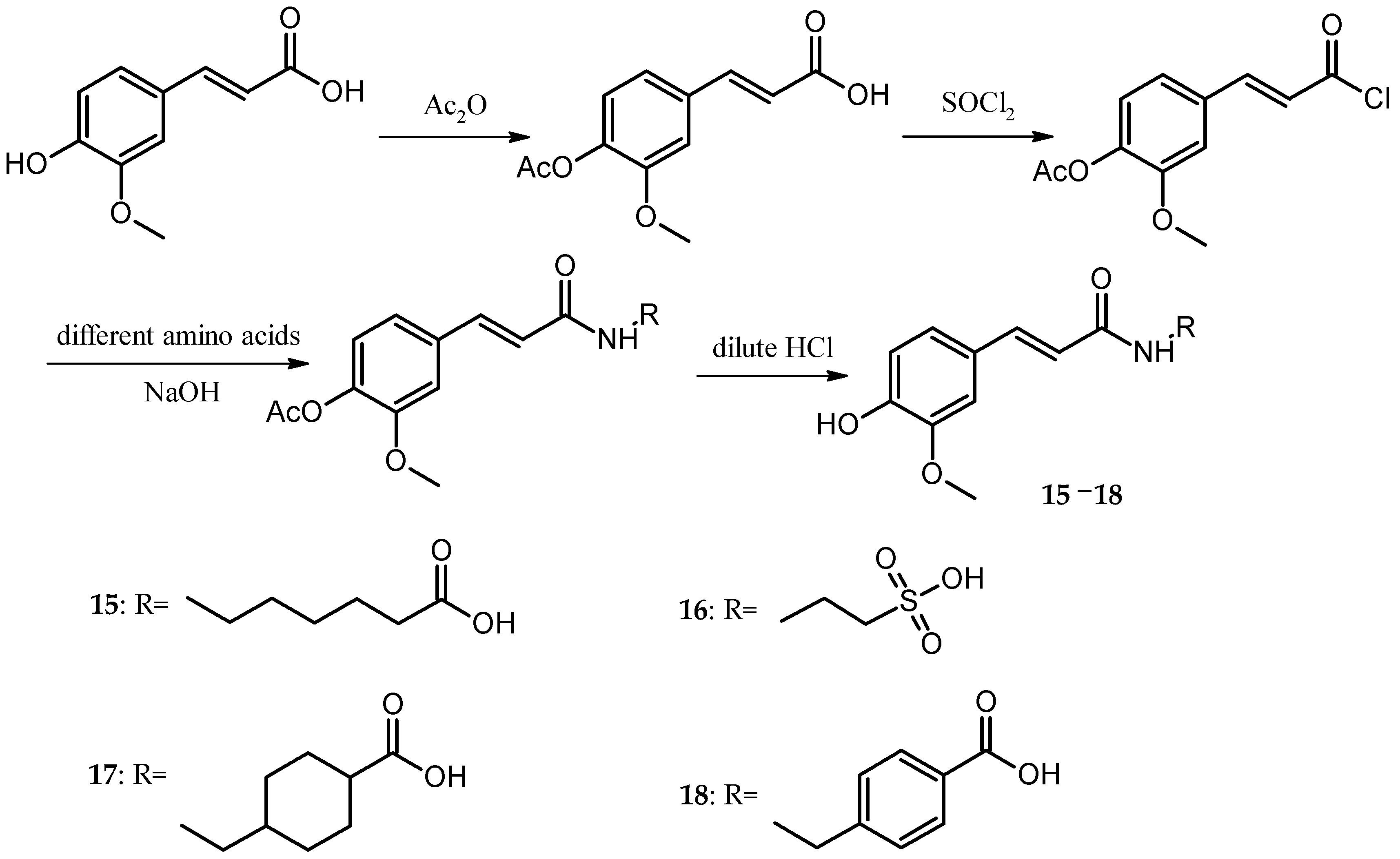
For the caffeic acid derivative series (Cds. 1–7), the binding energies of Cd. 5 with protein targets were lower than those of the other caffeic acid derivatives. Cds. 8–14 were p-hydroxycinnamic acid derivatives, and the binding energy data for some compounds revealed three important findings: The docking energies of Cd. 8 with the seven protein targets were very low (six of the seven energy values were the lowest compared with those of the other 17 compounds, and the remaining value was the second to the lowest value). In contrast, Cd. 12 had the best docking results among the 18 compounds (although the binding energy with 1QRZ was not the highest value, this value was still the second highest value). Except for Cd. 8, all other p-hydroxycinnamic acid derivatives had better docking results with 3GHG and 2ANT, and binding with these two protein targets would release greater than −20 kJ∙mol−1 of energy. For the four ferulic acid derivatives (Cds. 15–18), one derivative, Cd. 16, showed lower binding energies with the protein targets.
Three different series of PADs, Cds. 5, 8, and 16, had worse docking results and a similar structure fraction, the sulfonic acid group. These results suggest that in the docking calculation, the sulfonic acid group could affect the binding between PADs and protein targets, particularly in connection with the benzene ring in the PAD structure. Since the sulfonic acid group could withdraw electrons out of the benzene ring, the low electron density generated worse docking results for Cd. 8.
A comparison of the average of each protein target with the 18 PADs revealed four coagulation targets with better docking results, particularly 3RLW with caffeic acid derivatives and 3GHG with p-hydroxycinnamic acid derivatives.
3.3. Surface Plasmon Resonance Results
Based on the AutoDock docking results, thrombin and fibrinogen were the protein targets used in SPR analysis.
Autodock can predict the combination of small molecules and proteins by theoretical calculation. Moreover, to obtain the results, the calculation is based on the electronic distribution of the molecule itself (such that the electron-rich regions in the electrostatic potential map of the compound will combine with the electron deficient regions of the protein) and intermolecular interactions, including van der Waals, electrostatic, and bond interactions. During the calculation, the effects of the external environment are considered. Thus, the AutoDock results were not consistent with the results of surface plasmon resonance (SPR), listed in
Table 8.
In
Table 8, 12 of the 18 PADs could combine with thrombin and show better results than those obtained with fibrinogen. However, in AutoDock (
Table 7), the averages of PADs with the two proteins were very close. AutoDock predicted that Cd.
12 had the lowest binding energy with thrombin, but in fact Cd.
13 had the smallest
KD, indicating that this compound had the best combination with thrombin.
For
p-hydroxycinnamic acid derivatives, AutoDock predicted that Cds.
9–
12 would have a better combination with thrombin when the side chain was longer. Indeed, as shown in
Table 8, Cd.
9 had a smaller
KD due to its shorter side chain. The reason for this finding was that when the side chain got increasingly longer, the hydrophobicity of the molecule increased, and because the conditions of the assays and the environment in actual human bodies involved water as the major solvent, increasing hydrophobicity would decrease the interactions between small molecules and proteins. This phenomenon was very obvious in the SPR results for Cd.
13. Since the structure of Cd.
13 had two additional hydroxyl groups, which could decrease the hydrophobicity and increase its hydrophilicity, Cd.
13 would have better combination results with proteins under the assay conditions (water was the major solvent).
In AutoDock, p-hydroxycinnamic acid derivatives had the lowest average binding energy with fibrinogen compared with the other two derivatives. In SPR, although only two p-hydroxycinnamic acid derivatives had KD values, p-hydroxycinnamic acid derivatives remained the best among the three PADs series.
Although SPR provided a realistic combination of small molecules and proteins, the results did not show the effects on protein activities. To examine the changes (enhance or inhibit activity) of protein activities after combination, plasma recalcification time (PRT), PT, APTT, FIB, and thrombin time (TT) were examined.
3.4. PRT, PT, APTT, FIB, and TT Results
Five blood assays were used to determine the procoagulant and anticoagulation activities of 18 PADs, and the results are listed in
Table 9.
Cds.
8 and
13 had the best combination results with thrombin in SPR and showed significant anticoagulation activities in all five assays, which might indicate that these compounds could inhibit thrombin activity in combination and make the conversion from soluble fibrinogen to insoluble fibrin without thrombin promotion. Cd.
16 was the second compound in the 18 PADs, which had significant anticoagulation activity in all five blood assays. However, this compound did not combine with thrombin and showed weak combination with fibrinogen (large
KD value). These results showed that the anticoagulation activity of Cd.
16 might be caused by the combination with other proteins or factors in the coagulation (clotting) cascade (See
Figure 1). Cd.
9 showed combined results with thrombin and fibrinogen in SPR, but the results with fibrinogen had a large
KD value, indicating that the combination was extremely weak. No obvious effect in the FIB assay confirmed that Cd.
9 had little effect on fibrinogen, and its anticoagulation activity might be caused by the combination with thrombin.
Although Cds. 8, 9, 13, and 16 all had the better anticoagulation activities among the 18 PADs, these compounds belonged to two different phenolic acid derivatives: Cds. 8, 9, and 13 were members of the p-hydroxycinnamic acid series and Cd. 16 was a member of the ferulic acid series, indicating that the anticoagulation activities of these compounds affected different proteins or factors in the coagulation cascade. Even in the same phenolic acid derivative, Cds. 8 and 13, showing the best anticoagulation activity, had different SPR results.
Both Cds.
16 and
5 had good anticoagulation activity, and the former was the best among the 18 PADs (See
Table 9). This activity difference was triggered by the only difference in their structures, the 3-position of the benzene ring. The hydroxyl group in the 3-position of the benzene ring was changed to a methoxy group. This change made Cd.
16 bigger and much easier to fit the receptor (proteins or factors in the coagulation cascade).
Cd.
4 had the best procoagulant activity among the 18 PADs. However, the combination (see
Table 8) with thrombin was very weak, and no combination was observed with fibrinogen. Cds.
6 and
7 had similar results compared to Cd.
4. These phenomena indicated that the procoagulant activities of Cds.
4,
6, and
7 were achieved via combining with other proteins or factors in the coagulation cascade rather than with thrombin and fibrinogen. This finding might be due to their structures. All three compounds had very long R groups in their structures, and the distances between the acylamino N and the carbonyl non-hydrogen atoms were very close (see
Table 4 and the
Supplementary Information). Cd.
4 showed the best procoagulant activity among the 18 PADs, indicating that its R structure was suitable to combine with some proteins or factors in the coagulation cascade and subsequently promote procoagulation. However, the three key distances of Cds.
6 and
7 were shorter than those of Cd.
4, particularly N–hydroxyl O of −COOH and N–C of −COOH, suggesting that the combinations between Cds.
6 or
7 and the proteins or factors were less efficient compared with the combination between Cd.
4 and the proteins or factors.
Mackman [
50] classified anticoagulants into three main types: Vitamin K antagonists, heparins, and direct inhibitors of factor Xa and thrombin. For the first type, Vitamin K antagonists function by inhibiting the enzyme vitamin K epoxide reductase, which uses vitamin K to post-translationally modify several coagulation proteins (factor VII, factor IX, factor X and prothrombin). The second type, heparin, binds to the protein antithrombin and markedly increases the ability of this protein to inhibit factor Xa and thrombin. The third type has two typical examples, lepirudin and desirudin. However, Correia-da-Silva and co-workers [
51] demonstrated that some persulfated small molecules might be a fourth type of anticoagulant, with dual anticoagulant and antiplatelet effects. The structures of the persulfated compounds in [
50] and the PADs synthesized in the present study showed some similarities, such as benzene rings, phenolic hydroxyl groups, carbonyl groups, and sulfo groups. This evidence suggests that the PADs might have similar functions with dual effects.