Exploring the Degradation of Ibuprofen by Bacillus thuringiensis B1(2015b): The New Pathway and Factors Affecting Degradation
Abstract
:1. Introduction
2. Results and Discussion
2.1. Factors Affecting Ibuprofen Degradation
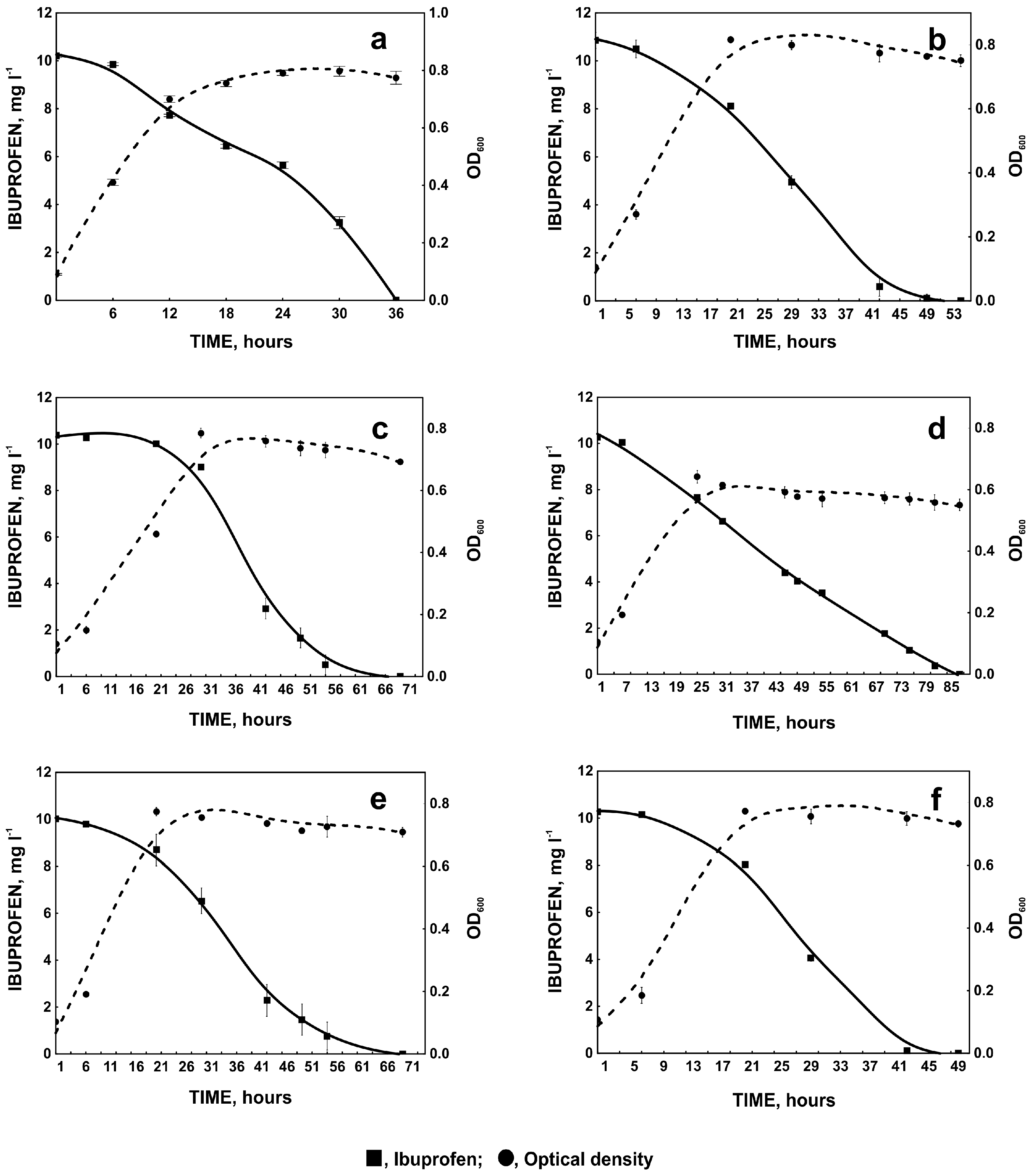
2.1.1. Influence of pH and Temperature on Ibuprofen Degradation
2.1.2. Heavy Metals and Ibuprofen Degradation
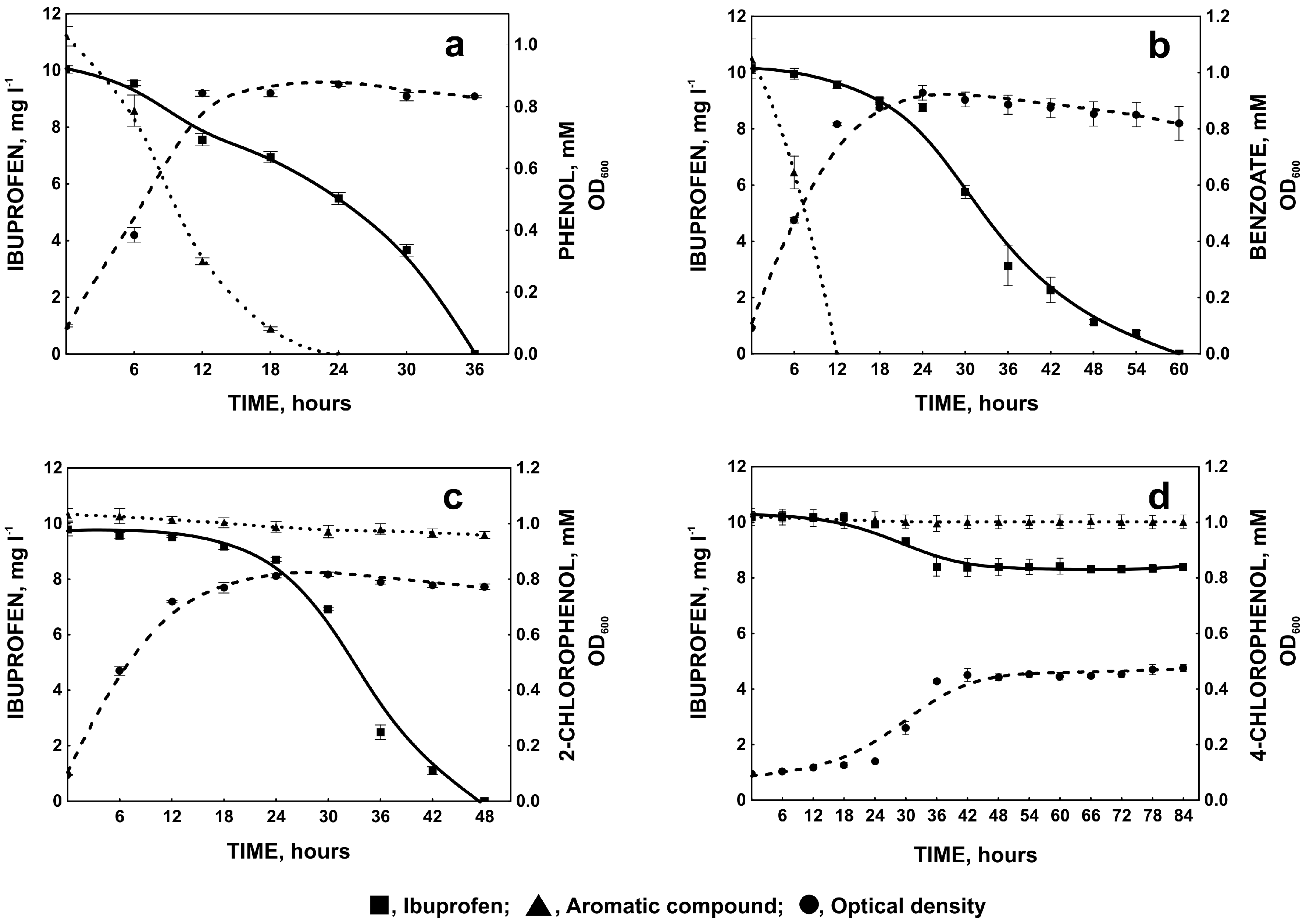
2.1.3. Degradation of Ibuprofen in the Presence of Selected Aromatic Compounds
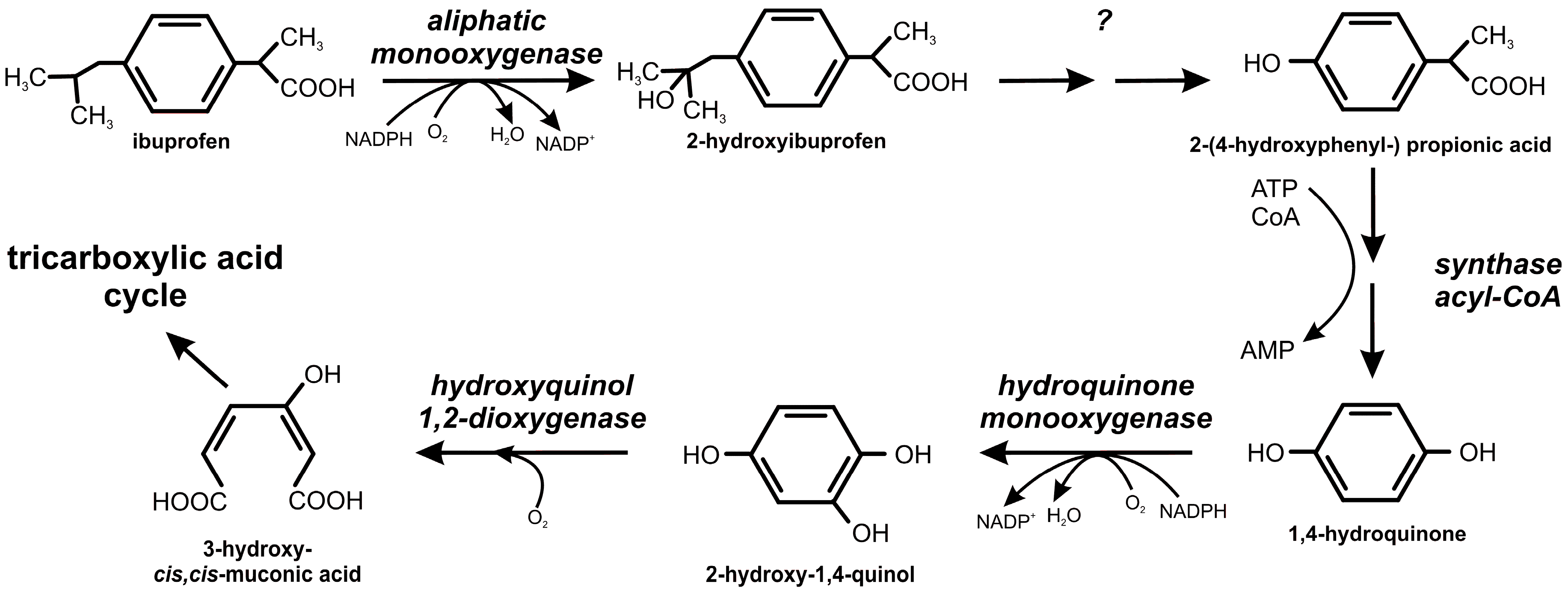
2.2. Ibuprofen Degradation Pathway
3. Materials and Methods
3.1. Bacterial Strain and Growth Conditions
3.2. Ibuprofen Degradation Experiments
3.3. Determination of No-Observed-Effect Concentrations (NOECs) of Heavy Metals
3.4. HPLC Analysis
3.5. Enzyme Assays
3.6. Intermediates Identification
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Nowak, K.M.; Girardi, C.; Miltner, A.; Gehre, M.; Schaffer, A.; Kastner, M. Contribution of microorganisms to non-extractable residue formation during biodegradation of ibuprofen in soil. Sci. Total Environ. 2013, 445-446, 377–384. [Google Scholar] [CrossRef] [PubMed]
- Araujo, L.; Troconis, M.E.; Espina, M.B.; Prieto, A. Persistence of ibuprofen, ketoprofen, diclofenac and clofibric acid in natural waters. J. Environ. Hum. 2014, 1, 32–38. [Google Scholar] [CrossRef]
- Kruglova, A.; Ahlgren, P.; Korhonen, N.; Rantanen, P.; Mikola, A.; Vahala, R. Biodegradation of ibuprofen, diclofenac and carbamazepine in nitifying activated sludge under 12 °C temperature conditions. Sci. Total Environ. 2014, 499, 394–401. [Google Scholar] [CrossRef] [PubMed]
- Joss, A.; Zabczynski, S.; Gobel, A.; Hoffmann, B.; Loffler, D.; McArdell, Ch.S.; Ternes, T.A.; Thomsen, A.; Siegrist, H. Biological degradation of pharmaceuticals in municipal wastewater treatment: Proposing a classification scheme. Water Res. 2006, 40, 1686–1696. [Google Scholar] [CrossRef] [PubMed]
- Murdoch, R.W.; Hay, A.G. Formation of catechols via removal of acid side chains from ibuprofen and related aromatic acids. Appl. Environ. Microbiol. 2005, 71, 6121–6125. [Google Scholar] [CrossRef] [PubMed]
- Murdoch, R.W.; Hay, A.G. Genetic and chemical characterization of ibuprofen degradation by Sphingomonas Ibu-2. Microbiology 2013, 159, 621–632. [Google Scholar] [CrossRef] [PubMed]
- Murdoch, R.W.; Hay, A.G. The biotransformation of ibuprofen to trihydroxyibuprofen in activated sludge and by Variovorax Ibu-1. Biodegradation 2015, 26, 105–113. [Google Scholar] [CrossRef] [PubMed]
- Almeida, B.; Kjeldal, H.; Lolas, I.; Knudsen, A.D.; Carvalho, G.; Nielsen, K.L.; Barreto Crespo, M.T.; Stensballe, A.; Nielsen, J.L. Quantitative proteomic analysis of ibuprofen-degrading Patulibacter sp. strain I11. Biodegradation 2013, 24, 615–630. [Google Scholar] [CrossRef] [PubMed]
- Zwiener, C.; Seeger, S.; Glauner, T.; Frimmel, F.H. Metabolites from the biodegradation of pharmaceutical residues of ibuprofen in biofilm reactors and batch experiments. Anal. Bioanal. Chem. 2002, 372, 569–575. [Google Scholar] [CrossRef] [PubMed]
- Quintana, J.B.; Weiss, S.; Reemtsma, T. Pathways and metabolites of microbial degradation of selected acidic pharmaceutical and their occurrence in municipal wastewater treated by a membrane bioreactor. Water Res. 2005, 39, 2654–2664. [Google Scholar] [CrossRef] [PubMed]
- Olaniran, A.O.; Balgobind, A.; Pillay, B. Bioavailability of heavy metals in soil: Impact on microbial biodegradation of organic compounds and possible improvement strategies. Int. J. Mol. Sci. 2013, 14, 10197–10228. [Google Scholar] [CrossRef] [PubMed]
- Al_Defiery, M.E.J.; Reddy, G. Influence of metal ions concentration on phenol degradation by Rhodococcus pyridinivorans GM3. Mesopo. Environ. J. 2014, 1, 30–38. [Google Scholar]
- Kuo, C.-W.; Genthner, B.R.S. Effect of added heavy metal ions on biotransformation and biodegradation of 2-chlorophenol and 3-chlorobenzoate in anaerobic bacterial consortia. Appl. Environ. Microbiol. 1996, 62, 2317–2323. [Google Scholar] [PubMed]
- Sandrin, T.R.; Maier, R.M. Impact of metals on the biodegradation of organic pollutants. Environ. Health Perspect. 2003, 111, 1093–1101. [Google Scholar] [CrossRef] [PubMed]
- Yeom, S.H.; Yoo, Y.J. Overcoming the inhibition effects of metal ions in the degradation of benzene and toluene by Alcaligenes xylosoxidans Y234. Korean J. Chem. Eng. 1997, 14, 204–208. [Google Scholar] [CrossRef]
- Monsalvo, V.M.; Mohedano, A.F.; Casas, J.A.; Rodrigguez, J.J. Cometabolic biodegradation of 4-chlorophenol by sequencing bath reactors at different temperatures. Biores. Technol. 2009, 100, 4572–4578. [Google Scholar] [CrossRef] [PubMed]
- Greń, I.; Wojcieszyńska, D.; Guzik, U.; Perkosz, M.; Hupert-Kocurek, K. Enhanced biotransformation of mononitrophenols by Stenotrophomonas maltophilia KB2 in the presence of aromatic compounds of plant origin. World J. Microbiol. Biotechnol. 2010, 26, 289–295. [Google Scholar] [CrossRef]
- Wojcieszyńska, D.; Hupert-Kocurek, K.; Guzik, U. Factors affecting activity of catechol 2,3-dioxygenase from 2-chlorophenol-degrading Stenotrophomonas maltophilia KB2. Biocatal. Biotransfor. 2013, 31, 141–147. [Google Scholar] [CrossRef]
- Marchlewicz, A.; Domaradzka, D.; Guzik, U.; Wojcieszyńska, D. Bacillus thuringiensis B1(2015b) is a gram-positive bacteria able to degrade naproxen and ibuprofen. Water Air Soil Pollut. 2016, 227, 197. [Google Scholar] [CrossRef] [PubMed]
- Marchlewicz, A.; Guzik, U.; Hupert-Kocurek, K.; Nowak, A.; Wilczyńska, S.; Wojcieszyńska, D. Toxicity and biodegradation of ibuprofen by Bacillus thiringiensis B1(2015b). Environ. Sci. Pollut. Res. 2017, 24, 7572–7584. [Google Scholar] [CrossRef] [PubMed]
- Sanguanpak, S.; Chiemchaisri, C.; Chiemchaisri, W.; Yamamoto, K. Influence of operating pH on biodegradation performance and fouling propensity in membrane bioreactors for landfill leachate treatment. Int. Biodeter. Biodegrad. 2015, 102, 64–72. [Google Scholar] [CrossRef]
- Boström, M.L.; Berglund, O. Influence of pH-dependent aquatic toxicity of ionizable pharmaceuticals on risk assessments over environmental pH ranges. Water Res. 2015, 72, 154–161. [Google Scholar] [CrossRef] [PubMed]
- Aksu, A.; Gönen, F. Biosorption of phenol by immobilized activated sludge in a continuous packed bed: Prediction of breakthrough curves. Process Biochem. 2004, 39, 599–613. [Google Scholar] [CrossRef]
- Moonen, M.J.H.; Fraaije, M.W.; Rietjens, I.M.C.M.; Laane, C.; Berkel, W.J.H. Flavoenzyme-catalyzed oxygenation and oxidations of phenolic compounds. Adv. Synth. Catal. 2002, 344, 1023–1035. [Google Scholar] [CrossRef]
- Horel, A.; Schiewer, S. Influence of constant and fluctuating temperature on biodegradation rates of fish biodiesel blends contaminating Alaskan sand. Chemosphere 2011, 83, 652–660. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Cho, K.-J.; Han, G.; Lee, C.; Hwang, S. Effects of temperature and pH on the biokinetic properties of thiocyanate biodegradation under autotrophic conditions. Water Res. 2013, 47, 251–258. [Google Scholar] [CrossRef] [PubMed]
- Delille, D.; Pelletier, E.; Rodriguez-Blanco, A.; Ghiglione, J.-F. Effects of nutrient and temperature on degradation of petroleum hydrocarbons in sub-Antarctic coastal seawater. Polar Biol. 2009, 32, 1521–1528. [Google Scholar] [CrossRef]
- Lenartova, V.; Holovska, K.; Javorsky, P. The influence of mercury on the antioxidant enzyme activity of rumen bacteria Streptococcus bovis and Selenomonas ruminantium. FEMS Microbiol. Ecol. 1998, 27, 319–325. [Google Scholar] [CrossRef]
- Bodaly, R.A.; Rudd, J.W.M.; Flett, R.J. Effect of urban sewage treatment on total and methyl mercury concentrations in effluents. Biogeochemistry 1998, 40, 279–291. [Google Scholar] [CrossRef]
- Vishnivetskaya, T.A.; Mosher, J.J.; Palumbo, A.V.; Yang, Z.K.; Podar, M.; Brown, S.D.; Brooks, S.C.; Gu, B.; Southworth, G.R.; Drake, M.M.; et al. Mercury and other heavy metals influence bacterial community structure in contaminated Tennessee streams. Appl. Environ. Microbiol. 2011, 77, 302–311. [Google Scholar] [CrossRef] [PubMed]
- Said, W.A.; Lewis, D.L. Quantitative assessment of the effects of metals on microbial degradation of organic chemicals. Appl. Environ. Microbiol. 1991, 57, 1498–1503. [Google Scholar] [PubMed]
- Hupert-Kocurek, K.; Saczyńska, A.; Piotrowska-Seget, Z. Cadmium increases catechol 2,3-dioxygenase activity in Variovorax sp. 12S, a metal-tolerant and phenol-degrading strain. Anton. Leeuw. Int. J. G. 2013, 104, 845–853. [Google Scholar] [CrossRef] [PubMed]
- Ruparelia, J.P.; Chatterjee, A.K.; Duttagupta, S.P.; Mukherji, S. Strain specificity in antimicrobial activity of silver and copper nanoparticles. Acta Biomater. 2008, 4, 707–716. [Google Scholar] [CrossRef] [PubMed]
- Guzik, U.; Hupert-Kocurek, K.; Sałek, K.; Wojcieszyńska, D. Influence of metal ions on bioremediation activity of protocatechuate 3,4-dioxygenase from Stenotrophomonas maltophilia KB2. World J. Microbiol. Biotechnol. 2013, 29, 267–273. [Google Scholar] [CrossRef] [PubMed]
- Argüello, J.M.; Raimunda, D.; Padilla-Benavides, T. Mechanisms of copper homeostasis in bacteria. Front. Cell. Infect. Microbiol. 2013, 3, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Banci, L.; Bertini, I.; Ciofi-Baffoni, S.; Del Conte, R.; Gonnelli, L. Understanding copper trafficking in bacteria: interaction between the copper transport protein CopZ and the N-terminal domain of the copper ATPase CopA from Bacillus subtilis. Biochemistry 2003, 42, 1939–1949. [Google Scholar] [CrossRef] [PubMed]
- Ron, E.Z.; Minz, D.; Finkelstein, N.P.; Rosenberg, E. Interaction of bacteria with cadmium. Biodegradation 1992, 3, 161–170. [Google Scholar] [CrossRef]
- Begg, S.L.; Eijkelkamp, B.A.; Luo, Z.; Counago, R.M.; Morey, J.R.; Maher, M.J.; Ong, C.Y.; McEwan, A.G.; Kobe, B.; O’Mara, M.L.; et al. Dysregulation of transition metal ion homeostasis is the molecular basis for cadmium toxicity in Streptococcus pneumonia. Nat. Commun. 2015, 6, 6418. [Google Scholar] [CrossRef] [PubMed]
- Guzik, U.; Hupert-Kocurek, K.; Wojcieszyńska, D. Intradiol dioxygenases- the key enzymes in xenobiotics degradation. In Biodegradation of Hazardous and Special Products, 1st ed.; Chamy, R., Rosenkranz, F., Eds.; Intech: Rijeka, Croatia, 2013; pp. 129–153. ISBN 978-953-51-1155-9. [Google Scholar]
- Ranquet, C.; Ollagnier-de-Choudens, S.; Loiseau, L.; Barras, F.; Fontecave, M. Cobalt stress in Escherichia coli. The effect on the iron-sulfur proteins. J. Biol. Chem. 2007, 282, 30442–30451. [Google Scholar] [CrossRef] [PubMed]
- Thorgersen, M.P.; Downs, D.M. Cobalt targets multiple metabolic processes in Salmonella enterica. J. Bacteriol. 2007, 189, 7774–7781. [Google Scholar] [CrossRef] [PubMed]
- Stasinakis, A.S.; Mamais, D.; Thomaidis, N.S.; Lekkas, T.D. Effect of chromium (VI) on bacterial kinetics of heterotrophic biomass of activated sludge. Water Res. 2002, 36, 3341–3349. [Google Scholar] [CrossRef]
- Pal, A.; Paul, A.K. Aerobic chromate reduction by chromium-resistant bacteria isolated from serpentine soil. Microbiol. Res. 2004, 159, 347–354. [Google Scholar] [CrossRef] [PubMed]
- Cervantes, C.; Campos-Garcia, J.; Devars, S.; Gutierrez-Corona, F.; Loza-Tavera, H.; Torres-Guzman, J.C.; Moreno-Sanchez, R. Interactions of chromium with microorganisms and plants. FEMS Microbiol. Rev. 2001, 25, 335–347. [Google Scholar] [CrossRef] [PubMed]
- Viti, C.; Marchi, E.; Decorosi, F.; Giovannetti, L. Molecular mechanisms of Cr(VI) resistance in becteria and fungi. FEMS Microbiol. Rev. 2014, 38, 633–659. [Google Scholar] [CrossRef] [PubMed]
- Pierre, L.L.; Moses, N.M.; Peter, M. Structure activity relationship (SAR) of some benzoic acid derivatives from plant origin that exhibit anti-sickling properties in vitro- (review). Nat. Prod. Chem. Res. 2015, 3, 176–179. [Google Scholar]
- Jamshidian, H.; Khatami, S.; Mogharei, A.; Vahabzadeha, F.; Nickzad, A. Cometabolic degradation of para-nitrophenol and phenol by Ralstonia eutropha in a Kissiris-immobilized cell bioreactor. Korean J. Chem. Eng. 2013, 30, 2052–2058. [Google Scholar] [CrossRef]
- Li, Y.; Li, B.; Wang, C.P.; Fan, J.Z.; Sun, H.W. Aerobic degradation of trichloroethylene by co-metabolism using phenol and gasoline as growth substrates. Int. J. Mol. Sci. 2014, 15, 9134–9148. [Google Scholar] [CrossRef] [PubMed]
- Nowak, A.; Mrozik, A. Facilitation of co-metabolic transformation and degradation of monochlorophenols by Pseudomonas sp. CF600 and changes in its fatty acid composition. Water Air Soil Pollut. 2016, 227, 83. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Li, Y.; Wang, Y.; Wang, C.; Wang, P. Experimental and kinetic study on the comatbolic biodegradation of phenol and 4-chlorophenol by psychrotropic Pseudomonas putida LY1. Environ. Sci. Pollut. Res. 2015, 22, 565–573. [Google Scholar] [CrossRef] [PubMed]
- Wojcieszyńska, D.; Guzik, U.; Greń, I.; Perkosz, M.; Hupert-Kocurek, K. Induction of aromatic ring-cleavage dioxygenases in Stenotrophomonas maltophilia strain KB2 in cometabolic systems. World J. Microbiol. Biotechnol. 2011, 27, 805–811. [Google Scholar] [CrossRef] [PubMed]
- Penttinen, O. Chlorophenols in aquatic environments: structure-activity correlations. Ann. Zool. Fenn. 1995, 32, 287–294. [Google Scholar]
- Czaplicka, M. Sources and transformations of chlorophenols in the natural environment. Sci. Total Environ. 2004, 322, 21–39. [Google Scholar] [CrossRef] [PubMed]
- Ding, T.; Yang, M.; Zhang, J.; Yang, B.; Lin, K.; Li, J. Toxicity, degradation and metabolic fate of ibuprofen on freshwater diatom Navicula sp. J. Hazard. Mater. 2017, 330, 127–134. [Google Scholar] [CrossRef] [PubMed]
- Wojcieszyńska, D.; Greń, I.; Hupert-Kocurek, K.; Guzik, U. Modulation of FAD-dependent monooxygenase activity from aromatic compounds-degrading Stenotrophomonas maltophilia KB2. Acta Biochim. Pol. 2011, 58, 421–426. [Google Scholar] [PubMed]
- Colby, J.; Stirling, D.I.; Dalton, H. The soluble methane mono-oxygenase of Methylococcus capsulatus (Bath). Its ability to oxygenate n-alkanes, n-alkenes, ethers, and alicyclic, aromatic and heterocyclic compounds. Biochem. J. 1977, 165, 395–402. [Google Scholar] [CrossRef] [PubMed]
- Hou, Ch.T.; Lillard, M.O.; Schwartz, R.D. Protocatechuate 3,4-dioxygenase from Acinetobacter calcoaceticus. Biochemistry 1976, 15, 582–588. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Khoo, H.E.; Poh, C.L. Purification and characterization of gentisate 1,2-dioxygenases from Pseudomonas alcaligenes NCIB 9867 and Pseudomonas putida NCIB 9869. Appl. Microbiol. Biotechnol. 1999, 65, 946–950. [Google Scholar]
- Wei, M.; Zhang, J.J.; Liu, H.; Zhou, N.Y. para-Nitrophenol 4-monooxygenase and hydroxyquinol 1,2-dioxygenase catalyze sequential transformation of 4-nitrocatechol in Pseudomonas sp. strain WBC-3. Biodegradation 2010, 21, 915–921. [Google Scholar] [CrossRef] [PubMed]
- Moonen, M.J.H.; Synowsky, S.A.; Van den Berg, W.A.M.; Westphal, A.H.; Heck, A.J.R.; Van den Heuvel, R.H.H.; Fraaije, M.W.; Van Berkel, W.J.H. Hydroquinone dioxygenase from Pseudomonas fluorescens ACB: A novel member of the family of nonheme-iron(II)-dependent dioxygenases. J. Bacteriol. 2008, 190, 5199–5209. [Google Scholar] [CrossRef] [PubMed]
- Castano-Cerezo, S.; Bernal, V.; Blanco-Catala, J.; Iborra, J.L.; Canovas, M. cAMP-CRP co-odinates the expression of the protein acetylation pathway with central metabolism in Escherichia Coli. Mol. Microbiol. 2011, 82, 1110–1128. [Google Scholar] [CrossRef] [PubMed]
- Castano-Cerezo, S.; Bernal, V.; Canovas, M. Acetyl-coenzyme A synthetase (ACS) assay. Bio-protocol 2012, 2, 1–5. [Google Scholar]
- Leonowicz, A.; Grzywnowicz, K. Quantitative estimation of laccase forms in some white-rot fungi using syringaldazide as a substrate. Enzyme Microb. Technol. 1981, 3, 55–58. [Google Scholar] [CrossRef]
Sample Availability: Samples of the compounds are not available from the authors. |





| Metal Ion | Metal Ion Concentration (mM) | Optical Density (OD600) | NOEC (mM) |
|---|---|---|---|
| Cu2+ | 0.00000082 | 0.150 ± 0.040 | 0.00256 |
| 0.0000041 | 0.146 ± 0.026 | ||
| 0.00002048 | 0.138 ± 0.033 | ||
| 0.0001024 | 0.139 ± 0.019 | ||
| 0.000512 | 0.145 ± 0.016 | ||
| 0.00256 | 0.156 ± 0.012 | ||
| 0.128 * | 0.092 ± 0.015 | ||
| 0.064 * | 0.109 ± 0.015 | ||
| 0.32 * | 0.094 ± 0.034 | ||
| 1.6 * | 0.077 ± 0.019 | ||
| Cd2+ | 0.000001 | 0.111 ± 0.009 | 0.003 |
| 0.000004 | 0.106 ± 0.028 | ||
| 0.00002 | 0.113 ± 0.025 | ||
| 0.000102 | 0.118 ± 0.032 | ||
| 0.001 | 0.120 ± 0.034 | ||
| 0.003 | 0.117 ± 0.030 | ||
| 0.013 * | 0.108 ± 0.001 | ||
| 0.064 * | 0.119 ± 0.011 | ||
| 0.32 * | 0.124 ± 0.027 | ||
| 1.6 * | 0.079 ± 0.008 | ||
| 8.0 * | 0.061 ± 0.008 | ||
| Co2+ | 0.17 | 0.149 ± 0.008 | 0.21 |
| 0.21 | 0.162 ± 0.015 | ||
| 0.27 * | 0.120 ± 0.014 | ||
| 0.34 * | 0.108 ± 0.022 | ||
| 0.42 * | 0.095 ± 0.010 | ||
| 0.52 * | 0.050 ± 0.016 | ||
| 0.66 * | 0.035 ± 0.009 | ||
| 0.82 * | 0.019 ± 0.008 | ||
| 1.02 * | 0.020 ± 0.004 | ||
| 1.28 * | 0.017 ± 0.008 | ||
| 1.60 * | 0.017 ± 0.008 | ||
| Cr6+ | 0.00000082 * | 0.079 ± 0.021 | 0.32 |
| 0.0000041 * | 0.092 ± 0.030 | ||
| 0.00002048 * | 0.095 ± 0.030 | ||
| 0.0001024 * | 0.127 ± 0.023 | ||
| 0.000512 * | 0.170 ± 0.021 | ||
| 0.00256 * | 0.138 ± 0.021 | ||
| 0.128 | 0.065 ± 0.008 | ||
| 0.064 | 0.009 ± 0.004 | ||
| 0.32 | 0.004 ± 0.003 | ||
| 1.6 * | 0.003 ± 0.003 | ||
| 8.0 * | 0.000 ± 0.000 | ||
| Hg2+ | 0.000000492 | 0.104 ± 0.015 | 0.000307 |
| 0.000002 | 0.103 ± 0.023 | ||
| 0.000012 | 0.113 ± 0.002 | ||
| 0.000061 | 0.108 ± 0.006 | ||
| 0.000307 | 0.121 ± 0.013 | ||
| 0.001536 * | 0.047 ± 0.014 | ||
| 0.00768 * | 0.000 ± 0.000 | ||
| 0.0384 * | 0.000 ± 0.000 | ||
| 0.192 * | 0.000 ± 0.000 | ||
| 0.96 * | 0.000 ± 0.000 | ||
| 4.8 * | 0.000 ± 0.000 |
| Enzyme | Specific Enzyme Activity (mU/mg Protein) | Enzyme | Specific Enzyme Activity (mU/mg Protein) |
|---|---|---|---|
| Methane monooxygenase | 831.39 ± 167.97 | Catechol 2,3-dioxygenase | 0.00 ± 0.00 |
| Butane/isobutane monooxygenase | 150.94 ± 21.95 | Protocatechuate 3,4-dioxygenase | 0.00 ± 0.00 |
| Propane/butane monooxygenase | 311.77 ± 62.99 | Protocatechuate 4,5-dioxygenase | 0.00 ± 0.00 |
| Pentane monooxygenase | 0.00 ± 0.00 | Gentisate 1,2-dioxygenase | 0.00 ± 0.00 |
| Phenol monooxygenase | 77.40 ± 4.50 | Hydroxyquinol 1,2-dioxygenase | 99.12 ± 12.92 |
| Hydroquinone monooxygenase | 102.68 ± 14.24 | Hydroquinone 1,2-dioxygenase | 0.00 ± 0.00 |
| Catechol 1,2-dioxygenase | 34.99 ± 2.91 | Acetyl-coenzyme A synthetase | 249.47 ± 24.16 |
| Laccase | 0.00 ± 0.00 | Peroxidase | 0.00 ± 0.00 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Marchlewicz, A.; Guzik, U.; Smułek, W.; Wojcieszyńska, D. Exploring the Degradation of Ibuprofen by Bacillus thuringiensis B1(2015b): The New Pathway and Factors Affecting Degradation. Molecules 2017, 22, 1676. https://doi.org/10.3390/molecules22101676
Marchlewicz A, Guzik U, Smułek W, Wojcieszyńska D. Exploring the Degradation of Ibuprofen by Bacillus thuringiensis B1(2015b): The New Pathway and Factors Affecting Degradation. Molecules. 2017; 22(10):1676. https://doi.org/10.3390/molecules22101676
Chicago/Turabian StyleMarchlewicz, Ariel, Urszula Guzik, Wojciech Smułek, and Danuta Wojcieszyńska. 2017. "Exploring the Degradation of Ibuprofen by Bacillus thuringiensis B1(2015b): The New Pathway and Factors Affecting Degradation" Molecules 22, no. 10: 1676. https://doi.org/10.3390/molecules22101676
APA StyleMarchlewicz, A., Guzik, U., Smułek, W., & Wojcieszyńska, D. (2017). Exploring the Degradation of Ibuprofen by Bacillus thuringiensis B1(2015b): The New Pathway and Factors Affecting Degradation. Molecules, 22(10), 1676. https://doi.org/10.3390/molecules22101676









