A SILAC-Based Approach Elicits the Proteomic Responses to Vancomycin-Associated Nephrotoxicity in Human Proximal Tubule Epithelial HK-2 Cells
Abstract
:1. Introduction
2. Results
2.1. Overview of Proteomic Response to Vancomycin Treatment in HK-2 Cells
2.2. Vancomycin Alters a Number of Cellular Functions in HK-2 cells
| Protein ID | Protein Name | Gene Names | Average(H/L) |
|---|---|---|---|
| P04264 | Keratin, type II cytoskeletal 1 | KRT1 | 0.172905 |
| P35527 | Keratin, type I cytoskeletal 9 | KRT9 | 0.224193 |
| E9PCX2 | Aldose reductase | AKR1B1 | 0.272265 |
| P02751 | Fibronectin | FN1 | 0.288053 |
| Q5T7C4 | High mobility group protein B1 | HMGB1; HMGB1P1 | 0.315275 |
| Q13185 | Chromobox protein homolog 3 | CBX3 | 0.34326 |
| P40261 | Nicotinamide N-methyltransferase | NNMT | 0.397895 |
| P29966 | Myristoylated alanine-rich C-kinase substrate | MARCKS | 0.400377 |
| F5H7V9 | Tenascin | TNC | 0.40316 |
| Q9Y2V2 | Calcium-regulated heat stable protein 1 | CARHSP1 | 0.41335 |
| K7ERT7 | Synaptic vesicle membrane protein VAT-1 homolog | VAT1 | 0.434395 |
| P51858 | Hepatoma-derived growth factor | HDGF | 0.453443 |
| E9PEJ4 | Dihydrolipoyllysine-residue acetyltransferase component of pyruvate dehydrogenase complex, mitochondrial | DLAT | 0.4645 |
| H0YK49 | Electron transfer flavoprotein subunit α, mitochondrial | ETFA | 0.46645 |
| E7ES10 | Calpastatin | CAST | 0.46797 |
| Q06830 | Peroxiredoxin-1 | PRDX1 | 0.486445 |
| Q9HB71 | Calcyclin-binding protein | CACYBP | 0.49843 |
| H0YJX6 | Protein arginine N-methyltransferase 5 | PRMT5 | 0.50622 |
| O75937 | DNAJ homolog subfamily C member 8 | DNAJC8 | 0.50726 |
| P49915 | GMP synthase [glutamine-hydrolyzing] | GMPS | 0.513415 |
| H0YFA4 | Cysteine-rich protein 2 | CRIP2 | 0.51363 |
| P04792 | Heat shock protein β-1 | HSPB1 | 0.51522 |
| E9PLK3 | Puromycin-sensitive aminopeptidase | NPEPPS | 0.535463 |
| P49773 | Histidine triad nucleotide-binding protein 1 | HINT1 | 0.5365 |
| Q06323 | Proteasome activator complex subunit 1 | PSME1 | 0.541965 |
| B8ZZQ6 | Prothymosin α | PTMA | 0.547615 |
| Q9Y696 | Chloride intracellular channel protein 4 | CLIC4 | 0.55066 |
| P16403 | Histone H1.2 | HIST1H1C; HIST1H1E; HIST1H1D; HIST1H1T; HIST1H1A | 0.55312 |
| H7C469 | Cathepsin D | CTSD | 0.554823 |
| Q12792 | Twinfilin-1 | TWF1 | 0.55623 |
| J3KTF8 | Rho GDP-dissociation inhibitor 1 | ARHGDIA | 0.5657 |
| P28482 | Mitogen-activated protein kinase 1 | MAPK1 | 0.57503 |
| Q9C005 | Protein DPY-30 homolog | DPY30 | 0.57574 |
| Q9UKY7 | Protein CDV3 homolog | CDV3 | 0.582217 |
| B4E022 | Transketolase | TKT | 0.58253 |
| Q99497 | Protein DJ-1 | PARK7 | 0.584365 |
| P18669 | Phosphoglycerate mutase 1 | PGAM1 | 0.587947 |
| P09104 | γ-Enolase | ENO2 | 0.58879 |
| P20042 | Eukaryotic translation initiation factor 2 subunit 2 | EIF2S2 | 0.60232 |
| Q01995 | Transgelin | TAGLN | 0.60334 |
| C9J9W2 | LIM and SH3 domain protein 1 | LASP1 | 0.60464 |
| P37837 | Transaldolase | TALDO1 | 0.60516 |
| F8VZ29 | Ubiquitin-conjugating enzyme E2 N | UBE2N | 0.60613 |
| Q9H4M9 | EH domain-containing protein 1 | EHD1 | 0.60702 |
| Q96QK1 | Vacuolar protein sorting-associated protein 35 | VPS35 | 0.609495 |
| P49321 | Nuclear autoantigenic sperm protein | NASP | 0.6098 |
| P40925 | Malate dehydrogenase, cytoplasmic | MDH1 | 0.61013 |
| P31150 | Rab GDP dissociation inhibitor α | GDI1 | 0.61355 |
| Q16836 | Hydroxyacyl-coenzyme A dehydrogenase, mitochondrial | HADH | 0.61513 |
| P60174 | Triosephosphate isomerase | TPI1 | 0.617187 |
| Q92616 | Translational activator GCN1 | GCN1L1 | 0.623255 |
| Q99714 | 3-Hydroxyacyl-CoA dehydrogenase type-2 | HSD17B10 | 0.630585 |
| P31947 | 14-3-3 protein σ | SFN | 0.630687 |
| P12956 | X-ray repair cross-complementing protein 6 | XRCC6 | 0.63482 |
| Q00341 | Vigilin | HDLBP | 0.63632 |
| O75367 | Core histone macro-H2A.1 | H2AFY | 0.63646 |
| P07814 | Bifunctional glutamate/proline-tRNA ligase | EPRS | 0.6377 |
| P27816 | Microtubule-associated protein 4 | MAP4 | 0.64062 |
| P05141 | ADP/ATP translocase 2 | SLC25A5 | 0.641425 |
| Q04917 | 14-3-3 protein η | YWHAH | 0.641525 |
| O43175 | d-3-phosphoglycerate dehydrogenase | PHGDH | 0.644225 |
| P30086 | Phosphatidylethanolamine-binding protein 1 | PEBP1 | 0.644533 |
| Q01082 | Spectrin β chain, non-erythrocytic 1 | SPTBN1 | 0.64709 |
| Q13907 | Isopentenyl-diphosphate δ-isomerase 1 | IDI1 | 0.647705 |
| P53621 | Coatomer subunit α | COPA | 0.65046 |
| O60884 | DNAJ homolog subfamily A member 2 | DNAJA2 | 0.65208 |
| C9JNR4 | Transforming protein RhoA | RHOA | 0.65726 |
| P11413 | Glucose-6-phosphate 1-dehydrogenase | G6PD | 0.664673 |
| P26196 | Probable ATP-dependent RNA helicase DDX6 | DDX6 | 0.66701 |
| Q08945 | FACT complex subunit SSRP1 | SSRP1 | 0.66756 |
| Q13283 | Ras GTPase-activating protein-binding protein 1 | G3BP1 | 0.66807 |
| Q15365 | Poly(rC)-binding protein 1 | PCBP1 | 0.66997 |
| P42166 | Lamina-associated polypeptide 2, isoform α | TMPO | 0.6721 |
| K7ELL7 | Glucosidase 2 subunit β | PRKCSH | 0.67451 |
| P32119 | Peroxiredoxin-2 | PRDX2 | 0.67591 |
| P23381 | Tryptophan-tRNA ligase, cytoplasmic | WARS | 0.677045 |
| P33991 | DNA replication licensing factor MCM4 | MCM4 | 0.6794 |
| P05787 | Keratin, type II cytoskeletal 8 | KRT8 | 0.680803 |
| C9JJ34 | Ran-specific GTPase-activating protein | RANBP1 | 0.68427 |
| P49327 | Fatty acid synthase | FASN | 0.687997 |
| P09936 | Ubiquitin carboxyl-terminal hydrolase isozyme L1 | UCHL1 | 0.688333 |
| A2A2D0 | Stathmin | STMN1 | 0.69224 |
| Q9ULV4 | Coronin-1C | CORO1C | 0.695523 |
| Q9Y4L1 | Hypoxia up-regulated protein 1 | HYOU1 | 0.696023 |
| P07195 | l-lactate dehydrogenase B chain | LDHB | 0.700453 |
| F8VWS0 | 60S acidic ribosomal protein P0 | RPLP0 | 0.703667 |
| P25205 | DNA replication licensing factor MCM3 | MCM3 | 0.708945 |
| K7EJE8 | Lon protease homolog, mitochondrial | LONP1 | 0.70909 |
| P23528 | Cofilin-1 | CFL1 | 0.71004 |
| P07737 | Profilin-1 | PFN1 | 0.71163 |
| E5RIW3 | Tubulin-specific chaperone A | TBCA | 0.71218 |
| P63010 | AP-2 complex subunit β | AP2B1 | 0.714287 |
| P52209 | 6-Phosphogluconate dehydrogenase, decarboxylating | PGD | 0.715343 |
| O00151 | PDZ and LIM domain protein 1 | PDLIM1 | 0.71603 |
| P36871 | Phosphoglucomutase-1 | PGM1 | 0.717643 |
| Q15056 | Eukaryotic translation initiation factor 4H | EIF4H | 0.721007 |
| P27797 | Calreticulin | CALR | 0.722223 |
| P37802 | Transgelin-2 | TAGLN2 | 0.723653 |
| Q16777 | Histone H2A type 2-C | HIST2H2AC; HIST2H2AA3 | 0.72469 |
| Q92973 | Transportin-1 | TNPO1 | 0.72551 |
| O60506 | Heterogeneous nuclear ribonucleoprotein Q | SYNCRIP | 0.7259 |
| O14579 | Coatomer subunit ε | COPE | 0.72719 |
| P14324 | Farnesyl pyrophosphate synthase | FDPS | 0.7323 |
| B4DQU5 | Ras-related protein Rab-11A | RAB11A; RAB11B | 0.73251 |
| P00338 | l-lactate dehydrogenase A chain | LDHA | 0.735003 |
| P24752 | Acetyl-CoA acetyltransferase, mitochondrial | ACAT1 | 0.73511 |
| O75369 | Filamin-B | FLNB | 0.73513 |
| P14866 | Heterogeneous nuclear ribonucleoprotein L | HNRNPL | 0.735237 |
| B4DDD8 | Histidine—tRNA ligase, cytoplasmic | HARS | 0.73643 |
| E9PBS1 | Multifunctional protein ADE2 | PAICS | 0.738453 |
| P42224 | Signal transducer and activator of transcription 1-α/β | STAT1 | 0.738843 |
| P30084 | Enoyl-CoA hydratase, mitochondrial | ECHS1 | 0.740183 |
| B7Z972 | Protein-l-isoaspartate O-methyltransferase | PCMT1 | 0.7404 |
| P53618 | Coatomer subunit β | COPB1 | 0.74088 |
| Q04760 | Lactoylglutathione lyase | GLO1 | 0.7413 |
| D6RG15 | Twinfilin-2 | TWF2 | 0.74232 |
| P40939 | Trifunctional enzyme subunit α, mitochondrial | HADHA | 0.744977 |
| P14618 | Pyruvate kinase PKM | PKM | 0.745617 |
| P07237 | Protein disulfide-isomerase | P4HB | 0.7458 |
| P30447 | HLA class I histocompatibility antigen, A-23 α chain | HLA-A; HLA-H; HLA-C | 0.74708 |
| Q10567 | AP-1 complex subunit β-1 | AP1B1 | 0.74868 |
| P21796 | Voltage-dependent anion-selective channel protein 1 | VDAC1 | 0.748783 |
| E9PMH2 | Peptidyl-prolyl cis-trans isomerase | AIP | 0.75192 |
| F8VUA6 | 60S ribosomal protein L18 | RPL18 | 0.75214 |
| P06744 | Glucose-6-phosphate isomerase | GPI | 0.755337 |
| O60888 | Protein CutA | CUTA | 0.7555 |
| P35606 | Coatomer subunit β | COPB2 | 0.755705 |
| P10768 | S-formylglutathione hydrolase | ESD | 0.75572 |
| P27695 | DNA-(apurinic or apyrimidinic site) lyase | APEX1 | 0.75672 |
| P12004 | Proliferating cell nuclear antigen | PCNA | 0.75721 |
| Q6ZR64 | MXRA7 | 0.75738 | |
| Q9UQ80 | Proliferation-associated protein 2G4 | PA2G4 | 0.75995 |
| E7EUY0 | DNA-dependent protein kinase catalytic subunit | PRKDC | 0.761483 |
| Q15185 | Prostaglandin E synthase 3 | PTGES3 | 0.762053 |
| P43243 | Matrin-3 | MATR3 | 0.765323 |
| Q96FW1 | Ubiquitin thioesterase OTUB1 | OTUB1 | 0.766887 |
| P31939 | Bifunctional purine biosynthesis protein PURH | ATIC | 0.766895 |
| Q99733 | Nucleosome assembly protein 1-like 4 | NAP1L4 | 0.769025 |
| P62191 | 26S protease regulatory subunit 4 | PSMC1 | 0.77016 |
| P40926 | Malate dehydrogenase, mitochondrial | MDH2 | 0.770383 |
| Q32Q12 | Nucleoside diphosphate kinase | NME1-NME2; NME2; NME2P1; NME1 | 0.770707 |
| P30044 | Peroxiredoxin-5, mitochondrial | PRDX5 | 0.77087 |
| P34897 | Serine hydroxymethyltransferase, mitochondrial | SHMT2 | 0.77093 |
| P08758 | Annexin A5 | ANXA5 | 0.772893 |
| P00558 | Phosphoglycerate kinase 1 | PGK1 | 0.776877 |
| Q92945 | Far upstream element-binding protein 2 | KHSRP | 0.778823 |
| Q01105 | Protein SET | SET | 0.779857 |
| F8W1N5 | Nascent polypeptide-associated complex subunit α | NACA | 0.78007 |
| P05386 | 60S acidic ribosomal protein P1 | RPLP1 | 0.784843 |
| Q9BWD1 | Acetyl-CoA acetyltransferase, cytosolic | ACAT2 | 0.7849 |
| Q96I24 | Far upstream element-binding protein 3 | FUBP3 | 0.78507 |
| F8VQE1 | LIM domain and actin-binding protein 1 | LIMA1 | 0.786983 |
| O43852 | Calumenin | CALU | 0.787113 |
| Q02790 | Peptidyl-prolyl cis-trans isomerase FKBP4 | FKBP4 | 0.790175 |
| J3KN67 | Tropomyosin α-3 chain | TPM3 | 0.790573 |
| P00441 | Superoxide dismutase [Cu-Zn] | SOD1 | 0.79129 |
| Q15181 | Inorganic pyrophosphatase | PPA1 | 0.79164 |
| P20700 | Lamin-B1 | LMNB1 | 0.798267 |
| E9PK47 | Phosphorylase | PYGL | 0.79884 |
| P62136 | Serine/threonine-protein phosphatase PP1-α catalytic subunit | PPP1CA; PPP1CC | 0.79982 |
| P43686 | 26S protease regulatory subunit 6B | PSMC4 | 0.800465 |
| P07384 | Calpain-1 catalytic subunit | CAPN1 | 0.806005 |
| O14773 | Tripeptidyl-peptidase 1 | TPP1 | 0.807353 |
| Q14566 | DNA replication licensing factor MCM6 | MCM6 | 0.80749 |
| Q09666 | Neuroblast differentiation-associated protein AHNAK | AHNAK | 0.81098 |
| P84077 | ADP-ribosylation factor 1 | ARF1; ARF3; ARF4; ARF5 | 0.81248 |
| Q14444 | Caprin-1 | CAPRIN1 | 0.812877 |
| F5H018 | GTP-binding nuclear protein Ran | RAN | 0.813607 |
| Q92499 | ATP-dependent RNA helicase DDX1 | DDX1 | 0.813953 |
| Q99832 | T-complex protein 1 subunit η | CCT7 | 0.81562 |
| P68431 | Histone H3.1 | HIST1H3A; H3F3B; H3F3A; HIST3H3; H3F3C | 0.816783 |
| P33993 | DNA replication licensing factor MCM7 | MCM7 | 0.81733 |
| Q01518 | Adenylyl cyclase-associated protein 1 | CAP1 | 0.818003 |
| P62258 | 14-3-3 protein ε | YWHAE | 0.822137 |
| P63241 | Eukaryotic translation initiation factor 5A-1 | EIF5A; EIF5AL1 | 0.82674 |
| P26368 | Splicing factor U2AF 65 kDa subunit | U2AF2 | 0.827353 |
| P49588 | Alanine-tRNA ligase, cytoplasmic | AARS | 0.827427 |
| P24534 | Elongation factor 1-β | EEF1B2 | 0.82769 |
| P68104 | Elongation factor 1-α 1 | EEF1A1; EEF1A1P5 | 0.828823 |
| P39023 | 60S ribosomal protein L3 | RPL3 | 0.829365 |
| O75083 | WD repeat-containing protein 1 | WDR1 | 0.82946 |
| H0YLC2 | Proteasome subunit α type | PSMA4 | 0.83034 |
| P46778 | 60S ribosomal protein L21 | RPL21 | 0.83041 |
| P40227 | T-complex protein 1 subunit ζ | CCT6A | 0.83204 |
| Q14980 | Nuclear mitotic apparatus protein 1 | NUMA1 | 0.832107 |
| Q13263 | Transcription intermediary factor 1-β | TRIM28 | 0.832473 |
| P06733 | α-Enolase | ENO1 | 0.833683 |
| Q9UK76 | Hematological and neurological expressed 1 protein | HN1 | 0.833713 |
| P13010 | X-ray repair cross-complementing protein 5 | XRCC5 | 0.835137 |
| P67809 | Nuclease-sensitive element-binding protein 1 | YBX1 | 0.83629 |
| Q13492 | Phosphatidylinositol-binding clathrin assembly protein | PICALM | 0.83656 |
| P26639 | Threonine-tRNA ligase, cytoplasmic | TARS | 0.837765 |
| P30153 | Serine/threonine-protein phosphatase 2A 65 kDa regulatory subunit A α isoform | PPP2R1A | 0.839417 |
| P61978 | Heterogeneous nuclear ribonucleoprotein K | HNRNPK | 0.839853 |
| G3V1A1 | 60S ribosomal protein L8 | RPL8 | 0.83992 |
| R4GN98 | Protein S100-A6 | S100A6 | 0.840685 |
| P62081 | 40S ribosomal protein S7 | RPS7 | 0.84122 |
| P16989 | Y-box-binding protein 3 | YBX3 | 0.84325 |
| P67936 | Tropomyosin α-4 chain | TPM4 | 0.84394 |
| P10809 | 60 kDa heat shock protein, mitochondrial | HSPD1 | 0.84574 |
| E9PLD0 | Ras-related protein Rab-1B | RAB1B; RAB1C | 0.84636 |
| P54136 | Arginine-tRNA ligase, cytoplasmic | RARS | 0.84759 |
| P50454 | Serpin H1 | SERPINH1 | 0.848623 |
| P11021 | 78 kDa glucose-regulated protein | HSPA5 | 0.84892 |
| P61769 | β2-microglobulin | B2M | 0.849333 |
| F5GY37 | Prohibitin-2 | PHB2 | 0.85107 |
| Q07666 | KH domain-containing, RNA-binding, signal transduction-associated protein 1 | KHDRBS1 | 0.85112 |
| P18206 | Vinculin | VCL | 0.851217 |
| P62314 | Small nuclear ribonucleoprotein Sm D1 | SNRPD1 | 0.85213 |
| P49411 | Elongation factor Tu, mitochondrial | TUFM | 0.85245 |
| H0Y4R1 | Inosine-5-monophosphate dehydrogenase 2 | IMPDH2 | 0.853715 |
| P04844 | Dolichyl-diphosphooligosaccharide-protein glycosyltransferase subunit 2 | RPN2 | 0.85403 |
| Q71DI3 | Histone H3.2 | HIST2H3A | 0.854437 |
| P50395 | Rab GDP dissociation inhibitor β | GDI2 | 0.855377 |
| Q00839 | Heterogeneous nuclear ribonucleoprotein U | HNRNPU | 0.8554 |
| Q13347 | Eukaryotic translation initiation factor 3 subunit I | EIF3I | 0.85975 |
| P40222 | α-Taxilin | TXLNA | 0.860857 |
| P09382 | Galectin-1 | LGALS1 | 0.861283 |
| Q15084 | Protein disulfide-isomerase A6 | PDIA6 | 0.861437 |
| Q14247 | Src substrate cortactin | CTTN | 0.86161 |
| P21333 | Filamin-A | FLNA | 0.86397 |
| O14818 | Proteasome subunit α type-7 | PSMA7; PSMA8 | 0.86409 |
| P13667 | Protein disulfide-isomerase A4 | PDIA4 | 0.86544 |
| Q9UMS4 | Pre-mRNA-processing factor 19 | PRPF19 | 0.866023 |
| E7EPW6 | Programmed cell death protein 6 | PDCD6 | 0.87172 |
| E9PIR7 | Thioredoxin reductase 1, cytoplasmic | TXNRD1 | 0.872513 |
| Q9UHX1 | Poly(U)-binding-splicing factor PUF60 | PUF60 | 0.872915 |
| P05556 | Integrin β-1 | ITGB1 | 0.873165 |
| P40121 | Macrophage-capping protein | CAPG | 0.87371 |
| Q03135 | Caveolin-1 | CAV1 | 0.87472 |
| P13797 | Plastin-3 | PLS3 | 0.87507 |
| P27824 | Calnexin | CANX | 0.87692 |
| Q15019 | Septin-2 | 42249 | 0.87721 |
| Q9UNZ2 | NSFL1 cofactor p47 | NSFL1C | 0.878297 |
| Q9Y5B9 | FACT complex subunit SPT16 | SUPT16H | 0.879483 |
| B1AK85 | F-actin-capping protein subunit β | CAPZB | 0.88193 |
| P68133 | Actin, α skeletal muscle | ACTA1; ACTC1; ACTG2; ACTA2 | 0.88416 |
| P06493 | Cyclin-dependent kinase 1 | CDK1 | 0.88483 |
| Q3ZCM7 | Tubulin β-8 chain | TUBB8 | 0.88649 |
| P07858 | Cathepsin B | CTSB | 0.889247 |
| Q16555 | Dihydropyrimidinase-related protein 2 | DPYSL2 | 0.88961 |
| P63104 | 14-3-3 protein ζ/δ | YWHAZ | 0.890777 |
| Q14697 | Neutral α-glucosidase AB | GANAB | 0.89087 |
| P05783 | Keratin, type I cytoskeletal 18 | KRT18 | 0.891143 |
| P26640 | Valine-tRNA ligase | VARS | 0.89307 |
| B4DUR8 | T-complex protein 1 subunit γ | CCT3 | 0.895913 |
| Q96P70 | Importin-9 | IPO9 | 0.89849 |
| Q7KZF4 | Staphylococcal nuclease domain-containing protein 1 | SND1 | 0.899653 |
| Q16643 | Drebrin | DBN1 | 0.90207 |
| P35579 | Myosin-9 | MYH9 | 0.90543 |
| P22626 | Heterogeneous nuclear ribonucleoproteins A2/B1 | HNRNPA2B1 | 0.907457 |
| Q15942 | Zyxin | ZYX | 0.909143 |
| Q9NQC3 | Reticulon-4 | RTN4 | 0.90961 |
| P07108 | Acyl-CoA-binding protein | DBI | 0.91033 |
| P11216 | Glycogen phosphorylase, brain form | PYGB | 0.914587 |
| A6PVH9 | Copine-1 | CPNE1 | 0.914685 |
| O14980 | Exportin-1 | XPO1 | 0.91646 |
| P04075 | Fructose-bisphosphate aldolase A | ALDOA | 0.917223 |
| O43390 | Heterogeneous nuclear ribonucleoprotein R | HNRNPR | 0.91729 |
| P61158 | Actin-related protein 3 | ACTR3 | 0.91757 |
| P13639 | Elongation factor 2 | EEF2 | 0.918273 |
| P07942 | Laminin subunit β-1 | LAMB1 | 0.91949 |
| P31948 | Stress-induced-phosphoprotein 1 | STIP1 | 0.920767 |
| Q14204 | Cytoplasmic dynein 1 heavy chain 1 | DYNC1H1 | 0.92284 |
| O60664 | Perilipin-3 | PLIN3 | 0.922903 |
| P62158 | Calmodulin | CALM1; CALM2; CALM3 | 0.923537 |
| Q96QD8 | Sodium-coupled neutral amino acid transporter 2 | SLC38A2 | 0.92438 |
| Q92575 | UBX domain-containing protein 4 | UBXN4 | 0.924495 |
| M0R0F0 | 40S ribosomal protein S5 | RPS5 | 0.925887 |
| P08133 | Annexin A6 | ANXA6 | 0.92615 |
| P62906 | 60S ribosomal protein L10a | RPL10A | 0.930083 |
| O15144 | Actin-related protein 2/3 complex subunit 2 | ARPC2 | 0.932825 |
| Q9Y490 | Talin-1 | TLN1 | 0.933233 |
| P04406 | Glyceraldehyde-3-phosphate dehydrogenase | GAPDH | 0.934323 |
| P25786 | Proteasome subunit α type-1 | PSMA1 | 0.93491 |
| P25787 | Proteasome subunit α type-2 | PSMA2 | 0.93574 |
| P34932 | Heat shock 70 kDa protein 4 | HSPA4 | 0.935993 |
| E7EQR4 | Ezrin | EZR | 0.93788 |
| P46777 | 60S ribosomal protein L5 | RPL5 | 0.939327 |
| P05387 | 60S acidic ribosomal protein P2 | RPLP2 | 0.94137 |
| P25398 | 40S ribosomal protein S12 | RPS12 | 0.941813 |
| P42704 | Leucine-rich PPR motif-containing protein, mitochondrial | LRPPRC | 0.944687 |
| P55072 | Transitional endoplasmic reticulum ATPase | VCP | 0.945267 |
| P62937 | Peptidyl-prolyl cis-trans isomerase A | PPIA | 0.94628 |
| P68363 | Tubulin α-1B chain | TUBA1B | 0.94723 |
| Q01628 | Interferon-induced transmembrane protein 3 | IFITM3 | 0.950665 |
| O00299 | Chloride intracellular channel protein 1 | CLIC1 | 0.952683 |
| P30040 | Endoplasmic reticulum resident protein 29 | ERP29 | 0.954027 |
| Q8N8S7 | Protein enabled homolog | ENAH | 0.954545 |
| P41252 | Isoleucine-tRNA ligase, cytoplasmic | IARS | 0.95526 |
| P61981 | 14-3-3 protein γ | YWHAG | 0.9565 |
| F8VPF3 | Myosin light polypeptide 6 | MYL6 | 0.95902 |
| O00571 | ATP-dependent RNA helicase DDX3X | DDX3X; DDX3Y | 0.959895 |
| P45880 | Voltage-dependent anion-selective channel protein 2 | VDAC2 | 0.960875 |
| P19338 | Nucleolin | NCL | 0.963977 |
| P14625 | Endoplasmin | HSP90B1 | 0.96555 |
| O43707 | α-Actinin-4 | ACTN4 | 0.968487 |
| Q5JP53 | Tubulin β chain | TUBB | 0.971943 |
| Q13409 | Cytoplasmic dynein 1 intermediate chain 2 | DYNC1I2 | 0.976747 |
| P55060 | Exportin-2 | CSE1L | 0.978383 |
| Q13838 | Spliceosome RNA helicase DDX39B | DDX39B | 0.979103 |
| P02786 | Transferrin receptor protein 1 | TFRC | 0.982483 |
| P00387 | NADH-cytochrome b5 reductase 3 | CYB5R3 | 0.98267 |
| O75947 | ATP synthase subunit d, mitochondrial | ATP5H | 0.985415 |
| G8JLD5 | Dynamin-1-like protein | DNM1L | 0.98906 |
| P04080 | Cystatin-B | CSTB | 0.98954 |
| P05198 | Eukaryotic translation initiation factor 2 subunit 1 | EIF2S1 | 0.98961 |
| Q9UL46 | Proteasome activator complex subunit 2 | PSME2 | 0.990545 |
| P55884 | Eukaryotic translation initiation factor 3 subunit B | EIF3B | 0.991517 |
| P46783 | 40S ribosomal protein S10 | RPS10; RPS10P5 | 0.99156 |
| F2Z2Y4 | Pyridoxal kinase | PDXK | 0.99206 |
| P09211 | Glutathione S-transferase P | GSTP1 | 0.992743 |
| P38646 | Stress-70 protein, mitochondrial | HSPA9 | 0.995527 |
| P80723 | Brain acid soluble protein 1 | BASP1 | 0.998527 |
| O15143 | Actin-related protein 2/3 complex subunit 1B | ARPC1B | 1.003 |
| Q8NC51 | Plasminogen activator inhibitor 1 RNA-binding protein | SERBP1 | 1.003403 |
| P54727 | UV excision repair protein RAD23 homolog B | RAD23B | 1.005245 |
| P51991 | Heterogeneous nuclear ribonucleoprotein A3 | HNRNPA3 | 1.006415 |
| P07355 | Annexin A2 | ANXA2; ANXA2P2 | 1.007367 |
| P23246 | Splicing factor, proline- and glutamine-rich | SFPQ | 1.009463 |
| P08107 | Heat shock 70 kDa protein 1A/1B | HSPA1A | 1.01165 |
| P67775 | Serine/threonine-protein phosphatase 2A catalytic subunit α isoform | PPP2CA; PPP2CB | 1.011925 |
| P21291 | Cysteine and glycine-rich protein 1 | CSRP1 | 1.01607 |
| P28066 | Proteasome subunit α type-5 | PSMA5 | 1.01847 |
| P68036 | Ubiquitin-conjugating enzyme E2 L3 | UBE2L3 | 1.02085 |
| Q9BUF5 | Tubulin β-6 chain | TUBB6 | 1.021827 |
| Q15459 | Splicing factor 3A subunit 1 | SF3A1 | 1.02408 |
| Q9P0L0 | Vesicle-associated membrane protein-associated protein A | VAPA | 1.026153 |
| C9JD32 | 60S ribosomal protein L23 | RPL23 | 1.026225 |
| Q15293 | Reticulocalbin-1 | RCN1 | 1.02665 |
| P55735 | Protein SEC13 homolog | SEC13 | 1.0291 |
| Q99880 | Histone H2B type 1-l | HIST1H2BL; HIST1H2BM; HIST1H2BN; HIST1H2BH; HIST2H2BF; HIST1H2BC; HIST1H2BD; H2BFS; HIST1H2BK | 1.02927 |
| D6RFM5 | Succinate dehydrogenase [ubiquinone] flavoprotein subunit, mitochondrial | SDHA | 1.034553 |
| P46821 | Microtubule-associated protein 1B | MAP1B | 1.03558 |
| Q5W0X3 | Peptidyl-prolyl cis-trans isomerase | FKBP1A | 1.0359 |
| K7EK07 | Histone H3 | H3F3B; H3F3A | 1.037015 |
| P52272 | Heterogeneous nuclear ribonucleoprotein M | HNRNPM | 1.03914 |
| P05023 | Sodium/potassium-transporting ATPase subunit α-1 | ATP1A1; ATP1A3 | 1.043157 |
| P36578 | 60S ribosomal protein L4 | RPL4 | 1.044293 |
| P08238 | Heat shock protein HSP 90-β | HSP90AB1 | 1.044997 |
| P68371 | Tubulin β-4B chain | TUBB4B | 1.045547 |
| Q01081 | Splicing factor U2AF 35 kDa subunit | U2AF1; U2AF1L4 | 1.0472 |
| P35637 | RNA-binding protein FUS | FUS | 1.049003 |
| P56192 | Methionine-tRNA ligase, cytoplasmic | MARS | 1.0493 |
| P06576 | ATP synthase subunit β, mitochondrial | ATP5B | 1.049537 |
| P04083 | Annexin A1 | ANXA1 | 1.0517 |
| P41250 | Glycine-tRNA ligase | GARS | 1.05216 |
| Q13813 | Spectrin αα chain, non-erythrocytic 1 | SPTAN1 | 1.0563 |
| Q9Y678 | Coatomer subunit γ-1 | COPG1 | 1.0569 |
| M0QZS6 | SUMO-activating enzyme subunit 1 | SAE1 | 1.0578 |
| Q9UHD8 | Septin-9 | 42256 | 1.063053 |
| P31946 | 14-3-3 protein β/α | YWHAB | 1.063833 |
| P63261 | Actin, cytoplasmic 2 | ACTG1 | 1.063967 |
| P13489 | Ribonuclease inhibitor | RNH1 | 1.064393 |
| P47897 | Glutamine-tRNA ligase | QARS | 1.0658 |
| B4DJV2 | Citrate synthase | CS | 1.0662 |
| P04843 | Dolichyl-diphosphooligosaccharide-protein glycosyltransferase subunit 1 | RPN1 | 1.066757 |
| Q14974 | Importin subunit β-1 | KPNB1 | 1.072667 |
| Q15121 | Astrocytic phosphoprotein PEA-15 | PEA15 | 1.072933 |
| Q9NYL9 | Tropomodulin-3 | TMOD3 | 1.0755 |
| O00410 | Importin-5 | IPO5 | 1.0767 |
| P26038 | Moesin | MSN | 1.078713 |
| Q9P2E9 | Ribosome-binding protein 1 | RRBP1 | 1.081993 |
| P68402 | Platelet-activating factor acetylhydrolase IB subunit β | PAFAH1B2 | 1.0887 |
| H0YEN5 | 40S ribosomal protein S2 | RPS2 | 1.091785 |
| Q5VU59 | TPM3 | 1.0933 | |
| P02545 | Prelamin-A/C | LMNA | 1.095077 |
| P35221 | Catenin α-1 | CTNNA1 | 1.09635 |
| Q08211 | ATP-dependent RNA helicase A | DHX9 | 1.096937 |
| P50991 | T-complex protein 1 subunit δ | CCT4 | 1.103 |
| O60493 | Sorting nexin-3 | SNX3 | 1.1169 |
| P61160 | Actin-related protein 2 | ACTR2 | 1.1175 |
| O95433 | Activator of 90 kDa heat shock protein ATPase homolog 1 | AHSA1 | 1.1199 |
| P62805 | Histone H4 | HIST1H4A | 1.1241 |
| P18465 | HLA class I histocompatibility antigen, B-57 α chain | HLA-B | 1.12765 |
| Q8TAT6 | Nuclear protein localization protein 4 homolog | NPLOC4 | 1.135 |
| P46940 | Ras GTPase-activating-like protein IQGAP1 | IQGAP1 | 1.135853 |
| P09972 | Fructose-bisphosphate aldolase C | ALDOC | 1.1406 |
| P62424 | 60S ribosomal protein L7a | RPL7A | 1.14125 |
| Q00610 | Clathrin heavy chain 1 | CLTC | 1.142133 |
| O15355 | Protein phosphatase 1G | PPM1G | 1.1437 |
| Q16222 | UDP-N-acetylhexosamine pyrophosphorylase | UAP1 | 1.145295 |
| P25705 | ATP synthase subunit α, mitochondrial | ATP5A1 | 1.14579 |
| E7ETU9 | Procollagen-lysine, 2-oxoglutarate 5-dioxygenase 2 | PLOD2 | 1.150217 |
| K7EJ78 | 40S ribosomal protein S15 | RPS15 | 1.1525 |
| P52292 | Importin subunit α-1 | KPNA2 | 1.153933 |
| Q15417 | Calponin-3 | CNN3 | 1.154837 |
| E7ETK0 | 40S ribosomal protein S24 | RPS24 | 1.1574 |
| P51149 | Ras-related protein Rab-7a | RAB7A | 1.158243 |
| Q9NTK5 | OBG-like ATPase 1 | OLA1 | 1.1594 |
| Q5URX0 | β-Hexosaminidase subunit β | HEXB | 1.159533 |
| P12814 | α-Actinin-1 | ACTN1 | 1.165233 |
| F8VVM2 | Phosphate carrier protein, mitochondrial | SLC25A3 | 1.165257 |
| Q9Y224 | UPF0568 protein C14orf166 | C14orf166 | 1.1655 |
| P06748 | Nucleophosmin | NPM1 | 1.1659 |
| P30101 | Protein disulfide-isomerase A3 | PDIA3 | 1.167067 |
| P26641 | Elongation factor 1-γ | EEF1G | 1.1681 |
| P78371 | T-complex protein 1 subunit β | CCT2 | 1.169767 |
| F8VY35 | Nucleosome assembly protein 1-like 1 | NAP1L1 | 1.17258 |
| P56537 | Eukaryotic translation initiation factor 6 | EIF6 | 1.172633 |
| E7EV56 | Pericentriolar material 1 protein | PCM1 | 1.1749 |
| P26599 | Polypyrimidine tract-binding protein 1 | PTBP1 | 1.1785 |
| P27348 | 14-3-3 protein θ | YWHAQ | 1.180407 |
| P22314 | Ubiquitin-like modifier-activating enzyme 1 | UBA1 | 1.182133 |
| D6RG13 | 40S ribosomal protein S3a | RPS3A | 1.184367 |
| P00367 | Glutamate dehydrogenase 1, mitochondrial | GLUD1; GLUD2 | 1.1844 |
| E9PNW4 | CD59 glycoprotein | CD59 | 1.1867 |
| P11717 | Cation-independent mannose-6-phosphate receptor | IGF2R | 1.1869 |
| P17987 | T-complex protein 1 subunit α | TCP1 | 1.1904 |
| P60842 | Eukaryotic initiation factor 4A–I | EIF4A1 | 1.191733 |
| P04632 | Calpain small subunit 1 | CAPNS1 | 1.19265 |
| P07900 | Heat shock protein HSP 90-α | HSP90AA1 | 1.195633 |
| P20618 | Proteasome subunit β type-1 | PSMB1 | 1.2078 |
| J3KPE3 | Guanine nucleotide-binding protein subunit β-2-like 1 | GNB2L1 | 1.209467 |
| P35613 | Basigin | BSG | 1.21668 |
| P31930 | Cytochrome b-c1 complex subunit 1, mitochondrial | UQCRC1 | 1.217533 |
| P31942 | Heterogeneous nuclear ribonucleoprotein H3 | HNRNPH3 | 1.223433 |
| E9PCY7 | Heterogeneous nuclear ribonucleoprotein H | HNRNPH1 | 1.2236 |
| F8W7C6 | RPL10 | 1.225633 | |
| Q07021 | Complement component 1 Q subcomponent-binding protein, mitochondrial | C1QBP | 1.2273 |
| Q96AG4 | Leucine-rich repeat-containing protein 59 | LRRC59 | 1.227793 |
| Q14315 | Filamin-C | FLNC | 1.2332 |
| P48643 | T-complex protein 1 subunit ε | CCT5 | 1.234767 |
| E7EQV3 | Polyadenylate-binding protein 1 | PABPC1; PABPC4 | 1.235133 |
| P62701 | 40S ribosomal protein S4, X isoform | RPS4X | 1.2362 |
| P29692 | Elongation factor 1-δ | EEF1D | 1.239133 |
| P53396 | ATP-citrate synthase | ACLY | 1.241963 |
| Q12906 | Interleukin enhancer-binding factor 3 | ILF3 | 1.245833 |
| P23396 | 40S ribosomal protein S3 | RPS3 | 1.2509 |
| P50990 | T-complex protein 1 subunit θ | CCT8 | 1.257 |
| P11142 | Heat shock cognate 71 kDa protein | HSPA8 | 1.258433 |
| Q12905 | Interleukin enhancer-binding factor 2 | ILF2 | 1.2642 |
| H0Y3Y4 | Septin-7 | 42254 | 1.2656 |
| F8VZX2 | Poly(rC)-binding protein 2 | PCBP2 | 1.265963 |
| C9J9K3 | 40S ribosomal protein SA | RPSA; RPSAP58 | 1.2668 |
| P35580 | Myosin-10 | MYH10 | 1.275013 |
| P23526 | Adenosylhomocysteinase | AHCY | 1.276 |
| P30050 | 60S ribosomal protein L12 | RPL12 | 1.287833 |
| H3BT13 | Small nuclear ribonucleoprotein Sm D3 | SNRPD3 | 1.288 |
| I7HJJ0 | ADP/ATP translocase 3 | SLC25A6; SLC25A4 | 1.30365 |
| E7EMC7 | Sequestosome-1 | SQSTM1 | 1.3099 |
| P52907 | F-actin-capping protein subunit α-1 | CAPZA1 | 1.3127 |
| Q01813 | 6-Phosphofructokinase type C | PFKP | 1.312933 |
| Q02878 | 60S ribosomal protein L6 | RPL6 | 1.3141 |
| P19105 | Myosin regulatory light chain 12A | MYL12A; MYL12B; MYL9 | 1.315967 |
| Q14019 | Coactosin-like protein | COTL1 | 1.318367 |
| Q5TCI8 | LMNA | 1.318467 | |
| P20290 | Transcription factor BTF3 | BTF3 | 1.324333 |
| Q9UJZ1 | Stomatin-like protein 2, mitochondrial | STOML2 | 1.3251 |
| Q15149 | Plectin | PLEC | 1.325767 |
| Q5T8U5 | Surfeit locus protein 4 | SURF4 | 1.3268 |
| F8W6I7 | Heterogeneous nuclear ribonucleoprotein A1 | HNRNPA1; HNRNPA1L2 | 1.329333 |
| J3QSX4 | Mitotic checkpoint protein BUB3 | BUB3 | 1.3315 |
| P33176 | Kinesin-1 heavy chain | KIF5B | 1.3317 |
| P62280 | 40S ribosomal protein S11 | RPS11 | 1.33505 |
| P17812 | CTP synthase 1 | CTPS1 | 1.3383 |
| P35268 | 60S ribosomal protein L22 | RPL22 | 1.338767 |
| Q13200 | 26S proteasome non-ATPase regulatory subunit 2 | PSMD2 | 1.33905 |
| P18124 | 60S ribosomal protein L7 | RPL7 | 1.340267 |
| Q13162 | Peroxiredoxin-4 | PRDX4 | 1.3457 |
| O95373 | Importin-7 | IPO7 | 1.355167 |
| O75533 | Splicing factor 3B subunit 1 | SF3B1 | 1.35925 |
| O00622 | Protein CYR61 | CYR61 | 1.36705 |
| P61353 | 60S ribosomal protein L27 | RPL27 | 1.373033 |
| Q15233 | Non-POU domain-containing octamer-binding protein | NONO | 1.3738 |
| Q99613 | Eukaryotic translation initiation factor 3 subunit C | EIF3C; EIF3CL | 1.37415 |
| P08574 | Cytochrome c1, heme protein, mitochondrial | CYC1 | 1.37425 |
| P98179 | Putative RNA-binding protein 3 | RBM3 | 1.37635 |
| P17655 | Calpain-2 catalytic subunit | CAPN2 | 1.386833 |
| Q86UY0 | Thioredoxin domain-containing protein 5 | TXNDC5 | 1.390233 |
| P62979 | Ubiquitin-40S ribosomal protein S27a | RPS27A; UBB; UBC; UBA52; UBBP4 | 1.399267 |
| Q86VP6 | Cullin-associated NEDD8-dissociated protein 1 | CAND1 | 1.4086 |
| Q15691 | Microtubule-associated protein RP/EB family member 1 | MAPRE1 | 1.4179 |
| Q7L2H7 | Eukaryotic translation initiation factor 3 subunit M | EIF3M | 1.45085 |
| E7EX73 | Eukaryotic translation initiation factor 4 γ 1 | EIF4G1 | 1.4581 |
| X1WI28 | 60S ribosomal protein L10 | RPL10 | 1.4608 |
| P17844 | Probable ATP-dependent RNA helicase DDX5 | DDX5 | 1.470067 |
| P04899 | Guanine nucleotide-binding protein Gi subunit α-2 | GNAI2 | 1.5439 |
| Q16891 | Mitochondrial inner membrane protein | IMMT | 1.5646 |
| P08670 | Vimentin | VIM | 1.578767 |
| P62241 | 40S ribosomal protein S8 | RPS8 | 1.5809 |
| Q14764 | Major vault protein | MVP | 1.63639 |
| P31689 | DNAJ homolog subfamily A member 1 | DNAJA1 | 1.6699 |
| Q9H299 | SH3 domain-binding glutamic acid-rich-like protein 3 | SH3BGRL3 | 1.68785 |
| Q07065 | Cytoskeleton-associated protein 4 | CKAP4 | 1.728867 |
| Q86V81 | THO complex subunit 4 | ALYREF | 1.7965 |
| Q5TCU3 | Tropomyosin β chain | TPM2 | 1.8451 |
| O00505 | Importin subunit α-4 | KPNA3 | 2.1267 |
| Q6NZI2 | Polymerase I and transcript release factor | PTRF | 2.1653 |
| H0YD13 | CD44 antigen | CD44 | 2.2206 |
| Q9BSJ8 | Extended synaptotagmin-1 | ESYT1 | 2.36957 |
| P10619 | Lysosomal protective protein | CTSA | 3.0967 |
| ID | Symbol | Entrez Gene Name | Location | Type(s) | Fold Change | |
|---|---|---|---|---|---|---|
| P49588 | AARS | Alanyl-tRNA synthetase | Cytoplasm | Enzyme | −1.209 | |
| P24752 | ACAT1 | Acetyl-CoA acetyltransferase 1 | Cytoplasm | Enzyme | −1.360 | |
| Q9BWD1 | ACAT2 | Acetyl-CoA acetyltransferase 2 | Cytoplasm | Enzyme | −1.274 | |
| P53396 | ACLY | ATP citrate lyase | Cytoplasm | Enzyme | 1.242 | |
| P68133 | ACTA1 | Actin, α 1, skeletal muscle | Cytoplasm | Other | −1.131 | |
| P63261 | ACTG1 | Actin, γ 1 | Cytoplasm | Other | 1.064 | |
| P12814 | ACTN1 | Actinin, α 1 | Cytoplasm | Other | 1.165 | |
| O43707 | ACTN4 | Actinin, α 4 | Cytoplasm | Other | −1.033 | |
| P61160 | ACTR2 | ARP2 actin-related protein 2 homolog (yeast) | Plasma Membrane | Other | 1.118 | |
| P61158 | ACTR3 | ARP3 actin-related protein 3 homolog (yeast) | Plasma Membrane | Other | −1.090 | |
| P23526 | AHCY | Adenosylhomocysteinase | Cytoplasm | Enzyme | 1.276 | |
| Q09666 | AHNAK | AHNAK nucleoprotein | Nucleus | Other | −1.233 | |
| O95433 | AHSA1 | AHA1, activator of heat shock 90 kDa protein ATPase homolog 1 (yeast) | Cytoplasm | Other | 1.120 | |
| E9PMH2 | AIP | Aryl hydrocarbon receptor interacting protein | Nucleus | Transcription regulator | −1.330 | |
| E9PCX2 | AKR1B1 | Aldo-keto reductase family 1, member B1 (aldose reductase) | Cytoplasm | Enzyme | −3.673 | |
| P04075 | ALDOA | Aldolase A, fructose-bisphosphate | Cytoplasm | Enzyme | −1.090 | |
| P09972 | ALDOC | Aldolase C, fructose-bisphosphate | Cytoplasm | Enzyme | 1.141 | |
| Q86V81 | ALYREF | Aly/REF export factor | Nucleus | Transcription regulator | 1.796 | |
| P04083 | ANXA1 | Annexin A1 | Plasma Membrane | Enzyme | 1.052 | |
| P07355 | ANXA2 | Annexin A2 | Plasma Membrane | Other | 1.007 | |
| P08758 | ANXA5 | Annexin A5 | Plasma Membrane | Other | −1.294 | |
| P08133 | ANXA6 | Annexin A6 | Plasma Membrane | Ion channel | −1.080 | |
| Q10567 | AP1B1 | Adaptor-related protein complex 1, β 1 subunit | Cytoplasm | Transporter | −1.336 | |
| P63010 | AP2B1 | Adaptor-related protein complex 2, β 1 subunit | Plasma Membrane | Transporter | −1.400 | |
| P27695 | APEX1 | APEX nuclease (multifunctional DNA repair enzyme) 1 | Nucleus | Enzyme | −1.321 | |
| P84077 | ARF1 | ADP-ribosylation factor 1 | Cytoplasm | Enzyme | −1.231 | |
| J3KTF8 | ARHGDIA | Rho GDP dissociation inhibitor (GDI) α | Cytoplasm | Other | −1.768 | |
| O15143 | ARPC1B | Actin related protein 2/3 complex, subunit 1B, 41 kDa | Cytoplasm | Other | 1.003 | |
| O15144 | ARPC2 | Actin related protein 2/3 complex, subunit 2, 34 kDa | Cytoplasm | Other | −1.072 | |
| P31939 | ATIC | 5-Aminoimidazole-4-carboxamide ribonucleotide formyltransferase/IMP cyclohydrolase | Cytoplasm | Enzyme | −1.304 | |
| P05023 | ATP1A1 | ATPase, Na+/K+ transporting, α1 polypeptide | Plasma Membrane | Transporter | 1.043 | |
| P25705 | ATP5A1 | ATP synthase, H+ transporting, mitochondrial F1 complex, α subunit 1, cardiac muscle | Cytoplasm | Transporter | 1.146 | |
| P06576 | ATP5B | ATP synthase, H+ transporting, mitochondrial F1 complex, β polypeptide | Cytoplasm | Transporter | 1.050 | |
| O75947 | ATP5H | ATP synthase, H+ transporting, mitochondrial Fo complex, subunit d | Cytoplasm | Enzyme | −1.015 | |
| P61769 | B2M | β2-Microglobulin | Plasma Membrane | Transmembrane receptor | −1.177 | |
| P80723 | BASP1 | Brain abundant, membrane attached signal protein 1 | Nucleus | Transcription regulator | −1.001 | |
| P35613 | BSG | Basigin (Ok blood group) | Plasma Membrane | Transporter | 1.217 | |
| P20290 | BTF3 | Basic transcription factor 3 | Nucleus | Transcription regulator | 1.324 | |
| J3QSX4 | BUB3 | BUB3 mitotic checkpoint protein | Nucleus | Other | 1.332 | |
| Q9Y224 | C14orf166 | Chromosome 14 open reading frame 166 | Nucleus | Other | 1.166 | |
| Q07021 | C1QBP | Complement component 1, q subcomponent binding protein | Cytoplasm | Transcription regulator | 1.227 | |
| Q9HB71 | CACYBP | Calcyclin binding protein | Nucleus | Other | −2.006 | |
| P27797 | CALR | Calreticulin | Cytoplasm | Transcription regulator | −1.385 | |
| O43852 | CALU | Calumenin | Cytoplasm | Other | −1.270 | |
| Q86VP6 | CAND1 | Cullin-associated and neddylation-dissociated 1 | Cytoplasm | Transcription regulator | 1.409 | |
| P27824 | CANX | calnexin | Cytoplasm | Other | −1.140 | |
| Q01518 | CAP1 | CAP, adenylate cyclase-associated protein 1 (yeast) | Plasma Membrane | Other | −1.222 | |
| P40121 | CAPG | Capping protein (actin filament), gelsolin-like | Nucleus | Other | −1.145 | |
| P07384 | CAPN1 | Calpain 1, (mu/I) large subunit | Cytoplasm | Peptidase | −1.241 | |
| P17655 | CAPN2 | Calpain 2, (m/II) large subunit | Cytoplasm | Peptidase | 1.387 | |
| P04632 | CAPNS1 | Calpain, small subunit 1 | Cytoplasm | Peptidase | 1.193 | |
| Q14444 | CAPRIN1 | Cell cycle associated protein 1 | Plasma Membrane | Other | −1.230 | |
| P52907 | CAPZA1 | Capping protein (actin filament) muscle Z-line, α 1 | Cytoplasm | Other | 1.313 | |
| B1AK85 | CAPZB | Capping protein (actin filament) muscle Z-line, β | Cytoplasm | Other | −1.134 | |
| Q9Y2V2 | CARHSP1 | Calcium regulated heat stable protein 1, 24 kDa | Cytoplasm | Other | −2.419 | |
| E7ES10 | CAST | Calpastatin | Cytoplasm | Peptidase | −2.137 | |
| Q03135 | CAV1 | Caveolin 1, caveolae protein, 22 kDa | Plasma Membrane | Transmembrane receptor | −1.143 | |
| Q13185 | CBX3 | Chromobox homolog 3 | Nucleus | Transcription regulator | −2.913 | |
| P78371 | CCT2 | Chaperonin containing TCP1, subunit 2 (β) | Cytoplasm | Kinase | 1.170 | |
| B4DUR8 | CCT3 | Chaperonin containing TCP1, subunit 3 (γ) | Cytoplasm | Other | −1.116 | |
| P50991 | CCT4 | Chaperonin containing TCP1, subunit 4 (δ) | Cytoplasm | Other | 1.103 | |
| P48643 | CCT5 | Chaperonin containing TCP1, subunit 5 (ε) | Cytoplasm | Other | 1.235 | |
| P40227 | CCT6A | Chaperonin containing TCP1, subunit 6A (ζ 1) | Cytoplasm | Other | −1.202 | |
| Q99832 | CCT7 | Chaperonin containing TCP1, subunit 7 (η) | Cytoplasm | Other | −1.226 | |
| P50990 | CCT8 | Chaperonin containing TCP1, subunit 8 (θ) | Cytoplasm | Enzyme | 1.257 | |
| H0YD13 | CD44 | CD44 molecule (Indian blood group) | Plasma Membrane | Enzyme | 2.221 | |
| E9PNW4 | CD59 | CD59 molecule, complement regulatory protein | Plasma Membrane | Other | 1.187 | |
| P06493 | CDK1 | Cyclin-dependent kinase 1 | Nucleus | Kinase | −1.130 | |
| Q9UKY7 | CDV3 | CDV3 homolog (mouse) | Cytoplasm | Other | −1.718 | |
| P23528 | CFL1 | Cofilin 1 (non-muscle) | Nucleus | Other | −1.408 | |
| Q07065 | CKAP4 | Cytoskeleton-associated protein 4 | Cytoplasm | Other | 1.729 | |
| O00299 | CLIC1 | Chloride intracellular channel 1 | Nucleus | Ion channel | -1.050 | |
| Q9Y696 | CLIC4 | Chloride intracellular channel 4 | Plasma Membrane | Ion channel | −1.816 | |
| Q00610 | CLTC | Clathrin, heavy chain (Hc) | Plasma Membrane | Other | 1.142 | |
| Q15417 | CNN3 | Calponin 3, acidic | Cytoplasm | Other | 1.155 | |
| P53621 | COPA | Coatomer protein complex, subunit α | Cytoplasm | Transporter | −1.537 | |
| P53618 | COPB1 | Coatomer protein complex, subunit β 1 | Cytoplasm | Transporter | −1.350 | |
| P35606 | COPB2 | Coatomer protein complex, subunit β 2 (β prime) | Cytoplasm | Transporter | −1.323 | |
| O14579 | COPE | Coatomer protein complex, subunit ε | Cytoplasm | Transporter | −1.375 | |
| Q9Y678 | COPG1 | Coatomer protein complex, subunit γ 1 | Cytoplasm | Transporter | 1.057 | |
| Q9ULV4 | CORO1C | Coronin, actin binding protein, 1C | Cytoplasm | Other | −1.438 | |
| Q14019 | COTL1 | Coactosin-like F-actin binding protein 1 | Cytoplasm | Other | 1.318 | |
| A6PVH9 | CPNE1 | Copine I | Nucleus | Transporter | −1.093 | |
| H0YFA4 | CRIP2 | Cysteine-rich protein 2 | Other | Other | −1.947 | |
| B4DJV2 | CS | Citrate synthase | Cytoplasm | Enzyme | 1.066 | |
| P55060 | CSE1L | CSE1 chromosome segregation 1-like (yeast) | Nucleus | Transporter | −1.022 | |
| P21291 | CSRP1 | Cysteine and glycine-rich protein 1 | Nucleus | Other | 1.016 | |
| P04080 | CSTB | Cystatin B (stefin B) | Cytoplasm | Peptidase | −1.011 | |
| P35221 | CTNNA1 | Catenin (cadherin-associated protein), α 1, 102 kDa | Plasma Membrane | Other | 1.096 | |
| P17812 | CTPS1 | CTP synthase 1 | Nucleus | Enzyme | 1.338 | |
| P10619 | CTSA | Cathepsin A | Cytoplasm | Peptidase | 3.097 | |
| P07858 | CTSB | Cathepsin B | Cytoplasm | Peptidase | −1.125 | |
| Q14247 | CTTN | Cortactin | Plasma Membrane | Other | −1.161 | |
| O60888 | CUTA | CutA divalent cation tolerance homolog (E. coli) | Cytoplasm | Other | −1.324 | |
| P00387 | CYB5R3 | Cytochrome b5 reductase 3 | Cytoplasm | Enzyme | −1.018 | |
| P08574 | CYC1 | Cytochrome c-1 | Cytoplasm | Enzyme | 1.374 | |
| O00622 | CYR61 | Cysteine-rich, angiogenic inducer, 61 | Extracellular Space | Other | 1.367 | |
| P07108 | DBI | Diazepam binding inhibitor (GABA receptor modulator, acyl-CoA binding protein) | Cytoplasm | Other | −1.099 | |
| Q16643 | DBN1 | Drebrin 1 | Cytoplasm | Other | −1.109 | |
| Q92499 | DDX1 | DEAD (Asp-Glu-Ala-Asp) box helicase 1 | Nucleus | Enzyme | −1.229 | |
| Q13838 | DDX39B | DEAD (Asp-Glu-Ala-Asp) box polypeptide 39B | Nucleus | Enzyme | −1.021 | |
| O00571 | DDX3X | DEAD (Asp-Glu-Ala-Asp) box helicase 3, X-linked | Cytoplasm | Enzyme | −1.042 | |
| P17844 | DDX5 | DEAD (Asp-Glu-Ala-Asp) box helicase 5 | Nucleus | Enzyme | 1.470 | |
| P26196 | DDX6 | DEAD (Asp-Glu-Ala-Asp) box helicase 6 | Nucleus | Enzyme | −1.499 | |
| Q08211 | DHX9 | DEAH (Asp-Glu-Ala-His) box helicase 9 | Nucleus | Enzyme | 1.097 | |
| E9PEJ4 | DLAT | Dihydrolipoamide S-acetyltransferase | Cytoplasm | Enzyme | −2.153 | |
| P31689 | DNAJA1 | DnaJ (Hsp40) homolog, subfamily A, member 1 | Nucleus | Other | 1.670 | |
| O60884 | DNAJA2 | DnaJ (Hsp40) homolog, subfamily A, member 2 | Nucleus | Enzyme | −1.534 | |
| O75937 | DNAJC8 | DnaJ (Hsp40) homolog, subfamily C, member 8 | Nucleus | Other | −1.971 | |
| G8JLD5 | DNM1L | Dynamin 1-like | Cytoplasm | Enzyme | −1.011 | |
| Q9C005 | DPY30 | Dpy-30 homolog (C. elegans) | Nucleus | Other | −1.737 | |
| Q16555 | DPYSL2 | Dihydropyrimidinase-like 2 | Cytoplasm | Enzyme | −1.124 | |
| Q14204 | DYNC1H1 | Dynein, cytoplasmic 1, heavy chain 1 | Cytoplasm | Peptidase | −1.084 | |
| Q13409 | DYNC1I2 | Dynein, cytoplasmic 1, intermediate chain 2 | Cytoplasm | Other | −1.024 | |
| P30084 | ECHS1 | Enoyl CoA hydratase, short chain, 1, mitochondrial | Cytoplasm | Enzyme | −1.351 | |
| P68104 | EEF1A1 | Eukaryotic translation elongation factor 1 α 1 | Cytoplasm | Transcription regulator | −1.207 | |
| P24534 | EEF1B2 | Eukaryotic translation elongation factor 1 β 2 | Cytoplasm | Transcription regulator | −1.208 | |
| P29692 | EEF1D | Eukaryotic translation elongation factor 1 delta (guanine nucleotide exchange protein) | Cytoplasm | Transcription regulator | 1.239 | |
| P26641 | EEF1G | Eukaryotic translation elongation factor 1 γ | Cytoplasm | Transcription regulator | 1.168 | |
| P13639 | EEF2 | Eukaryotic translation elongation factor 2 | Cytoplasm | Transcription regulator | −1.089 | |
| Q9H4M9 | EHD1 | EH-domain containing 1 | Cytoplasm | Other | −1.647 | |
| P05198 | EIF2S1 | Eukaryotic translation initiation factor 2, subunit 1 α, 35 kDa | Cytoplasm | Transcription regulator | −1.010 | |
| P20042 | EIF2S2 | Eukaryotic translation initiation factor 2, subunit 2 β, 38 kDa | Cytoplasm | Transcription regulator | −1.660 | |
| P55884 | EIF3B | Eukaryotic translation initiation factor 3, subunit B | Cytoplasm | Transcription regulator | −1.009 | |
| Q99613 | EIF3C | Eukaryotic translation initiation factor 3, subunit C | Other | Transcription regulator | 1.374 | |
| Q13347 | EIF3I | Eukaryotic translation initiation factor 3, subunit I | Cytoplasm | Transcription regulator | −1.163 | |
| Q7L2H7 | EIF3M | Eukaryotic translation initiation factor 3, subunit M | Other | Other | 1.451 | |
| P60842 | EIF4A1 | Eukaryotic translation initiation factor 4A1 | Cytoplasm | Transcription regulator | 1.192 | |
| E7EX73 | EIF4G1 | Eukaryotic translation initiation factor 4 γ, 1 | Cytoplasm | Transcription regulator | 1.458 | |
| Q15056 | EIF4H | Eukaryotic translation initiation factor 4H | Cytoplasm | Transcription regulator | −1.387 | |
| P63241 | EIF5A | Eukaryotic translation initiation factor 5A | Cytoplasm | Transcription regulator | −1.210 | |
| P56537 | EIF6 | Eukaryotic translation initiation factor 6 | Cytoplasm | Transcription regulator | 1.173 | |
| Q8N8S7 | ENAH | Enabled homolog (Drosophila) | Plasma Membrane | Other | −1.048 | |
| P06733 | ENO1 | Enolase 1, (α) | Cytoplasm | Enzyme | −1.199 | |
| P09104 | ENO2 | Enolase 2 (γ, neuronal) | Cytoplasm | Enzyme | −1.698 | |
| P07814 | EPRS | Glutamyl-prolyl-tRNA synthetase | Cytoplasm | Enzyme | −1.568 | |
| P30040 | ERP29 | Endoplasmic reticulum protein 29 | Cytoplasm | Transporter | −1.048 | |
| P10768 | ESD | Esterase D | Cytoplasm | Enzyme | −1.323 | |
| Q9BSJ8 | ESYT1 | Extended synaptotagmin-like protein 1 | Cytoplasm | Other | 2.370 | |
| H0YK49 | ETFA | Electron-transfer-flavoprotein, α polypeptide | Cytoplasm | Transporter | −2.144 | |
| E7EQR4 | EZR | Ezrin | Plasma Membrane | Other | −1.066 | |
| P49327 | FASN | Fatty acid synthase | Cytoplasm | Enzyme | −1.453 | |
| P14324 | FDPS | Farnesyl diphosphate synthase | Cytoplasm | Enzyme | −1.366 | |
| Q5W0X3 | FKBP1A | FK506 binding protein 1A, 12 kDa | Cytoplasm | Enzyme | 1.036 | |
| Q02790 | FKBP4 | FK506 binding protein 4, 59 kDa | Nucleus | Enzyme | −1.266 | |
| P21333 | FLNA | Filamin A, α | Cytoplasm | Other | −1.157 | |
| O75369 | FLNB | Filamin B, β | Cytoplasm | Other | −1.360 | |
| Q14315 | FLNC | Filamin C, γ | Cytoplasm | Other | 1.233 | |
| P02751 | FN1 | Fibronectin 1 | Extracellular Space | Enzyme | −3.472 | |
| Q96I24 | FUBP3 | Far upstream element (FUSE) binding protein 3 | Nucleus | Transcription regulator | −1.274 | |
| P35637 | FUS | FUS RNA binding protein | Nucleus | Transcription regulator | 1.049 | |
| Q13283 | G3BP1 | GTPase activating protein (SH3 domain) binding protein 1 | Nucleus | Enzyme | −1.497 | |
| P11413 | G6PD | Flucose-6-phosphate dehydrogenase | Cytoplasm | Enzyme | −1.504 | |
| Q14697 | GANAB | Flucosidase, α; neutral AB | Cytoplasm | Enzyme | −1.122 | |
| P04406 | GAPDH | Flyceraldehyde-3-phosphate dehydrogenase | Cytoplasm | Enzyme | −1.070 | |
| P41250 | GARS | Flycyl-tRNA synthetase | Cytoplasm | Enzyme | 1.052 | |
| Q92616 | GCN1L1 | GCN1 general control of amino-acid synthesis 1-like 1 (yeast) | Cytoplasm | Transcription regulator | −1.604 | |
| P31150 | GDI1 | GDP dissociation inhibitor 1 | Cytoplasm | Other | −1.630 | |
| P50395 | GDI2 | GDP dissociation inhibitor 2 | Cytoplasm | Other | −1.169 | |
| Q04760 | GLO1 | Flyoxalase I | Cytoplasm | Enzyme | −1.349 | |
| P00367 | GLUD1 | Flutamate dehydrogenase 1 | Cytoplasm | Enzyme | 1.184 | |
| E9PIR7 | GML | Flycosylphosphatidylinositol anchored molecule like | Plasma Membrane | Other | −1.146 | |
| P49915 | GMPS | Fuanine monphosphate synthase | Nucleus | Enzyme | −1.948 | |
| P04899 | GNAI2 | Fuanine nucleotide binding protein (G protein), α inhibiting activity polypeptide 2 | Plasma Membrane | Other | 1.544 | |
| J3KPE3 | GNB2L1 | Fuanine nucleotide binding protein (G protein), β polypeptide 2-like 1 | Cytoplasm | Enzyme | 1.209 | |
| P06744 | GPI | Flucose-6-phosphate isomerase | Extracellular Space | Enzyme | −1.324 | |
| P09211 | GSTP1 | Flutathione S-transferase pi 1 | Cytoplasm | Enzyme | −1.007 | |
| O75367 | H2AFY | H2A histone family, member Y | Nucleus | Other | −1.571 | |
| K7EK07 | H3F3A/H3F3B | H3 histone, family 3A | Nucleus | Other | 1.037 | |
| Q16836 | HADH | Hydroxyacyl-CoA dehydrogenase | Cytoplasm | Enzyme | −1.626 | |
| P40939 | HADHA | Hydroxyacyl-CoA dehydrogenase/3-ketoacyl-CoA thiolase/enoyl-CoA hydratase (trifunctional protein), α subunit | Cytoplasm | Enzyme | −1.342 | |
| B4DDD8 | HARS | Histidyl-tRNA synthetase | Cytoplasm | Enzyme | −1.358 | |
| P51858 | HDGF | Hepatoma-derived growth factor | Extracellular Space | Growth factor | −2.205 | |
| Q00341 | HDLBP | High density lipoprotein binding protein | Nucleus | Transporter | −1.572 | |
| Q5URX0 | HEXB | Hexosaminidase B (β polypeptide) | Cytoplasm | Enzyme | 1.160 | |
| P49773 | HINT1 | Histidine triad nucleotide binding protein 1 | Nucleus | Enzyme | −1.864 | |
| P16403 | HIST1H1C | Histone cluster 1, H1c | Nucleus | Other | −1.808 | |
| Q99880 | HIST1H2BL | Histone cluster 1, H2bl | Nucleus | Other | 1.029 | |
| Q16777 | HIST2H2AC | Histone cluster 2, H2ac | Nucleus | Other | −1.380 | |
| P30447 | HLA-A | Major histocompatibility complex, class I, A | Plasma Membrane | Other | −1.339 | |
| P18465 | HLA-B | Major histocompatibility complex, class I, B | Plasma Membrane | Transmembrane receptor | 1.128 | |
| Q5T7C4 | HMGB1 | High mobility group box 1 | Nucleus | Transcription regulator | −3.172 | |
| Q9UK76 | HN1 | Hematological and neurological expressed 1 | Nucleus | Other | −1.199 | |
| F8W6I7 | HNRNPA1 | Heterogeneous nuclear ribonucleoprotein A1 | Nucleus | Other | 1.329 | |
| P22626 | HNRNPA2B1 | Heterogeneous nuclear ribonucleoprotein A2/B1 | Nucleus | Other | −1.102 | |
| P51991 | HNRNPA3 | Heterogeneous nuclear ribonucleoprotein A3 | Nucleus | Other | 1.006 | |
| E9PCY7 | HNRNPH1 | Heterogeneous nuclear ribonucleoprotein H1 (H) | Nucleus | Other | 1.224 | |
| P31942 | HNRNPH3 | Heterogeneous nuclear ribonucleoprotein H3 (2H9) | Nucleus | Other | 1.223 | |
| P61978 | HNRNPK | Heterogeneous nuclear ribonucleoprotein K | Nucleus | Other | −1.191 | |
| P14866 | HNRNPL | Heterogeneous nuclear ribonucleoprotein L | Nucleus | Other | −1.360 | |
| P52272 | HNRNPM | Heterogeneous nuclear ribonucleoprotein M | Nucleus | Other | 1.039 | |
| O43390 | HNRNPR | Heterogeneous nuclear ribonucleoprotein R | Nucleus | Other | −1.090 | |
| Q00839 | HNRNPU | Heterogeneous nuclear ribonucleoprotein U (scaffold attachment factor A) | Nucleus | Transporter | −1.169 | |
| Q99714 | HSD17B10 | Hydroxysteroid (17-β) dehydrogenase 10 | Cytoplasm | Enzyme | −1.586 | |
| P07900 | HSP90AA1 | Heat shock protein 90 kDa α (cytosolic), class A member 1 | Cytoplasm | Enzyme | 1.196 | |
| P08238 | HSP90AB1 | Heat shock protein 90 kDa α (cytosolic), class B member 1 | Cytoplasm | Enzyme | 1.045 | |
| P14625 | HSP90B1 | Heat shock protein 90 kDa β (GRP94), member 1 | Cytoplasm | Other | −1.036 | |
| P34932 | HSPA4 | Heat shock 70 kDa protein 4 | Cytoplasm | Other | −1.068 | |
| P11021 | HSPA5 | Heat shock 70 kDa protein 5 (glucose-regulated protein, 78 kDa) | Cytoplasm | Enzyme | −1.178 | |
| P11142 | HSPA8 | Heat shock 70 kDa protein 8 | Cytoplasm | Enzyme | 1.258 | |
| P38646 | HSPA9 | Heat shock 70 kDa protein 9 (mortalin) | Cytoplasm | Other | −1.004 | |
| P04792 | HSPB1 | Heat shock 27 kDa protein 1 | Cytoplasm | Other | −1.941 | |
| P10809 | HSPD1 | Heat shock 60 kDa protein 1 (chaperonin) | Cytoplasm | Enzyme | −1.182 | |
| Q9Y4L1 | HYOU1 | Hypoxia up-regulated 1 | Cytoplasm | Other | −1.437 | |
| P41252 | IARS | Isoleucyl-tRNA synthetase | Cytoplasm | Enzyme | −1.047 | |
| Q13907 | IDI1 | Isopentenyl-diphosphate delta isomerase 1 | Cytoplasm | Enzyme | −1.544 | |
| Q01628 | IFITM3 | Interferon induced transmembrane protein 3 | Plasma Membrane | Other | −1.052 | |
| P11717 | IGF2R | Insulin-like growth factor 2 receptor | Plasma Membrane | Transmembrane receptor | 1.187 | |
| Q12905 | ILF2 | Interleukin enhancer binding factor 2 | Nucleus | Transcription regulator | 1.264 | |
| Q12906 | ILF3 | Interleukin enhancer binding factor 3, 90 kDa | Nucleus | Transcription regulator | 1.246 | |
| Q16891 | IMMT | Inner membrane protein, mitochondrial | Cytoplasm | Other | 1.565 | |
| H0Y4R1 | IMPDH2 | IMP (inosine 5′-monophosphate) dehydrogenase 2 | Cytoplasm | Enzyme | −1.171 | |
| O00410 | IPO5 | Importin 5 | Nucleus | Transporter | 1.077 | |
| O95373 | IPO7 | Importin 7 | Nucleus | Transporter | 1.355 | |
| Q96P70 | IPO9 | Importin 9 | Nucleus | Transporter | −1.113 | |
| P46940 | IQGAP1 | IQ motif containing GTPase activating protein 1 | Cytoplasm | Other | 1.136 | |
| P05556 | ITGB1 | Integrin, β 1 (fibronectin receptor, β polypeptide, antigen CD29 includes MDF2, MSK12) | Plasma Membrane | Transmembrane receptor | −1.145 | |
| Q07666 | KHDRBS1 | KH domain containing, RNA binding, signal transduction associated 1 | Nucleus | Transcription regulator | −1.175 | |
| Q92945 | KHSRP | KH-type splicing regulatory protein | Nucleus | Enzyme | −1.284 | |
| P33176 | KIF5B | Kinesin family member 5B | Cytoplasm | Other | 1.332 | |
| P52292 | KPNA2 | Karyopherin α 2 (RAG cohort 1, importin α 1) | Nucleus | Transporter | 1.154 | |
| O00505 | KPNA3 | Karyopherin α 3 (importin α 4) | Nucleus | Transporter | 2.127 | |
| Q14974 | KPNB1 | Karyopherin (importin) β 1 | Nucleus | Transporter | 1.073 | |
| P04264 | KRT1 | Keratin 1 | Cytoplasm | Other | −5.784 | |
| P05783 | KRT18 | Keratin 18 | Cytoplasm | Other | −1.122 | |
| P05787 | KRT8 | Keratin 8 | Cytoplasm | Other | −1.469 | |
| P35527 | KRT9 | Keratin 9 | Other | Other | −4.460 | |
| P07942 | LAMB1 | Laminin, β 1 | Extracellular Space | Other | −1.088 | |
| C9J9W2 | LASP1 | LIM and SH3 protein 1 | Cytoplasm | Transporter | −1.654 | |
| P00338 | LDHA | Lactate dehydrogenase A | Cytoplasm | Enzyme | −1.361 | |
| P07195 | LDHB | Lactate dehydrogenase B | Cytoplasm | Enzyme | −1.428 | |
| P09382 | LGALS1 | Lectin, galactoside-binding, soluble, 1 | Extracellular Space | Other | −1.161 | |
| P02545 | LMNA | Lamin A/C | Nucleus | Other | 1.095 | |
| P20700 | LMNB1 | Lamin B1 | Nucleus | Other | −1.253 | |
| Q01081 | LOC102724594/U2AF1 | U2 small nuclear RNA auxiliary factor 1 | Nucleus | Other | 1.047 | |
| K7EJE8 | LONP1 | Lon peptidase 1, mitochondrial | Cytoplasm | Peptidase | −1.410 | |
| P42704 | LRPPRC | Leucine-rich pentatricopeptide repeat containing | Cytoplasm | Other | −1.059 | |
| Q96AG4 | LRRC59 | Leucine rich repeat containing 59 | Cytoplasm | Other | 1.228 | |
| P46821 | MAP1B | Microtubule-associated protein 1B | Cytoplasm | Other | 1.036 | |
| P27816 | MAP4 | Microtubule-associated protein 4 | Cytoplasm | Other | −1.561 | |
| P28482 | MAPK1 | Mitogen-activated protein kinase 1 | Cytoplasm | Kinase | −1.739 | |
| Q15691 | MAPRE1 | Microtubule-associated protein, RP/EB family, member 1 | Cytoplasm | Other | 1.418 | |
| P29966 | MARCKS | Myristoylated alanine-rich protein kinase C substrate | Plasma Membrane | Other | −2.498 | |
| P56192 | MARS | Methionyl-tRNA synthetase | Cytoplasm | Enzyme | 1.049 | |
| P43243 | MATR3 | Matrin 3 | Nucleus | Other | −1.307 | |
| P25205 | MCM3 | Minichromosome maintenance complex component 3 | Nucleus | Enzyme | −1.411 | |
| P33991 | MCM4 | Minichromosome maintenance complex component 4 | Nucleus | Enzyme | −1.472 | |
| Q14566 | MCM6 | Minichromosome maintenance complex component 6 | Nucleus | Enzyme | −1.238 | |
| P33993 | MCM7 | Minichromosome maintenance complex component 7 | Nucleus | Enzyme | −1.223 | |
| P40925 | MDH1 | Malate dehydrogenase 1, NAD (soluble) | Cytoplasm | Enzyme | −1.639 | |
| P40926 | MDH2 | Malate dehydrogenase 2, NAD (mitochondrial) | Cytoplasm | Enzyme | −1.298 | |
| P26038 | MSN | Moesin | Plasma Membrane | Other | 1.079 | |
| Q14764 | MVP | Major vault protein | Nucleus | Other | 1.636 | |
| P35580 | MYH10 | Myosin, heavy chain 10, non-muscle | Cytoplasm | Other | 1.275 | |
| P35579 | MYH9 | Myosin, heavy chain 9, non-muscle | Cytoplasm | Enzyme | −1.104 | |
| P19105 | MYL12A | Myosin, light chain 12A, regulatory, non-sarcomeric | Cytoplasm | Other | 1.316 | |
| F8W1N5 | NACA | Nascent polypeptide-associated complex α subunit | Cytoplasm | Transcription regulator | −1.282 | |
| F8VY35 | NAP1L1 | Nucleosome assembly protein 1-like 1 | Nucleus | Other | 1.173 | |
| Q99733 | NAP1L4 | Nucleosome assembly protein 1-like 4 | Cytoplasm | Other | −1.300 | |
| P49321 | NASP | Nuclear autoantigenic sperm protein (histone-binding) | Nucleus | Other | −1.640 | |
| P19338 | NCL | Nucleolin | Nucleus | Other | −1.037 | |
| Q32Q12 | NME1-NME2 | NME1-NME2 readthrough | Cytoplasm | Other | −1.298 | |
| P40261 | NNMT | Nicotinamide N-methyltransferase | Cytoplasm | Enzyme | −2.513 | |
| Q15233 | NONO | Non-POU domain containing, octamer-binding | Nucleus | Other | 1.374 | |
| E9PLK3 | NPEPPS | Aminopeptidase puromycin sensitive | Cytoplasm | Peptidase | −1.868 | |
| Q8TAT6 | NPLOC4 | Nuclear protein localization 4 homolog (S. cerevisiae) | Nucleus | Other | 1.135 | |
| P06748 | NPM1 | Nucleophosmin (nucleolar phosphoprotein B23, numatrin) | Nucleus | Transcription regulator | 1.166 | |
| Q9UNZ2 | NSFL1C | NSFL1 (p97) cofactor (p47) | Cytoplasm | Other | −1.139 | |
| Q14980 | NUMA1 | Nuclear mitotic apparatus protein 1 | Nucleus | Other | −1.202 | |
| Q9NTK5 | OLA1 | Obg-like ATPase 1 | Cytoplasm | Other | 1.159 | |
| Q96FW1 | OTUB1 | OTU deubiquitinase, ubiquitin aldehyde binding 1 | Cytoplasm | Enzyme | −1.304 | |
| P07237 | P4HB | Prolyl 4-hydroxylase, β polypeptide | Cytoplasm | Enzyme | −1.341 | |
| Q9UQ80 | PA2G4 | Proliferation-associated 2G4, 38kDa | Nucleus | Transcription regulator | −1.316 | |
| E7EQV3 | PABPC1 | Poly(A) binding protein, cytoplasmic 1 | Cytoplasm | Transcription regulator | 1.235 | |
| P68402 | PAFAH1B2 | Platelet-activating factor acetylhydrolase 1b, catalytic subunit 2 (30 kDa) | Cytoplasm | Enzyme | 1.089 | |
| E9PBS1 | PAICS | Phosphoribosylaminoimidazole carboxylase, phosphoribosylaminoimidazole succinocarboxamide synthetase | Cytoplasm | Enzyme | −1.354 | |
| Q99497 | PARK7 | Parkinson protein 7 | Nucleus | Enzyme | −1.711 | |
| Q15365 | PCBP1 | Poly(rC) binding protein 1 | Nucleus | Transcription regulator | −1.493 | |
| F8VZX2 | PCBP2 | Poly(rC) binding protein 2 | Nucleus | Other | 1.266 | |
| E7EV56 | PCM1 | Pericentriolar material 1 | Cytoplasm | Other | 1.175 | |
| B7Z972 | PCMT1 | Protein-l-isoaspartate (d-aspartate) O-methyltransferase | Cytoplasm | Enzyme | −1.351 | |
| P12004 | PCNA | Proliferating cell nuclear antigen | Nucleus | Enzyme | −1.321 | |
| E7EPW6 | PDCD6 | Programmed cell death 6 | Cytoplasm | Other | −1.147 | |
| F8VPF3 | PDE6H | Phosphodiesterase 6H, cGMP-specific, cone, γ | Cytoplasm | Enzyme | −1.043 | |
| P30101 | PDIA3 | Protein disulfide isomerase family A, member 3 | Cytoplasm | Peptidase | 1.167 | |
| P13667 | PDIA4 | Protein disulfide isomerase family A, member 4 | Cytoplasm | Enzyme | −1.155 | |
| Q15084 | PDIA6 | Protein disulfide isomerase family A, member 6 | Cytoplasm | Enzyme | −1.161 | |
| O00151 | PDLIM1 | PDZ and LIM domain 1 | Cytoplasm | Transcription regulator | −1.397 | |
| F2Z2Y4 | PDXK | Pyridoxal (pyridoxine, vitamin B6) kinase | Cytoplasm | Kinase | −1.008 | |
| Q15121 | PEA15 | Phosphoprotein enriched in astrocytes 15 | Cytoplasm | Transporter | 1.073 | |
| P30086 | PEBP1 | Phosphatidylethanolamine binding protein 1 | Cytoplasm | Other | −1.552 | |
| Q01813 | PFKP | Phosphofructokinase, platelet | Cytoplasm | Kinase | 1.313 | |
| P07737 | PFN1 | Profilin 1 | Cytoplasm | Other | −1.405 | |
| P18669 | PGAM1 | Phosphoglycerate mutase 1 (brain) | Cytoplasm | Phosphatase | −1.701 | |
| P52209 | PGD | Phosphogluconate dehydrogenase | Cytoplasm | Enzyme | −1.398 | |
| P00558 | PGK1 | Phosphoglycerate kinase 1 | Cytoplasm | Kinase | −1.287 | |
| P36871 | PGM1 | Phosphoglucomutase 1 | Cytoplasm | Enzyme | −1.393 | |
| F5GY37 | PHB2 | Prohibitin 2 | Cytoplasm | Transcription regulator | −1.175 | |
| O43175 | PHGDH | Phosphoglycerate dehydrogenase | Cytoplasm | Enzyme | −1.552 | |
| Q13492 | PICALM | Phosphatidylinositol binding clathrin assembly protein | Cytoplasm | Other | −1.195 | |
| P14618 | PKM | Pyruvate kinase, muscle | Cytoplasm | Kinase | −1.341 | |
| Q15149 | PLEC | Plectin | Cytoplasm | Other | 1.326 | |
| O60664 | PLIN3 | Perilipin 3 | Cytoplasm | Other | −1.084 | |
| E7ETU9 | PLOD2 | Procollagen-lysine, 2-oxoglutarate 5-dioxygenase 2 | Cytoplasm | Enzyme | 1.150 | |
| P13797 | PLS3 | Plastin 3 | Cytoplasm | Other | −1.143 | |
| Q15181 | PPA1 | Pyrophosphatase (inorganic) 1 | Cytoplasm | Enzyme | −1.263 | |
| P62937 | PPIA | Peptidylprolyl isomerase A (cyclophilin A) | Cytoplasm | Enzyme | −1.057 | |
| O15355 | PPM1G | Protein phosphatase, Mg2+/Mn2+ dependent, 1G | Nucleus | Phosphatase | 1.144 | |
| P62136 | PPP1CA | Protein phosphatase 1, catalytic subunit, α isozyme | Cytoplasm | Phosphatase | −1.250 | |
| P67775 | PPP2CA | Protein phosphatase 2, catalytic subunit, α isozyme | Cytoplasm | Phosphatase | 1.012 | |
| P30153 | PPP2R1A | Protein phosphatase 2, regulatory subunit A, α | Cytoplasm | Phosphatase | −1.191 | |
| Q06830 | PRDX1 | Peroxiredoxin 1 | Cytoplasm | Enzyme | −2.056 | |
| P32119 | PRDX2 | Peroxiredoxin 2 | Cytoplasm | Enzyme | −1.479 | |
| Q13162 | PRDX4 | Peroxiredoxin 4 | Cytoplasm | Enzyme | 1.346 | |
| P30044 | PRDX5 | Peroxiredoxin 5 | Cytoplasm | Enzyme | −1.297 | |
| K7ELL7 | PRKCSH | Protein kinase C substrate 80K–H | Cytoplasm | Enzyme | −1.483 | |
| E7EUY0 | PRKDC | Protein kinase, DNA-activated, catalytic polypeptide | Nucleus | Kinase | −1.313 | |
| H0YJX6 | PRMT5 | Protein arginine methyltransferase 5 | Cytoplasm | Enzyme | −1.975 | |
| Q9UMS4 | PRPF19 | Pre-mRNA processing factor 19 | Nucleus | Enzyme | −1.155 | |
| P25786 | PSMA1 | Proteasome (prosome, macropain) subunit, α type, 1 | Cytoplasm | Peptidase | −1.070 | |
| P25787 | PSMA2 | Proteasome (prosome, macropain) subunit, α type, 2 | Cytoplasm | Peptidase | −1.069 | |
| H0YLC2 | PSMA4 | Proteasome (prosome, macropain) subunit, α type, 4 | Cytoplasm | Peptidase | -1.204 | |
| P28066 | PSMA5 | Proteasome (prosome, macropain) subunit, α type, 5 | Cytoplasm | Peptidase | 1.018 | |
| O14818 | PSMA7 | Proteasome (prosome, macropain) subunit, α type, 7 | Cytoplasm | Peptidase | −1.157 | |
| P20618 | PSMB1 | Proteasome (prosome, macropain) subunit, β type, 1 | Cytoplasm | Peptidase | 1.208 | |
| P62191 | PSMC1 | Proteasome (prosome, macropain) 26S subunit, ATPase, 1 | Nucleus | Peptidase | −1.298 | |
| P43686 | PSMC4 | Proteasome (prosome, macropain) 26S subunit, ATPase, 4 | Nucleus | Peptidase | −1.249 | |
| Q13200 | PSMD2 | Proteasome (prosome, macropain) 26S subunit, non−ATPase, 2 | Cytoplasm | Other | 1.339 | |
| Q06323 | PSME1 | Proteasome (prosome, macropain) activator subunit 1 (PA28 α) | Cytoplasm | Other | −1.845 | |
| Q9UL46 | PSME2 | Proteasome (prosome, macropain) activator subunit 2 (PA28 β) | Cytoplasm | Peptidase | −1.010 | |
| P26599 | PTBP1 | Polypyrimidine tract binding protein 1 | Nucleus | Enzyme | 1.178 | |
| Q15185 | PTGES3 | Prostaglandin E synthase 3 (cytosolic) | Cytoplasm | Enzyme | −1.312 | |
| B8ZZQ6 | PTMA | Prothymosin, α | Nucleus | Other | −1.826 | |
| Q6NZI2 | PTRF | Polymerase I and transcript release factor | Nucleus | Transcription regulator | 2.165 | |
| Q9UHX1 | PUF60 | Poly-U binding splicing factor 60 kDa | Nucleus | Other | −1.146 | |
| P11216 | PYGB | Phosphorylase, glycogen; brain | Cytoplasm | Enzyme | −1.093 | |
| E9PK47 | PYGL | Phosphorylase, glycogen, liver | Cytoplasm | Enzyme | −1.252 | |
| P47897 | QARS | Glutaminyl-tRNA synthetase | Cytoplasm | Enzyme | 1.066 | |
| B4DQU5 | RAB11A | RAB11A, member RAS oncogene family | Cytoplasm | Enzyme | −1.365 | |
| E9PLD0 | RAB1B | RAB1B, member RAS oncogene family | Cytoplasm | Other | −1.182 | |
| P51149 | RAB7A | RAB7A, member RAS oncogene family | Cytoplasm | Enzyme | 1.158 | |
| P54727 | RAD23B | RAD23 homolog B (S. cerevisiae) | Nucleus | Other | 1.005 | |
| F5H018 | RAN | RAN, member RAS oncogene family | Nucleus | Enzyme | −1.229 | |
| C9JJ34 | RANBP1 | RAN binding protein 1 | Nucleus | Other | −1.461 | |
| P54136 | RARS | Arginyl-tRNA synthetase | Cytoplasm | Enzyme | −1.180 | |
| P98179 | RBM3 | RNA binding motif (RNP1, RRM) protein 3 | Cytoplasm | Other | 1.376 | |
| Q15293 | RCN1 | Reticulocalbin 1, EF-hand calcium binding domain | Cytoplasm | Other | 1.027 | |
| C9JNR4 | RHOA | Ras homolog family member A | Cytoplasm | Enzyme | −1.521 | |
| P13489 | RNH1 | Ribonuclease/angiogenin inhibitor 1 | Cytoplasm | Other | 1.064 | |
| X1WI28 | RPL10 | Ribosomal protein L10 | Cytoplasm | Other | 1.461 | |
| P62906 | RPL10A | Ribosomal protein L10a | Nucleus | Other | −1.075 | |
| P30050 | RPL12 | Ribosomal protein L12 | Nucleus | Other | 1.288 | |
| F8VUA6 | RPL18 | Ribosomal protein L18 | Cytoplasm | Other | −1.330 | |
| P46778 | RPL21 | Ribosomal protein L21 | Cytoplasm | Other | −1.204 | |
| P35268 | RPL22 | Ribosomal protein L22 | Nucleus | Other | 1.339 | |
| C9JD32 | RPL23 | Ribosomal protein L23 | Cytoplasm | Other | 1.026 | |
| P61353 | RPL27 | Ribosomal protein L27 | Cytoplasm | Other | 1.373 | |
| P39023 | RPL3 | Ribosomal protein L3 | Cytoplasm | Other | −1.206 | |
| P36578 | RPL4 | Ribosomal protein L4 | Cytoplasm | Enzyme | 1.044 | |
| P46777 | RPL5 | Ribosomal protein L5 | Cytoplasm | Other | −1.065 | |
| Q02878 | RPL6 | Ribosomal protein L6 | Cytoplasm | Other | 1.314 | |
| P18124 | RPL7 | Ribosomal protein L7 | Nucleus | Transcription regulator | 1.340 | |
| P62424 | RPL7A | Ribosomal protein L7a | Cytoplasm | Other | 1.141 | |
| G3V1A1 | RPL8 | Ribosomal protein L8 | Other | Other | −1.191 | |
| F8VWS0 | RPLP0 | Ribosomal protein, large, P0 | Cytoplasm | Other | −1.421 | |
| P05386 | RPLP1 | Ribosomal protein, large, P1 | Cytoplasm | Other | −1.274 | |
| P05387 | RPLP2 | Ribosomal protein, large, P2 | Cytoplasm | Other | −1.062 | |
| P04843 | RPN1 | Ribophorin I | Cytoplasm | Enzyme | 1.067 | |
| P04844 | RPN2 | Ribophorin II | Cytoplasm | Enzyme | −1.171 | |
| P46783 | RPS10 | Ribosomal protein S10 | Cytoplasm | Other | −1.009 | |
| P62280 | RPS11 | Ribosomal protein S11 | Cytoplasm | Other | 1.335 | |
| P25398 | RPS12 | Ribosomal protein S12 | Cytoplasm | Other | −1.062 | |
| K7EJ78 | RPS15 | Ribosomal protein S15 | Cytoplasm | Other | 1.152 | |
| H0YEN5 | RPS2 | Ribosomal protein S2 | Cytoplasm | Other | 1.092 | |
| E7ETK0 | RPS24 | Ribosomal protein S24 | Cytoplasm | Other | 1.157 | |
| P62979 | RPS27A | Ribosomal protein S27a | Cytoplasm | Other | 1.399 | |
| P23396 | RPS3 | Ribosomal protein S3 | Cytoplasm | Enzyme | 1.251 | |
| D6RG13 | RPS3A | Ribosomal protein S3A | Nucleus | Other | 1.184 | |
| P62701 | RPS4X | Ribosomal protein S4, X-linked | Cytoplasm | Other | 1.236 | |
| M0R0F0 | RPS5 | Ribosomal protein S5 | Cytoplasm | Other | −1.080 | |
| P62081 | RPS7 | Ribosomal protein S7 | Cytoplasm | Other | −1.189 | |
| P62241 | RPS8 | Ribosomal protein S8 | Cytoplasm | Other | 1.581 | |
| C9J9K3 | RPSA | Ribosomal protein SA | Cytoplasm | Transcription regulator | 1.267 | |
| Q9P2E9 | RRBP1 | Ribosome binding protein 1 | Cytoplasm | Other | 1.082 | |
| Q9NQC3 | RTN4 | Reticulon 4 | Cytoplasm | Other | −1.099 | |
| R4GN98 | S100A6 | S100 calcium binding protein A6 | Cytoplasm | Transporter | −1.190 | |
| M0QZS6 | SAE1 | SUMO1 activating enzyme subunit 1 | Cytoplasm | Enzyme | 1.058 | |
| D6RFM5 | SDHA | uccinate dehydrogenase complex, subunit A, flavoprotein (Fp) | Cytoplasm | Enzyme | 1.035 | |
| P55735 | SEC13 | SEC13 homolog (S. cerevisiae) | Cytoplasm | Transporter | 1.029 | |
| Q15019 | SEPT2 | Septin 2 | Cytoplasm | Enzyme | −1.140 | |
| H0Y3Y4 | SEPT7 | Septin 7 | Cytoplasm | Other | 1.266 | |
| Q9UHD8 | SEPT9 | Septin 9 | Cytoplasm | Enzyme | 1.063 | |
| Q8NC51 | SERBP1 | SERPINE1 mRNA binding protein 1 | Cytoplasm | Other | 1.003 | |
| P50454 | SERPINH1 | Serpin peptidase inhibitor, clade H (heat shock protein 47), member 1, (collagen binding protein 1) | Extracellular Space | Other | −1.178 | |
| Q01105 | SET | SET nuclear proto-oncogene | Nucleus | Phosphatase | −1.282 | |
| Q15459 | SF3A1 | Splicing factor 3a, subunit 1, 120 kDa | Nucleus | Other | 1.024 | |
| O75533 | SF3B1 | Splicing factor 3b, subunit 1, 155 kDa | Nucleus | Other | 1.359 | |
| P31947 | SFN | Stratifin | Cytoplasm | Other | −1.586 | |
| P23246 | SFPQ | Splicing factor proline/glutamine-rich | Nucleus | Other | 1.009 | |
| Q9H299 | SH3BGRL3 | SH3 domain binding glutamate-rich protein like 3 | Nucleus | Other | 1.688 | |
| P34897 | SHMT2 | Serine hydroxymethyltransferase 2 (mitochondrial) | Cytoplasm | Enzyme | −1.297 | |
| F8VVM2 | SLC25A3 | Solute carrier family 25 (mitochondrial carrier; phosphate carrier), member 3 | Cytoplasm | Transporter | 1.165 | |
| P05141 | SLC25A5 | Solute carrier family 25 (mitochondrial carrier; adenine nucleotide translocator), member 5 | Cytoplasm | Transporter | −1.559 | |
| I7HJJ0 | SLC25A6 | Solute carrier family 25 (mitochondrial carrier; adenine nucleotide translocator), member 6 | Cytoplasm | Transporter | 1.304 | |
| Q96QD8 | SLC38A2 | Solute carrier family 38, member 2 | Plasma Membrane | Transporter | −1.082 | |
| Q7KZF4 | SND1 | Staphylococcal nuclease and tudor domain containing 1 | Nucleus | Enzyme | −1.112 | |
| P62314 | SNRPD1 | Small nuclear ribonucleoprotein D1 polypeptide 16 kDa | Nucleus | Other | −1.174 | |
| H3BT13 | SNRPD3 | Small nuclear ribonucleoprotein D3 polypeptide 18 kDa | Nucleus | Other | 1.288 | |
| O60493 | SNX3 | Sorting nexin 3 | Cytoplasm | Transporter | 1.117 | |
| P00441 | SOD1 | Superoxide dismutase 1, soluble | Cytoplasm | Enzyme | −1.264 | |
| Q13813 | SPTAN1 | Spectrin, α, non-erythrocytic 1 | Plasma Membrane | Other | 1.056 | |
| Q01082 | SPTBN1 | Spectrin, β, non-erythrocytic 1 | Plasma Membrane | Other | −1.545 | |
| E7EMC7 | SQSTM1 | Sequestosome 1 | Cytoplasm | Transcription regulator | 1.310 | |
| Q08945 | SSRP1 | Structure specific recognition protein 1 | Nucleus | Other | −1.498 | |
| P42224 | STAT1 | Signal transducer and activator of transcription 1, 91 kDa | Nucleus | Transcription regulator | −1.353 | |
| P31948 | STIP1 | Stress-induced phosphoprotein 1 | Cytoplasm | Other | −1.086 | |
| A2A2D0 | STMN1 | Stathmin 1 | Cytoplasm | Other | −1.445 | |
| Q9UJZ1 | STOML2 | Stomatin (EPB72)-like 2 | Plasma Membrane | Other | 1.325 | |
| Q9Y5B9 | SUPT16H | Suppressor of Ty 16 homolog (S. cerevisiae) | Nucleus | Transcription regulator | −1.137 | |
| Q5T8U5 | SURF4 | Surfeit 4 | Cytoplasm | Other | 1.327 | |
| O60506 | SYNCRIP | Synaptotagmin binding, cytoplasmic RNA interacting protein | Nucleus | Other | −1.378 | |
| Q01995 | TAGLN | Transgelin | Cytoplasm | Other | −1.657 | |
| P37802 | TAGLN2 | Transgelin 2 | Cytoplasm | Other | −1.382 | |
| P37837 | TALDO1 | Transaldolase 1 | Cytoplasm | Enzyme | −1.652 | |
| P26639 | TARS | Threonyl-tRNA synthetase | Nucleus | Enzyme | −1.194 | |
| E5RIW3 | TBCA | Tubulin folding cofactor A | Cytoplasm | Other | −1.404 | |
| P17987 | TCP1 | T-complex 1 | Cytoplasm | Other | 1.190 | |
| P02786 | TFRC | Transferrin receptor | Plasma Membrane | Transporter | −1.018 | |
| B4E022 | TKT | Transketolase | Cytoplasm | Enzyme | −1.717 | |
| Q9Y490 | TLN1 | Talin 1 | Plasma Membrane | Other | −1.072 | |
| Q9NYL9 | TMOD3 | Tropomodulin 3 (ubiquitous) | Cytoplasm | Other | 1.076 | |
| P42166 | TMPO | Thymopoietin | Nucleus | Other | −1.488 | |
| F5H7V9 | TNC | Tenascin C | Extracellular Space | Other | −2.480 | |
| Q92973 | TNPO1 | Transportin 1 | Nucleus | Transporter | −1.378 | |
| P60174 | TPI1 | Triosephosphate isomerase 1 | Cytoplasm | Enzyme | −1.620 | |
| Q5TCU3 | TPM2 | Tropomyosin 2 (β) | Other | Other | 1.845 | |
| J3KN67 | TPM3 | Tropomyosin 3 | Cytoplasm | Other | −1.265 | |
| P67936 | TPM4 | Tropomyosin 4 | Cytoplasm | Other | −1.185 | |
| O14773 | TPP1 | Tripeptidyl peptidase I | Cytoplasm | Peptidase | −1.239 | |
| Q13263 | TRIM28 | Tripartite motif containing 28 | Nucleus | Transcription regulator | −1.201 | |
| F8VQE1 | TRMT1 | tRNA methyltransferase 1 homolog (S. cerevisiae) | Extracellular Space | Enzyme | −1.271 | |
| P68363 | TUBA1B | Tubulin, α 1b | Cytoplasm | Other | −1.056 | |
| Q5JP53 | TUBB | Tubulin, β class I | Cytoplasm | Other | −1.029 | |
| P68371 | TUBB4B | Tubulin, β 4B class IVb | Cytoplasm | Other | 1.046 | |
| Q9BUF5 | TUBB6 | Tubulin, β 6 class V | Cytoplasm | Other | 1.022 | |
| Q3ZCM7 | TUBB8 | Tubulin, β 8 class VIII | Cytoplasm | Other | −1.128 | |
| P49411 | TUFM | Tu translation elongation factor, mitochondrial | Cytoplasm | Transcription regulator | −1.173 | |
| Q12792 | TWF1 | Twinfilin actin-binding protein 1 | Cytoplasm | Kinase | −1.798 | |
| D6RG15 | TWF2 | Twinfilin actin-binding protein 2 | Cytoplasm | Kinase | −1.347 | |
| P40222 | TXLNA | Taxilin α | Extracellular Space | Cytokine | −1.162 | |
| Q86UY0 | TXNDC5 | Thioredoxin domain containing 5 (endoplasmic reticulum) | Cytoplasm | Enzyme | 1.390 | |
| P26368 | U2AF2 | U2 small nuclear RNA auxiliary factor 2 | Nucleus | Other | −1.209 | |
| Q16222 | UAP1 | UDP-N-acetylglucosamine pyrophosphorylase 1 | Nucleus | Enzyme | 1.145 | |
| P22314 | UBA1 | Ubiquitin-like modifier activating enzyme 1 | Cytoplasm | Enzyme | 1.182 | |
| P68036 | UBE2L3 | Ubiquitin-conjugating enzyme E2L 3 | Nucleus | Enzyme | 1.021 | |
| F8VZ29 | UBE2N | Ubiquitin-conjugating enzyme E2N | Cytoplasm | Enzyme | −1.650 | |
| Q92575 | UBXN4 | UBX domain protein 4 | Extracellular Space | Other | −1.082 | |
| P09936 | UCHL1 | Ubiquitin carboxyl-terminal esterase L1 (ubiquitin thiolesterase) | Cytoplasm | Peptidase | −1.453 | |
| P31930 | UQCRC1 | Ubiquinol-cytochrome c reductase core protein I | Cytoplasm | Enzyme | 1.218 | |
| Q9P0L0 | VAPA | VAMP (vesicle-associated membrane protein)-associated protein A, 33 kDa | Plasma Membrane | Other | 1.026 | |
| P26640 | VARS | Valyl-tRNA synthetase | Cytoplasm | Enzyme | −1.120 | |
| K7ERT7 | VAT1 | Vesicle amine transport 1 | Plasma Membrane | Transporter | −2.302 | |
| P18206 | VCL | Vinculin | Plasma Membrane | Enzyme | −1.175 | |
| P55072 | VCP | Valosin containing protein | Cytoplasm | Enzyme | −1.058 | |
| P21796 | VDAC1 | Voltage-dependent anion channel 1 | Cytoplasm | Ion channel | −1.335 | |
| P45880 | VDAC2 | Voltage-dependent anion channel 2 | Cytoplasm | Ion channel | −1.041 | |
| P08670 | VIM | Vimentin | Cytoplasm | Other | 1.579 | |
| Q96QK1 | VPS35 | Vacuolar protein sorting 35 homolog (S. cerevisiae) | Cytoplasm | Transporter | −1.641 | |
| P23381 | WARS | Tryptophanyl-tRNA synthetase | Cytoplasm | Enzyme | −1.477 | |
| O75083 | WDR1 | WD repeat domain 1 | Extracellular Space | Other | −1.206 | |
| O14980 | XPO1 | exportin 1 | Nucleus | Transporter | −1.091 | |
| P13010 | XRCC5 | X-ray repair complementing defective repair in Chinese hamster cells 5 (double-strand-break rejoining) | Nucleus | Enzyme | −1.197 | |
| P12956 | XRCC6 | X-ray repair complementing defective repair in Chinese hamster cells 6 | Nucleus | Enzyme | −1.575 | |
| P67809 | YBX1 | Y box binding protein 1 | Nucleus | Transcription regulator | −1.196 | |
| P16989 | YBX3 | Y box binding protein 3 | Nucleus | Transcription regulator | −1.186 | |
| P31946 | YWHAB | Tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, β | Cytoplasm | Transcription regulator | 1.064 | |
| P62258 | YWHAE | Tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, epsilon | Cytoplasm | Other | −1.216 | |
| P61981 | YWHAG | Tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, γ | Cytoplasm | Other | −1.045 | |
| Q04917 | YWHAH | Tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, η | Cytoplasm | Transcription regulator | −1.559 | |
| P27348 | YWHAQ | Tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, θ | Cytoplasm | Other | 1.180 | |
| P63104 | YWHAZ | Tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, ζ | Cytoplasm | Enzyme | −1.123 | |
| Q15942 | ZYX | Zyxin | Plasma Membrane | Other | −1.100 |
| INGENUITY Canonical Pathways | −log(p-Value) a | Ratio b | z-Score c | Molecules |
|---|---|---|---|---|
| eIF2 signaling | 29.5 | 2.32 × 10−1 | −0.229 | RPL22, EIF3C, RPLP1, MAPK1, RPS3A, RPLP2, RPS8, EIF2S1, EIF4G1, RPL7, RPS11, RPS4X, RPS7, RPL6, RPL7A, EIF3B, PPP1CA, RPS3, RPS5, RPL18, RPS24, PABPC1, RPL4, RPL3, RPS2, RPL27, RPS10, RPL21, RPL23, RPL12, RPLP0, RPL10A, EIF3M, RPS12, EIF2S2, RPL8, EIF4A1, EIF3I, RPS15, RPL10, RPS27A, RPL5, RPSA |
| Remodeling of epithelial adherent junctions | 16.2 | 2.94 × 10−1 | 0.000 | TUBA1B, ACTR2, ARPC1B, TUBB4B, MAPRE1, RAB7A, CTNNA1, IQGAP1, TUBB, ACTR3, TUBB6, TUBB8, ARPC2, ZYX, VCL, ACTN4, DNM1L, ACTG1, ACTN1, ACTA1 |
| Regulation of eIF4 and p70S6K signaling | 16.0 | 1.85 × 10−1 | −1.000 | EIF3C, RPS3A, MAPK1, PPP2CA, RPS8, EIF4G1, EIF2S1, RPS4X, RPS11, RPS7, EIF3B, RPS3, RPS5, RPS24, ITGB1, PABPC1, RPS2, RPS10, RPS12, EIF3M, EIF2S2, PPP2R1A, EIF4A1, EIF3I, RPS15, RPS27A, RPSA |
| mTOR signaling | 11.4 | 1.33 × 10−1 | −0.447 | EIF3C, RPS3A, MAPK1, PPP2CA, RPS8, FKBP1A, EIF4G1, RPS4X, RPS11, RPS7, EIF3B, RPS3, RPS5, RPS24, RPS2, RPS10, RPS12, EIF3M, PPP2R1A, RHOA, EIF4A1, RPS15, EIF3I, RPS27A, RPSA |
| Protein ubiquitination pathway | 11.4 | 1.14 × 10−1 | B2M, PSMA7, HLA-A, UBE2N, HLA-B, HSPA5, DNAJA1, UCHL1, HSPA4, HSP90B1, HSP90AB1, DNAJC8, PSMA2, HSPA9, PSME2, PSMC4, PSMA1, HSPD1, HSPA8, PSME1, PSMC1, UBE2L3, PSMD2, PSMA5, PSMA4, PSMB1, HSP90AA1, UBA1, HSPB1 | |
| Glycolysis I | 11.4 | 4.40 × 10−1 | PGK1, ENO1, GPI, TPI1, PGAM1, PKM, ENO2, ALDOA, GAPDH, PFKP, ALDOC | |
| Caveolar-mediated endocytosis signaling | 11.0 | 2.22 × 10−1 | ITGB1, B2M, FLNB, HLA-A, HLA-B, COPA, COPE, COPB2, COPB1, COPG1, FLNA, FLNC, CAV1, PTRF, ACTG1, ACTA1 | |
| Ran signaling | 10.3 | 5.29 × 10−1 | KPNB1, KPNA3, CSE1L, KPNA2, TNPO1, RAN, XPO1, RANBP1, IPO5 | |
| Gluconeogenesis I | 9.88 | 4.00 × 10−1 | PGK1, ENO1, GPI, PGAM1, ENO2, ALDOA, GAPDH, MDH1, MDH2, ALDOC | |
| Epithelial adherent junction signaling | 9.51 | 1.37 × 10−1 | TUBA1B, MYH10, ACTR2, MYH9, ARPC1B, TUBB4B, CTNNA1, IQGAP1, TUBB, ACTR3, TUBB6, TUBB8, ARPC2, RHOA, ZYX, VCL, ACTN4, ACTG1, ACTN1, ACTA1 | |
| tRNA charging | 8.93 | 2.82 × 10−1 | WARS, RARS, GARS, HARS, TARS, AARS, VARS, MARS, IARS, EPRS, QARS | |
| Actin cytoskeleton signaling | 8.59 | 1.06 × 10−1 | ITGB1, ACTR2, MYH10, MYH9, FN1, PFN1, CFL1, MAPK1, ARPC1B, TLN1, IQGAP1, ACTR3, FLNA, EZR, ARPC2, RHOA, VCL, ACTN4, ACTG1, ACTA1, ACTN1, MSN, MYL12A | |
| Regulation of cellular mechanics by calpain protease | 8.10 | 2.11 × 10−1 | 0.378 | ITGB1, CAPNS1, MAPK1, EZR, CAPN1, TLN1, CAPN2, VCL, CAST, ACTN4, CDK1, ACTN1 |
| Integrin signaling | 7.74 | 1.04 × 10−1 | −0.894 | ITGB1, ACTR2, ARPC1B, MAPK1, ARF1, TLN1, ACTR3, CAPNS1, CAPN1, RHOA, ARPC2, CAV1, ZYX, CAPN2, VCL, ACTN4, CTTN, ACTG1, ACTN1, ACTA1, MYL12A |
| Virus entry via endocytic pathways | 7.62 | 1.57 × 10−1 | B2M, ITGB1, FLNB, FLNC, FLNA, HLA-A, AP2B1, HLA-B, CLTC, CAV1, TFRC, AP1B1, ACTG1, ACTA1 | |
| Unfolded protein response | 7.31 | 2.04 × 10−1 | HSPA8, HSPA4, CALR, P4HB, HSP90B1, UBXN4, HSPA9, VCP, CANX, DNAJA2, HSPA5 | |
| ILK signaling | 6.94 | 1.02 × 10−1 | −1.414 | ITGB1, FLNB, MYH10, MYH9, FN1, CFL1, MAPK1, PPP2CA, VIM, PPP2R1A, FLNA, FLNC, RHOA, KRT18, ACTN4, ACTG1, ACTN1, ACTA1, NACA |
| 14-3-3-mediated signaling | 6.89 | 1.28 × 10−1 | TUBA1B, YWHAG, YWHAH, YWHAE, MAPK1, TUBB4B, PDIA3, YWHAB, YWHAZ, VIM, TUBB, YWHAQ, TUBB6, TUBB8, SFN | |
| Rhogdi signaling | 6.73 | 1.04 × 10−1 | −1.000 | ITGB1, GDI1, ACTR2, CFL1, ARPC1B, GNB2L1, GDI2, GNAI2, ACTR3, RHOA, ARPC2, EZR, CD44, ARHGDIA, ACTG1, ACTA1, MSN, MYL12A |
| RhoA signaling | 6.65 | 1.23 × 10−1 | −0.258 | ACTR2, PFN1, SEPT9, ARPC1B, CFL1, SEPT7, ACTR3, ARPC2, EZR, RHOA, ACTG1, ACTA1, SEPT2, MSN, MYL12A |
| Signaling by Rho family GTPases | 6.64 | 8.97 × 10−2 | 0.471 | ITGB1, ACTR2, CFL1, MAPK1, SEPT9, ARPC1B, SEPT7, GNB2L1, VIM, IQGAP1, GNAI2, STMN1, ACTR3, EZR, ARPC2, RHOA, ACTG1, ACTA1, SEPT2, MSN, MYL12A |
| PI3K/Akt signaling | 5.84 | 1.14 × 10−1 | −0.535 | YWHAQ, ITGB1, PPP2R1A, HSP90B1, YWHAG, HSP90AB1, YWHAH, MAPK1, YWHAE, YWHAB, PPP2CA, YWHAZ, HSP90AA1, SFN |
| Germ cell-sertoli cell junction signaling | 5.82 | 1.00 × 10−1 | ITGB1, TUBA1B, MAPK1, CFL1, TUBB4B, CTNNA1, TUBB, IQGAP1, TUBB6, TUBB8, RHOA, ZYX, ACTN4, ACTG1, ACTN1, ACTA1 | |
| Cell cycle: G2/M DNA damage checkpoint regulation | 5.69 | 1.84 × 10−1 | −0.447 | YWHAQ, PRKDC, YWHAG, YWHAE, YWHAH, YWHAB, YWHAZ, SFN, CDK1 |
| Clathrin-mediated endocytosis signaling | 5.63 | 9.19 × 10−2 | ITGB1, ACTR2, ARPC1B, PICALM, AP2B1, CLTC, RAB7A, AP1B1, HSPA8, ACTR3, ARPC2, RAB11A, TFRC, DNM1L, CTTN, ACTG1, ACTA1 | |
| Glutaryl-CoA degradation | 5.51 | 4.55 × 10−1 | HSD17B10, ACAT2, ACAT1, HADHA, HADH | |
| p70S6K signaling | 5.26 | 1.09 × 10−1 | 0.000 | GNAI2, YWHAQ, PPP2R1A, YWHAG, YWHAH, MAPK1, YWHAE, EEF2, YWHAB, PDIA3, PPP2CA, YWHAZ, SFN |
| Hippo signaling | 5.20 | 1.28 × 10−1 | 0.632 | YWHAQ, PPP2R1A, YWHAG, YWHAE, YWHAH, PPP2CA, YWHAB, CD44, YWHAZ, SFN, PPP1CA |
| Regulation of actin-based motility by rho | 4.96 | 1.21 × 10−1 | −1.265 | ITGB1, ACTR2, PFN1, ACTR3, ARPC1B, CFL1, ARPC2, RHOA, ARHGDIA, ACTA1, MYL12A |
| Isoleucine degradation I | 4.89 | 3.57 × 10−1 | HSD17B10, ECHS1, ACAT2, ACAT1, HADHA | |
| Aldosterone signaling in epithelial cells | 4.76 | 9.21 × 10−2 | MAPK1, PDIA3, HSPA9, HSPD1, DNAJA1, HSPA5, HSPA8, HSPA4, HSP90B1, HSP90AB1, DNAJC8, HSP90AA1, AHCY, HSPB1 | |
| Protein kinase a signaling | 4.67 | 6.22 × 10−2 | FLNB, HIST1H1C, MYH10, YWHAG, YWHAH, MAPK1, YWHAE, YWHAB, PDIA3, GNB2L1, YWHAZ, PYGL, PYGB, PDE6H, GNAI2, YWHAQ, H3F3A/H3F3B, FLNA, FLNC, RHOA, SFN, PPP1CA, APEX1, MYL12A | |
| Sertoli cell-sertoli cell junction signaling | 4.59 | 8.43 × 10−2 | ITGB1, SPTBN1, TUBA1B, MAPK1, TUBB4B, CTNNA1, YBX3, TUBB, TUBB6, TUBB8, ACTN4, SPTAN1, ACTG1, ACTN1, ACTA1 | |
| Lipid antigen presentation by CD1 | 4.56 | 2.31 × 10−1 | B2M, CALR, PDIA3, AP2B1, CANX, AP1B1 | |
| Superpathway of geranylgeranyldiphosphate biosynthesis I (via mevalonate) | 4.43 | 2.94 × 10−1 | FDPS, ACAT2, IDI1, ACAT1, HADHA | |
| VEGF signaling | 4.20 | 1.10 × 10−1 | 0.000 | EIF2S2, YWHAE, MAPK1, VCL, ACTN4, EIF2S1, SFN, ACTG1, ACTA1, ACTN1 |
| Fcγ receptor-mediated phagocytosis in macrophages and monocytes | 4.12 | 1.08 × 10−1 | −1.265 | ACTR2, ACTR3, ARPC1B, MAPK1, ARPC2, EZR, RAB11A, TLN1, ACTG1, ACTA1 |
| Tryptophan degradation III (eukaryotic) | 4.06 | 2.50 × 10−1 | HSD17B10, ACAT2, ACAT1, HADHA, HADH | |
| Pentose phosphate pathway | 4.03 | 3.64 × 10−1 | PGD, TKT, TALDO1, G6PD | |
| Nrf2-mediated oxidative stress response | 3.95 | 7.78 × 10−2 | −1.890 | SOD1, MAPK1, PRDX1, DNAJA1, ERP29, DNAJC8, STIP1, VCP, CCT7, DNAJA2, SQSTM1, ACTG1, ACTA1, GSTP1 |
| Purine nucleotides de novo biosynthesis II | 3.86 | 3.33 × 10−1 | GMPS, IMPDH2, PAICS, ATIC | |
| Huntington’s disease signaling | 3.86 | 6.96 × 10−2 | SDHA, MAPK1, GNB2L1, HSPA9, CLTC, PSME2, HSPA5, HSPA8, HSPA4, DYNC1I2, PSME1, CAPNS1, ATP5B, CAPN1, CAPN2, DNM1L | |
| Erk/MAPK signaling | 3.77 | 7.49 × 10−2 | −1.000 | ITGB1, YWHAG, YWHAH, MAPK1, PPP2CA, YWHAB, YWHAZ, TLN1, YWHAQ, PPP2R1A, H3F3A/H3F3B, PPP1CA, STAT1, HSPB1 |
| Rac signaling | 3.71 | 9.62 × 10−2 | −0.333 | ITGB1, ACTR2, ACTR3, ARPC1B, MAPK1, CFL1, ARPC2, RHOA, CD44, IQGAP1 |
| Mevalonate pathway I | 3.71 | 3.08 × 10−1 | ACAT2, IDI1, ACAT1, HADHA | |
| Breast cancer regulation by stathmin1 | 3.68 | 7.33 × 10−2 | TUBA1B, MAPK1, PPP2CA, TUBB4B, GNB2L1, TUBB, CDK1, GNAI2, STMN1, PPP2R1A, TUBB6, TUBB8, RHOA, PPP1CA | |
| Antigen presentation pathway | 3.65 | 1.62 × 10−1 | B2M, CALR, HLA-A, PDIA3, HLA-B, CANX | |
| FAK signaling | 3.64 | 1.03 × 10−1 | ITGB1, CAPNS1, MAPK1, CAPN1, TLN1, CAPN2, VCL, ACTG1, ACTA1 | |
| Leukocyte extravasation signaling | 3.52 | 7.07 × 10−2 | 0.000 | ITGB1, MAPK1, CTNNA1, GNAI2, EZR, RHOA, CD44, ACTN4, VCL, CTTN, ACTG1, ACTN1, ACTA1, MSN |
| Telomere extension by telomerase | 3.45 | 2.67 × 10−1 | HNRNPA1, XRCC6, HNRNPA2B1, XRCC5 | |
| Gap junction signaling | 3.45 | 7.74 × 10−2 | GNAI2, DBN1, TUBA1B, TUBB6, MAPK1, TUBB8, TUBB4B, PDIA3, CAV1, TUBB, ACTG1, ACTA1 | |
| Myc mediated apoptosis signaling | 3.35 | 1.21 × 10−1 | YWHAQ, YWHAG, YWHAE, YWHAH, YWHAB, YWHAZ, SFN | |
| Superpathway of cholesterol biosynthesis | 3.32 | 1.79 × 10−1 | FDPS, ACAT2, IDI1, ACAT1, HADHA | |
| IGF-1 signaling | 3.28 | 9.28 × 10−2 | YWHAQ, YWHAG, YWHAE, MAPK1, YWHAH, YWHAB, YWHAZ, SFN, CYR61 | |
| Spliceosomal cycle | 3.24 | 1.00 | LOC102724594/U2AF1, U2AF2 | |
| Tight junction signaling | 3.16 | 7.19 × 10−2 | MYH10, PPP2R1A, MYH9, PPP2CA, RHOA, VAPA, CTNNA1, YBX3, SPTAN1, VCL, ACTG1, ACTA1 | |
| Cdc42 signaling | 3.16 | 7.19 × 10−2 | −0.707 | ITGB1, B2M, ACTR2, ACTR3, ARPC1B, MAPK1, CFL1, HLA-A, ARPC2, HLA-B, IQGAP1, MYL12A |
| Erk5 signaling | 3.13 | 1.11 × 10−1 | YWHAQ, YWHAG, YWHAE, YWHAH, YWHAB, YWHAZ, SFN | |
| Paxillin signaling | 3.12 | 8.82 × 10−2 | −1.667 | ITGB1, MAPK1, ARF1, TLN1, VCL, ACTN4, ACTG1, ACTA1, ACTN1 |
| Mitochondrial dysfunction | 3.07 | 7.02 × 10−2 | HSD17B10, SDHA, ATP5H, ATP5B, PARK7, PRDX5, ATP5A1, CYC1, VDAC1, UQCRC1, CYB5R3, VDAC2 | |
| Hypoxia signaling in the cardiovascular system | 3.05 | 1.08 × 10−1 | P4HB, HSP90B1, UBE2L3, HSP90AB1, UBE2N, HSP90AA1, LDHA | |
| Mitotic roles of polo-like kinase | 3.01 | 1.06 × 10−1 | HSP90B1, PPP2R1A, HSP90AB1, PPP2CA, CAPN1, HSP90AA1, CDK1 | |
| Sucrose degradation V (mammalian) | 2.99 | 3.33 × 10−1 | TPI1, ALDOA, ALDOC | |
| Ketolysis | 2.99 | 3.33 × 10−1 | ACAT2, ACAT1, HADHA | |
| CTLA4 signaling in cytotoxic T lymphocytes | 2.92 | 9.09 × 10−2 | B2M, PPP2R1A, PPP2CA, HLA-A, AP2B1, CLTC, HLA-B, AP1B1 | |
| Endoplasmic reticulum stress pathway | 2.86 | 1.90 × 10−1 | CALR, HSP90B1, EIF2S1, HSPA5 | |
| Ketogenesis | 2.84 | 3.00 × 10−1 | ACAT2, ACAT1, HADHA | |
| Glycogen degradation II | 2.84 | 3.00 × 10−1 | PGM1, PYGB, PYGL | |
| Inosine-5′-phosphate biosynthesis II | 2.77 | 6.67 × 10−1 | PAICS, ATIC | |
| Ephrin b signaling | 2.75 | 9.59 × 10−2 | −0.447 | GNAI2, MAPK1, CFL1, RHOA, GNB2L1, CAP1, HNRNPK |
| TCA cycle II (eukaryotic) | 2.70 | 1.74 × 10−1 | SDHA, CS, MDH1, MDH2 | |
| Actin nucleation by ARP-WASP complex | 2.67 | 1.07 × 10−1 | −0.816 | ITGB1, ACTR2, ACTR3, ARPC1B, ARPC2, RHOA |
| Glycogen degradation III | 2.60 | 2.50 × 10−1 | PGM1, PYGB, PYGL | |
| Telomerase signaling | 2.59 | 8.08 × 10−2 | 0.000 | HSP90B1, PPP2R1A, MAPK1, HSP90AB1, PPP2CA, HSP90AA1, TPP1, PTGES3 |
| Cell cycle control of chromosomal replication | 2.44 | 1.48 × 10−1 | MCM3, MCM6, MCM4, MCM7 | |
| Axonal guidance signaling | 2.40 | 4.62 × 10−2 | DPYSL2, ITGB1, TUBA1B, ACTR2, PFN1, CFL1, MAPK1, ARPC1B, TUBB4B, PDIA3, GNB2L1, TUBB, GNAI2, ACTR3, TUBB6, TUBB8, ARPC2, RHOA, RTN4, MYL12A | |
| DNA double-strand break repair by non-homologous end joining | 2.39 | 2.14 × 10−1 | PRKDC, XRCC6, XRCC5 | |
| Glucocorticoid receptor signaling | 2.37 | 5.36 × 10−2 | MAPK1, YWHAH, HSPA9, HSPA5, PTGES3, HSPA8, HMGB1, HSPA4, HSP90B1, HSP90AB1, ANXA1, FKBP4, HSP90AA1, STAT1 | |
| Fatty acid β-oxidation I | 2.27 | 1.33 × 10−1 | HSD17B10, ECHS1, HADHA, HADH | |
| Pentose phosphate pathway (oxidative branch) | 2.27 | 4.00 × 10−1 | PGD, G6PD | |
| Trans, trans-farnesyl diphosphate biosynthesis | 2.27 | 4.00 × 10−1 | FDPS, IDI1 | |
| Apoptosis signaling | 2.26 | 7.87 × 10−2 | 1.134 | CAPNS1, MAPK1, CAPN1, LMNA, CAPN2, SPTAN1, CDK1 |
| Granzyme B signaling | 2.22 | 1.88 × 10−1 | PRKDC, NUMA1, LMNB1 | |
| Parkinson’s signaling | 2.22 | 1.88 × 10−1 | UCHL1, MAPK1, PARK7 | |
| Agrin interactions at neuromuscular junction | 2.22 | 8.70 × 10−2 | −1.342 | ITGB1, MAPK1, LAMB1, CTTN, ACTG1, ACTA1 |
| Aryl hydrocarbon receptor signaling | 2.18 | 6.43 × 10−2 | −0.378 | HSP90B1, MAPK1, HSP90AB1, HSP90AA1, GSTP1, PTGES3, HSPB1, MCM7, AIP |
| Pyruvate fermentation to lactate | 2.10 | 3.33 × 10−1 | LDHA, LDHB | |
| Pentose phosphate pathway (non-oxidative branch) | 2.10 | 3.33 × 10−1 | TKT, TALDO1 | |
| Semaphorin signaling in neurons | 2.07 | 9.43 × 10−2 | ITGB1, DPYSL2, MAPK1, CFL1, RHOA | |
| Ephrin receptor signaling | 2.04 | 5.75 × 10−2 | ITGB1, GNAI2, ACTR2, ACTR3, ARPC1B, MAPK1, CFL1, ARPC2, RHOA, GNB2L1 | |
| CDK5 signaling | 2.02 | 7.07 × 10−2 | 0.000 | ITGB1, PPP2R1A, MAPK1, PPP2CA, CAPN1, LAMB1, PPP1CA |
| Superpathway of serine and glycine biosynthesisI | 1.96 | 2.86 × 10−1 | PHGDH, SHMT2 | |
| Aspartate degradation II | 1.96 | 2.86 × 10−1 | MDH1, MDH2 | |
| Granzyme A signaling | 1.94 | 1.50 × 10−1 | HIST1H1C, SET, APEX1 | |
| Prostate cancer signaling | 1.86 | 7.32 × 10−2 | HSP90B1, MAPK1, PA2G4, HSP90AB1, HSP90AA1, GSTP1 | |
| fMLP signaling in neutrophils | 1.82 | 6.48 × 10−2 | 0.000 | GNAI2, ACTR2, ACTR3, ARPC1B, MAPK1, ARPC2, GNB2L1 |
| Mechanisms of viral exit from host cells | 1.79 | 9.76 × 10−2 | XPO1, LMNB1, ACTG1, ACTA1 | |
| UDP-N-acetyl-d-galactosamine biosynthesis II | 1.74 | 2.22 × 10−1 | GPI, UAP1 | |
| eNOS signaling | 1.70 | 5.67 × 10−2 | 0.000 | HSPA8, HSPA4, HSP90B1, HSP90AB1, HSPA9, CAV1, HSP90AA1, HSPA5 |
| Acetyl-CoA biosynthesis III (from citrate) | 1.62 | 1.00 | ACLY | |
| Ber pathway | 1.49 | 1.67 × 10−1 | PCNA, APEX1 | |
| Amyloid processing | 1.48 | 7.84 × 10−2 | 0.000 | CAPNS1, MAPK1, CAPN1, CAPN2 |
| Agranulocyte adhesion and diapedesis | 1.42 | 4.76 × 10−2 | ITGB1, GNAI2, MYH10, FN1, MYH9, EZR, ACTG1, ACTA1, MSN | |
| Cytotoxic T lymphocyte-mediated apoptosis of target cells | 1.40 | 9.38 × 10−2 | B2M, HLA-A, HLA-B | |
| Role of CHK proteins in cell cycle checkpoint control | 1.37 | 7.27 × 10−2 | 0.000 | PCNA, PPP2R1A, PPP2CA, CDK1 |
| Oxidative phosphorylation | 1.33 | 5.50 × 10−2 | SDHA, ATP5H, ATP5B, ATP5A1, CYC1, UQCRC1 | |
| Palmitate biosynthesis I (animals) | 1.33 | 5.00 × 10−1 | FASN | |
| Fatty acid biosynthesis initiation II | 1.33 | 5.00 × 10−1 | FASN | |
| Formaldehyde oxidation II (glutathione-dependent) | 1.33 | 5.00 × 10−1 | ESD | |
| Glycine biosynthesis I | 1.33 | 5.00 × 10−1 | SHMT2 | |
| Glutamate biosynthesis II | 1.33 | 5.00 × 10−1 | GLUD1 | |
| Glutamate degradation X | 1.33 | 5.00 × 10−1 | GLUD1 | |
| Chondroitin sulfate degradation (metazoa) | 1.31 | 1.33 × 10−1 | CD44, HEXB | |
| Androgen signaling | 1.30 | 5.41 × 10−2 | GNAI2, HSPA4, CALR, MAPK1, GNB2L1, HSP90AA1 | |
| Dermatan sulfate degradation (metazoa) | 1.26 | 1.25 × 10−1 | CD44, HEXB | |
| α-Adrenergic signaling | 1.24 | 5.75 × 10−2 | −1.000 | GNAI2, MAPK1, GNB2L1, PYGB, PYGL |
| Neuregulin signaling | 1.22 | 5.68 × 10−2 | ITGB1, HSP90B1, MAPK1, HSP90AB1, HSP90AA1 | |
| Methionine degradation I (to homocysteine) | 1.21 | 1.18 × 10−1 | PRMT5, AHCY | |
| Crosstalk between dendritic cells and natural killer cells | 1.21 | 5.62 × 10−2 | HLA-A, HLA-B, TLN1, ACTG1, ACTA1 | |
| Calcium signaling | 1.19 | 4.49 × 10−2 | MYH10, CALR, MYH9, MAPK1, TPM3, TPM4, TPM2, ACTA1 | |
| PPARα/RXRα activation | 1.18 | 4.47 × 10−2 | 0.378 | CAND1, HSP90B1, MAPK1, HSP90AB1, PDIA3, FASN, HSP90AA1, AIP |
| Valine degradation I | 1.17 | 1.11 × 10−1 | ECHS1, HADHA | |
| Death receptor signaling | 1.16 | 5.43 × 10−2 | 1.342 | LMNA, SPTAN1, ACTG1, ACTA1, HSPB1 |
| Neuroprotective role of THOP1 in Alzheimer’s disease | 1.16 | 7.50 × 10−2 | YWHAE, HLA-A, HLA-B | |
| Diphthamide biosynthesis | 1.16 | 3.33 × 10−1 | EEF2 | |
| NADH repair | 1.16 | 3.33 × 10−1 | GAPDH | |
| Methylglyoxal degradation I | 1.16 | 3.33 × 10−1 | GLO1 | |
| Hypusine biosynthesis | 1.16 | 3.33 × 10−1 | EIF5A | |
| Oxidized GTP and DGTP detoxification | 1.16 | 3.33 × 10−1 | DDX6 | |
| Gadd45 signaling | 1.13 | 1.05 × 10−1 | PCNA, CDK1 | |
| DNA damage-induced 14-3-3σ signaling | 1.13 | 1.05 × 10−1 | SFN, CDK1 | |
| Cysteine biosynthesis III (mammalia) | 1.13 | 1.05 × 10−1 | PRMT5, AHCY | |
| PPAR signaling | 1.13 | 5.32 × 10−2 | 0.447 | HSP90B1, MAPK1, HSP90AB1, HSP90AA1, AIP |
| Tec kinase signaling | 1.07 | 4.43 × 10−2 | −1.342 | ITGB1, GNAI2, RHOA, GNB2L1, STAT1, ACTG1, ACTA1 |
| Nitric oxide signaling in the cardiovascular system | 1.05 | 5.05 × 10−2 | 0.447 | HSP90B1, MAPK1, HSP90AB1, CAV1, HSP90AA1 |
| Cellular effects of sildenafil (viagra) | 1.05 | 4.65 × 10−2 | MYH10, MYH9, PDIA3, ACTG1, ACTA1, MYL12A | |
| Chemokine signaling | 1.05 | 5.63 × 10−2 | 0.000 | GNAI2, MAPK1, CFL1, RHOA |
| Uracil degradation II (reductive) | 1.04 | 2.50 × 10−1 | DPYSL2 | |
| Thymine degradation | 1.04 | 2.50 × 10−1 | DPYSL2 | |
| Geranylgeranyldiphosphate biosynthesis | 1.04 | 2.50 × 10−1 | FDPS | |
| Rapoport-luebering glycolytic shunt | 1.04 | 2.50 × 10−1 | PGAM1 | |
| Polyamine regulation in colon cancer | 1.02 | 9.09 × 10−2 | PSME1, PSME2 | |
| HIF1α signaling | 1.01 | 4.90 × 10−2 | MAPK1, HSP90AA1, LDHA, APEX1, LDHB | |
| Cardiac β-adrenergic signaling | 1.00 | 4.51 × 10−2 | −0.447 | PPP2R1A, PPP2CA, GNB2L1, PPP1CA, APEX1, PDE6H |
| nNOS signaling in neurons | 9.92 × 10−1 | 6.38 × 10−2 | CAPNS1, CAPN1, CAPN2 | |
| AMPK signaling | 9.90 × 10−1 | 4.48 × 10−2 | −0.447 | PPP2R1A, MAPK1, PPP2CA, FASN, PFKP, PPM1G |
| IL-22 signaling | 9.53 × 10−1 | 8.33 × 10−2 | MAPK1, STAT1 | |
| Serine biosynthesis | 9.44 × 10−1 | 2.00 × 10−1 | PHGDH | |
| DTMP de novo biosynthesis | 9.44 × 10−1 | 2.00 × 10−1 | SHMT2 | |
| Folate polyglutamylation | 9.44 × 10−1 | 2.00 × 10−1 | SHMT2 | |
| Xenobiotic metabolism signaling | 9.35 × 10−1 | 3.69 × 10−2 | HSP90B1, PPP2R1A, MAPK1, HSP90AB1, PPP2CA, HSP90AA1, GSTP1, PTGES3, ESD, AIP | |
| Cyclins and cell cycle regulation | 9.34 × 10−1 | 5.13 × 10−2 | PPP2R1A, PA2G4, PPP2CA, CDK1 | |
| Role of JAK family kinases in IL-6-type cytokine signaling | 9.24 × 10−1 | 8.00 × 10−2 | MAPK1, STAT1 | |
| Type I diabetes mellitus signaling | 9.08 × 10−1 | 4.55 × 10−2 | MAPK1, HLA-A, HLA-B, HSPD1, STAT1 | |
| Role of tissue factor in cancer | 9.08 × 10−1 | 4.55 × 10−2 | ITGB1, P4HB, MAPK1, CFL1, CYR61 | |
| Antiproliferative role of Tob in T cell signaling | 8.95 × 10−1 | 7.69 × 10−2 | PABPC1, MAPK1 | |
| Arginine biosynthesis IV | 8.70 × 10−1 | 1.67 × 10−1 | GLUD1 | |
| Superoxide radicals degradation | 8.70 × 10−1 | 1.67 × 10−1 | SOD1 | |
| UDP-N-acetyl-d-glucosamine biosynthesis II | 8.70 × 10−1 | 1.67 × 10−1 | UAP1 | |
| GDP-mannose biosynthesis | 8.70 × 10−1 | 1.67 × 10−1 | GPI | |
| IL-15 production | 8.69 × 10−1 | 7.41 × 10−2 | TWF1, STAT1 | |
| CCR3 signaling in eosinophils | 8.28 × 10−1 | 4.27 × 10−2 | GNAI2, MAPK1, CFL1, RHOA, GNB2L1 | |
| IL-8 signaling | 8.25 × 10−1 | 3.83 × 10−2 | −0.816 | GNAI2, MAPK1, RHOA, GNB2L1, IQGAP1, LASP1, CSTB |
| Glioma invasiveness signaling | 8.09 × 10−1 | 5.26 × 10−2 | MAPK1, RHOA, CD44 | |
| Acetyl-CoA biosynthesis I (pyruvate dehydrogenase complex) | 8.08 × 10−1 | 1.43 × 10−1 | DLAT | |
| GDP-glucose biosynthesis | 8.08 × 10−1 | 1.43 × 10−1 | PGM1 | |
| G β γ signaling | 7.98 × 10−1 | 4.55 × 10−2 | GNAI2, MAPK1, GNB2L1, CAV1 | |
| Role of p14/p19arf in tumor suppression | 7.95 × 10−1 | 6.67 × 10−2 | NPM1, SF3A1 | |
| PAK signaling | 7.86 × 10−1 | 4.49 × 10−2 | ITGB1, MAPK1, CFL1, MYL12A | |
| Induction of apoptosis by HIV1 | 7.63 × 10−1 | 5.00 × 10−2 | SLC25A6, SLC25A3, SLC25A5 | |
| Glucose and glucose-1-phosphate degradation | 7.55 × 10−1 | 1.25 × 10−1 | PGM1 | |
| Superpathway of methionine degradation | 7.51 × 10−1 | 6.25 × 10−2 | PRMT5, AHCY | |
| GM-CSFsignaling | 7.34 × 10−1 | 4.84 × 10−2 | MAPK1, GNB2L1, STAT1 | |
| Estrogen receptor signaling | 7.26 × 10−1 | 3.94 × 10−2 | PRKDC, DDX5, H3F3A/H3F3B, MAPK1, PHB2 | |
| PCP pathway | 7.20 × 10−1 | 4.76 × 10−2 | PFN1, RHOA, HSPB1 | |
| Oncostatin m signaling | 7.11 × 10−1 | 5.88 × 10−2 | MAPK1, STAT1 | |
| Interferon signaling | 7.11 × 10−1 | 5.88 × 10−2 | IFITM3, STAT1 | |
| Prostanoid biosynthesis | 7.09 × 10−1 | 1.11 × 10−1 | PTGES3 | |
| Folate transformations I | 7.09 × 10−1 | 1.11 × 10−1 | SHMT2 | |
| Pyridoxal 5′-phosphate salvage pathway | 7.06 × 10−1 | 4.69 × 10−2 | PDXK, MAPK1, CDK1 | |
| Cell cycle regulation by btg family proteins | 6.92 × 10−1 | 5.71 × 10−2 | PPP2R1A, PPP2CA | |
| Trna splicing | 6.92 × 10−1 | 5.71 × 10−2 | APEX1, PDE6H | |
| Amyotrophic lateral sclerosis signaling | 6.84 × 10−1 | 4.08 × 10−2 | SOD1, CAPNS1, CAPN1, CAPN2 | |
| p53 signaling | 6.84 × 10−1 | 4.08 × 10−2 | PRKDC, PCNA, SFN, GML | |
| Role of PI3K/Akt signaling in the pathogenesis of influenza | 6.80 × 10−1 | 4.55 × 10−2 | GNAI2, KPNA3, MAPK1 | |
| Phospholipase c signaling | 6.70 × 10−1 | 3.35 × 10−2 | −1.633 | ITGB1, PEBP1, MARCKS, AHNAK, MAPK1, RHOA, GNB2L1, MYL12A |
| Glycine βine degradation | 6.68 × 10−1 | 1.00 × 10−1 | SHMT2 | |
| Complement system | 6.56 × 10−1 | 5.41 × 10−2 | CD59, C1QBP | |
| Macropinocytosis signaling | 6.55 × 10−1 | 4.41 × 10−2 | ITGB1, RHOA, ACTN4 | |
| Relaxin signaling | 6.54 × 10−1 | 3.70 × 10−2 | GNAI2, MAPK1, GNB2L1, APEX1, PDE6H | |
| CCR5 signaling in macrophages | 6.42 × 10−1 | 4.35 × 10−2 | GNAI2, MAPK1, GNB2L1 | |
| Melatonin signaling | 6.30 × 10−1 | 4.29 × 10−2 | GNAI2, MAPK1, PDIA3 | |
| Role of PKR in interferon induction and antiviral response | 6.06 × 10−1 | 5.00 × 10−2 | STAT1, EIF2S1 | |
| Synaptic long term depression | 6.06 × 10−1 | 3.55 × 10−2 | 0.447 | GNAI2, PPP2R1A, MAPK1, PPP2CA, PDIA3 |
| Dendritic cell maturation | 5.90 × 10−1 | 3.35 × 10−2 | B2M, MAPK1, HLA-A, PDIA3, HLA-B, STAT1 | |
| Production of nitric oxide and reactive oxygen species in macrophages | 5.83 × 10−1 | 3.33 × 10−2 | −0.816 | PPP2R1A, MAPK1, PPP2CA, RHOA, STAT1, PPP1CA |
| Sphingosine-1-phosphate signaling | 5.79 × 10−1 | 3.67 × 10−2 | 0.000 | GNAI2, MAPK1, PDIA3, RHOA |
| Assembly of rna polymerase III complex | 5.69 × 10−1 | 7.69 × 10−2 | SF3A1 | |
| Systemic lupus erythematosus signaling | 5.62 × 10−1 | 3.18 × 10−2 | PRPF19, MAPK1, HLA-A, HNRNPA2B1, HLA-B, SNRPD1, SNRPD3 | |
| PDGF signaling | 5.54 × 10−1 | 3.90 × 10−2 | MAPK1, CAV1, STAT1 | |
| iNOS signaling | 5.48 × 10−1 | 4.55 × 10−2 | MAPK1, STAT1 | |
| Cardiac hypertrophy signaling | 5.45 × 10−1 | 3.14 × 10−2 | −0.816 | GNAI2, MAPK1, PDIA3, RHOA, GNB2L1, HSPB1, MYL12A |
| Dopamine receptor signaling | 5.44 × 10−1 | 3.85 × 10−2 | PPP2R1A, PPP2CA, PPP1CA | |
| Urate biosynthesis/inosine 5′-phosphate degradation | 5.42 × 10−1 | 7.14 × 10−2 | IMPDH2 | |
| Colanic acid building blocks biosynthesis | 5.42 × 10−1 | 7.14 × 10−2 | GPI | |
| CXCR4 signaling | 5.25 × 10−1 | 3.29 × 10−2 | 0.000 | GNAI2, MAPK1, RHOA, GNB2L1, MYL12A |
| MSP-RON signaling pathway | 5.21 × 10−1 | 4.35 × 10−2 | ACTG1, ACTA1 | |
| Methylglyoxal degradation III | 5.17 × 10−1 | 6.67 × 10−2 | AKR1B1 | |
| Thrombin signaling | 5.14 × 10−1 | 3.14 × 10−2 | 0.447 | GNAI2, MAPK1, PDIA3, RHOA, GNB2L1, MYL12A |
| CD28 signaling in Thelper cells | 5.06 × 10−1 | 3.39 × 10−2 | 0.000 | ACTR2, ACTR3, ARPC1B, ARPC2 |
| PTEN signaling | 5.06 × 10−1 | 3.39 × 10−2 | ITGB1, MAPK1, YWHAH, IGF2R | |
| P2Y purigenic receptor signaling pathway | 4.99 × 10−1 | 3.36 × 10−2 | GNAI2, MAPK1, PDIA3, GNB2L1 | |
| Graft-versus-host disease signaling | 4.97 × 10−1 | 4.17 × 10−2 | HLA-A, HLA-B | |
| Ephrin a signaling | 4.97 × 10−1 | 4.17 × 10−2 | CFL1, RHOA | |
| Mismatch repair in eukaryotes | 4.94 × 10−1 | 6.25 × 10−2 | PCNA | |
| Gαi signaling | 4.92 × 10−1 | 3.33 × 10−2 | GNAI2, MAPK1, GNB2L1, CAV1 | |
| Autoimmune thyroid disease signaling | 4.85 × 10−1 | 4.08 × 10−2 | HLA-A, HLA-B | |
| TR/RXR activation | 4.79 × 10−1 | 3.53 × 10−2 | ENO1, FASN, PFKP | |
| Γ-linolenate biosynthesis II (animals) | 4.72 × 10−1 | 5.88 × 10−2 | CYB5R3 | |
| Allograft rejection signaling | 4.71 × 10−1 | 3.49 × 10−2 | B2M, HLA-A, HLA-B | |
| Dopamine-DARPP32 feedback in camp signaling | 4.68 × 10−1 | 3.11 × 10−2 | GNAI2, PPP2R1A, PPP2CA, PDIA3, PPP1CA | |
| UVa-induced MAPK signaling | 4.54 × 10−1 | 3.41 × 10−2 | MAPK1, PDIA3, STAT1 | |
| CNTF signaling | 4.51 × 10−1 | 3.85 × 10−2 | MAPK1, STAT1 | |
| Endometrial cancer signaling | 4.51 × 10−1 | 3.85 × 10−2 | MAPK1, CTNNA1 | |
| OX40 signaling pathway | 4.46 × 10−1 | 3.37 × 10−2 | B2M, HLA-A, HLA-B | |
| UVb-induced MAPK signaling | 4.41 × 10−1 | 3.77 × 10−2 | H3F3A/H3F3B, MAPK1 | |
| Communication between innate and adaptive immune cells | 4.31 × 10−1 | 3.30 × 10−2 | B2M, HLA-A, HLA-B | |
| IL-1 signaling | 4.31 × 10−1 | 3.30 × 10−2 | GNAI2, MAPK1, GNB2L1 | |
| Thrombopoietin signaling | 4.21 × 10−1 | 3.64 × 10−2 | MAPK1, STAT1 | |
| Purine nucleotides degradation II (aerobic) | 4.16 × 10−1 | 5.00 × 10−2 | IMPDH2 | |
| Wnt/Ca++ pathway | 4.11 × 10−1 | 3.57 × 10−2 | PDIA3, PDE6H | |
| EGF signaling | 4.11 × 10−1 | 3.57 × 10−2 | MAPK1, STAT1 | |
| Glioma signaling | 4.01 × 10−1 | 3.16 × 10−2 | MAPK1, PA2G4, IGF2R | |
| Maturity onset diabetes of young (mody) signaling | 3.84 × 10−1 | 4.55 × 10−2 | GAPDH | |
| ATM signaling | 3.84 × 10−1 | 3.39 × 10−2 | TRIM28, CDK1 | |
| Granulocyte adhesion and diapedesis | 3.81 × 10−1 | 2.82 × 10−2 | ITGB1, GNAI2, EZR, HSPB1, MSN | |
| Estrogen-dependent breast cancer signaling | 3.59 × 10−1 | 3.23 × 10−2 | HSD17B10, MAPK1 | |
| Role of JAK1, JAK2 and TYK2 in interferon signaling | 3.56 × 10−1 | 4.17 × 10−2 | STAT1 | |
| Estrogen-mediated s-phase entry | 3.56 × 10−1 | 4.17 × 10−2 | CDK1 | |
| Antiproliferative role of somatostatin receptor 2 | 3.51 × 10−1 | 3.17 × 10−2 | MAPK1, GNB2L1 | |
| Role of JAK1 and JAK3 in γc cytokine signaling | 3.51 × 10−1 | 3.17 × 10−2 | MAPK1, STAT1 | |
| Role of lipids/lipid rafts in the pathogenesis of influenza | 3.43 × 10−1 | 4.00 × 10−2 | FDPS | |
| IL-17a signaling in gastric cells | 3.43 × 10−1 | 4.00 × 10−2 | MAPK1 | |
| Cell cycle: G1/S checkpoint regulation | 3.43 × 10−1 | 3.12 × 10−2 | PA2G4, RPL5 | |
| Non-small cell lung cancer signaling | 3.35 × 10−1 | 3.08 × 10−2 | MAPK1, PA2G4 | |
| Pancreatic adenocarcinoma signaling | 3.31 × 10−1 | 2.83 × 10−2 | MAPK1, PA2G4, STAT1 | |
| GABA receptor signaling | 3.21 × 10−1 | 2.99 × 10−2 | AP2B1, AP1B1 | |
| Pyrimidine ribonucleotides interconversion | 3.19 × 10−1 | 3.70 × 10−2 | CTPS1 | |
| Growth hormone signaling | 3.07 × 10−1 | 2.90 × 10−2 | MAPK1, STAT1 | |
| Corticotropin releasing hormone signaling | 3.04 × 10−1 | 2.70 × 10−2 | GNAI2, MAPK1, KRT1 | |
| Pyrimidine ribonucleotides de novo biosynthesis | 2.97 × 10−1 | 3.45 × 10−2 | CTPS1 | |
| IL-3 signaling | 2.94 × 10−1 | 2.82 × 10−2 | MAPK1, STAT1 | |
| PEDF signaling | 2.94 × 10−1 | 2.82 × 10−2 | MAPK1, RHOA | |
| GPCR-mediated integration of enteroendocrine signaling exemplified by an L cell | 2.94 × 10−1 | 2.82 × 10−2 | GNAI2, PDIA3 | |
| JAK/STAT signaling | 2.87 × 10−1 | 2.78 × 10−2 | MAPK1, STAT1 | |
| Glutathione-mediated detoxification | 2.87 × 10−1 | 3.33 × 10−2 | GSTP1 | |
| Sonic hedgehog signaling | 2.87 × 10−1 | 3.33 × 10−2 | CDK1 | |
| Hereditary breast cancer signaling | 2.84 × 10−1 | 2.61 × 10−2 | NPM1, SFN, CDK1 | |
| NF-κB activation by viruses | 2.81 × 10−1 | 2.74 × 10−2 | ITGB1, MAPK1 | |
| Prolactin signaling | 2.81 × 10−1 | 2.74 × 10−2 | MAPK1, STAT1 | |
| STAT3 pathway | 2.81 × 10−1 | 2.74 × 10−2 | MAPK1, IGF2R | |
| 4-1BB signaling in T lymphocytes | 2.78 × 10−1 | 3.23 × 10−2 | MAPK1 | |
| Flt3 signaling in hematopoietic progenitor cells | 2.75 × 10−1 | 2.70 × 10−2 | MAPK1, STAT1 | |
| Leptin signaling in obesity | 2.75 × 10−1 | 2.70 × 10−2 | MAPK1, PDIA3 | |
| Gα12/13 signaling | 2.74 × 10−1 | 2.56 × 10−2 | MAPK1, RHOA, MYL12A | |
| p38 MAPK signaling | 2.74 × 10−1 | 2.56 × 10−2 | H3F3A/H3F3B, STAT1, HSPB1 | |
| TREM1 signaling | 2.69 × 10−1 | 2.67 × 10−2 | ITGB1, MAPK1 | |
| Synaptic long term potentiation | 2.65 × 10−1 | 2.52 × 10−2 | MAPK1, PDIA3, PPP1CA | |
| HMGB1 signaling | 2.60 × 10−1 | 2.50 × 10−2 | HMGB1, MAPK1, RHOA | |
| G protein signaling mediated by tubby | 2.60 × 10−1 | 3.03 × 10−2 | GNB2L1 | |
| MIF-mediated glucocorticoid regulation | 2.60 × 10−1 | 3.03 × 10−2 | MAPK1 | |
| Retinol biosynthesis | 2.60 × 10−1 | 3.03 × 10−2 | ESD | |
| Ethanol degradation ii | 2.60 × 10−1 | 3.03 × 10−2 | HSD17B10 | |
| LXR/RXR activation | 2.56 × 10−1 | 2.48 × 10−2 | ECHS1, FASN, HADH | |
| IL-9 signaling | 2.51 × 10−1 | 2.94 × 10−2 | STAT1 | |
| Inhibition of angiogenesis by TSP1 | 2.51 × 10−1 | 2.94 × 10−2 | MAPK1 | |
| Reelin signaling in neurons | 2.47 × 10−1 | 2.53 × 10−2 | ITGB1, PAFAH1B2 | |
| IL-17a signaling in fibroblasts | 2.43 × 10−1 | 2.86 × 10−2 | MAPK1 | |
| Role of JAK2 in hormone-like cytokine signaling | 2.43 × 10−1 | 2.86 × 10−2 | STAT1 | |
| Stearate biosynthesis I (animals) | 2.43 × 10−1 | 2.86 × 10−2 | FASN | |
| Noradrenaline and adrenaline degradation | 2.43 × 10−1 | 2.86 × 10−2 | HSD17B10 | |
| Nucleotide excision repair pathway | 2.43 × 10−1 | 2.86 × 10−2 | RAD23B | |
| Ceramide signaling | 2.42 × 10−1 | 2.50 × 10−2 | PPP2R1A, PPP2CA | |
| Estrogen biosynthesis | 2.28 × 10−1 | 2.70 × 10−2 | HSD17B10 | |
| April mediated signaling | 2.21 × 10−1 | 2.63 × 10−2 | MAPK1 | |
| Netrin signaling | 2.14 × 10−1 | 2.56 × 10−2 | ENAH | |
| B cell activating factor signaling | 2.07 × 10−1 | 2.50 × 10−2 | MAPK1 | |
| Thyroid cancer signaling | 2.07 × 10−1 | 2.50 × 10−2 | MAPK1 | |
| Transcriptional regulatory network in embryonic stem cells | 2.07 × 10−1 | 2.50 × 10−2 | SET | |
| MIF regulation of innate immunity | 2.01 × 10−1 | 2.44 × 10−2 | MAPK1 |
| ID | Molecules in Network | Score | Molecules | Top Diseases and Functions |
|---|---|---|---|---|
| 1 | ALDO, ALDOA, ALDOC, ATP synthase, C14orf166, DDX1, ENO1, ENO2, enolase, FUS, HIST2H2AC, HNRNPU, NAP1L4, NCL, NONO, PA2G4, PGK1, Ras, RPL5, RPL6, RPL7, RPL8, RPL12, RPL18, RPL22, RPL23, RPL10A, RPL7A, RPLP1, RPLP2, SFPQ, SQSTM1, SYNCRIP, T3-TR-RXR, TARS | 47 | 30 | RNA post-transcriptional modification, carbohydrate metabolism, small molecule biochemistry |
| 2 | 14-3-3, 14-3-3 (β, ε, ζ), 14-3-3(β, γ, θ, η, ζ), 14-3-3(η, θ, ζ), adenosine-tetraphosphatase, ATP5A1, ATP5B, C1QBP, CAPZB, CLTC, dishevelled, DYNC1H1, EPRS, ESYT1, GNAI2, GNB2L1, GSK3, HSP90AA1, HSPA9, KIF5B, MARS, PDCD6, RNH1, SFN, SLC25A3, SLC25A5, SLC25A6, TPI1, TUFM, YWHAB, YWHAE, YWHAG, YWHAH, YWHAQ, YWHAZ | 42 | 28 | Protein trafficking, cell death and survival, nucleic acid metabolism |
| 3 | 60S ribosomal subunit, CAND1, CLIC1, EIF2, ERK, HNRNPH3, IPO9, Ku, LAMB1, laminin1, laminin2, NPM1, PDXK, PYGB, ribosomal 40s subunit, RNR, RPL3, RPL4, RPL10, RPL21, RPL27, RPLP0, RPS2, RPS3, RPS5, RPS7, RPS8, RPS10, RPS11, RPS12, RPS15, RPS24, RPS27A, RPS4X, RPSA | 40 | 27 | Hematological disease, organismal injury and abnormalities, RNA post-transcriptional modification |
| 4 | α-Tubulin, Ant, ATP5H, β-tubulin, CCT2, CCT3, CCT4, CCT5, CCT7, CCT8, CCT6A, DPYSL2, dynein, EHD1, ENAH, ERP29, GCN1L1, integrin α5 β1, MAP1B, MAPK1, NUMA1, PPP2CA, PPP2R1A, TCP1, TUBA1B, TUBB6, TUBB8, TUBB4B, tubulin (complex), tubulin (family), VAPA, VDAC, VDAC1, VDAC2, ZYX | 40 | 27 | Cellular assembly and organization, cell-to-cell signaling and interaction, reproductive system development and function |
| 5 | ACLY, aconitase, Akt, angiotensin II receptor type 1, Arf, ARF1, CNN3, COP I, COPA, COPB1, COPB2, COPE, COPG1, CS, EIF3, EIF6, EIF3C, EIF3M, HN1, LRRC59, malate dehydrogenase, MAP4, MDH1, MDH2, N-cadherin, PARK7, RAB1B, SEPT2, SEPT7, SEPT9, Septin, SHMT2, TALDO1, TXNDC5, WARS | 38 | 26 | Infectious disease, small molecule biochemistry, reproductive system disease |
| 6 | 19S proteasome, 20S proteasome, 26S proteasome, BUB3, CPNE1, ECHS1, immunoproteasome Pa28/20s, KPNA3, MHC CLASS I (family), NF-kB (complex), NPLOC4, NSFL1C, OTUB1, PDLIM1, PRDX2, Proteasome PA700/20s, PSMA, PSMA1, PSMA2, PSMA4, PSMA5, PSMA7, PSMB1, PSMC1, PSMC4, PSMD2, PSME1, PSME2, RAD23B, RPN2, UBA1, UBE2, UBE2L3, UBE2N, ubiquitin | 36 | 25 | Infectious disease, organismal injury and abnormalities, immunological disease |
| 7 | AHCY, calpain, CYR61, DDX6, DDX3X, EIF2S2, EIF3B, EIF3I, EIF4A, EIF4A1, EIF4F, EIF4G, EIF4G1, EIF4H, fascin, filamin, FLNA, FLNB, FLNC, G3BP1, HNRNPA1, HNRNPL, ILF2, ILF3, LFA-1, MIRLET7, OLA1, PABPC1, PCBP1, PCBP2, PKC(s), PUF60, SAPK, SNX3, VPS35 | 36 | 25 | Protein synthesis, cellular movement, gene expression |
| 8 | CALU, CBX3, CD59, CDK1, DBI, DDX5, DNA-methyltransferase, DPY30, GML, GSTP1, H3F3A/H3F3B, histone H3, histone H4, HNRNPA3, HNRNPK, HSP70, IFITM3, IgA, MATR3, NAP1L1, NASP, NOS, PKM, PPM1G, PRMT5, secretase γ, SH3BGRL3, Sod, SOD1, SRC (family), TFRC, thymidine kinase, TRIM28, UBXN4, YBX1 | 36 | 25 | Cellular growth and proliferation, infectious disease, cellular development |
| 9 | APEX1, c-Src, CLIC4, CSE1L, H2AFY, HDGF, HIST1H1C, histone H1, HNRNP H, HNRNPH1, HNRNPM, importin α, importin β, integrin α3β1, IPO5, IPO7, karyopherin β, KPNA2, KPNB1, LOC102724594/U2AF1, PHGDH, PRPF19, PTMA, RAN, RANBP1, SAE1, SF3B1, snRNP, SNRPD1, SNRPD3, TNPO1, transportin, U2AF, U2AF2, VEGF | 34 | 24 | Molecular transport, protein trafficking, RNA post-transcriptional modification |
| 10 | Adaptor protein 1, AIP, AP1γ, AP1B1, CALR, CAP1, CD1, CD1D-CANX-CALR-ERp57, CFL1, CRIP2, cytoplasmic dynein, DNAJ, DNAJA1, DNAJA2, DNAJC8, FKBP4, G-actin, HSP, HSP90, HSP22/HSP40/HSP90, HSP90B1, HYOU1, JNK, P4HB, PCMT1, PDIA3, PDIA4, PDIA6, PRDX4, PTGES3, PYGL, STIP1, TMSB4, TPM2, TPM3 | 32 | 23 | Post-translational modification, protein folding, drug metabolism |
| 11 | ACTG1, ADRB, AP2B1, ATPase, CAPZA1, casein, caspase, creatine kinase, DLAT, DYNC1I2, EEF2, EEF1A1, HSP90AB1, HSPA4, HSPA5, HSPA8, HSPB1, HSPD1, IDI1, IFNβ, LONP1, mediator, MHC Class II (complex), MMP, NACA, PHB2, PPA1, PPP2C, PRKDC, RPN1, RSK, STMN1, TLR, TXLNA, VCP | 32 | 23 | Post-translational modification, protein folding, cell morphology |
| 12 | Actin, ACTN1, ACTN4, α-actinin, α-catenin, ANXA6, ARP2/3, cadherin, CAPG, CAPN2, Cdc2, CSRP1, CTNNA1, cytokeratin, F actin, GPI, IMPDH2, integrinβ, KRT1, KRT8, KRT9, KRT18, lamin b, LMNB1, PI3K (complex), PLEC, spectrin, SPTAN1, SPTBN1, talin, TLN1, TPM4, TUBB, VCL, VIM | 30 | 22 | Cellular compromise, cellular assembly and organization, cellular function and maintenance |
| 13 | ACTR2, ACTR3, α-actin, ARHGDIA, ARPC2, ARPC1B, BCR (complex), CD44, CTTN, ERM, EZR, GDI1, GDI2, IQGAP1, MAPK, MIR124, MSN, PAFAH1B2, PAK, PLIN3, PLS3, RAB5, RAB11, RAB11A, RAB7A, Rac, Ras homolog, Rho GDI, RTN4, SEC13, Sos, TMOD3, TWF2, VAT1, VAV | 30 | 22 | Cellular assembly and organization, cellular function and maintenance, cell morphology |
| 14 | 3-Hydroxyacyl-CoA dehydrogenase, ACAT1, ACAT2, acetyl-CoA C-acetyltransferase, AHNAK, AKR1B1, ANXA1, ANXA2, calmodulin, CAPN1, CAPNS1, CAST, CAV1, caveolin, CKAP4, CYC1, cytochrome bc1, cytochrome C, cytochrome-c oxidase, DNM1L, dynamin, fibrinogen, GAPDH, GLUD1, HADH, HADHA, HSD17B10, IMMT, integrin, MVP, PARP, Smad, tyrosine kinase, UQCRC1, VLDL-cholesterol | 28 | 21 | Endocrine system development and function, lipid metabolism, small molecule biochemistry |
| 15 | AARS, AHSA1, B2M, B2m-MHC1a, CACYBP, CANX, CAPRIN1, chymotrypsin, EEF1B2, EEF1D, EEF1G, ENaC, ERK1/2, ESD, GARS, GC-GCR dimer, GMPS, HARS, HLA Class I, HLA-A, HLA-abc, HLA-B, HLA-B27, KIR, MHC Class I (complex), MHC I-α, Na+/K+-ATPase, NPEPPS, PDE6H, PDGF (family), PKI, RPS3A, TAP, TBCA, VARS | 26 | 20 | Post-translational modification, protein folding, dermatological diseases and conditions |
| 16 | BTF3, Cbp/p300, CDK1/2, cyclin A, cyclin D, cyclin E, DHX9, E2f, Gap, HDLBP, Holo RNA polymerase II, MCM, MCM3, MCM4, MCM6, MCM7, PCM1, PCNA, PDE, Raf, Rb, RHOA, RPA, S100A6, SERBP1, SND1, SSRP1, SUPT16H, TAGLN, TIP60, TMPO, tropomyosin, XRCC5, XRCC6, YBX3 | 26 | 20 | DNA replication, recombination, and repair, cell morphology, cellular function and maintenance |
| 17 | ACTA1, BASP1, CaMKII, cofilin, CORO1C, COTL1, CTPS1, DBN1, FKBP1A, GLO1, glycogen phosphorylase, ITPR, LDH (complex), LDHA, LDHB, MARCKS, MEF2, Mlc, MYH9, MYH10, MYL12A, myosin2, PFN1, PICALM, PKA, PP1 protein complex group, PP2A, profilin, PTRF, RAP1, RBM3, RNA polymerase I, Rock, voltage-gated calcium channel, WDR1 | 24 | 19 | Cellular movement, connective tissue disorders, hematological disease |
| 18 | ALYREF, ATP1A1, BSG, calcineurin A, calcineurin protein(s), CK2, DDX39B, EIF2S1, EIF5A, ETFA, FCER1, focal adhesion kinase, HEXB, HINT1, IgE, IGF2R, Ikb, JINK1/2, MAP2K1/2, mitochondrial complex 1, NFAT (complex), NFAT (family), NF-kB (family), peptidylprolyl isomerase, phosphatase, PLCγ, PP1-C, PP1/PP2A, PPIA, PPP1CA, RCN1, SET, STOML2, VLA-4, XPO1 | 19 | 16 | Molecular transport, RNA trafficking, cell death and survival |
| 19 | CARHSP1, CDV3, CECR2, CLDN12, CTSA, CUTA, CYB5R3, DMXL2, FADS1, FAM57A, FBXO11, HEBP2, HM13, KHSRP, LRPPRC, NEU1, NME1-NME2, NNMT, PARN, PFKP, PGM1, POLRMT, SDHA, SDHAF2, SDHC, SIRT5, SLC25A25, SURF4, TIPARP, TKT, TRMT1, UAP1, UBC, VPS37A, ZCCHC6 | 19 | 16 | Developmental disorder, hereditary disorder, metabolic disease |
| 20 | ANXA5, collagen, collagen α1, collagen type I, collagen type II, collagen type III, collagen type IV, collagen(s), complement component 1, CPLA2, CSTB, CTSB, eotaxin, FDPS, fibrin, FUBP3, G6PD, GPIIB-IIIA, Hsp27, IARS, laminin, P38 MAPK, PDGF (complex), PDGF-AA, peroxidase (miscellaneous), PKG, PLOD2, PRDX1, PRDX5, QARS, RRBP1, SERPINH1, TNC, TPP1, trypsin | 17 | 15 | Cancer, endocrine system disorders, organismal injury and abnormalities |
| 21 | AMPK, caspase 3/7, CD3, CD8, cyclin B, FASN, HNRNPA2B1, HNRNPR, IFN, IFNγ, IFNAR, IL-2R, IL12 (complex), interferon-α, JAK, KHDRBS1, LGALS1, Mek, MHC, MTORC1, p70 S6K, p85 (PIK3R), PEBP1, PGAM1, PGD, PRKAA, PTBP1, PTK, RARS, SF3A1, SREBP, STAT1, STAT5a/b, TCR, TWF1 | 14 | 13 | RNA post-transcriptional modification, dermatological diseases and conditions, cell death and survival |
| 22 | ARPIN/C15orf38-AP3S2, ATIC, ESD, FAM169A, γ-tubulin, GANAB, GBA3, GPATCH1, HEATR5A, KIAA1551, KIF18B, LMNA, MAPRE1, MUM1L1, NAPG, PAICS, PRKCSH, RCCD1, RNA polymerase II, SLC22A23, SLC33A1, SLC41A2, TAGLN2, TCF, THUMPD3, TMEM64, TMEM164, TMEM201, TMEM183A, TRAF6, UBC, UCHL1, WDR70, ZNF397, ZNF280C | 8 | 9 | Carbohydrate metabolism, small molecule biochemistry, cell morphology |
| 23 | ALP, Ap1, C1q, Creb, FGF, FGFR, FN1, HDL, hemoglobin, HIST1H2BL, histon, HMGB1, IgG, IgG1, IgG2a, IgM, IL1, IL12 (family), immunoglobulin, insulin, ITGB1, LASP1, LDL, LRP, NADPH oxidase, NR1H, PDGF BB, PEA15, PI3K (family), PLD, pro-inflammatory cytokine, Shc, SLC38A2, TGFβ, TSH | 6 | 7 | Cell-to-cell signaling and interaction, tissue development, cardiovascular system development and function |
| 24 | Adaptor protein 2, ADCY, βARK, CAP2, CG, chemokine, clathrin, CRH, estrogen receptor, FPR, FSH, G protein, G protein α, G protein αi, G protein β γ, Gi-coupled receptor, growth hormone, HINT1, IKK (complex), IL8r, LH, LPAR2, metalloprotease, MMP19, myosin2, NMDA receptor, PDGFR, PDXK, PLC, PPIH, proinsulin, S1PR5, SFK, TNF (family), TRHR | 1 | 2 | Connective tissue development and function, tissue morphology, behavior |
| Categories | Diseases or Functions Annotation | p-Value | Activation z-Score | Molecules | Number of Molecules |
|---|---|---|---|---|---|
| Cell death and survival | Cell death | 2.14 × 10−35 | 0.626 | AARS, ACAT1, ACLY, ACTN4, AHSA1, AKR1B1, ALDOA, ALDOC, ANXA1, ANXA2, ANXA5, AP2B1, APEX1, ARHGDIA, ATP1A1, ATP5A1, B2M, BASP1, BSG, BTF3, C1QBP, CACYBP, CALR, CANX, CAPN1, CAPN2, CAPNS1, CAPRIN1, CAST, CAV1, CCT2, CCT3, CCT4, CCT5, CCT6A, CCT7, CCT8, CD44, CD59, CDK1, CFL1, CLIC4, CSE1L, CSTB, CTNNA1, CTSB, CTTN, CYR61, DDX3X, DDX5, DHX9, DNAJA1, DNM1L, DYNC1H1, EEF1A1, EEF1D, EHD1, EIF2S1, EIF3B, EIF3C, EIF3I, EIF4G1, EIF5A, EIF6, ENO1, EZR, FASN, FDPS, FKBP1A, FKBP4, FLNA, FLNB, FN1, FUS, G6PD, GAPDH, GLO1, GLUD1, GML, GNAI2, GNB2L1, GPI, GSTP1, HADHA, HDGF, HEXB, HINT1, HIST1H1C, HLA-A, HLA-B, HMGB1, HNRNPA1, HNRNPK, HSD17B10, HSP90AA1, HSP90AB1, HSP90B1, HSPA4, HSPA5, HSPA8, HSPA9, HSPB1, HSPD1, HYOU1, IGF2R, ILF2, ILF3, IMMT, IMPDH2, ITGB1, KHDRBS1, KPNA2, KPNB1, KRT18, KRT8, LDHA, LGALS1, LMNA, LMNB1, LONP1, MAP1B, MAP4, MAPK1, MCM7, MDH1, MSN, MVP, MYH9, NCL, | 232 |
| NPM1, NUMA1, OTUB1, P4HB, PA2G4, PAFAH1B2, PARK7, PCBP2, PCMT1, PCNA, PDCD6, PDIA3, PDXK, PEA15, PEBP1, PHB2, PKM, PLEC, PLIN3, PPIA, PPP1CA, PPP2CA, PPP2R1A, PRDX1, PRDX2, PRDX4, PRDX5, PRKDC, PRMT5, PRPF19, PSMB1, PSMC1, PSMC4, PSMD2, PSME2, PTGES3, PTMA, QARS, RAD23B, RAN, RANBP1, RBM3, RHOA, RPL10, RPLP0, RPS24, RPS3, RPS3A, RPSA, RRBP1, RTN4, S100A6, SDHA, SERPINH1, SET, SF3A1, SFN, SFPQ, SH3BGRL3, SLC25A5, SLC25A6, SND1, SOD1, SPTBN1, SQSTM1, SSRP1, STAT1, STIP1, STMN1, STOML2, TAGLN2, TCP1, TFRC, TNC, TPM3, TPP1, TRIM28, TUBB, TUBB6, TUFM, TXNDC5, UBA1, UBE2L3, UBE2N, UCHL1, VAPA, VCL, VCP, VDAC1, VDAC2, VIM, XPO1, XRCC5, XRCC6, YBX1, YBX3, YWHAB, YWHAE, YWHAG, YWHAH, YWHAQ, YWHAZ, ZYX | |||||
| Cellular growth and proliferation | Proliferation of cells | 5.39 × 10−34 | −1.530 | AARS, ACAT1, ACLY, ACTG1, ACTN1, ACTN4, AHCY, AHNAK, AHSA1, AKR1B1, ALDOA, ANXA1, ANXA2, ANXA6, APEX1, ARF1, ARHGDIA, ATIC, ATP5A1, ATP5B, B2M, BASP1, BSG, BUB3, C1QBP, CACYBP, CALR, CAPN1, CAPN2, CAPNS1, CAPRIN1, CAPZA1, CAST, CAV1, CCT2, CCT3, CCT5, CCT7, CD44, CD59, CDK1, CFL1, CLIC1, CLTC, COPE, CSE1L, CSRP1, CTNNA1, CTSB, CTTN, CYR61, DBI, DBN1, DDX3X, DDX5, DNAJA1, DNAJA2, DNM1L, DPY30, DPYSL2, DYNC1H1, EEF1A1, EEF1B2, EEF1D, EIF3B, EIF3C, EIF3I, EIF4A1, EIF4G1, EIF5A, EIF6, ENO1, EZR, FASN, FKBP1A, FKBP4, FLNA, FN1, FUS, G3BP1, G6PD, GAPDH, GML, GNAI2, GNB2L1, GPI, GSTP1, H2AFY, H3F3A/H3F3B, HADHA, HDGF, HEXB, HINT1, HMGB1, HNRNPA1, HNRNPA2B1, HNRNPK, HNRNPL, HNRNPM, HNRNPR, HNRNPU, HSP90AA1, HSP90AB1, HSP90B1, HSPA4, HSPA5, HSPA8, HSPB1, HSPD1, IFITM3, IGF2R, ILF2, ILF3, IMMT, IMPDH2, IPO7, IQGAP1, ITGB1, KHDRBS1, KPNA2, KRT8, LAMB1, LASP1, LDHA, LGALS1, LMNA, LMNB1, LONP1, MAP1B, MAPK1, MAPRE1, MARCKS, MCM3, MCM4, MCM7, MVP, MYH10, MYH9, NACA, NAP1L1, NASP, NCL, NNMT, NPM1, NUMA1, PA2G4, PARK7, PCNA, PDCD6, PDIA3, PDXK, PEA15, PEBP1, PFKP, PFN1, PGK1, PICALM, PKM, PLEC, PLIN3, PLS3, PPIA, PPM1G, PPP1CA, PPP2CA, PPP2R1A, PRDX1, PRDX2, PRDX4, PRKCSH, PRKDC, PRMT5, PRPF19, PSMA4, PSMC1, PSMC4, PSMD2, PSME2, PTBP1, PTGES3, PTMA, RAB11A, RAN, RANBP1, RBM3, RHOA, RNH1, RPS3A, RPS4X, RPSA, RTN4, S100A6, SAE1, SEPT9, SERPINH1, SET, SFN, SFPQ, SHMT2, SLC25A5, SLC25A6, SND1, SNX3, SOD1, SPTAN1, SPTBN1, SQSTM1, STAT1, STIP1, STMN1, SURF4, TAGLN, TAGLN2, TCP1, TFRC, TLN1, TMPO, TNC, TPM3, TRIM28, TUBB, TUBB4B, TXLNA, TXNDC5, UBA1, UBE2L3, UBE2N, UCHL1, VAPA, VCL, VCP, VDAC1, VIM, WARS, XRCC5, XRCC6, YBX1, YBX3, YWHAG, YWHAQ, YWHAZ, ZYX | 242 |
| Cell death and survival | Apoptosis | 8.59 × 10−31 | 0.054 | AARS, ACLY, ACTN4, AHSA1, AKR1B1, ALDOA, ALDOC, ANXA1, ANXA2, ANXA5, APEX1, ARHGDIA, ATP1A1, B2M, BASP1, BSG, BTF3, C1QBP, CALR, CANX, CAPN1, CAPN2, CAPNS1, CAPRIN1, CAST, CAV1, CCT2, CCT4, CD44, CD59, CDK1, CFL1, CLIC4, CSE1L, CSTB, CTNNA1, CTSB, CTTN, CYR61, DDX3X, DDX5, DHX9, DNAJA1, DNM1L, DYNC1H1, EEF1A1, EEF1D, EHD1, EIF2S1, EIF3B, EIF3C, EIF3I, EIF4G1, EIF5A, EIF6, ENO1, EZR, FASN, FKBP1A, FKBP4, FLNA, FLNB, FN1, FUS, G6PD, GAPDH, GLO1, GLUD1, GML, GNAI2, GNB2L1, GPI, GSTP1, HDGF, HEXB, HINT1, HIST1H1C, HLA-B, | 192 |
| HMGB1, HNRNPA1, HNRNPK, HSD17B10, HSP90AA1, HSP90AB1, HSP90B1, HSPA4, HSPA5, HSPA8, HSPA9, HSPB1, HSPD1, HYOU1, IGF2R, ILF3, IMMT, ITGB1, KHDRBS1, KPNA2, KRT18, KRT8, LDHA, LGALS1, LMNA, LMNB1, MAP1B, MAP4, MAPK1, MDH1, MSN, MVP, NCL, NPM1, NUMA1, OTUB1, P4HB, PA2G4, PAFAH1B2, PARK7, PCBP2, PCMT1, PCNA, PDCD6, PDIA3, PDXK, PEA15, PEBP1, PHB2, PKM, PPIA, PPP1CA, PPP2CA, PPP2R1A, PRDX1, PRDX2, PRDX4, PRDX5, PRKDC, PRMT5, PRPF19, PSMB1, PTMA, QARS, RAD23B, RANBP1, RBM3, RHOA, RPL10, RPLP0, RPS24, RPS3, RPS3A, RRBP1, RTN4, S100A6, SERPINH1, SET, SFN, SFPQ, SH3BGRL3, SLC25A5, SLC25A6, SND1, SOD1, SPTBN1, SQSTM1, SSRP1, STAT1, STIP1, STMN1, STOML2, TAGLN2, TCP1, TFRC, TNC, TRIM28, TXNDC5, UBA1, UCHL1, VCL, VCP, VDAC1, VDAC2, VIM, XPO1, XRCC5, XRCC6, YBX1, YBX3, YWHAB, YWHAE, YWHAQ, YWHAZ | |||||
| Cell death and survival | Necrosis | 1.65 × 10−30 | 1.232 | AARS, ACAT1, ACLY, AHSA1, AKR1B1, ALDOA, ANXA1, ANXA2, AP2B1, APEX1, ARHGDIA, ATP1A1, ATP5A1, B2M, BSG, CALR, CAPN1, CAPN2, CAPNS1, CAPRIN1, CAST, CAV1, CCT2, CCT3, CCT4, CCT5, CCT6A, CCT7, CCT8, CD44, CD59, CDK1, CLIC4, CSE1L, CTNNA1, CTSB, CTTN, CYR61, DDX3X, DNM1L, DYNC1H1, EEF1A1, EEF1D, EIF2S1, EIF3B, EIF3C, EIF3I, EIF6, ENO1, EZR, FASN, FDPS, FKBP1A, FKBP4, FLNA, FLNB, FN1, FUS, G6PD, GAPDH, GLO1, GLUD1, GML, GNAI2, GNB2L1, GPI, GSTP1, HADHA, HDGF, HINT1, HIST1H1C, HLA-B, HMGB1, HNRNPA1, HNRNPK, HSD17B10, HSP90AA1, HSP90AB1, HSP90B1, HSPA4, HSPA5, HSPA8, HSPA9, HSPB1, HSPD1, HYOU1, IGF2R, ILF2, IMMT, ITGB1, KHDRBS1, KPNA2, KPNB1, KRT18, KRT8, LDHA, LGALS1, LMNA, LMNB1, LONP1, MAP1B, MAP4, MAPK1, MCM7, MDH1, MSN, MVP, NCL, NPM1, OTUB1, P4HB, PA2G4, PARK7, PCBP2, PCNA, PDIA3, PEA15, PEBP1, PKM, PLEC, PPIA, PPP1CA, PPP2CA, PPP2R1A, PRDX1, PRDX2, PRKDC, PRMT5, PRPF19, PSMB1, PSMD2, PTMA, RAD23B, RAN, RBM3, RHOA, RPL10, RPLP0, RPS24, RPS3, RPS3A, RPSA, RTN4, S100A6, SDHA, SERPINH1, SET, SF3A1, SFN, SFPQ, SLC25A5, SLC25A6, SND1, SOD1, SPTBN1, SQSTM1, SSRP1, STAT1, STIP1, STMN1, STOML2, TAGLN2, TCP1, TFRC, TPP1, TRIM28, TUBB, TUBB6, TUFM, UBA1, UBE2L3, UCHL1, VAPA, VCP, VDAC1, VDAC2, VIM, XPO1, XRCC5, XRCC6, YBX1, YWHAB, YWHAE, YWHAG, YWHAH, YWHAQ, YWHAZ, ZYX | 188 |
| Infectious disease | Viral infection | 2.33 × 10−30 | −1.968 | ACTN1, ACTR2, ANXA1, ANXA2, ANXA5, ANXA6, AP1B1, AP2B1, ARF1, ARPC1B, ATP5B, B2M, BSG, BTF3, C14orf166, CAV1, CCT2, CD44, CLIC4, CLTC, COPA, COPB1, COPB2, COPG1, CSE1L, CTSB, DDX3X, DDX5, DDX6, DHX9, DNAJA1, DNAJA2, DNM1L, EEF1A1, EIF2S1, EIF3C, EIF3I, FASN, FDPS, FKBP1A, FLNA, FN1, G3BP1, GANAB, GAPDH, GML, GPI, H3F3A/H3F3B, HIST1H1C, HLA-A, HLA-B, HMGB1, HNRNPH1, HNRNPK, HNRNPM, HNRNPU, HSP90AA1, HSP90AB1, HSP90B1, HSPA5, HSPA9, HSPD1, IFITM3, IGF2R, ILF3, IMPDH2, ITGB1, KHDRBS1, KHSRP, KPNB1, KRT18, KRT8, LGALS1, LMNA, LONP1, MAP4, MAPK1, MVP, NCL, NPM1, PCBP2, PDIA3, PDIA6, PDXK, PGM1, PICALM, PLIN3, PLOD2, PPIA, PRDX1, PRDX2, PSMA1, PSMA2, PSMA5, PSMA7, PSMC4, PSMD2, PSME2, PTGES3, PYGL, RAB11A, RAB1B, RAB7A, RAN, RANBP1, RHOA, RNH1, RPL10A, RPL12, RPL18, RPL3, RPL5, RPS10, RPS27A, RPS5, RPSA, SAE1, SF3A1, SF3B1, SFN, SFPQ, | 142 |
| SNRPD3, SPTAN1, SPTBN1, SSRP1, STAT1, STIP1, SUPT16H, TAGLN2, TALDO1, TFRC, TKT, TMPO, TUBB, TUBB4B, TWF1, UAP1, UBE2L3, UQCRC1, XPO1, YBX1, ZYX | |||||
| Cell death and survival | Cell death of tumor cell lines | 1.13 × 10−27 | 0.949 | AHSA1, AKR1B1, ANXA2, APEX1, ARHGDIA, ATP1A1, ATP5A1, B2M, CALR, CAPN1, CAPN2, CAPNS1, CAST, CAV1, CCT2, CCT3, CCT4, CCT5, CCT6A, CCT7, CCT8, CD44, CD59, CDK1, CSE1L, CTSB, CTTN, CYR61, DDX3X, DNM1L, DYNC1H1, EIF2S1, ENO1, EZR, FASN, FKBP1A, FLNB, FN1, G6PD, GAPDH, GLO1, GML, GNB2L1, GPI, GSTP1, HINT1, HNRNPA1, HNRNPK, HSP90AB1, HSPA4, HSPA5, HSPA8, HSPA9, HSPB1, HSPD1, IGF2R, ILF2, IMMT, ITGB1, KHDRBS1, KPNA2, KPNB1, KRT18, LGALS1, LMNA, LMNB1, LONP1, MAPK1, MCM7, MSN, MVP, NCL, NPM1, OTUB1, P4HB, PA2G4, PARK7, PCBP2, PCNA, PEA15, PEBP1, PKM, PPIA, PPP1CA, PPP2CA, PPP2R1A, PRDX1, PRDX2, PRKDC, PRMT5, PRPF19, PSMD2, PTMA, RAD23B, RHOA, RPLP0, RPS24, RPS3, RTN4, S100A6, SF3A1, SFN, SFPQ, SLC25A5, SLC25A6, SND1, SOD1, SQSTM1, SSRP1, STAT1, STMN1, STOML2, TAGLN2, TCP1, TFRC, TRIM28, TUBB6, TUFM, UBE2L3, UCHL1, VCP, VDAC1, VDAC2, XPO1, XRCC5, XRCC6, YBX1, YWHAE, YWHAG, YWHAH, YWHAZ | 131 |
| Protein synthesis | Translation | 6.09 × 10−25 | 0.792 | APEX1, CALR, CAPRIN1, DDX3X, DDX6, DHX9, EEF1A1, EEF1B2, EEF2, EIF2S1, EIF3B, EIF3C, EIF3I, EIF3M, EIF4A1, EIF4G1, EIF4H, EIF5A, EPRS, FN1, GAPDH, GNB2L1, HNRNPK, HSPA5, HSPB1, ILF3, ITGB1, MARS, NACA, NCL, PABPC1, PCBP1, PPP1CA, PTBP1, RBM3, RPL23, RPS24, RPS3, RPS3A, RPS4X, RPS5, RPS7, RRBP1, SYNCRIP, TUFM, WARS, YBX1 | 47 |
| Protein synthesis | Translation of protein | 2.00 × 10−23 | 0.860 | APEX1, CALR, CAPRIN1, DDX3X, DDX6, DHX9, EEF1A1, EEF1B2, EIF2S1, EIF3B, EIF3C, EIF3I, EIF3M, EIF4A1, EIF4G1, EIF4H, EIF5A, EPRS, FN1, GAPDH, GNB2L1, HNRNPK, HSPA5, HSPB1, ILF3, ITGB1, MARS, NACA, NCL, PABPC1, PCBP1, PPP1CA, PTBP1, RBM3, RPL23, RPS24, RPS3, RPS3A, RPS4X, RPS5, RPS7, RRBP1, SYNCRIP, WARS, YBX1 | 45 |
| Protein synthesis | Expression of protein | 2.95 × 10−22 | 0.229 | ANXA1, APEX1, CALR, CAPRIN1, CAV1, DDX3X, DDX6, DHX9, EEF1A1, EEF1B2, EIF2S1, EIF3B, EIF3C, EIF3I, EIF3M, EIF4A1, EIF4G1, EIF4H, EIF5A, EPRS, FN1, GAPDH, GNB2L1, HNRNPK, HSPA5, HSPB1, ILF3, ITGB1, KHDRBS1, MARS, NACA, NCL, PABPC1, PCBP1, PPM1G, PPP1CA, PTBP1, RBM3, RPL23, RPS24, RPS3, RPS3A, RPS4X, RPS5, RPS7, RRBP1, SOD1, SYNCRIP, WARS, YBX1 | 50 |
| Protein synthesis | Synthesis of protein | 3.25 × 10−22 | −0.265 | ANXA1, APEX1, CALR, CAPRIN1, CAV1, DDX3X, DDX6, DHX9, EEF1A1, EEF1B2, EEF2, EIF2S1, EIF3B, EIF3C, EIF3I, EIF3M, EIF4A1, EIF4G1, EIF4H, EIF5A, EIF6, EPRS, FN1, GAPDH, GNB2L1, HNRNPK, HSPA5, HSPB1, ILF3, ITGB1, KHDRBS1, MAPK1, MARS, NACA, NCL, NPM1, PABPC1, PARK7, PCBP1, PPM1G, PPP1CA, PTBP1, RBM3, RPL23, RPL5, RPS24, RPS3, RPS3A, RPS4X, RPS5, RPS7, RRBP1, SOD1, STIP1, SYNCRIP, TUFM, WARS, YBX1 | 58 |
| Cell death and survival | Apoptosis of tumor cell lines | 3.17 × 10−21 | 0.816 | AHSA1, AKR1B1, ANXA2, ARHGDIA, B2M, CALR, CAPN1, CAPN2, CAPNS1, CAST, CAV1, CCT2, CCT4, CD44, CD59, CDK1, CSE1L, CTSB, CTTN, CYR61, DNM1L, DYNC1H1, ENO1, EZR, FASN, FLNB, FN1, G6PD, GAPDH, GLO1, GML, GNB2L1, GSTP1, HINT1, HNRNPA1, HNRNPK, HSP90AB1, HSPA4, HSPA5, HSPA8, HSPA9, HSPB1, HSPD1, IGF2R, IMMT, ITGB1, KHDRBS1, KPNA2, KRT18, LGALS1, LMNB1, MAPK1, MSN, MVP, NCL, NPM1, OTUB1, P4HB, PA2G4, PARK7, PCBP2, PCNA, PEA15, PEBP1, PPIA, PPP1CA, PPP2CA, PPP2R1A, PRDX1, PRKDC, PRMT5, PRPF19, PTMA, RAD23B, RHOA, RPLP0, RPS24, RPS3, RTN4, S100A6, SFN, SFPQ, SLC25A5, SND1, SOD1, SSRP1, STAT1, | 102 |
| STMN1, STOML2, TAGLN2, TCP1, TFRC, TRIM28, UCHL1, VCP, VDAC1, VDAC2, XPO1, XRCC5, YBX1, YWHAE, YWHAZ | |||||
| Cellular assembly and organization, cellular function and maintenance | Organization of cytoplasm | 4.80 × 10−20 | −1.198 | ACTG1, ACTN4, ACTR2, ACTR3, ALDOA, ANXA1, ARHGDIA, ARPC2, BASP1, BSG, C1QBP, CALR, CANX, CAP1, CAPG, CAPN2, CAPNS1, CAPRIN1, CAPZB, CAST, CAV1, CD44, CDK1, CFL1, CKAP4, CLTC, COPB2, CORO1C, CSRP1, CTTN, CYR61, DBN1, DNM1L, DPYSL2, EEF1A1, EHD1, ENAH, EZR, FASN, FKBP4, FLNA, FLNB, FLNC, FN1, GAPDH, GDI1, HDGF, HEXB, HMGB1, HNRNPK, HSP90AA1, HSPB1, IQGAP1, ITGB1, KIF5B, KPNB1, KRT18, KRT9, LAMB1, LASP1, LONP1, MAP1B, MAP4, MAPK1, MAPRE1, MARCKS, MSN, MYH10, MYH9, NNMT, NPLOC4, NSFL1C, NUMA1, PARK7, PCM1, PDIA3, PFN1, PHGDH, PICALM, PKM, PLEC, PLS3, PPP2CA, PRKCSH, PRKDC, RAB11A, RAN, RANBP1, RHOA, RRBP1, RTN4, SEPT2, SEPT7, SEPT9, SOD1, SPTAN1, SPTBN1, SSRP1, STIP1, STMN1, STOML2, SURF4, TLN1, TMOD3, TNC, TPM3, TPP1, TUBB, TWF2, UCHL1, VAPA, VCL, VIM, XPO1, YBX1, YWHAH | 116 |
| Protein synthesis | Metabolism of protein | 8.33 × 10−20 | 0.064 | ANXA1, APEX1, CALR, CANX, CAPN1, CAPN2, CAPNS1, CAPRIN1, CAST, CAV1, COPG1, CSTB, CTSB, DDX3X, DDX6, DHX9, EEF1A1, EEF1B2, EEF2, EIF2S1, EIF3B, EIF3C, EIF3I, EIF3M, EIF4A1, EIF4G1, EIF4H, EIF5A, EIF6, EPRS, FLNA, FN1, GAPDH, GNB2L1, HEXB, HNRNPK, HSP90B1, HSPA5, HSPB1, HSPD1, ILF3, IPO9, ITGB1, KHDRBS1, LONP1, MAPK1, MARS, MYH9, NACA, NCL, NPEPPS, NPM1, PABPC1, PARK7, PCBP1, PDIA3, PPM1G, PPP1CA, PPP2CA, PSMC4, PSMD2, PTBP1, RBM3, RPL23, RPL5, RPS24, RPS3, RPS3A, RPS4X, RPS5, RPS7, RRBP1, SNX3, SOD1, SQSTM1, STIP1, SYNCRIP, TPP1, TUFM, UBE2L3, UBE2N, UBXN4, UCHL1, VCP, WARS, XPO1, YBX1 | 87 |
| Cancer, organismal injury and abnormalities, reproductive system disease | Mammary tumor | 1.12 × 10−19 | 1.847 | AKR1B1, ALDOA, ALDOC, ALYREF, ANXA1, ARHGDIA, ATP5A1, BSG, C1QBP, CAV1, CBX3, CCT3, CD44, CDK1, CSE1L, CTTN, CYR61, DDX5, DYNC1H1, EEF1A1, EIF3B, EIF3C, EIF4A1, ENAH, ENO1, ETFA, FASN, FDPS, FKBP1A, FLNA, FLNB, FN1, FUS, GCN1L1, GLO1, GSTP1, H2AFY, H3F3A/H3F3B, HDLBP, HLA-A, HLA-B, HNRNPA1, HSP90AA1, HSP90AB1, HSP90B1, HSPA5, HSPD1, ILF3, ITGB1, KHSRP, KPNA2, KRT18, KRT8, LAMB1, LASP1, LGALS1, LMNA, LONP1, MAPRE1, MARS, MCM4, MCM6, MYH10, MYH9, NAP1L4, NCL, NPEPPS, P4HB, PAFAH1B2, PCNA, PDCD6, PFN1, PGK1, PGM1, PKM, PLEC, PRDX1, PRDX5, PRKCSH, PTBP1, PTRF, RANBP1, RHOA, RPL4, RPL6, RPLP0, RPS24, RPS3, RPS4X, S100A6, SF3B1, SFPQ, SHMT2, SLC25A5, SLC25A6, SOD1, STAT1, STIP1, TAGLN, TAGLN2, TCP1, TLN1, TNC, TPI1, TUBA1B, TUBB, TUBB4B, TUBB6, UBA1, UCHL1, VCL, VDAC2, WDR1, XPO1, XRCC5, YWHAH, YWHAQ, YWHAZ, ZYX | 119 |
| Dermatological diseases and conditions | Psoriasis | 2.52 × 10−19 | ANXA1, ANXA2, ARHGDIA, ARPC1B, C1QBP, CALR, CAV1, CBX3, CCT5, CFL1, COPB2, CSTB, CTSB, CYB5R3, DBN1, EEF1A1, EIF2S1, EIF2S2, EIF5A, EIF6, FASN, FKBP1A, FN1, GAPDH, GARS, GSTP1, H2AFY, HSPA5, HSPA8, KPNA2, KPNB1, KRT1, KRT18, LGALS1, MARCKS, OTUB1, P4HB, PCBP2, PCMT1, PCNA, PGAM1, PGD, PKM, PPP2CA, PSME2, PTRF, RAN, RANBP1, SERBP1, SFN, SLC25A5, SPTAN1, STAT1, SYNCRIP, TAGLN, TNC, TUBB, TWF1, UBE2N, VDAC1, YWHAB, YWHAE, YWHAQ | 63 | |
| Infectious disease | Infection of cells | 2.88 × 10−19 | −2.041 | ACTR2, ANXA2, AP1B1, AP2B1, ARF1, ARPC1B, ATP5B, BSG, C1QBP, CAV1, CCT2, CD44, CLTC, COPA, COPB1, COPB2, COPG1, CTSB, DDX3X, DHX9, DNAJA2, EIF3I, FN1, G3BP1, | 78 |
| GANAB, GML, GPI, H3F3A/H3F3B, HMGB1, HNRNPH1, HNRNPK, HNRNPU, HSPA5, HSPA9, IFITM3, IGF2R, ITGB1, KHDRBS1, KPNB1, KRT18, MAP4, PCBP2, PDIA3, PDIA6, PGM1, PICALM, PLOD2, PSMA1, PSMA2, PSMA5, PSMA7, PSMC4, PSME2, PTGES3, RAB1B, RANBP1, RNH1, RPL10A, RPL12, RPL18, RPL3, RPL5, SF3A1, SF3B1, SNRPD3, SPTAN1, SPTBN1, STAT1, STIP1, TAGLN2, TFRC, TWF1, UAP1, UBE2L3, UQCRC1, XPO1, YBX1, ZYX | |||||
| Infectious disease | Replication of virus | 5.92 × 10−19 | −0.358 | ANXA5, ANXA6, ATP5B, B2M, BTF3, C14orf166, CAV1, CLIC4, COPA, COPB1, COPB2, COPG1, CSE1L, DDX3X, DDX5, DDX6, DHX9, DNAJA1, EEF1A1, EIF2S1, EIF3C, FASN, FDPS, FKBP1A, G3BP1, HMGB1, HNRNPM, HSP90AA1, HSP90AB1, HSP90B1, HSPD1, IFITM3, ILF3, KPNB1, KRT8, LONP1, MAPK1, MVP, NCL, NPM1, PCBP2, PLIN3, PPIA, PRDX1, PRDX2, PSMA1, PSMD2, RAB11A, RPS10, RPS27A, RPS5, RPSA, SAE1, SF3A1, SF3B1, SFPQ, SSRP1, STAT1, TMPO, TUBB, XPO1, YBX1 | 62 |
| Neurological disease | Progressive motor neuropathy | 9.90 × 10−19 | ANXA1, ANXA2, ANXA5, CAPN2, CAPZB, CCT2, COPE, CSE1L, CYR61, DBI, EEF1A1, EIF4A1, EIF4G1, ENO2, EZR, FASN, FUS, GSTP1, H3F3A/H3F3B, HLA-B, HN1, HNRNPA1, HSPA5, HSPB1, IMPDH2, LDHA, LDHB, MAP1B, MAPK1, MDH1, NNMT, PARK7, PCNA, PDIA3, PEBP1, PFKP, PFN1, PGK1, PRDX2, PSMC1, PTGES3, RAN, RHOA, RPL3, RPL5, RPS3A, RPS4X, RTN4, S100A6, SLC25A6, SOD1, STMN1, TFRC, TUBA1B, UCHL1, VCP, VIM, VPS35 | 58 | |
| Cellular assembly and organization, cellular function and maintenance | Organization of cytoskeleton | 1.01 × 10−18 | −1.190 | ACTG1, ACTN4, ACTR2, ACTR3, ALDOA, ANXA1, ARHGDIA, ARPC2, BASP1, BSG, C1QBP, CALR, CANX, CAP1, CAPG, CAPN2, CAPNS1, CAPRIN1, CAPZB, CAST, CAV1, CD44, CDK1, CFL1, CKAP4, COPB2, CORO1C, CSRP1, CTTN, CYR61, DBN1, DNM1L, DPYSL2, EEF1A1, EHD1, ENAH, EZR, FASN, FKBP4, FLNA, FLNB, FLNC, FN1, GAPDH, GDI1, HDGF, HMGB1, HNRNPK, HSP90AA1, HSPB1, IQGAP1, ITGB1, KIF5B, KPNB1, KRT18, KRT9, LAMB1, LASP1, MAP1B, MAP4, MAPK1, MAPRE1, MARCKS, MSN, MYH10, MYH9, NNMT, NUMA1, PCM1, PDIA3, PFN1, PHGDH, PICALM, PKM, PLEC, PLS3, PPP2CA, PRKCSH, PRKDC, RAB11A, RAN, RANBP1, RHOA, RRBP1, RTN4, SEPT2, SEPT7, SEPT9, SOD1, SPTAN1, SPTBN1, SSRP1, STIP1, STMN1, TLN1, TMOD3, TNC, TPM3, TUBB, TWF2, UCHL1, VAPA, VCL, VIM, XPO1, YBX1, YWHAH | 107 |
| Post-translational modification, protein folding | Folding of protein | 1.21 × 10−18 | 0.988 | AARS, AIP, B2M, CALR, CANX, CCT4, DNAJA1, DNAJA2, ERP29, FKBP1A, FKBP4, HSP90AA1, HSP90AB1, HSPA5, HSPA8, HSPB1, HSPD1, P4HB, PDIA6, PPIA, PRDX4, RAB7A, TBCA, TCP1 | 24 |
| Neurological disease, psychological disorders | Disorder of basal ganglia | 2.74 × 10−18 | ACAT1, AHCY, ANXA2, B2M, BASP1, CAPNS1, CAPZB, CAV1, CD44, COPE, CSE1L, CYC1, DDX1, DNAJA1, EEF1A1, EIF4G1, ENO2, FKBP4, GAPDH, GPI, GSTP1, H3F3A/H3F3B, HADH, HINT1, HLA-B, HNRNPU, HSP90AA1, HSPA5, HSPA8, HSPB1, LAMB1, LDHA, LDHB, MAP1B, MDH1, MYL12A, NPM1, PARK7, PCMT1, PCNA, PDE6H, PDLIM1, PEBP1, PFN1, PGK1, PKM, PLOD2, PPIA, PRDX2, PSMC1, PSME1, PTGES3, RAB11A, RAN, RANBP1, RPL3, RPS3A, RPS4X, RPSA, RTN4, SDHA, SLC25A6, SOD1, STMN1, TPI1, TPM3, TUBA1B, TUBB4B, UCHL1, UQCRC1, VIM, VPS35, XRCC6, YWHAZ | 74 | |
| Cellular assembly and organization, cellular function and maintenance | Microtubule dynamics | 4.81 × 10−18 | −0.705 | ACTG1, ACTN4, ACTR2, ACTR3, ARPC2, BASP1, BSG, C1QBP, CANX, CAP1, CAPG, CAPN2, CAPNS1, CAPRIN1, CAPZB, CAST, CAV1, CD44, CDK1, CFL1, CKAP4, COPB2, CSRP1, CTTN, CYR61, DBN1, DNM1L, DPYSL2, EEF1A1, EHD1, ENAH, EZR, FASN, FKBP4, FLNA, FN1, GAPDH, GDI1, HDGF, HMGB1, HNRNPK, HSP90AA1, HSPB1, IQGAP1, ITGB1, KIF5B, KPNB1, KRT18, LAMB1, LASP1, MAP1B, MAP4, MAPK1, MAPRE1, MARCKS, MSN, MYH10, MYH9, NNMT, NUMA1, PCM1, PDIA3, PFN1, PHGDH, PICALM, PKM, PLEC, PPP2CA, PRKCSH, RAB11A, RAN, RANBP1, RHOA, RRBP1, RTN4, SEPT2, SEPT7, SEPT9, SOD1, SPTBN1, SSRP1, STIP1, STMN1, TLN1, TNC, TPM3, TUBB, TWF2, UCHL1, VAPA, VCL, VIM, XPO1, YBX1, YWHAH | 95 |
| Infectious disease | Replication of RNA virus | 4.87 × 10−18 | −0.056 | ANXA5, ANXA6, B2M, BTF3, C14orf166, CAV1, CLIC4, COPA, COPB1, COPB2, COPG1, CSE1L, DDX3X, DDX5, DDX6, DHX9, DNAJA1, EEF1A1, EIF2S1, EIF3C, FASN, FDPS, G3BP1, HMGB1, HNRNPM, HSP90AA1, HSP90AB1, HSPD1, IFITM3, ILF3, KPNB1, LONP1, MAPK1, MVP, NCL, NPM1, PCBP2, PLIN3, PPIA, PRDX1, PRDX2, PSMA1, PSMD2, RAB11A, RPS10, RPS27A, RPS5, RPSA, SAE1, SF3A1, SF3B1, SFPQ, STAT1, TMPO, TUBB, XPO1, YBX1 | 57 |
| Infectious disease | Infection by RNA virus | 7.40 × 10−18 | −2.209 | ACTR2, ANXA1, ANXA2, AP1B1, AP2B1, ARF1, ARPC1B, ATP5B, B2M, BSG, CCT2, CD44, CLTC, COPA, COPB1, COPB2, COPG1, CTSB, DDX3X, DHX9, DNAJA2, EIF3I, FDPS, FN1, G3BP1, GANAB, GML, GPI, H3F3A/H3F3B, HMGB1, HNRNPH1, HNRNPK, HNRNPU, HSPA5, HSPA9, IFITM3, IMPDH2, ITGB1, KHDRBS1, KPNB1, MAP4, PCBP2, PDIA3, PDIA6, PGM1, PICALM, PLOD2, PSMA1, PSMA2, PSMA5, PSMA7, PSMC4, PSME2, PTGES3, RAB1B, RANBP1, RHOA, RNH1, RPL10A, RPL12, RPL18, RPL3, RPL5, SF3A1, SF3B1, SNRPD3, SPTAN1, SPTBN1, STAT1, STIP1, TAGLN2, TFRC, TWF1, UAP1, UBE2L3, UQCRC1, XPO1, YBX1, ZYX | 79 |
| Neurological disease, skeletal and muscular disorders | Neuromuscular disease | 2.20 × 10−17 | ACAT1, AHCY, ANXA1, ANXA2, B2M, BASP1, CAPNS1, CAPZB, CD44, COPE, CSE1L, CYC1, DDX1, DNAJA1, EEF1A1, EIF4G1, ENO2, FKBP1A, FKBP4, GAPDH, GPI, GSTP1, H3F3A/H3F3B, HADH, HINT1, HLA-B, HNRNPA1, HNRNPU, HSP90AA1, HSPA5, HSPA8, HSPB1, IMPDH2, LAMB1, LDHA, LDHB, MAP1B, MDH1, MYL12A, NPM1, PARK7, PCMT1, PCNA, PDE6H, PDIA3, PDLIM1, PEBP1, PFN1, PGK1, PKM, PLOD2, PPIA, PRDX2, PSMC1, PSME1, PTGES3, RAB11A, RAN, RANBP1, RPL3, RPL5, RPS3A, RPS4X, RPSA, RTN4, SDHA, SLC25A6, SOD1, STMN1, TPI1, TPM3, TUBA1B, UCHL1, UQCRC1, VIM, VPS35, XRCC6, YWHAZ | 78 | |
| Cancer, organismal injury and abnormalities, reproductive system disease | Breast cancer | 6.17 × 10−17 | AKR1B1, ALDOA, ALDOC, ANXA1, ARHGDIA, ATP5A1, BSG, C1QBP, CAV1, CBX3, CCT3, CD44, CDK1, CSE1L, CYR61, DDX5, DYNC1H1, EEF1A1, EIF3B, EIF3C, EIF4A1, ENAH, ENO1, ETFA, FASN, FDPS, FKBP1A, FLNA, FLNB, FN1, FUS, GCN1L1, GLO1, GSTP1, H2AFY, H3F3A/H3F3B, HLA-A, HLA-B, HNRNPA1, HSP90AA1, HSP90AB1, HSP90B1, HSPD1, ILF3, ITGB1, KHSRP, KPNA2, KRT8, LAMB1, LASP1, LGALS1, LMNA, LONP1, MAPRE1, MARS, MCM6, MYH10, MYH9, NAP1L4, NCL, NPEPPS, P4HB, PAFAH1B2, PCNA, PDCD6, PFN1, PGK1, PGM1, PKM, PLEC, PRDX1, PRDX5, PRKCSH, PTBP1, PTRF, RANBP1, RPL4, RPL6, RPLP0, RPS24, RPS3, S100A6, SF3B1, SFPQ, SHMT2, SLC25A5, SLC25A6, SOD1, STAT1, TAGLN, TAGLN2, TCP1, TLN1, TNC, TPI1, TUBA1B, TUBB, TUBB4B, TUBB6, UBA1, VCL, VDAC2, WDR1, XPO1, YWHAH, YWHAQ, YWHAZ, ZYX | 108 | |
| Neurological disease | Movement disorders | 1.91 × 10−16 | 0.283 | ACAT1, AHCY, ANXA2, B2M, BASP1, CANX, CAPNS1, CAPZB, CAV1, CD44, COPE, CSE1L, CSTB, CTSA, CTSB, CYC1, DDX1, DNAJA1, EEF1A1, EEF2, EIF4G1, ENO2, FDPS, FKBP1A, FKBP4, FUS, GAPDH, GPI, GSTP1, H3F3A/H3F3B, HADH, HEXB, HINT1, HLA-B, HNRNPU, HSP90AA1, HSPA5, HSPA8, HSPB1, IGF2R, LAMB1, LDHA, LDHB, LMNA, MAP1B, MDH1, MYL12A, NPM1, PARK7, PCMT1, PCNA, PDE6H, PDLIM1, PEBP1, PFN1, PGK1, PKM, PLOD2, PPIA, PRDX2, PSMC1, PSME1, PTGES3, RAB11A, RAN, RANBP1, RPL3, RPS3A, RPS4X, RPSA, RTN4, SDHA, SLC25A6, SOD1, SQSTM1, STMN1, TPI1, TPM3, TPP1, TUBA1B, TUBB4B, UCHL1, UQCRC1, VIM, VPS35, XRCC6, YWHAZ | 87 |
| Cancer | Breast or ovarian cancer | 2.54 × 10−16 | AKR1B1, ALDOA, ALDOC, ANXA1, ARHGDIA, ATIC, ATP5A1, B2M, BSG, C1QBP, CAV1, CBX3, CCT3, CCT5, CD44, CDK1, CSE1L, CYR61, DDX5, DYNC1H1, EEF1A1, EIF3B, EIF3C, EIF4A1, ENAH, ENO1, ETFA, FASN, FDPS, FKBP1A, FLNA, FLNB, FN1, FUS, GAPDH, GCN1L1, GLO1, GSTP1, H2AFY, H3F3A/H3F3B, HLA-A, HLA-B, HN1, HNRNPA1, HSP90AA1, HSP90AB1, HSP90B1, HSPD1, ILF3, ITGB1, KHSRP, KPNA2, KRT8, LAMB1, LASP1, LGALS1, LMNA, LONP1, MAPRE1, MARS, MCM6, MYH10, MYH9, NAP1L4, NCL, NPEPPS, P4HB, PAFAH1B2, PCNA, PDCD6, PDIA3, PEA15, PFN1, PGK1, PGM1, PKM, PLEC, PPP2R1A, PRDX1, PRDX5, PRKCSH, PTBP1, PTRF, RANBP1, RPL4, RPL6, RPLP0, RPS24, RPS3, S100A6, SF3B1, SFPQ, SHMT2, SLC25A5, SLC25A6, SOD1, SSRP1, STAT1, TAGLN, TAGLN2, TCP1, TLN1, TNC, TPI1, TUBA1B, TUBB, TUBB4B, TUBB6, UBA1, VCL, VDAC2, VIM, WDR1, XPO1, YWHAH, YWHAQ, YWHAZ, ZYX | 118 | |
| Cellular movement | Cell movement | 4.44 × 10−15 | −1.559 | ACTN4, ACTR2, ACTR3, AHCY, AKR1B1, ALDOA, ANXA1, ANXA2, ANXA5, ARF1, ARHGDIA, ARPC1B, ARPC2, BSG, C1QBP, CALR, CAP1, CAPG, CAPN1, CAPN2, CAPNS1, CAST, CAV1, CD44, CD59, CDK1, CFL1, CLIC4, CORO1C, CRIP2, CSE1L, CTNNA1, CTSB, CTTN, CYR61, DBN1, DDX3X, DNAJA1, DPYSL2, EHD1, ENAH, EZR, FASN, FKBP4, FLNA, FLNB, FLNC, FN1, G6PD, GDI1, GNAI2, GNB2L1, GPI, HARS, HDGF, HLA-A, HMGB1, HNRNPA2B1, HNRNPK, HNRNPL, HSP90AA1, HSP90AB1, HSP90B1, HSPA5, HSPB1, HSPD1, HYOU1, IGF2R, ILF3, IQGAP1, ITGB1, KHDRBS1, KPNA2, KRT8, LAMB1, LASP1, LGALS1, LMNA, LMNB1, MAP1B, MAPK1, MAPRE1, MARCKS, MCM3, MCM7, MSN, MYH10, MYH9, MYL12A, NACA, NCL, NPM1, PA2G4, PARK7, PDCD6, PEBP1, PFN1, PHB2, PKM, PLEC, PPIA, PRDX1, PRDX2, PRMT5, PTMA, RHOA, RNH1, RPSA, RTN4, S100A6, SEPT7, SEPT9, SERPINH1, SFN, SOD1, SQSTM1, STAT1, STMN1, TAGLN2, TARS, TLN1, TMOD3, TNC, TPM3, VCL, VCP, VIM, WARS, YBX1, YWHAE, YWHAZ, ZYX | 132 |
| Dermatological diseases and conditions | Chronic psoriasis | 5.65 × 10−15 | C1QBP, CCT5, COPB2, CSTB, CTSB, EIF2S1, EIF2S2, EIF5A, GARS, GSTP1, HSPA8, KPNA2, KPNB1, MARCKS, PCMT1, PGAM1, PGD, PKM, PPP2CA, PSME2, RANBP1, SLC25A5, STAT1, TAGLN, TWF1, UBE2N | 26 | |
| Nucleic acid metabolism, small molecule biochemistry | Metabolism of nucleoside triphosphate | 1.02 × 10−14 | −2.160 | ACLY, ALDOA, ANXA1, ATP1A1, ATP5A1, ATP5B, ATP5H, CAV1, CCT8, CDK1, CTPS1, DDX1, DDX39B, DDX3X, DDX5, DHX9, DNM1L, G3BP1, HMGB1, HSP90AA1, HSPA8, HSPD1, LONP1, MCM7, MYH10, MYH9, NNMT, OLA1, PKM, PSMC1, PSMC4, SLC25A5, SOD1, VCP, VDAC1 | 35 |
| Neurological disease, psychological disorders, skeletal and muscular disorders | Parkinson’s disease | 1.16 × 10−14 | ANXA2, CAPZB, COPE, CSE1L, EEF1A1, EIF4G1, ENO2, GSTP1, H3F3A/H3F3B, HLA-B, HSPB1, LDHA, LDHB, MAP1B, MDH1, PARK7, PCNA, PEBP1, PGK1, PRDX2, PSMC1, PTGES3, RAN, RPL3, RPS3A, RPS4X, RTN4, SLC25A6, SOD1, STMN1, TUBA1B, UCHL1, VIM, VPS35 | 34 | |
| Cellular development, cellular growth and proliferation | Proliferation of tumor cell lines | 3.28 × 10−14 | −0.292 | ACAT1, ACLY, ACTN4, AHSA1, ALDOA, ANXA1, ANXA2, ANXA6, APEX1, ARF1, BSG, C1QBP, CACYBP, CALR, CAV1, CD44, CD59, CDK1, CSE1L, CTSB, CTTN, CYR61, DDX5, DYNC1H1, EEF1A1, EEF1B2, EIF3C, EIF4A1, EIF5A, EZR, FASN, FLNA, FN1, FUS, G6PD, GAPDH, GNB2L1, H2AFY, HDGF, HMGB1, HNRNPA1, HNRNPA2B1, HNRNPK, HSP90AA1, HSPA5, IGF2R, ILF3, IMMT, IPO7, IQGAP1, ITGB1, KPNA2, KRT8, LGALS1, MAP1B, MAPK1, MAPRE1, MCM7, NASP, NCL, NPM1, NUMA1, PA2G4, PCNA, PDIA3, PEBP1, PFKP, PFN1, PKM, PPP2R1A, PRDX2, PRKCSH, PRMT5, PSMA4, PTBP1, PTMA, RAN, RHOA, S100A6, SEPT9, SET, SFN, SFPQ, SLC25A6, SND1, SOD1, SPTAN1, STAT1, STMN1, TAGLN2, TCP1, TFRC, TRIM28, TUBB, UBA1, UCHL1, XRCC5, XRCC6, YBX1, YWHAG, YWHAQ | 101 |
| Molecular transport, protein trafficking | Transport of protein | 3.57 × 10−14 | 0.362 | AP1B1, ARF1, CALR, CAV1, CFL1, CSE1L, CTSA, DNAJA1, DNAJA2, EHD1, EIF5A, ERP29, GDI2, HSPA9, IGF2R, IPO5, IPO7, IPO9, KPNA2, KPNB1, MAP1B, MYH9, NPM1, PDCD6, PDIA3, PICALM, RAB11A, RAB7A, RAN, SNX3, SOD1, SPTBN1, SQSTM1, TNPO1, VAPA, XPO1, YWHAH | 37 |
| Immunological disease | Allergy | 7.01 × 10−14 | AHCY, AHNAK, ALDOA, ANXA1, ANXA5, CAPG, CAPN1, CAPZB, CD44, CFL1, CNN3, CPNE1, DPYSL2, EEF1A1, ENO1, FKBP1A, FKBP4, FLNA, GAPDH, H3F3A/H3F3B, HN1, HNRNPR, HSPA5, IDI1, KRT1, LGALS1, MSN, MYH9, P4HB, PDIA3, PHGDH, PPIA, PRDX1, STAT1, TFRC, TKT, TPI1, TPM3, TUBB, VCL, VCP, VDAC2, XRCC6, ZYX | 44 | |
| Cellular movement | Cell movement of tumor cell lines | 7.21 × 10−14 | 0.345 | ACTN4, ANXA1, ANXA2, ARF1, ARPC1B, ARPC2, BSG, C1QBP, CALR, CAP1, CAPN1, CAPN2, CAPNS1, CAV1, CD44, CSE1L, CTSB, CTTN, CYR61, DPYSL2, ENAH, EZR, FLNA, FLNB, FLNC, FN1, GDI1, GNB2L1, GPI, HDGF, HNRNPA2B1, HNRNPK, HSP90AA1, ILF3, IQGAP1, ITGB1, KHDRBS1, KPNA2, KRT8, LASP1, LGALS1, MAPK1, MSN, MYH10, MYH9, NCL, PA2G4, PEBP1, PHB2, PPIA, PRDX2, PRMT5, RHOA, SEPT9, SFN, SQSTM1, STAT1, STMN1, TAGLN2, TLN1, TNC, TPM3, VCL, VCP, VIM, YBX1, ZYX | 67 |
| Cell death and survival | Cell survival | 1.06 × 10−13 | −1.997 | ACLY, ANXA5, AP2B1, APEX1, ATP5H, B2M, CALR, CAPN2, CAV1, CD44, CD59, CDK1, COPB2, CSE1L, CTSB, CYR61, DDX3X, DDX5, DHX9, DNM1L, EEF2, EIF2S1, EIF3C, EIF4A1, EIF4G1, EZR, FLNA, FN1, GLUD1, GNB2L1, GSTP1, HDGF, HINT1, HMGB1, HNRNPU, HSD17B10, HSP90AB1, HSP90B1, HSPA4, HSPA5, HSPB1, HSPD1, HYOU1, IGF2R, IQGAP1, ITGB1, LDHA, LMNA, MAPK1, MVP, P4HB, PARK7, PCNA, PDIA3, PEA15, PKM, PPIA, PPM1G, PPP1CA, PPP2CA, PPP2R1A, PRDX2, PRKDC, PRPF19, PSMA1, PSMA4, PSMA7, PSMC4, RAB11A, RHOA, RPL27, RPSA, S100A6, SET, SF3A1, SFN, SHMT2, SND1, SNRPD1, SOD1, SQSTM1, STAT1, STMN1, TCP1, TRIM28, TUBB, TXNDC5, UCHL1, VCL, VCP, VDAC1, VIM, XPO1, XRCC5, YBX1, YWHAZ | 96 |
| Cellular movement | Migration of cells | 1.22 × 10−13 | −2.069 | ACTN4, ACTR3, AHCY, ALDOA, ANXA1, ANXA2, ANXA5, ARF1, ARHGDIA, ARPC2, BSG, C1QBP, CALR, CAP1, CAPG, CAPN2, CAPNS1, CAST, CAV1, CD44, CDK1, CFL1, CLIC4, CORO1C, CRIP2, CSE1L, CTNNA1, CTSB, CTTN, CYR61, DBN1, DDX3X, DPYSL2, EHD1, EZR, FASN, FKBP4, FLNA, FLNB, FLNC, FN1, G6PD, GNAI2, GNB2L1, GPI, HARS, HDGF, HLA-A, HMGB1, HNRNPA2B1, HNRNPK, HNRNPL, HSP90AA1, HSP90AB1, HSP90B1, HSPA5, HSPB1, HSPD1, HYOU1, IGF2R, ILF3, IQGAP1, ITGB1, KHDRBS1, KPNA2, KRT8, LAMB1, LASP1, LGALS1, LMNA, LMNB1, MAP1B, MAPK1, MAPRE1, MARCKS, MCM3, MCM7, MSN, MYH10, MYH9, MYL12A, NACA, NCL, NPM1, PA2G4, PARK7, PDCD6, PFN1, PHB2, PKM, PLEC, PPIA, PRDX1, PRDX2, PRMT5, PTMA, RHOA, RNH1, RPSA, RTN4, S100A6, SEPT7, SFN, SOD1, STAT1, STMN1, TARS, TLN1, TMOD3, TNC, TPM3, VCL, VCP, VIM, WARS, YBX1, YWHAE, YWHAZ, ZYX | 119 |
| Gene expression, protein synthesis | Translation of mRNA | 2.07 × 10−13 | 1.632 | CALR, CAPRIN1, DDX3X, EEF1B2, EIF2S1, EIF3B, EIF3C, EIF3I, EIF3M, EIF4G1, EIF4H, EPRS, GAPDH, GNB2L1, HSPB1, ILF3, NACA, PPP1CA, RBM3, RPL23, RPS24, RPS3A, RPS4X, RPS5, RRBP1, SYNCRIP, WARS | 27 |
| Cancer, respiratory disease | Respiratory system tumor | 3.89 × 10−13 | 0.316 | ACLY, AHNAK, AKR1B1, ALDOA, ALDOC, ALYREF, ANXA1, ANXA2, APEX1, ATIC, B2M, CAPRIN1, CAV1, CD44, CDK1, CYR61, DHX9, EEF1A1, EEF1B2, EIF4A1, ENAH, ENO1, ENO2, EZR, FASN, FDPS, FN1, G3BP1, GNB2L1, GPI, GSTP1, H2AFY, HIST2H2AC, HLA-A, HSP90AA1, HSP90AB1, HSP90B1, HSPD1, IMPDH2, ITGB1, KRT8, LDHA, LGALS1, LOC102724594/U2AF1, MAP4, MSN, MYH9, MYL12A, NACA, NCL, NSFL1C, PABPC1, PCNA, PDCD6, PFKP, PKM, PPIA, PRDX1, PRDX2, PRPF19, RHOA, RPL27, RPL7, RPS11, RPS27A, SERBP1, SFN, SHMT2, SND1, SQSTM1, STAT1, STMN1, TNC, TPI1, TUBB, TUBB4B, VIM, YWHAE, ZYX | 79 |
| Cancer | Malignant neoplasm of thorax | 4.70 × 10−13 | 0.949 | ACLY, AHNAK, ALDOA, ALDOC, ANXA2, APEX1, ATIC, B2M, CAPRIN1, CAV1, CD44, CDK1, CYR61, DHX9, EEF1A1, EEF1B2, EIF4A1, ENAH, ENO1, ENO2, EZR, FASN, FDPS, FN1, G3BP1, GNB2L1, GPI, GSTP1, H2AFY, HIST2H2AC, HLA-A, HSP90AA1, HSP90AB1, HSP90B1, HSPD1, IMPDH2, ITGB1, KRT8, LDHA, LGALS1, LOC102724594/U2AF1, MAP4, MSN, MYH9, MYL12A, NACA, NCL, NSFL1C, PABPC1, PCNA, PDCD6, PFKP, PKM, PPIA, PRDX1, PRDX2, PRKDC, PRPF19, RHOA, RPL27, RPL7, RPS11, RPS27A, SERBP1, SHMT2, SND1, SQSTM1, STAT1, STMN1, TPI1, TUBB, TUBB4B, VIM, XRCC5, XRCC6, YWHAE | 76 |
| Gene expression | Expression of mRNA | 6.12 × 10−13 | 0.865 | CALR, CAPRIN1, DDX3X, EEF1B2, EIF2S1, EIF3B, EIF3C, EIF3I, EIF3M, EIF4G1, EIF4H, EPRS, GAPDH, GNB2L1, HMGB1, HSPB1, ILF3, KHDRBS1, NACA, PFN1, PPP1CA, RBM3, RPL23, RPS24, RPS3A, RPS4X, RPS5, RRBP1, STAT1, SYNCRIP, WARS | 31 |
| Cell morphology, cellular assembly and organization, cellular function and maintenance | Formation of cellular protrusions | 6.14 × 10−13 | −0.837 | ACTN4, ACTR2, ACTR3, ARPC2, BASP1, BSG, C1QBP, CAP1, CAPG, CAPN2, CAPNS1, CAPRIN1, CAPZB, CAST, CAV1, CD44, CFL1, CSRP1, CTTN, CYR61, DBN1, DNM1L, DPYSL2, EEF1A1, EHD1, ENAH, EZR, FASN, FLNA, FN1, GDI1, HMGB1, HNRNPK, HSP90AA1, HSPB1, IQGAP1, ITGB1, KIF5B, LAMB1, LASP1, MAP1B, MARCKS, MSN, MYH10, NNMT, PCM1, PDIA3, PFN1, PHGDH, PICALM, PLEC, PPP2CA, PRKCSH, RAB11A, RANBP1, RHOA, RTN4, SEPT2, SOD1, SPTBN1, STIP1, STMN1, TNC, TPM3, TWF2, UCHL1, VAPA, VCL, VIM, YWHAH | 70 |
| Cancer, respiratory disease | Lung tumor | 7.06 × 10−13 | 0.316 | ACLY, AHNAK, AKR1B1, ALDOA, ALDOC, ALYREF, ANXA2, APEX1, ATIC, B2M, CAPRIN1, CAV1, CD44, CDK1, CYR61, DHX9, EEF1A1, EEF1B2, EIF4A1, ENAH, ENO1, ENO2, EZR, FASN, FDPS, FN1, G3BP1, GNB2L1, GPI, GSTP1, H2AFY, HIST2H2AC, HLA-A, HSP90AA1, HSP90AB1, HSP90B1, HSPD1, IMPDH2, ITGB1, KRT8, LDHA, LGALS1, LOC102724594/U2AF1, MAP4, MSN, MYH9, MYL12A, NACA, NCL, NSFL1C, PABPC1, PCNA, PDCD6, PFKP, PKM, PPIA, PRDX1, PRDX2, PRPF19, RHOA, RPL27, RPL7, RPS11, RPS27A, SERBP1, SHMT2, SND1, SQSTM1, STAT1, STMN1, TNC, TPI1, TUBB, TUBB4B, VIM, YWHAE, ZYX | 77 |
| Organismal survival | Organismal death | 9.82 × 10−13 | 2.742 | ACLY, ACTA1, ACTG1, ACTN4, AIP, ANXA1, APEX1, ARHGDIA, ATP1A1, B2M, BSG, BUB3, C1QBP, CALR, CANX, CAP1, CAPN1, CAPN2, CAPNS1, CAPRIN1, CAPZB, CAST, CAV1, CD44, CD59, CDK1, CFL1, CLIC4, CSE1L, CTSA, CTSB, CTTN, CYR61, DBI, DDX1, DDX5, DHX9, DNM1L, DYNC1H1, EHD1, EIF2S1, EIF3M, EIF4A1, EIF6, ENAH, FASN, FKBP1A, FKBP4, FLNA, FLNB, FLNC, FN1, FUS, G6PD, GNAI2, GSTP1, H3F3A/H3F3B, HADHA, HEXB, HIST1H1C, HMGB1, HNRNPK, HSP90AA1, HSP90AB1, HSP90B1, HSPA5, HYOU1, IGF2R, ILF2, ILF3, IMPDH2, ITGB1, KHDRBS1, KIF5B, KRT1, KRT8, LMNA, LMNB1, LRPPRC, MAP1B, MAPK1, MSN, MYH10, MYH9, NASP, NPM1, NUMA1, PCNA, PDIA3, PEA15, PFN1, PHB2, PHGDH, PICALM, PLEC, PLS3, PPIA, PPP2CA, PRDX1, PRKDC, PRMT5, PRPF19, PSMC1, PSMC4, PTBP1, PTGES3, RAD23B, RHOA, RPL4, RPL6, RPSA, RTN4, SERPINH1, SF3B1, SHMT2, SOD1, SPTBN1, SQSTM1, SSRP1, STAT1, STIP1, TFRC, TKT, TLN1, TPM3, TPP1, TRIM28, UBE2L3, UBE2N, VCL, VCP, VDAC1, VIM, WDR1, XRCC5, XRCC6, YBX1, YBX3, YWHAE | 139 |
| Infectious disease, renal and urological disease | Infection of kidney cell lines | 1.19 × 10−12 | −1.281 | BSG, CTSB, GANAB, HMGB1, HNRNPH1, HSPA5, IFITM3, KHDRBS1, KPNB1, KRT18, MAP4, PCBP2, PGM1, PLOD2, PSMA1, PSMA2, PSMA5, PSMA7, PSMC4, PTGES3, RNH1, RPL10A, RPL12, RPL18, RPL3, RPL5, SF3A1, SF3B1, SNRPD3, TAGLN2, TFRC, UBE2L3, YBX1, ZYX | 34 |
| Inflammatory disease, skeletal and muscular disorders | Inclusion body myopathy | 2.03 × 10−12 | CALR, CANX, HNRNPA1, HNRNPA2B1, HSP90B1, HSPA5, PSMA2, PSMA4, SQSTM1, VCP | 10 | |
| Cancer, respiratory disease | Lung cancer | 2.29 × 10−12 | 0.205 | ACLY, AHNAK, ALDOA, ALDOC, ANXA2, APEX1, ATIC, B2M, CAPRIN1, CAV1, CD44, CDK1, CYR61, DHX9, EEF1A1, EEF1B2, EIF4A1, ENAH, ENO1, ENO2, EZR, FASN, FDPS, FN1, G3BP1, GNB2L1, GPI, GSTP1, H2AFY, HIST2H2AC, HLA-A, HSP90AA1, HSP90AB1, HSP90B1, HSPD1, IMPDH2, ITGB1, KRT8, LDHA, LGALS1, LOC102724594/U2AF1, MAP4, MSN, MYH9, MYL12A, NACA, NCL, NSFL1C, PABPC1, PCNA, PDCD6, PFKP, PKM, PPIA, PRDX1, PRDX2, PRPF19, RHOA, RPL27, RPL7, RPS11, RPS27A, SERBP1, SHMT2, SND1, SQSTM1, STAT1, STMN1, TPI1, TUBB, TUBB4B, VIM, YWHAE | 73 |
| Dermatological diseases and conditions, immunological disease, inflammatory disease, inflammatory response | Atopic dermatitis | 2.95 × 10−12 | AHCY, AHNAK, ANXA1, ANXA5, CAPG, CAPZB, CD44, CFL1, CNN3, CPNE1, DPYSL2, EEF1A1, ENO1, FKBP1A, FKBP4, FLNA, H3F3A/H3F3B, HN1, HNRNPR, IDI1, KRT1, LGALS1, MSN, PHGDH, STAT1, TPI1, TPM3, VCL, VCP, VDAC2, XRCC6, ZYX | 32 | |
| Developmental disorder, hereditary disorder, skeletal and muscular disorders | Distal myopathy | 4.20 × 10−12 | CALR, CANX, FLNC, HSP90B1, HSPA5, MATR3, PSMA2, PSMA4, SQSTM1, VCP | 10 | |
| RNA post-transcriptional modification | Processing of RNA | 4.45 × 10−12 | 0.640 | AARS, AHNAK, C1QBP, DDX39B, DDX5, FUS, HNRNPA1, HNRNPA2B1, HNRNPH1, HNRNPH3, HNRNPK, HNRNPL, HNRNPM, HNRNPU, KHDRBS1, KHSRP, LOC102724594/U2AF1, NPM1, PABPC1, PCBP1, PRPF19, PTBP1, RBM3, RPL5, RPL7, RPS15, RPS24, RPS7, SF3A1, SF3B1, SFPQ, SNRPD1, SYNCRIP, U2AF2 | 34 |
| Nucleic acid metabolism | Metabolism of nucleic acid component or derivative | 7.96 × 10-12 | −1.213 | ACLY, AHCY, ALDOA, ANXA1, ATIC, ATP1A1, ATP5A1, ATP5B, ATP5H, CAV1, CCT8, CDK1, CS, CTPS1, DBI, DDX1, DDX39B, DDX3X, DDX5, DHX9, DNM1L, DPYSL2, FASN, FDPS, G3BP1, G6PD, GMPS, GNAI2, HMGB1, HSP90AA1, HSPA8, HSPD1, IMPDH2, LONP1, MAPK1, MCM7, MDH1, MDH2, MYH10, MYH9, NNMT, OLA1, PGK1, PKM, PPA1, PSMC1, PSMC4, SET, SLC25A5, SOD1, TALDO1, UAP1, VCP, VDAC1 | 54 |
| Infectious disease, organismal injury and abnormalities | Infection of embryonic cell lines | 1.06 × 10-11 | −1.045 | BSG, GANAB, HMGB1, HNRNPH1, HSPA5, IFITM3, KHDRBS1, KPNB1, MAP4, PCBP2, PGM1, PLOD2, PSMA1, PSMA2, PSMA5, PSMA7, PSMC4, PTGES3, RNH1, RPL10A, RPL12, RPL18, RPL3, RPL5, SF3A1, SF3B1, SNRPD3, TAGLN2, TFRC, UBE2L3, YBX1, ZYX | 32 |
| Infectious disease | Infection of epithelial cell lines | 1.06 × 10-11 | −1.045 | BSG, GANAB, HMGB1, HNRNPH1, HSPA5, IFITM3, KHDRBS1, KPNB1, MAP4, PCBP2, PGM1, PLOD2, PSMA1, PSMA2, PSMA5, PSMA7, PSMC4, PTGES3, RNH1, RPL10A, RPL12, RPL18, RPL3, RPL5, SF3A1, SF3B1, SNRPD3, TAGLN2, TFRC, UBE2L3, YBX1, ZYX | 32 |
| Cellular movement | Invasion of cells | 1.64 × 10-11 | −0.711 | ACAT1, AHCY, ANXA1, ANXA2, APEX1, ARHGDIA, BSG, C1QBP, CALR, CAP1, CAPN2, CAPNS1, CAV1, CD44, CDK1, CSE1L, CTSB, CTTN, CYR61, ENAH, EZR, FKBP1A, FLNA, FN1, GNAI2, GNB2L1, GPI, HDGF, HDLBP, HMGB1, HSP90AA1, HSP90AB1, HSP90B1, HSPA5, ILF3, IQGAP1, ITGB1, KRT8, LASP1, LGALS1, MAPK1, MARCKS, MYH10, NPM1, OTUB1, PA2G4, PARK7, PEBP1, PICALM, PKM, PTGES3, RHOA, RNH1, RPSA, S100A6, SEPT9, SQSTM1, STMN1, TAGLN, TAGLN2, VCP, VIM, YWHAQ | 63 |
| Cell death and survival | Cell viability | 3.09 × 10-11 | −1.538 | ACLY, ANXA5, AP2B1, APEX1, ATP5H, B2M, CALR, CAPN2, CAV1, CD44, CD59, CDK1, COPB2, CTSB, CYR61, DDX5, DHX9, DNM1L, EEF2, EIF3C, EIF4A1, EIF4G1, EZR, FLNA, FN1, GLUD1, GNB2L1, GSTP1, HDGF, HINT1, HMGB1, HNRNPU, HSD17B10, HSP90AB1, HSP90B1, HSPA5, HSPB1, HSPD1, HYOU1, IGF2R, IQGAP1, ITGB1, LDHA, LMNA, MAPK1, MVP, P4HB, PARK7, PCNA, PDIA3, PPIA, PPM1G, PPP1CA, PPP2CA, PPP2R1A, PRDX2, PRKDC, PRPF19, PSMA1, PSMA4, PSMC4, RAB11A, RHOA, RPL27, RPSA, S100A6, SF3A1, SFN, SHMT2, SND1, SNRPD1, SOD1, SQSTM1, STAT1, STMN1, TCP1, TRIM28, TXNDC5, UCHL1, VCP, VDAC1, XPO1, XRCC5, YBX1, YWHAZ | 85 |
| Cancer, organismal injury and abnormalities, reproductive system disease | Cervical tumor | 3.68 × 10-11 | ACTR3, ALYREF, ANXA1, ANXA2, ANXA5, ATIC, CTSB, ENO1, GSTP1, H2AFY, HIST1H2BL, HIST2H2AC, HLA-A, HLA-B, HSP90AA1, HSP90AB1, HSP90B1, HSPB1, KRT1, KRT9, MAP4, MCM7, RPS12, TAGLN, TFRC, TPM2, TPM3, TUBB, TUBB4B, XPO1, YWHAE | 31 | |
| Developmental disorder, hereditary disorder, inflammatory disease, neurological disease, skeletal and muscular disorders | Nonaka myopathy | 4.55 × 10-11 | CALR, CANX, HSP90B1, HSPA5, PSMA2, PSMA4, SQSTM1, VCP | 8 | |
| Dermatological diseases and conditions, inflammatory disease, inflammatory response | Dermatitis | 5.49 × 10-11 | AHCY, AHNAK, ANXA1, ANXA5, ARHGDIA, ATP1A1, CALR, CAPG, CAPZB, CD44, CFL1, CNN3, CPNE1, DPYSL2, EEF1A1, ENO1, FKBP1A, FKBP4, FLNA, GSTP1, H3F3A/H3F3B, HN1, HNRNPR, IDI1, KRT1, LGALS1, MSN, PHGDH, SFN, STAT1, TPI1, TPM3, TUBB, TUBB4B, TUBB6, TUBB8, VCL, VCP, VDAC2, XRCC6, ZYX | 41 | |
| Infectious disease | Replication of influenza A virus | 6.00 × 10-11 | −0.322 | ANXA6, B2M, C14orf166, CLIC4, COPA, COPB1, COPB2, COPG1, CSE1L, EEF1A1, EIF3C, FDPS, HSP90AA1, HSPD1, IFITM3, ILF3, KPNB1, LONP1, MAPK1, PSMA1, PSMD2, RPS10, RPS27A, RPS5, RPSA, SAE1, SF3A1, SF3B1, SFPQ, STAT1, TUBB, XPO1 | 32 |
| Cell cycle | M phase | 6.05 × 10-11 | −0.578 | ACTN4, BUB3, CAP1, CAPN2, CDK1, CFL1, FLNA, FN1, GNAI2, ITGB1, LMNA, MAPRE1, MCM4, MCM7, MSN, MYH10, MYH9, NPM1, NUMA1, PFN1, RAB11A, RHOA, SEPT2, SEPT7, SEPT9, SSRP1, XRCC5, YBX1 | 28 |
| DNA replication, recombination, and repair, energy production, nucleic acid metabolism, small molecule biochemistry | Catabolism of ATP | 7.68 × 10-11 | ACLY, ANXA1, ATP1A1, ATP5A1, ATP5B, ATP5H, CCT8, DDX1, DDX39B, DDX3X, DDX5, DHX9, G3BP1, HSP90AA1, HSPA8, LONP1, MCM7, MYH10, MYH9, OLA1, PSMC1, PSMC4, VCP | 23 | |
| Cell death and survival | Cell death of neuroblastoma cell lines | 1.35 × 10-10 | −1.206 | ATP5A1, CAPN1, CAPN2, CAST, CCT2, CCT3, CCT4, CCT5, CCT6A, CCT7, CCT8, ENO1, FKBP1A, GAPDH, HSPA8, ITGB1, PARK7, PEBP1, PRDX2, S100A6, SOD1, SQSTM1, TCP1, YWHAE | 24 |
| Hematological disease, immunological disease, inflammatory disease, inflammatory response, respiratory disease | Allergic pulmonary eosinophilia | 1.55 × 10-10 | ALDOA, ENO1, GAPDH, HSPA5, MYH9, P4HB, PDIA3, PRDX1, TKT, TPI1, TUBB | 11 | |
| Cell morphology | Cell spreading | 1.72 × 10-10 | −0.739 | CAP1, CAPN2, CAST, CD44, CYR61, EHD1, FLNA, FLNB, FLNC, FN1, GNB2L1, IGF2R, IPO9, ITGB1, LGALS1, MAP4, MAPK1, MARCKS, PFN1, PTBP1, RHOA, RRBP1, RTN4, TLN1, TNC, UCHL1, VCL, VIM, YWHAZ, ZYX | 30 |
| Skeletal and muscular disorders | Myopathy | 1.81 × 10-10 | −2.236 | AARS, ACTA1, AHNAK, ATP1A1, BUB3, CALR, CANX, CAST, DYNC1H1, ECHS1, FKBP1A, FLNC, FN1, GARS, HARS, HEXB, HINT1, HLA-A, HMGB1, HNRNPA1, HNRNPA2B1, HSP90B1, HSPA5, HSPB1, ITGB1, LMNA, LRPPRC, MATR3, PLEC, PPP1CA, PPP2CA, PSMA2, PSMA4, RAB7A, RHOA, SDHA, SOD1, SQSTM1, STMN1, TPM2, TPM3, TUBB, TUBB4B, UBA1, VCL, VCP | 46 |
| Molecular transport, protein trafficking | Internalization of protein | 2.17 × 10-10 | 1.900 | AP1B1, AP2B1, CAP1, CFL1, CLTC, DNAJA1, DNAJA2, IPO5, IPO7, IPO9, KPNA2, KPNB1, PDIA3, RAN, RHOA, SPTBN1, XPO1 | 17 |
| Cell death and survival | Cell death of cervical cancer cell lines | 2.24 × 10-10 | −0.340 | ATP1A1, CALR, CCT4, CDK1, CTSB, DNM1L, DYNC1H1, EZR, FLNB, HNRNPK, IMMT, KHDRBS1, KPNB1, KRT18, LMNA, MAPK1, MSN, PCBP2, PKM, PPIA, PPP2R1A, PRDX2, PRKDC, PRPF19, PTMA, RPS24, SOD1, SQSTM1, STAT1, TCP1, UCHL1, VCP, VDAC1, YWHAH | 34 |
| Cellular development, cellular growth and proliferation, connective tissue development and function | Proliferation of fibroblast cell lines | 2.34 × 10-10 | −0.983 | ACAT1, AKR1B1, ALDOA, APEX1, ATIC, CAPRIN1, CAST, CAV1, CLTC, DDX3X, DPY30, EEF1D, EIF3B, EIF3C, EIF3I, EIF6, FN1, G6PD, GNAI2, GNB2L1, HDGF, HINT1, HMGB1, ITGB1, KHDRBS1, LMNA, MAPK1, NPM1, PRPF19, PSMA4, PTMA, RHOA, S100A6, STAT1, STIP1, STMN1, TFRC, TPM3, XRCC6 | 39 |
| Cancer, respiratory disease | Stage 1-2 non-small cell lung cancer | 2.36 × 10-10 | ANXA2, B2M, EEF1A1, EIF4A1, FN1, MYL12A, PPIA, RPL27, RPL7, RPS11, RPS27A, SERBP1 | 12 | |
| Immunological disease | Immediate hypersensitivity | 2.58 × 10-10 | AHCY, AHNAK, ANXA1, ANXA5, CAPG, CAPZB, CD44, CFL1, CNN3, CPNE1, DPYSL2, EEF1A1, ENO1, FKBP1A, FKBP4, FLNA, H3F3A/H3F3B, HN1, HNRNPR, IDI1, KRT1, LGALS1, MSN, PHGDH, PPIA, STAT1, TPI1, TPM3, VCL, VCP, VDAC2, XRCC6, ZYX | 33 | |
| DNA replication, recombination, and repair | Metabolism of DNA | 2.73 × 10-10 | −1.611 | APEX1, BSG, CACYBP, CALR, CAPN2, CAST, CAV1, CDK1, DHX9, ENO1, HMGB1, HNRNPA1, HSD17B10, HSPB1, KPNA2, KRT8, LMNA, MAPK1, MCM7, NAP1L1, NASP, NCL, NPM1, PCM1, PCNA, PEA15, PFN1, PPIA, RAN, RHOA, SET, SOD1, SSRP1, STAT1, SUPT16H, TMPO, XRCC5, XRCC6 | 38 |
| Cellular assembly and organization | Organization of organelle | 3.33 × 10-10 | −1.732 | ALDOA, ANXA2, CAPZB, CAV1, CD44, CFL1, CLTC, CTNNA1, DBN1, DNM1L, DPYSL2, ENAH, FKBP1A, FLNA, FN1, GAPDH, H3F3A/H3F3B, HEXB, ITGB1, KRT18, KRT9, LMNA, LONP1, MAPRE1, MSN, MYH9, NPLOC4, NPM1, NSFL1C, NUMA1, PARK7, PCM1, PLS3, RHOA, RRBP1, SERPINH1, SOD1, SPTBN1, STOML2, SURF4, TPP1, TUBB, VCL, VIM | 44 |
| Cancer, respiratory disease | Carcinoma in lung | 3.34 × 10-10 | 0.555 | AHNAK, ALDOA, ALDOC, ANXA2, APEX1, ATIC, B2M, CAPRIN1, CAV1, CD44, CDK1, DHX9, EEF1A1, EEF1B2, EIF4A1, ENO1, ENO2, EZR, FASN, FDPS, FN1, G3BP1, GNB2L1, GPI, GSTP1, HIST2H2AC, HLA-A, HSP90AA1, HSP90AB1, HSP90B1, HSPD1, IMPDH2, ITGB1, LDHA, LOC102724594/U2AF1, MAP4, MSN, MYH9, MYL12A, NACA, NSFL1C, PABPC1, PCNA, PDCD6, PFKP, PKM, PPIA, PRDX1, PRPF19, RPL27, RPL7, RPS11, RPS27A, SERBP1, STAT1, STMN1, TPI1, TUBB, TUBB4B, VIM, YWHAE | 61 |
| Nucleic acid metabolism, small molecule biochemistry | Metabolism of nucleotide | 4.07 × 10-10 | −1.647 | ACLY, ALDOA, ANXA1, ATP1A1, ATP5A1, ATP5B, ATP5H, CAV1, CCT8, CDK1, CTPS1, DDX1, DDX39B, DDX3X, DDX5, DHX9, DNM1L, FASN, FDPS, G3BP1, G6PD, GMPS, GNAI2, HMGB1, HSP90AA1, HSPA8, HSPD1, IMPDH2, LONP1, MCM7, MDH1, MDH2, MYH10, MYH9, NNMT, OLA1, PKM, PPA1, PSMC1, PSMC4, SET, SLC25A5, SOD1, TALDO1, VCP, VDAC1 | 46 |
| DNA replication, recombination, and repair, nucleic acid metabolism, small molecule biochemistry | Hydrolysis of nucleotide | 4.19 × 10-10 | −1.306 | ATP1A1, CALR, CCT4, CCT5, CDK1, DNM1L, GNAI2, HMGB1, HSPA5, HSPD1, IPO5, MAPK1, RAB7A, RAN, RANBP1, RHOA, STMN1, UBA1, XPO1 | 19 |
| Infectious disease | HIV infection | 4.22 × 10-10 | −1.958 | ANXA2, ARF1, ATP5B, B2M, CCT2, CD44, DDX3X, DHX9, DNAJA2, FDPS, FN1, GANAB, GML, GPI, H3F3A/H3F3B, HMGB1, HNRNPH1, HNRNPU, IMPDH2, ITGB1, KHDRBS1, KPNB1, MAP4, PDIA3, PDIA6, PGM1, PLOD2, PSMA1, PSMA2, PSMA5, PSMA7, PSMC4, PSME2, PTGES3, RAB1B, RANBP1, RNH1, RPL10A, RPL12, RPL18, SF3A1, SF3B1, SNRPD3, SPTAN1, SPTBN1, STIP1, TAGLN2, TFRC, TWF1, UAP1, UBE2L3, UQCRC1, XPO1, YBX1, ZYX | 55 |
| Nucleic acid metabolism, small molecule biochemistry | Metabolism of purine nucleotide | 4.60 × 10-10 | ACLY, ANXA1, ATP1A1, ATP5A1, ATP5B, ATP5H, CCT8, DDX1, DDX39B, DDX3X, DDX5, DHX9, G3BP1, G6PD, HSP90AA1, HSPA8, LONP1, MCM7, MDH1, MDH2, MYH10, MYH9, OLA1, PSMC1, PSMC4, VCP | 26 | |
| Cancer, respiratory disease | Non-small cell lung cancer | 4.91 × 10-10 | AHNAK, ALDOA, ALDOC, ANXA2, APEX1, ATIC, B2M, CAPRIN1, CAV1, CD44, CDK1, DHX9, EEF1A1, EEF1B2, EIF4A1, ENO1, EZR, FASN, FDPS, FN1, G3BP1, GNB2L1, GPI, GSTP1, HIST2H2AC, HLA-A, HSP90AA1, HSP90AB1, HSP90B1, HSPD1, IMPDH2, ITGB1, LDHA, LOC102724594/U2AF1, MAP4, MSN, MYH9, MYL12A, NACA, NSFL1C, PABPC1, PCNA, PFKP, PKM, PPIA, PRDX1, PRPF19, RPL27, RPL7, RPS11, RPS27A, SERBP1, STAT1, STMN1, TPI1, TUBB, TUBB4B, VIM | 58 | |
| Immunological disease | Hypersensitive reaction | 5.39 × 10-10 | 0.404 | AHCY, AHNAK, ANXA1, ANXA5, CALR, CAPG, CAPZB, CD44, CFL1, CNN3, CPNE1, DPYSL2, EEF1A1, ENO1, FKBP1A, FKBP4, FLNA, GARS, H3F3A/H3F3B, HLA-A, HLA-B, HN1, HNRNPR, IDI1, KRT1, LGALS1, MSN, PHGDH, PPIA, PSME2, SOD1, STAT1, TPI1, TPM3, VCL, VCP, VDAC2, XRCC6, ZYX | 39 |
| Cancer, endocrine system disorders, organismal injury and abnormalities | Benign cold thyroid nodule | 1.09 × 10-9 | ANXA5, CALR, CTSB, GSTP1, HSP90AB1, PARK7, PRDX2, PRDX5 | 8 | |
| Cancer, organismal injury and abnormalities, reproductive system disease | Cervical cancer | 1.13 × 10-9 | ACTR3, ANXA1, ANXA2, ANXA5, ATIC, CTSB, ENO1, GSTP1, H2AFY, HIST1H2BL, HIST2H2AC, HSP90AA1, HSP90AB1, HSP90B1, HSPB1, KRT1, KRT9, MAP4, MCM7, RPS12, TAGLN, TFRC, TPM2, TPM3, TUBB, TUBB4B, XPO1, YWHAE | 28 | |
| Cellular assembly and organization, cellular function and maintenance | Organization of filaments | 1.22 × 10-9 | −1.929 | ALDOA, ANXA2, CFL1, DBN1, DPYSL2, ENAH, FKBP1A, FLNA, FN1, GAPDH, ITGB1, KRT18, KRT9, MAPRE1, MSN, NUMA1, PCM1, PLS3, RHOA, RRBP1, SERPINH1, SOD1, TUBB, VIM | 24 |
| Cancer, respiratory disease | Pulmonary metastasis | 1.30 × 10-9 | ATIC, B2M, EEF1A1, EIF4A1, FN1, MYL12A, PPIA, RHOA, RPL27, RPL7, RPS11, RPS27A, YWHAE | 13 | |
| Cellular assembly and organization | Stabilization of filaments | 1.55 × 10-9 | −0.454 | CFL1, COPB2, CTTN, DNM1L, DPYSL2, FLNA, IQGAP1, KRT18, KRT8, MAP1B, MAP4, MAPRE1, NUMA1, PKM, RHOA, SEPT7, STMN1 | 17 |
| Cancer, respiratory disease | Metastatic lung carcinoma | 1.60 × 10-9 | ATIC, B2M, EEF1A1, EIF4A1, FN1, MYL12A, PPIA, RPL27, RPL7, RPS11, RPS27A, YWHAE | 12 | |
| Cancer | Metastasis | 1.88 × 10-9 | −1.645 | ANXA1, ANXA5, ATIC, B2M, C1QBP, CAPG, CAPN2, CAV1, CD44, CSE1L, CTSB, CTTN, CYR61, DPYSL2, EEF1A1, EIF4A1, ENAH, EZR, FASN, FDPS, FKBP1A, FLNA, FN1, FUS, GNB2L1, HMGB1, HSP90AA1, HSP90AB1, HSP90B1, HYOU1, ITGB1, KHDRBS1, KRT18, KRT8, LGALS1, MAPK1, MYL12A, PLOD2, PPIA, PRDX2, RHOA, RNH1, RPL27, RPL7, RPS11, RPS27A, SND1, SQSTM1, STAT1, TUBB, TUBB4B, UBE2N, VIM, YWHAE, YWHAZ | 55 |
| Cellular movement | Migration of tumor cell lines | 1.96 × 10-9 | −0.054 | ACTN4, ANXA1, ANXA2, ARF1, ARPC2, BSG, C1QBP, CAPN2, CAPNS1, CAV1, CD44, CSE1L, CTTN, CYR61, DPYSL2, EZR, FLNA, FLNB, FLNC, FN1, GNB2L1, HDGF, HNRNPA2B1, HNRNPK, HSP90AA1, ILF3, IQGAP1, ITGB1, KHDRBS1, KPNA2, KRT8, LASP1, LGALS1, MAPK1, MSN, MYH10, MYH9, NCL, PHB2, PRDX2, PRMT5, RHOA, SFN, STAT1, STMN1, TNC, TPM3, VCP, VIM, ZYX | 50 |
| Cell morphology, connective tissue development and function | Cell spreading of fibroblast cell lines | 2.05 × 10-9 | 0.201 | CAST, EHD1, FLNA, FLNC, FN1, GNB2L1, IPO9, ITGB1, PTBP1, RTN4, VCL, VIM | 12 |
| Cancer, respiratory disease | Metastatic lung cancer | 2.05 × 10-9 | ATIC, B2M, EEF1A1, EIF4A1, FN1, MYL12A, PPIA, RPL27, RPL7, RPS11, RPS27A, YWHAE | 12 | |
| Gene expression | Expression of RNA | 2.21 × 10-9 | 1.310 | ACTR2, ACTR3, ALYREF, BASP1, BTF3, C14orf166, C1QBP, CALR, CAND1, CAPRIN1, CAV1, CBX3, CD44, CDK1, CSE1L, CYR61, DDX3X, DDX5, DHX9, EEF1B2, EEF1D, EEF2, EIF2S1, EIF3B, EIF3C, EIF3I, EIF3M, EIF4G1, EIF4H, ENO1, EPRS, EZR, FKBP1A, FLNA, FN1, FUBP3, GAPDH, GLO1, GNB2L1, GSTP1, H2AFY, HDGF, HEXB, HINT1, HIST1H1C, HMGB1, HNRNPA1, HNRNPA2B1, HNRNPK, HSPA4, HSPA8, HSPB1, ILF2, ILF3, IQGAP1, KHDRBS1, KPNA2, LGALS1, LMNA, LRPPRC, MAPK1, MATR3, MCM7, MSN, NACA, NONO, NPM1, OTUB1, PA2G4, PARK7, PCNA, PDLIM1, PEA15, PEBP1, PFN1, PHB2, PHGDH, PICALM, PPP1CA, PPP2CA, PRKDC, PRMT5, PTGES3, PTMA, PTRF, RBM3, RHOA, RPL23, RPL6, RPS24, RPS3A, RPS4X, RPS5, RRBP1, SET, SFPQ, SQSTM1, SSRP1, STAT1, SYNCRIP, TAGLN, TMPO, TNC, TRIM28, UBE2L3, VAPA, VIM, WARS, XPO1, XRCC5, XRCC6, YBX1, YBX3, YWHAB, YWHAH, YWHAQ, YWHAZ | 117 |
| Inflammatory response | Inflammation of organ | 2.36 × 10-9 | 0.481 | ACTN4, AHCY, AHNAK, ALDOA, ANXA1, ANXA5, ARHGDIA, ATP1A1, B2M, BSG, CALR, CAPG, CAPZB, CAV1, CD44, CFL1, CNN3, CPNE1, CTSB, CYR61, DDX5, DPYSL2, EEF1A1, ENO1, FKBP1A, FKBP4, FLNA, FUS, GAPDH, GNAI2, GSTP1, H3F3A/H3F3B, HARS, HLA-A, HMGB1, HN1, HNRNPR, HSP90B1, HSPA5, IDI1, IMPDH2, KRT1, KRT18, KRT8, LGALS1, MSN, MYH10, MYH9, P4HB, PDIA3, PDLIM1, PHGDH, PPIA, PPP2CA, PRDX1, PSMB1, PSMD2, SFN, SOD1, STAT1, TKT, TPI1, TPM3, TUBB, TUBB4B, TUBB6, TUBB8, VCL, VCP, VDAC2, VIM, XRCC6, YBX1, ZYX | 74 |
| Embryonic development, organismal survival | Death of embryo | 2.51 × 10-9 | 0.942 | BSG, CAPN2, CAPNS1, EIF6, FASN, IMPDH2, KIF5B, LRPPRC, MAP1B, NASP, PCNA, PFN1, PHB2, PRPF19, PSMC4, RPSA, SF3B1, TKT, TPM3, VCP | 20 |
| Cell cycle | Cell cycle progression | 2.53 × 10-9 | −1.218 | BUB3, C1QBP, CALR, CAPN2, CAST, CAV1, CCT4, CD44, CD59, CDK1, CLTC, CSE1L, CYR61, DBI, DHX9, DYNC1H1, EIF6, EZR, FASN, FKBP1A, FLNA, GNB2L1, GPI, H3F3A/H3F3B, HMGB1, HSPA8, HSPB1, ITGB1, KHDRBS1, KPNB1, KRT18, KRT8, LMNA, MAP4, MAPK1, MARCKS, MCM7, MYH10, NASP, NPM1, NUMA1, PA2G4, PCM1, PCNA, PEBP1, PPM1G, PPP1CA, PPP2CA, PRDX1, PRMT5, PTMA, RAN, RBM3, RHOA, RPS24, SEPT9, SFN, SFPQ, SSRP1, STAT1, STMN1, TCP1, TMPO, TNC, TUBB, VCP, XRCC6, YWHAB, YWHAE | 69 |
| Cell morphology | Morphology of cells | 2.61 × 10-9 | ACTA1, ACTN4, ACTR2, ANXA1, ANXA2, ARF1, ARHGDIA, ARPC2, B2M, BUB3, CALR, CAP1, CAPN1, CAPN2, CAPRIN1, CAPZB, CAST, CAV1, CD44, CD59, CDK1, CFL1, CLIC4, CLTC, CSTB, CTSA, CTSB, CTTN, DNAJA1, DNM1L, DPY30, DPYSL2, EHD1, EIF6, EZR, FASN, FLNA, FLNC, FN1, GPI, H3F3A/H3F3B, HADH, HADHA, HEXB, HLA-A, HMGB1, HSP90AA1 | 110 | |
| HSP90B1, HSPA4, IGF2R, ILF3, IMPDH2, IQGAP1, ITGB1, KIF5B, KRT1, KRT18, KRT8, LAMB1, LGALS1, LMNA, LMNB1, MAP1B, MAP4, MAPK1, MARCKS, MYH10, MYH9, NPEPPS, NPM1, PAFAH1B2, PEA15, PEBP1, PLEC, PLIN3, PLS3, PRDX1, PRDX2, PRKDC, PTBP1, RAB11A, RAD23B, RAN, RHOA, RPSA, RTN4, SEPT7, SEPT9, SERPINH1, SNX3, SOD1, SPTBN1, STAT1, STMN1, SYNCRIP, TALDO1, TFRC, TLN1, TMOD3, TNC, TPM3, TPP1, VCL, VIM, XRCC5, XRCC6, YBX1, YWHAG, YWHAZ, ZYX | |||||
| Cancer, respiratory disease | Stage 1 non-small-cell lung carcinoma | 2.83 × 10-9 | B2M, EEF1A1, EIF4A1, FN1, MYL12A, PPIA, RPL27, RPL7, RPS11, RPS27A, SERBP1 | 11 | |
| Cellular assembly and organization, tissue development | Formation of filaments | 3.49 × 10-9 | −0.462 | ACTA1, ACTR3, ARF1, ARPC2, CAPN1, CAPZB, CAV1, CD44, CFL1, CTTN, DPYSL2, FKBP4, FN1, GNAI2, GPI, HSPA5, HSPB1, ITGB1, KRT18, MAP1B, MAPK1, MAPRE1, NPM1, PFN1, RHOA, SERPINH1, SOD1, STMN1, TCP1, TNC, TPM2, TUBB, TWF1, TWF2, VIM, ZYX | 36 |
| Infectious disease | Infection by HIV-1 | 4.09 × 10-9 | −1.626 | ARF1, ATP5B, CCT2, CD44, DDX3X, DHX9, DNAJA2, GANAB, GML, H3F3A/H3F3B, HMGB1, HNRNPH1, HNRNPU, ITGB1, KHDRBS1, KPNB1, MAP4, PDIA3, PDIA6, PGM1, PLOD2, PSMA1, PSMA2, PSMA5, PSMA7, PSMC4, PSME2, PTGES3, RAB1B, RANBP1, RNH1, RPL10A, RPL12, RPL18, SF3A1, SF3B1, SNRPD3, SPTAN1, SPTBN1, STIP1, TAGLN2, TWF1, UAP1, UBE2L3, UQCRC1, XPO1, YBX1, ZYX | 48 |
| Cancer | Malignant neoplasm of heart, mediastinum and pleura | 4.65 × 10-9 | 1.982 | ATIC, B2M, EEF1A1, EIF4A1, FN1, MYL12A, PPIA, PRDX1, PRKDC, RHOA, RPL27, RPL7, RPS11, RPS27A, TUBB, TUBB4B, XRCC5, XRCC6, YWHAE | 19 |
| Cancer | Growth of tumor | 5.06 × 10-9 | −0.277 | ACLY, ACTN4, AHCY, AKR1B1, ANXA1, ANXA2, BSG, CACYBP, CALR, CAPN2, CAV1, CD44, CD59, CTSB, CTTN, CYR61, EEF1A1, EZR, FASN, FLNA, FN1, GAPDH, GNB2L1, HDGF, HMGB1, HSPA4, HSPA5, HSPA8, HSPD1, HYOU1, ILF3, ITGB1, LAMB1, LDHA, LGALS1, MAPK1, NPM1, PARK7, PKM, PLEC, PPP2CA, RHOA, RPL22, RPS4X, S100A6, SET, SQSTM1, STAT1, STMN1, TNC, TXLNA, UCHL1, XRCC5, YBX1 | 54 |
| RNA post-transcriptional modification | Splicing of RNA | 5.23 × 10-9 | 0.239 | AHNAK, C1QBP, DDX39B, DDX5, FUS, HNRNPA2B1, HNRNPH1, HNRNPH3, HNRNPK, HNRNPM, KHSRP, NPM1, PRPF19, PTBP1, SF3A1, SF3B1, SFPQ, SNRPD1, SYNCRIP, U2AF2 | 20 |
| Cell death and survival | Cell death of connective tissue cells | 5.91 × 10-9 | −0.258 | CAPNS1, CD44, CDK1, CLIC4, CTSB, CYR61, DDX3X, DNM1L, EEF1A1, EIF2S1, EIF3B, EIF3C, EIF3I, EIF6, ENO1, FKBP4, FLNA, FN1, FUS, GLO1, GPI, GSTP1, HINT1, HSPA5, HSPD1, IGF2R, ITGB1, KHDRBS1, KRT18, KRT8, LMNA, MAP4, MAPK1, PARK7, PRKDC, RHOA, RPL10, SERPINH1, STAT1, STMN1, TPP1, UBA1, VCP, VIM, XRCC6, YWHAZ | 46 |
| Cancer, hematological disease | Blood tumor | 6.43 × 10-9 | 1.121 | ACTG1, ANXA1, ANXA2, ANXA6, ATIC, B2M, CALR, CAV1, CD44, CDK1, CFL1, CSE1L, CYR61, DNM1L, EEF1A1, FASN, FDPS, FKBP1A, FLNA, FN1, FUS, G3BP1, GMPS, GNB2L1, GSTP1, HIST1H1C, HLA-A, HMGB1, HSPD1, IMPDH2, IQGAP1, KPNA2, KRT1, LDHA, LGALS1, LOC102724594/U2AF1, LONP1, MARS, MYH10, NNMT, NONO, NPM1, PCNA, PDIA6, PICALM, PLOD2, PPP2CA, PRDX1, PRKDC, PSMA2, PSMB1, PSMD2, PSME1, RHOA, RPL3, RPL6, RPS12, RPS2, RPS24, RPS4X, SF3B1, SHMT2, SNRPD3, SPTBN1, STIP1, STMN1, TUBB, TUBB4B, TUBB6, TUBB8, TXLNA, UQCRC1, VIM, XPO1, XRCC5, XRCC6, YWHAE, YWHAZ | 78 |
| Cell-to-cell signaling and interaction | Binding of cells | 6.68 × 10-9 | −0.441 | ANXA2, ANXA5, BSG, C1QBP, CALR, CAV1, CCT2, CCT3, CCT4, CCT5, CCT6A, CCT7, CCT8, CD44, CKAP4, CYR61, DDX3X, FN1, HLA-A, HMGB1, HSPA5, IGF2R, ITGB1, KRT1, MAP4, MSN, MYH9, NCL, PDIA3, PPP2CA, PRMT5, RHOA, RPSA, STIP1, STMN1, TCP1, TFRC, TLN1, TNC, VCL | 40 |
| Cancer, hematological disease | Hematologic cancer | 7.44 × 10-9 | 1.121 | ACTG1, ANXA1, ANXA2, ANXA6, ATIC, B2M, CALR, CAV1, CD44, CDK1, CFL1, CSE1L, CYR61, DNM1L, EEF1A1, FASN, FDPS, FKBP1A, FLNA, FN1, FUS, G3BP1, GMPS, GNB2L1, GSTP1, HIST1H1C, HLA-A, HMGB1, HSPD1, IMPDH2, IQGAP1, KPNA2, KRT1, LDHA, LGALS1, LOC102724594/U2AF1, LONP1, MARS, MYH10, NNMT, NONO, NPM1, PCNA, PDIA6, PICALM, PPP2CA, PRDX1, PRKDC, PSMA2, PSMB1, PSMD2, PSME1, RHOA, RPL3, RPL6, RPS12, RPS2, RPS24, RPS4X, SF3B1, SHMT2, SNRPD3, SPTBN1, STIP1, STMN1, TUBB, TUBB4B, TUBB6, TUBB8, TXLNA, UQCRC1, VIM, XPO1, XRCC5, XRCC6, YWHAE, YWHAZ | 77 |
| Cellular assembly and organization | Development of cytoplasm | 8.43 × 10-9 | −0.761 | ACTR3, ANXA1, ARF1, ARPC2, CAPN1, CAPZB, CAV1, CD44, CFL1, CORO1C, CTTN, DNM1L, DPYSL2, ERP29, FKBP4, FLNA, FN1, GDI1, GNAI2, GPI, HSPA8, IPO9, ITGB1, MAP1B, MAPK1, MAPRE1, NPM1, PARK7, PFN1, RHOA, STMN1, STOML2, TNC, TPM2, TUBB, TWF1, TWF2, ZYX | 38 |
| Neurological disease | Amyotrophic lateral sclerosis | 9.09 × 10-9 | ANXA1, ANXA5, CAPN2, CCT2, CYR61, DBI, EEF1A1, EIF4A1, EZR, FASN, FUS, HN1, HNRNPA1, MAPK1, NNMT, PFKP, PFN1, RHOA, RTN4, S100A6, SOD1, TFRC, VCP, VIM | 24 | |
| Cell death and survival | Cell viability of tumor Cell lines | 9.74 × 10-9 | −0.894 | APEX1, ATP5H, B2M, CAPN2, CAV1, CD44, COPB2, DHX9, EEF2, EIF3C, EIF4A1, EIF4G1, FLNA, FN1, GLUD1, GNB2L1, GSTP1, HINT1, HMGB1, HSP90AB1, HSP90B1, HSPA5, HSPB1, IGF2R, ITGB1, P4HB, PARK7, PCNA, PPM1G, PPP1CA, PPP2CA, PPP2R1A, PRDX2, PRKDC, PRPF19, PSMA1, PSMA4, PSMC4, RAB11A, RHOA, RPL27, RPSA, S100A6, SF3A1, SFN, SNRPD1, SOD1, SQSTM1, TCP1, TRIM28, VCP, XPO1, YBX1 | 53 |
| Hereditary disorder, neurological disease, psychological disorders, skeletal and muscular disorders | Huntington’s disease | 1.18 × 10-8 | ACAT1, AHCY, B2M, BASP1, CAPNS1, CD44, CYC1, DDX1, DNAJA1, ENO2, FKBP4, GAPDH, GPI, HADH, HINT1, HNRNPU, HSP90AA1, HSPA5, HSPA8, LAMB1, LDHA, LDHB, MYL12A, NPM1, PCMT1, PCNA, PDE6H, PDLIM1, PFN1, PGK1, PKM, PLOD2, PPIA, PRDX2, PSME1, RAB11A, RANBP1, RPSA, SDHA, TPI1, TPM3, UCHL1, UQCRC1, XRCC6, YWHAZ | 45 | |
| Cellular development, cellular growth and proliferation | Proliferation of breast cancer cell lines | 1.19 × 10-8 | 0.389 | ANXA2, ARF1, C1QBP, CAV1, CD44, CD59, CDK1, CSE1L, CYR61, EEF1A1, EEF1B2, EIF3C, FN1, GNB2L1, HNRNPK, HSPA5, ILF3, ITGB1, KPNA2, KRT8, LGALS1, MAPK1, PDIA3, PFKP, PFN1, PPP2R1A, PRDX2, PTMA, SEPT9, SFN, SOD1, TRIM28, UBA1, YBX1, YWHAQ | 35 |
| Carbohydrate metabolism | Glycolysis of cells | 1.30 × 10-8 | −1.474 | ALDOA, BSG, C1QBP, CAV1, ENO1, ENO2, GAPDH, GPI, LDHA, PFKP, PGAM1, PGK1, PGM1, PKM, TPI1 | 15 |
| Cancer, respiratory disease | Metastatic non-small cell lung cancer | 1.31 × 10-8 | ATIC, B2M, EEF1A1, EIF4A1, FN1, MYL12A, PPIA, RPL27, RPL7, RPS11, RPS27A | 11 | |
| Protein synthesis | Polymerization of protein | 1.44 × 10-8 | ACAT1, AHNAK, ALDOC, ANXA2, ANXA5, ANXA6, ARPC2, CAV1, CTNNA1, CUTA, DNM1L, EEF1A1, EHD1, ENAH, HSD17B10, IMPDH2, LONP1, NPM1, PFKP, PFN1, PRKCSH, RHOA, SEPT2, SEPT7, SEPT9, SHMT2, SQSTM1, STOML2, TRIM28, VCP, YWHAB | 31 | |
| Cell death and survival | Cell death of fibroblast cell lines | 1.61 × 10-8 | −0.361 | CAPNS1, CDK1, CLIC4, CTSB, CYR61, DDX3X, DNM1L, EEF1A1, EIF2S1, EIF3B, EIF3C, EIF3I, EIF6, ENO1, FKBP4, FN1, FUS, GLO1, GPI, GSTP1, HINT1, HSPA5, HSPD1, ITGB1, KRT18, KRT8, MAPK1, PARK7, RHOA, RPL10, STAT1, STMN1, UBA1, VCP, VIM, YWHAZ | 36 |
| Cellular assembly and organization, cellular function and maintenance | Organization of actin cytoskeleton | 2.12 × 10-8 | −1.937 | ACTR2, ALDOA, ARHGDIA, CALR, CAP1, CFL1, CORO1C, CSRP1, DBN1, DPYSL2, ENAH, EZR, FLNA, FLNB, FLNC, FN1, ITGB1, MSN, MYH10, MYH9, PLS3, RAN, RHOA, SEPT2, SPTAN1, TLN1, TMOD3, TWF2 | 28 |
| Cancer | Malignant neoplasm of mediastinum | 2.27 × 10-8 | 1.982 | ATIC, B2M, EEF1A1, EIF4A1, FN1, MYL12A, PPIA, PRDX1, PRKDC, RHOA, RPL27, RPL7, RPS11, RPS27A, TUBB4B, XRCC5, XRCC6, YWHAE | 18 |
| Metabolic disease | Amyloidosis | 2.45 × 10-8 | ACLY, ATP5A1, B2M, CANX, CAPN1, CAST, CAV1, CDK1, CTSB, DHX9, DNM1L, DPYSL2, EEF1G, EEF2, EIF2S1, EPRS, FDPS, G3BP1, GAPDH, GNB2L1, HNRNPA1, HNRNPA2B1, HNRNPU, HSPA5, HSPD1, LGALS1, PCNA, PICALM, PRDX1, PRKDC, PSMB1, PSMD2, RAN, SET, SOD1, STIP1, TAGLN, TUBB, UBXN4, UCHL1, VIM, VPS35, YWHAZ | 43 | |
| Nucleic acid metabolism, small molecule biochemistry | Biosynthesis of purine ribonucleotide | 2.57 × 10-8 | −2.378 | ALDOA, ATP5A1, ATP5B, CAV1, CDK1, DNM1L, HMGB1, HSPD1, NNMT, PKM, PPA1, SLC25A5, SOD1, VCP, VDAC1 | 15 |
| Neurological disease | Neurological signs | 2.58 × 10-8 | ACAT1, AHCY, B2M, BASP1, CAPNS1, CAV1, CD44, CYC1, DDX1, DNAJA1, ENO2, FKBP4, GAPDH, GPI, HADH, HINT1, HNRNPA2B1, HNRNPU, HSP90AA1, HSPA5, HSPA8, LAMB1, LDHA, LDHB, MYL12A, NPM1, PCMT1, PCNA, PDE6H, PDLIM1, PFN1, PGK1, PKM, PLOD2, PPIA, PRDX2, PSME1, RAB11A, RANBP1, RPSA, SDHA, SOD1, TPI1, TPM3, UCHL1, UQCRC1, XRCC6, YWHAZ | 48 | |
| Cancer | Epithelial cancer | 3.20 × 10-8 | −0.050 | AARS, ACAT1, ACAT2, ACLY, ACTG1, ACTN1, ACTN4, ACTR2, ACTR3, AHNAK, AKR1B1, ALDOA, ALDOC, ANXA1, ANXA2, ANXA5, AP1B1, APEX1, ARF1, ARHGDIA, ARPC2, ATIC, ATP5A1, ATP5B, B2M, BASP1, BSG, C14orf166, C1QBP, CACYBP, CAND1, CANX, CAPG, CAPN2, CAPRIN1, CAST, CAV1, CBX3, CCT2, CCT4, CCT5, CCT6A, CCT7, CCT8, CD44, CD59, CDK1, CLIC1, CLTC, COPA, COPB1, COPB2, COPE, COPG1, CORO1C, COTL1, CRIP2, CSE1L, CTNNA1, CTPS1, CTSA, CTSB, CTTN, CUTA, CYC1, CYR61, DBI, DBN1, DDX1, DDX39B, DDX3X, DDX5, DHX9, DLAT, DNAJA1, DNAJA2, DNAJC8, DNM1L, DPYSL2, DYNC1H1, DYNC1I2, ECHS1, EEF1A1, EEF1B2, EEF1D, EEF2, EIF2S1, EIF2S2, EIF3I, EIF4A1, EIF4G1, ENAH, ENO1, ENO2, EPRS, ESYT1, ETFA, EZR, FASN, FDPS, FKBP1A, FLNA, FLNB, FLNC, FN1, FUBP3, FUS, G3BP1, G6PD, GANAB, GARS, GCN1L1, GDI2, GLUD1, GML, GMPS, GNAI2, GNB2L1, GPI, GSTP1, H2AFY, H3F3A/H3F3B, HADHA, HARS, HDGF, HDLBP, HEXB, HINT1, HIST1H1C, HIST1H2BL, HIST2H2AC, HLA-A, HLA-B, HMGB1, HN1, HNRNPA1, HNRNPA2B1, HNRNPA3, HNRNPH1, HNRNPH3, HNRNPK, HNRNPL, HNRNPM, HNRNPU, HSP90AA1, HSP90AB1, HSP90B1, HSPA5, HSPA8, HSPA9, HSPD1, HYOU1, IARS, IDI1, IFITM3, IGF2R, ILF2, ILF3, IMMT, IMPDH2, IPO5, IPO7, IPO9, IQGAP1, ITGB1, KHDRBS1, KHSRP, KIF5B, KPNA2, KPNA3, KPNB1, KRT1, KRT18, KRT8, KRT9, LAMB1, LASP1, LDHA, LGALS1, LMNA, LMNB1, LOC102724594/U2AF1, LONP1, LRPPRC, LRRC59, MAP1B, MAP4, MAPK1, MAPRE1, MARS, MATR3, MCM3, MCM4, MCM6, MCM7, MDH1, MDH2, MSN, MVP, MYH10, MYH9, MYL12A, NACA, NAP1L1, NAP1L4, NASP, NCL, NNMT, NONO, NPEPPS | 350 |
| NPLOC4, NPM1, NSFL1C, NUMA1, OLA1, P4HB, PA2G4, PABPC1, PAFAH1B2, PAICS, PCBP1, PCBP2, PCM1, PCMT1, PCNA, PDCD6, PDIA3, PDIA4, PDLIM1, PDXK, PEA15, PFKP, PFN1, PGAM1, PGD, PGK1, PGM1, PHGDH, PICALM, PKM, PLEC, PLOD2, PPIA, PPM1G, PPP1CA, PPP2CA, PPP2R1A, PRDX1, PRDX2, PRKCSH, PRKDC, PRMT5, PRPF19, PSMA1, PSMC1, PSMC4, PSMD2, PTBP1, PTMA, PTRF, PUF60, PYGB, PYGL, QARS, RANBP1, RARS, RHOA, RPL10, RPL12, RPL22, RPL27, RPL4, RPL5, RPL7, RPL7A, RPL8, RPLP0, RPN1, RPN2, RPS11, RPS12, RPS2, RPS24, RPS27A, RPS7, RRBP1, RTN4, S100A6, SDHA, SEPT9, SERBP1, SERPINH1, SF3A1, SF3B1, SFN, SFPQ, SH3BGRL3, SLC25A3, SLC25A5, SLC38A2, SND1, SOD1, SPTAN1, SPTBN1, SSRP1, STAT1, STIP1, STMN1, SYNCRIP, TAGLN, TALDO1, TARS, TCP1, TFRC, TLN1, TMPO, TNC, TNPO1, TPI1, TPM4, TUBB, TUBB4B, TUBB6, TUBB8, TWF2, TXNDC5, UAP1, UBA1, UBE2N, UBXN4, UCHL1, VARS, VCL, VCP, VIM, VPS35, WARS, WDR1, XPO1, XRCC5, XRCC6, YBX1, YBX3, YWHAB, YWHAE, YWHAG, YWHAH, YWHAQ, YWHAZ, ZYX | |||||
| Cancer | Metastatic carcinoma | 3.23 × 10-8 | ATIC, B2M, CSE1L, EEF1A1, EIF4A1, FKBP1A, FN1, HSP90AA1, HSP90AB1, HSP90B1, MYL12A, PPIA, RPL27, RPL7, RPS11, RPS27A, TUBB, TUBB4B, YWHAE | 19 | |
| Neurological disease | Dyskinesia | 3.44 × 10-8 | ACAT1, AHCY, B2M, BASP1, CAPNS1, CAV1, CD44, CYC1, DDX1, DNAJA1, ENO2, FKBP4, GAPDH, GPI, HADH, HINT1, HNRNPU, HSP90AA1, HSPA5, HSPA8, LAMB1, LDHA, LDHB, MYL12A, NPM1, PCMT1, PCNA, PDE6H, PDLIM1, PFN1, PGK1, PKM, PLOD2, PPIA, PRDX2, PSME1, RAB11A, RANBP1, RPSA, SDHA, TPI1, TPM3, UCHL1, UQCRC1, XRCC6, YWHAZ | 46 | |
| Cellular assembly and organization | Formation of cytoskeleton | 3.46 × 10-8 | −0.867 | ACTR3, ARF1, ARPC2, CAPN1, CAPZB, CAV1, CD44, CFL1, CORO1C, CTTN, DPYSL2, ERP29, FKBP4, FLNA, FN1, GDI1, GNAI2, GPI, ITGB1, MAP1B, MAPK1, MAPRE1, NPM1, PFN1, RHOA, STMN1, TNC, TPM2, TUBB, TWF1, TWF2, ZYX | 32 |
| Inflammatory disease | Chronic inflammatory disorder | 3.62 × 10-8 | ACLY, ACTA1, ACTN4, ALDOA, ANXA1, ARF1, ATIC, B2M, BSG, CALR, CAPG, CD59, COTL1, CTSB, DDX39B, DDX5, EEF1G, EEF2, ENO1, FDPS, FKBP1A, H3F3A/H3F3B, HLA-A, HMGB1, HNRNPA1, HNRNPA3, HSP90B1, HSPA8, HSPD1, IMPDH2, KHSRP, KRT18, KRT8, LDHB, LGALS1, MAPRE1, MYL12A, NONO, P4HB, PCM1, PDIA3, PDLIM1, PGK1, PKM, PRDX1, PRDX2, PRDX5, PSMB1, PSMD2, PTMA, RHOA, RPS24, RPS3, RPSA, SND1, STAT1, TFRC, TPM2, TRIM28, UCHL1, VARS, VIM | 62 | |
| Cell morphology, connective tissue development and function | Shape change of fibroblast cell lines | 3.74 × 10-8 | −0.342 | CAST, EHD1, FLNA, FLNC, FN1, GNB2L1, IPO9, ITGB1, MARCKS, PTBP1, RHOA, RTN4, VCL, VIM | 14 |
| Cell morphology, cellular assembly and organization, cellular function and maintenance | Formation of lamellipodia | 3.90 × 10-8 | 0.308 | ACTN4, ACTR3, ARPC2, BSG, C1QBP, CAP1, CAPZB, CD44, CFL1, CYR61, DNM1L, EZR, FN1, HSP90AA1, HSPB1, LASP1, RHOA, TWF2, VCL | 19 |
| Cell cycle, cell morphology, cellular assembly and organization, cellular movement | Elongation of filaments | 4.03 × 10-8 | −1.000 | ACTN4, CAP1, FN1, MAPRE1, PFN1, SEPT7, SEPT9 | 7 |
| Cell death and survival | Neuronal cell death | 4.08 × 10-8 | 0.202 | AARS, AP2B1, CAPN1, CAPNS1, CAPRIN1, CAST, CDK1, CTSB, DNM1L, FKBP1A, FN1, FUS, G6PD, GAPDH, GLUD1, GPI, HDGF, HMGB1, HSD17B10, HSP90AB1, HSPA5, HSPB1, HSPD1, HYOU1, ITGB1, LDHA, LGALS1, MAP1B, MAPK1, NPM1, P4HB, PARK7, PDIA3, PEA15, PRDX2, RHOA, RPS3, SDHA, SET, SOD1, STAT1, STIP1, TCP1, UCHL1, VAPA, XRCC5, XRCC6, YWHAB, YWHAZ | 49 |
| Cell-to-cell signaling and interaction, cellular assembly and organization, tissue development | Quantity of focal adhesions | 4.08 × 10-8 | 1.463 | CAPNS1, CAV1, FLNA, FLNB, FN1, GNB2L1, ITGB1, MYH9, RHOA, SPTAN1 | 10 |
| Carbohydrate metabolism | Glycolysis | 4.46 × 10-8 | −1.946 | ALDOA, BSG, C1QBP, CAV1, DNM1L, ENO1, ENO2, GAPDH, GPI, LDHA, PFKP, PGAM1, PGK1, PGM1, PKM, TPI1 | 16 |
| RNA post-transcriptional modification | Processing of mRNA | 4.53 × 10-8 | −0.128 | C1QBP, DDX39B, DDX5, HNRNPA1, HNRNPA2B1, HNRNPH3, HNRNPK, HNRNPM, KHDRBS1, LOC102724594/U2AF1, NPM1, PABPC1, PCBP1, PRPF19, PTBP1, SF3A1, SF3B1, SFPQ, SNRPD1, U2AF2 | 20 |
| Molecular transport | Transport of molecule | 4.60 × 10-8 | −1.226 | ACAT2, ALYREF, ANXA1, ANXA2, ANXA6, AP1B1, ARF1, ATP1A1, ATP5B, B2M, BSG, CALR, CANX, CAV1, CD44, CFL1, CLIC1, CLIC4, CPNE1, CSE1L, CTSA, DDX39B, DDX3X, DNAJA1, DNAJA2, DNM1L, DPYSL2, EHD1, EIF5A, ERP29, FKBP4, FN1, FUS, GDI2, GNAI2, HNRNPA2B1, HSPA8, HSPA9, IGF2R, IPO5, IPO7, IPO9, ITGB1, KHDRBS1, KIF5B, KPNA2, KPNB1, KRT8, LASP1, LGALS1, MAP1B, MYH9, NPM1, P4HB, PDCD6, PDIA3, PDIA4, PEA15, PICALM, PLIN3, PPIA, RAB11A, RAB7A, RAN, RHOA, RRBP1, RTN4, S100A6, SEC13, SEPT2, SLC25A3, SLC25A5, SLC38A2, SNX3, SOD1, SPTBN1, SQSTM1, STAT1, STOML2, TFRC, TNPO1, U2AF2, VAPA, VDAC1, XPO1, YWHAB, YWHAE, YWHAH, YWHAZ | 89 |
| Cellular development | Differentiation of cells | 4.66 × 10-8 | −1.742 | ACLY, ACTR3, ALYREF, ANXA1, ANXA2, ANXA6, ATP5B, B2M, BASP1, BSG, C1QBP, CACYBP, CALR, CAND1, CAPNS1, CAPZB, CAST, CAV1, CBX3, CD44, CDK1, CFL1, CLIC1, CLIC4, CLTC, CSRP1, CTSB, CYR61, DBN1, DDX5, DHX9, DNM1L, DPYSL2, EIF5A, ENO1, EZR, FASN, FLNB, FLNC, FN1, GAPDH, GLO1, GNB2L1, H2AFY, H3F3A/H3F3B, HMGB1, HNRNPA2B1, HNRNPK, HNRNPL, HNRNPU, HSP90AA1, HSP90AB1, HSP90B1, HSPA5, HSPD1, IARS, IGF2R, ITGB1, KHDRBS1, KRT1, KRT8, LGALS1, LMNA, LMNB1, LONP1, MAP1B, MAPK1, NAP1L1, NNMT, NPEPPS, NPM1, PA2G4, PDIA3, PFN1, PHGDH, PICALM, PKM, PPIA, PPP1CA, PRDX2, PRKDC, PRMT5, PRPF19, RHOA, RNH1, RPL22, RPS11, RPS15, RPS3A, RPS7, RRBP1, RTN4, SFN, SFPQ, SND1, SOD1, SQSTM1, STAT1, STMN1, SYNCRIP, TAGLN, TAGLN2, TFRC, TLN1, TNC, TPM2, TPM4, VIM, XPO1, XRCC5, XRCC6, YBX1, YBX3, YWHAG, YWHAQ | 115 |
| Cancer | Neoplasia of epithelial tissue | 4.68 × 10-8 | 0.453 | AARS, ACAT1, ACAT2, ACLY, ACTG1, ACTN1, ACTN4, ACTR2, ACTR3, AHNAK, AIP, AKR1B1, ALDOA, ALDOC, ANXA1, ANXA2, ANXA5, AP1B1, APEX1, ARF1, ARHGDIA, ARPC2, ATIC, ATP5A1, ATP5B, B2M, BASP1, BSG, C14orf166, C1QBP, CACYBP, CALR, CAND1, CANX, CAPG, CAPN2, CAPRIN1, CAST, CAV1, CBX3, CCT2, CCT4, CCT5, CCT6A, CCT7, CCT8, CD44, CD59, CDK1, CLIC1, CLTC, COPA, COPB1, COPB2, COPE, COPG1, CORO1C, COTL1, CRIP2, CSE1L, CTNNA1, CTPS1, CTSA, CTSB, CTTN, CUTA, CYC1, CYR61 | 355 |
| DBI, DBN1, DDX1, DDX39B, DDX3X, DDX5, DHX9, DLAT, DNAJA1, DNAJA2, DNAJC8, DNM1L, DPYSL2, DYNC1H1, DYNC1I2, ECHS1, EEF1A1, EEF1B2, EEF1D, EEF2, EIF2S1, EIF2S2, EIF3I, EIF4A1, EIF4G1, ENAH, ENO1, ENO2, EPRS, ESYT1, ETFA, EZR, FASN, FDPS, FKBP1A, FLNA, FLNB, FLNC, FN1, FUBP3, FUS, G3BP1, G6PD, GANAB, GARS, GCN1L1, GDI2, GLUD1, GML, GMPS, GNAI2, GNB2L1, GPI, GSTP1, H2AFY, H3F3A/H3F3B, HADHA, HARS, HDGF, HDLBP, HEXB, HINT1, HIST1H1C, HIST1H2BL, HIST2H2AC, HLA-A, HLA-B, HMGB1, HN1, HNRNPA1, HNRNPA2B1, HNRNPA3, HNRNPH1, HNRNPH3, HNRNPK, HNRNPL, HNRNPM, HNRNPU, HSP90AA1, HSP90AB1, HSP90B1, HSPA5, HSPA8, HSPA9, HSPD1, HYOU1, IARS, IDI1, IFITM3, IGF2R, ILF2, ILF3, IMMT, IMPDH2, IPO5, IPO7, IPO9, IQGAP1, ITGB1, KHDRBS1, KHSRP, KIF5B, KPNA2, KPNA3, KPNB1, KRT1, KRT18, KRT8, KRT9, LAMB1, LASP1, LDHA, LGALS1, LMNA, LMNB1, LOC102724594/U2AF1, LONP1, LRPPRC, LRRC59, MAP1B, MAP4, MAPK1, MAPRE1, MARCKS, MARS, MATR3, MCM3, MCM4, MCM6, MCM7, MDH1, MDH2, MSN, MVP, MYH10, MYH9, MYL12A, NACA, NAP1L1, NAP1L4, NASP, NCL, NNMT, NONO, NPEPPS, NPLOC4, NPM1, NSFL1C, NUMA1, OLA1, P4HB, PA2G4, PABPC1, PAFAH1B2, PAICS, PARK7, PCBP1, PCBP2, PCM1, PCMT1, PCNA, PDCD6, PDIA3, PDIA4, PDLIM1, PDXK, PEA15, PFKP, PFN1, PGAM1, PGD, PGK1, PGM1, PHGDH, PICALM, PKM, PLEC, PLOD2, PPIA, PPM1G, PPP1CA, PPP2CA, PPP2R1A, PRDX1, PRDX2, PRKCSH, PRKDC, PRMT5, PRPF19, PSMA1, PSMC1, PSMC4, PSMD2, PTBP1, PTMA, PTRF, PUF60, PYGB, PYGL, QARS, RANBP1, RARS, RHOA, RPL10, RPL12, RPL21, RPL22, RPL27, RPL4, RPL5, RPL7, RPL7A, RPL8, RPLP0, RPN1, RPN2, RPS11, RPS12, RPS2, RPS24, RPS27A, RPS7, RRBP1, RTN4, S100A6, SDHA, SEPT9, SERBP1, SERPINH1, SF3A1, SF3B1, SFN, SFPQ, SH3BGRL3, SLC25A3, SLC25A5, SLC38A2, SND1, SOD1, SPTAN1, SPTBN1, SSRP1, STAT1, STIP1, STMN1, SYNCRIP, TAGLN, TALDO1, TARS, TCP1, TFRC, TLN1, TMPO, TNC, TNPO1, TPI1, TPM4, TUBB, TUBB4B, TUBB6, TUBB8, TWF2, TXNDC5, UAP1, UBA1, UBE2N, UBXN4, UCHL1, VARS, VCL, VCP, VIM, VPS35, WARS, WDR1, XPO1, XRCC5, XRCC6, YBX1, YBX3, YWHAB, YWHAE, YWHAG, YWHAH, YWHAQ, YWHAZ, ZYX | |||||
| Hereditary disorder, skeletal and muscular disorders | Autosomal recessive myopathy | 4.96 × 10-8 | AHNAK, CALR, CANX, FN1, HINT1, HSP90B1, HSPA5, ITGB1, LMNA, PLEC, PSMA2, PSMA4, SQSTM1, VCP | 14 | |
| Infectious disease | Infection of tumor cell lines | 5.02 × 10-8 | −1.728 | ACTR2, AP1B1, AP2B1, ARF1, ARPC1B, ATP5B, CCT2, CLTC, COPA, COPB1, COPB2, COPG1, DDX3X, DNAJA2, EIF3I, G3BP1, GML, H3F3A/H3F3B, HMGB1, HNRNPK, HNRNPU, HSPA9, IFITM3, IGF2R, MAP4, PDIA3, PDIA6, PICALM, PSME2, RAB1B, RANBP1, SPTAN1, SPTBN1, STAT1, STIP1, TFRC, TWF1, UAP1, UQCRC1, XPO1 | 40 |
| Cancer | Abdominal neoplasm | 5.10 × 10-8 | −1.025 | AARS, ACAT1, ACAT2, ACLY, ACTG1, ACTN1, ACTN4, ACTR2, ACTR3, AHCY, AHNAK, AKR1B1, ALDOA, ALYREF, ANXA1, ANXA2, ANXA5, AP1B1, APEX1, ARF1, ARHGDIA, ARPC2, ATIC, ATP5A1, ATP5B, B2M, BASP1, BSG, C14orf166, CACYBP, CALR, CAND1, CANX, CAP1, CAPG, CAPN2, CAST, CAV1, CBX3, CCT2, CCT4, CCT5, CCT6A, CCT8, CD44, CD59, CDK1, CLIC1, CLTC, COPA, COPB1, COPB2, COPE, COPG1, CORO1C, COTL1, CRIP2, CSE1L, CTNNA1, CTPS1, CTSA, CTSB, CTTN, CUTA, CYC1, CYR61, DBI, DBN1, DDX1, DDX39B, DDX3X, DDX5, DDX6, DHX9, DLAT, DNAJA1, DNAJA2, DNAJC8, DNM1L, DPYSL2 | 350 |
| DYNC1H1, DYNC1I2, ECHS1, EEF1A1, EEF1B2, EEF1D, EEF2, EIF2S1, EIF2S2, EIF3I, EIF4A1, EIF4G1, ENAH, ENO1, ENO2, EPRS, ESYT1, FASN, FDPS, FKBP1A, FLNA, FLNB, FLNC, FN1, FUBP3, FUS, G3BP1, G6PD, GANAB, GAPDH, GARS, GCN1L1, GDI2, GLUD1, GML, GMPS, GNAI2, GNB2L1, GPI, GSTP1, H2AFY, H3F3A/H3F3B, HADHA, HARS, HDGF, HDLBP, HEXB, HINT1, HIST1H1C, HIST1H2BL, HIST2H2AC, HLA-A, HLA-B, HMGB1, HN1, HNRNPA1, HNRNPA2B1, HNRNPA3, HNRNPH1, HNRNPH3, HNRNPK, HNRNPL, HNRNPM, HNRNPU, HSD17B10, HSP90AA1, HSP90AB1, HSP90B1, HSPA5, HSPA8, HSPA9, HSPB1, HSPD1, HYOU1, IARS, IDI1, IFITM3, IGF2R, ILF2, ILF3, IMMT, IMPDH2, IPO5, IPO7, IPO9, IQGAP1, ITGB1, KHDRBS1, KHSRP, KIF5B, KPNA2, KPNA3, KPNB1, KRT1, KRT18, KRT8, KRT9, LAMB1, LDHA, LGALS1, LMNA, LMNB1, LOC102724594/U2AF1, LONP1, LRPPRC, LRRC59, MAP1B, MAP4, MAPK1, MAPRE1, MARS, MATR3, MCM3, MCM4, MCM6, MCM7, MDH1, MDH2, MSN, MVP, MYH10, MYH9, NACA, NAP1L1, NAP1L4, NASP, NCL, NNMT, NONO, NPEPPS, NPLOC4, NPM1, NSFL1C, NUMA1, OLA1, P4HB, PA2G4, PABPC1, PAICS, PCBP1, PCBP2, PCM1, PCNA, PDIA3, PDIA4, PDLIM1, PDXK, PEA15, PFKP, PFN1, PGAM1, PGD, PGK1, PGM1, PHGDH, PICALM, PKM, PLEC, PLIN3, PLOD2, PLS3, PPM1G, PPP1CA, PPP2CA, PPP2R1A, PRDX1, PRDX2, PRKCSH, PRKDC, PSMA1, PSMA7, PSMC1, PSMC4, PSMD2, PTBP1, PTMA, PTRF, PUF60, PYGB, PYGL, QARS, RAN, RANBP1, RARS, RBM3, RHOA, RPL12, RPL22, RPL27, RPL4, RPL5, RPL6, RPL7A, RPL8, RPLP0, RPN1, RPN2, RPS12, RPS15, RPS2, RPS24, RPS4X, RPS7, RRBP1, RTN4, S100A6, SDHA, SEPT9, SERBP1, SERPINH1, SET, SF3A1, SF3B1, SFPQ, SH3BGRL3, SHMT2, SLC25A3, SLC25A5, SLC38A2, SND1, SOD1, SPTAN1, SPTBN1, SQSTM1, SSRP1, STAT1, STIP1, STMN1, SYNCRIP, TAGLN, TALDO1, TARS, TCP1, TFRC, TLN1, TMPO, TNC, TNPO1, TPI1, TPM2, TPM3, TPM4, TUBB, TUBB4B, TUBB8, TWF1, TWF2, TXNDC5, UAP1, UBA1, UBE2N, UBXN4, UCHL1, VARS, VCL, VCP, VIM, VPS35, WARS, XPO1, XRCC5, XRCC6, YBX1, YBX3, YWHAE, YWHAG, YWHAH, YWHAQ, YWHAZ, ZYX | |||||
| RNA post-transcriptional modification | Splicing of mRNA | 5.56 × 10-8 | 0.239 | C1QBP, DDX39B, DDX5, HNRNPA2B1, HNRNPH3, HNRNPK, HNRNPM, NPM1, PRPF19, PTBP1, SF3A1, SF3B1, SFPQ, SNRPD1, U2AF2 | 15 |
| Cell morphology | Shape of cells | 7.25 × 10-8 | ACTN4, CAPN1, FLNA, GPI, IGF2R, ITGB1, MYH10, RHOA, TPM3 | 9 | |
| Cell morphology, cellular assembly and organization, cellular development, cellular function and maintenance | Formation of plasma membrane projections | 7.91 × 10-8 | −0.331 | ACTR3, BASP1, CAPNS1, CAPRIN1, CAPZB, CAV1, CD44, CSRP1, CYR61, DBN1, DNM1L, DPYSL2, EHD1, EZR, GDI1, HMGB1, HNRNPK, ITGB1, KIF5B, LAMB1, MAP1B, MSN, MYH10, NNMT, PDIA3, PFN1, PHGDH, PICALM, PPP2CA, PRKCSH, RAB11A, RHOA, RTN4, SEPT2, SOD1, SPTBN1, STIP1, STMN1, TNC, UCHL1, VAPA, VIM, YWHAH | 43 |
| Cell cycle, cellular movement | Cytokinesis | 8.89 × 10-8 | −0.943 | ACTN4, CAP1, CFL1, FN1, GNAI2, ITGB1, LMNA, MAPRE1, MSN, MYH10, MYH9, NPM1, PFN1, RAB11A, RHOA, SEPT2, SEPT7, SEPT9, SSRP1, YBX1 | 20 |
| Cellular movement | Cell movement of breast cancer cell lines | 1.01 × 10-7 | 0.407 | ANXA1, ANXA2, ARF1, ARPC1B, CAPN2, CAV1, CSE1L, CTTN, ENAH, FLNA, FN1, GNB2L1, GPI, ILF3, ITGB1, KHDRBS1, KPNA2, KRT8, LASP1, MAPK1, MYH9, RHOA, SEPT9, SFN, VIM | 25 |
| Post-translational modification, protein folding | Refolding of protein | 1.08 × 10-7 | B2M, DNAJA1, DNAJA2, FKBP1A, HSP90AA1, HSPA8, HSPD1 | 7 | |
| Cell death and survival | Cell viability of myeloma cell lines | 1.08 × 10-7 | −0.376 | COPB2, EIF3C, EIF4A1, EIF4G1, HSP90B1, PSMA1, PSMA4, PSMC4, RAB11A, RPL27, SF3A1, SOD1, XPO1 | 13 |
| Cell morphology, cellular assembly and organization, cellular development, cellular function and maintenance, nervous system development and function, tissue development | Neuritogenesis | 1.09 × 10-7 | −0.401 | ACTR3, BASP1, CAPNS1, CAPRIN1, CAPZB, CAV1, CD44, CSRP1, CYR61, DBN1, DNM1L, DPYSL2, EHD1, EZR, GDI1, HMGB1, HNRNPK, ITGB1, LAMB1, MAP1B, MSN, MYH10, NNMT, PDIA3, PFN1, PHGDH, PICALM, PPP2CA, PRKCSH, RAB11A, RHOA, RTN4, SEPT2, SOD1, SPTBN1, STIP1, STMN1, TNC, UCHL1, VAPA, VIM, YWHAH | 42 |
| Free radical scavenging | Synthesis of reactive oxygen species | 1.15 × 10-7 | −0.019 | AKR1B1, ANXA1, ARHGDIA, CAV1, CD44, CLIC1, CTTN, CYR61, DNM1L, FN1, G6PD, GNB2L1, GSTP1, HMGB1, HNRNPK, HSD17B10, HSP90AB1, HSPA9, HSPB1, IMMT, IQGAP1, ITGB1, LDHA, LONP1, MAPK1, PARK7, PPIA, PRDX1, PRDX2, RHOA, S100A6, SOD1, SQSTM1, TAGLN, TFRC, VDAC1, YWHAZ | 37 |
| Free radical scavenging | Metabolism of reactive oxygen species | 1.17 × 10-7 | −0.644 | AKR1B1, ANXA1, ARHGDIA, CAV1, CD44, CLIC1, CTTN, CYR61, DNM1L, FN1, G6PD, GNB2L1, GSTP1, HMGB1, HNRNPK, HSD17B10, HSP90AB1, HSPA9, HSPB1, IMMT, IQGAP1, ITGB1, LDHA, LONP1, MAPK1, PARK7, PPIA, PRDX1, PRDX2, PRDX5, RHOA, S100A6, SOD1, SQSTM1, TAGLN, TFRC, VDAC1, YWHAZ | 38 |
| Cancer | Breast or colorectal cancer | 1.22 × 10-7 | 0.555 | AARS, ACAT1, ACAT2, ACLY, ACTN1, AHCY, AHNAK, AKR1B1, ALDOA, ALDOC, ANXA1, ANXA2, ARHGDIA, ATIC, ATP5A1, B2M, BSG, C1QBP, CACYBP, CAPG, CAPN2, CAV1, CBX3, CCT3, CCT4, CCT5, CD44, CDK1, COPA, COPB2, CSE1L, CTSB, CYC1, CYR61, DBI, DBN1, DDX1, DDX3X, DDX5, DLAT, DNAJA1, DNAJC8, DPYSL2, DYNC1H1, DYNC1I2, ECHS1, EEF1A1, EEF1D, EEF2, EIF3B, EIF3C, EIF4A1, EIF4G1, ENAH, ENO1, ESYT1, ETFA, FASN, FDPS, FKBP1A, FLNA, FLNB, FLNC, FN1, FUBP3, FUS, G3BP1, G6PD, GARS, GCN1L1, GLO1, GML, GMPS, GNAI2, GSTP1, H2AFY, H3F3A/H3F3B, HDGF, HIST1H1C, HLA-A, HLA-B, HNRNPA1, HNRNPA3, HNRNPH3, HNRNPL, HNRNPM, HNRNPU, HSP90AA1, HSP90AB1, HSP90B1, HSPA8, HSPA9, HSPD1, IDI1, IFITM3, IGF2R, ILF2, ILF3, IMMT, IPO7, IQGAP1, ITGB1, KHSRP, KIF5B, KPNA2, KPNA3, KRT1, KRT18, KRT8, LAMB1, LASP1, LDHA, LGALS1, LMNA, LMNB1, LONP1, LRPPRC, MAP1B, MAPRE1, MARS, MATR3, MCM3, MCM4, MCM6, MDH1, MDH2, MVP, MYH10, MYH9, NACA, NAP1L1, NAP1L4, NCL, NNMT, NPEPPS, OLA1, P4HB, PAFAH1B2, PAICS, PCBP1, PCNA, PDCD6, PFKP, PFN1, PGAM1, PGD, PGK1, PGM1, PKM, PLEC, PLIN3, PLOD2, PLS3, PPP1CA, PPP2CA, PPP2R1A, PRDX1, PRDX5, PRKCSH, PRKDC, PSMA7, PTBP1, PTMA, PTRF, PYGB, RANBP1, RARS, RHOA, RPL4, RPL5, RPL6, RPLP0, RPN1, RPN2, RPS24, RPS3, RTN4, S100A6, SEPT9, SERBP1, SERPINH1, SET, SF3B1, SFPQ, SHMT2, SLC25A5, SLC25A6, SOD1, SPTAN1, SSRP1, STAT1, STIP1, SYNCRIP, TAGLN, TAGLN2, TARS, TCP1, TFRC, TLN1, TMPO, TNC, TNPO1, TPI1, TPM4, TUBA1B, TUBB, TUBB4B, TUBB6, TUBB8, UBA1, VCL, VDAC2, VIM, VPS35, WDR1, XPO1, XRCC6, YBX1, YWHAE, YWHAH, YWHAQ, YWHAZ, ZYX | 223 |
| Cellular function and maintenance | Endocytosis | 1.22 × 10-7 | −0.427 | ACTN4, ANXA5, ANXA6, ATP5B, CANX, CAP1, CAV1, CD44, CLTC, CORO1C, CTTN, DPYSL2, EHD1, EZR, GNB2L1, HNRNPK, HSPA8, HSPA9, HYOU1, IGF2R, IQGAP1, ITGB1, MAP1B, NCL, PICALM, RAB7A, RHOA, SFPQ, TLN1 | 29 |
| Neurological disease, psychological disorders | Dementia | 1.30 × 10-7 | ACLY, ATP5A1, CANX, CAPN1, CAV1, CDK1, CTSB, DHX9, DNM1L, DPYSL2, EEF1G, EEF2, EIF2S1, EPRS, FDPS, G3BP1, GAPDH, GNB2L1, HNRNPA1, HNRNPA2B1, HNRNPU, HSPA5, HSPD1, LGALS1, PARK7, PCNA, PICALM, PRDX1, PRKDC, RAN, SET, SOD1, STIP1, TAGLN, TUBB, UBXN4, UCHL1, VCP, VIM, VPS35, YWHAZ | 41 | |
| Cancer, respiratory disease | Non-squamous non-small cell lung cancer | 1.39 × 10-7 | AHNAK, ALDOA, ALDOC, ATIC, CAPRIN1, CAV1, CD44, CDK1, DHX9, EEF1B2, EIF4A1, ENO1, EZR, G3BP1, GPI, HIST2H2AC, HLA-A, HSP90AA1, HSP90AB1, HSP90B1, IMPDH2, LDHA, LOC102724594/U2AF1, MAP4, MSN, MYH9, NACA, NSFL1C, PCNA, PFKP, PKM, PRDX1, PRPF19, SERBP1, TPI1, TUBB4B | 36 | |
| Skeletal and muscular disorders | Caveolinopathy | 1.42 × 10-7 | ACTA1, AHNAK, CALR, CANX, FLNC, HSP90B1, HSPA5, LMNA, MATR3, PSMA2, PSMA4, SQSTM1, VCL, VCP | 14 | |
| Cellular movement | Invasion of tumor cell lines | 1.49 × 10-7 | −0.733 | ACAT1, ANXA1, APEX1, ARHGDIA, BSG, CALR, CAP1, CAPNS1, CAV1, CD44, CSE1L, CTSB, CTTN, ENAH, EZR, FN1, GNB2L1, GPI, HDGF, HMGB1, HSP90AA1, ILF3, IQGAP1, ITGB1, KRT8, LGALS1, MAPK1, MARCKS, PA2G4, PEBP1, PKM, PTGES3, RHOA, RPSA, S100A6, SEPT9, SQSTM1, STMN1, TAGLN, TAGLN2, VCP, VIM, YWHAQ | 43 |
| Cellular movement | Cell movement of connective tissue cells | 1.54 × 10-7 | −0.391 | CAPN1, CAPN2, CAPNS1, CAV1, CD44, CYR61, EHD1, ENAH, EZR, FLNB, FN1, GNB2L1, HMGB1, IGF2R, ITGB1, LGALS1, MAPK1, MYH10, PLEC, STMN1, VCL, ZYX | 22 |
| Cancer | Follicular adenoma | 1.67 × 10-7 | ANXA5, CALR, CTSB, GSTP1, HSP90AB1, PARK7, PRDX2 | 7 | |
| Neurological disease, psychological disorders | Tauopathy | 1.70 × 10-7 | ACLY, ATP5A1, CANX, CAPN1, CAV1, CDK1, CTSB, DHX9, DNM1L, DPYSL2, EEF1G, EEF2, EIF2S1, EPRS, FDPS, G3BP1, GAPDH, GNB2L1, HNRNPA1, HNRNPA2B1, HNRNPU, HSPA5, HSPD1, LGALS1, PARK7, PCNA, PICALM, PRDX1, PRKDC, RAN, SET, SOD1, STIP1, TAGLN, TUBB, TUBB4B, UBXN4, UCHL1, VIM, VPS35, YWHAZ | 41 | |
| Cancer | Metastatic malignant solid tumor | 1.70 × 10-7 | ATIC, B2M, CAV1, CSE1L, EEF1A1, EIF4A1, FKBP1A, FN1, HSP90AA1, HSP90AB1, HSP90B1, MYL12A, PPIA, RPL27, RPL7, RPS11, RPS27A, TUBB, TUBB4B, YWHAE, YWHAZ | 21 | |
| Cell death and survival | Cell death of central nervous system cells | 1.85 × 10-7 | 0.374 | CAPN1, CAPNS1, CAST, CDK1, DNM1L, FUS, GAPDH, HMGB1, HSP90AA1, HSPA5, HSPB1, HSPD1, HYOU1, MAP1B, MAPK1, NPM1, P4HB, PARK7, PEA15, RHOA, RPS3, SOD1, STIP1, TCP1, VAPA, YWHAB | 26 |
| Molecular transport, protein trafficking | Import of protein | 2.01 × 10-7 | 0.816 | CFL1, DNAJA1, DNAJA2, IPO5, IPO7, IPO9, KPNA2, KPNB1, PDIA3, RAN, SPTBN1, XPO1 | 12 |
| Nucleic acid metabolism, small molecule biochemistry | Synthesis of purine nucleotide | 2.04 × 10-7 | −2.561 | ALDOA, ATP5A1, ATP5B, CAV1, CDK1, DNM1L, FASN, G6PD, GMPS, HMGB1, HSPD1, IMPDH2, NNMT, PKM, PPA1, SLC25A5, SOD1, VCP, VDAC1 | 19 |
| Cancer, organismal injury and abnormalities | Lymphatic neoplasia | 2.22 × 10-7 | 0.882 | ACTG1, ANXA1, ATIC, B2M, CALR, CD44, CFL1, CSE1L, CYR61, DNM1L, EEF1A1, FKBP1A, FN1, FUS, GMPS, GNB2L1, GSTP1, HIST1H1C, HMGB1, HSP90AA1, HSP90AB1, HSP90B1, HSPD1, IMPDH2, IQGAP1, ITGB1, KHDRBS1, KRT1, LDHA, LGALS1, LOC102724594/U2AF1, LONP1, MARS, MYH10, NNMT, NPM1, PICALM, PLOD2, PRDX1, PRKDC, PSMB1, PSMD2, PSME1, RHOA, RPL10, RPL3, RPL6, RPS12, RPS2, RPS24, RPS4X, SF3B1, SHMT2, SNRPD3, SPTBN1, STMN1, TUBB, TUBB4B, TUBB6, TUBB8, TXLNA, UQCRC1, XPO1, XRCC5, XRCC6, YWHAZ | 66 |
| Energy production, nucleic acid metabolism, small molecule biochemistry | Synthesis of ATP | 2.47 × 10-7 | −2.160 | ALDOA, ATP5B, CAV1, CDK1, DNM1L, HMGB1, HSPD1, NNMT, PKM, SLC25A5, SOD1, VCP, VDAC1 | 13 |
| Cell morphology, cellular assembly and organization, cellular development, cellular function and maintenance, nervous system development and function, tissue development | Morphogenesis of neurites | 2.71 × 10-7 | −0.815 | ACTR3, CAPNS1, CAPRIN1, CAPZB, CAV1, CD44, CSRP1, CYR61, DBN1, DNM1L, DPYSL2, EHD1, EZR, HMGB1, HNRNPK, ITGB1, LAMB1, MAP1B, MYH10, NNMT, PDIA3, PFN1, PHGDH, PICALM, RAB11A, RHOA, RTN4, SEPT2, SOD1, TNC, VAPA, YWHAH | 32 |
| Cell death and survival | Apoptosis of neurons | 2.84 × 10-7 | 1.726 | AARS, CAPN1, CAPRIN1, CAST, CDK1, CTSB, DNM1L, FN1, G6PD, GAPDH, GLUD1, GPI, HSD17B10, HSP90AB1, HSPA5, HSPB1, HSPD1, HYOU1, LGALS1, MAP1B, MAPK1, NPM1, P4HB, PARK7, PEA15, PRDX2, RHOA, RPS3, SET, SOD1, STAT1, XRCC5, XRCC6, YWHAB | 34 |
| Nucleic acid metabolism, small molecule biochemistry | Biosynthesis of nucleoside triphosphate | 2.93 × 10-7 | −2.160 | ALDOA, ATP5B, CAV1, CDK1, CTPS1, DNM1L, HMGB1, HSPD1, NNMT, PKM, SLC25A5, SOD1, VCP, VDAC1 | 14 |
| Cellular development, skeletal and muscular system development and function, tissue development | Differentiation of osteoblasts | 3.01 × 10-7 | −1.093 | ALYREF, ATP5B, CAPNS1, CLIC1, CLTC, CYR61, DDX5, DHX9, FASN, FN1, GNB2L1, H3F3A/H3F3B, HNRNPU, IARS, MAPK1, RPS11, RPS15, RRBP1, SND1, STAT1, STMN1, SYNCRIP, TNC, TPM4, VIM | 25 |
| Cell-to-cell signaling and interaction, cellular movement, hematological system development and function, immune cell trafficking, tissue development | Detachment of granulocytes | 3.22 × 10-7 | 0.254 | ANXA1, ANXA5, ITGB1, RHOA | 4 |
| Cellular movement, connective tissue development and function | Cell movement of fibroblasts | 3.30 × 10-7 | −0.142 | CAPN2, CAPNS1, CAV1, CD44, CYR61, EHD1, ENAH, EZR, FLNB, FN1, HMGB1, IGF2R, ITGB1, MAPK1, MYH10, PLEC, STMN1, VCL, ZYX | 19 |
| Cancer, organismal injury and abnormalities | Lymphoid cancer | 3.34 × 10-7 | 0.818 | ACTG1, ANXA1, ATIC, B2M, CALR, CD44, CFL1, CSE1L, CYR61, DNM1L, EEF1A1, FKBP1A, FN1, FUS, GMPS, GNB2L1, GSTP1, HIST1H1C, HMGB1, HSP90AA1, HSP90AB1, HSP90B1, HSPD1, IMPDH2, IQGAP1, ITGB1, KHDRBS1, KRT1, LDHA, LGALS1, LOC102724594/U2AF1, LONP1, MARS, MYH10, NNMT, NPM1, PICALM, PRDX1, PRKDC, PSMB1, PSMD2, PSME1, RHOA, RPL3, RPL6, RPS12, RPS2, RPS24, RPS4X, SF3B1, SHMT2, SNRPD3, SPTBN1, STMN1, TUBB, TUBB4B, TUBB6, TUBB8, TXLNA, UQCRC1, XPO1, XRCC5, XRCC6, YWHAZ | 64 |
| Metabolic disease, neurological disease, psychological disorders | Alzheimer’s disease | 3.37 × 10-7 | ACLY, ATP5A1, CANX, CAPN1, CAV1, CDK1, CTSB, DHX9, DNM1L, DPYSL2, EEF1G, EEF2, EIF2S1, EPRS, FDPS, G3BP1, GAPDH, GNB2L1, HNRNPA1, HNRNPA2B1, HNRNPU, HSPA5, HSPD1, LGALS1, PCNA, PICALM, PRDX1, PRKDC, RAN, SET, SOD1, STIP1, TAGLN, TUBB, UBXN4, UCHL1, VIM, VPS35, YWHAZ | 39 | |
| Cancer, tumor morphology | Invasion of tumor | 3.40 × 10-7 | −1.402 | AHCY, ANXA2, BSG, CAPN2, CAV1, CD44, CTSB, CTTN, EZR, FASN, FKBP1A, FLNA, FN1, HDLBP, HMGB1, HNRNPA1, HSPA5, ITGB1, LGALS1, MAPK1, PARK7, RHOA, VIM | 23 |
| Hereditary disorder, neurological disease | Autosomal dominant neuropathy | 4.17 × 10-7 | AARS, DYNC1H1, FUS, GARS, HSPB1, HSPD1, RAB7A, SOD1, UCHL1 | 9 | |
| Gene expression | Binding of DNA | 4.32 × 10-7 | −1.045 | ALYREF, APEX1, CALR, CAV1, CDK1, CFL1, FN1, GAPDH, GNB2L1, GSTP1, H3F3A/H3F3B, HMGB1, LGALS1, LMNA, MAPK1, NCL, NPM1, PCNA, PPIA, PRDX1, PRDX4, PRKDC, RHOA, SET, SFPQ, SND1, SOD1, STAT1, TAGLN, TRIM28, UBE2N, XRCC5, YBX1, YBX3, YWHAB, YWHAE, YWHAG, YWHAZ | 38 |
| Cell-to-cell signaling and interaction, tissue development | Adhesion of connective tissue cells | 5.10 × 10-7 | −1.487 | ANXA2, CALR, CD44, CYR61, FN1, IGF2R, IPO9, IQGAP1, ITGB1, MAPK1, PLEC, RHOA, RPL22, TLN1, TNC, VCL, ZYX | 17 |
| Cellular movement | Cell movement of fibrosarcoma cell lines | 5.20 × 10-7 | −1.621 | ARPC2, CTTN, FLNA, FLNB, FLNC, FN1, GPI, HNRNPK, TPM3 | 9 |
| Lipid metabolism, small molecule biochemistry | Binding of lipid | 5.48 × 10-7 | 0.823 | ANXA2, DNAJA1, FKBP4, FN1, HMGB1, HSP90AB1, MAP4, PTGES3, RPSA, SFN, STIP1, STMN1 | 12 |
| DNA replication, recombination, and repair | Double-stranded DNA break repair | 5.75 × 10-7 | 0.116 | CDK1, DDX1, FUS, KPNA2, NPM1, OTUB1, PCNA, PRKDC, PRPF19, TRIM28, UBE2N, VCP, XRCC5, XRCC6 | 14 |
| Cell signaling, DNA replication, recombination, and repair, nucleic acid metabolism, small molecule biochemistry | Hydrolysis of GTP | 5.79 × 10-7 | −1.580 | CALR, CDK1, DNM1L, GNAI2, IPO5, RAB7A, RAN, RANBP1, RHOA, STMN1, XPO1 | 11 |
| Cell death and survival | Cell death of breast cancer Cell lines | 6.10 × 10-7 | −0.856 | ARHGDIA, B2M, CAV1, CD44, CDK1, CSE1L, CYR61, FASN, FN1, HINT1, HSPA5, HSPB1, HSPD1, KPNA2, KRT18, MAPK1, PEA15, PEBP1, PRKDC, RHOA, SLC25A6, SND1, SSRP1, STAT1, STMN1, TFRC, VCP | 27 |
| Connective tissue disorders, skeletal and muscular disorders | Arthropathy | 7.25 × 10-7 | −0.277 | ACLY, ACTA1, ALDOA, ANXA1, ARF1, ATIC, B2M, CALR, CAPN2, CD44, CD59, CTSB, DDX39B, EEF1G, EEF2, ENO1, FASN, FDPS, FKBP1A, FN1, GNAI2, GPI, H3F3A/H3F3B, HLA-A, HLA-B, HMGB1, HNRNPA1, HNRNPA3, HSP90B1, HSPA5, HSPA8, HSPD1, KHSRP, LDHB, LGALS1, MAPRE1, MYL12A, NONO, PCM1, PDIA3, PGK1, PHB2, PRDX1, PRDX2, PRDX5, PTMA, RPS24, RPS3, RPSA, SND1, STAT1, TFRC, TPM2, TRIM28, TUBB, TUBB4B, UBA1, VARS, VIM | 59 |
| Cancer, gastrointestinal disease, hepatic system disease | Cholangiocarcinoma | 7.33 × 10-7 | ANXA1, ANXA2, GNB2L1, HSP90AA1, HSP90AB1, PGK1, PKM, RPL4, RPLP0, VIM | 10 | |
| Endocrine system disorders, gastrointestinal disease, immunological disease, inflammatory disease | Autoimmune pancreatitis | 7.47 × 10-7 | CALR, CAPG, COTL1, P4HB, PKM, PRDX2, UCHL1 | 7 | |
| Cellular assembly and organization, cellular compromise | Formation of cytoplasmic inclusions | 7.50 × 10-7 | FUS, KRT18, KRT8, SOD1, SQSTM1, UCHL1 | 6 | |
| Cell death and survival | Cell death of colon cancer Cell lines | 7.75 × 10-7 | 1.899 | AHSA1, CAPN2, CD44, CDK1, CSE1L, FASN, GSTP1, HINT1, HSPD1, IGF2R, ITGB1, KRT18, LMNA, PARK7, PPIA, RHOA, SFN, SFPQ, SQSTM1, STAT1, VCP, XRCC5, YWHAE, YWHAH | 24 |
| Cancer | Abdominal cancer | 8.21 × 10-7 | −0.593 | AARS, ACAT1, ACAT2, ACLY, ACTG1, ACTN1, ACTN4, ACTR2, ACTR3, AHCY, AHNAK, AKR1B1, ANXA1, ANXA2, ANXA5, AP1B1, APEX1, ARF1, ARHGDIA, ARPC2, ATIC, ATP5A1, ATP5B, B2M, BASP1, BSG, C14orf166, CACYBP, CALR, CAND1, CANX, CAP1, CAPG, CAPN2, CAST, CAV1, CBX3, CCT2, CCT4, CCT5, CCT6A, CCT8, CD44, CD59, CDK1, CLIC1, CLTC, COPA, COPB1, COPB2, COPE, COPG1, CORO1C, COTL1, CRIP2, CSE1L, CTNNA1, CTPS1, CTSA, CTSB, CTTN, CUTA, CYC1, CYR61, DBI, DBN1, DDX1, DDX39B, DDX3X, DDX5, DHX9, DLAT, DNAJA1, DNAJA2, DNAJC8, DNM1L, DPYSL2, DYNC1H1, DYNC1I2, ECHS1, EEF1A1, EEF1B2, EEF1D, EEF2, EIF2S1, EIF2S2, EIF3I, EIF4A1, EIF4G1, ENAH, ENO1, ENO2, EPRS, ESYT1, FASN, FDPS, FKBP1A, FLNA, FLNB, FLNC, FN1, FUBP3, FUS, G3BP1, G6PD, GANAB, GAPDH, GARS, GCN1L1, GDI2, GLUD1, GML, GMPS, GNAI2, GNB2L1, GPI, GSTP1, H2AFY, H3F3A/H3F3B, HADHA, HARS, HDGF, HDLBP, HEXB, HINT1, HIST1H1C, HIST1H2BL, HIST2H2AC, HLA-A, HLA-B, HMGB1, HN1, HNRNPA1, HNRNPA2B1, HNRNPA3, HNRNPH1, HNRNPH3, HNRNPK, HNRNPL, HNRNPM, HNRNPU, HSP90AA1, HSP90AB1, HSP90B1, HSPA5, HSPA8, HSPA9, HSPB1, HSPD1, HYOU1, IARS, IDI1, IFITM3, IGF2R, ILF2, ILF3, IMMT, IMPDH2, IPO5, IPO7, IPO9, IQGAP1, ITGB1, KHDRBS1, KHSRP, KIF5B, KPNA2, KPNA3, KPNB1, KRT1, KRT18, KRT8, KRT9, LAMB1, LDHA, LGALS1, LMNA, LMNB1, LOC102724594/U2AF1, LONP1, LRPPRC, LRRC59, MAP1B, MAP4, MAPK1, MAPRE1, MARS, MATR3, MCM3, MCM4, MCM6, MCM7, MDH1, MDH2, MSN, MVP, MYH10, MYH9, NACA, NAP1L1, NAP1L4, NASP, NCL, NNMT, NONO, NPEPPS, NPLOC4, NPM1, NSFL1C, NUMA1, OLA1, P4HB, PA2G4, PABPC1, PAICS, PCBP1, PCBP2, PCM1, PCNA, PDIA3, PDIA4, PDLIM1, PDXK, PEA15, PFKP, PFN1, PGAM1, PGD, PGK1, PGM1, PHGDH, PICALM, PKM, PLEC, PLIN3, PLOD2, PLS3, PPM1G, PPP1CA, PPP2CA, PPP2R1A, PRDX1, PRDX2, PRKCSH, PRKDC, PSMA1, PSMA7, PSMC1, PSMC4, PSMD2, PTBP1, PTMA, PTRF, PUF60, PYGB, PYGL, QARS, RAN, RANBP1, RARS, RBM3, RHOA, RPL12, RPL22, RPL4, RPL5, RPL6, RPL7A, RPL8, RPLP0, RPN1, RPN2, RPS12, RPS2, RPS24, RPS4X, RPS7, RRBP1, RTN4, S100A6, SDHA, SEPT9, SERBP1, SERPINH1, SET, SF3A1, SF3B1, SFPQ, SH3BGRL3, SHMT2, SLC25A3, SLC25A5, SLC38A2, SND1, SOD1, SPTAN1, SPTBN1, SQSTM1, SSRP1, STAT1, STIP1, SYNCRIP, TAGLN, TALDO1, TARS, TCP1, TFRC, TLN1, TMPO, TNC, TNPO1, TPI1, TPM2, TPM3, TPM4, TUBB, TUBB4B, TUBB8, TWF2, TXNDC5, UAP1, UBA1, UBE2N, UBXN4, UCHL1, VARS, VCL, VCP, VIM, VPS35, WARS, XPO1, XRCC5, XRCC6, YBX1, YBX3, YWHAE, YWHAG, YWHAH, YWHAQ, YWHAZ, ZYX | 342 |
| Drug metabolism, endocrine system development and function, lipid metabolism, small molecule biochemistry | Binding of progesterone | 8.87 × 10-7 | −0.277 | DNAJA1, FKBP4, HSP90AB1, PTGES3, STIP1 | 5 |
| Connective tissue disorders, inflammatory disease, skeletal and muscular disorders | Arthritis | 8.98 × 10-7 | −0.277 | ACLY, ACTA1, ALDOA, ANXA1, ARF1, ATIC, B2M, CALR, CAPN2, CD44, CD59, CTSB, DDX39B, EEF1G, EEF2, ENO1, FASN, FDPS, FKBP1A, FN1, GNAI2, GPI, H3F3A/H3F3B, HLA-A, HLA-B, HMGB1, HNRNPA1, HNRNPA3, HSP90B1, HSPA5, HSPA8, HSPD1, KHSRP, LDHB, LGALS1, MAPRE1, MYL12A, NONO, PCM1, PDIA3, PGK1, PHB2, PRDX1, PRDX2, PRDX5, PTMA, RPS24, RPS3, RPSA, SND1, STAT1, TFRC, TPM2, TRIM28, TUBB, TUBB4B, VARS, VIM | 58 |
| Cellular compromise, cellular function and maintenance | Endoplasmic reticulum stress response | 9.28 × 10-7 | 1.069 | AARS, CALR, CTSB, DNAJA1, HSP90AA1, HSP90AB1, HSP90B1, HSPA5, HSPD1, HYOU1, SERPINH1, SLC38A2, SQSTM1, UCHL1, VCP | 15 |
| Cell death and survival | Cell death of cerebral cortex cells | 9.41 × 10-7 | 1.181 | CAPN1, CAPNS1, CAST, CDK1, FUS, GAPDH, HMGB1, HSPA5, HSPB1, HSPD1, HYOU1, MAP1B, P4HB, PARK7, PEA15, RPS3, SOD1, STIP1, TCP1, VAPA, YWHAB | 21 |
| Connective tissue disorders, immunological disease, inflammatory disease, skeletal and muscular disorders | Rheumatoid arthritis | 9.82 × 10-7 | ACLY, ACTA1, ALDOA, ANXA1, ARF1, ATIC, B2M, CALR, CTSB, DDX39B, EEF1G, EEF2, ENO1, FDPS, FKBP1A, H3F3A/H3F3B, HLA-A, HMGB1, HNRNPA1, HNRNPA3, HSP90B1, HSPA8, HSPD1, KHSRP, LDHB, LGALS1, MAPRE1, MYL12A, NONO, PCM1, PDIA3, PGK1, PRDX1, PRDX2, PRDX5, PTMA, RPS24, RPS3, RPSA, SND1, STAT1, TFRC, TPM2, TRIM28, VARS, VIM | 46 | |
| Cellular assembly and organization, cellular function and maintenance | Bundling of microtubules | 1.03 × 10-6 | 0.218 | DNM1L, MAP1B, MAPRE1, NUMA1, RRBP1, SEPT9, SSRP1 | 7 |
| Cellular movement | Migration of breast cancer Cell lines | 1.21 × 10-6 | 0.280 | ANXA1, ANXA2, ARF1, CAPN2, CAV1, CSE1L, CTTN, FLNA, FN1, GNB2L1, ILF3, ITGB1, KHDRBS1, KPNA2, KRT8, LASP1, MAPK1, MYH9, RHOA, SFN, VIM | 21 |
| Cellular compromise | Collapse of intermediate filaments | 1.58 × 10-6 | KRT18, KRT8, PLEC, RHOA | 4 | |
| Connective tissue disorders, inflammatory disease, skeletal and muscular disorders | Rheumatic disease | 1.66 × 10-6 | −0.277 | ACLY, ACTA1, ALDOA, ANXA1, ARF1, ATIC, B2M, CALR, CAPN2, CD44, CD59, CTSA, CTSB, DDX39B, EEF1G, EEF2, ENO1, FASN, FDPS, FKBP1A, FN1, GNAI2, GPI, H3F3A/H3F3B, HLA-A, HLA-B, HMGB1, HNRNPA1, HNRNPA3, HSP90B1, HSPA5, HSPA8, HSPD1, KHSRP, LDHB, LGALS1, MAPRE1, MYL12A, NONO, PCM1, PDIA3, PGK1, PHB2, PPP2CA, PRDX1, PRDX2, PRDX5, PTMA, RHOA, RPS24, RPS3, RPSA, SND1, STAT1, TFRC, TPM2, TRIM28, TUBB, TUBB4B, UBE2L3, VARS, VIM | 62 |
| Organismal injury and abnormalities, reproductive system disease | Adenomyosis | 1.67 × 10-6 | ALDOA, ANXA2, DDX6, GPI, IQGAP1, ITGB1, LDHA, MYH10, PRDX5, VDAC1 | 10 | |
| DNA replication, recombination, and repair, energy production, nucleic acid metabolism, small molecule biochemistry | Hydrolysis of ATP | 1.72 × 10-6 | −0.164 | ATP1A1, CCT4, CCT5, CDK1, HMGB1, HSPA5, HSPD1, MAPK1, UBA1 | 9 |
| Cell-to-cell signaling and interaction, tissue development | Detachment of blood cells | 1.74 × 10-6 | 0.293 | ANXA1, ANXA5, FN1, ITGB1, RHOA | 5 |
| Cell death and survival | Cell death of muscle cells | 1.79 × 10-6 | 1.087 | ALDOA, CALR, CAPN1, CAV1, CYR61, EEF1A1, EEF1D, GAPDH, GNAI2, HADHA, HLA-B, HMGB1, HSPB1, HSPD1, LMNA, MAPK1, MDH1, NCL, PARK7, PLEC, PSMB1, RBM3, RHOA, RPSA, S100A6, STAT1, ZYX | 27 |
| Cell-to-cell signaling and interaction, tissue development | Adhesion of tumor cell lines | 1.86 × 10-6 | −0.235 | ANXA1, ANXA2, BSG, C1QBP, CAV1, CD44, CD59, CYR61, ERP29, FLNA, FN1, GNB2L1, HMGB1, ITGB1, MAPK1, MARCKS, MYH9, PTGES3, RHOA, RPSA, UCHL1, VCL, ZYX | 23 |
| Cardiovascular system development and function, cellular movement | Migration of endothelial cells | 1.90 × 10-6 | −2.222 | ANXA2, CAV1, CYR61, FLNA, FLNB, FN1, G6PD, HMGB1, HSP90AB1, HSPA5, HSPB1, IGF2R, ITGB1, LGALS1, MAPK1, MARCKS, NCL, PDCD6, PKM, RHOA, RTN4, SEPT7, TARS, VIM, WARS, YWHAZ | 26 |
| Cancer, cellular development, cellular growth and proliferation, tumor morphology | Proliferation of tumor cells | 2.04 × 10-6 | −1.588 | ACLY, ACTN4, AHCY, AKR1B1, ANXA1, ANXA2, BSG, CACYBP, CAV1, CD44, CTSB, CYR61, EEF1A1, EZR, FASN, FN1, HDGF, HMGB1, HSPA5, LDHA, LGALS1, MAPK1, NPM1, PARK7, PPP2CA, RHOA, RPS4X, S100A6, SQSTM1, STAT1, STMN1, TXLNA, UCHL1, XRCC5 | 34 |
| Cellular assembly and organization | Binding of zona pellucida | 2.10 × 10-6 | CCT2, CCT3, CCT4, CCT5, CCT6A, CCT7, CCT8, TCP1 | 8 | |
| Cellular function and maintenance | Engulfment of cells | 2.23 × 10-6 | −0.600 | ACTN4, ANXA1, ANXA5, CALR, CAPG, CAV1, CD44, CLIC4, CLTC, CORO1C, EHD1, EZR, FN1, GNB2L1, HMGB1, HYOU1, IQGAP1, ITGB1, MAP1B, MAPK1, MYH9, NCL, NPM1, PFN1, PICALM, RAB7A, RHOA, SFPQ, SNX3, VIM | 30 |
| Cancer, respiratory disease | Non small cell lung adenocarcinoma | 2.24 × 10-6 | AHNAK, ALDOA, ALDOC, ATIC, CAPRIN1, CAV1, CD44, CDK1, DHX9, EEF1B2, EIF4A1, ENO1, EZR, G3BP1, GPI, HIST2H2AC, HLA-A, IMPDH2, LDHA, LOC102724594/U2AF1, MAP4, MSN, MYH9, NACA, NSFL1C, PCNA, PFKP, PKM, PRDX1, PRPF19, SERBP1, TPI1, TUBB4B | 33 | |
| Cancer, hematological disease, immunological disease, organismal injury and abnormalities | Peripheral t-cell lymphoma | 2.47 × 10-6 | CYR61, FN1, NNMT, PSMB1, PSMD2, TUBB, TUBB4B, TUBB6, TUBB8 | 9 | |
| Cell death and survival | Cell death of brain | 2.52 × 10-6 | 0.904 | CAPN1, CAPNS1, CAST, CDK1, DNM1L, FUS, GAPDH, HMGB1, HSPA5, HSPB1, HSPD1, HYOU1, MAP1B, P4HB, PARK7, PEA15, RPS3, SDHA, SOD1, STAT1, STIP1, TCP1, VAPA, YWHAB | 24 |
| Cancer | Metastatic cancer | 2.58 × 10-6 | ATIC, B2M, CAPG, CAV1, CSE1L, DPYSL2, EEF1A1, EIF4A1, FDPS, FKBP1A, FLNA, FN1, HSP90AA1, HSP90AB1, HSP90B1, KRT18, MYL12A, PLOD2, PPIA, RPL27, RPL7, RPS11, RPS27A, STAT1, TUBB, TUBB4B, YWHAE | 27 | |
| Free radical scavenging | Production of reactive oxygen species | 2.60 × 10-6 | 0.372 | AKR1B1, ANXA1, ARHGDIA, CAV1, CD44, CTTN, DNM1L, FN1, G6PD, GNB2L1, HMGB1, HNRNPK, HSP90AB1, HSPA9, IMMT, LDHA, LONP1, MAPK1, PARK7, PPIA, PRDX1, PRDX2, RHOA, SOD1, SQSTM1, TAGLN, VDAC1, YWHAZ | 28 |
| Cellular movement | Migration of fibrosarcoma cell lines | 2.60 × 10-6 | −1.394 | ARPC2, CTTN, FLNA, FLNB, FLNC, FN1, HNRNPK, TPM3 | 8 |
| Cellular growth and proliferation | Outgrowth of cells | 2.75 × 10-6 | −0.580 | APEX1, BASP1, CDK1, CYR61, DNM1L, DPYSL2, EZR, FKBP4, FN1, HMGB1, HSP90AA1, IQGAP1, ITGB1, LGALS1, MAP1B, MAPK1, MARCKS, MYH10, MYH9, NPM1, PDIA3, RAB11A, RHOA, RTN4, SLC25A5, SNX3, SOD1, SPTBN1, TNC, VAPA, VIM, YWHAZ | 32 |
| Cancer, hematological disease, immunological disease | Waldenstrom’s macroglobulinemia | 2.79 × 10-6 | ANXA2, ANXA6, CAV1, CDK1, FASN, FKBP1A, FN1, FUS, G3BP1, KPNA2, NONO, PCNA, PPP2CA, PSMB1, PSMD2, STIP1, XRCC6, YWHAE | 18 | |
| Cancer, gastrointestinal disease | Upper gastrointestinal tract tumor | 2.84 × 10-6 | AKR1B1, ALYREF, ANXA1, ATIC, BSG, CANX, CAV1, CD44, COPB2, CSTB, CTNNA1, CTTN, DDX39B, EZR, FASN, FKBP1A, FN1, FUS, GSTP1, H3F3A/H3F3B, HDGF, HINT1, HLA-B, HNRNPH1, HSP90AA1, HSP90AB1, HSP90B1, IGF2R, IPO5, LGALS1, NPM1, P4HB, PA2G4, PCNA, PLOD2, PPP1CA, RAN, RHOA, RPL10, RPL22, SFN, TAGLN, TNC, TUBB, TUBB4B, XRCC5 | 46 | |
| Molecular transport, RNA trafficking | Transport of RNA | 3.04 × 10-6 | −1.969 | ALYREF, DDX39B, DDX3X, EIF5A, FUS, HNRNPA2B1, KHDRBS1, RAN, U2AF2, XPO1 | 10 |
| Cancer, hematological disease, immunological disease | Plasma cell dyscrasia | 3.09 × 10-6 | ANXA2, ANXA6, CAV1, CDK1, FASN, FDPS, FKBP1A, FLNA, FN1, FUS, G3BP1, HLA-A, KPNA2, NONO, PCNA, PDIA6, PPP2CA, PSMA2, PSMB1, PSMD2, STIP1, VIM, XRCC6, YWHAE, YWHAZ | 25 | |
| Cell cycle | Arrest in interphase | 3.10 × 10-6 | ANXA2, CAV1, CD44, CDK1, CSE1L, CYR61, FASN, FKBP1A, FLNA, FN1, ITGB1, LGALS1, LMNA, MAPK1, MCM7, PDCD6, PPP1CA, PRKDC, PTGES3, RHOA, RPL23, RPL5, RPL7A, SFN, SPTAN1, STAT1, TCP1, TFRC, TMPO, YWHAG | 30 | |
| Cell signaling, post-translational modification, protein synthesis | Actin capping of filament barbed ends | 3.12 × 10-6 | CAPG, CAPZA1, CAPZB, TWF1, TWF2 | 5 | |
| Cellular movement | Cell movement of bone cancer cell lines | 3.21 × 10-6 | −1.584 | ACTN4, CAPN2, CAV1, CTSB, FN1, STMN1, TLN1, VCP | 8 |
| Hematological system development and function, inflammatory response, tissue development | Aggregation of blood platelets | 3.42 × 10-6 | −0.977 | ACTG1, AKR1B1, CAPN1, CAST, CLIC1, CSRP1, FLNA, GNAI2, HSPB1, ITGB1, LGALS1, MAPK1, MYH9, MYL12A, P4HB, PDIA3, TLN1, VCL | 18 |
| Cancer | Cancer | 3.45 × 10-6 | −1.084 | AARS, ACAT1, ACAT2, ACLY, ACTG1, ACTN1, ACTN4, ACTR2, ACTR3, AHCY, AHNAK, AKR1B1, ALDOA, ALDOC, ANXA1, ANXA2, ANXA5, ANXA6, AP1B1, APEX1, ARF1, ARHGDIA, ARPC2, ATIC, ATP5A1, ATP5B, B2M, BASP1, BSG, C14orf166, C1QBP, CACYBP, CALR, CAND1, CANX, CAP1, CAPG, CAPN1, CAPN2, CAPRIN1, CAPZA1, CAST, CAV1, CBX3, CCT2, CCT3, CCT4, CCT5, CCT6A, CCT7, CCT8, CD44, CD59, CDK1, CFL1, CLIC1, CLTC, CNN3, COPA, COPB1, COPB2, COPE, COPG1, CORO1C, COTL1, CRIP2, CSE1L, CSTB, CTNNA1, CTPS1, CTSA, CTSB | 406 |
| CTTN, CUTA, CYB5R3, CYC1, CYR61, DBI, DBN1, DDX1, DDX39B, DDX3X, DDX5, DHX9, DLAT, DNAJA1, DNAJA2, DNAJC8, DNM1L, DPYSL2, DYNC1H1, DYNC1I2, ECHS1, EEF1A1, EEF1B2, EEF1D, EEF2, EHD1, EIF2S1, EIF2S2, EIF3B, EIF3C, EIF3I, EIF3M, EIF4A1, EIF4G1, EIF4H, ENAH, ENO1, ENO2, EPRS, ESYT1, ETFA, EZR, FASN, FDPS, FKBP1A, FLNA, FLNB, FLNC, FN1, FUBP3, FUS, G3BP1, G6PD, GANAB, GAPDH, GARS, GCN1L1, GDI1, GDI2, GLO1, GLUD1, GML, GMPS, GNAI2, GNB2L1, GPI, GSTP1, H2AFY, H3F3A/H3F3B, HADHA, HARS, HDGF, HDLBP, HEXB, HINT1, HIST1H1C, HIST1H2BL, HIST2H2AC, HLA-A, HLA-B, HMGB1, HN1, HNRNPA1, HNRNPA2B1, HNRNPA3, HNRNPH1, HNRNPH3, HNRNPK, HNRNPL, HNRNPM, HNRNPR, HNRNPU, HSP90AA1, HSP90AB1, HSP90B1, HSPA4, HSPA5, HSPA8, HSPA9, HSPB1, HSPD1, HYOU1, IARS, IDI1, IFITM3, IGF2R, ILF2, ILF3, IMMT, IMPDH2, IPO5, IPO7, IPO9, IQGAP1, ITGB1, KHDRBS1, KHSRP, KIF5B, KPNA2, KPNA3, KPNB1, KRT1, KRT18, KRT8, KRT9, LAMB1, LASP1, LDHA, LGALS1, LMNA, LMNB1, LOC102724594/U2AF1, LONP1, LRPPRC, LRRC59, MAP1B, MAP4, MAPK1, MAPRE1, MARCKS, MARS, MATR3, MCM3, MCM4, MCM6, MCM7, MDH1, MDH2, MSN, MVP, MYH10, MYH9, MYL12A, NACA, NAP1L1, NAP1L4, NASP, NCL, NNMT, NONO, NPEPPS, NPLOC4, NPM1, NSFL1C, NUMA1, OLA1, P4HB, PA2G4, PABPC1, PAFAH1B2, PAICS, PCBP1, PCBP2, PCM1, PCMT1, PCNA, PDCD6, PDE6H, PDIA3, PDIA4, PDIA6, PDLIM1, PDXK, PEA15, PFKP, PFN1, PGAM1, PGD, PGK1, PGM1, PHGDH, PICALM, PKM, PLEC, PLIN3, PLOD2, PLS3, PPIA, PPM1G, PPP1CA, PPP2CA, PPP2R1A, PRDX1, PRDX2, PRDX5, PRKCSH, PRKDC, PRMT5, PRPF19, PSMA1, PSMA2, PSMA7, PSMB1, PSMC1, PSMC4, PSMD2, PSME1, PTBP1, PTMA, PTRF, PUF60, PYGB, PYGL, QARS, RAN, RANBP1, RARS, RBM3, RCN1, RHOA, RNH1, RPL10, RPL12, RPL22, RPL27, RPL3, RPL4, RPL5, RPL6, RPL7, RPL7A, RPL8, RPLP0, RPN1, RPN2, RPS11, RPS12, RPS2, RPS24, RPS27A, RPS3, RPS3A, RPS4X, RPS5, RPS7, RRBP1, RTN4, S100A6, SDHA, SEPT9, SERBP1, SERPINH1, SET, SF3A1, SF3B1, SFN, SFPQ, SH3BGRL3, SHMT2, SLC25A3, SLC25A5, SLC25A6, SLC38A2, SND1, SNRPD3, SOD1, SPTAN1, SPTBN1, SQSTM1, SSRP1, STAT1, STIP1, STMN1, STOML2, SYNCRIP, TAGLN, TAGLN2, TALDO1, TARS, TCP1, TFRC, TLN1, TMPO, TNC, TNPO1, TPI1, TPM2, TPM3, TPM4, TUBA1B, TUBB, TUBB4B, TUBB6, TUBB8, TWF2, TXLNA, TXNDC5, UAP1, UBA1, UBE2N, UBXN4, UCHL1, UQCRC1, VARS, VAT1, VCL, VCP, VDAC2, VIM, VPS35, WARS, WDR1, XPO1, XRCC5, XRCC6, YBX1, YBX3, YWHAB, YWHAE, YWHAG, YWHAH, YWHAQ, YWHAZ, ZYX | |||||
| Cell morphology | Blebbing | 3.47 × 10-6 | 0.053 | CTTN, DPYSL2, EZR, HSPB1, LMNA, LMNB1, MARCKS, RHOA, SPTAN1 | 9 |
| Hereditary disorder, neurological disease, organismal injury and abnormalities, skeletal and muscular disorders | Charcot-Marie-tooth disease type 2 | 3.75 × 10-6 | AARS, DYNC1H1, GARS, HSPB1, LMNA, RAB7A | 6 | |
| Endocrine system development and function, small molecule biochemistry | Binding of hormone | 3.75 × 10-6 | 0.250 | DNAJA1, FKBP4, HSP90AB1, PTGES3, SFN, STIP1 | 6 |
| Cancer, hematological disease, immunological disease, organismal injury and abnormalities | Classic Hodgkin disease | 3.75 × 10-6 | FKBP1A, LGALS1, TUBB, TUBB4B, TUBB6, TUBB8 | 6 | |
| Cellular assembly and organization, cellular function and maintenance | Stabilization of microtubules | 3.80 × 10-6 | 0.247 | COPB2, DNM1L, DPYSL2, IQGAP1, MAP1B, MAP4, MAPRE1, NUMA1, PKM, RHOA, SEPT7, STMN1 | 12 |
| Cell morphology, cellular function and maintenance | Transmembrane potential of mitochondria | 4.05 × 10-6 | −0.508 | ANXA6, B2M, CLIC1, CLIC4, HSPA4, HSPB1, HSPD1, IMMT, LDHA, LGALS1, LONP1, PARK7, PHB2, SLC25A6, SOD1, STOML2, VCP, VDAC1, YWHAE | 19 |
| Cancer, gastrointestinal disease | Digestive organ tumor | 4.26 × 10-6 | −0.321 | AARS, ACAT1, ACAT2, ACLY, ACTG1, ACTN1, ACTR2, ACTR3, AHCY, AHNAK, AKR1B1, ALYREF, ANXA1, ANXA2, AP1B1, APEX1, ARF1, ARHGDIA, ATIC, ATP5A1, ATP5B, B2M, BSG, CACYBP, CANX, CAPG, CAPN2, CAST, CAV1, CBX3, CCT4, CCT5, CCT6A, CCT8, CD44, CD59, CLIC1, CLTC, COPA, COPB2, COPG1, COTL1, CRIP2, CSE1L, CSTB, CTNNA1, CTPS1, CTSA, CTSB, CTTN, CUTA, CYC1, CYR61, DBI, DBN1, DDX1, DDX39B, DDX3X, DDX5, DHX9, DLAT, DNAJA1, DNAJC8, DNM1L, DPYSL2, DYNC1H1, DYNC1I2, ECHS1, EEF1A1, EEF1D, EEF2, EIF3I, EIF4A1, EIF4G1, ENAH, ENO1, ENO2, EPRS, ESYT1, EZR, FASN, FKBP1A, FLNA, FLNB, FLNC, FN1, FUBP3, FUS, G3BP1, G6PD, GANAB, GARS, GCN1L1, GDI2, GLUD1, GML, GMPS, GNAI2, GNB2L1, GPI, GSTP1, H2AFY, H3F3A/H3F3B, HADHA, HDGF, HDLBP, HEXB, HINT1, HIST1H1C, HLA-A, HLA-B, HMGB1, HNRNPA1, HNRNPA2B1, HNRNPA3, HNRNPH1, HNRNPH3, HNRNPK, HNRNPL, HNRNPM, HNRNPU, HSP90AA1, HSP90AB1, HSP90B1, HSPA5, HSPA8, HSPA9, HSPD1, HYOU1, IARS, IDI1, IFITM3, IGF2R, ILF2, ILF3, IMMT, IMPDH2, IPO5, IPO7, IQGAP1, ITGB1, KHDRBS1, KHSRP, KIF5B, KPNA2, KPNA3, KPNB1, KRT1, KRT18, KRT8, KRT9, LAMB1, LDHA, LGALS1, LMNA, LMNB1, LOC102724594/U2AF1, LONP1, LRPPRC, LRRC59, MAP1B, MAP4, MAPRE1, MARS, MATR3, MCM3, MCM4, MCM6, MCM7, MDH1, MDH2, MVP, MYH10, MYH9, NACA, NAP1L1, NAP1L4, NASP, NCL, NNMT, NPEPPS, NPLOC4, NPM1, NUMA1, OLA1, P4HB, PA2G4, PAICS, PCBP1, PCBP2, PCM1, PCNA, PDIA4, PFKP, PFN1, PGAM1, PGD, PGK1, PGM1, PHGDH, PKM, PLEC, PLIN3, PLOD2, PLS3, PPM1G, PPP1CA, PPP2CA, PPP2R1A, PRDX1, PRKCSH, PRKDC, PSMA7, PSMC1, PSMD2, PTBP1, PTMA, PTRF, PUF60, PYGB, PYGL, QARS, RAN, RANBP1, RARS, RHOA, RPL10, RPL12, RPL22, RPL4, RPL5, RPL7A, RPLP0, RPN1, RPN2, RPS2, RPS24, RPS4X, RRBP1, RTN4, S100A6, SDHA, SEPT9, SERBP1, SERPINH1, SET, SF3A1, SF3B1, SFN, SFPQ, SH3BGRL3, SLC38A2, SND1, SOD1, SPTAN1, SPTBN1, SQSTM1, SSRP1, STAT1, STIP1, STMN1, SYNCRIP, TAGLN, TALDO1, TARS, TCP1, TFRC, TLN1, TMPO, TNC, TNPO1, TPI1, TPM2, TPM4, TUBB, TUBB4B, TUBB8, TWF2, UAP1, UBA1, UBE2N, UBXN4, UCHL1, VARS, VCL, VCP, VIM, VPS35, WARS, XRCC5, XRCC6, YBX1, YBX3, YWHAE, YWHAG, YWHAH, YWHAZ | 297 |
| Cellular assembly and organization, tissue development | Polymerization of filaments | 4.45 × 10-6 | 0.934 | ACTR3, ARPC2, CAPZB, CAV1, CFL1, FKBP4, MAP1B, MAPRE1, PFN1, RHOA, STMN1, TUBB, TWF1, TWF2 | 14 |
| Cellular assembly and organization, cellular development, cellular growth and proliferation, nervous system development and function, tissue development | Growth of neurites | 4.55 × 10-6 | −1.344 | APEX1, BASP1, CAV1, CDK1, CSRP1, DNM1L, DPYSL2, EZR, FKBP4, FN1, HMGB1, IQGAP1, ITGB1, LGALS1, MAP1B, MAPK1, MARCKS, MYH10, MYH9, PDIA3, RAB11A, RHOA, RTN4, SEPT9, SLC25A5, SNX3, SOD1, SPTBN1, TNC, VAPA, VCL, VIM, YWHAZ | 33 |
| Hematological disease | Blood protein disorder | 4.60 × 10-6 | ANXA2, ANXA6, ARHGDIA, B2M, CAV1, CDK1, FASN, FDPS, FKBP1A, FLNA, FN1, FUS, G3BP1, HLA-A, KPNA2, NONO, PCNA, PDIA6, PPP2CA, PSMA2, PSMB1, PSMD2, STIP1, VIM, XRCC6, YWHAE, YWHAZ | 27 | |
| Hereditary disorder | Autosomal dominant disease | 4.66 × 10-6 | AARS, ACTA1, ACTG1, ACTN4, AHNAK, CAV1, DNM1L, DYNC1H1, EEF2, EIF4G1, FKBP1A, FLNB, FLNC, FN1, FUS, GARS, HNRNPA1, HNRNPA2B1, HSPB1, HSPD1, KRT1, KRT9, LMNA, LMNB1, MYH9, PARK7, RAB7A, SOD1, STAT1, TNC, TPM2, TPM3, UCHL1, VCP, VPS35 | 35 | |
| Cancer, immunological disease, organismal injury and abnormalities | Lymphatic node tumor | 4.71 × 10-6 | 0.000 | ANXA1, ATIC, B2M, CSE1L, CYR61, FKBP1A, FN1, FUS, HIST1H1C, HMGB1, IQGAP1, ITGB1, KHDRBS1, LONP1, NNMT, PRDX1, PRKDC, PSMB1, PSMD2, RHOA, RPS2, SF3B1, SHMT2, STMN1, TUBB, TUBB4B, TUBB6, TUBB8, TXLNA, UQCRC1, XPO1, XRCC5, YWHAZ | 33 |
| Molecular transport | Nuclear export | 4.97 × 10-6 | ALYREF, CALR, CSE1L, DDX39B, EIF5A, HNRNPA1, HSPA9, KHDRBS1, NPM1, RAN, XPO1 | 11 | |
| Cell death and survival | Apoptosis of colon cancer cell lines | 5.12 × 10-6 | 1.294 | AHSA1, CD44, CDK1, CSE1L, FASN, GSTP1, HINT1, HSPD1, IGF2R, ITGB1, KRT18, PARK7, PPIA, RHOA, SFN, SFPQ, STAT1, VCP, XRCC5, YWHAE | 20 |
| Cellular movement | Cell movement of pericytes | 5.37 × 10-6 | −0.928 | CD44, FN1, GNB2L1, HMGB1, LGALS1, MAPK1, MYH10 | 7 |
| Molecular transport, protein synthesis, protein trafficking | Localization of protein | 5.46 × 10-6 | −0.324 | APEX1, ARHGDIA, BTF3, C1QBP, CAV1, CUTA, FKBP1A, FKBP4, GNB2L1, HLA-A, HYOU1, ITGB1, NPM1, PICALM, SEPT2, SQSTM1, TFRC | 17 |
| Cellular movement | Migration of connective tissue cells | 5.46 × 10-6 | 0.420 | CAPNS1, CAV1, CD44, CYR61, EHD1, FLNB, FN1, GNB2L1, HMGB1, ITGB1, LGALS1, MAPK1, MYH10, PLEC, STMN1, VCL, ZYX | 17 |
| Hereditary disorder, skeletal and muscular disorders | Autosomal dominant myopathy | 6.79 × 10-6 | AARS, AHNAK, DYNC1H1, GARS, HSPB1, LMNA, RAB7A | 7 | |
| Neurological disease | Lower motor neuron disease | 6.79 × 10-6 | DYNC1H1, GARS, HNRNPA1, HNRNPA2B1, HSPB1, UBA1, VCP | 7 | |
| Cancer, endocrine system disorders | Thyroid adenoma | 6.79 × 10-6 | ANXA5, CALR, CTSB, GSTP1, HSP90AB1, PARK7, PRDX2 | 7 | |
| Cell death and survival | Apoptosis of cervical cancer cell lines | 7.05 × 10-6 | −0.772 | CCT4, CDK1, CTSB, DNM1L, DYNC1H1, EZR, FLNB, HNRNPK, IMMT, KHDRBS1, MAPK1, MSN, PCBP2, PPIA, PRKDC, PRPF19, PTMA, RPS24, STAT1, TCP1, UCHL1, VDAC1 | 22 |
| Cell morphology | Shape change of tumor cell lines | 7.13 × 10-6 | −1.937 | ANXA5, CAP1, CAPN2, DPYSL2, FLNA, FLNB, FN1, ITGB1, MAPK1, MARCKS, MYH9, PFN1, RHOA, TNC, UCHL1 | 15 |
| Energy production, molecular transport, nucleic acid metabolism, small molecule biochemistry | Concentration of ATP | 7.22 × 10-6 | −1.019 | ALDOA, ATP5B, CAV1, DNM1L, ENO1, HSD17B10, KRT8, LDHA, LRPPRC, PKM, SQSTM1, STOML2, VDAC1 | 13 |
| Cell death and survival | Cell death of brain cells | 7.42 × 10-6 | 1.086 | CAPN1, CAPNS1, CAST, CDK1, DNM1L, FUS, GAPDH, HMGB1, HSPA5, HSPB1, HSPD1, HYOU1, MAP1B, P4HB, PARK7, PEA15, RPS3, SOD1, STIP1, TCP1, VAPA, YWHAB | 22 |
| Cancer, gastrointestinal disease | Digestive tract cancer | 7.58 × 10-6 | 0.345 | AARS, ACAT1, ACAT2, ACLY, ACTG1, ACTN1, ACTR2, ACTR3, AHCY, AHNAK, AKR1B1, ANXA1, ANXA2, AP1B1, APEX1, ARF1, ARHGDIA, ATIC, ATP5A1, ATP5B, B2M, BSG, CACYBP, CANX, CAPG, CAPN2, CAST, CAV1, CBX3, CCT4, CCT5, CCT6A, CCT8, CD44, CD59, CLIC1, CLTC, COPA, COPB2, COPG1, COTL1, CRIP2, CSE1L, CSTB, CTNNA1, CTPS1, CTSA, CTSB, CTTN, CUTA, CYC1, CYR61, DBI, DBN1, DDX1, DDX39B, DDX3X, DDX5, DHX9, DLAT, DNAJA1, DNAJC8, DNM1L, DPYSL2, DYNC1H1, DYNC1I2, ECHS1, EEF1A1, EEF1D, EEF2, EIF3I, EIF4A1, EIF4G1, ENAH, ENO1, ENO2, EPRS, ESYT1, EZR, FASN, FKBP1A, FLNA, FLNB, FLNC, FN1, FUBP3, FUS, G3BP1, G6PD, GANAB, GARS, GCN1L1, GDI2, GLUD1, GML, GMPS, GNAI2, GNB2L1, GPI, GSTP1, H2AFY, H3F3A/H3F3B, HADHA, HDGF, HDLBP, HEXB, HINT1, HIST1H1C, HLA-A, HLA-B, HMGB1, HNRNPA1, HNRNPA2B1, HNRNPA3, HNRNPH1, HNRNPH3, HNRNPK, HNRNPL, HNRNPM, HNRNPU, HSP90AA1, HSP90AB1, HSP90B1, HSPA5, HSPA8, HSPA9, HSPD1, HYOU1, IARS, IDI1, IFITM3, IGF2R, ILF2, ILF3, IMMT, IMPDH2, IPO5, IPO7, IQGAP1, ITGB1, KHDRBS1, KHSRP, KIF5B, KPNA2, KPNA3, KPNB1, KRT1, KRT18, KRT8, KRT9, LAMB1, LDHA, LGALS1, LMNA, LMNB1, LOC102724594/U2AF1, LONP1, LRPPRC, LRRC59, MAP1B, MAP4, MAPRE1, MARS, MATR3, MCM3, MCM4, MCM6, MCM7, MDH1, MDH2, MVP, MYH10, MYH9, NACA, NAP1L1, NAP1L4, NASP, NCL, NNMT, NPEPPS, NPLOC4, NPM1, NUMA1, OLA1, P4HB, PA2G4, PAICS, PCBP1, PCBP2, PCM1, PCNA, PDIA4, PFKP, PFN1, PGAM1, PGD, PGK1, PGM1, PHGDH, PKM, PLEC, PLIN3, PLOD2, PLS3, PPM1G, PPP1CA, PPP2CA, PPP2R1A, PRDX1, PRKCSH, PRKDC, PSMA7, PSMC1, PSMD2, PTBP1, PTMA, PTRF, PUF60, PYGB, PYGL, QARS, RAN, RANBP1, RARS, RHOA, RPL10, RPL12, RPL22, RPL4, RPL5, RPL7A, RPLP0, RPN1, RPN2, RPS2, RPS24, RPS4X, RRBP1, RTN4, S100A6, SDHA, SEPT9, SERBP1, SERPINH1, SET, SF3A1, SF3B1, SFN, SFPQ, SH3BGRL3, SLC38A2, SND1, SOD1, SPTAN1, SPTBN1, SQSTM1, SSRP1, STAT1, STIP1, SYNCRIP, TAGLN, TALDO1, TARS, TCP1, TFRC, TLN1, TMPO, TNC, TNPO1, TPI1, TPM2, TPM4, TUBB, TUBB4B, TUBB8, TWF2, UAP1, UBA1, UBE2N, UBXN4, UCHL1, VARS, VCL, VCP, VIM, VPS35, WARS, XRCC6, YBX1, YBX3, YWHAE, YWHAG, YWHAH, YWHAZ | 294 |
| Cancer, gastrointestinal disease | Malignant neoplasm of digestive system | 7.58 × 10-6 | 0.345 | AARS, ACAT1, ACAT2, ACLY, ACTG1, ACTN1, ACTR2, ACTR3, AHCY, AHNAK, AKR1B1, ANXA1, ANXA2, AP1B1, APEX1, ARF1, ARHGDIA, ATIC, ATP5A1, ATP5B, B2M, BSG, CACYBP, CANX, CAPG, CAPN2, CAST, CAV1, CBX3, CCT4, CCT5, CCT6A, CCT8, CD44, CD59, CLIC1, CLTC, COPA, COPB2, COPG1, COTL1, CRIP2, CSE1L, CSTB, CTNNA1, CTPS1, CTSA, CTSB, CTTN, CUTA, CYC1, CYR61, DBI, DBN1, DDX1, DDX39B, DDX3X, DDX5, DHX9, DLAT, DNAJA1, DNAJC8, DNM1L, DPYSL2, DYNC1H1, DYNC1I2, ECHS1, EEF1A1, EEF1D, EEF2, EIF3I, EIF4A1, EIF4G1, ENAH, ENO1, ENO2, EPRS, ESYT1, EZR, FASN, FKBP1A, FLNA, FLNB, FLNC, FN1, FUBP3, FUS, G3BP1, G6PD, GANAB, GARS, GCN1L1, GDI2, GLUD1, GML, GMPS, GNAI2, | 294 |
| GNB2L1, GPI, GSTP1, H2AFY, H3F3A/H3F3B, HADHA, HDGF, HDLBP, HEXB, HINT1, HIST1H1C, HLA-A, HLA-B, HMGB1, HNRNPA1, HNRNPA2B1, HNRNPA3, HNRNPH1, HNRNPH3, HNRNPK, HNRNPL, HNRNPM, HNRNPU, HSP90AA1, HSP90AB1, HSP90B1, HSPA5, HSPA8, HSPA9, HSPD1, HYOU1, IARS, IDI1, IFITM3, IGF2R, ILF2, ILF3, IMMT, IMPDH2, IPO5, IPO7, IQGAP1, ITGB1, KHDRBS1, KHSRP, KIF5B, KPNA2, KPNA3, KPNB1, KRT1, KRT18, KRT8, KRT9, LAMB1, LDHA, LGALS1, LMNA, LMNB1, LOC102724594/U2AF1, LONP1, LRPPRC, LRRC59, MAP1B, MAP4, MAPRE1, MARS, MATR3, MCM3, MCM4, MCM6, MCM7, MDH1, MDH2, MVP, MYH10, MYH9, NACA, NAP1L1, NAP1L4, NASP, NCL, NNMT, NPEPPS, NPLOC4, NPM1, NUMA1, OLA1, P4HB, PA2G4, PAICS, PCBP1, PCBP2, PCM1, PCNA, PDIA4, PFKP, PFN1, PGAM1, PGD, PGK1, PGM1, PHGDH, PKM, PLEC, PLIN3, PLOD2, PLS3, PPM1G, PPP1CA, PPP2CA, PPP2R1A, PRDX1, PRKCSH, PRKDC, PSMA7, PSMC1, PSMD2, PTBP1, PTMA, PTRF, PUF60, PYGB, PYGL, QARS, RAN, RANBP1, RARS, RHOA, RPL10, RPL12, RPL22, RPL4, RPL5, RPL7A, RPLP0, RPN1, RPN2, RPS2, RPS24, RPS4X, RRBP1, RTN4, S100A6, SDHA, SEPT9, SERBP1, SERPINH1, SET, SF3A1, SF3B1, SFN, SFPQ, SH3BGRL3, SLC38A2, SND1, SOD1, SPTAN1, SPTBN1, SQSTM1, SSRP1, STAT1, STIP1, SYNCRIP, TAGLN, TALDO1, TARS, TCP1, TFRC, TLN1, TMPO, TNC, TNPO1, TPI1, TPM2, TPM4, TUBB, TUBB4B, TUBB8, TWF2, UAP1, UBA1, UBE2N, UBXN4, UCHL1, VARS, VCL, VCP, VIM, VPS35, WARS, XRCC6, YBX1, YBX3, YWHAE, YWHAG, YWHAH, YWHAZ | |||||
| Molecular transport | Export of molecule | 7.90 × 10-6 | −1.118 | ALYREF, ANXA6, CALR, CSE1L, DDX39B, EIF5A, HSPA9, KHDRBS1, RAN, SEC13, U2AF2, XPO1, YWHAE | 13 |
| Cellular assembly and organization | Quantity of filaments | 8.03 × 10-6 | −0.914 | CFL1, FN1, GNB2L1, KRT18, MAP1B, MAP4, MARCKS, PLEC, RHOA, SERPINH1, SPTAN1, STMN1, TMOD3, VIM | 14 |
| Cell-to-cell signaling and interaction, reproductive system development and function | Binding of sperm | 8.32 × 10-6 | CCT2, CCT3, CCT4, CCT5, CCT6A, CCT7, CCT8, TCP1 | 8 | |
| Connective tissue disorders, developmental disorder, metabolic disease, skeletal and muscular disorders | Paget’s disease of bone | 8.37 × 10-6 | FDPS, HNRNPA1, HNRNPA2B1, SQSTM1, VCP | 5 | |
| Cancer, hematological disease, immunological disease, organismal injury and abnormalities | Refractory classical hodgkin lymphoma | 8.37 × 10-6 | FKBP1A, TUBB, TUBB4B, TUBB6, TUBB8 | 5 | |
| Cancer, hematological disease, immunological disease, organismal injury and abnormalities | Relapsed classical hodgkin lymphoma | 8.37 × 10-6 | FKBP1A, TUBB, TUBB4B, TUBB6, TUBB8 | 5 | |
| Infectious disease | Infection of hepatoma cell lines | 8.85 × 10-6 | −0.425 | ACTR2, AP1B1, AP2B1, ARPC1B, CLTC, G3BP1, HMGB1, IFITM3, TFRC | 9 |
| Cancer, gastrointestinal disease | Upper gastrointestinal tract cancer | 8.86 × 10-6 | ANXA1, ATIC, BSG, CANX, CAV1, CD44, COPB2, CSTB, CTNNA1, CTTN, DDX39B, EZR, FASN, FKBP1A, FN1, FUS, GSTP1, H3F3A/H3F3B, HDGF, HINT1, HLA-B, HNRNPH1, HSP90AA1, HSP90AB1, HSP90B1, IGF2R, IPO5, LGALS1, NPM1, P4HB, PA2G4, PLOD2, RAN, RHOA, RPL10, RPL22, SFN, TAGLN, TNC, TUBB, TUBB4B | 41 | |
| Hematological disease, organismal injury and abnormalities | Myelodysplastic syndrome | 9.44 × 10-6 | CALR, LOC102724594/U2AF1, NPM1, PDIA3, RPL3, RPL4, RPL5, RPL6, RPL7, RPLP0, RPLP1, RPS27A, RPS4X, SF3B1 | 14 | |
| Cancer | Osteosarcoma | 9.56 × 10-6 | CD44, FASN, FDPS, HINT1, HSP90AA1, HSP90AB1, HSP90B1, IMPDH2, PRDX1, STOML2, TUBB, TUBB4B | 12 | |
| Cellular function and maintenance | Uptake of cells | 9.91 × 10-6 | 0.555 | C1QBP, CAV1, ITGB1, NCL, OTUB1, RHOA, RPSA, SNX3 | 8 |
| Cellular development, cellular growth and proliferation | Proliferation of carcinoma Cell lines | 1.06 × 10-5 | −0.759 | ACLY, ANXA1, ANXA2, BSG, C1QBP, CAV1, CYR61, DYNC1H1, GAPDH, H2AFY, HMGB1, HNRNPA2B1, IGF2R, ITGB1, MAPK1, MAPRE1, PTBP1, S100A6, SET, SLC25A6, SOD1, STMN1, TRIM28, UCHL1 | 24 |
| Cell death and survival | Apoptosis of breast cancer Cell lines | 1.16 × 10-5 | −1.394 | ARHGDIA, B2M, CAV1, CD44, CDK1, CSE1L, CYR61, FASN, FN1, HINT1, HSPA5, HSPB1, HSPD1, KPNA2, KRT18, MAPK1, PEA15, PEBP1, SND1, STAT1, TFRC, VCP | 22 |
| Cell-to-cell signaling and interaction, connective tissue development and function, tissue development | Adhesion of fibroblast Cell lines | 1.18 × 10-5 | −1.591 | CALR, CD44, CYR61, FN1, IQGAP1, ITGB1, MAPK1, RHOA, VCL | 9 |
| Cellular assembly and organization, cellular compromise | Formation of cellular inclusion bodies | 1.21 × 10-5 | 0.246 | CTSB, DNAJA1, FUS, HSPA4, KRT18, KRT8, PSMC4, SOD1, SQSTM1, TCP1, UCHL1 | 11 |
| Cell-to-cell signaling and interaction | Binding of lymphoma cell lines | 1.31 × 10-5 | −0.881 | ANXA2, CD44, FN1, ITGB1, NCL, RHOA, TFRC | 7 |
| Organismal injury and abnormalities, renal and urological disease | Nephrosis | 1.35 × 10-5 | ACTN4, ANXA1, ARHGDIA, CLTC, FKBP1A, IMPDH2, ITGB1, LAMB1, PDLIM1 | 9 | |
| Cancer, gastrointestinal disease, respiratory disease | CDKN2A positive oropharyngeal squamous cell carcinoma | 1.35 × 10-5 | HSP90AA1, HSP90AB1, HSP90B1 | 3 | |
| Cancer, respiratory disease | Advanced stage primary laryngeal cancer | 1.35 × 10-5 | HSP90AA1, HSP90AB1, HSP90B1 | 3 | |
| Gene expression | Binding of Y box | 1.35 × 10-5 | APEX1, YBX1, YBX3 | 3 | |
| Connective tissue disorders, developmental disorder, hereditary disorder, metabolic disease, skeletal and muscular disorders | Familial Paget’s disease of bone | 1.35 × 10-5 | HNRNPA1, HNRNPA2B1, VCP | 3 | |
| Cellular movement | Initiation of migration of fibrosarcoma cell lines | 1.35 × 10-5 | FLNA, FLNB, FLNC | 3 | |
| Cancer, organismal injury and abnormalities, reproductive system disease | Locally advanced cervical cancer | 1.35 × 10-5 | HSP90AA1, HSP90AB1, HSP90B1 | 3 | |
| Cancer, immunological disease, organismal injury and abnormalities, reproductive system disease | Node positive cervical cancer | 1.35 × 10-5 | HSP90AA1, HSP90AB1, HSP90B1 | 3 | |
| Cell morphology, cellular assembly and organization, cellular development, cellular growth and proliferation, nervous system development and function, tissue development | Outgrowth of neurites | 1.36 × 10-5 | −1.232 | APEX1, BASP1, CDK1, DNM1L, DPYSL2, EZR, FKBP4, FN1, HMGB1, IQGAP1, ITGB1, LGALS1, MAP1B, MAPK1, MARCKS, MYH10, MYH9, PDIA3, RAB11A, RHOA, RTN4, SLC25A5, SNX3, SOD1, SPTBN1, TNC, VAPA, VIM, YWHAZ | 29 |
| Cancer, tumor morphology | Invasion of malignant tumor | 1.38 × 10-5 | −2.111 | AHCY, BSG, CAPN2, CD44, CTSB, CTTN, EZR, FASN, HMGB1, HNRNPA1, HSPA5, ITGB1, LGALS1, MAPK1, VIM | 15 |
| Cellular assembly and organization, cellular function and maintenance | Quantity of cellular protrusions | 1.38 × 10-5 | 0.905 | ACTG1, ACTR2, ARPC2, BSG, CANX, CAPZB, CTTN, ENAH, EZR, FN1, MAP1B, MSN, SOD1, TLN1, TPM3 | 15 |
| Cell cycle | Interphase | 1.57 × 10-5 | 0.828 | ANXA2, CAPNS1, CAV1, CD44, CDK1, CSE1L, CYR61, DDX3X, FASN, FKBP1A, FLNA, FN1, GNB2L1, GPI, ITGB1, LGALS1, LMNA, MAPK1, MCM7, NASP, NPM1, PCNA, PDCD6, PEA15, PPP1CA, PRKDC, PTGES3, RHOA, RPL23, RPL5, RPL7A, SFN, SPTAN1, STAT1, TCP1, TFRC, TMPO, XRCC5, YWHAG, YWHAQ | 40 |
| Cell morphology, cellular function and maintenance | Repair of cells | 1.62 × 10-5 | 0.131 | APEX1, KPNA2, NPM1, OTUB1, PCNA, PRKDC, UBE2N, XRCC5, XRCC6, YBX1 | 10 |
| Protein synthesis | Oligomerization of protein | 1.63 × 10-5 | ACAT1, AHNAK, ANXA5, ANXA6, CAV1, CTNNA1, CUTA, EEF1A1, EHD1, LONP1, NPM1, PRKCSH, SEPT7, SEPT9, SQSTM1, STOML2, TRIM28, VCP, YWHAB | 19 | |
| Cellular movement | Migration of pericytes | 1.68 × 10-5 | −0.600 | CD44, FN1, GNB2L1, HMGB1, LGALS1, MYH10 | 6 |
| Cellular development, nervous system development and function, tissue development | Development of neurons | 1.85 × 10-5 | −0.428 | ACTR3, BASP1, CAPNS1, CAPRIN1, CAPZB, CAV1, CD44, CSRP1, CYR61, DBN1, DNM1L, DPYSL2, EHD1, ENAH, EZR, GDI1, HMGB1, HNRNPK, HSPB1, ITGB1, LAMB1, MAP1B, MSN, MYH10, NNMT, PDIA3, PFN1, PHGDH, PICALM, PPP2CA, PRKCSH, RAB11A, RHOA, RTN4, SEPT2, SOD1, SPTBN1, STIP1, STMN1, TNC, UCHL1, VAPA, VIM, YWHAG, YWHAH | 45 |
| Hereditary disorder, neurological disease, organismal injury and abnormalities, skeletal and muscular disorders | Autosomal dominant Charcot-Marie-Tooth disease type 2 | 1.88 × 10-5 | AARS, DYNC1H1, GARS, HSPB1, RAB7A | 5 | |
| Cell morphology, hair and skin development and function | Cell spreading of epithelial cell lines | 1.88 × 10-5 | −1.446 | CYR61, FLNA, FN1, ITGB1, VIM | 5 |
| Cell morphology, cellular function and maintenance, DNA replication, recombination, and repair | Double-stranded DNA break repair of cells | 1.91 × 10-5 | 0.131 | KPNA2, NPM1, OTUB1, PCNA, PRKDC, UBE2N, XRCC5, XRCC6 | 8 |
| Cell death and survival | Apoptosis of central nervous system cells | 1.96 × 10-5 | −0.963 | CAPN1, CAST, CDK1, DNM1L, GAPDH, HSP90AA1, HSPA5, MAP1B, NPM1, P4HB, PEA15, RHOA, RPS3, SOD1, YWHAB | 15 |
| Cell cycle | Arrest in G1 phase | 1.97 × 10-5 | CYR61, FASN, FKBP1A, FN1, ITGB1, LGALS1, LMNA, MAPK1, MCM7, PPP1CA, PRKDC, PTGES3, RHOA, RPL23, RPL5, RPL7A, SPTAN1, TFRC, TMPO, YWHAG | 20 | |
| Cellular function and maintenance | Cellular homeostasis | 1.98 × 10-5 | 0.822 | AKR1B1, ALDOA, ANXA1, ANXA2, ANXA6, ARF1, ATP1A1, B2M, BSG, CALR, CAPN1, CAPNS1, CAST, CAV1, CLIC1, CLIC4, CLTC, CTTN, CYR61, DNM1L, EEF1D, EIF2S1, EIF4G1, FKBP1A, FN1, GPI, H2AFY, HADH, HEXB, HMGB1, HNRNPL, HSP90AA1, HSP90B1, HSPA4, HSPA5, HSPB1, HSPD1, IGF2R, IMMT, ITGB1, KIF5B, KRT18, KRT8, LDHA, LGALS1, LONP1, MAPK1, PAFAH1B2, PARK7, PGM1, PHB2, PICALM, PPIA, PPP1CA, PPP2CA, PRDX2, PRKDC, PSMD2, PYGL, RAB11A, RAB7A, RPL22, SEPT9, SERPINH1, SLC25A6, SOD1, SQSTM1, STAT1, STOML2, TFRC, TUFM, UBE2N, VCP, VDAC1, XRCC5, XRCC6, YWHAE | 77 |
| Tissue development | Growth of connective tissue | 2.07 × 10-5 | 0.381 | ANXA2, ATIC, CAPNS1, CAV1, CD44, CLTC, CTSB, CYR61, DDX5, DNM1L, FN1, GNAI2, GPI, GSTP1, HDGF, HNRNPA2B1, HSPB1, ITGB1, KHDRBS1, LGALS1, LMNA, LMNB1, MAPK1, MYH10, MYH9, NPM1, PA2G4, PGK1, PRDX4, PRKDC, PTGES3, RBM3, RHOA, SERPINH1, SFN, STAT1, STMN1, TPM3, VCL, YBX1 | 40 |
| Post-translational modification, protein folding | Folding maturation of protein | 2.09 × 10-5 | AIP, CALR, FKBP1A, PRDX4 | 4 | |
| Cellular movement | Invasion of breast cancer cell lines | 2.24 × 10-5 | 0.051 | BSG, CALR, CAV1, CD44, CSE1L, CTSB, CTTN, ENAH, EZR, FN1, HSP90AA1, ILF3, IQGAP1, KRT8, MAPK1, PTGES3, RHOA, SEPT9, STMN1 | 19 |
| Cellular function and maintenance, inflammatory response | Phagocytosis | 2.34 × 10-5 | −0.334 | ANXA1, ANXA5, CALR, CAPG, CAV1, CD44, CLIC4, CLTC, CORO1C, EHD1, GNB2L1, HMGB1, ITGB1, MAPK1, MSN, MYH9, NPM1, PFN1, RAB7A, RHOA, SNX3, VIM | 22 |
| Cellular development, cellular growth and proliferation, connective tissue development and function, tissue development | Proliferation of fibroblasts | 2.34 × 10-5 | 0.428 | ANXA2, CAV1, CD44, DDX5, DNM1L, FN1, GPI, GSTP1, HDGF, HSPB1, ITGB1, LMNA, LMNB1, MAPK1, NPM1, PA2G4, PGK1, PRDX4, PRKDC, PTGES3, RBM3, RHOA, STAT1, VCL, YBX1 | 25 |
| Protein trafficking | Targeting of protein | 2.35 × 10-5 | AIP, CALR, CSE1L, EIF5A, HSPA9, RAB7A, RAN, XPO1, YWHAB, YWHAE, YWHAG, YWHAQ, YWHAZ | 13 | |
| Hematological system development and function, tissue development | Aggregation of blood cells | 2.36 × 10-5 | −1.292 | ACTG1, AKR1B1, CAPN1, CAST, CD44, CLIC1, CSRP1, FLNA, GNAI2, HSPB1, ITGB1, LGALS1, MAPK1, MYH9, MYL12A, P4HB, PDIA3, TLN1, VCL | 19 |
| Cellular movement, connective tissue development and function | Cell movement of fibroblast Cell lines | 2.39 × 10-5 | −0.697 | ACTR2, ACTR3, ARPC2, CD44, CLIC4, CTTN, DDX3X, EHD1, FN1, GPI, HMGB1, ITGB1, MAPRE1, NPM1, RHOA | 15 |
| Cellular development, tissue development | Differentiation of bone cells | 2.43 × 10-5 | −1.093 | ALYREF, ATP5B, CAPNS1, CLIC1, CLTC, CYR61, DDX5, DHX9, FASN, FN1, GLO1, GNB2L1, H3F3A/H3F3B, HNRNPU, IARS, MAPK1, RPS11, RPS15, RRBP1, SND1, SQSTM1, STAT1, STMN1, SYNCRIP, TFRC, TNC, TPM4, VIM | 28 |
| Cancer, hematological disease, immunological disease, organismal injury and abnormalities | Lymphocytic cancer | 2.48 × 10-5 | 1.450 | ANXA1, ATIC, B2M, CD44, CSE1L, CYR61, FKBP1A, FN1, HIST1H1C, HMGB1, IQGAP1, LGALS1, LONP1, NNMT, NPM1, PRDX1, PRKDC, PSMB1, PSMD2, RHOA, RPS2, SF3B1, SHMT2, SPTBN1, STMN1, TUBB, TUBB4B, TUBB6, TUBB8, TXLNA, UQCRC1, XPO1, XRCC5, XRCC6, YWHAZ | 35 |
| Cancer, cellular development, cellular growth and proliferation, tumor morphology | Proliferation of cancer cells | 2.63 × 10-5 | −1.887 | AHCY, AKR1B1, BSG, CACYBP, CAV1, CD44, CTSB, CYR61, EEF1A1, EZR, FASN, HDGF, HMGB1, LDHA, LGALS1, MAPK1, NPM1, PPP2CA, RHOA, RPS4X, S100A6, SQSTM1, STAT1, STMN1, TXLNA, XRCC5 | 26 |
| Cell-to-cell signaling and interaction | Fusion of cells | 2.74 × 10-5 | 1.580 | ANXA1, ANXA5, CAPN2, CAV1, CD44, CTSB, FLNC, LGALS1, MAPK1, MYH9, RHOA, SOD1, STAT1 | 13 |
| Cell morphology, cellular assembly and organization, cellular development, cellular function and maintenance, nervous system development and function, tissue development | Branching of neurons | 2.75 × 10-5 | −0.900 | ACTR3, BASP1, CAPNS1, CAPZB, CAV1, CSRP1, CYR61, DBN1, DNM1L, DPYSL2, EZR, HMGB1, HNRNPK, ITGB1, MAP1B, NNMT, PDIA3, PFN1, RHOA, RTN4, SOD1 | 21 |
| Cell morphology | Cell spreading of tumor Cell lines | 2.96 × 10-5 | −1.711 | CAP1, CAPN2, FLNA, FLNB, FN1, ITGB1, MARCKS, PFN1, RHOA, TNC, UCHL1 | 11 |
| Dermatological diseases and conditions, immunological disease, inflammatory disease | Lichen planus | 2.96 × 10-5 | B2M, CFL1, EEF1A1, EIF5A, ENO1, FKBP1A, HLA-A, HLA-B, IFITM3, PFN1, PSME2 | 11 | |
| Cellular assembly and organization, cellular function and maintenance | Formation of vesicles | 3.07 × 10-5 | 0.707 | ANXA2, ANXA5, ARF1, CAST, CLTC, FLNA, IQGAP1, MARCKS, PICALM, RAB11A, RHOA, SNX3 | 12 |
| Immunological disease | Systemic autoimmune syndrome | 3.10 × 10-5 | ACLY, ACTA1, ALDOA, ANXA1, ARF1, ATIC, B2M, CALR, CD44, CTSA, CTSB, DDX39B, EEF1G, EEF2, ENAH, ENO1, FDPS, FKBP1A, H3F3A/H3F3B, HLA-A, HLA-B, HMGB1, HNRNPA1, HNRNPA3, HSP90B1, HSPA8, HSPD1, KHSRP, LDHB, LGALS1, MAPRE1, MYH10, MYH9, MYL12A, NONO, PCM1, PDIA3, PGK1, PGM1, PPP2CA, PRDX1, PRDX2, PRDX5, PTMA, RPS24, RPS3, RPSA, S100A6, SND1, STAT1, TFRC, TPM2, TRIM28, TUBB, UBE2L3, VARS, VIM | 57 | |
| Cell morphology, cellular assembly and organization, cellular function and maintenance | Formation of filopodia | 3.11 × 10-5 | −2.546 | ACTN4, ACTR3, CAPNS1, CSRP1, CYR61, EEF1A1, ENAH, EZR, FN1, IQGAP1, ITGB1, MAP1B, MARCKS, RHOA, TNC | 15 |
| Cell-to-cell signaling and interaction, cellular assembly and organization, cellular function and maintenance, tissue development | Formation of focal adhesions | 3.12 × 10-5 | −2.245 | ACTN1, CAPN1, CAV1, CTTN, EHD1, EZR, FN1, GNAI2, GNB2L1, ITGB1, RHOA, STMN1, VCL, VIM | 14 |
| Cellular growth and proliferation, tissue development | Proliferation of connective tissue cells | 3.14 × 10-5 | 0.548 | ANXA2, ATIC, CAPNS1, CAV1, CD44, CLTC, CTSB, CYR61, DDX5, DNM1L, FN1, GNAI2, GPI, GSTP1, HDGF, HSPB1, ITGB1, KHDRBS1, LGALS1, LMNA, LMNB1, MAPK1, NPM1, PA2G4, PGK1, PRDX4, PRKDC, PTGES3, RBM3, RHOA, SERPINH1, SFN, STAT1, STMN1, TPM3, VCL, YBX1 | 37 |
| Hematological disease, immunological disease | Eosinophilia | 3.45 × 10-5 | ALDOA, ENO1, GAPDH, HLA-A, HLA-B, HSPA5, MYH9, P4HB, PDIA3, PRDX1, TKT, TPI1, TUBB | 13 | |
| Cellular assembly and organization, cellular function and maintenance | Organization of actin filaments | 3.56 × 10-5 | −1.633 | ALDOA, CFL1, DBN1, DPYSL2, ENAH, FLNA, FN1, ITGB1, MSN, PLS3, RHOA | 11 |
| Cancer | Benign neoplasia | 3.63 × 10-5 | 1.591 | ACAT1, AIP, ALDOA, ALDOC, ANXA1, ANXA2, ANXA5, CALR, CD44, CTSB, CYR61, DBN1, DDX5, DDX6, FKBP1A, GSTP1, HINT1, HSD17B10, HSP90AB1, IFITM3, IQGAP1, KPNA3, LGALS1, MAPK1, MDH1, MYH10, NPEPPS, PARK7, PCNA, PEA15, PRDX1, PRDX2, PRDX5, RPL27, RPS15, SF3A1, SPTBN1, TNC, TWF1, VCP, ZYX | 41 |
| Cancer, gastrointestinal disease | Oral cancer | 3.66 × 10-5 | ANXA1, ATIC, BSG, EZR, FASN, FN1, H3F3A/H3F3B, HSP90AA1, HSP90AB1, HSP90B1, LGALS1, NPM1, PLOD2, RPL10, TAGLN, TNC, TUBB, TUBB4B | 18 | |
| Cell death and survival | Cell death of lymphoma Cell lines | 3.67 × 10-5 | 1.961 | ANXA2, ARHGDIA, CD59, EZR, GLO1, HNRNPA1, HSPB1, LGALS1, LMNB1, LONP1, MAPK1, MSN, NCL, RAD23B, RPLP0, YWHAG, YWHAZ | 17 |
| Cell cycle, cell morphology, cellular assembly and organization, cellular movement | Elongation of actin filaments | 3.68 × 10-5 | −1.969 | ACTN4, CAP1, FN1, PFN1 | 4 |
| Infectious disease | Infection by Marburg virus | 3.68 × 10-5 | −0.000 | HSPA5, RPL18, RPL3, RPL5 | 4 |
| Carbohydrate metabolism, nucleic acid metabolism | Pentose shunt of monosaccharide | 3.68 × 10-5 | G6PD, PGD, TALDO1, TKT | 4 | |
| Cancer, hematological disease, immunological disease, organismal injury and abnormalities | Non-Hodgkin’s disease | 3.72 × 10-5 | ANXA1, ATIC, B2M, CSE1L, CYR61, FKBP1A, FN1, HIST1H1C, HMGB1, IQGAP1, LONP1, NNMT, PRDX1, PRKDC, PSMB1, PSMD2, RHOA, RPS2, SF3B1, SHMT2, STMN1, TUBB, TUBB4B, TUBB6, TUBB8, TXLNA, UQCRC1, XPO1, XRCC5, YWHAZ | 30 | |
| Organismal injury and abnormalities | Nodule | 3.73 × 10-5 | ANXA5, CALR, CTSB, GSTP1, HMGB1, HSP90AB1, PARK7, PRDX2, PRDX5 | 9 | |
| Cell death and survival | Cell death of cortical neurons | 3.77 × 10-5 | 0.671 | CAPN1, CAPNS1, CAST, CDK1, FUS, GAPDH, HMGB1, HSPA5, HSPB1, HSPD1, MAP1B, PARK7, PEA15, TCP1, YWHAB | 15 |
| Cell morphology, connective tissue development and function | Morphology of fibroblast cell lines | 3.85 × 10-5 | CAV1, CTTN, DPY30, FN1, KRT18, KRT8, MARCKS, PTBP1, RHOA, RTN4 | 10 | |
| Infectious disease | Replication of flaviviridae | 3.85 × 10-5 | −0.905 | DDX3X, DDX6, DNAJA1, FASN, G3BP1, HMGB1, IFITM3, MVP, PPIA, YBX1 | 10 |
| Cell death and survival | Cell viability of neuroblastoma cell lines | 3.96 × 10-5 | 0.537 | APEX1, HSP90AB1, HSPB1, P4HB, S100A6, SOD1, SQSTM1, VCP | 8 |
| Cellular compromise | Stress response of cells | 4.00 × 10-5 | 0.174 | CALR, CTSB, DNAJA1, HNRNPA1, HSD17B10, HSP90B1, HSPA5, PRDX1, SLC38A2, SOD1, SQSTM1, VCP, YBX1 | 13 |
| Cancer, respiratory disease | Laryngeal tumor | 4.00 × 10-5 | ANXA1, CD44, HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 7 | |
| DNA replication, recombination, and repair | DNA damage | 4.01 × 10-5 | 1.228 | CALR, FN1, G6PD, GNAI2, GSTP1, HSPB1, LONP1, NPM1, PARK7, PDIA3, PRDX1, PRKDC, SOD1, SQSTM1, XRCC5, YWHAE | 16 |
| Cellular development, cellular growth and proliferation, nervous system development and function, tissue development | Proliferation of neuronal cells | 4.31 × 10-5 | −1.654 | APEX1, B2M, BASP1, CAV1, CDK1, CFL1, CSRP1, CTNNA1, DNM1L, DPYSL2, EZR, FKBP4, FN1, HMGB1, IQGAP1, ITGB1, LGALS1, MAP1B, MAPK1, MARCKS, MYH10, MYH9, PDIA3, RAB11A, RHOA, RTN4, SEPT9, SLC25A5, SNX3, SOD1, SPTBN1, TNC, VAPA, VCL, VIM, YWHAZ | 36 |
| Cellular movement | Cell movement of hepatic stellate cells | 4.34 × 10-5 | −0.600 | CD44, GNB2L1, HMGB1, LGALS1, MAPK1, MYH10 | 6 |
| Cancer, respiratory disease | Laryngeal squamous Cell carcinoma | 4.34 × 10-5 | CD44, HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 6 | |
| Cellular movement | Migration of bone cancer cell lines | 4.34 × 10-5 | −1.091 | ACTN4, CAPN2, CAV1, FN1, STMN1, VCP | 6 |
| Cell morphology | Shape change of epithelial cell lines | 4.34 × 10-5 | −1.000 | CYR61, FLNA, FN1, ITGB1, MAPK1, VIM | 6 |
| Molecular transport, protein trafficking | Nuclear transport of protein | 4.54 × 10-5 | CALR, CSE1L, EIF5A, HSPA9, KPNB1, RAN, TNPO1, XPO1 | 8 | |
| Cancer | Growth of malignant tumor | 4.65 × 10-5 | −1.175 | AHCY, AKR1B1, BSG, CACYBP, CAV1, CD44, CTSB, CYR61, EEF1A1, EZR, FASN, GNB2L1, HDGF, HMGB1, LDHA, LGALS1, MAPK1, NPM1, PKM, PPP2CA, RHOA, RPS4X, S100A6, SQSTM1, STAT1, STMN1, TNC, TXLNA, XRCC5 | 29 |
| Cell morphology, cellular assembly and organization, cellular development, cellular function and maintenance, embryonic development, nervous system development and function, tissue development | Branching of neurites | 4.69 × 10-5 | −0.668 | ACTR3, CAPNS1, CAPZB, CAV1, CSRP1, CYR61, DBN1, DNM1L, DPYSL2, EZR, HMGB1, HNRNPK, ITGB1, MAP1B, NNMT, PDIA3, PFN1, RHOA, RTN4, SOD1 | 20 |
| Cell-to-cell signaling and interaction, cellular function and maintenance, inflammatory response | Phagocytosis of cells | 4.69 × 10-5 | −0.618 | ANXA1, ANXA5, CALR, CAPG, CAV1, CD44, CLIC4, CLTC, CORO1C, EHD1, GNB2L1, HMGB1, ITGB1, MAPK1, MYH9, NPM1, PFN1, RHOA, SNX3, VIM | 20 |
| Cell-to-cell signaling and interaction | Binding of stem cells | 5.32 × 10-5 | C1QBP, ITGB1, TLN1 | 3 | |
| Tissue morphology | Collapse of epithelial tissue | 5.32 × 10-5 | KRT18, KRT8, RHOA | 3 | |
| Cell-to-cell signaling and interaction, tissue development | Growth of focal adhesions | 5.32 × 10-5 | FLNA, FLNB, VIM | 3 | |
| Molecular transport, protein trafficking | Import of green fluorescent protein | 5.32 × 10-5 | KPNA2, KPNB1, RAN | 3 | |
| Cell-to-cell signaling and interaction, hair and skin development and function | Binding of epithelial cell lines | 5.36 × 10-5 | −0.294 | ANXA2, ANXA5, CAV1, ITGB1, PPP2CA, PRMT5 | 6 |
| Cancer, gastrointestinal disease, respiratory disease | Oropharyngeal tumor | 5.36 × 10-5 | HSP90AA1, HSP90AB1, HSP90B1, PCNA, TUBB, TUBB4B | 6 | |
| Protein degradation, protein synthesis | Catabolism of protein | 5.52 × 10-5 | 0.237 | CALR, CANX, CAPN1, CAPN2, CAPNS1, CAST, CAV1, COPG1, CSTB, CTSB, FLNA, GAPDH, HSP90B1, HSPA5, HSPD1, IPO9, ITGB1, LONP1, MYH9, NACA, NPEPPS, PARK7, PDIA3, PPP2CA, PSMC4, PSMD2, SNX3, SOD1, SQSTM1, TPP1, UBE2L3, UBE2N, UBXN4, UCHL1, VCP, XPO1 | 36 |
| Cell cycle | Arrest in cell cycle progression | 5.60 × 10-5 | C1QBP, CALR, CAV1, CD44, FASN, FKBP1A, KHDRBS1, LMNA, MAP4, MAPK1, NPM1, PA2G4, PCM1, PPM1G, PRMT5, RHOA, SFN, SSRP1, STAT1, TCP1, TNC, XRCC6, YWHAE | 23 | |
| Infectious disease | Infection by flaviviridae | 5.71 × 10-5 | 0.460 | ACTR2, AP1B1, AP2B1, ARPC1B, CLTC, G3BP1, HMGB1, IFITM3, STAT1, TFRC | 10 |
| Cell cycle | G1 phase | 5.92 × 10-5 | −0.221 | CAPNS1, CDK1, CYR61, FASN, FKBP1A, FN1, GNB2L1, GPI, ITGB1, LGALS1, LMNA, MAPK1, MCM7, NASP, PPP1CA, PRKDC, PTGES3, RHOA, RPL23, RPL5, RPL7A, SPTAN1, TFRC, TMPO, YWHAG, YWHAQ | 26 |
| Cancer, hematological disease, immunological disease, organismal injury and abnormalities | CD30-positive peripheral t-Cell lymphoma | 6.02 × 10-5 | TUBB, TUBB4B, TUBB6, TUBB8 | 4 | |
| Cardiovascular system development and function, organ morphology | Contraction of left ventricle | 6.02 × 10-5 | 0.849 | CAPNS1, ITGB1, LMNA, RHOA | 4 |
| Cellular assembly and organization, cellular function and maintenance | Formation of caveolae | 6.02 × 10-5 | −1.009 | ANXA6, CAV1, PICALM, PTRF | 4 |
| Cancer, hematological disease, immunological disease, organismal injury and abnormalities | Systemic large-cell KI-1 lymphoma | 6.02 × 10-5 | TUBB, TUBB4B, TUBB6, TUBB8 | 4 | |
| Cancer, gastrointestinal disease | Stomach tumor | 6.30 × 10-5 | AKR1B1, ALYREF, ANXA1, CANX, CAV1, CD44, COPB2, CTNNA1, DDX39B, FKBP1A, FUS, GSTP1, HDGF, HINT1, HNRNPH1, IGF2R, IPO5, P4HB, PA2G4, RHOA, RPL22, TUBB4B, XRCC5 | 23 | |
| Cell morphology, cellular function and maintenance, DNA replication, recombination, and repair | Double-stranded DNA break repair of tumor cell lines | 6.57 × 10-5 | 0.878 | KPNA2, NPM1, OTUB1, UBE2N, XRCC5, XRCC6 | 6 |
| Cancer | Head and neck cancer | 6.57 × 10-5 | 1.756 | ACTN1, ANXA1, ATIC, BSG, CAV1, CD44, CTTN, DDX3X, EZR, FASN, FLNA, FN1, GSTP1, H3F3A/H3F3B, HNRNPK, HSP90AA1, HSP90AB1, HSP90B1, HSPA5, LDHA, LGALS1, MARCKS, MCM7, NPM1, PCM1, PKM, PLOD2, PRDX1, PRKDC, PRMT5, RPL10, SET, SF3B1, SFN, SPTBN1, STAT1, TAGLN, TNC, TUBB, TUBB4B, VIM, XRCC5, XRCC6 | 43 |
| Cellular assembly and organization, cellular function and maintenance | Formation of actin cytoskeleton | 6.71E-05 | CORO1C, ERP29, FLNA, FN1, ITGB1 | 5 | |
| Gene expression | Transcription | 7.48 × 10-5 | 0.968 | ACTR2, ACTR3, ALYREF, BASP1, BTF3, C14orf166, C1QBP, CALR, CAND1, CAV1, CBX3, CD44, CDK1, CSE1L, CYR61, DDX3X, DDX5, DHX9, EEF1D, EIF2S1, ENO1, FKBP1A, FLNA, FN1, FUBP3, GLO1, GSTP1, H2AFY, HDGF, HEXB, HINT1, HIST1H1C, HMGB1, HNRNPA1, HNRNPA2B1, HNRNPK, HSPA8, ILF2, ILF3, IQGAP1, KHDRBS1, KPNA2, LGALS1, LMNA, LRPPRC, MAPK1, MATR3, MCM7, NONO, NPM1, OTUB1, PA2G4, PARK7, PCNA, PDLIM1, PEA15, PEBP1, PFN1, PHB2, PHGDH, PICALM, PPP2CA, PRKDC, PRMT5, PTGES3, PTMA, PTRF, RHOA, RPL12, RPL6, SET, SFPQ, SQSTM1, SSRP1, STAT1, SUPT16H, TAGLN2, TMPO, TRIM28, UBE2L3, VAPA, XPO1, XRCC5, XRCC6, YBX1, YBX3, YWHAB, YWHAH, YWHAQ, YWHAZ | 90 |
| Cell-to-cell signaling and interaction, connective tissue development and function, tissue development | Adhesion of fibroblasts | 7.57 × 10-5 | −0.896 | CD44, FN1, IGF2R, PLEC, TNC, VCL, ZYX | 7 |
| Organismal injury and abnormalities, skeletal and muscular disorders | Neurogenic muscular atrophy | 7.57 × 10-5 | AARS, DYNC1H1, GARS, HINT1, HSPB1, LMNA, RAB7A | 7 | |
| DNA replication, recombination, and repair | DNA replication | 7.63 × 10-5 | −1.248 | CACYBP, CALR, CAV1, CDK1, HMGB1, MAPK1, MCM7, NAP1L1, NASP, NCL, NPM1, PCM1, PCNA, PEA15, SET, SSRP1, SUPT16H, TMPO, XRCC5 | 19 |
| Molecular transport | Nuclear export of molecule | 8.10 × 10-5 | ALYREF, CALR, CSE1L, DDX39B, EIF5A, HSPA9, KHDRBS1, RAN, XPO1 | 9 | |
| Organismal injury and abnormalities, renal and urological disease | Chronic kidney disease | 8.13 × 10-5 | CDV3, DBI, EEF1B2, FDPS, FKBP1A, H3F3A/H3F3B, HMGB1, HSP90AA1, IMPDH2, MYH9, NPM1, PSMB1, PSMD2, RPL23, RPL7, TUBB, TUBB4B | 17 | |
| Cellular movement | Cell movement of melanoma cell lines | 8.37 × 10-5 | −0.364 | ARF1, BSG, CAPN1, FLNA, ITGB1, KHDRBS1, KRT8, MSN, PEBP1, PRDX2, RHOA | 11 |
| Cell death and survival | Cytolysis | 8.37 × 10-5 | 0.280 | ALDOA, ANXA1, B2M, CALR, CAV1, CD44, CD59, G6PD, GPI, HLA-A, IMPDH2, KRT18, LGALS1, PRDX1, PRDX2, PSME2, PTMA, SH3BGRL3, STAT1, STMN1 | 20 |
| Cell morphology, cellular function and maintenance | Transmembrane potential | 8.37 × 10-5 | −0.508 | ANXA6, B2M, CLIC1, CLIC4, HSPA4, HSPB1, HSPD1, IMMT, KIF5B, LDHA, LGALS1, LONP1, PARK7, PHB2, SLC25A6, SOD1, STOML2, VCP, VDAC1, YWHAE | 20 |
| Protein synthesis | Initiation of translation of protein | 8.60E-05 | EIF2S1, EIF3B, EIF3C, EIF3I, EIF4G1, EIF4H, HSPB1, RPS3A | 8 | |
| Hematological disease, infectious disease | Infection by Zaire ebolavirus | 8.77 × 10-5 | −2.216 | CTSB, HSPA5, RPL18, RPL3, RPL5 | 5 |
| Cellular assembly and organization | Formation of nucleus | 8.78 × 10-5 | −2.364 | CDK1, FLNA, FN1, GNAI2, LMNA, LMNB1, RAN | 7 |
| Cellular assembly and organization, cellular function and maintenance, tissue development | Polymerization of microtubules | 8.78 × 10-5 | 1.141 | CAPZB, CAV1, FKBP4, MAP1B, MAPRE1, STMN1, TUBB | 7 |
| Inflammatory response | Immune response of cells | 9.25 × 10-5 | −1.018 | ANXA1, ANXA5, CALR, CAPG, CAV1, CD44, CD59, CLIC4, CLTC, CORO1C, CTSB, DNM1L, EHD1, FN1, GNB2L1, HLA-A, HMGB1, HSP90AA1, HSPB1, ITGB1, LGALS1, MAPK1, MYH9, NPM1, OTUB1, PFN1, PSME2, RHOA, SF3A1, SNX3, STAT1, TNPO1, UBE2L3, VIM | 34 |
| Cell-to-cell signaling and interaction, tissue development | Adhesion of myeloma cell lines | 9.28 × 10-5 | ANXA2, CD44, FN1, ITGB1 | 4 | |
| Cell morphology, embryonic development | Cell spreading of embryonic cell lines | 9.28 × 10-5 | −1.067 | FLNA, FN1, ITGB1, VIM | 4 |
| Gene expression | Replication of RNA | 9.28 × 10-5 | 0.152 | G3BP1, NPM1, VAPA, YBX1 | 4 |
| Cardiovascular system development and function | Angiogenesis | 9.48 × 10-5 | −1.134 | AKR1B1, ANXA2, ARHGDIA, ATP5B, CALR, CAPZB, CAV1, CD44, CLIC4, CTSB, CYR61, FKBP1A, FLNA, FLNB, FN1, G6PD, HADHA, HMGB1, HSP90B1, HSPB1, HSPD1, IGF2R, LDHA, LGALS1, MAPK1, MYH10, MYH9, NCL, PDCD6, PGK1, PKM, PLEC, RHOA, RNH1, RPSA, RTN4, SPTBN1, STAT1, TNC, VCL, VIM, WARS, YWHAE, YWHAZ | 44 |
| Cancer, organismal injury and abnormalities, reproductive system disease | Metastatic cervical cancer | 9.62 × 10-5 | ATIC, HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 6 | |
| Molecular transport, protein trafficking | Nuclear export of protein | 9.62 × 10-5 | CALR, CSE1L, EIF5A, HSPA9, RAN, XPO1 | 6 | |
| Cancer, organismal injury and abnormalities, reproductive system disease | Recurrent cervical cancer | 9.62 × 10-5 | ATIC, HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 6 | |
| Cell morphology, renal and urological system development and function | Shape change of kidney Cell lines | 9.70 × 10-5 | 0.050 | ANXA2, FLNA, FLNC, FN1, ITGB1, MAPK1, RHOA, VIM | 8 |
| Cellular development, cellular growth and proliferation | Proliferation of lung cancer cell lines | 1.00 × 10-4 | −1.422 | ACLY, ANXA1, ANXA2, C1QBP, CAV1, CYR61, GAPDH, H2AFY, HNRNPA2B1, ITGB1, MAPK1, NASP, PKM, PTBP1, SET, SLC25A6, SOD1, STMN1, TRIM28 | 19 |
| Cancer | Mucoepidermoid carcinoma | 1.01 × 10-4 | ATIC, FN1, HNRNPK, KRT18, SFN, SLC25A5, TNC | 7 | |
| DNA replication, recombination, and repair | Degradation of DNA | 1.03 × 10-4 | −0.941 | APEX1, BSG, CAPN2, CAST, DHX9, ENO1, HNRNPA1, HSD17B10, HSPB1, KRT8, LMNA, NPM1, PPIA, RHOA, SOD1, STAT1 | 16 |
| Cellular development | Branching of cells | 1.12 × 10-4 | −1.379 | ACTR3, ANXA2, BASP1, CAPNS1, CAPZB, CAV1, CSRP1, CYR61, DBN1, DNM1L, DPYSL2, EZR, FN1, HMGB1, HNRNPK, ITGB1, MAP1B, NNMT, PDIA3, PFN1, RHOA, RTN4, SOD1, TNC | 24 |
| Cell morphology, embryonic development | Shape change of embryonic cell lines | 1.13 × 10-4 | −0.555 | FLNA, FN1, ITGB1, MAPK1, VIM | 5 |
| Cardiovascular system development and function, organismal development | Vasculogenesis | 1.15 × 10-4 | −1.808 | AKR1B1, ANXA2, ARHGDIA, ATP5A1, CALR, CAPNS1, CAPZB, CAV1, CD44, CLIC4, CTSB, CYR61, FKBP1A, FLNA, FN1, G6PD, HADHA, HMGB1, HSP90B1, IGF2R, ITGB1, KRT1, LDHA, LGALS1, MAPK1, MYH10, PDCD6, PKM, PLEC, PPIA, PRKDC, RAD23B, RHOA, SEPT9, SERPINH1, SOD1, SPTBN1, STAT1, TNC, VCL, VIM, WARS, YWHAE, YWHAG, YWHAZ | 45 |
| Cancer, cellular movement, organismal injury and abnormalities, reproductive system disease, tumor morphology | Invasion of mammary tumor cells | 1.17 × 10-4 | 0.156 | CD44, CTTN, FLNA, FN1, HDLBP, ITGB1, RHOA | 7 |
| Cancer | Solid tumor | 1.21 × 10-4 | 0.779 | AARS, ACAT1, ACAT2, ACLY, ACTG1, ACTN1, ACTN4, ACTR2, ACTR3, AHCY, AHNAK, AIP, AKR1B1, ALDOA, ALDOC, ANXA1, ANXA2, ANXA5, AP1B1, APEX1, ARF1, ARHGDIA, ARPC2, ATIC, ATP5A1, ATP5B, B2M, BASP1, BSG, C14orf166, C1QBP, CACYBP, CALR, CAND1, CANX, CAP1, CAPG, CAPN1, CAPN2, CAPRIN1, CAPZA1, CAST, CAV1, CBX3, CCT2, CCT3, CCT4, CCT5, CCT6A, CCT7, CCT8, CD44, CD59, CDK1, CFL1, CLIC1, CLTC, CNN3, COPA, COPB1, COPB2, COPE, COPG1, CORO1C, COTL1, CRIP2, CSE1L, CTNNA1, CTPS1, CTSA, CTSB, CTTN, CUTA, CYB5R3, CYC1, CYR61, DBI, DBN1, DDX1, DDX39B, DDX3X, DDX5, DHX9, DLAT, DNAJA1, DNAJA2, DNAJC8, DNM1L, DPYSL2, DYNC1H1, DYNC1I2, ECHS1, EEF1A1, EEF1B2, EEF1D, EEF2, EHD1, EIF2S1, EIF2S2, EIF3B, EIF3C, EIF3I, EIF3M, EIF4A1, EIF4G1, EIF4H, ENAH, ENO1, ENO2, EPRS, ESYT1, ETFA, EZR, FASN, FDPS, FKBP1A, FLNA, FLNB, FLNC, FN1, FUBP3, FUS, G3BP1, G6PD, GANAB, GARS, GCN1L1, GDI1, GDI2, GLUD1, GML, GMPS, GNAI2, GNB2L1, GPI, GSTP1, H2AFY, H3F3A/H3F3B, HADHA, HARS, HDGF, HDLBP, HEXB, HINT1, HIST1H1C, HIST1H2BL, HIST2H2AC, HLA-A, HLA-B, HMGB1, HN1, HNRNPA1, HNRNPA2B1, HNRNPA3, HNRNPH1, HNRNPH3, HNRNPK, HNRNPL, HNRNPM, HNRNPR, HNRNPU, HSP90AA1, HSP90AB1, HSP90B1, HSPA4, HSPA5, HSPA8, HSPA9, HSPD1, HYOU1, IARS, IDI1, IFITM3, IGF2R, ILF2, ILF3, IMMT, IMPDH2, IPO5, IPO7, IPO9, IQGAP1, ITGB1, KHDRBS1, KHSRP, KIF5B, KPNA2, KPNA3, KPNB1, KRT1, KRT18, KRT8, KRT9, LAMB1, LASP1, LDHA, LGALS1, LMNA, LMNB1, LOC102724594/U2AF1, LONP1, LRPPRC, LRRC59, MAP1B, MAP4, MAPK1, MAPRE1, MARCKS, MARS, MATR3, MCM3, MCM4, MCM6, MCM7, MDH1, MDH2, MSN, MVP, MYH10, MYH9, MYL12A, NACA, NAP1L1, NAP1L4, NASP, NCL, NNMT, NONO, NPEPPS, NPLOC4, NPM1, NSFL1C, NUMA1, OLA1, P4HB, PA2G4, PABPC1, PAFAH1B2, PAICS, PARK7, PCBP1, PCBP2, PCM1, PCMT1, PCNA, PDCD6, PDE6H, PDIA3, PDIA4, PDLIM1, PDXK, PEA15, PFKP, PFN1, PGAM1, PGD, PGK1, PGM1, PHGDH, PICALM, PKM, PLEC, PLOD2, PLS3, PPIA, PPM1G, PPP1CA, PPP2CA, PPP2R1A, PRDX1, PRDX2, PRDX5, PRKCSH, PRKDC, PRMT5, PRPF19, PSMA1, PSMB1, PSMC1, PSMC4, PSMD2, PSME1, PTBP1, PTMA, PTRF, PUF60, PYGB, PYGL, QARS, RANBP1, RARS, RCN1, RHOA, RNH1, RPL10, RPL12, RPL21, RPL22, RPL27, RPL4, RPL5, RPL7, RPL7A, RPL8, RPLP0, RPN1, RPN2, RPS11, RPS12, RPS2, RPS24, RPS27A, RPS3A, RPS5, RPS7, RRBP1, RTN4, S100A6, SDHA, SEPT9, SERBP1, SERPINH1, SF3A1, SF3B1, SFN, SFPQ, SH3BGRL3 | 387 |
| SHMT2, SLC25A3, SLC25A5, SLC38A2, SND1, SOD1, SPTAN1, SPTBN1, SSRP1, STAT1, STIP1, STMN1, STOML2, SYNCRIP, TAGLN, TALDO1, TARS, TCP1, TFRC, TLN1, TMPO, TNC, TNPO1, TPI1, TPM2, TPM3, TPM4, TUBB, TUBB4B, TUBB6, TUBB8, TWF2, TXLNA, TXNDC5, UAP1, UBA1, UBE2N, UBXN4, UCHL1, UQCRC1, VARS, VAT1, VCL, VCP, VIM, VPS35, WARS, WDR1, XPO1, XRCC5, XRCC6, YBX1, YBX3, YWHAB, YWHAE, YWHAG, YWHAH, YWHAQ, YWHAZ, ZYX | |||||
| Cancer, cellular movement, tumor morphology | Invasion of tumor cells | 1.29 × 10-4 | −1.586 | AHCY, CAPN2, CD44, CTSB, CTTN, EZR, FKBP1A, FLNA, FN1, HDLBP, ITGB1, LGALS1, MAPK1, PARK7, RHOA | 15 |
| Embryonic development, organ development, organismal development, skeletal and muscular system development and function, tissue development | Formation of muscle | 1.29 × 10-4 | −0.543 | ACTA1, ACTG1, ACTN4, ANXA1, ANXA5, CALR, CAPN2, CAPZB, CAV1, FKBP1A, FLNB, FLNC, FN1, HMGB1, HSP90B1, ITGB1, KRT8, MAPK1, MYH10, MYH9, PLEC, RHOA, STIP1, UCHL1, VCL, VIM | 26 |
| Cell-to-cell signaling and interaction, embryonic development | Binding of embryonic cells | 1.31 × 10-4 | C1QBP, ITGB1, TLN1 | 3 | |
| Cell death and survival, connective tissue disorders, developmental disorder, hematological disease, hereditary disorder | Hereditary nonspherocytic hemolytic anemia | 1.31 × 10-4 | ALDOA, G6PD, GPI | 3 | |
| Cell morphology, endocrine system development and function, organ morphology, organismal development | Morphology of thyroid cells | 1.31 × 10-4 | CAV1, CTSB, FN1 | 3 | |
| Cellular assembly and organization, cellular function and maintenance | Quantity of filopodia-like projection | 1.31 × 10-4 | ACTR2, ARPC2, CAPZB | 3 | |
| Amino acid metabolism, small molecule biochemistry | Synthesis of l-serine | 1.31 × 10-4 | PHGDH, PKM, SHMT2 | 3 | |
| Cellular assembly and organization, cellular function and maintenance, tissue development | Formation of actin filaments | 1.34 × 10-4 | −1.689 | ACTR3, ARF1, ARPC2, CAPN1, CAV1, CD44, CFL1, CTTN, FN1, GNAI2, GPI, ITGB1, MAPK1, NPM1, PFN1, RHOA, TNC, TPM2, TWF1, TWF2, ZYX | 21 |
| Cell cycle, cellular movement | Cytokinesis of cervical cancer cell lines | 1.34 × 10-4 | 0.218 | GNAI2, LMNA, NPM1, RHOA, SEPT7, SEPT9, SSRP1 | 7 |
| Cancer, respiratory disease | Metastatic non-small-cell lung cancer | 1.34 × 10-4 | ATIC, EIF4A1, HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 7 | |
| Cell-to-cell signaling and interaction, tissue development | Adhesion of hepatoma cell lines | 1.37 × 10-4 | BSG, C1QBP, ITGB1, RHOA | 4 | |
| Cardiovascular system development and function, cell morphology, cell-to-cell signaling and interaction, organ morphology, organismal development, skeletal and muscular system development and function, tissue morphology | Morphology of intercalated disks | 1.37 × 10-4 | CALR, CAPZB, PLEC, VCL | 4 | |
| Cellular compromise | Stress response of cervical cancer cell lines | 1.37 × 10-4 | CALR, HNRNPA1, HSP90B1, HSPA5 | 4 | |
| Cancer | Metastatic occult primary cancer of head and neck | 1.43 × 10-4 | HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 5 | |
| Cancer, connective tissue disorders, respiratory disease, skeletal and muscular disorders | Metastatic squamous cell cancer of the ethmoid sinus | 1.43 × 10-4 | HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 5 | |
| Cancer, respiratory disease | Metastatic squamous cell cancer of the glottis | 1.43 × 10-4 | HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 5 | |
| Cancer, gastrointestinal disease | Metastatic squamous cell cancer of the lip | 1.43 × 10-4 | HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 5 | |
| Cancer, respiratory disease | Metastatic squamous cell cancer of the maxillary sinus | 1.43 × 10-4 | HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 5 | |
| Cancer, gastrointestinal disease, respiratory disease | Metastatic squamous cell cancer of the oropharynx | 1.43 × 10-4 | HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 5 | |
| Cancer, respiratory disease | Metastatic squamous cell cancer of the supraglottis | 1.43 × 10-4 | HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 5 | |
| Cellular movement, connective tissue development and function, hepatic system development and function | Migration of hepatic stellate cells | 1.43 × 10-4 | −0.218 | CD44, GNB2L1, HMGB1, LGALS1, MYH10 | 5 |
| Cancer | Recurrent occult primary cancer of head and neck | 1.43 × 10-4 | HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 5 | |
| Cancer, connective tissue disorders, respiratory disease, skeletal and muscular disorders | Recurrent squamous cell cancer of the ethmoid sinus | 1.43 × 10-4 | HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 5 | |
| Cancer, respiratory disease | Recurrent squamous cell cancer of the glottis | 1.43 × 10-4 | HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 5 | |
| Cancer, gastrointestinal disease | Recurrent squamous cell cancer of the lip | 1.43 × 10-4 | HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 5 | |
| Cancer, respiratory disease | Recurrent squamous cell cancer of the maxillary sinus | 1.43 × 10-4 | HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 5 | |
| Cancer, respiratory disease | Recurrent squamous cell cancer of the supraglottis | 1.43 × 10-4 | HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 5 | |
| Cancer | Unresectable occult primary cancer of head and neck | 1.43 × 10-4 | HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 5 | |
| Cancer, connective tissue disorders, respiratory disease, skeletal and muscular disorders | Unresectable squamous Cell cancer of the ethmoid sinus | 1.43 × 10-4 | HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 5 | |
| Cancer, respiratory disease | Unresectable squamous Cell cancer of the glottis | 1.43 × 10-4 | HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 5 | |
| Cancer, gastrointestinal disease, respiratory disease | Unresectable squamous Cell cancer of the hypopharynx | 1.43 × 10-4 | HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 5 | |
| Cancer, gastrointestinal disease | Unresectable squamous Cell cancer of the lip | 1.43 × 10-4 | HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 5 | |
| Cancer, respiratory disease | Unresectable squamous Cell cancer of the maxillary sinus | 1.43 × 10-4 | HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 5 | |
| Cancer, gastrointestinal disease, respiratory disease | Unresectable squamous Cell cancer of the oropharynx | 1.43 × 10-4 | HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 5 | |
| Cancer, respiratory disease | Unresectable squamous Cell cancer of the supraglottis | 1.43 × 10-4 | HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 5 | |
| Cellular movement | Cell movement of carcinoma Cell lines | 1.48 × 10-4 | 0.447 | C1QBP, CAV1, CD44, CTTN, CYR61, HNRNPA2B1, ITGB1, LGALS1, STMN1, TAGLN2, VIM, ZYX | 12 |
| Cellular development, cellular growth and proliferation | Proliferation of colon cancer cell lines | 1.48 × 10-4 | 1.556 | AHSA1, CACYBP, CALR, CDK1, CSE1L, DDX5, EIF3C, FN1, HSP90AA1, IGF2R, IPO7, NCL, PKM, RHOA, SFN, SFPQ, UBA1, XRCC5 | 18 |
| Cardiovascular disease, organismal injury and abnormalities | Dilated cardiomyopathy | 1.50 × 10-4 | −0.192 | CAST, DNM1L, FKBP1A, LMNA, MYH9, PGK1, PPP1CA, SDHA, TLN1, TMPO, TPI1, VCL, VDAC1, VDAC2 | 14 |
| Cardiovascular system development and function, organ morphology, organismal development, skeletal and muscular system development and function, tissue morphology | Morphology of cardiac muscle | 1.50 × 10-4 | CALR, CAPZB, CAV1, FKBP1A, FN1, HADHA, HSP90B1, IGF2R, MAPK1, MYH10, PLEC, SPTBN1, VCL, YWHAE | 14 | |
| Infectious disease | Infection by dengue virus 2 | 1.53 × 10-4 | 0.896 | ACTR2, AP1B1, AP2B1, ARPC1B, CLTC, IFITM3, STAT1 | 7 |
| Cell morphology, cellular assembly and organization, cellular function and maintenance | Reorganization of cytoskeleton | 1.59 × 10-4 | −1.953 | ARHGDIA, CD44, CFL1, DPYSL2, EZR, FLNA, FLNC, FN1, HMGB1, MARCKS, MSN, MYH9, PLS3, RHOA, SPTAN1 | 15 |
| Cell-to-cell signaling and interaction, tissue development | Cell-cell adhesion of tumor Cell lines | 1.62 × 10-4 | 0.128 | C1QBP, ERP29, GNB2L1, ITGB1, MAPK1, ZYX | 6 |
| Cellular assembly and organization | Formation of cytoplasmic aggregates | 1.62 × 10-4 | −1.131 | CAV1, DDX6, LMNA, RHOA, SOD1, YWHAZ | 6 |
| Tissue development | Aggregation of cells | 1.65 × 10-4 | −0.939 | ACTG1, AKR1B1, BSG, CAPN1, CAST, CD44, CLIC1, CSRP1, FLNA, FN1, GNAI2, HNRNPA2B1, HSPB1, ITGB1, LGALS1, MAPK1, MYH9, MYL12A, P4HB, PDIA3, TLN1, VCL | 22 |
| Cell death and survival | Apoptosis of fibroblast Cell lines | 1.65 × 10-4 | −0.651 | CAPNS1, CDK1, CLIC4, CYR61, DDX3X, DNM1L, EEF1A1, EIF2S1, EIF3B, EIF3C, EIF3I, EIF6, FN1, FUS, GPI, HSPA5, HSPD1, ITGB1, RHOA, RPL10, STAT1, VIM | 22 |
| Infectious disease, reproductive system disease | Infection of cervical cancer cell lines | 1.69 × 10-4 | −2.295 | ARF1, ATP5B, CCT2, COPA, COPB1, COPB2, COPG1, DDX3X, DNAJA2, EIF3I, GML, H3F3A/H3F3B, HNRNPK, HNRNPU, HSPA9, MAP4, PDIA3, PDIA6, PSME2, RAB1B, RANBP1, SPTAN1, SPTBN1, STIP1, TWF1, UAP1, UQCRC1, XPO1 | 28 |
| Cell cycle | Mitosis | 1.76 × 10-4 | −1.833 | BUB3, CAPN2, CCT4, CDK1, CLTC, CSE1L, CYR61, DBI, DHX9, DYNC1H1, EIF6, KPNB1, KRT18, MAPK1, MARCKS, MYH10, NUMA1, PEBP1, PPP1CA, PRMT5, PTMA, RAN, RPS24, SEPT9, STAT1, STMN1, TCP1, TNC, TUBB, YWHAE | 30 |
| Cellular movement | Migration of tumor cells | 1.76 × 10-4 | −0.843 | ACTN4, AHCY, CAV1, CD44, CDK1, CSE1L, CTSB, CTTN, EZR, FN1, HMGB1, IQGAP1, ITGB1, RHOA, S100A6, TNC, VIM | 17 |
| Cell morphology, cellular function and maintenance | Autophagy of cells | 1.77 × 10-4 | 0.609 | CAPN1, CAPNS1, CAST, DNM1L, EIF2S1, EIF4G1, FKBP1A, HMGB1, HSPA5, KRT18, PAFAH1B2, PRKDC, RAB7A, SERPINH1, SOD1, SQSTM1, STAT1, TUFM | 18 |
| Cardiovascular system development and function, organismal development | Development of blood vessel | 1.78 × 10-4 | −1.788 | AKR1B1, ANXA2, ARHGDIA, ATP5A1, CALR, CAPNS1, CAPZB, CAV1, CD44, CLIC4, CTSB, CYR61, FKBP1A, FLNA, FN1, G6PD, HADHA, HMGB1, HSP90B1, HSPB1, IGF2R, ITGB1, KRT1, LDHA, LGALS1, MAPK1, MYH10, PDCD6, PGK1, PKM, PLEC, PPIA, PRKDC, RAD23B, RHOA, RNH1, SEPT9, SERPINH1, SOD1, SPTBN1, STAT1, TNC, VCL, VIM, WARS, YWHAE, YWHAG, YWHAZ | 48 |
| Cardiovascular system development and function, cell morphology | Cell spreading of endothelial cells | 1.79 × 10-4 | −1.000 | FLNA, FN1, ITGB1, VIM, YWHAZ | 5 |
| Cell morphology, renal and urological system development and function | Cell spreading of kidney Cell lines | 1.79 × 10-4 | −0.678 | FLNA, FLNC, FN1, ITGB1, VIM | 5 |
| Cell death and survival, DNA replication, recombination, and repair | Fragmentation of DNA | 1.88 × 10-4 | −0.910 | APEX1, BSG, CAPN2, CAST, ENO1, HSD17B10, HSPB1, KRT8, LMNA, NPM1, PPIA, RHOA, SOD1, STAT1 | 14 |
| Cancer, gastrointestinal disease, respiratory disease | Nasopharyngeal cancer | 1.91 × 10-4 | HSP90AA1, HSP90AB1, HSP90B1, SFN, TUBB, TUBB4B | 6 | |
| Energy production, nucleic acid metabolism, small molecule biochemistry | Binding of ATP | 1.94 × 10-4 | −0.068 | MAPK1, PCNA, PTGES3, STIP1 | 4 |
| Free radical scavenging | Modification of hydrogen peroxide | 1.94 × 10-4 | −1.710 | PRDX1, PRDX2, PRDX5, SOD1 | 4 |
| Cell-to-cell signaling and interaction, cellular assembly and organization, tissue development | Turnover of focal adhesions | 1.94 × 10-4 | −0.254 | CAV1, ITGB1, PLEC, VCL | 4 |
| Cancer, cell death and survival, tumor morphology | Cell death of tumor cells | 1.95 × 10-4 | 0.142 | ACLY, ANXA1, ANXA2, B2M, BSG, CAV1, CD44, CD59, CDK1, CTSB, CYR61, EIF3C, ENO1, FASN, GNB2L1, HMGB1, HSP90AB1, LGALS1, MAPK1, PPP2CA, RHOA, RPS3A, RTN4, SOD1 | 24 |
| Cellular function and maintenance, tissue development | Organization of muscle cells | 1.98 × 10-4 | ACTG1, CALR, CAV1, CTNNA1, FN1, ITGB1, KRT8 | 7 | |
| RNA post-transcriptional modification | Processing of rRNA | 1.98 × 10-4 | DDX5, NPM1, RPL5, RPL7, RPS15, RPS24, RPS7 | 7 | |
| Cancer, gastrointestinal disease | Oral cavity carcinoma | 2.06 × 10-4 | ATIC, EZR, FASN, FN1, HSP90AA1, HSP90AB1, HSP90B1, LGALS1, NPM1, PLOD2, RPL10, TAGLN, TNC, TUBB, TUBB4B | 15 | |
| Inflammatory response | Inflammation of body cavity | 2.07 × 10-4 | 0.920 | ACTN4, ALDOA, ANXA1, ANXA5, ARHGDIA, B2M, BSG, CAV1, CD44, CYR61, DDX5, ENO1, FKBP1A, GAPDH, GSTP1, HLA-A, HMGB1, HSPA5, IMPDH2, KRT18, KRT8, LGALS1, MYH10, MYH9, P4HB, PDIA3, PDLIM1, PPIA, PPP2CA, PRDX1, PSMB1, PSMD2, SOD1, STAT1, TKT, TPI1, TUBB, YBX1 | 38 |
| Cancer | Malignant solid tumor | 2.08 × 10-4 | 0.397 | AARS, ACAT1, ACAT2, ACLY, ACTG1, ACTN1, ACTN4, ACTR2, ACTR3, AHCY, AHNAK, AKR1B1, ALDOA, ALDOC, ANXA1, ANXA2, ANXA5, AP1B1, APEX1, ARF1, ARHGDIA, ARPC2, ATIC, ATP5A1, ATP5B, B2M, BASP1, BSG, C14orf166, C1QBP, CACYBP, CAND1, CANX, CAP1, CAPG, CAPN1, CAPN2, CAPRIN1, CAPZA1, CAST, CAV1, CBX3, CCT2, CCT3, CCT4, CCT5, CCT6A, CCT7, CCT8, CD44, CD59, CDK1, CFL1, CLIC1, CLTC, CNN3, COPA, COPB1, COPB2, COPE, COPG1, CORO1C, COTL1, CRIP2, CSE1L, CTNNA1, CTPS1, CTSA, CTSB, CTTN, CUTA, CYB5R3, CYC1, CYR61, DBI, DBN1, DDX1, DDX39B, DDX3X, DDX5, DHX9, DLAT, DNAJA1, DNAJA2, DNAJC8, DNM1L, DPYSL2, DYNC1H1, DYNC1I2, ECHS1, EEF1A1, EEF1B2, EEF1D, EEF2, EHD1, EIF2S1, EIF2S2, EIF3B, EIF3C, EIF3I, EIF3M, EIF4A1, EIF4G1, EIF4H, ENAH, ENO1, ENO2, EPRS, ESYT1, ETFA, EZR, FASN, FDPS, FKBP1A, FLNA, FLNB, FLNC, FN1, FUBP3, FUS, G3BP1, G6PD, GANAB, GARS, GCN1L1, GDI1, GDI2, GLUD1, GML, GMPS, GNAI2, GNB2L1, GPI, GSTP1, H2AFY, H3F3A/H3F3B, HADHA, HARS, HDGF, HDLBP, HEXB, HINT1, HIST1H1C | 383 |
| HIST1H2BL, HIST2H2AC, HLA-A, HLA-B, HMGB1, HN1, HNRNPA1, HNRNPA2B1, HNRNPA3, HNRNPH1, HNRNPH3, HNRNPK, HNRNPL, HNRNPM, HNRNPR, HNRNPU, HSP90AA1, HSP90AB1, HSP90B1, HSPA4, HSPA5, HSPA8, HSPA9, HSPD1, HYOU1, IARS, IDI1, IFITM3, IGF2R, ILF2, ILF3, IMMT, IMPDH2, IPO5, IPO7, IPO9, IQGAP1, ITGB1, KHDRBS1, KHSRP, KIF5B, KPNA2, KPNA3, KPNB1, KRT1, KRT18, KRT8, KRT9, LAMB1, LASP1, LDHA, LGALS1, LMNA, LMNB1, LOC102724594/U2AF1, LONP1, LRPPRC, LRRC59, MAP1B, MAP4, MAPK1, MAPRE1, MARCKS, MARS, MATR3, MCM3, MCM4, MCM6, MCM7, MDH1, MDH2, MSN, MVP, MYH10, MYH9, MYL12A, NACA, NAP1L1, NAP1L4, NASP, NCL, NNMT, NONO, NPEPPS, NPLOC4, NPM1, NSFL1C, NUMA1, OLA1, P4HB, PA2G4, PABPC1, PAFAH1B2, PAICS, PCBP1, PCBP2, PCM1, PCMT1, PCNA, PDCD6, PDE6H, PDIA3, PDIA4, PDLIM1, PDXK, PEA15, PFKP, PFN1, PGAM1, PGD, PGK1, PGM1, PHGDH, PICALM, PKM, PLEC, PLOD2, PLS3, PPIA, PPM1G, PPP1CA, PPP2CA, PPP2R1A, PRDX1, PRDX2, PRDX5, PRKCSH, PRKDC, PRMT5, PRPF19, PSMA1, PSMB1, PSMC1, PSMC4, PSMD2, PSME1, PTBP1, PTMA, PTRF, PUF60, PYGB, PYGL, QARS, RANBP1, RARS, RCN1, RHOA, RNH1, RPL10, RPL12, RPL22, RPL27, RPL4, RPL5, RPL7, RPL7A, RPL8, RPLP0, RPN1, RPN2, RPS11, RPS12, RPS2, RPS24, RPS27A, RPS3A, RPS5, RPS7, RRBP1, RTN4, S100A6, SDHA, SEPT9, SERBP1, SERPINH1, SF3A1, SF3B1, SFN, SFPQ, SH3BGRL3, SHMT2, SLC25A3, SLC25A5, SLC38A2, SND1, SOD1, SPTAN1, SPTBN1, SSRP1, STAT1, STIP1, STMN1, STOML2, SYNCRIP, TAGLN, TALDO1, TARS, TCP1, TFRC, TLN1, TMPO, TNC, TNPO1, TPI1, TPM2, TPM3, TPM4, TUBB, TUBB4B, TUBB6, TUBB8, TWF2, TXLNA, TXNDC5, UAP1, UBA1, UBE2N, UBXN4, UCHL1, UQCRC1, VARS, VAT1, VCL, VCP, VIM, VPS35, WARS, WDR1, XPO1, XRCC5, XRCC6, YBX1, YBX3, YWHAB, YWHAE, YWHAG, YWHAH, YWHAQ, YWHAZ, ZYX | |||||
| Cell cycle | M phase of cervical cancer Cell lines | 2.10 × 10-4 | 0.218 | GNAI2, LMNA, MCM7, NPM1, RHOA, SEPT7, SEPT9, SSRP1 | 8 |
| Cardiovascular system development and function, organismal development, tissue morphology | Morphology of cardiovascular tissue | 2.11 × 10-4 | ARHGDIA, CALR, CAPZB, CAV1, CD44, CLIC4, FKBP1A, FN1, HADHA, HSP90B1, IGF2R, MAPK1, MYH10, PLEC, SPTBN1, VCL, YWHAE | 17 | |
| Cell cycle | Interphase of tumor cell lines | 2.11 × 10-4 | 1.067 | ANXA2, CDK1, CSE1L, CYR61, FASN, ITGB1, LGALS1, MCM7, NASP, NPM1, PDCD6, PTGES3, RHOA, RPL23, RPL5, RPL7A, SFN, SPTAN1, STAT1, TCP1, TFRC, YWHAG, YWHAQ | 23 |
| Infectious disease | Replication of retroviridae | 2.19 × 10-4 | 0.989 | ANXA5, BTF3, CAV1, DDX5, DHX9, IFITM3, PLIN3, PPIA, PRDX1, PRDX2, RAB11A, TMPO | 12 |
| Cell morphology | Sprouting | 2.22 × 10-4 | −1.379 | ACTR3, ANXA2, BASP1, CAPNS1, CAPZB, CAV1, CSRP1, CYR61, DBN1, DNM1L, DPYSL2, EZR, FN1, HMGB1, HNRNPK, ITGB1, MAP1B, NNMT, PDIA3, PFN1, RHOA, RTN4, SOD1, TNC | 24 |
| Connective tissue disorders, developmental disorder, hereditary disorder | Marfan’s syndrome | 2.22 × 10-4 | ALDOC, FLNA, MYH10, TAGLN, VCL | 5 | |
| Cancer, gastrointestinal disease, respiratory disease | Metastatic squamous cell cancer of the hypopharynx | 2.22 × 10-4 | HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 5 | |
| Cellular function and maintenance | Pinocytosis | 2.22 × 10-4 | −1.000 | ACTN4, CAV1, EZR, NCL, RHOA | 5 |
| Cancer, gastrointestinal disease, respiratory disease | Recurrent squamous cell cancer of the hypopharynx | 2.22 × 10-4 | HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 5 | |
| Cancer, gastrointestinal disease, respiratory disease | Recurrent squamous cell cancer of the oropharynx | 2.22 × 10-4 | HSP90AA1, HSP90AB1, HSP90B1, TUBB, TUBB4B | 5 | |
| Embryonic development, tissue morphology | Morphology of visceral endoderm | 2.23 × 10-4 | DHX9, ITGB1, MAPK1, MYH9, RAB7A, SERPINH1 | 6 | |
| Cellular movement | Cell movement of lung cancer cell lines | 2.26 × 10-4 | 0.276 | C1QBP, CAV1, CD44, FN1, HNRNPA2B1, ITGB1, STMN1, VIM, YBX1, ZYX | 10 |
| Cellular movement | Migration of cancer cells | 2.28 × 10-4 | −0.743 | AHCY, CAV1, CD44, CDK1, CSE1L, CTSB, EZR, FN1, HMGB1, IQGAP1, ITGB1, RHOA, S100A6, TNC, VIM | 15 |
| Cell cycle | Arrest in interphase of tumor cell lines | 2.29 × 10-4 | ANXA2, CDK1, CSE1L, CYR61, FASN, ITGB1, LGALS1, PDCD6, PTGES3, RHOA, RPL23, RPL5, RPL7A, SFN, SPTAN1, TCP1, TFRC, YWHAG | 18 | |
| Cellular function and maintenance | Engulfment of tumor cell lines | 2.48 × 10-4 | 0.222 | ANXA1, CAV1, CD44, CLTC, EZR, GNB2L1, ITGB1, NCL, PFN1, RHOA, SFPQ, VIM | 12 |
| Cell-to-cell signaling and interaction, tissue development | Detachment of cells | 2.52 × 10-4 | 0.851 | ANXA1, ANXA5, CAPN2, FLNA, FN1, ITGB1, MAPK1, RHOA, TLN1 | 9 |
| Cell morphology, cell-to-Cell signaling and interaction | Morphology of intercellular junctions | 2.52 × 10-4 | B2M, CALR, CAPRIN1, CAPZB, CAV1, LGALS1, PLEC, VCL, YWHAG | 9 | |
| Gene expression, protein synthesis | Initiation of translation of mRNA | 2.53 × 10-4 | EIF3B, EIF3C, EIF3I, EIF4G1, EIF4H, HSPB1, RPS3A | 7 | |
| Cell-to-cell signaling and interaction, connective tissue development and function, tissue development | Attachment of fibroblast Cell lines | 2.57 × 10-4 | FN1, IPO9, ITGB1 | 3 | |
| Cell morphology | Cell spreading of fibrosarcoma cell lines | 2.57 × 10-4 | FLNA, FLNB, ITGB1 | 3 | |
| Organ morphology, skeletal and muscular system development and function | Contraction of airway smooth muscle | 2.57 × 10-4 | CAV1, GNAI2, RHOA | 3 | |
| Cellular assembly and organization, DNA replication, recombination, and repair | Formation of nuclear envelope | 2.57 × 10-4 | CDK1, LMNA, RAN | 3 | |
| Carbohydrate metabolism | Metabolism of fructose-1, 6-diphosphate | 2.57 × 10-4 | ALDOA, ALDOC, PFKP | 3 | |
| Cancer, endocrine system disorders, organismal injury and abnormalities, reproductive system disease | Metastatic ovarian tumor | 2.57 × 10-4 | HSP90AA1, HSP90AB1, HSP90B1 | 3 | |
| Cellular development, cellular growth and proliferation | Outgrowth of breast cancer Cell lines | 2.57 × 10-4 | CYR61, FN1, ITGB1 | 3 | |
| Cellular assembly and organization, cellular function and maintenance, protein trafficking | Sequestration of g-actin | 2.57 × 10-4 | PFN1, TWF1, TWF2 | 3 | |
| Cell cycle | Abnormal cell cycle | 2.58 × 10-4 | CAV1, H3F3A/H3F3B, HSPB1, PRDX1, RBM3, STAT1, TMPO, YWHAE | 8 | |
| Cellular movement, connective tissue development and function | Migration of fibroblasts | 2.64 × 10-4 | 0.975 | CAPNS1, CAV1, CYR61, EHD1, FLNB, FN1, MAPK1, MYH10, PLEC, STMN1, VCL, ZYX | 12 |
| Developmental disorder, hematological disease, hereditary disorder, organismal injury and abnormalities | Diamond-blackfan anemia | 2.66 × 10-4 | RPL5, RPS10, RPS24, RPS7 | 4 | |
| Organismal injury and abnormalities, renal and urological disease | Minimal change nephrotic syndrome | 2.66 × 10-4 | ACTN4, FKBP1A, IMPDH2, PDLIM1 | 4 | |
| Cancer, organismal injury and abnormalities, renal and urological disease | Transitional cell bladder cancer | 2.66 × 10-4 | GSTP1, HSP90AA1, HSP90AB1, HSP90B1 | 4 | |
| Cellular assembly and organization, cellular function and maintenance | Growth of microtubules | 2.72 × 10-4 | −1.710 | MAP4, MAPRE1, MYH9, SSRP1, STMN1 | 5 |
| Cell death and survival | Apoptosis of cerebral cortex cells | 2.84 × 10-4 | 0.050 | CAPN1, CAST, CDK1, HSPA5, MAP1B, P4HB, PEA15, RPS3, SOD1, YWHAB | 10 |
| Cell death and survival, renal and urological system development and function | Cell viability of kidney cell lines | 2.85 × 10-4 | −0.378 | ANXA5, CAV1, EZR, HNRNPU, HSP90B1, HYOU1, PPIA, VDAC1 | 8 |
| Cell-to-cell signaling and interaction, renal and urological system development and function | Binding of kidney cell lines | 2.85 × 10-4 | 0.055 | ANXA2, BSG, CAV1, ITGB1, PPP2CA, PRMT5, RPSA | 7 |
| Molecular transport, RNA trafficking | Transport of mRNA | 2.85 × 10-4 | ALYREF, DDX39B, EIF5A, FUS, KHDRBS1, U2AF2, XPO1 | 7 | |
| Gene expression | Transcription of RNA | 2.90 × 10-4 | 1.084 | ACTR2, ACTR3, ALYREF, BASP1, BTF3, C14orf166, C1QBP, CALR, CAND1, CAV1, CBX3, CD44, CDK1, CSE1L, CYR61, DDX3X, DDX5, EEF1D, EIF2S1, ENO1, FKBP1A, FLNA, FN1, FUBP3, GLO1, GSTP1, H2AFY, HDGF, HEXB, HINT1, HIST1H1C, HMGB1, HNRNPA1, HNRNPA2B1, HNRNPK, HSPA8, ILF2, ILF3, IQGAP1, KHDRBS1, KPNA2, LGALS1, LMNA, LRPPRC, MAPK1, MATR3, MCM7, NONO, NPM1, OTUB1, PA2G4, PARK7, PCNA, PDLIM1, PEA15, PEBP1, PFN1, PHB2, PHGDH, PICALM, PPP2CA, PRKDC, PRMT5, PTGES3, PTMA, PTRF, RHOA, RPL6, SET, SFPQ, SQSTM1, SSRP1, STAT1, TMPO, TRIM28, UBE2L3, VAPA, XPO1, XRCC5, XRCC6, YBX1, YBX3, YWHAB, YWHAH, YWHAQ, YWHAZ | 86 |
| Cancer, endocrine system disorders, respiratory disease | Small cell lung cancer | 2.99 × 10-4 | ENO2, HSP90AA1, HSP90AB1, HSP90B1, HSPD1, PDCD6, TUBB, TUBB4B, YWHAE | 9 | |
| Cellular development, cellular growth and proliferation, embryonic development, organ development, organismal development, skeletal and muscular system development and function, tissue development | Formation of muscle cells | 3.04 × 10-4 | −1.408 | ACTA1, ACTG1, ACTN4, CALR, CAPZB, FLNC, FN1, HMGB1, HSP90B1, ITGB1, KRT8, MYH10, PLEC, UCHL1 | 14 |
| Cellular assembly and organization | Quantity of actin filaments | 3.14 × 10-4 | −0.570 | CFL1, FN1, GNB2L1, MARCKS, PLEC, RHOA, SPTAN1, TMOD3 | 8 |
| DNA replication, recombination, and repair | Repair of DNA | 3.18 × 10-4 | −0.335 | APEX1, CDK1, DDX1, FUS, HMGB1, HNRNPU, KPNA2, NPM1, OTUB1, PCNA, PRKDC, PRPF19, RAD23B, TRIM28, UBE2N, VCP, XRCC5, XRCC6, YBX1 | 19 |
| Cell-to-cell signaling and interaction | Binding of leukemia cell lines | 3.20 × 10-4 | 0.415 | ANXA2, ANXA5, CD44, HLA-A, ITGB1, NCL, TNC | 7 |
| Molecular transport, RNA trafficking | Export of RNA | 3.20 × 10-4 | −1.974 | ALYREF, DDX39B, EIF5A, KHDRBS1, RAN, U2AF2, XPO1 | 7 |
| Cancer, hematological disease, immunological disease, organismal injury and abnormalities | Large-cell KI-1 lymphoma | 3.20 × 10-4 | HMGB1, PSMB1, PSMD2, TUBB, TUBB4B, TUBB6, TUBB8 | 7 | |
| Cellular assembly and organization, cellular function and maintenance | Organization of microtubules | 3.20 × 10-4 | −1.000 | DPYSL2, GAPDH, MAPRE1, NUMA1, PCM1, RRBP1, TUBB | 7 |
| Cancer, organismal injury and abnormalities, reproductive system disease | Metastatic breast cancer | 3.25 × 10-4 | FDPS, FKBP1A, FLNA, HSP90AA1, HSP90AB1, HSP90B1, STAT1, TUBB, TUBB4B | 9 | |
| Immunological disease, inflammatory disease, inflammatory response, organismal injury and abnormalities, renal and urological disease | IGA nephropathy | 3.31 × 10-4 | ACTN4, IMPDH2, PDLIM1, PSMB1, PSMD2 | 5 | |
| Cellular movement, hematological system development and function, immune cell trafficking | Cell rolling of granulocytes | 3.47 × 10-4 | 0.000 | ANXA1, CD44, CTTN, GNAI2, ITGB1, TLN1 | 6 |
| DNA replication, recombination, and repair | Conformational modification of DNA | 3.47 × 10-4 | HMGB1, MCM4, MCM6, MCM7, NCL, SSRP1 | 6 | |
| Cancer | Locally advanced malignant tumor | 3.47 × 10-4 | ATIC, FKBP1A, HSP90AA1, HSP90AB1, HSP90B1, TUBB4B | 6 | |
| Organismal survival | Viability | 3.53 × 10-4 | −1.671 | B2M, GNAI2, MAP1B, PFN1, PRKDC, RAN, TKT, VCL, XRCC5 | 9 |
| Cell-to-cell signaling and interaction, skeletal and muscular system development and function, tissue development | Adhesion of smooth muscle cells | 3.56 × 10-4 | CYR61, FN1, ITGB1, RHOA | 4 | |
| Cellular assembly and organization, cellular function and maintenance | Quantity of lamellipodia | 3.56 × 10-4 | 0.000 | ACTR2, ARPC2, ENAH, TPM3 | 4 |
| Cell morphology | Shape change of leukemia Cell lines | 3.56 × 10-4 | FN1, ITGB1, MAPK1, TNC | 4 | |
| Cardiovascular disease, organismal injury and abnormalities | Primary cardiomyopathy | 3.58 × 10-4 | LMNA, PGK1, SDHA, TPI1, VCL, VDAC1, VDAC2 | 7 | |
| Cellular growth and proliferation | Formation of cells | 3.63 × 10-4 | −2.209 | ACTA1, ACTG1, ACTN4, ANXA2, ARHGDIA, B2M, BSG, CALR, CAPZB, CD44, CTTN, DNAJA1, EEF1D, EHD1, ENAH, EZR, FLNA, FLNB, FLNC, FN1, HEXB, HMGB1, HSP90B1, HSPA4, ITGB1, KRT18, KRT8, LGALS1, MAPK1, MYH10, MYH9, NPEPPS, PAFAH1B2, PEBP1, PLEC, PLS3, PRDX4, RAD23B, RANBP1, RHOA, RPL22, SFN, SOD1, SQSTM1, STAT1, STIP1, STMN1, TCP1, UCHL1, XRCC6, YBX3 | 51 |
| Cancer, organismal injury and abnormalities, reproductive system disease | Uterine leiomyoma | 3.74 × 10-4 | ALDOA, CYR61, DDX6, HSD17B10, HSP90AB1, IQGAP1, KPNA3, MDH1, MYH10, NPEPPS, PCNA, RPL27, RPS15, SF3A1, TNC, TWF1, VCP, ZYX | 18 | |
| Cell morphology, cellular assembly and organization, cellular function and maintenance | Reorganization of actin cytoskeleton | 3.78 × 10-4 | −2.219 | ARHGDIA, CFL1, DPYSL2, EZR, FLNA, FLNC, FN1, MYH9, PLS3, RHOA, SPTAN1 | 11 |
| Cancer | Stage 3 cancer | 3.78 × 10-4 | ATIC, CAV1, CBX3, CCT3, FKBP1A, FN1, KRT8, MAPK1, PCNA, TUBB4B, TUBB6 | 11 | |
| Cancer, tumor morphology | Progression of tumor | 3.83 × 10-4 | −0.342 | ANXA1, ANXA2, BSG, CAV1, CD44, EZR, FASN, FUS, GSTP1, HMGB1, STAT1, TNC, YWHAZ | 13 |
| Categories | Disease or Function Annotation | p-Value | Activation z-Score | Molecules | Number of Molecules |
|---|---|---|---|---|---|
| Liver hyperplasia/hyperproliferation | Cholangiocarcinoma | 7.33 × 10−7 | ANXA1, ANXA2, GNB2L1, HSP90AA1, HSP90AB1, PGK1, PKM, RPL4, RPLP0, VIM | 10 | |
| Renal damage, renal tubule injury | Proximal tubular toxicity | 2.17 × 10−6 | ACAT1, FASN, FN1, G6PD, GSTP1, HADH, HADHA, HSP90AA1, HSPB1, LGALS1, YBX1, YWHAH | 12 | |
| Nephrosis | Nephrosis | 1.35 × 10−5 | ACTN4, ANXA1, ARHGDIA, CLTC, FKBP1A, IMPDH2, ITGB1, LAMB1, PDLIM1 | 9 | |
| Liver fibrosis | Migration of hepatic stellate cells | 1.43 × 10−4 | −0.218 | CD44, GNB2L1, HMGB1, LGALS1, MYH10 | 5 |
| Nephrosis | Minimal change nephrotic syndrome | 2.66 × 10−4 | ACTN4, FKBP1A, IMPDH2, PDLIM1 | 4 | |
| Renal necrosis/cell death | Cell viability of kidney cell lines | 2.85 × 10−4 | −0.378 | ANXA5, CAV1, EZR, HNRNPU, HSP90B1, HYOU1, PPIA, VDAC1 | 8 |
| Renal inflammation, renal nephritis | IgA nephropathy | 3.31 × 10−4 | ACTN4, IMPDH2, PDLIM1, PSMB1, PSMD2 | 5 | |
| Renal inflammation, renal nephritis | Fechtner syndrome | 5.69 × 10−4 | MYH10, MYH9 | 2 | |
| Liver cirrhosis | Cryptogenic cirrhosis | 5.69 × 10−4 | KRT18, KRT8 | 2 | |
| Renal inflammation, renal nephritis | Diffuse proliferative lupus nephritis | 5.69 × 10−4 | ACTN4, PDLIM1 | 2 | |
| Renal inflammation, renal nephritis | Epstein syndrome | 5.69 × 10−4 | MYH10, MYH9 | 2 | |
| Liver cirrhosis | Susceptibility to noncryptogenic cirrhosis | 5.69 × 10−4 | KRT18, KRT8 | 2 | |
| Liver hyperplasia/hyperproliferation | Liver cancer | 1.13 × 10−3 | 1.133 | ACLY, ACTG1, ACTR2, ACTR3, AHNAK, ANXA1, ANXA2, AP1B1, APEX1, ARF1, ARHGDIA, ATIC, ATP5A1, ATP5B, B2M, BSG, CACYBP, CAPG, CAST, CBX3, CCT6A, CCT8, CD44, CLIC1, CLTC, COPA, COPG1, COTL1, CRIP2, CSE1L, CTNNA1, CTPS1, CTSA, CTTN, CUTA, CYR61, DBI, DDX3X, DLAT, DNAJA1, DNM1L, DYNC1H1, EEF1A1, EEF2, EIF3I, EIF4A1, ENAH, ENO2, EPRS, FASN, FKBP1A, FLNA, FLNB, FLNC, FN1, GARS, GCN1L1, GDI2, GLUD1, GMPS, GNB2L1, GPI, GSTP1, H2AFY, H3F3A/H3F3B, HADHA, HDLBP, HEXB, HMGB1, HNRNPA1, HNRNPA2B1, HNRNPH1, HNRNPK, HNRNPM, HSP90AA1, HSP90AB1, HSPA5, HSPA8, HSPD1, IARS, IFITM3, IGF2R, ILF2, ILF3, IMPDH2, IPO5, IPO7, IQGAP1, ITGB1, KHSRP, KIF5B, KPNB1, KRT9, LAMB1, LDHA, LMNA, LOC102724594/U2AF1, LONP1, LRPPRC, LRRC59, MAP1B, MAPRE1, MARS, MCM3, MCM4, MCM6, MCM7, MDH1, MVP, MYH10, NACA, NAP1L4, NASP, NNMT, NPEPPS, NPLOC4, NPM1, NUMA1, P4HB, PA2G4, PAICS, PCBP2, PDIA4, PFKP, PFN1, PGK1, PGM1, PHGDH, PKM, PLEC, PPM1G, PPP2CA, PPP2R1A, PRDX1, PRKCSH, PRKDC, PSMC1, PSMD2, PTRF, PUF60, PYGL, QARS, RAN, RPL12, RPL4, RPL7A, RPLP0, RPN1, RPS2, RPS24, RPS4X, RRBP1, RTN4, SDHA, SF3A1, SF3B1, SFPQ, SH3BGRL3, SLC38A2, SND1, SOD1, SPTBN1, SQSTM1, STAT1, TPI1, TUBB4B, TWF2, UAP1, UBE2N, UBXN4, UCHL1, VARS, VCL, VCP, VIM, WARS, XRCC6, YBX3, YWHAG, YWHAH | 180 |
| Liver hyperplasia/hyperproliferation | Liver tumor | 1.16 × 10−3 | 0.590 | ACLY, ACTG1, ACTR2, ACTR3, AHNAK, ANXA1, ANXA2, AP1B1, APEX1, ARF1, ARHGDIA, ATIC, ATP5A1, ATP5B, B2M, BSG, CACYBP, CAPG, CAST, CBX3, CCT6A, CCT8, CD44, CLIC1, CLTC, COPA, COPG1, COTL1, CRIP2, CSE1L, CTNNA1, CTPS1, CTSA, CTTN, CUTA, CYR61, DBI, DDX3X, DLAT, DNAJA1, DNM1L, DYNC1H1, EEF1A1, EEF2, EIF3I, EIF4A1, ENAH, ENO2, EPRS, FASN, FKBP1A, FLNA, FLNB, FLNC, FN1, GARS, GCN1L1, GDI2, GLUD1, GMPS, GNB2L1, GPI, GSTP1, H2AFY, H3F3A/H3F3B, HADHA, HDLBP, HEXB, HMGB1, HNRNPA1, HNRNPA2B1, HNRNPH1, HNRNPK, HNRNPM, HSP90AA1, HSP90AB1, HSPA5, HSPA8, HSPD1, IARS, IFITM3, IGF2R, ILF2, ILF3, IMPDH2, IPO5, IPO7, IQGAP1, ITGB1, KHSRP, KIF5B, KPNB1, KRT9 | 181 |
| LAMB1, LDHA, LMNA, LOC102724594/U2AF1, LONP1, LRPPRC, LRRC59, MAP1B, MAPRE1, MARS, MCM3, MCM4, MCM6, MCM7, MDH1, MVP, MYH10, NACA, NAP1L4, NASP, NNMT, NPEPPS, NPLOC4, NPM1, NUMA1, P4HB, PA2G4, PAICS, PCBP2, PDIA4, PFKP, PFN1, PGK1, PGM1, PHGDH, PKM, PLEC, PPM1G, PPP2CA, PPP2R1A, PRDX1, PRKCSH, PRKDC, PSMC1, PSMD2, PTRF, PUF60, PYGL, QARS, RAN, RPL12, RPL4, RPL7A, RPLP0, RPN1, RPS2, RPS24, RPS4X, RRBP1, RTN4, S100A6, SDHA, SF3A1, SF3B1, SFPQ, SH3BGRL3, SLC38A2, SND1, SOD1, SPTBN1, SQSTM1, STAT1, TPI1, TUBB4B, TWF2, UAP1, UBE2N, UBXN4, UCHL1, VARS, VCL, VCP, VIM, WARS, XRCC6, YBX3, YWHAG, YWHAH | |||||
| Liver damage, liver inflammation/hepatitis | Susceptibility to hepatitis c virus | 1.68 × 10−3 | KRT18, KRT8 | 2 | |
| Renal necrosis/cell death | Cell death of kidney cell lines | 2.01 × 10−3 | 0.222 | CD44, CDK1, EZR, GAPDH, GSTP1, HSPA5, HSPB1, HYOU1, ITGB1, LDHA, PEA15, PRKDC, SOD1, SSRP1, TFRC, VCP, VDAC1, YWHAQ | 18 |
| Cardiac necrosis/cell death | Cell death of cardiomyocytes | 2.22 × 10−3 | 1.920 | CALR, CYR61, GAPDH, GNAI2, HADHA, HSPB1, HSPD1, MAPK1, NCL, PARK7, RHOA, RPSA, STAT1, ZYX | 14 |
| Renal necrosis/cell death | Cell death of kidney cells | 2.29 × 10−3 | 0.515 | ATP1A1, CD44, CDK1, EZR, GAPDH, GSTP1, HSPA5, HSPB1, HYOU1, ITGB1, LDHA, PEA15, PRKDC, SOD1, SSRP1, STMN1, TFRC, VCP, VDAC1, YWHAQ | 20 |
| Renal proliferation | Proliferation of kidney cell lines | 4.64 × 10−3 | −2.423 | HSP90AA1, HSP90AB1, HSPA4, HSPD1, ITGB1, LMNA, NCL, PPP2R1A, RHOA, VDAC1, YBX3 | 11 |
| Cardiac damage, cardiac degeneration | Degeneration of cardiomyocytes | 4.98 × 10−3 | CAV1, HADHA, MAPK1 | 3 | |
| Nephrosis | Autosomal recessive steroid-resistant nephrotic syndrome | 5.43 × 10−3 | ARHGDIA, IMPDH2 | 2 | |
| Decreased levels of albumin | Increased uptake of albumin | 5.43 × 10−3 | CAV1, FLNA | 2 | |
| Cardiac hypertrophy | Hypertrophy of heart cells | 5.90 × 10−3 | −0.243 | CAV1, CTSB, EEF1D, FN1, HMGB1, IGF2R, MAPK1, PPIA, RHOA, S100A6 | 10 |
| Hepatocellular carcinoma, liver hyperplasia/hyperproliferation | Hepatocellular carcinoma | 5.96 × 10−3 | 1.133 | ACTG1, ARF1, BSG, COPG1, CSE1L, CTTN, FKBP1A, GLUD1, GNB2L1, H2AFY, HMGB1, HNRNPA1, HSPA5, IFITM3, IGF2R, IPO7, IQGAP1, MAPRE1, NNMT, PFN1, PPM1G, PRDX1, SOD1, SPTBN1, TPI1, TUBB4B, YWHAG | 27 |
| Cardiac necrosis/cell death | Cell viability of cardiomyocytes | 6.07 × 10−3 | CALR, HSPD1, IQGAP1, RHOA | 4 | |
| Liver fibrosis, liver proliferation | Proliferation of hepatic stellate cells | 6.67 × 10−3 | −0.847 | CTSB, ITGB1, LGALS1, SERPINH1, STAT1 | 5 |
| Cardiac arrythmia | Chronic atrial fibrillation | 8.01 × 10−3 | PPP1CA, PPP2CA | 2 | |
| Renal dilation | Dilation of collecting tubule | 8.01 × 10−3 | ARHGDIA, PRKCSH | 2 | |
| Kidney failure | Acute renal failure | 9.94 × 10−3 | ATP1A1, CYR61, GSTP1, HEXB, STMN1 | 5 | |
| Liver damage, liver inflammation/hepatitis | Hepatitis c | 1.51 × 10−2 | DDX5, FKBP1A, IMPDH2, KRT18, KRT8 | 5 | |
| Liver damage | Damage of liver | 1.67 × 10−2 | 0.513 | BSG, CD44, CTSB, DDX5, FKBP1A, GSTP1, HLA-B, HMGB1, IMPDH2, KRT18, KRT8, SOD1, STAT1 | 13 |
| Cardiac inflammation | Inflammation of heart | 1.76 × 10−2 | 0.686 | B2M, CAV1, HMGB1, PPIA | 4 |
| Glomerular injury | Focal segmental glomerulosclerosis | 1.89 × 10−2 | ACTN4, MYH9, PDLIM1 | 3 | |
| Cardiac dilation | Dilation of left ventricle | 1.90 × 10−2 | −1.000 | CAPNS1, CAV1, LMNA, RHOA | 4 |
| Glomerular injury | Autosomal dominant focal segmental glomerulosclerosis | 2.39 × 10−2 | ACTN4 | 1 | |
| Liver inflammation/hepatitis | Chronic phase hepatitis | 2.39 × 10−2 | HLA-A | 1 | |
| Decreased levels of albumin | Decreased accumulation of albumin | 2.39 × 10−2 | CAV1 | 1 | |
| Renal degeneration | Degeneration of renal tubular epithelial cells | 2.39 × 10−2 | ARHGDIA | 1 | |
| Cardiac dysplasia | Dysplasia of right ventricle | 2.39 × 10−2 | RPSA | 1 | |
| Cardiac arrythmia, tachycardia | Familial ventricular tachycardia | 2.39 × 10−2 | GNAI2 | 1 | |
| Cardiac fibrosis | Fibrosis of right ventricle | 2.39 × 10−2 | CTNNA1 | 1 | |
| Kidney failure | Gentamicin-induced renal injury | 2.39 × 10−2 | HEXB | 1 | |
| Glomerular injury | Glomerulopathy with fibronectin deposits type 2 | 2.39 × 10−2 | FN1 | 1 | |
| Pulmonary hypertension | Hypertension of pulmonary artery | 2.39 × 10−2 | CAV1 | 1 | |
| Increased levels of albumin | Increased export of albumin | 2.39 × 10−2 | RAN | 1 | |
| Liver failure | Infantile liver failure syndrome type 2 | 2.39 × 10−2 | MARS | 1 | |
| Liver hyperplasia/hyperproliferation | Nodular hyperplasia of liver | 2.39 × 10−2 | SOD1 | 1 | |
| Pulmonary hypertension | Primary pulmonary hypertension 3 | 2.39 × 10−2 | CAV1 | 1 | |
| Cardiac damage | Rupture of ventricular septum | 2.39 × 10−2 | CTNNA1 | 1 | |
| Bradycardia, cardiac arrythmia | Severe bradycardia | 2.39 × 10−2 | RHOA | 1 | |
| Kidney failure | Failure of kidney | 2.48 × 10−2 | ACTN4, ARHGDIA, ATP1A1, CYR61, FDPS, FKBP1A, GSTP1, HEXB, IMPDH2, MYH9, STMN1 | 11 | |
| Renal inflammation, renal nephritis | Autoimmune glomerulonephritis | 2.85 × 10−2 | ACTN4, PDLIM1, PPP2CA | 3 | |
| Liver cirrhosis | Cirrhosis | 3.11 × 10−2 | B2M, BSG, FN1, GSTP1, HNRNPK, KRT18, KRT8, PCNA, TXLNA | 9 | |
| Liver necrosis/cell death | Necrosis of liver | 3.18 × 10−2 | 0.791 | CTSB, FKBP1A, FLNA, HMGB1, HSPD1, KRT8, NCL, SERPINH1, SLC25A5, SOD1, SPTBN1, STAT1 | 12 |
| Cardiac damage | Damage of heart | 3.20 × 10−2 | 0.156 | ANXA1, CAV1, CTNNA1, HADHA, MAPK1 | 5 |
| Liver inflammation/hepatitis | Inflammation of liver | 3.24 × 10−2 | 1.231 | CD44, CYR61, DDX5, FKBP1A, GSTP1, HLA-A, HMGB1, IMPDH2, KRT18, KRT8, PPIA, SOD1, STAT1 | 13 |
| Renal inflammation, renal nephritis | Lupus nephritis | 3.28 × 10−2 | ACTN4, FKBP1A, IMPDH2, PDLIM1, PSMB1, PSMD2 | 6 | |
| Cardiac arrythmia | Arrhythmia | 3.34 × 10−2 | −1.103 | ATP1A1, CALR, GNAI2, HMGB1, PPP1CA, PPP2CA, RHOA, TUBB, TUBB4B, VCL | 10 |
| Liver damage | Injury of liver | 3.41 × 10−2 | −0.447 | BSG, CTSB, HLA-B, HMGB1, KRT8, SOD1, STAT1 | 7 |
| Hepatocellular carcinoma, liver hyperplasia/hyperproliferation | Incidence of hepatocellular carcinoma | 3.54 × 10−2 | 1.718 | IQGAP1, PRDX1, SOD1, SPTBN1 | 4 |
| Glomerular injury, renal hypertrophy | Hypertrophy of mesangial cells | 3.73 × 10−2 | MAPK1, PARK7 | 2 | |
| Liver hepatomegaly | Hepatomegaly | 4.07 × 10−2 | CTSA, HADHA, IGF2R, KRT18, STAT1 | 5 | |
| Cardiac hypertrophy | Hypertrophy of cardiomyocytes | 4.35 × 10−2 | −0.243 | CAV1, FN1, HMGB1, IGF2R, MAPK1, PPIA, RHOA | 7 |
| Cardiac arrythmia | Supraventricular arrhythmia | 4.35 × 10−2 | ATP1A1, CALR, PPP1CA, PPP2CA, RHOA, TUBB, TUBB4B | 7 | |
| Congenital heart anomaly | Ventricular septal defect | 4.67 × 10−2 | 0.128 | AIP, FKBP1A, FLNA, MYH10, MYH9, YWHAE | 6 |
| Liver degeneration | Degeneration of hepatocytes | 4.72 × 10−2 | XRCC5 | 1 | |
| Increased levels of potassium | Increased efflux of k+ | 4.72 × 10−2 | MAPK1 | 1 | |
| Liver fibrosis, liver proliferation | Proliferation of hepatic stellate cells/myofibroblasts | 4.72 × 10−2 | CTSB | 1 | |
| Cardiac regeneration | Regeneration of cardiomyocytes | 4.72 × 10−2 | HMGB1 | 1 | |
| Cardiac regeneration | Regeneration of myocardium | 4.72 × 10−2 | HMGB1 | 1 | |
| Liver hyperplasia/hyperproliferation | Hyperplasia of liver | 4.87 × 10−2 | IGF2R, SOD1 | 2 | |
| Renal inflammation, renal nephritis | Membranous glomerulonephritis | 4.87 × 10−2 | ACTN4, PDLIM1 | 2 | |
| Renal necrosis/cell death | Apoptosis of kidney cell lines | 5.43 × 10−2 | −0.777 | CD44, CDK1, EZR, HSPA5, HSPB1, HYOU1, ITGB1, PEA15, TFRC, VDAC1, YWHAQ | 11 |
| Cardiac necrosis/cell death | Apoptosis of cardiomyocytes | 5.55 × 10−2 | 1.348 | CALR, CYR61, GAPDH, GNAI2, HSPD1, MAPK1, PARK7, RHOA, STAT1 | 9 |
| Cardiac hypertrophy | Hypertrophy of heart | 5.62 × 10−2 | −0.859 | CAV1, CTSB, EEF1D, FN1, GNAI2, HMGB1, IGF2R, MAPK1, MYH10, PARK7, PFN1, PPIA, RHOA, S100A6, TLN1 | 15 |
| Liver cirrhosis | Cirrhosis of liver | 6.13 × 10−2 | B2M, BSG, FN1, HNRNPK, KRT18, KRT8, TXLNA | 7 | |
| Cardiac arrythmia | Atrial fibrillation | 6.18 × 10−2 | ATP1A1, PPP1CA, PPP2CA, RHOA, TUBB, TUBB4B | 6 | |
| Cardiac inflammation | Pericarditis | 6.77 × 10−2 | TUBB, TUBB4B | 2 | |
| Liver steatosis | Advanced stage hepatic steatosis | 7.00 × 10−2 | CD44 | 1 | |
| Cardiac arteriopathy, cardiac hypertrophy | Hypertrophy of coronary artery | 7.00 × 10−2 | RHOA | 1 | |
| Glomerular injury, renal hypoplasia | Hypoplasia of mesangial cells | 7.00 × 10−2 | ARHGDIA | 1 | |
| Cardiac inflammation | Rheumatic carditis | 7.00 × 10−2 | ANXA1 | 1 | |
| Cardiac arteriopathy | Vasospasm of coronary artery | 7.00 × 10−2 | SOD1 | 1 | |
| Liver fibrosis | Fibrosis of liver | 8.42 × 10−2 | 0.391 | CTSB, GNB2L1, GSTP1, HDGF, HSPB1, STAT1 | 6 |
| Cardiac fibrosis | Fibrosis of heart | 9.03 × 10−2 | −0.956 | CAPNS1, CAV1, CTNNA1, FN1, ITGB1, LMNA, RHOA, TLN1 | 8 |
| Liver hypoplasia | Hypoplasia of liver | 9.21 × 10−2 | EIF6, SPTBN1, YBX1 | 3 | |
| Renal damage, renal tubule injury | Damage of tubular cells | 9.22 × 10−2 | YBX1 | 1 | |
| Glomerular injury, renal fibrosis | Fibrosis of renal tubule | 9.22 × 10−2 | STMN1 | 1 | |
| Renal inflammation, renal nephritis | Progressive crescentic glomerulonephritis | 9.22 × 10−2 | ACTN4 | 1 | |
| Renal inflammation, renal nephritis | Nephritis | 1.12 × 10−1 | ACTN4, ARHGDIA, FKBP1A, IMPDH2, MYH10, MYH9, PDLIM1, PPP2CA, PSMB1, PSMD2, YBX1 | 11 | |
| Renal damage | Damage of kidney | 1.13 × 10−1 | −0.447 | ARHGDIA, B2M, CD44, FN1, ITGB1, YBX1 | 6 |
| Renal proliferation | Arrest in growth of kidney cell lines | 1.14 × 10−1 | LMNA | 1 | |
| Cardiac enlargement | Enlargement of atrium | 1.14 × 10−1 | RHOA | 1 | |
| Cardiac arteriopathy, cardiac fibrosis | Fibrosis of coronary artery | 1.14 × 10−1 | RHOA | 1 | |
| Cardiac damage, cardiac degeneration | Injury of cardiomyocytes | 1.14 × 10−1 | MAPK1 | 1 | |
| Liver steatosis | Nonalcoholic fatty liver disease | 1.22 × 10−1 | FASN, HADHA, SOD1 | 3 | |
| Liver damage | Hepatotoxicity | 1.27 × 10−1 | AIP, HLA-B | 2 | |
| Heart failure | Failure of heart | 1.31 × 10−1 | −0.798 | ATP1A1, CAPNS1, CAV1, FKBP1A, HMGB1, IGF2R, MAPK1, MYH10, PARK7, PPP2R1A | 10 |
| Liver proliferation | Proliferation of liver cells | 1.34 × 10−1 | −1.461 | CAV1, CTSB, ITGB1, LGALS1, MAPK1, SERPINH1, SPTBN1, STAT1 | 8 |
| Kidney failure | Acute tubular necrosis | 1.35 × 10−1 | STMN1 | 1 | |
| Liver adhesion | Adhesion of hepatocytes | 1.35 × 10−1 | KRT8 | 1 | |
| Heart failure | End stage heart failure | 1.35 × 10−1 | PARK7 | 1 | |
| Liver enlargement | Enlargement of liver | 1.35 × 10−1 | LMNA | 1 | |
| Liver hyperplasia/hyperproliferation | Polycystic liver disease | 1.35 × 10−1 | PRKCSH | 1 | |
| Cardiac necrosis/cell death | Survival of ventricular myocytes | 1.35 × 10−1 | CALR | 1 | |
| Renal transformation | Transformation of kidney cells | 1.35 × 10−1 | EIF2S1 | 1 | |
| Cardiac dysfunction | Systolic dysfunction | 1.52 × 10−1 | ANXA5, FKBP1A | 2 | |
| Liver necrosis/cell death | Cell death of liver cells | 1.52 × 10−1 | 0.256 | CTSB, HMGB1, HSPD1, KRT8, NCL, SERPINH1, SLC25A5, SPTBN1 | 8 |
| Renal proliferation | Proliferation of renal tubular epithelial cells | 1.56 × 10−1 | STMN1 | 1 | |
| Cardiac arrythmia, tachycardia | Ventricular tachycardia | 1.61 × 10−1 | ATP1A1, GNAI2, VCL | 3 | |
| Liver hemorrhaging | Bleeding of liver | 1.69 × 10−1 | FKBP1A, KRT8 | 2 | |
| Renal inflammation, renal nephritis | Acute phase crescentic glomerulonephritis | 1.76 × 10−1 | ACTN4 | 1 | |
| Liver fibrosis | Chemotaxis of hepatic stellate cells | 1.76 × 10−1 | MAPK1 | 1 | |
| Increased levels of albumin | Increased localization of albumin | 1.76 × 10−1 | B2M | 1 | |
| Hepatocellular peroxisome proliferation | Quantity of peroxisomes | 1.76 × 10−1 | DNM1L | 1 | |
| Bradycardia, cardiac arrythmia | Sinus bradycardia | 1.76 × 10−1 | CALR | 1 | |
| Cardiac inflammation | Carditis | 1.85 × 10−1 | ANXA1, TUBB, TUBB4B | 3 | |
| Bradycardia, cardiac arrythmia | Bradycardia | 1.86 × 10−1 | CALR, RHOA | 2 | |
| Renal dilation | Dilation of kidney | 1.96 × 10−1 | PRKCSH | 1 | |
| Hepatocellular carcinoma, liver hyperplasia/hyperproliferation | Size of hepatocellular carcinoma | 1.96 × 10−1 | IQGAP1 | 1 | |
| Liver cirrhosis | Primary biliary cirrhosis | 1.97 × 10−1 | B2M, BSG, HNRNPK, TXLNA | 4 | |
| Liver necrosis/cell death | Cell death of hepatocytes | 2.11 × 10−1 | −0.537 | CTSB, HMGB1, KRT8, NCL, SLC25A5, SPTBN1 | 6 |
| Glomerular injury | Glomerulosclerosis | 2.13 × 10−1 | ACTN4, ARHGDIA, MYH9, PDLIM1, STMN1 | 5 | |
| Renal necrosis/cell death | Apoptosis of proximal tubule cells | 2.15 × 10−1 | ATP1A1 | 1 | |
| Cardiac infarction | Infarction of heart | 2.15 × 10−1 | CAV1 | 1 | |
| Kidney failure | Ischemic acute renal failure | 2.15 × 10−1 | ATP1A1 | 1 | |
| Renal necrosis/cell death | Apoptosis of tubular cells | 2.21 × 10−1 | ATP1A1, STMN1 | 2 | |
| Liver necrosis/cell death | Apoptosis of liver cells | 2.22 × 10−1 | 0.000 | CTSB, HMGB1, KRT8, NCL, SERPINH1, SPTBN1 | 6 |
| Increased levels of creatinine | Increased quantity of creatinine | 2.30 × 10−1 | AKR1B1, CD44 | 2 | |
| Glomerular injury | Cortical renal glomerulopathies | 2.34 × 10−1 | ACTN4 | 1 | |
| Cardiac hyperplasia/hyperproliferation | Hyperplasia of heart | 2.34 × 10−1 | IGF2R | 1 | |
| Liver necrosis/cell death | Apoptosis of hepatocytes | 2.48 × 10−1 | −0.492 | CTSB, HMGB1, KRT8, NCL, SPTBN1 | 5 |
| Decreased levels of albumin | Decreased excretion of albumin | 2.52 × 10−1 | LAMB1 | 1 | |
| Cardiac proliferation | Proliferation of cardiac fibroblasts | 2.52 × 10−1 | STAT1 | 1 | |
| Cardiac congestive cardiac failure, heart failure | Congestive heart failure | 2.58 × 10−1 | −1.103 | CAV1, FKBP1A, IGF2R, MYH10 | 4 |
| Cardiac damage | Injury of heart | 2.65 × 10−1 | ANXA1, MAPK1 | 2 | |
| Renal damage, renal tubule injury | Damage of tubulointerstitium | 2.70 × 10−1 | FN1 | 1 | |
| Increased levels of creatinine | Increased clearance of creatinine | 2.70 × 10−1 | ARHGDIA | 1 | |
| Renal damage, renal tubule injury | Damage of renal tubule | 2.74 × 10−1 | FN1, YBX1 | 2 | |
| Liver steatosis | Hepatic steatosis | 2.75 × 10−1 | 1.715 | CAV1, CD44, FASN, GSTP1, H2AFY, HADHA, KRT8, LMNA, SOD1 | 9 |
| Congenital heart anomaly | Congenital heart disease | 2.85 × 10−1 | 0.294 | AIP, ENO2, FKBP1A, FLNA, MYH10, MYH9, YWHAE | 7 |
| Liver inflammation/hepatitis | Acute hepatitis | 3.04 × 10−1 | CD44 | 1 | |
| Increased levels of blood urea nitrogen | Increased quantity of blood urea nitrogen | 3.04 × 10−1 | CD44 | 1 | |
| Liver fibrosis | Activation of hepatic stellate cells | 3.10 × 10−1 | FN1, STAT1 | 2 | |
| Liver damage, liver inflammation/hepatitis | Chronic hepatitis c | 3.10 × 10−1 | DDX5, IMPDH2 | 2 | |
| Kidney failure | End stage renal disease | 3.12 × 10−1 | FDPS, FKBP1A, IMPDH2, MYH9 | 4 | |
| Liver necrosis/cell death | Focal necrosis of liver | 3.21 × 10−1 | STAT1 | 1 | |
| Increased levels of potassium | Increased excretion of k+ | 3.21 × 10−1 | AKR1B1 | 1 | |
| Renal inflammation, renal nephritis | Glomerulonephritis | 3.23 × 10−1 | ACTN4, FKBP1A, IMPDH2, PDLIM1, PPP2CA, PSMB1, PSMD2 | 7 | |
| Cardiac arteriopathy | Coronary artery disease | 3.23 × 10−1 | ACAT2, EIF4H, FKBP1A, NNMT, RHOA, SOD1, TKT, TUBB, TUBB4B | 9 | |
| Cardiac infarction | Myocardial infarction | 3.27 × 10−1 | 2.000 | ACTA1, CCT7, CD44, MAPK1, SOD1, STAT1 | 6 |
| Renal damage | Injury of kidney | 3.32 × 10−1 | ARHGDIA, B2M, CD44 | 3 | |
| Renal inflammation, renal nephritis | Interstitial nephritis | 3.37 × 10−1 | ARHGDIA | 1 | |
| Renal necrosis/cell death | Apoptosis of kidney cells | 3.51 × 10−1 | ATP1A1, ITGB1, STMN1 | 3 | |
| Cardiac hypoplasia | Hypoplasia of heart ventricle | 3.62 × 10−1 | CALR, MYH10 | 2 | |
| Liver necrosis/cell death | Apoptosis of hepatic stellate cells | 3.68 × 10−1 | SERPINH1 | 1 | |
| Liver hyperbilirubinemia | Hyperbilirubinemia | 3.68 × 10−1 | GSTP1 | 1 | |
| Renal damage | Injury of renal glomerulus | 3.68 × 10−1 | B2M | 1 | |
| Glutathione depletion in liver | Conjugation of glutathione | 3.84 × 10−1 | GSTP1 | 1 | |
| Cardiac fibrosis | Fibrosis of myocardium | 3.84 × 10−1 | ITGB1 | 1 | |
| Increased levels of hematocrit | Increased hematocrit of organism | 4.03 × 10−1 | CD59, PRDX1, PRDX2 | 3 | |
| Cardiac hypertrophy | Hypertrophy of ventricular myocytes | 4.13 × 10−1 | RHOA | 1 | |
| Liver cholestasis | Progressive familial intrahepatic cholestasis type 1 | 4.13 × 10−1 | HDLBP, SLC25A6 | 2 | |
| Increased levels of red blood cells | Increased quantity of red blood cells | 4.22 × 10−1 | CD59, PICALM, SOD1 | 3 | |
| Renal necrosis/cell death | Apoptosis of mesangial cells | 4.27 × 10−1 | ITGB1 | 1 | |
| Renal dysfunction | Dysfunction of kidney | 4.40 × 10−1 | BSG | 1 | |
| Congenital heart anomaly | Perimembranous ventricular septal defect | 4.40 × 10−1 | MYH10 | 1 | |
| Cardiac hypertrophy | Hypertrophy of right ventricle | 4.54 × 10−1 | CAV1 | 1 | |
| Cardiac necrosis/cell death | Apoptosis of ventricular myocytes | 4.67 × 10−1 | RHOA | 1 | |
| Cardiac dilation | Dilation of heart | 4.67 × 10−1 | PPP2CA | 1 | |
| Cardiac fibrosis | Interstitial fibrosis of heart | 4.67 × 10−1 | CAPNS1 | 1 | |
| Cardiac arrythmia | Ventricular fibrillation | 4.67 × 10−1 | ATP1A1 | 1 | |
| Cardiac arrythmia | Arrhythmia of heart ventricle | 4.78 × 10−1 | ATP1A1, GNAI2 | 2 | |
| Liver hyperplasia/hyperproliferation | Proliferation of liver cancer cells | 4.80 × 10−1 | S100A6 | 1 | |
| Increased levels of alkaline phosphatase | Increased activation of alkaline phosphatase | 4.93 × 10−1 | ANXA6, ITGB1 | 2 | |
| Renal damage | Reperfusion injury of kidney | 5.04 × 10−1 | CD44 | 1 | |
| Cardiac arrythmia, tachycardia | Supraventricular tachycardia | 5.28 × 10−1 | ATP1A1 | 1 | |
| Liver inflammation/hepatitis | Alcoholic hepatitis | 5.39 × 10−1 | GSTP1 | 1 | |
| Renal proliferation | Proliferation of kidney cells | 5.45 × 10−1 | AKR1B1, STMN1 | 2 | |
| Congenital heart anomaly | Conotruncal heart malformations | 5.59 × 10−1 | AIP, FLNA | 2 | |
| Congenital heart anomaly | Persistent truncus arteriosus | 5.61 × 10−1 | FLNA | 1 | |
| Cardiac hypoplasia | Hypoplasia of trabeculae carne | 5.92 × 10−1 | CALR | 1 | |
| Cardiac output | Cardiac output | 6.01 × 10−1 | MAPK1 | 1 | |
| Liver inflammation/hepatitis, liver steatosis | Steatohepatitis | 6.29 × 10−1 | SOD1 | 1 | |
| Cardiac hypertrophy | Hypertrophy of left ventricle | 1.00E00 | CAV1 | 1 | |
| Liver proliferation | Proliferation of hepatocytes | 1.00E00 | CAV1, MAPK1, SPTBN1 | 3 | |
| Cardiac hypertrophy | Ventricular hypertrophy | 1.00E00 | CAV1, RHOA | 2 |

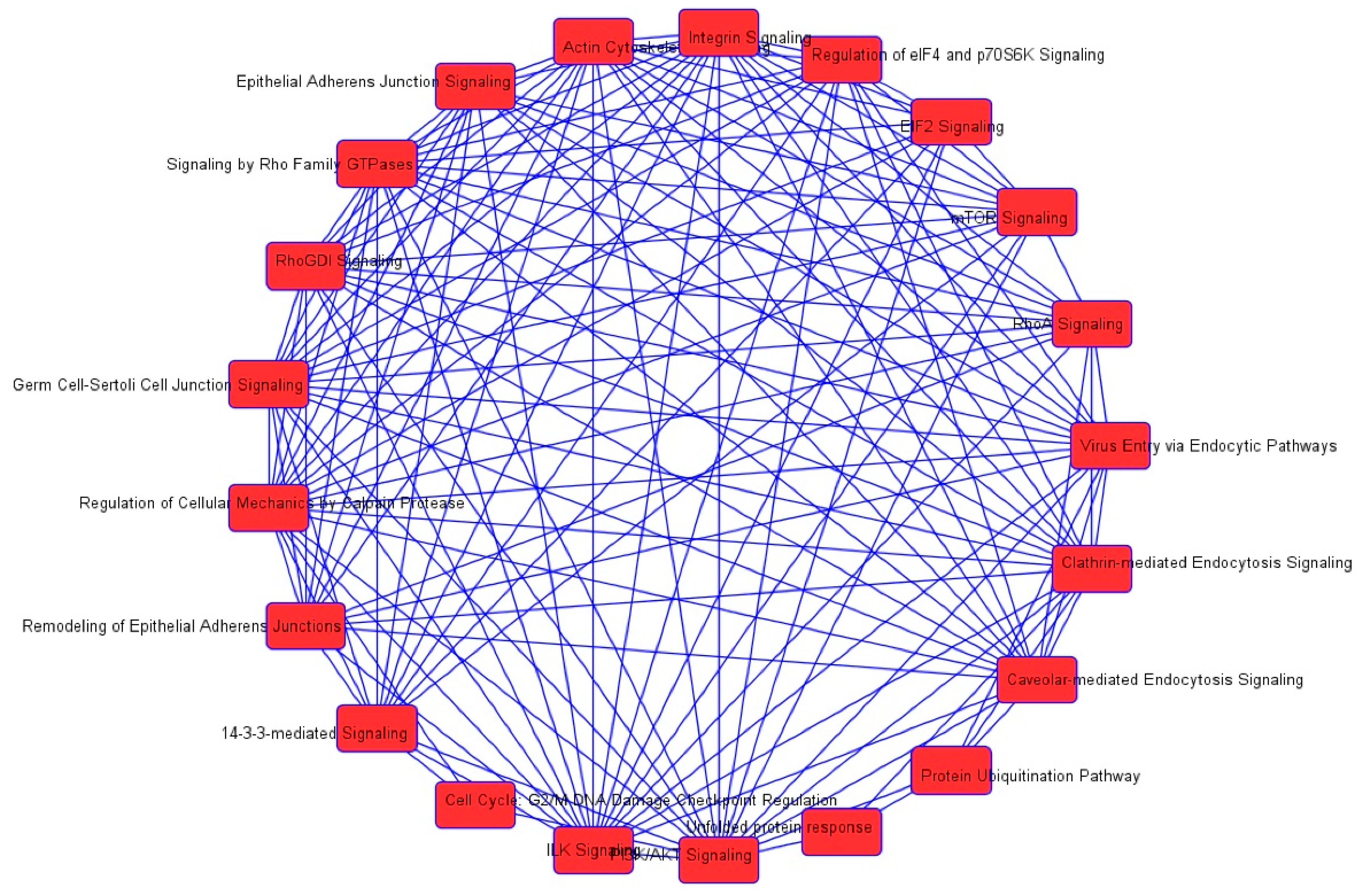
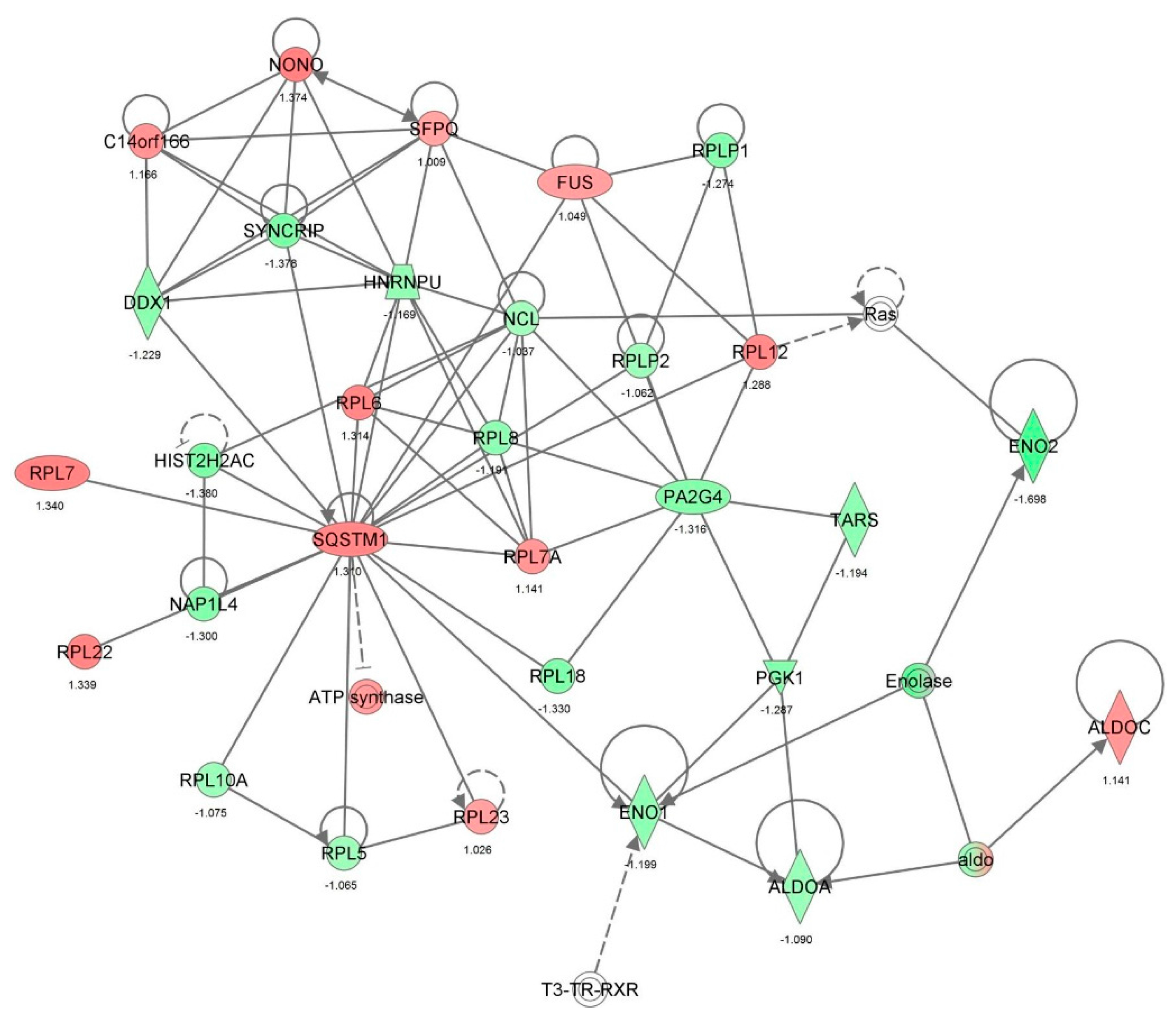
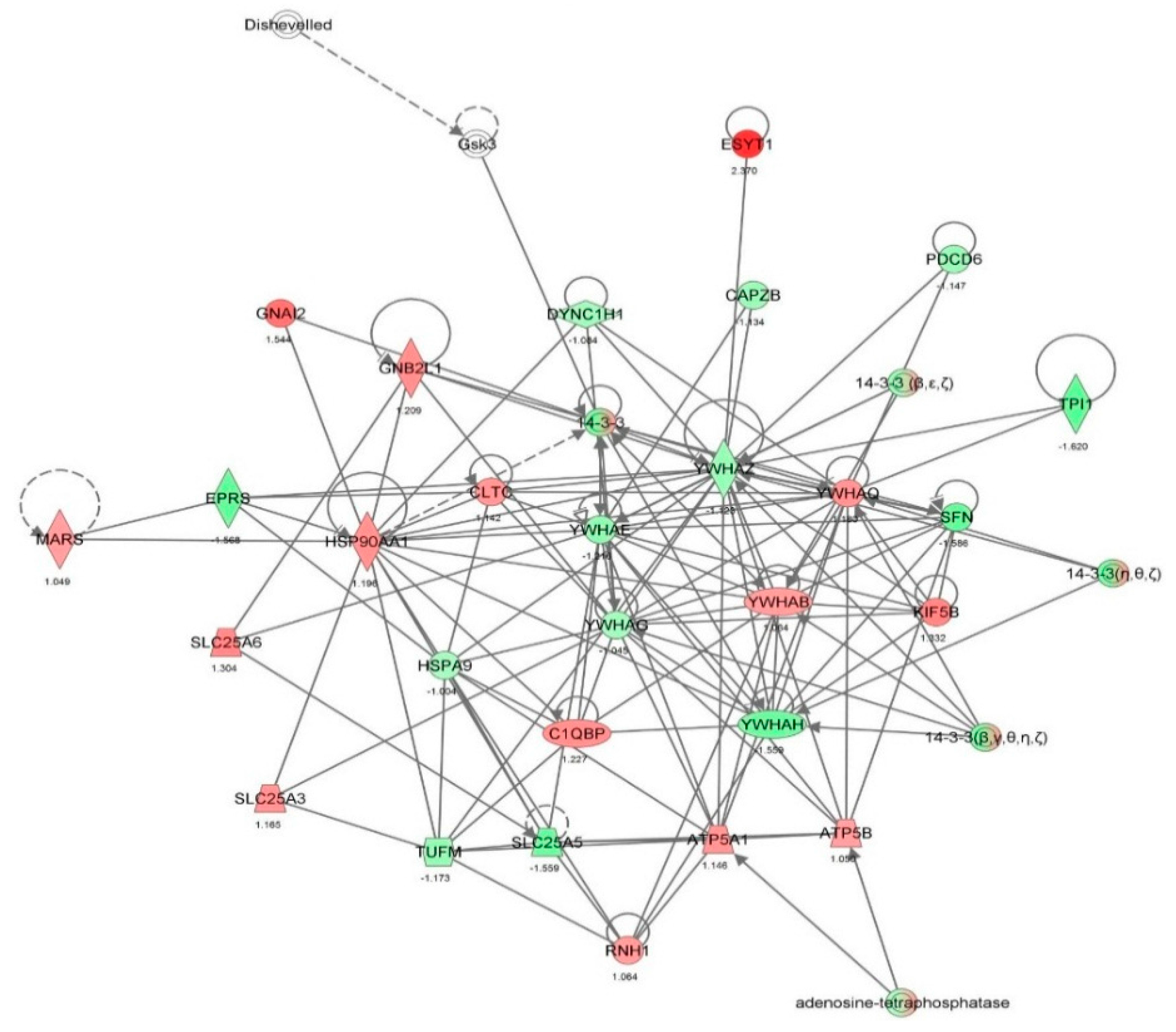
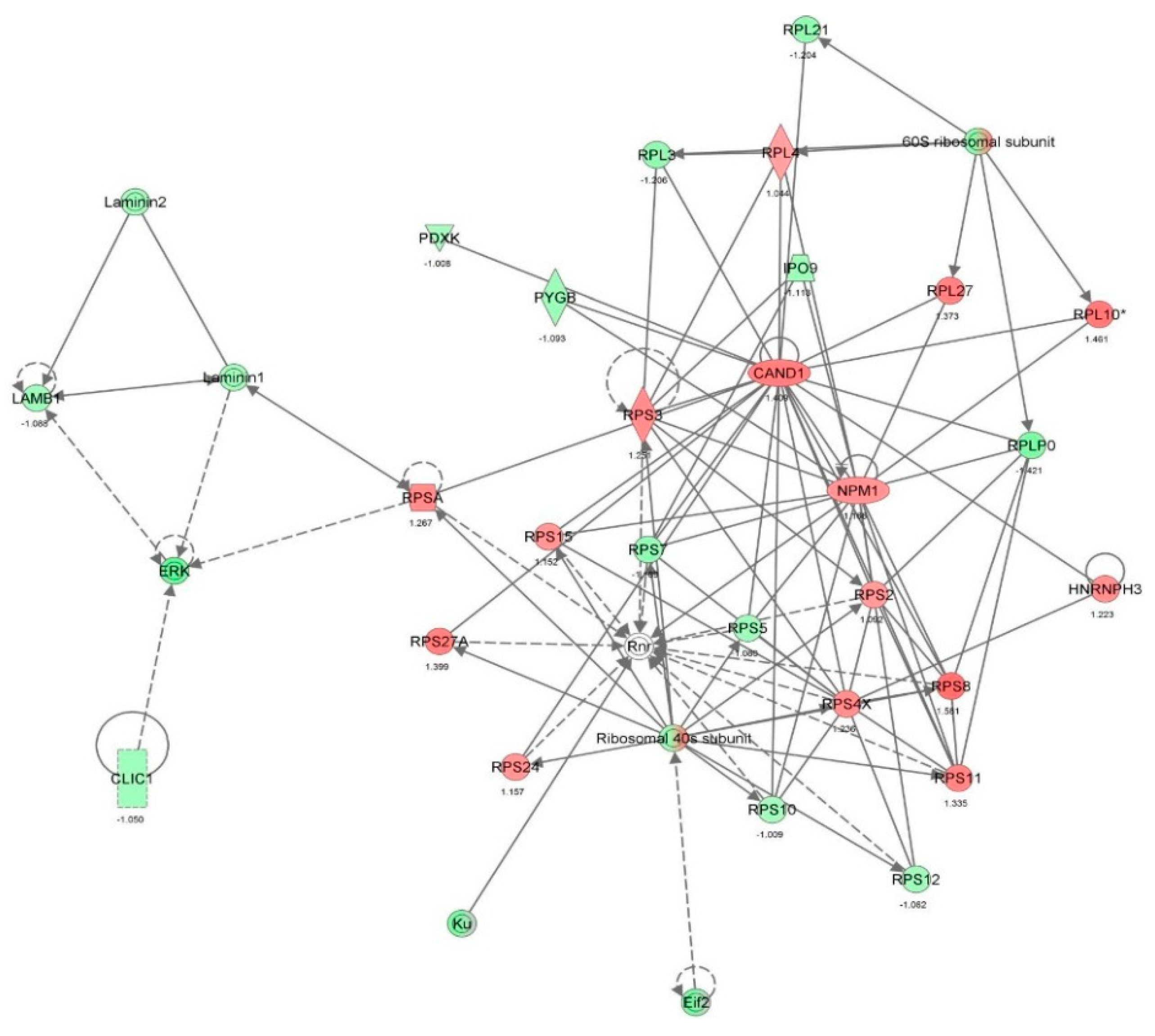
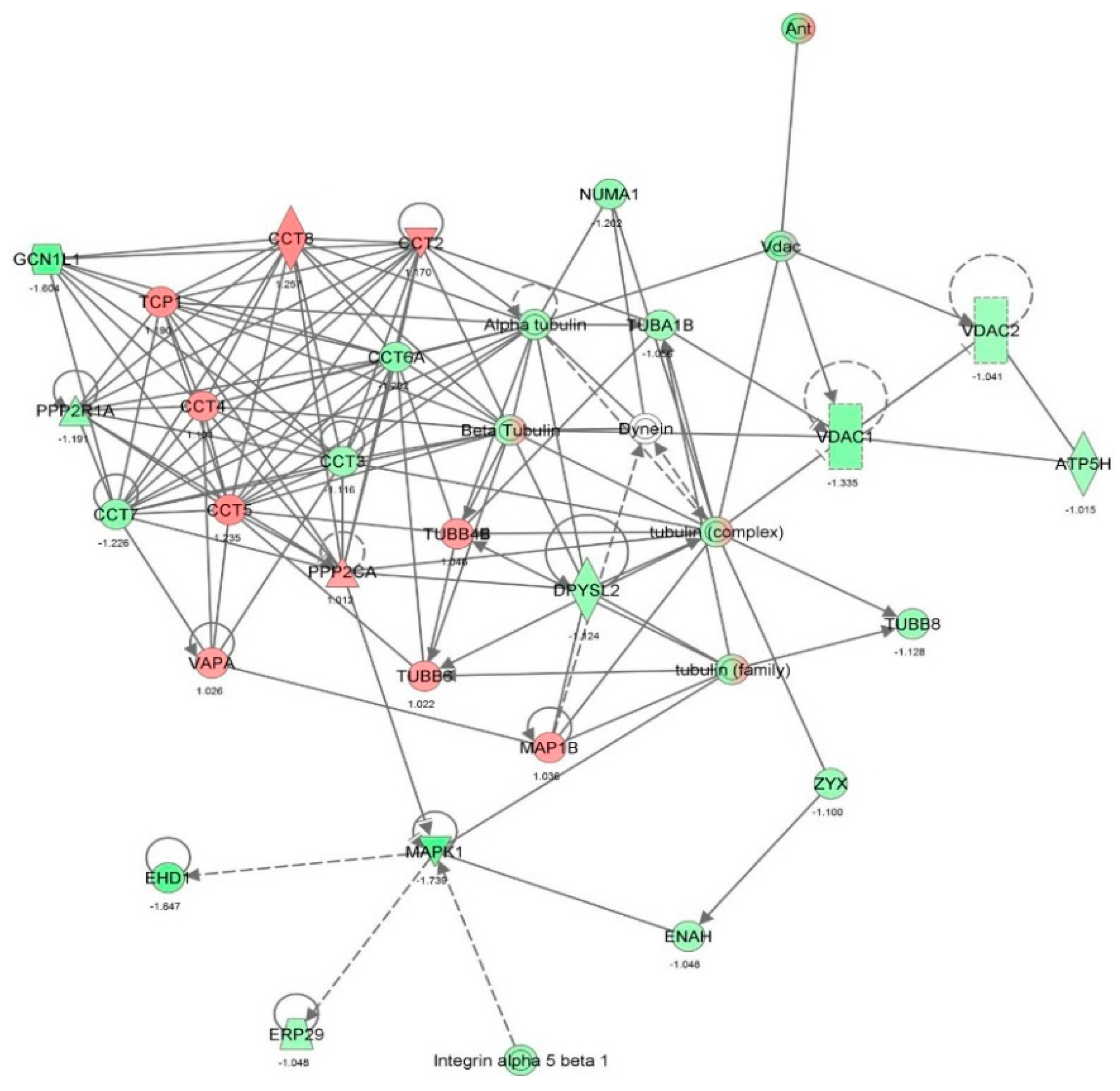
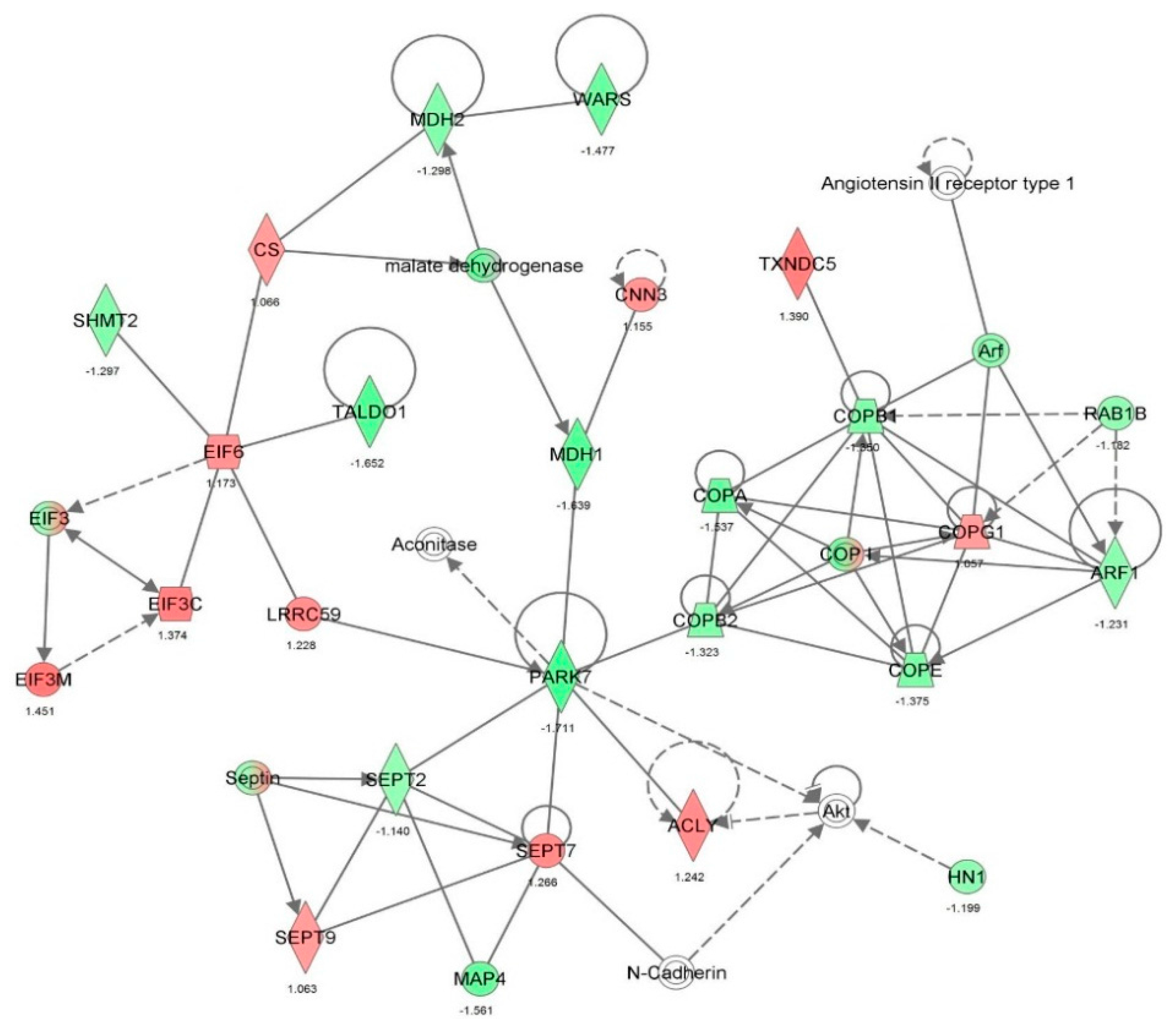

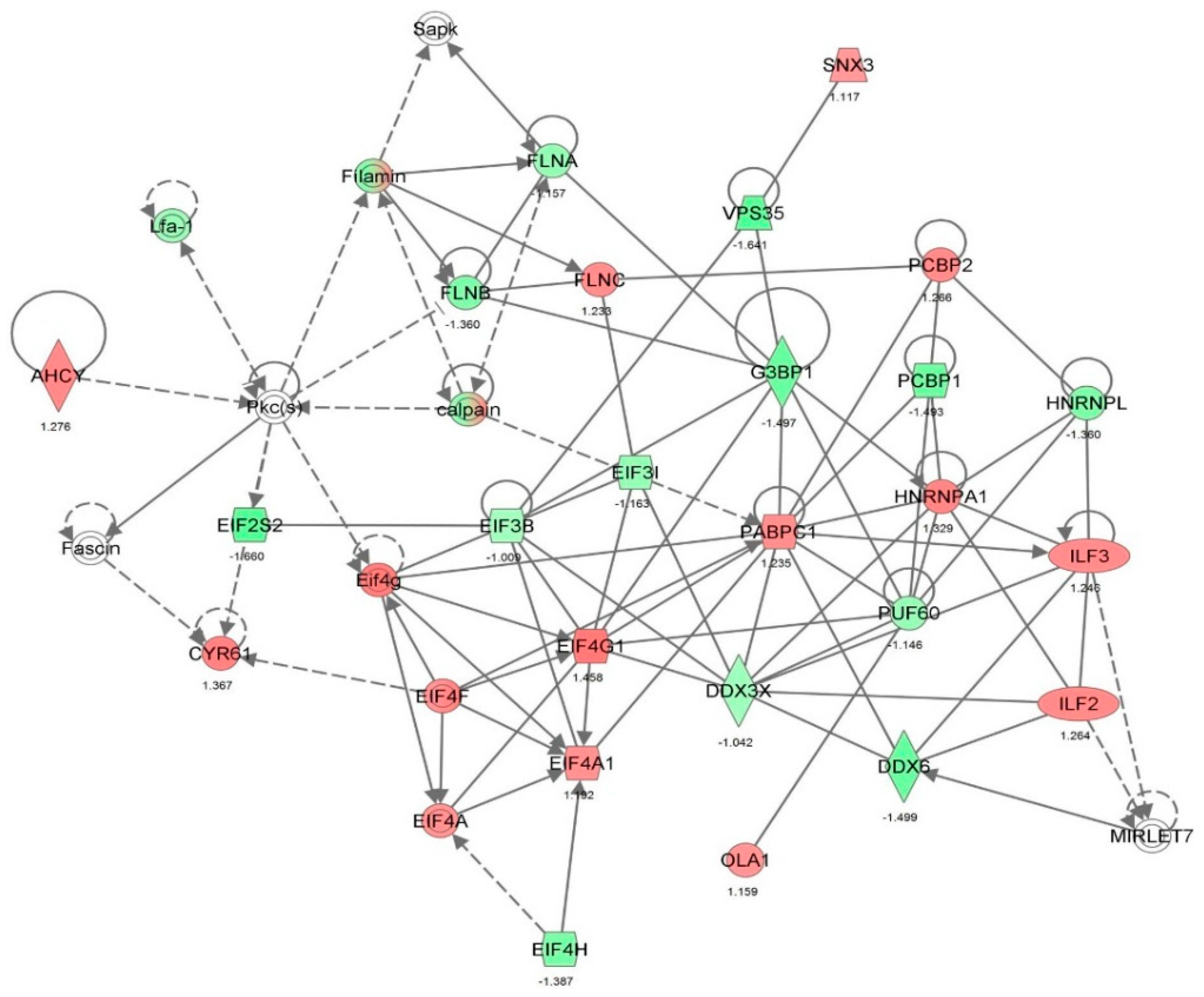
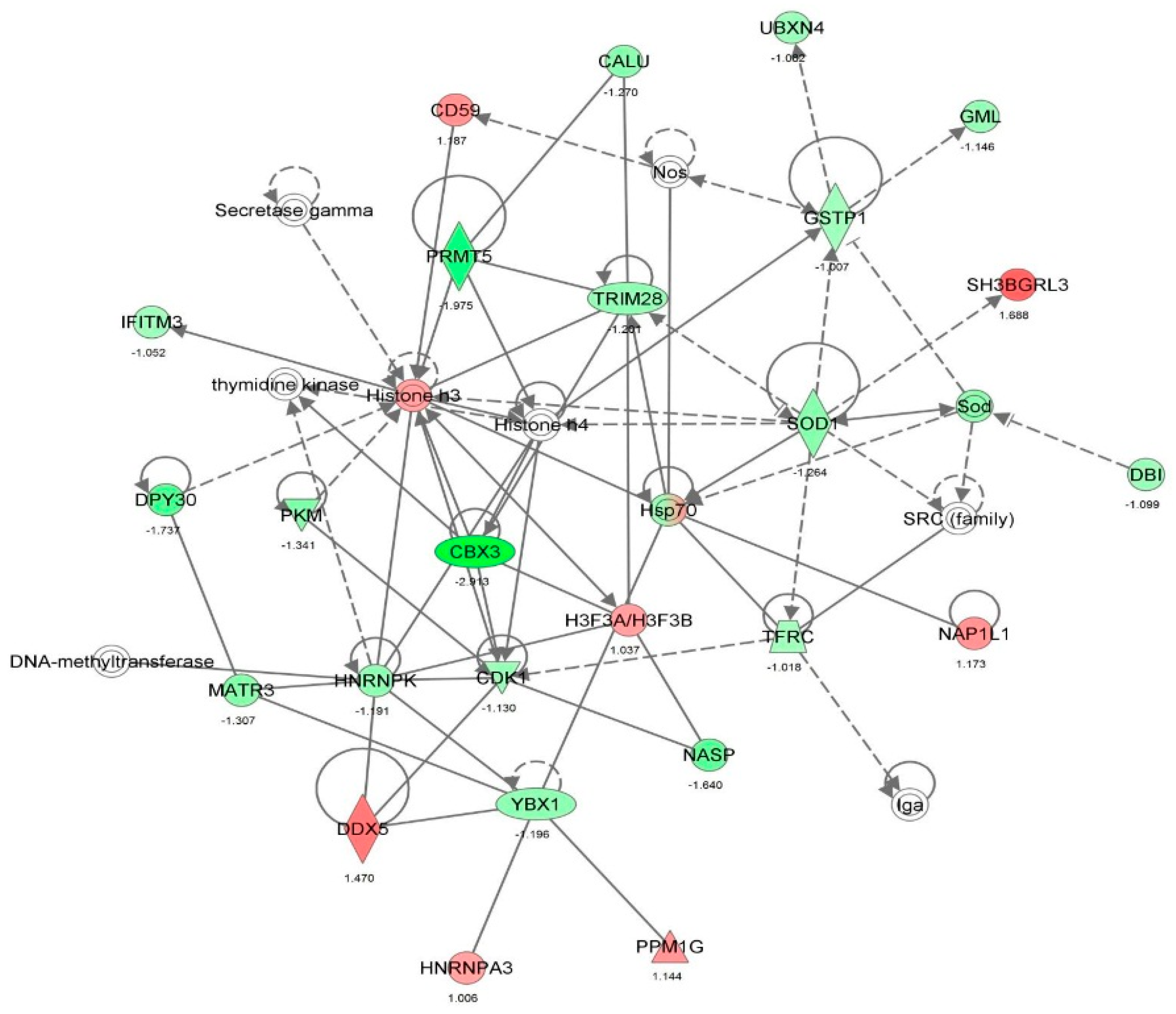
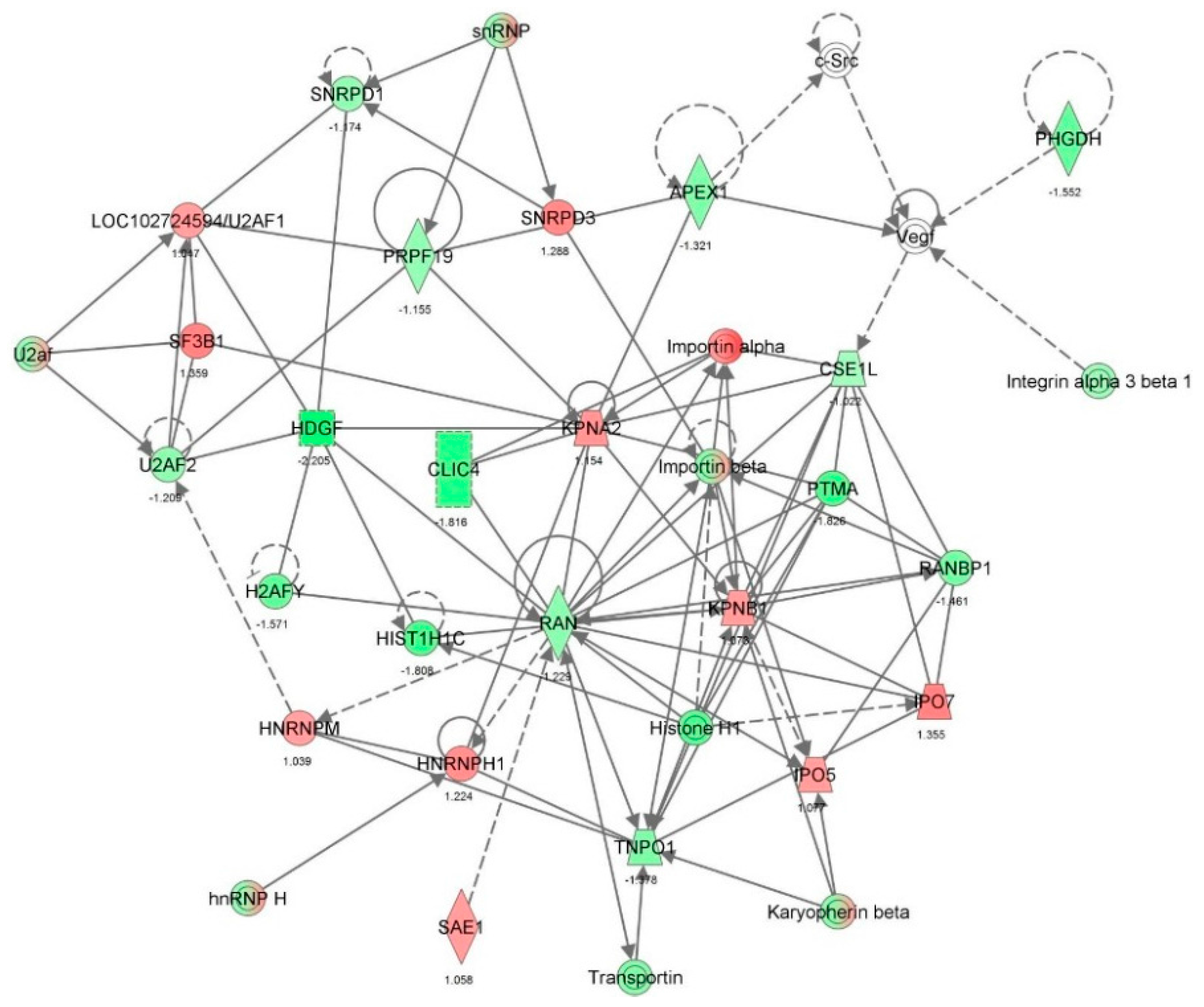
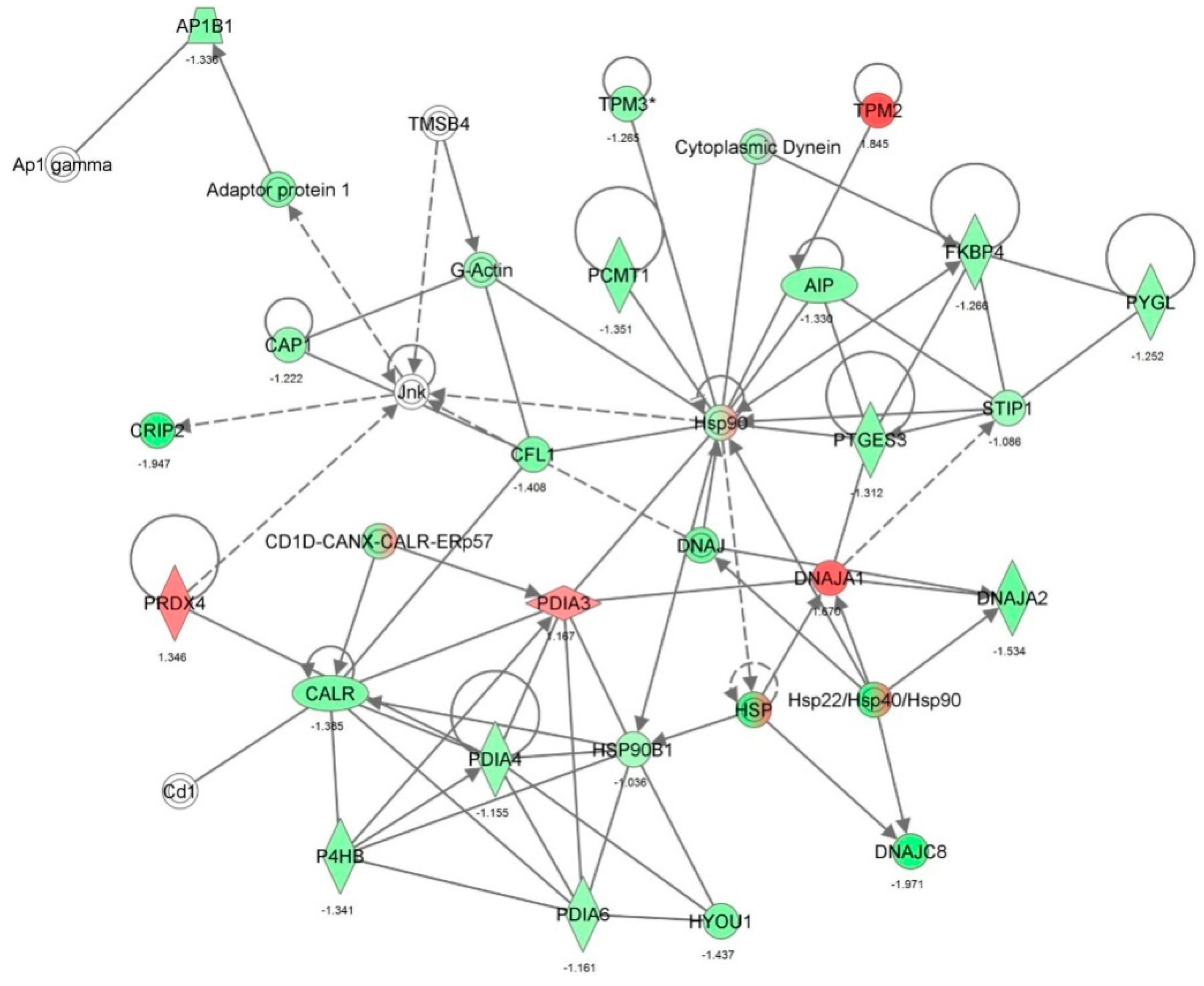

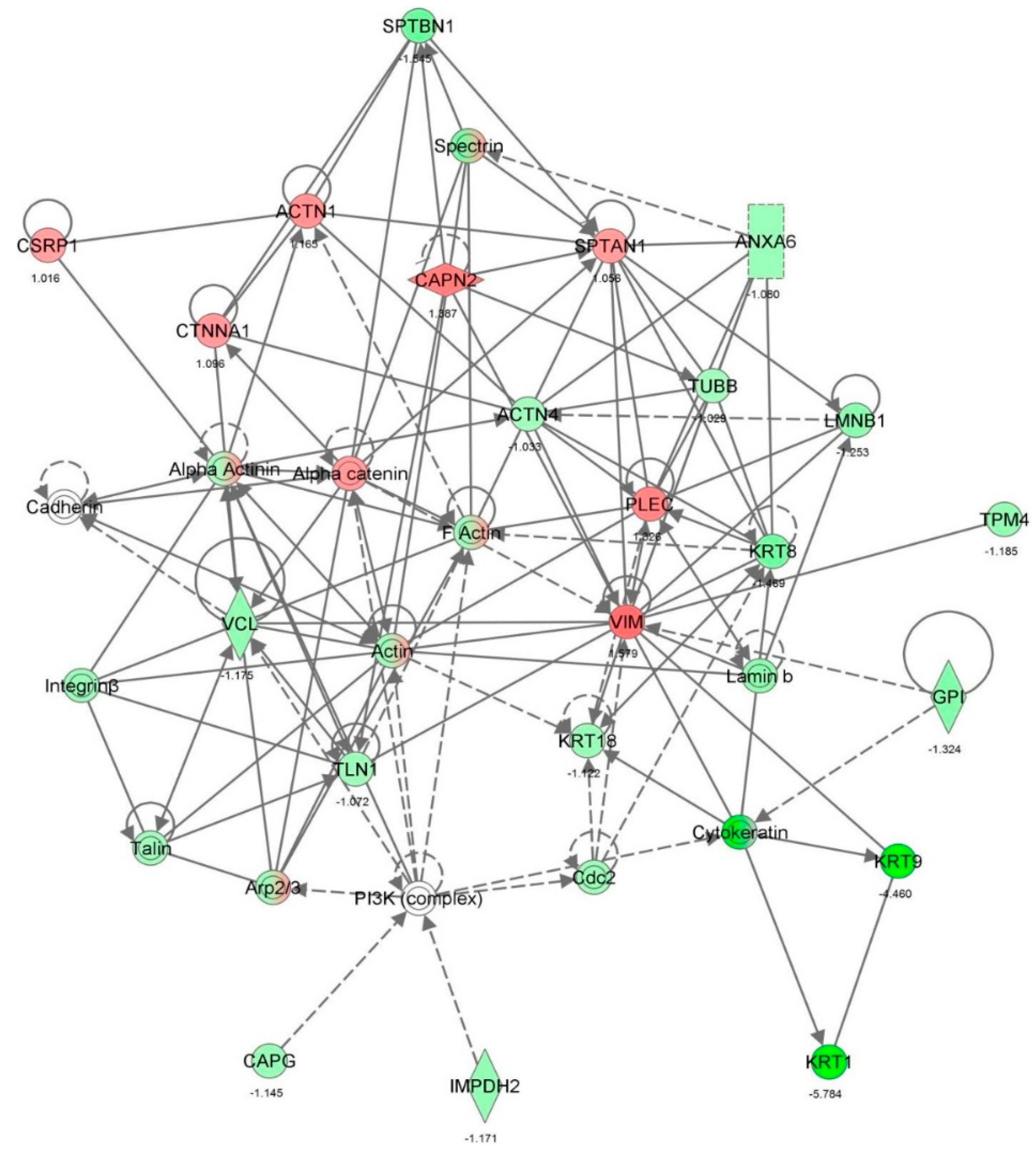
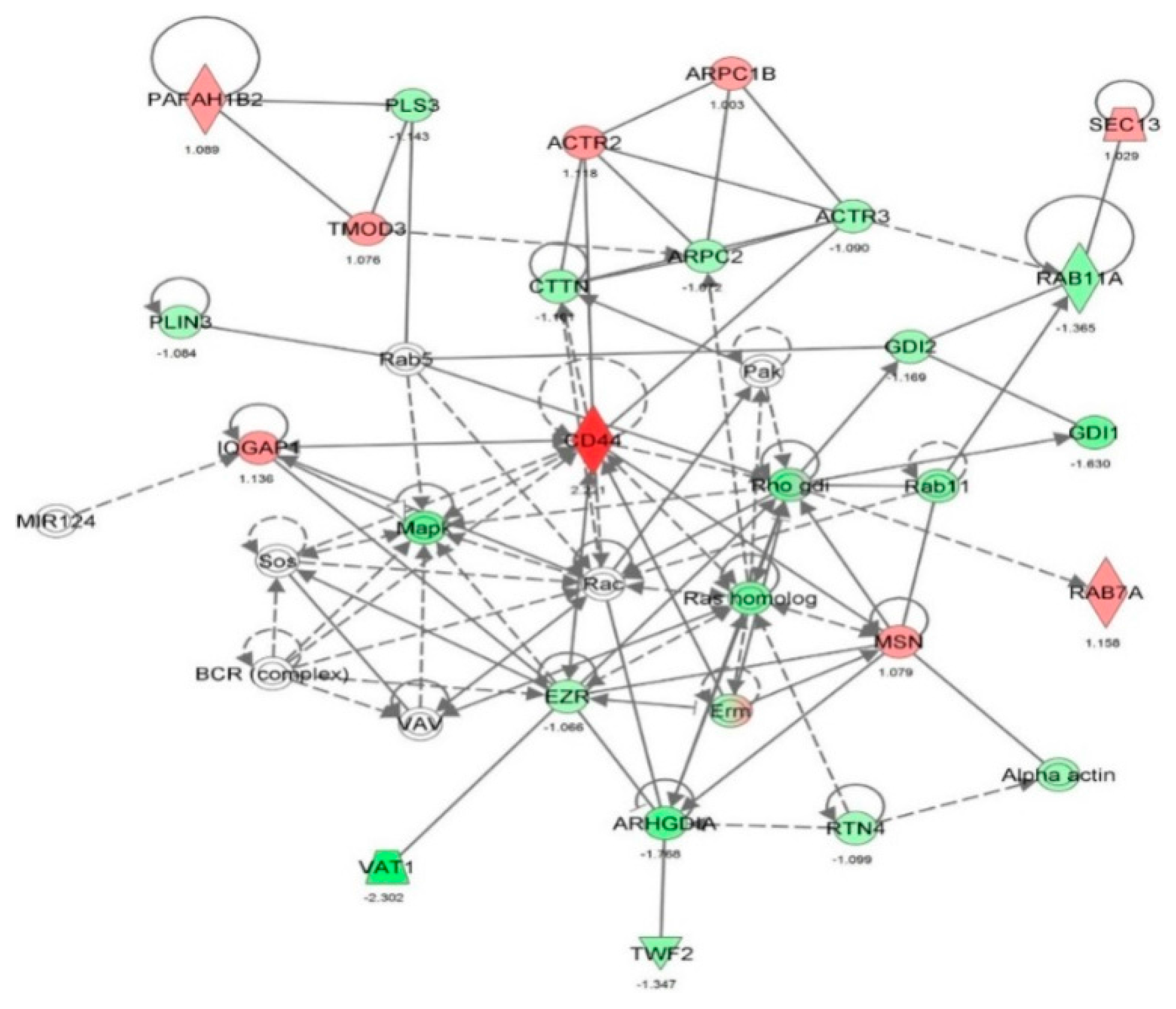
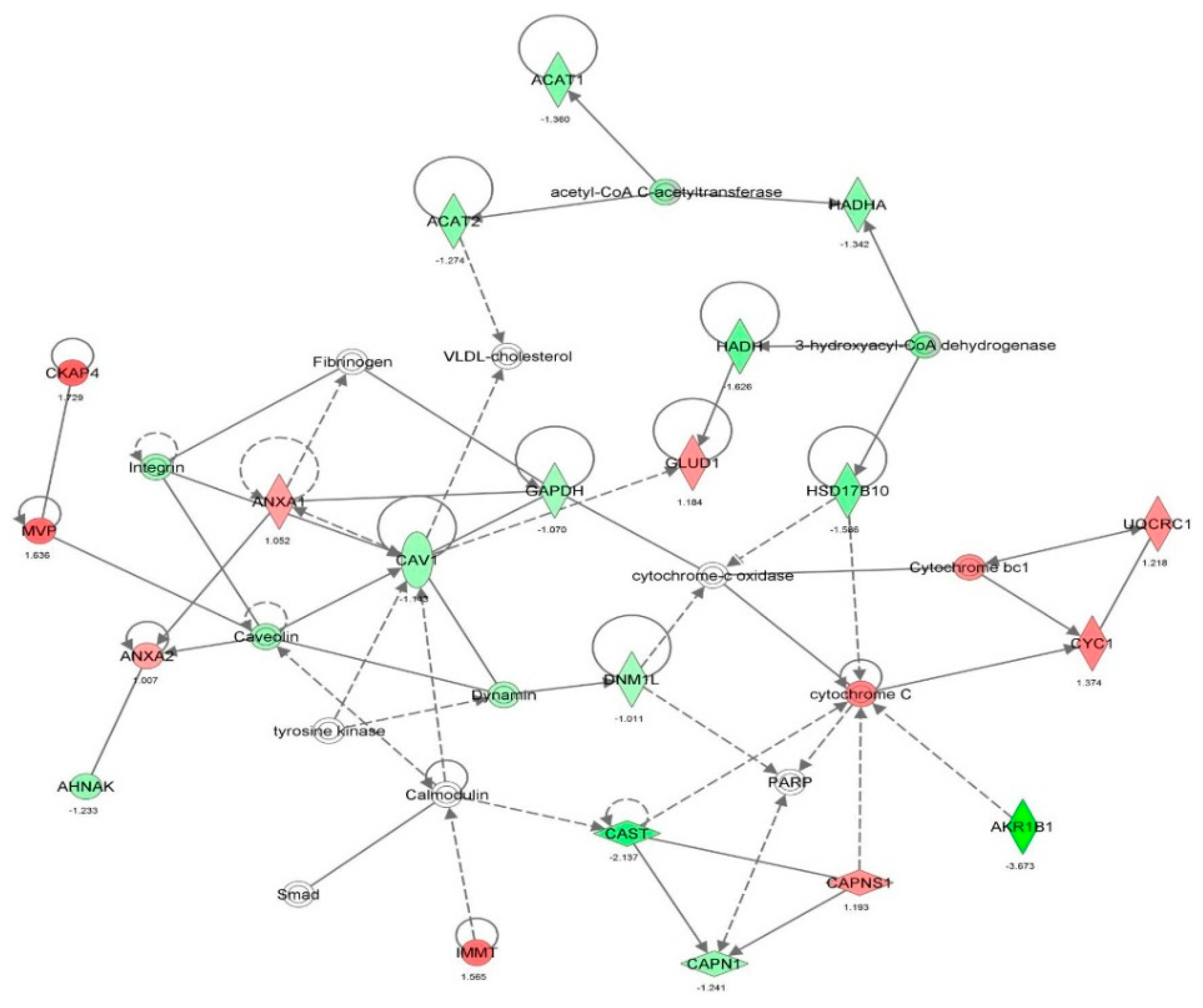
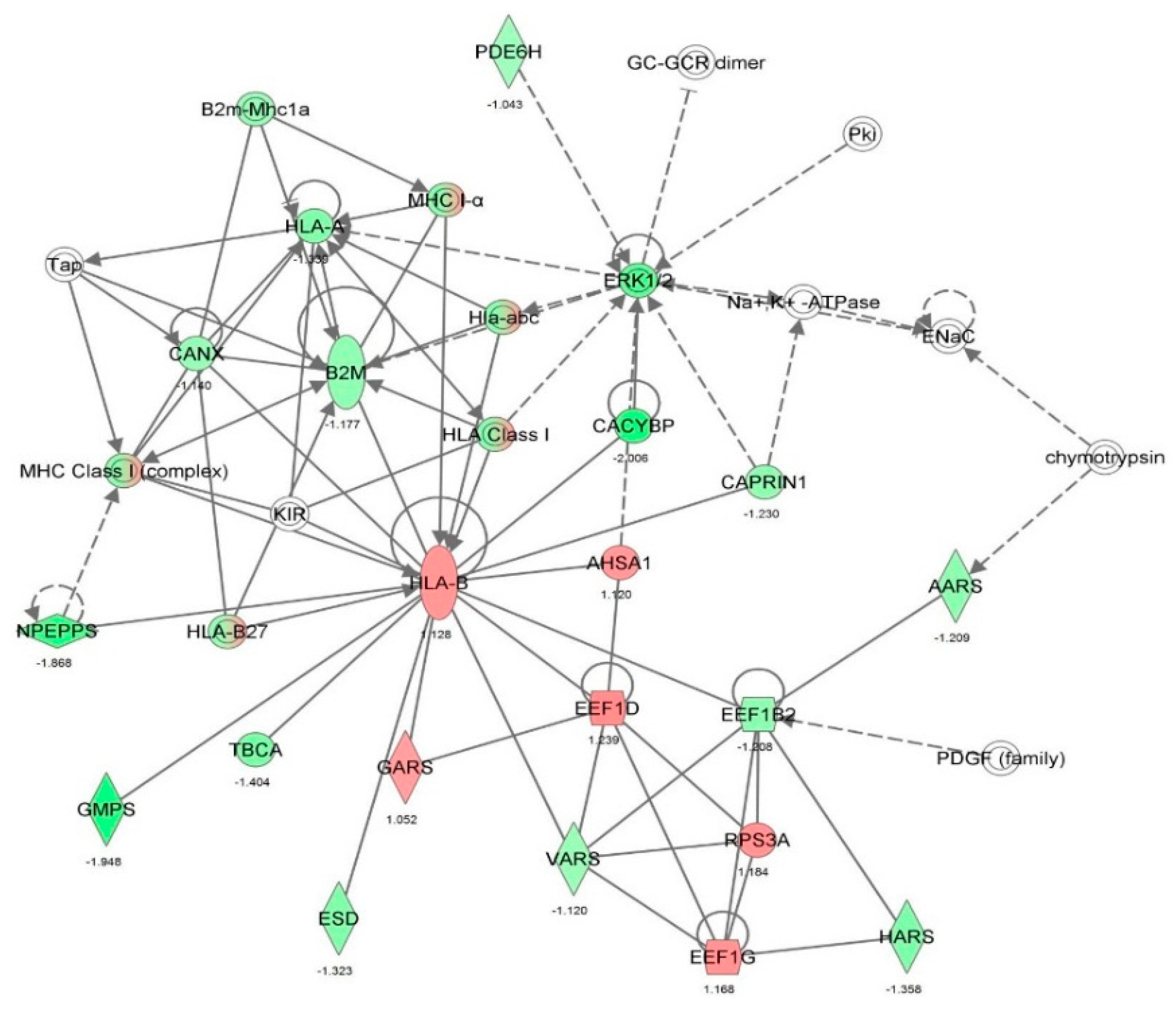

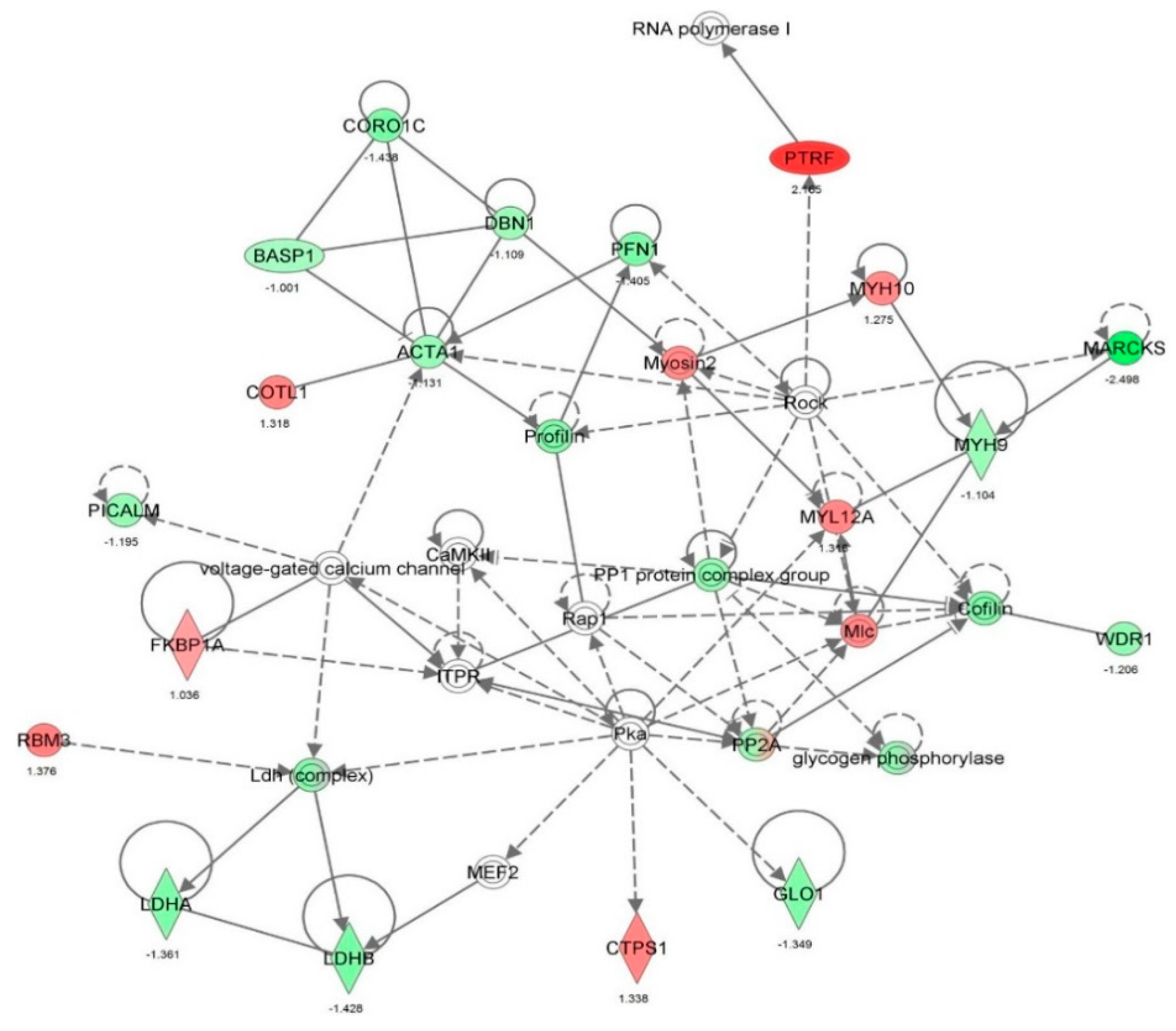
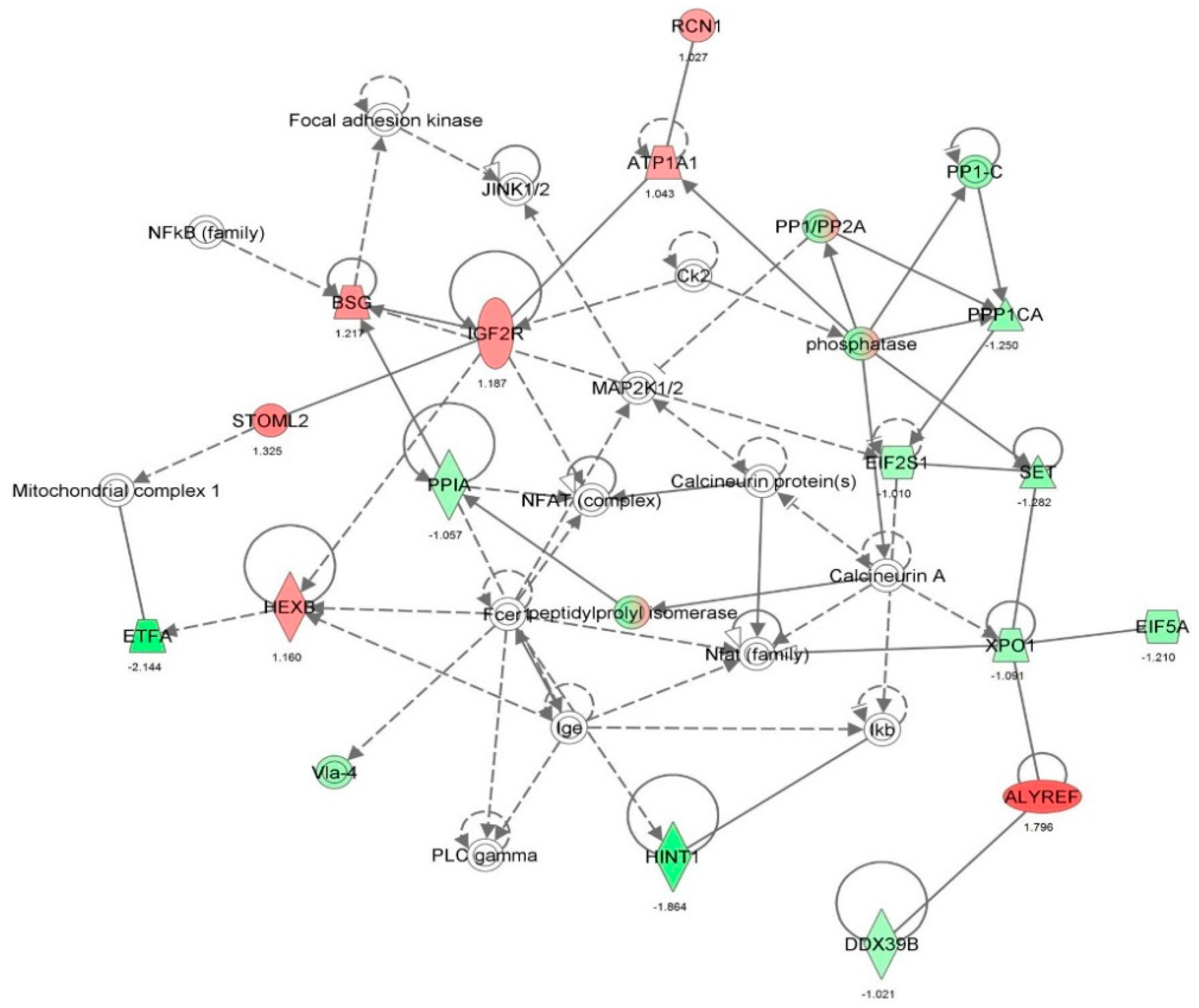
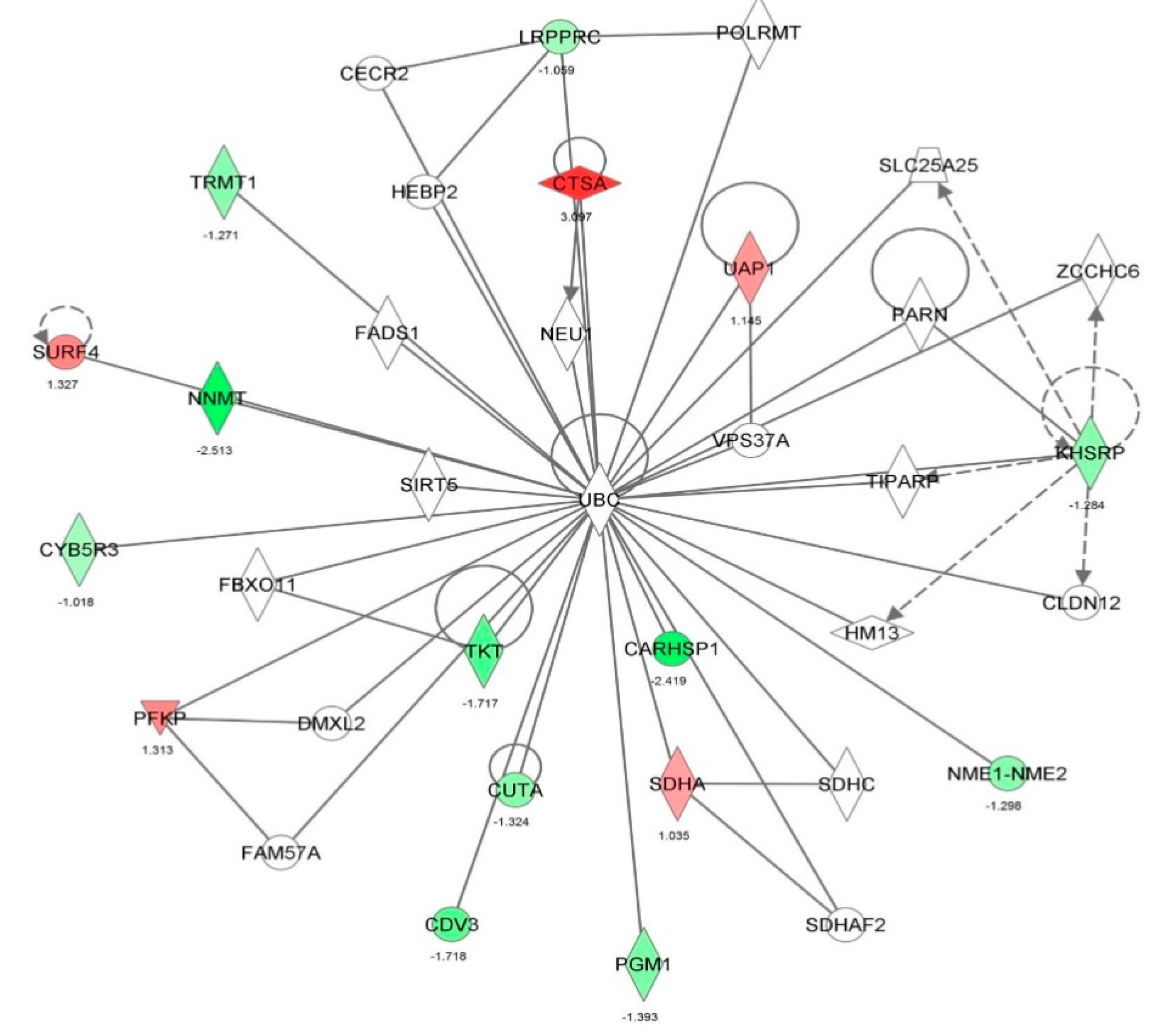
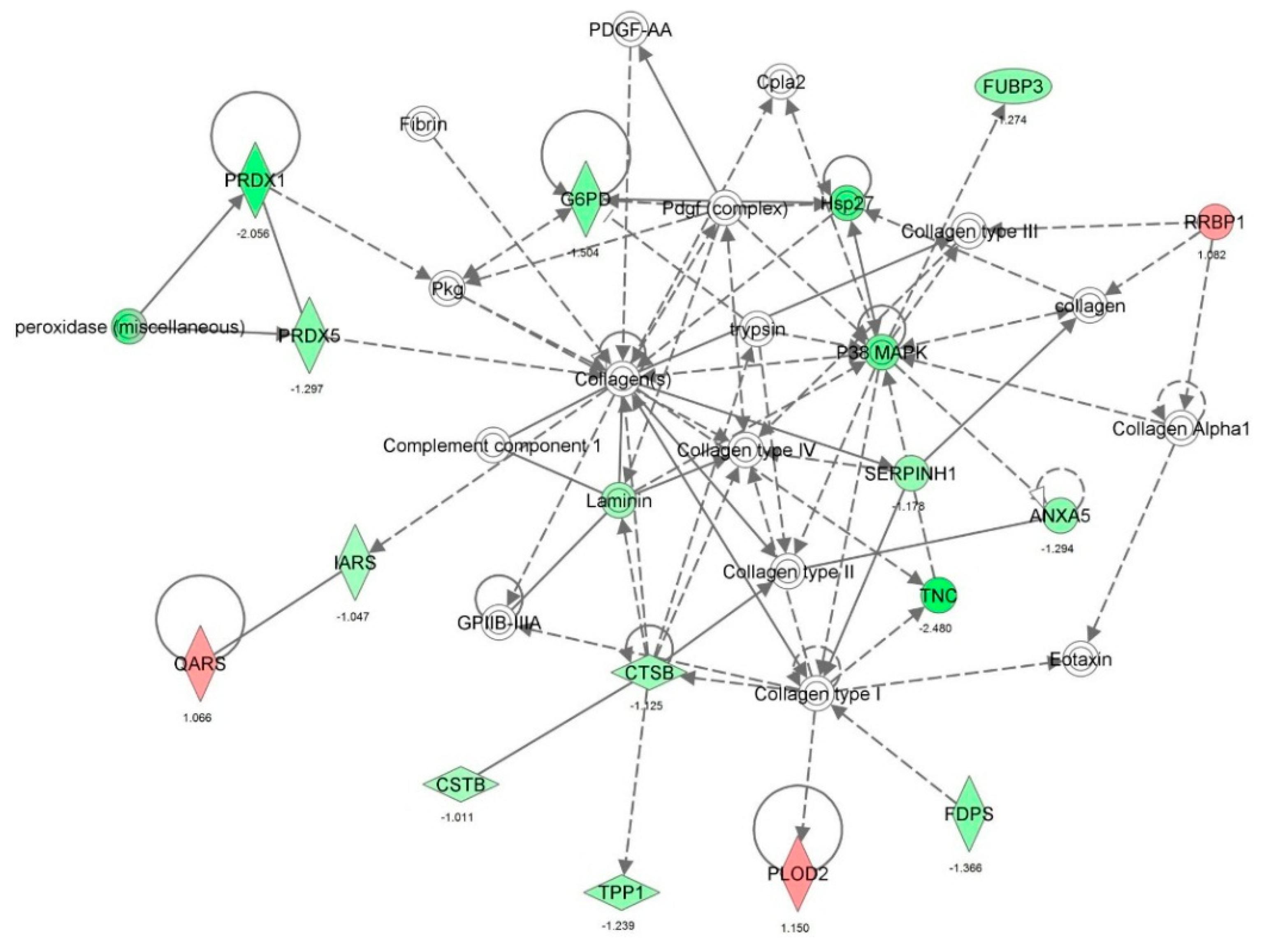

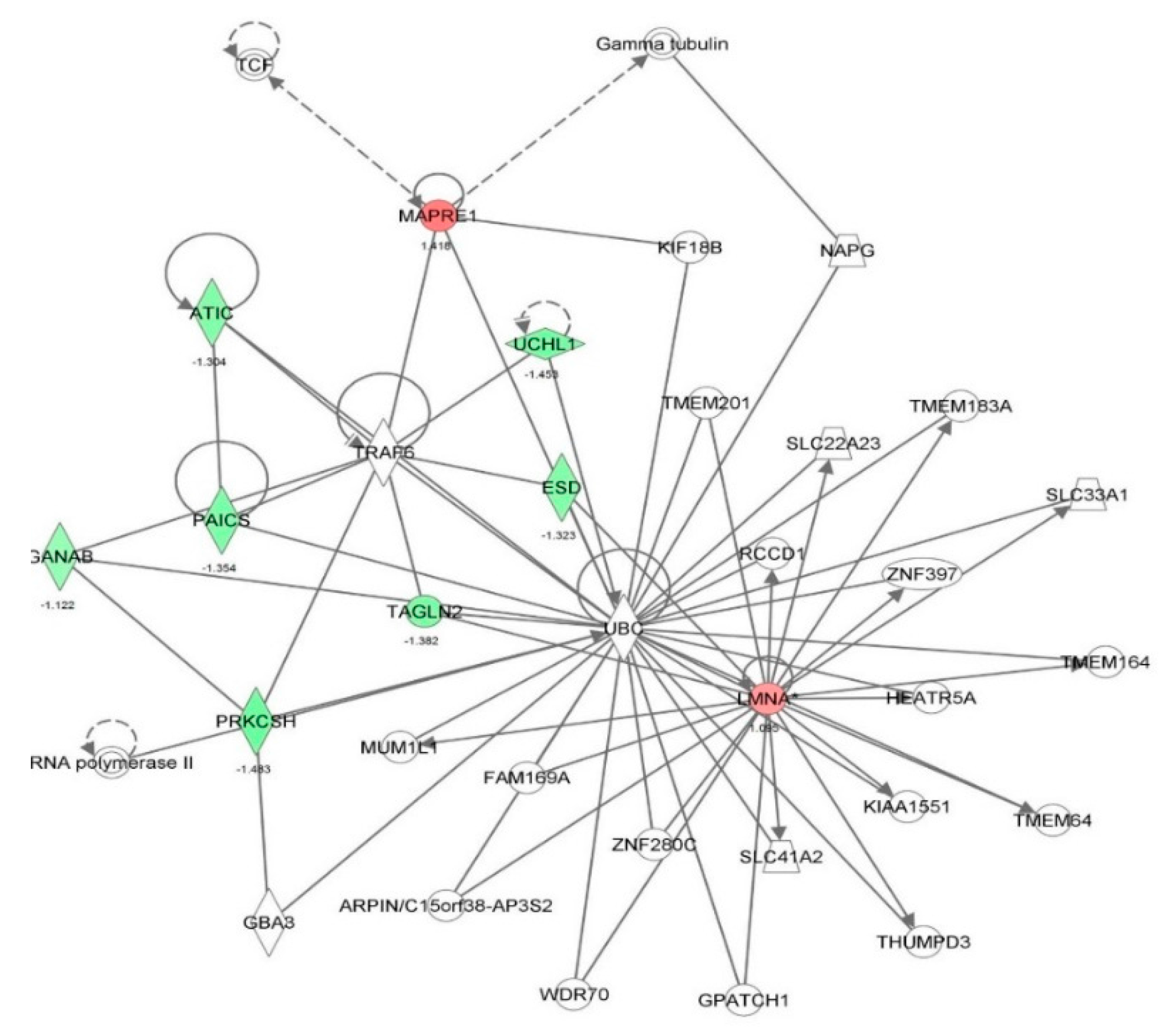
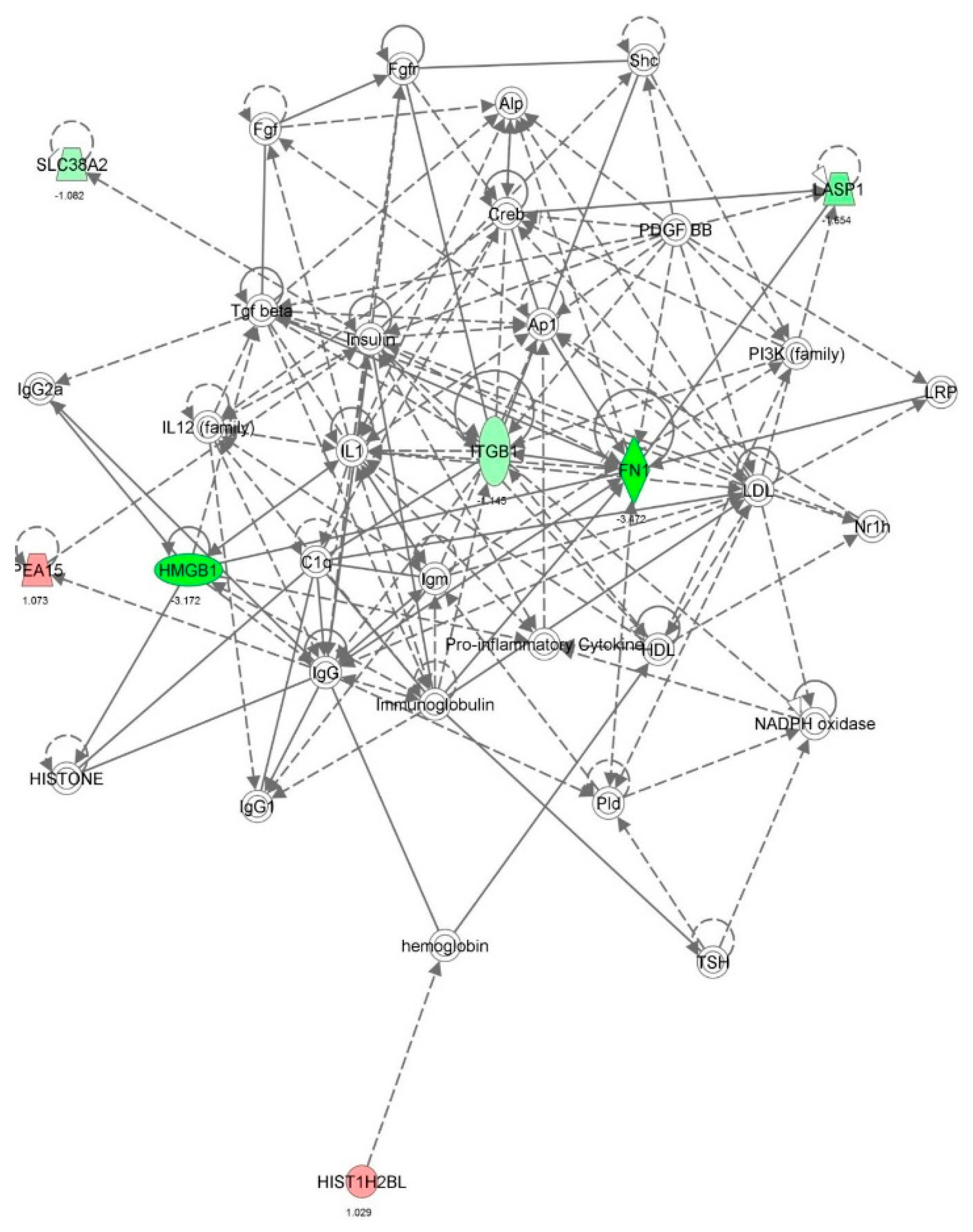
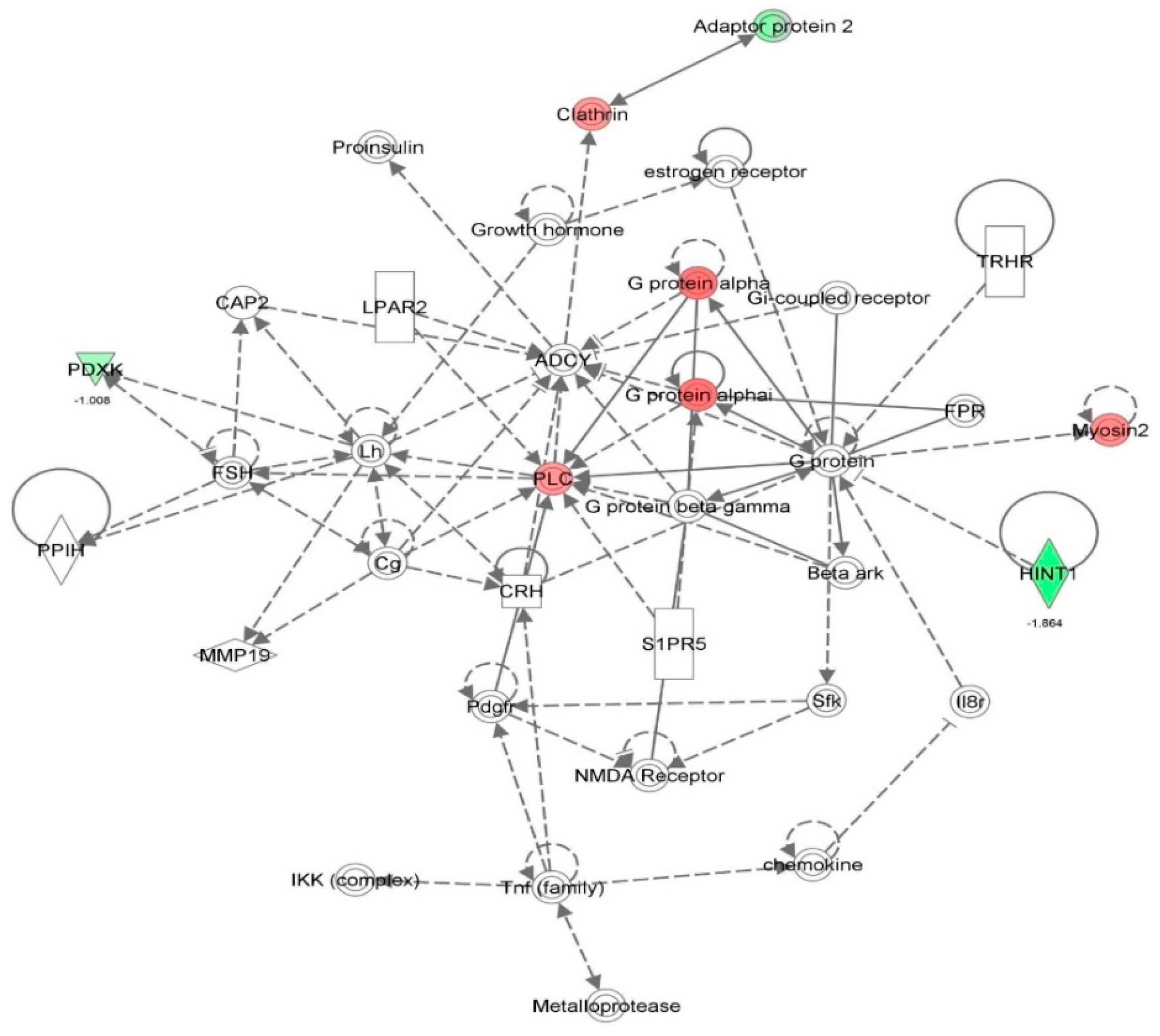
2.3. Vancomycin Induces Toxicity in HK-2 Cells
2.4. Vancomycin Regulates G2/M DNA Damage Check Point Signaling Pathway in HK-2 Cells
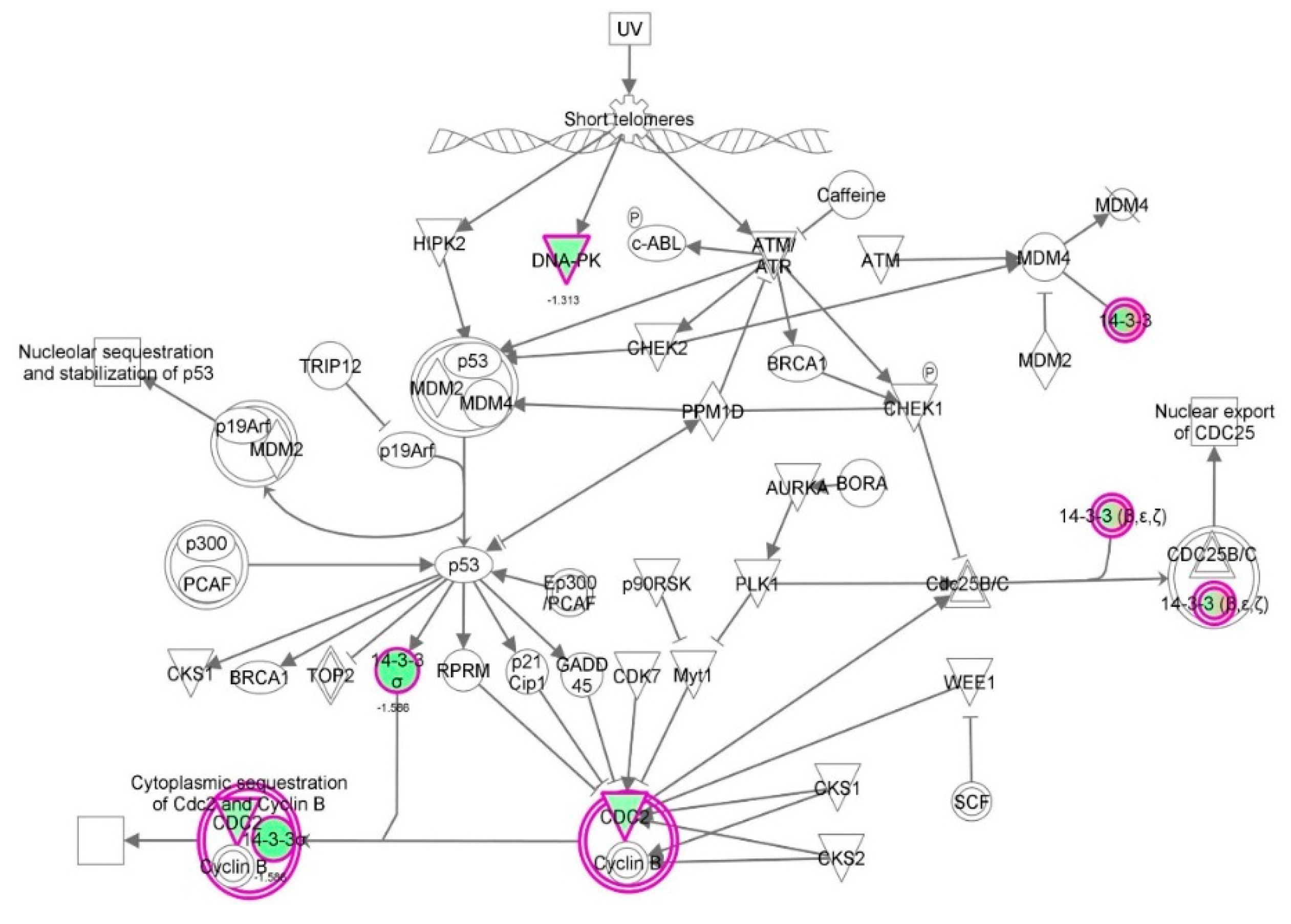
2.5. Vancomycin Regulates Apoptosis and Autophagy in HK-2 Cells
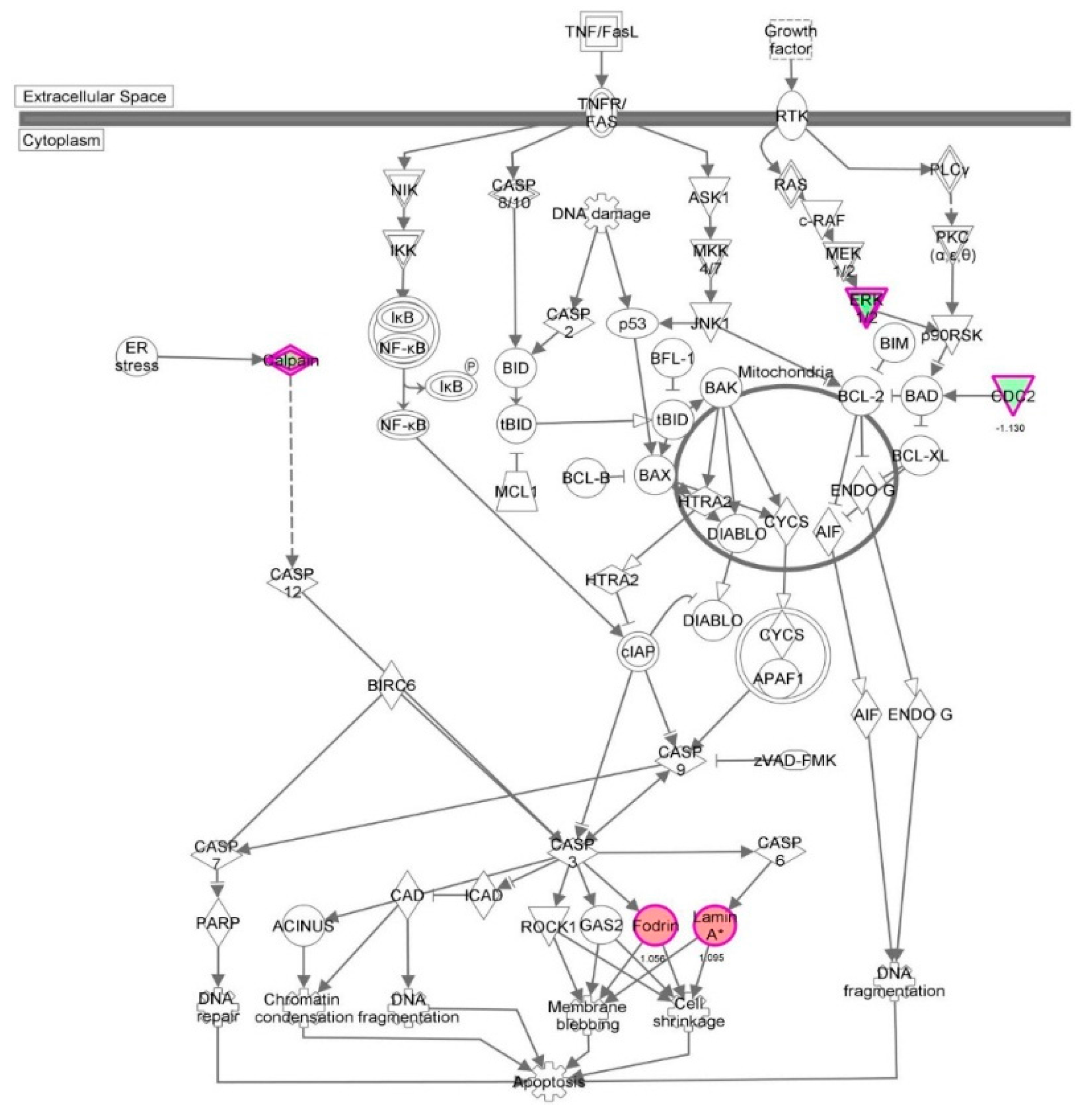
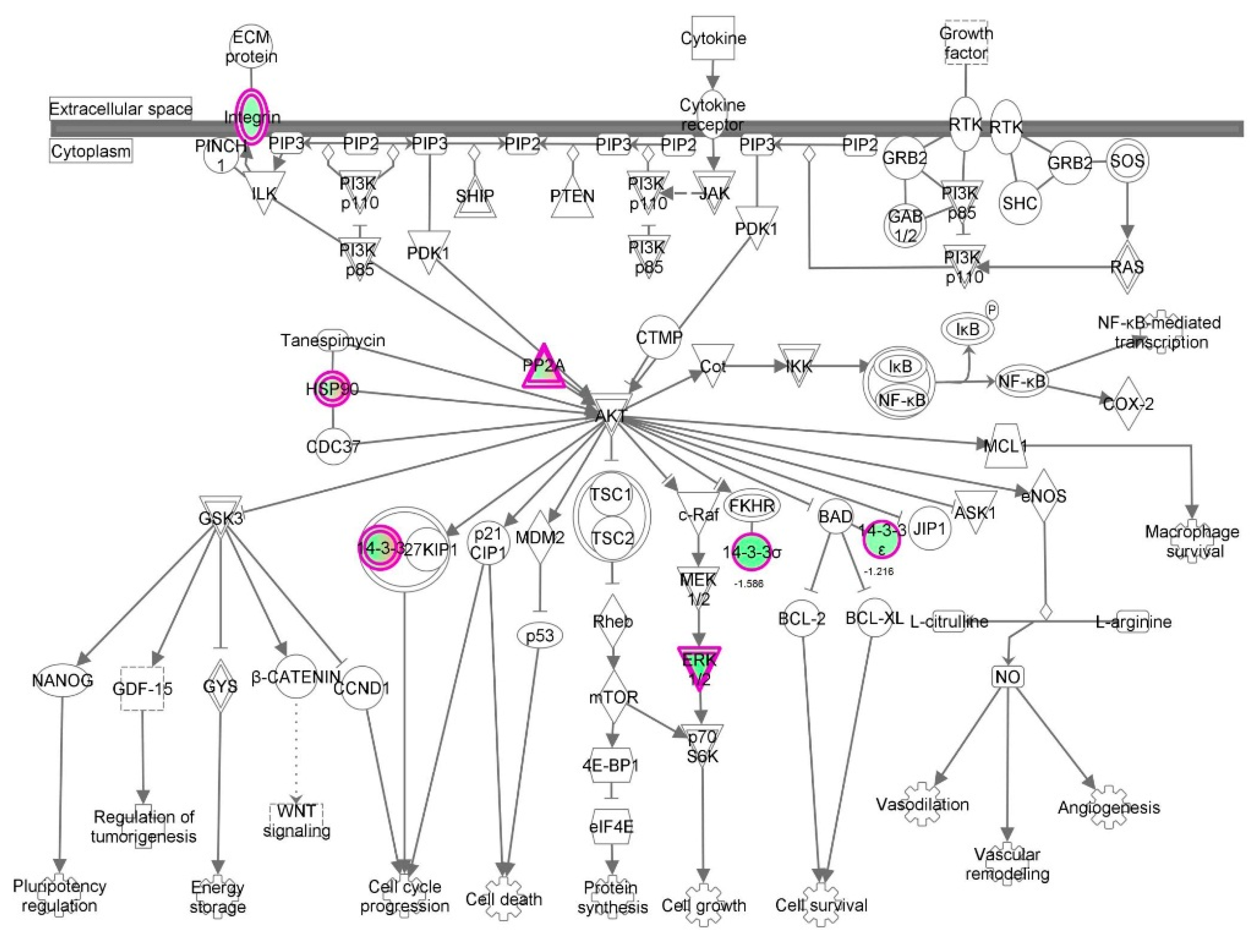


2.6. Vancomycin Regulates EMT Pathways in HK-2 cells
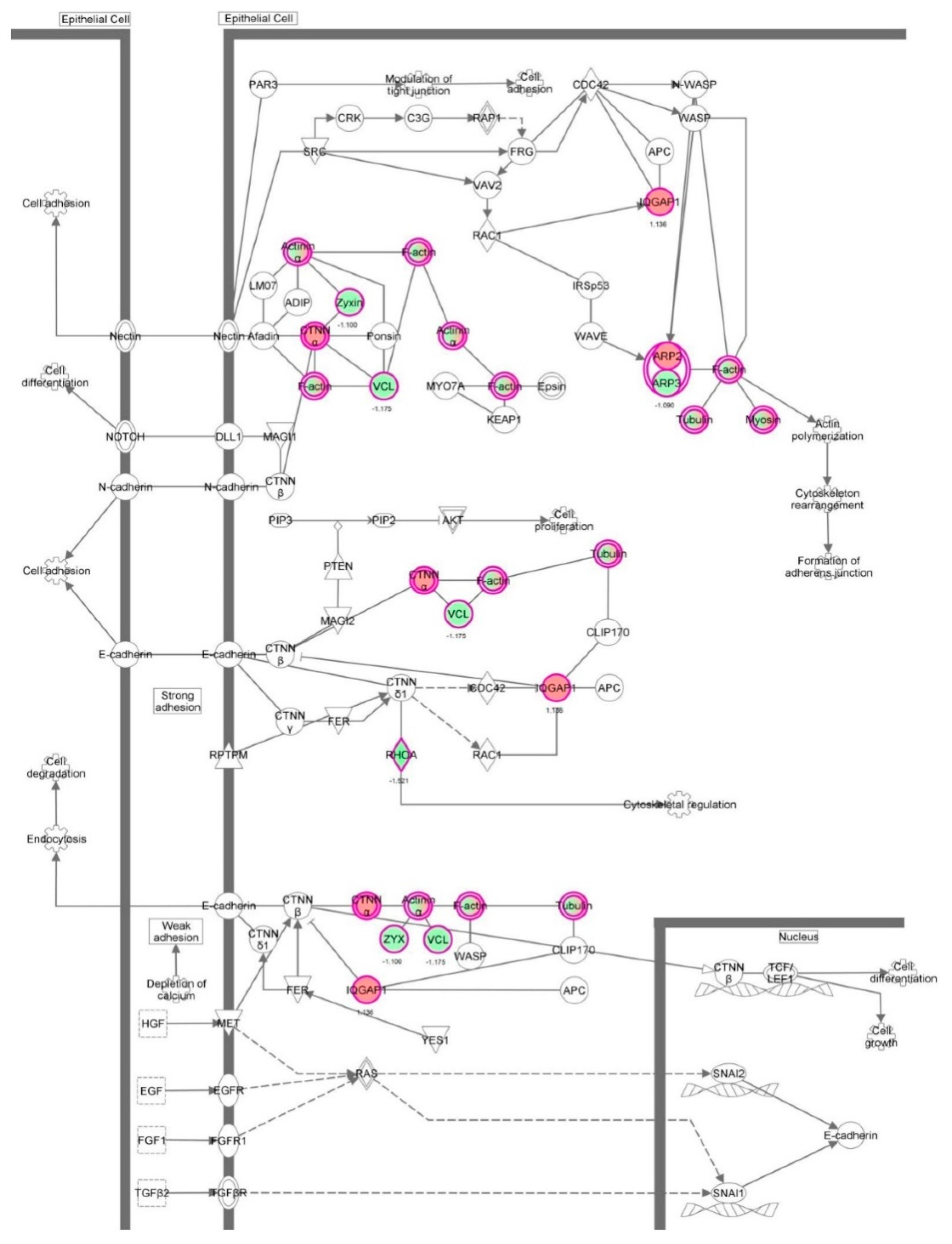
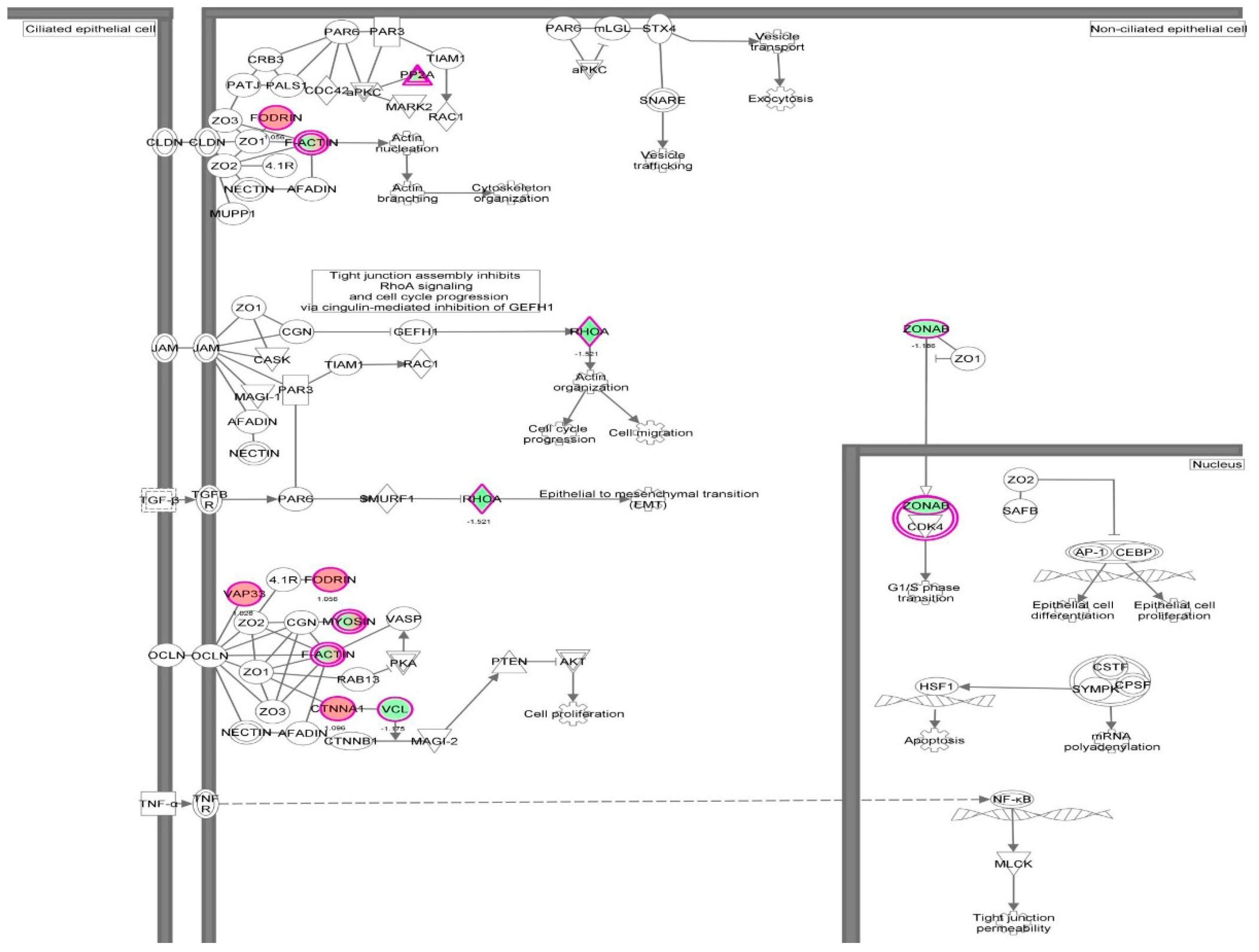
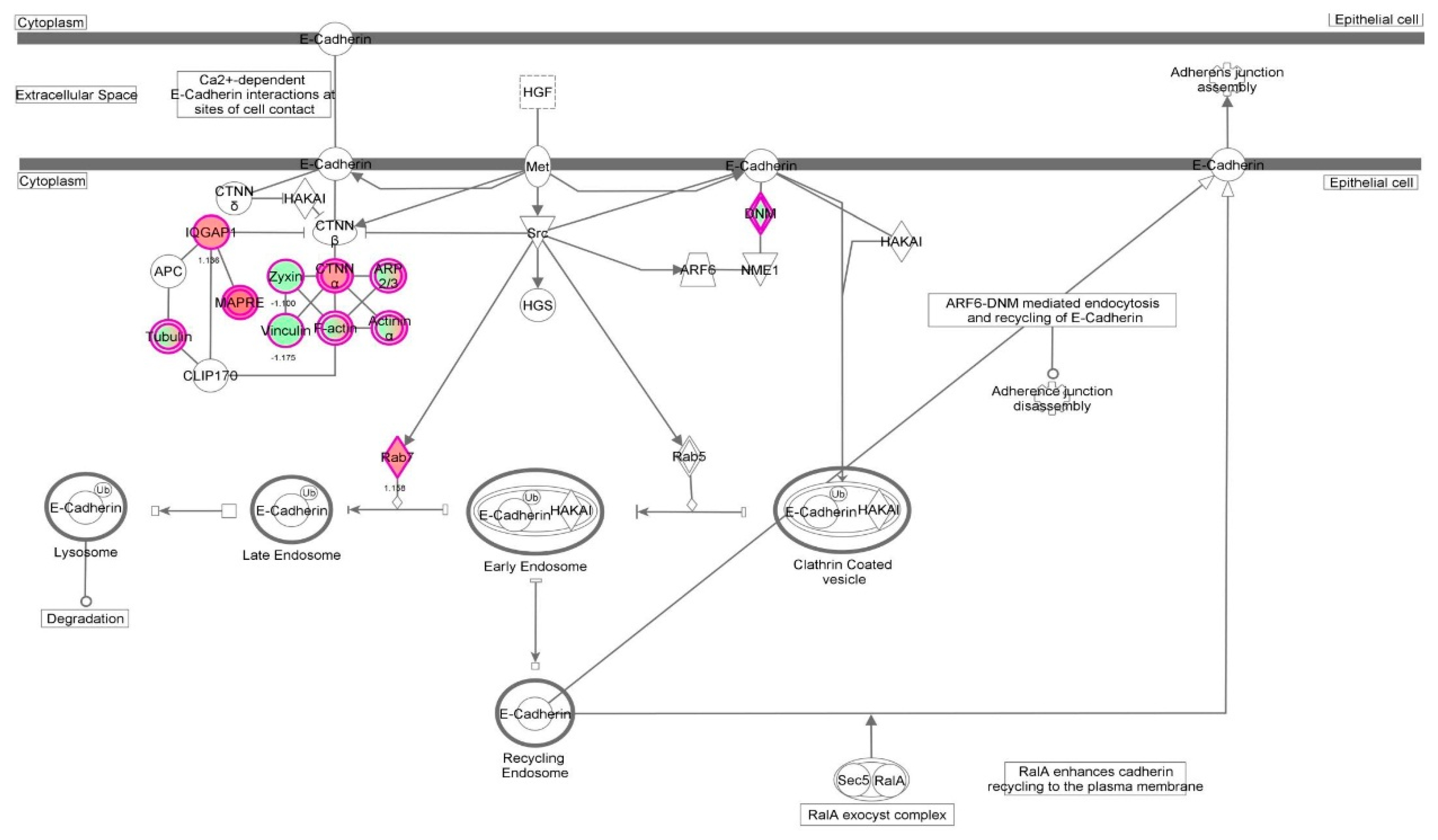
2.7. Vancomycin Regulates ER Stress Pathways in HK-2 cells
2.8. Vancomycin Regulates ERK-MAPK Signaling Pathway in HK-2 Cells
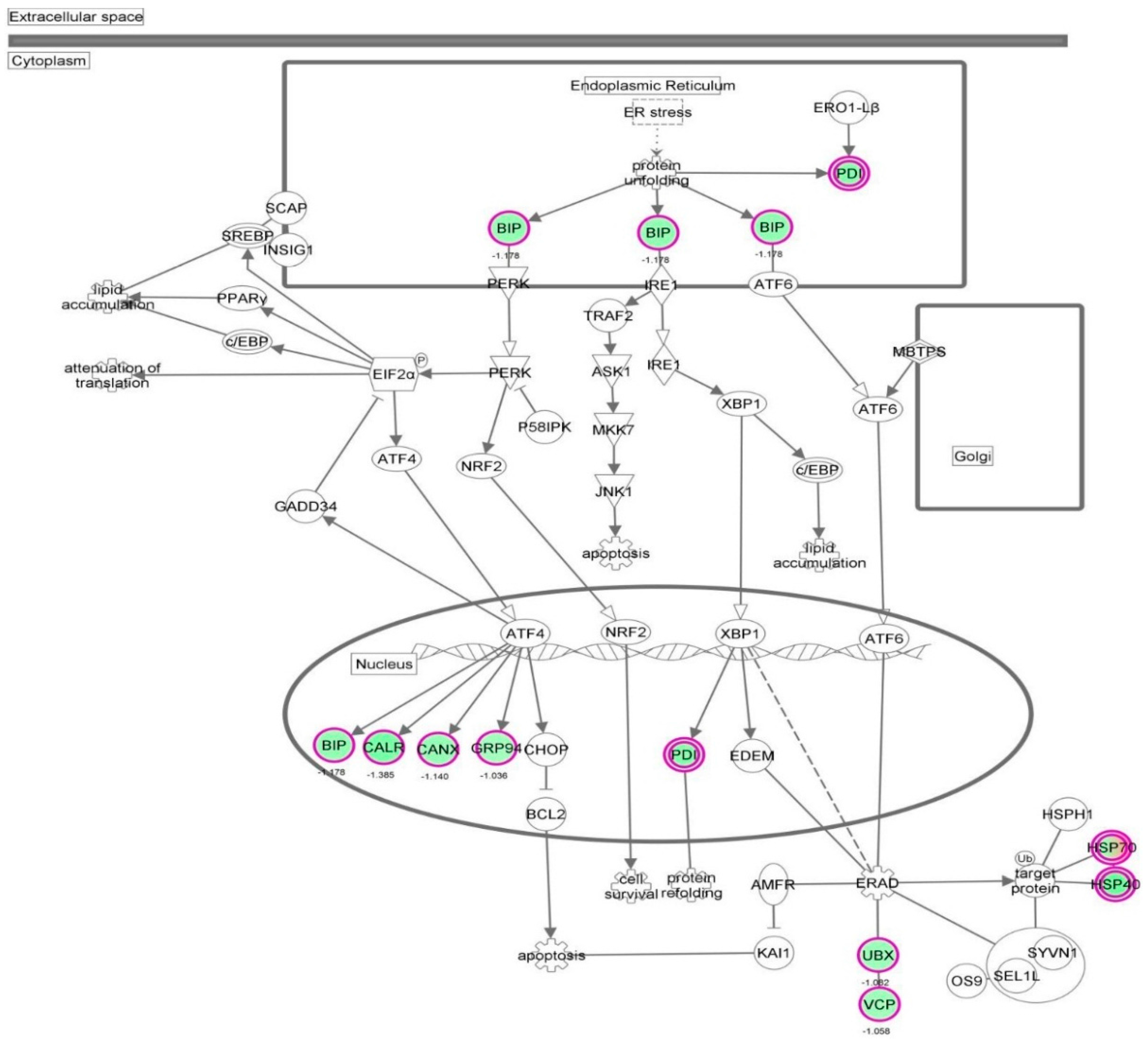
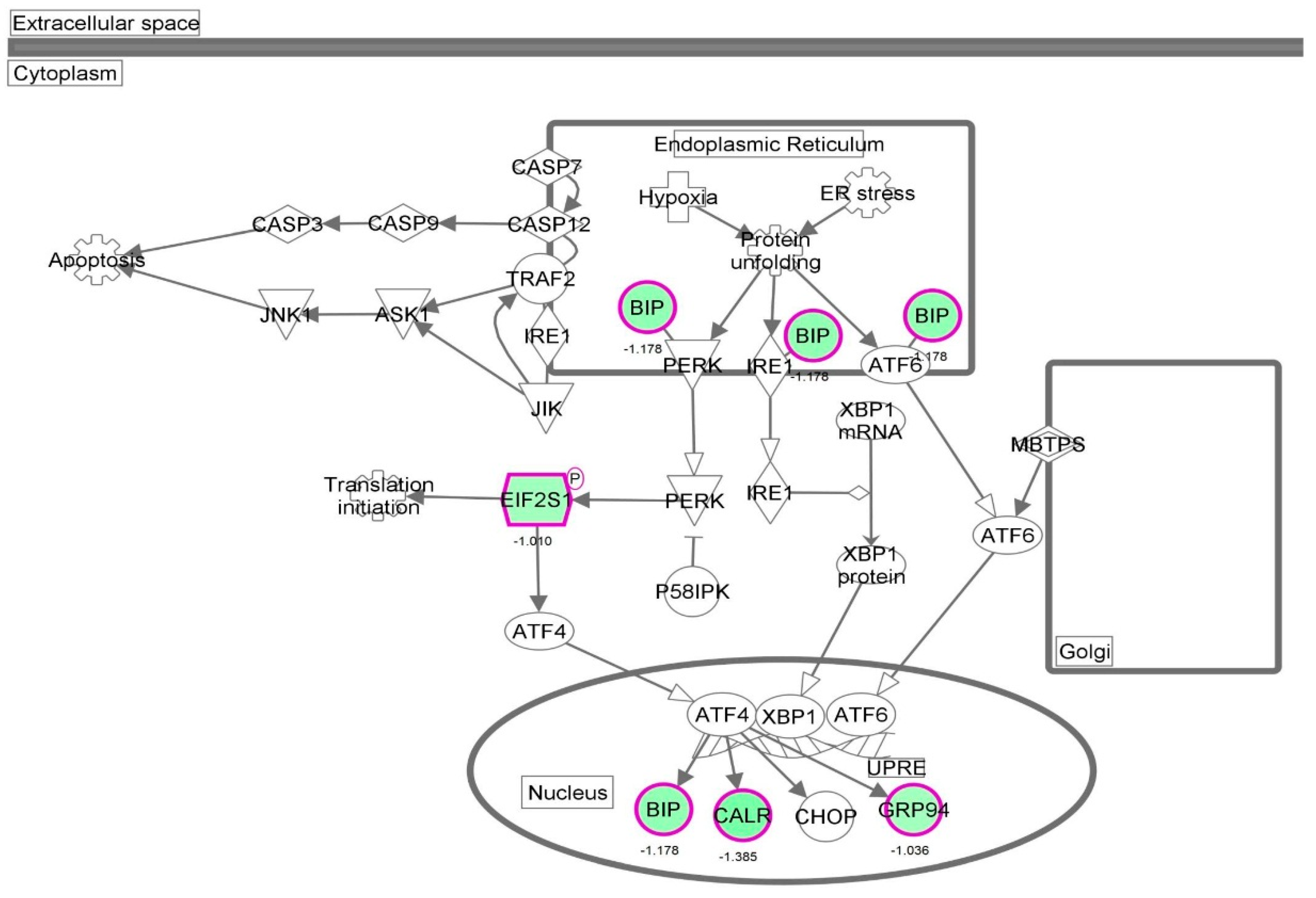

3. Discussion
4. Materials and Methods
4.1. Chemicals and Reagents
4.2. Cell Culture and Treatment
4.3. Digestion and Desalting SILAC Protein Samples
4.4. Liquid Chromatography-Tandem Mass Spectrometry (LC-MS/MS) and Statistical Analysis
4.5. Pathway Analysis and Bioinformatics
4.6. Statistical Analysis
5. Conclusions
Acknowledgments
Author contributions
Conflicts of Interest
References
- Camacho-Gonzalez, A.; Spearman, P.W.; Stoll, B.J. Neonatal infectious diseases: Evaluation of neonatal sepsis. Pediatr. Clin. N. Am. 2013, 60, 367–389. [Google Scholar] [CrossRef] [PubMed]
- Vandecasteele, S.J.; de Vriese, A.S. Recent changes in vancomycin use in renal failure. Kidney Int. 2010, 77, 760–764. [Google Scholar] [CrossRef] [PubMed]
- Pacifici, G.M.; Allegaert, K. Clinical pharmacokinetics of vancomycin in the neonate: A review. Clinics 2012, 67, 831–837. [Google Scholar] [CrossRef]
- Rostas, S.E.; Kubiak, D.W.; Calderwood, M.S. High-dose intravenous vancomycin therapy and the risk of nephrotoxicity. Clin. Ther. 2014, 36, 1098–1101. [Google Scholar] [CrossRef] [PubMed]
- Mergenhagen, K.A.; Borton, A.R. Vancomycin Nephrotoxicity: A Review. J. Pharm. Pract. 2014. [Google Scholar] [CrossRef] [PubMed]
- Elyasi, S.; Khalili, H.; Dashti-Khavidaki, S.; Mohammadpour, A. Vancomycin-induced nephrotoxicity: Mechanism, incidence, risk factors and special populations. A literature review. Eur. J. Clin. Pharmacol. 2012, 68, 1243–1255. [Google Scholar] [CrossRef] [PubMed]
- Moh’d, H.; Kheir, F.; Kong, L.; Du, P.; Farag, H.; Mohamad, A.; Zurlo, J.J. Incidence and predictors of vancomycin-associated nephrotoxicity. South. Med. J. 2014, 107, 383–388. [Google Scholar] [CrossRef] [PubMed]
- Ong, S.E.; Mann, M. Stable isotope labeling by amino acids in cell culture for quantitative proteomics. Methods Mol. Biol. 2007, 359, 37–52. [Google Scholar] [PubMed]
- Gruhler, S.; Kratchmarova, I. Stable isotope labeling by amino acids in cell culture (SILAC). Methods Mol. Biol. 2008, 424, 101–111. [Google Scholar] [PubMed]
- Mann, M. Functional and quantitative proteomics using SILAC. Nat. Rev. Mol. Cell Biol. 2006, 7, 952–968. [Google Scholar] [CrossRef] [PubMed]
- Ong, S.E. The expanding field of SILAC. Anal. Bioanal. Chem. 2012, 404, 967–976. [Google Scholar] [CrossRef] [PubMed]
- Shanware, N.P.; Bray, K.; Abraham, R.T. The PI3K, metabolic, and autophagy networks: Interactive partners in cellular health and disease. Annu. Rev. Pharmacol. Toxicol. 2013, 53, 89–106. [Google Scholar] [CrossRef] [PubMed]
- Morgensztern, D.; McLeod, H.L. PI3K/Akt/mTOR pathway as a target for cancer therapy. Anticancer Drugs 2005, 16, 797–803. [Google Scholar] [CrossRef] [PubMed]
- Wu, W.K.; Coffelt, S.B.; Cho, C.H.; Wang, X.J.; Lee, C.W.; Chan, F.K.; Yu, J.; Sung, J.J. The autophagic paradox in cancer therapy. Oncogene 2012, 31, 939–953. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, C.G.; Epping, M.; Kruyt, F.A.; Giaccone, G. Apoptosis: Target of cancer therapy. Clin. Cancer Res. 2002, 8, 2024–2034. [Google Scholar] [PubMed]
- Fulda, S.; Galluzzi, L.; Kroemer, G. Targeting mitochondria for cancer therapy. Nat. Rev. Drug Discov. 2010, 9, 447–464. [Google Scholar] [CrossRef] [PubMed]
- Laplante, M.; Sabatini, D.M. mTOR signaling in growth control and disease. Cell 2012, 149, 274–293. [Google Scholar] [CrossRef] [PubMed]
- Dancey, J. mTOR signaling and drug development in cancer. Nat. Rev. Clin. Oncol. 2010, 7, 209–219. [Google Scholar] [CrossRef] [PubMed]
- Lamouille, S.; Xu, J.; Derynck, R. Molecular mechanisms of epithelial-mesenchymal transition. Nat. Rev. Mol. Cell Biol. 2014, 15, 178–196. [Google Scholar] [CrossRef] [PubMed]
- Schroder, M.; Kaufman, R.J. ER stress and the unfolded protein response. Mutat. Res. 2005, 569, 29–63. [Google Scholar] [CrossRef] [PubMed]
- Hetz, C. The unfolded protein response: Controlling cell fate decisions under ER stress and beyond. Nat. Rev. Mol. Cell Biol. 2012, 13, 89–102. [Google Scholar] [CrossRef] [PubMed]
- Boehning, D.; Patterson, R.L.; Sedaghat, L.; Glebova, N.O.; Kurosaki, T.; Snyder, S.H. Cytochrome c binds to inositol (1,4,5) trisphosphate receptors, amplifying calcium-dependent apoptosis. Nat. Cell Biol. 2003, 5, 1051–1061. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Liu, X.; Bhalla, K.; Kim, C.N.; Ibrado, A.M.; Cai, J.; Peng, T.I.; Jones, D.P.; Wang, X. Prevention of apoptosis by Bcl-2: Release of cytochrome c from mitochondria blocked. Science 1997, 275, 1129–1132. [Google Scholar] [CrossRef] [PubMed]
- Jeffers, J.R.; Parganas, E.; Lee, Y.; Yang, C.; Wang, J.; Brennan, J.; MacLean, K.H.; Han, J.; Chittenden, T.; Ihle, J.N.; et al. Puma is an essential mediator of p53-dependent and -independent apoptotic pathways. Cancer Cell 2003, 4, 321–328. [Google Scholar] [CrossRef]
- Fesik, S.W.; Shi, Y. Structural biology. Controlling the caspases. Science 2001, 294, 1477–1478. [Google Scholar] [PubMed]
- Klionsky, D.J.; Emr, S.D. Autophagy as a regulated pathway of cellular degradation. Science 2000, 290, 1717–1721. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Yu, L. Autophagic lysosome reformation. Exp. Cell Res. 2013, 319, 142–146. [Google Scholar] [CrossRef] [PubMed]
- Denton, D.; Nicolson, S.; Kumar, S. Cell death by autophagy: Facts and apparent artefacts. Cell Death Differ. 2012, 19, 87–95. [Google Scholar] [CrossRef] [PubMed]
- Zheng, H.; Kang, Y. Multilayer control of the EMT master regulators. Oncogene 2014, 33, 1755–1763. [Google Scholar] [CrossRef] [PubMed]
- Ong, S.E.; Mann, M. A practical recipe for stable isotope labeling by amino acids in cell culture (SILAC). Nat. Protoc. 2006, 1, 2650–2660. [Google Scholar] [CrossRef] [PubMed]
- Sample Availability: Samples of vancomycin-tretaed HK-2 cells are available from the authors.
© 2016 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, Z.-L.; Zhou, S.-F. A SILAC-Based Approach Elicits the Proteomic Responses to Vancomycin-Associated Nephrotoxicity in Human Proximal Tubule Epithelial HK-2 Cells. Molecules 2016, 21, 148. https://doi.org/10.3390/molecules21020148
Li Z-L, Zhou S-F. A SILAC-Based Approach Elicits the Proteomic Responses to Vancomycin-Associated Nephrotoxicity in Human Proximal Tubule Epithelial HK-2 Cells. Molecules. 2016; 21(2):148. https://doi.org/10.3390/molecules21020148
Chicago/Turabian StyleLi, Zhi-Ling, and Shu-Feng Zhou. 2016. "A SILAC-Based Approach Elicits the Proteomic Responses to Vancomycin-Associated Nephrotoxicity in Human Proximal Tubule Epithelial HK-2 Cells" Molecules 21, no. 2: 148. https://doi.org/10.3390/molecules21020148
APA StyleLi, Z.-L., & Zhou, S.-F. (2016). A SILAC-Based Approach Elicits the Proteomic Responses to Vancomycin-Associated Nephrotoxicity in Human Proximal Tubule Epithelial HK-2 Cells. Molecules, 21(2), 148. https://doi.org/10.3390/molecules21020148





