Cytotoxic Quinones from the Roots of Aloe dawei
Abstract
:1. Introduction
2. Results and Discussion
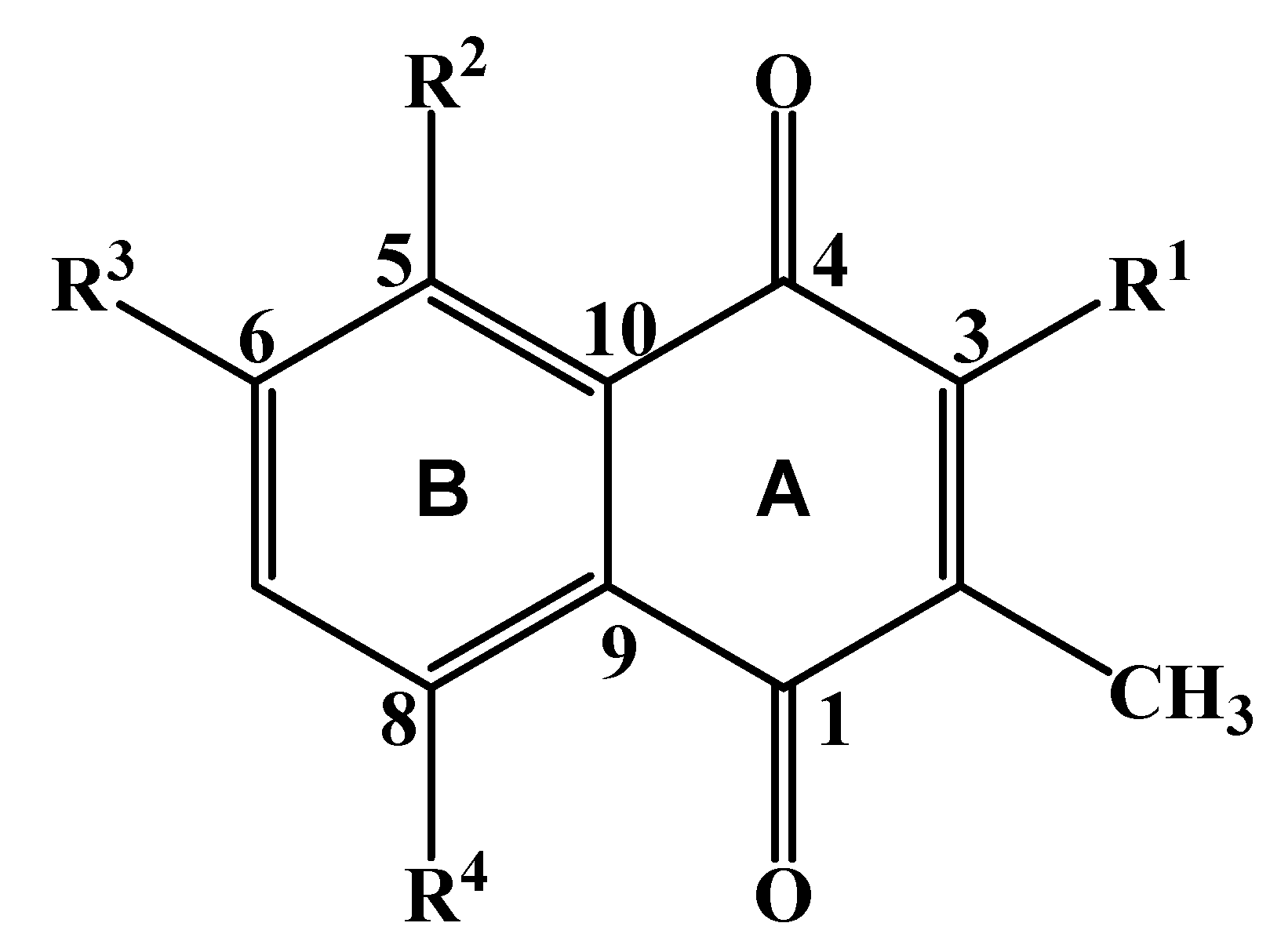
| δC | δH (I, m, J in Hz) | HMBC (2J, 3J) | |
|---|---|---|---|
| 1 | 187.2 | ||
| 2 | 131.5 | ||
| 3 | 161.4 | ||
| 4 | 183.0 | ||
| 5 | 150.2 | ||
| 6 | 160.0 | ||
| 7 | 123.6 | 7.19 (1H, d, 8.4) | C-6, C-5, C-9 |
| 8 | 126.5 | 7.65 (1H, d, 8.4) | C-1, C-10 |
| 9 | 127.4 | ||
| 10 | 127.9 | ||
| 2-CH3 | 12.0 | 1.91 (3H, s) | C-1, C-2, C-3 |
| 3-OCH3 | 63.5 | 3.95 (3H, s) | C-3 |
| 5-OCH3 | 63.6 | 3.77 (3H, s) | C-5 |
3. Experimental
3.1. General Information
| Isolated compound | IC50 (μM) a | ||
|---|---|---|---|
| MCF-7 b | MDA-MB-231 | ||
| 1 | 6-Hydroxy-3,5-dimethoxy-2-methyl-1,4-naphthoquinone | >403 | >403 |
| 2 | Ancistroquinone C | >370 | 330 |
| 3 | 5,8-Dihydroxy-3-methoxy-2-methylnaphthalene-1,4-dione | 1.15 | 408 |
| 4 | Malvone A | 222 | 65 |
| 5 | Droserone | >490 | >490 |
| 6 | Droserone-5-methyl ether | >459 | >459 |
| 7 | Hydroxydroserone | 432 | >455 |
| 8 | Chrysophanol | >394 | >394 |
| 9 | Helminthosporin | >370 | >370 |
| 10 | Aloesaponarin I | 211 | >357 |
| 11 | Aloesaponarin II | 157 | 72 |
| 12 | Laccaic acid D-methyl ester | >305 | 277 |
| 13 | Deoxyerythrolaccin | 178 | 140 |
| 14 | 1-Hydroxy-8-methoxy-3-methylanthraquinone | 4.85 | >100 |
| 15 | Aloesaponol I | >352 | 125 |
| 16 | Aloesaponol II-6-methyl ether | 261 | 131 |
3.2. Plant Material
3.3. Extraction and Isolation
3.4. Cytotoxicity Assay
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflictts of Interest
References
- Smith, G.F.; Vanwyk, B.E.; Mossmer, M.; Viljoen, A. The Taxonomy of Aloinella, Guillauminia and Lemee (Aloaceae). Taxon 1995, 44, 513–517. [Google Scholar] [CrossRef]
- Okamura, N.; Asai, M.; Hine, N.; Yagi, A. High-performance liquid chromatographic determination of phenolic compounds in Aloe species. J. Chromatogr. A 1996, 746, 225–231. [Google Scholar] [CrossRef]
- Mebberley, D.J. The Plant Book: A Portable Dictionary of Higher Plants; Cambridge University Press: Cambridge, UK, 1987. [Google Scholar]
- Demissew, S. The botany and chemistry of Aloes of Africa. Bull. Chem. Soc. Ethiop. 1996, 10, 73–88. [Google Scholar]
- Reynolds, G.W. Aloes of Tropical Africa and Madagascar; Aloe Book Fund: Mbabane, Swaziland, 1966. [Google Scholar]
- Viljoen, A.M.; van Wyk, B. The chemotaxonomic significance of the phenylpyrone aloenin in the genus Aloe. Biochem. Syst. Ecol. 2000, 28, 1009–1017. [Google Scholar] [CrossRef]
- Berger, A. Das Pflanzreich-Liliaceae-Asphodeloideae-Aloineae. Part. 33; Verlag von H.R. Engelmann: Weinheim, Germany, 1959. [Google Scholar]
- Schmelzer, G.H.; Gurib-Fakim, A. Plant. Resources of Tropical Africa II (1). Medicinal Plants 1; PROTA Foundation: Wogeningen/Leiden, The Netherlands, 2008. [Google Scholar]
- Bringmann, G.; Rudenauer, S.; Irmer, A.; Bruhn, T.; Brun, R.; Heimberger, T.; Stuhmer, T.; Bargou, R.; Chatterjee, M. Antitumoral and antileishmanial dioncoquinones and ancistroquinones from cell cultures of Triphyophyllum peltatum (Dioncophyllaceae) and Ancistrocladus abbreviatus (Ancistrocladaceae). Phytochemistry 2008, 69, 2501–2509. [Google Scholar] [CrossRef]
- Sankaram, A.V.B.; Rao, A.S.; Shoolery, J.N. Zeylanone and Isozeylanone, 2 Novel Quinones from Plumbagozeylanica. Tetrahedron 1979, 35, 1777–1782. [Google Scholar] [CrossRef]
- Rischer, H.; Hamm, A.; Bringmann, G. Nepenthes insignis uses a C2-portion of the carbon skeleton of L-alanine acquired via its carnivorous organs, to build up the allelochemical plumbagin. Phytochemistry 2002, 59, 603–609. [Google Scholar] [CrossRef]
- Bringmann, G.; Rischer, H.; Wohlfarth, M.; Schlauer, J.; Assi, L.A. Droserone from cell cultures of Triphyophyllum peltatum (Dioncophyllaceae) and its biosynthetic origin. Phytochemistry 2000, 53, 339–343. [Google Scholar] [CrossRef]
- Induli, M.; Cheloti, M.; Wasuna, A.; Wekesa, I.; Wanjohi, J.M.; Byamukama, R.; Heydenreich, M.; Makayoto, M.; Yenesew, A. Naphthoquinones from the roots of Aloe secundiflora. Phytochem. Lett. 2012, 5, 506–509. [Google Scholar] [CrossRef]
- Veshkurova, O.; Golubenko, Z.; Pshenichnov, E.; Arzanova, I.; Uzbekov, V.; Sultanova, E.; Salikhov, S.; Williams, H.J.; Reibenspies, J.H.; Puckhaber, L.S.; et al. Malvone A, a phytoalexin found in Malva sylvestris (family Malvaceae). Phytochemistry 2006, 67, 2376–2379. [Google Scholar] [CrossRef]
- Kreher, B.; Neszmelyi, A.; Wagner, H. Naphthoquinones from Dionaea-Muscipula. Phytochemistry 1990, 29, 605–606. [Google Scholar] [CrossRef]
- Likhitwitayawuid, K.; Kaewamatawong, R.; Ruangrungsi, N.; Krungkrai, J. Antimalarial naphthoquinones from Nepenthes thorelii. Planta Med. 1998, 64, 237–241. [Google Scholar] [CrossRef]
- Cooke, R.G.; Segal, W. Colouring Matters of Australian Plants. 1. The Structure of Droserone. Aust. J. Sci. Res. Ser. A 1950, 3, 628–634. [Google Scholar]
- Ghera, E.; Bendavid, Y. Annulation Reactions Leading to Naphthalene Derivatives—New Syntheses of Natural 1,2-Naphthoquinones and 1,4-Naphthoquinones. J. Org. Chem. 1985, 50, 3355–3359. [Google Scholar] [CrossRef]
- Winzor, F.L. The colouring matters of Drosera whittakeri Part III The synthesis of hydroxydroserone. J. Chem. Soc. 1935, 336–338. [Google Scholar] [CrossRef]
- Budzianowski, J. Naphthoquinone glucosides of Drosera gigantea from in vitro cultures. Planta Med. 2000, 66, 667–669. [Google Scholar] [CrossRef]
- Timmers, M. Marine and Terrestrial Natural Products Discovery; Royal Melbourne Institute of Technology: Melbourne, Australia, 2013. [Google Scholar]
- Todorova, G.; Lazarova, I.; Mikhova, B.; Kostova, I. Anthraquinone, Naphthalene, and Naphthoquinone Components of Asphodeline lutea. Chem. Nat. Compd. 2010, 46, 322–323. [Google Scholar] [CrossRef]
- Balestra, F.; Batusov, Y.A.; Bendiscioli, G.; Bossolasco, S.; Breivik, F.O.; Bunyatov, S.A.; Bussa, M.P.; Busso, L.; Danielsen, K.M.; Guaraldo, C.; et al. Experimental Results on Antiproton Interaction with Nuclei Obtained in the Ps179 Cern Experiment at Lear. AIP Conf. Proc. 1992, 243, 361–367. [Google Scholar] [CrossRef]
- Yagi, A.; Makino, K.; Nishioka, I. Studies on Constituents of Aloe saponaria Haw. 2. Structures of Tetrahydroanthracene Derivatives, Aloesaponol-III and Aloesaponol-IV. Chem. Pharm. Bull. 1977, 25, 1764–1770. [Google Scholar] [CrossRef]
- Yagi, A.; Makino, K.; Nishioka, I. Studies on Constituents of Aloe sapnaria Haw. 1. Structures of Tetrahydroanthracene Derivatives and Related Anthraquinones. Chem. Pharm. Bull. 1974, 22, 1159–1166. [Google Scholar] [CrossRef]
- Dagne, E.; Casser, I.; Steglich, W. Aloechrysone, a Dihydroanthracenone from Aloe berhana. Phytochemistry 1992, 31, 1791–1793. [Google Scholar] [CrossRef]
- Mehandale, A.R.; Rao, A.V.R.; Shaikh, I.N.; Venkataraman, K. Desoxyerythrolaccin and Laccaic Acid D. Tetrahedron Lett. 1968, 9, 2231–2234. [Google Scholar]
- Zaman, K.; Khan, M.R.; Ali, M.; Maitland, D.J. New anthraquinone dimer from the root bark of Cassia artemisioides (Gaudich. Ex. DC) Randell. J. Asian Nat. Prod. Res. 2011, 13, 62–67. [Google Scholar] [CrossRef]
- Makino, K.; Yagi, A.; Nishioka, I. Studies on the constituents of Aloe arborescens Mill. var. natalensis Berger. II. The structures of two new aloesin esters. Chem. Pharm. Bull. 1974, 22, 1565–1570. [Google Scholar] [CrossRef]
- Van wyk, B.E.; Yenesew, A.; Dagne, E. Chemotaxonomic Survey of Anthraquinones and Pre-Anthraquinones in Roots of Aloe species. Biochem. System. Ecol. 1995, 23, 267–275. [Google Scholar] [CrossRef]
- Monks, T.J.; Jones, D.C. The metabolism and toxicity of quinones, quinonimines, quinone methides, and quinone-thioethers. Curr. Drug Metab. 2002, 3, 425–438. [Google Scholar] [CrossRef]
- Huang, Q.; Lu, G.; Sben, H.M.; Cbung, M.C.M.; Ong, C.N. Anti-cancer properties of anthraquinones from rhubarb. Med. Res. Rev. 2007, 27, 609–630. [Google Scholar] [CrossRef]
- Endale, M.; Alao, J.P.; Akala, H.M.; Rono, N.K.; Eyase, F.L.; Derese, S.; Ndakala, A.; Mbugua, M.; Walsh, D.S.; Sunnerhagen, P.; et al. Antiplasmodial Quinones from Pentas longiflora and Pentas lanceolata. Planta Med. 2012, 78, 31–35. [Google Scholar] [CrossRef]
- Queiroz, M.L.; Valadares, M.C.; Torello, C.O.; Ramos, A.L.; Oliveira, A.B.; Rocha, F.D.; Arruda, V.A.; Accorci, W.R. Comparative studies of the effects of Tabebuia avellanedae bark extract and beta-lapachone on the hematopoietic response of tumour-bearing mice. J. Ethnopharmacol. 2008, 117, 228–235. [Google Scholar] [CrossRef]
- Lim, E.S.; Rhee, Y.H.; Park, M.K.; Shim, B.S.; Ahn, K.S.; Kang, H.; Yoo, H.S.; Kim, S.H. DMNQ S-64 induces apoptosis via caspase activation and cyclooxygenase-2 inhibition in human nonsmall lung cancer cells. Ann. N. Y. Acad. Sci. 2007, 1095, 7–18. [Google Scholar] [CrossRef]
- Lu, J.J.; Bao, J.L.; Wu, G.S.; Xu, W.S.; Huang, M.Q.; Chen, X.P.; Wang, Y.T. Quinones derived from plant secondary metabolites as anti-cancer agents. Anticancer Agents Med. Chem. 2013, 13, 456–463. [Google Scholar] [CrossRef]
- Dinér, P.; Alao, J.P.; Söderlund, J.; Sunnerhagen, P.; Grotli, M. Preparation of 3-substituted-1-isopropyl-1H-pyrazolo[3,4-d]pyrimidin-4-amines as RET kinase inhibitors. J. Med. Chem. 2012, 55, 4872–4876. [Google Scholar] [CrossRef]
- Wokaun, A.; Ernst, R.R. Selective Detection of multiple quantum transitions in NMR by 2-dimensional spectroscopy. Chem. Phys. Lett. 1977, 52, 407–412. [Google Scholar] [CrossRef]
- Kumar, A.; Ernst, R.R.; Wüthrich, K. A Two-dimensional nuclear Overhauser enhancement (2D NOE) Experiment for the elucidation of complete proton-proton cross-relaxation networks in biological macromolecules. Biochem. Biophys. Res. Commun. 1980, 95, 1–6. [Google Scholar] [CrossRef]
- Perpickdumont, M.; Reynolds, W.F.; Enriquez, R.G. C-13–H-1 shift correlation with full H-1-H-1 decoupling. Magn. Reson. Chem. 1988, 26, 358–361. [Google Scholar] [CrossRef]
- Hurd, R.E.; John, B.K. Gradient-enhanced proton-detected heteronuclear multiple-quantum coherence spectroscopy. J. Magn. Reson. 1991, 91, 648–653. [Google Scholar]
- Bbosa, G.S.; Kyegombe, D.B.; Lubega, A.; Musisi, N.; Ogwal-Okeng, J.; Odyek, O. Anti-Plasmodium falciparum activity of Aloe dawei and Justicia betonica. Afr. J. Pharm. Pharmacol. 2013, 7, 2258–2263. [Google Scholar] [CrossRef]
- Sample Availability: Samples of the compounds 1–16 are available from the authors.
© 2014 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Abdissa, N.; Induli, M.; Fitzpatrick, P.; Alao, J.P.; Sunnerhagen, P.; Landberg, G.; Yenesew, A.; Erdélyi, M. Cytotoxic Quinones from the Roots of Aloe dawei. Molecules 2014, 19, 3264-3273. https://doi.org/10.3390/molecules19033264
Abdissa N, Induli M, Fitzpatrick P, Alao JP, Sunnerhagen P, Landberg G, Yenesew A, Erdélyi M. Cytotoxic Quinones from the Roots of Aloe dawei. Molecules. 2014; 19(3):3264-3273. https://doi.org/10.3390/molecules19033264
Chicago/Turabian StyleAbdissa, Negera, Martha Induli, Paul Fitzpatrick, John Patrick Alao, Per Sunnerhagen, Göran Landberg, Abiy Yenesew, and Máté Erdélyi. 2014. "Cytotoxic Quinones from the Roots of Aloe dawei" Molecules 19, no. 3: 3264-3273. https://doi.org/10.3390/molecules19033264
APA StyleAbdissa, N., Induli, M., Fitzpatrick, P., Alao, J. P., Sunnerhagen, P., Landberg, G., Yenesew, A., & Erdélyi, M. (2014). Cytotoxic Quinones from the Roots of Aloe dawei. Molecules, 19(3), 3264-3273. https://doi.org/10.3390/molecules19033264





