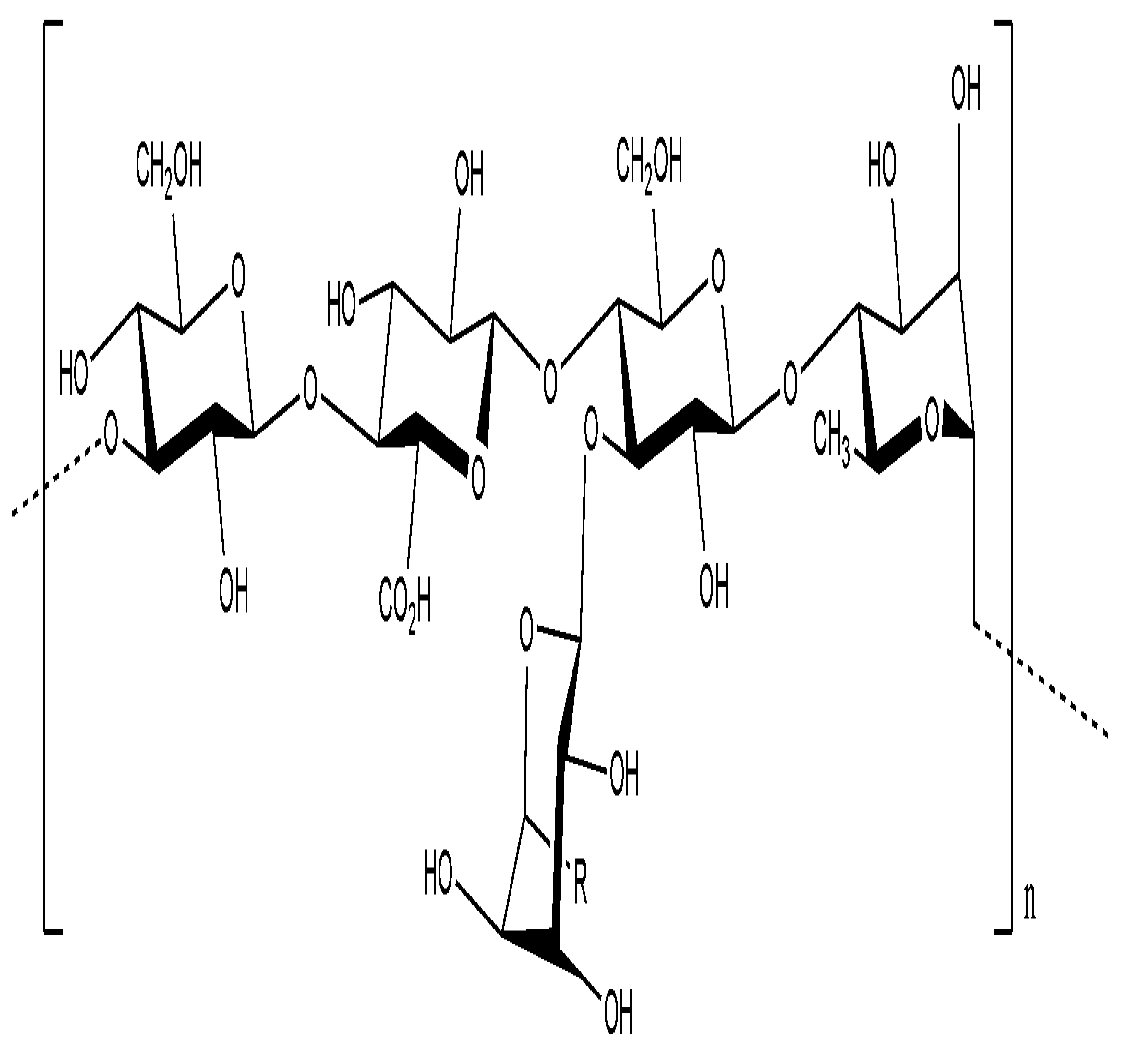
Bacterial Extracellular Polysaccharides Involved in Biofilm Formation
Abstract
:1. Introduction
2. Microbial extracellular polysaccharides as an integral part of bacterial biofilms
3. Role of EPS in the regulation of biofilm formation
4. Control of EPS production
5. EPS implications in biotechnology
6. EPS roles in bioleaching
7. Chemical composition of EPS produced by A. ferrooxidans
7.1. A. ferrooxidans EPS produced on pyrite
7.2. A. ferrooxidans EPS produced on sulfur
7.3. A. ferrooxidans EPS produced on ferrous sulfate
8. Extracellular polysaccharides produced by pathogenic microorganisms
9. Extracellular polysaccharides employed in biotechnology applications
9.1. Extracellular polysaccharides produced by gram-positive bacteria, Firmicutes
9.1.1. Dextran
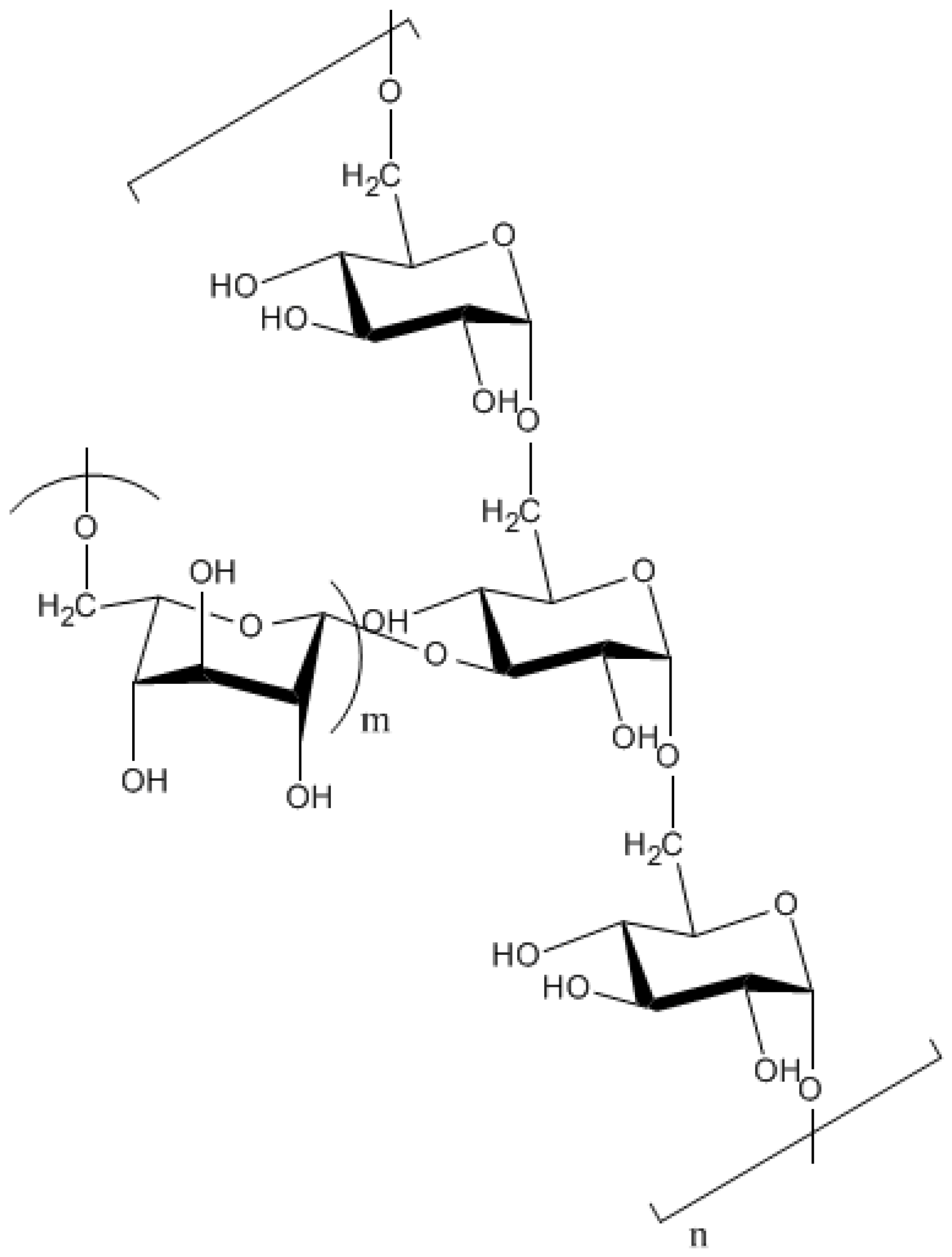
9.1.2. Kefiran
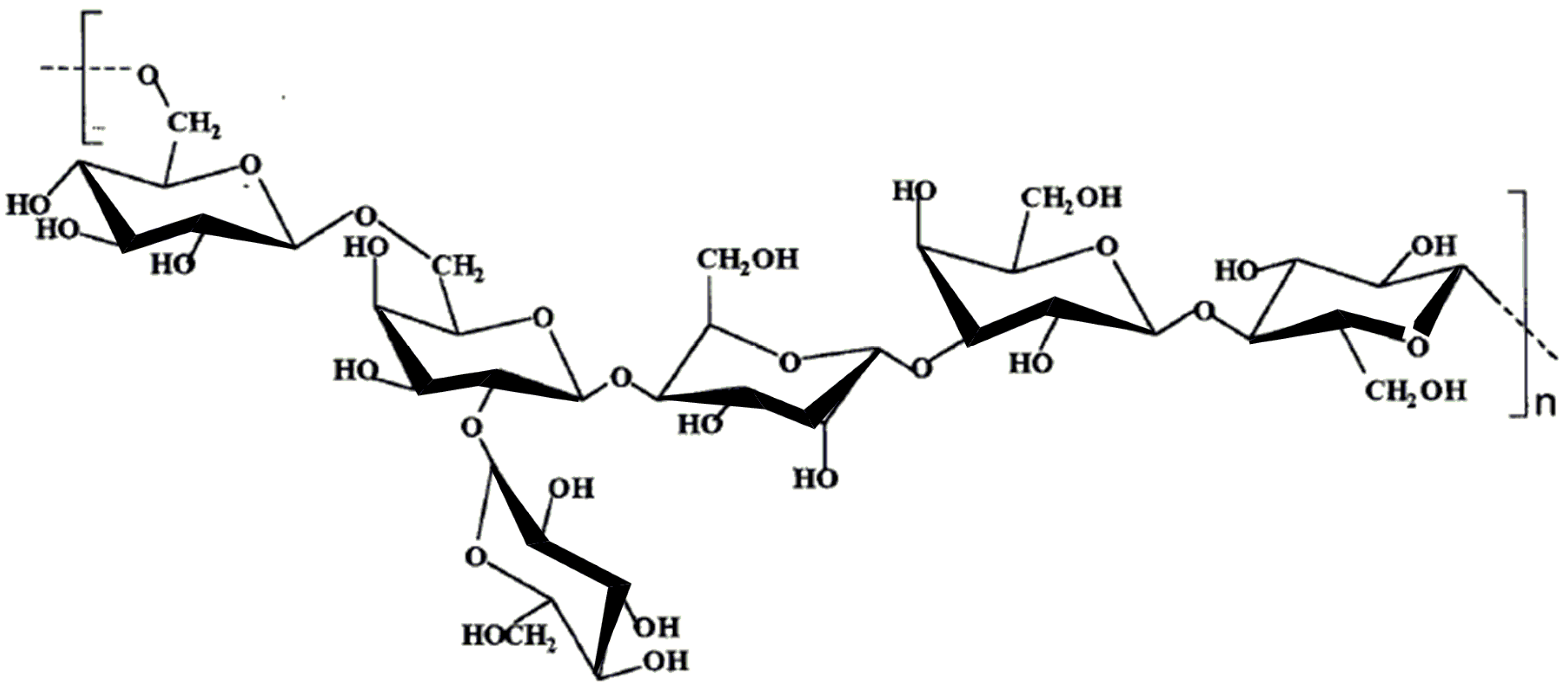
9.2. Extracellular polysaccharides produced by gram-negative bacteria, Proteobacteria
9.2.1. Alphaproteobacteria
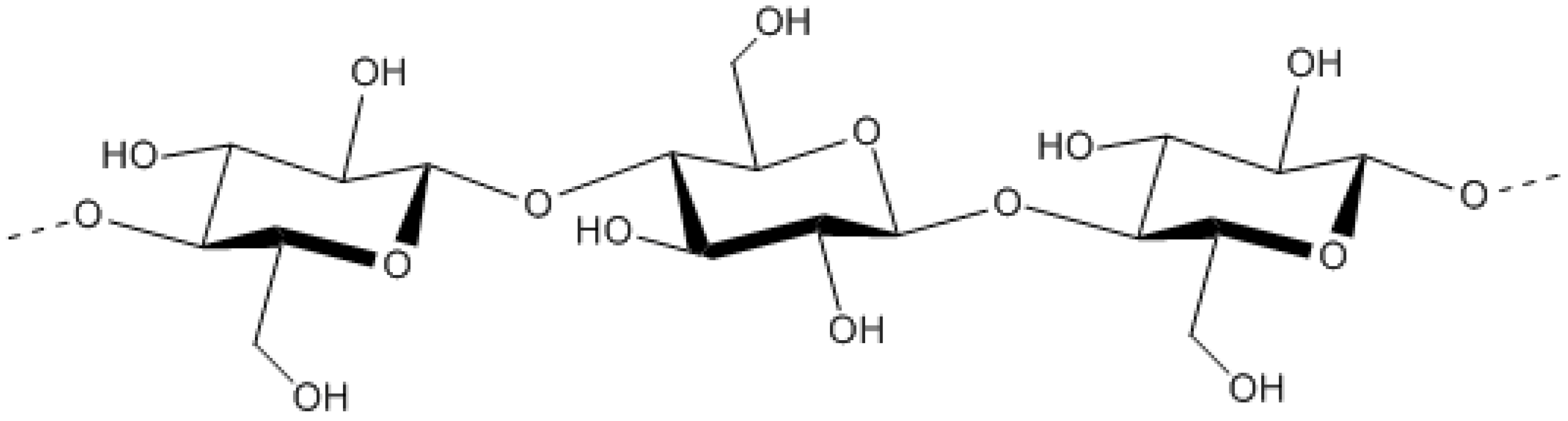
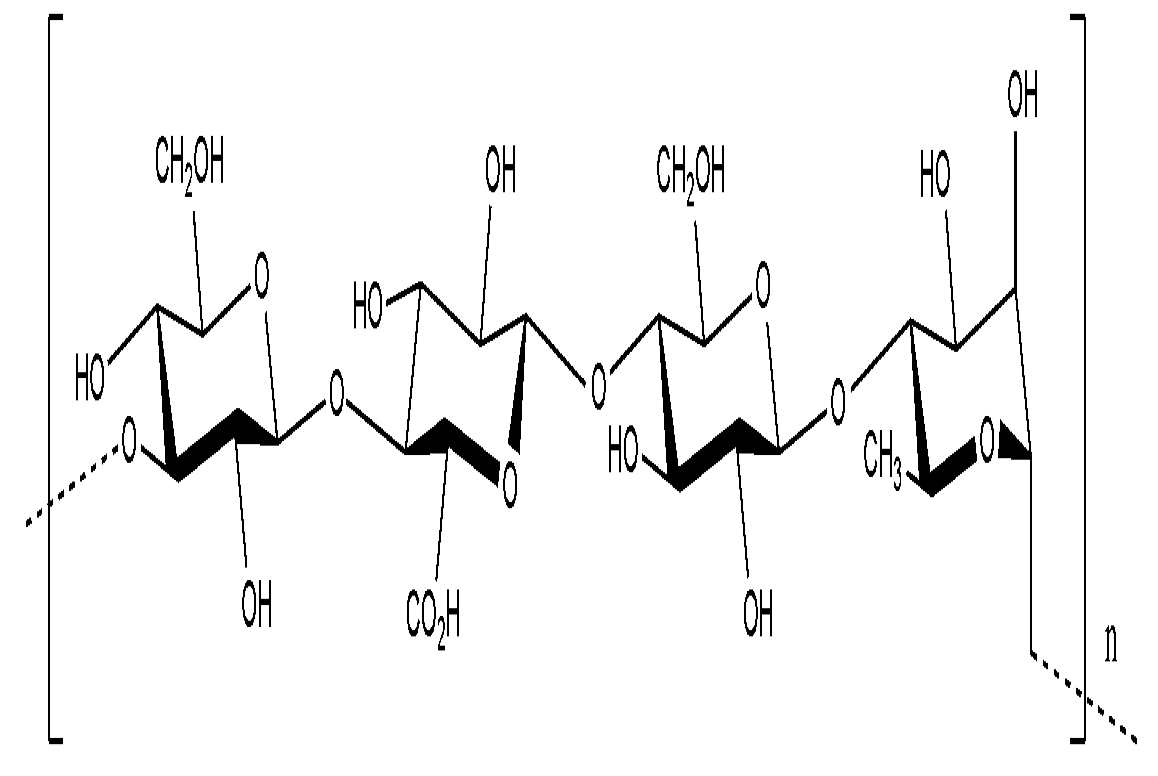
9.2.2. Betaproteobacteria
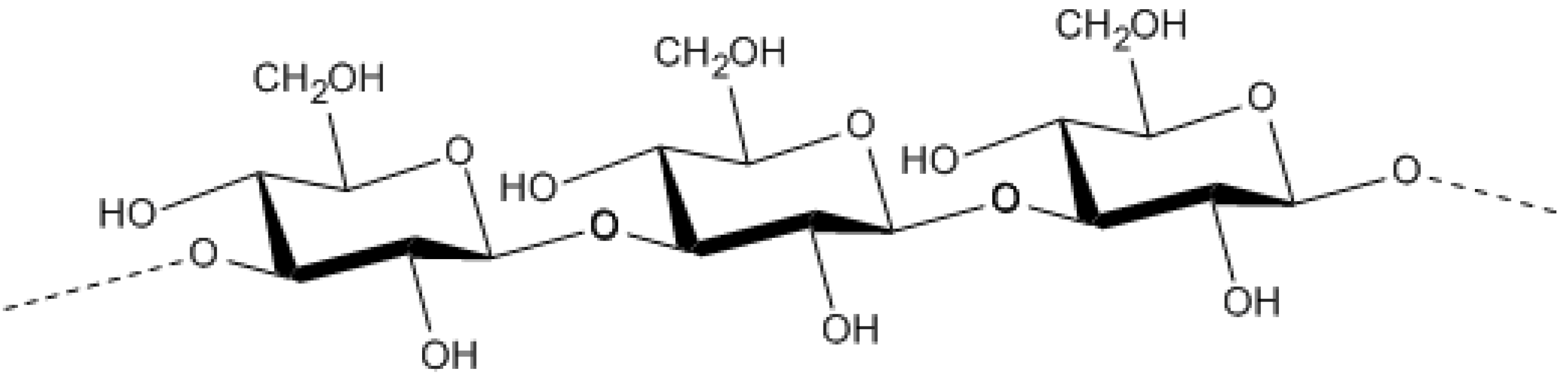
9.2.3. Gammaproteobacteria
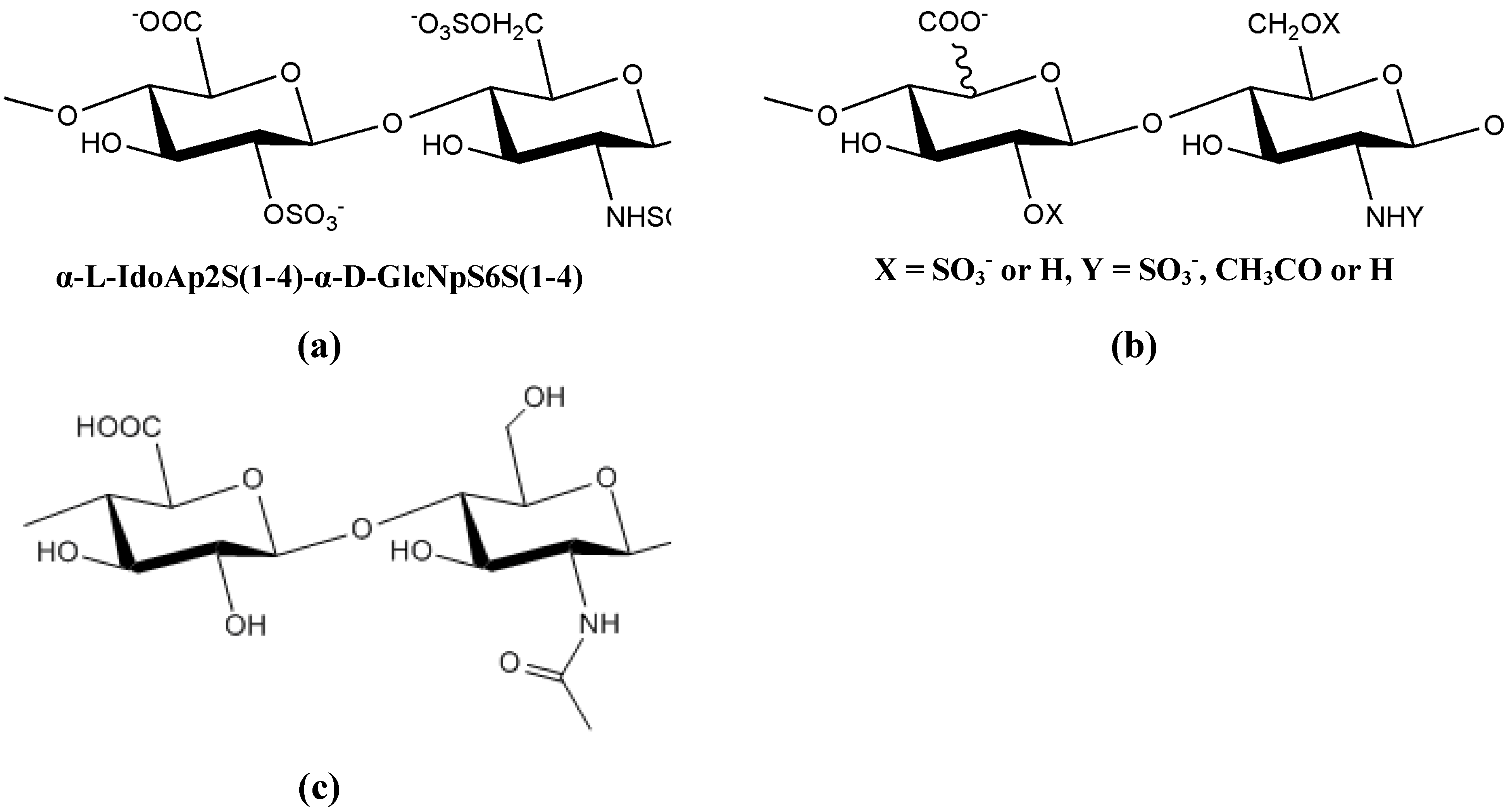
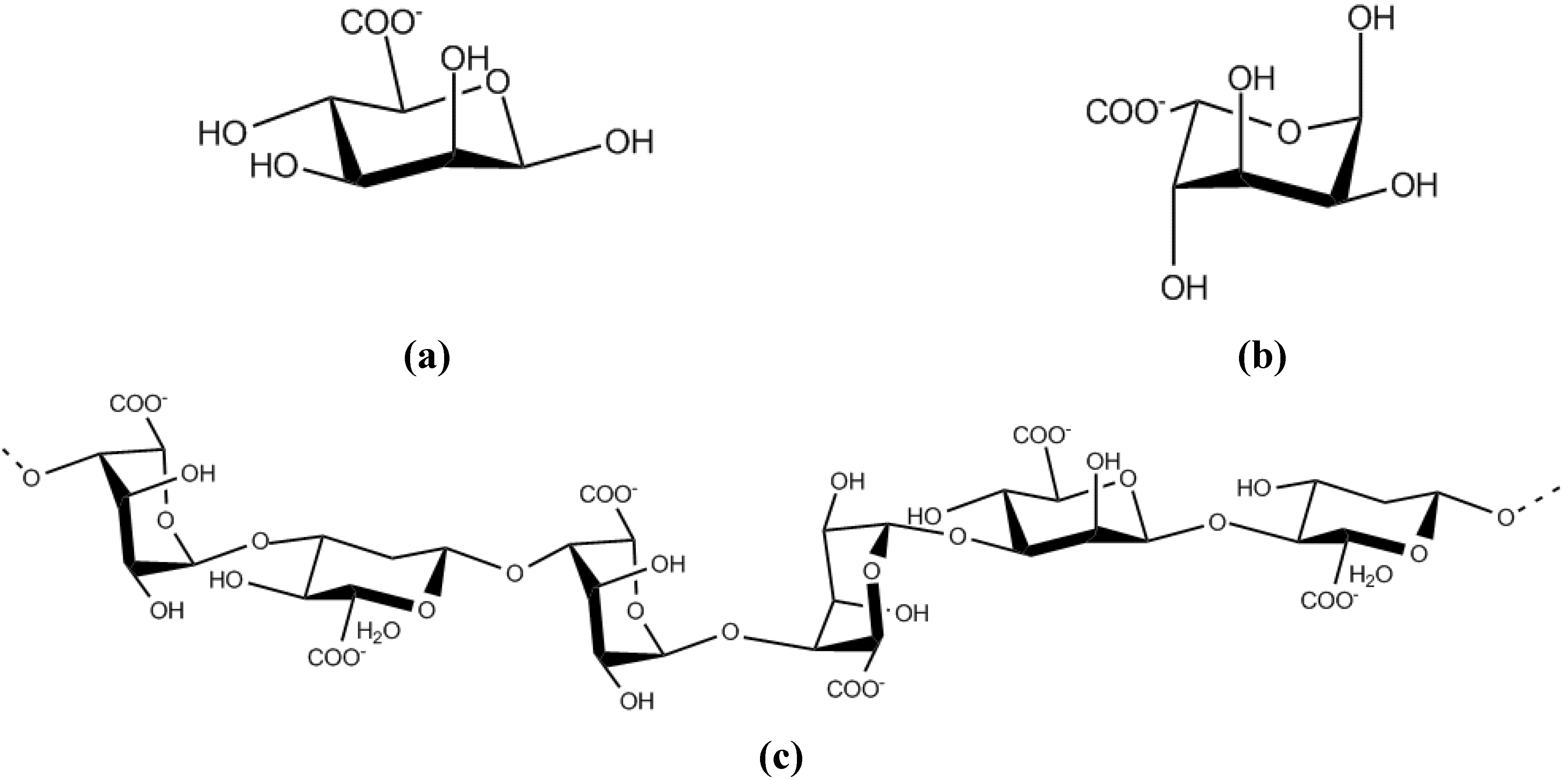
10. Novel techniques in the study of biofilm-specific polysaccharides
11. Conclusions
References
- Donlan, R.M. Biofilms: Microbial life on surfaces. Emerging Infect. Dis. 2002, 8, 881–890. [Google Scholar] [CrossRef]
- Costerton, J.W. Introduction to biofilm. Int. J. Antimicrob. Agents 1999, 11, 217–221. [Google Scholar] [CrossRef]
- Costerton, J.W.; Lewandowski, Z.; Caldwell, D.E.; Korber, D.R.; Lappin-Scott, H.M. Microbial biofilms. Annu. Rev. Microbiol. 1995, 49, 711–745. [Google Scholar] [CrossRef]
- Socransky, S.S.; Haffajee, A.D.; Cugini, M.A.; Smith, C.; Kent, R.L., Jr. Microbial complexes in subgingival plaque. J. Clin. Periodontol. 1998, 25, 134–144. [Google Scholar] [CrossRef]
- Costerton, J.W.; Stewart, P.S.; Greenberg, E.P. Bacterial biofilms: A common cause of persistent infections. Science 1999, 284, 1318–1322. [Google Scholar] [CrossRef]
- Heukelekian, H.; Heller, A. Relation between food concentration and surface for bacterial growth. J. Bacteriol. 1940, 40, 547–558. [Google Scholar]
- Zobell, C.E. The effect of solid surfaces upon bacterial activity. J. Bacteriol. 1943, 46, 39–56. [Google Scholar]
- Davey, M.E.; O'Toole, G.A. Microbial biofilms: from ecology to molecular genetics. Microbiol. Mol. Biol. Rev. 2000, 64, 847–867. [Google Scholar] [CrossRef]
- Costerton, J.W.; Geesey, G.G.; Cheng, K.J. How bacteria stick. Sci. Am. 1978, 238, 86–95. [Google Scholar] [CrossRef]
- Flemming, H.C.; Wingender, J. Relevance of microbial extracellular polymeric substances (EPSs) – Part I: Structural and ecological aspects. Water Sci. Technol. 43, 1–8.
- Singh, R.; Paul, D.; Jain, R.K. Biofilms: implications in bioremediation. Trends Microbiol. 2006, 14, 389–397. [Google Scholar] [CrossRef]
- Ahimou, F.; Semmens, M.J.; Haugstad, G.; Novak, P.J. Effect of protein, polysaccharide, and oxygen concentration profiles on biofilm cohesiveness. Appl. Environ. Microbiol. 2007, 73, 2905–2910. [Google Scholar] [CrossRef]
- Costerton, W.; Veeh, R.; Shirtliff, M.; Pasmore, M.; Post, C.; Ehrlich, G. The application of biofilm science to the study and control of chronic bacterial infections. J. Clin. Invest. 2003, 112, 1466–1477. [Google Scholar]
- Flemming, H.C.; Wingender, J. Relevance of microbial extracellular polymeric substances (EPSs) – Part II: Technical aspects. Water Sci. Technol. 43, 9–16.
- Sutherland, I.W. Biofilm exopolysaccharides: a strong and sticky framework. Microbiology 2001, 147, 3–9. [Google Scholar]
- van Hullebusch, E.D.; Zandvoort, M.H.; Lens, P.N.L. Metal immobilisation by biofilms: mechanisms and analytical tools. Rev. Environ. Sci. Biotechnol. 2003, 2, 9–33. [Google Scholar] [CrossRef]
- Mayer, C.; Moritz, R.; Kirschner, C.; Borchard, W.; Maibaum, R.; Wingender, J.; Flemming, H.-C. The role of intermolecular interactions: studies on model systems for bacterial biofilms. Int. J. Biol. Macromol. 1999, 26, 3–16. [Google Scholar] [CrossRef]
- Romani, A.M.; Fund, K.; Artigas, J.; Schwartz, T.; Sabater, S.; Obst, U. Relevance of polymeric matrix enzymes during biofilm formation. Microb. Ecol. 2008, 56, 427–436. [Google Scholar] [CrossRef]
- Beech, I.B. Corrosion of technical materials in the presence of biofilms--current understanding and state-of-the art methods of study. Int. Biodeterior. Biodegradation 2004, 53, 177–183. [Google Scholar] [CrossRef]
- Ruiz, L.M.; Valenzuela, S.; Castro, M.; Gonzalez, A.; Frezza, M.; Soulère, L.; Rohwerder, T.; Queneau, Y.; Doutheau, A.; Sand, W.; Jerez, C.A.; Guiliani, N. AHL communication is a widespread phenomenon in biomining bacteria and seems to be involved in mineral-adhesion efficiency. Hydrometallurgy 2008, 94, 133–137. [Google Scholar] [CrossRef]
- Waters, C.M.; Bassler, B.L. Quorum sensing: cell-to-cell communication in bacteria. Annu. Rev. Cell Dev. Biol. 2005, 21, 319–346. [Google Scholar] [CrossRef]
- von Bodman, S.B.; Majerczak, D.R.; Coplin, D.L. A negative regulator mediates quorum-sensing control of exopolysaccharide production in Pantoea stewartii subsp. stewartii. Proc. Natl. Acad. Sci. USA 1998, 95, 7687–7692. [Google Scholar] [CrossRef]
- Rivas, M.; Seeger, M.; Holmes, D.S.; Jedlicki, E. A Lux-like quorum sensing system in the extreme acidophile Acidithiobacillus ferrooxidans. Biol. Res. 2005, 38, 283–297. [Google Scholar]
- Davies, D.G.; Parsek, M.R.; Pearson, J.P.; Iglewski, B.H.; Costerton, J.W.; Greenberg, E.P. The involvement of cell-to-cell signals in the development of a bacterial biofilm. Science 1998, 280, 295–298. [Google Scholar] [CrossRef]
- Valenzuela, L.; Chi, A.; Beard, S.; Orell, A.; Guiliani, N.; Shabanowitz, J.; Hunt, D.F.; Jerez, C.A. Genomics, metagenomics and proteomics in biomining microorganisms. Biotechnol. Adv. 2006, 24, 197–211. [Google Scholar] [CrossRef]
- Hooshangi, S.; Bentley, W.E. From unicellular properties to multicellular behavior: bacteria quorum sensing circuitry and applications. Curr. Opin. Biotechnol. 2008, 19, 550–555. [Google Scholar] [CrossRef]
- Miller, M.B.; Bassler, B.L. Quorum sensing in bacteria. Annu. Rev. Microbiol. 2001, 55, 165–199. [Google Scholar] [CrossRef]
- Farah, C.; Vera, M.; Morin, D.; Haras, D.; Jerez, C.A.; Guiliani, N. Evidence for a functional quorum-sensing type AI-1 system in the extremophilic bacterium Acidithiobacillus ferrooxidans. Appl. Environ. Microbiol. 2005, 71, 7033–7040. [Google Scholar] [CrossRef]
- Urbanczyk, H.; Ast, J.C.; Higgins, M.J.; Carson, J.; Dunlap, P.V. Reclassification of Vibrio fischeri, Vibrio logei, Vibrio salmonicida and Vibrio wodanis as Aliivibrio fischeri gen. nov., comb. nov., Aliivibrio logei comb. nov., Aliivibrio salmonicida comb. nov. and Aliivibrio wodanis comb. nov. Int. J. Syst. Evol. Microbiol. 2007, 57, 2823–2829. [Google Scholar] [CrossRef]
- Hall-Stoodley, L.; Stoodley, P. Developmental regulation of microbial biofilms. Curr. Opin. Biotechnol. 2002, 13, 228–233. [Google Scholar] [CrossRef]
- Rashid, M.H.; Rumbaugh, K.; Passador, L.; Davies, D.G.; Hamood, A.N.; Iglewski, B.H.; Kornberg, A. Polyphosphate kinase is essential for biofilm development, quorum sensing, and virulence of Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 2000, 97, 9636–9641. [Google Scholar]
- Nakamura, S.; Higashiyama, Y.; Izumikawa, K.; Seki, M.; Kakeya, H.; Yamamoto, Y.; Yanagihara, K.; Miyazaki, Y.; Mizuta, Y.; Kohno, S. The roles of quorum-sensing system in the release of extracellular DNA, lipopolysaccharide, and membrane vesicles from Pseudomonas aeruginosa. Jpn. J. Infect. Dis. 2008, 61, 375–378. [Google Scholar]
- Li, J.; Attila, C.; Wang, L.; Wood, T.K.; Valdes, J.J.; Bentley, W.E. Quorum sensing in Escherichia coli is signaled by AI-2/LsrR: effects on small RNA and biofilm architecture. J. Bacteriol. 2007, 189, 6011–6020. [Google Scholar] [CrossRef]
- Fialho, A.M.; Moreira, L.M.; Granja, A.T.; Popescu, A.O.; Hoffmann, K.; Sá-Correia, I. Occurrence, production, and applications of gellan: current state and perspectives. Appl. Microbiol. Biotechnol. 2008, 79. [Google Scholar]
- Rehm, B. Microbial exopolysaccharides: Variety and potential applications. In Microbial production of biopolymers and polymer precursors: applications and perspectives; Caister Academic: Norfolk, UK, 2009; pp. 229–254. [Google Scholar]
- Rawlings, D.E.; Johnson, D.B. The microbiology of biomining: development and optimization of mineral-oxidizing microbial consortia. Microbiology 2007, 153, 315–324. [Google Scholar] [CrossRef]
- Gadd, G.M. Microbial influence on metal mobility and application for bioremediation. Geoderma 2004, 122, 109–119. [Google Scholar] [CrossRef]
- Liu, Y.G.; Zhou, M.; Zeng, G.M.; Wang, X.; Li, X.; Fan, T.; Xu, W.H. Bioleaching of heavy metals from mine tailings by indigenous sulfur-oxidizing bacteria: Effects of substrate concentration. Bioresour. Technol. 2008, 99, 4124–4129. [Google Scholar] [CrossRef]
- Pal, A.; Paul, A.K. Microbial extracellular polymeric substances: central elements in heavy metal bioremediation. Indian J. Microbiol. 2008, 48, 49–64. [Google Scholar] [CrossRef]
- Lynch, J.M.; Moffat, A.J. Bioremediation - prospects for the future application of innovative applied biological research. Ann. Appl. Biol. 2005, 146, 217–221. [Google Scholar] [CrossRef]
- Singh, P.; Cameotra, S.S. Enhancement of metal bioremediation by use of microbial surfactants. Biochem. Biophys. Res. Commun. 2004, 319, 291–297. [Google Scholar] [CrossRef]
- Sand, W.; Gehrke, T. Extracellular polymeric substances mediate bioleaching/biocorrosion via interfacial processes involving iron(III) ions and acidophilic bacteria. Res. Microbiol. 2006, 157, 49–56. [Google Scholar] [CrossRef]
- Sheng, X.; Ting, Y.P.; Pehkonen, S.O. The influence of ionic strength, nutrients and pH on bacterial adhesion to metals. J. Colloid Interface Sci. 2008, 321, 256–264. [Google Scholar] [CrossRef]
- Suzuki, I. Microbial leaching of metals from sulfide minerals. Biotechnol. Adv. 2001, 19, 119–132. [Google Scholar] [CrossRef]
- Rohwerder, T.; Gehrke, T.; Kinzler, K.; Sand, W. Bioleaching review part A: Progress in bioleaching: fundamentals and mechanisms of bacterial metal sulfide oxidation. Appl. Microbiol. Biotechnol. 2003, 63, 239–248. [Google Scholar] [CrossRef]
- Kelly, D.P.; Wood, A.P. Reclassification of some species of Thiobacillus to the newly designated genera Acidithiobacillus gen. nov., Halothiobacillus gen. nov. and Thermithiobacillus gen. nov. Int. J. Syst. Evol. Microbiol. 2000, 50, 511–516. [Google Scholar] [CrossRef]
- Bosecker, K. Bioleaching: metal solubilisation by microorganisms. FEMS Microbiol. Rev. 1997, 20, 591–604. [Google Scholar] [CrossRef]
- Ehrlich, H.L. Past, present and future of biohydrometallurgy. Hydrometallurgy 2001, 59, 127–134. [Google Scholar] [CrossRef]
- Olson, G.J.; Brierley, J.A.; Brierley, C.L. Bioleaching review part B: Progress in bioleaching: applications of microbial processes by the minerals industries. Appl. Environ. Microbiol. 2003, 63, 249–257. [Google Scholar]
- Gehrke, T.; Telegdi, J.; Thierry, D.; Sand, W. Importance of extracellular polymeric substances from Thiobacillus ferrooxidans for bioleaching. Appl. Environ. Microbiol. 1998, 64, 2743–2747. [Google Scholar]
- Barreto, M.; Jedlicki, E.; Holmes, D.S. Identification of a gene cluster for the formation of extracellular polysaccharide precursors in the chemolithoautotroph Acidithiobacillus ferrooxidans. Appl. Environ. Microbiol. 2005, 71, 2902–2909. [Google Scholar] [CrossRef]
- Gehrke, T.; Hallmann, R.; Kinzler, K.; Sand, W. The EPS of Acidithiobacillus ferrooxidans – a model for structure-function relationships of attached bacteria and their physiology. Water Sci. Technol. 2001, 43, 159–167. [Google Scholar]
- Kinzler, K.; Gehrke, T.; Telegdi, J.; Sand, W. Bioleaching—a result of interfacial processes caused by extracellular polymeric substances (EPS). Hydrometallurgy 2003, 71, 83–88. [Google Scholar] [CrossRef]
- Yu, R.L.; Tan, J.X.; Yang, P.; Sun, J.; Ouyang, X.J.; Dai, Y.J. EPS-contact-leaching mechanisms of chalcopyrite concentrates by A. ferrooxidans. Trans. Nonferrous Met. Soc. China 2008, 18, 1427–1432. [Google Scholar] [CrossRef]
- Das, T.; Ayyappan, S.; Roy Chaudhury, G. Factors affecting bioleaching kinetics of sulfide ores using acidophilic micro-organisms. BioMetals 1999, 12, 1–10. [Google Scholar] [CrossRef]
- Zimmerley, S.R.; Wilson, D.G.; Prater, J.D. Cyclic leaching process employing iron oxidizing bacteria. U.S. Pat. Off. 1958, 829, 964. [Google Scholar]
- Watling, H.R. The bioleaching of sulphide minerals with emphasis on copper sulphides -- A review. Hydrometallurgy 2006, 84, 81–108. [Google Scholar] [CrossRef]
- Devasia, P.; Natarajan, K.A.; Sathyanarayana, D.N.; Rao, G.R. Surface chemistry of Thiobacillus ferrooxidans relevant to adhesion on mineral surfaces. Appl. Environ. Microbiol. 1993, 59, 4051–4055. [Google Scholar]
- Escobar, B.; Huerta, G.; Rubio, J. Short Communication: Influence of lipopolysaccharides on the attachment of Thiobacillus ferrooxidans to minerals. World J. Microbiol. Biotechnol. 1997, 13, 593–594. [Google Scholar] [CrossRef]
- Harneit, K.; Goksel, A.; Kock, D.; Klock, J.H.; Gehrke, T.; Sand, W. Adhesion to metal sulfide surfaces by cells of Acidithiobacillus ferrooxidans, Acidithiobacillus thiooxidans and Leptospirillum ferrooxidans. Hydrometallurgy 2006, 83, 245–254. [Google Scholar] [CrossRef]
- Ghauri, M.A.; Okibe, N.; Johnson, D.B. Attachment of acidophilic bacteria to solid surfaces: The significance of species and strain variations. Hydrometallurgy 2007, 85, 72–80. [Google Scholar] [CrossRef]
- Arredondo, R.; Garcia, A.; Jerez, C.A. Partial removal of lipopolysaccharide from Thiobacillus ferrooxidans affects its adhesion to solids. Appl. Environ. Microbiol. 1994, 60, 2846–2851. [Google Scholar]
- Sand, W.; Gehrke, T.; Jozsa, P.G.; Schippers, A. (Bio)chemistry of bacterial leaching - direct vs. indirect bioleaching. Hydrometallurgy 2001, 59, 159–175. [Google Scholar] [CrossRef]
- Vuong, C.; Kocianova, S.; Voyich, J.M.; Yao, Y.; Fischer, E.R.; DeLeo, F.R.; Otto, M. A crucial role for exopolysaccharide modification in bacterial biofilm formation, immune evasion, and virulence. J. Biol. Chem. 2004, 279, 54881–54886. [Google Scholar]
- Sanford, P.A.; Cottrell, I.W.; Pettitt, D.J. Microbial polysaccharides: New products and their commercial applications. Pure Appl. Chem. 1984, 56, 879–892. [Google Scholar] [CrossRef]
- Sutherland, I.W. Novel and established applications of microbial polysaccharides. Trends Biotechnol. 1998, 16, 41–46. [Google Scholar] [CrossRef]
- Ertesvåg, H.; Valla, S. Biosynthesis and applications of alginates. Polym. Degrad. Stab. 1998, 59, 85–91. [Google Scholar] [CrossRef]
- Sutherland, I.W. Microbial polysaccharides from Gram-negative bacteria. Int. Dairy J. 2001, 11, 663–674. [Google Scholar] [CrossRef]
- Garrity, G.M. Bergey's Manual of Systematic Bacteriology: The Archaea and the deeply branching and phototrophic bacteria; Springer-Verlag: New York, NY, USA, 2001; pp. 155–166. [Google Scholar]
- Gibbons, N.E.; Murray, R.G.E. Proposals concerning the higher taxa of bacteria. Int. J. Syst. Bacteriol. 1978, 28, 1–6. [Google Scholar] [CrossRef]
- Wolf, M.; Muller, T.; Dandekar, T.; Pollack, J.D. Phylogeny of Firmicutes with special reference to Mycoplasma (Mollicutes) as inferred from phosphoglycerate kinase amino acid sequence data. Int. J. Syst. Evol. Microbiol. 2004, 54, 871–875. [Google Scholar] [CrossRef]
- Naessens, M.; Cerdobbel, A.; Soetaert, W.; Vandamme, E.J. Leuconostoc dextransucrase and dextran: production, properties and applications. J. Chem. Technol. Biotechnol. 2005, 80, 845–860. [Google Scholar] [CrossRef]
- Duboc, P.; Mollet, B. Applications of exopolysaccharides in the dairy industry. Int. Dairy J. 2001, 11, 759–768. [Google Scholar] [CrossRef]
- Lapasin, R.; Pricl, S. Rheology of industrial polysaccharides: Theory and applications. Aspen Publishers: New York, NY, USA, 1999; pp. 1–118. [Google Scholar]
- Micheli, L.; Uccelletti, D.; Palleschi, C.; Crescenzi, V. Isolation and characterisation of a ropy Lactobacillus strain producing the exopolysaccharide kefiran. Appl. Environ. Microbiol. 1999, 53, 69–74. [Google Scholar]
- Piermaria, J.A.; Pinotti, A.; Garcia, M.A.; Abraham, A.G. Films based on kefiran, an exopolysaccharide obtained from kefir grain: Development and characterization. Food Hydrocolloid. 2009, 23, 684–690. [Google Scholar] [CrossRef]
- Garrity, G.M.; Brenner, D.J.; Krieg, N.R.; Staley, J.R. Bergey's Manual of Systematic Bacteriology, Volume Two: The Proteobacteria, Parts A - C. Springer - Verlag: New York, NY, USA, 2005; p. 3001. [Google Scholar]
- Jonas, R.; Farah, L.F. Production and application of microbial cellulose. Polym. Degrad. Stab. 1998, 59, 101–106. [Google Scholar] [CrossRef]
- Römling, U. Molecular biology of cellulose production in bacteria. Res. Microbiol. 2002, 153, 205–212. [Google Scholar] [CrossRef]
- Klemm, D.; Schumann, D.; Udhardt, U.; Marsch, S. Bacterial synthesized cellulose -- artificial blood vessels for microsurgery. Prog. Polym. Sci. 2001, 26, 1561–1603. [Google Scholar] [CrossRef]
- Gummadi, S.N.; Kumar, K. Production of extracellular water insoluble β-1,3-glucan (curdlan) from Bacillus sp. SNC07. Biotechnol. Bioprocess Eng. 2005, 10, 546–551. [Google Scholar] [CrossRef]
- Dumitriu, S. Microbial exopolysaccharides. In Polysaccharides: Structural diversity and functional versatility; CRC Press: Marcel Dekker, New York, NY, USA, 2004; pp. 431–457. [Google Scholar]
- Jin, Y.; Zhang, H.; Yin, Y.; Nishinari, K. Conformation of curdlan as observed by tapping mode atomic force microscopy. Colloid Polym. Sci. 2006, 284, 1371–1377. [Google Scholar] [CrossRef]
- Chandrasekaran, R.; Radha, A. Molecular architectures and functional properties of gellan gum and related polysaccharides. Trends Food Sci. Technol. 1995, 6, 143–148. [Google Scholar]
- Linhardt, R.J.; Toida, T. Heparin oligosaccharides: New analogues development and applications. In Carbohydrates in Drug Design; Witczak, Z.B., Nieforth, K.A., Eds.; Marcel Dekker: New York, NY, USA, 1997; pp. 277–308. [Google Scholar]
- Warda, M.; Mao, W.; Toida, T.; Linhardt, R.J. Turkey intestine as a commercial source of heparin? Comparative structural studies of intestinal avian and mammalian glycosaminoglycans. Comp. Biochem. Physiol. B: Biochem. Mol. Biol. 2003, 134, 189–197. [Google Scholar] [CrossRef]
- Kusche, M.; Hannesson, H.H.; Lindahl, U. Biosynthesis of heparin. Use of Escherichia coli K5 capsular polysaccharide as a model substrate in enzymic polymer-modification reactions. Biochem. J. 1991, 275, 151–158. [Google Scholar]
- Wagner, M.; Ivleva, N.P.; Haisch, C.; Niessner, R.; Horn, H. Combined use of confocal laser scanning microscopy (CLSM) and Raman microscopy (RM): Investigations on EPS - Matrix. Water Res. 2009, 43, 63–76. [Google Scholar] [CrossRef]
- Denkhaus, E.; Meisen, S.; Telgheder, U.; Wingender, J. Chemical and physical methods for characterisation of biofilms. Microchim. Acta 2007, 158, 1–27. [Google Scholar] [CrossRef]
- Wolf, G.; Crespo, J.G.; Reis, M.A.M. Optical and spectroscopic methods for biofilm examination and monitoring. Rev. Environ. Sci. Biotechnol. 2002, 1, 227–251. [Google Scholar] [CrossRef]
- Howell, C.; Diesner, M.O.; Grunze, M.; Koelsch, P. Probing the extracellular matrix with sum-frequency-generation spectroscopy. Langmuir 2008, 24, 13819–13821. [Google Scholar] [CrossRef]
- Haisch, C.; Niessner, R. Visualisation of transient processes in biofilms by optical coherence tomography. Water Res. 2007, 41, 2467–2472. [Google Scholar] [CrossRef]
- Schmid, T.; Burkhard, J.; Yeo, B.S.; Zhang, W.; Zenobi, R. Towards chemical analysis of nanostructures in biofilms I: Imaging of biological nanostructures. Anal. Bioanal. Chem. 2008, 391, 1899–1905. [Google Scholar] [CrossRef]
- Pradhan, N.; Pradhan, S.K.; Nayak, B.B.; Mukherjee, P.S.; Sukla, L.B.; Mishra, B.K. Micro-Raman analysis and AFM imaging of Acidithiobacillus ferrooxidans biofilm grown on uranium ore. Res. Microbiol. 2008, 159, 557–561. [Google Scholar] [CrossRef]
- Mangold, S.; Harneit, K.; Rohwerder, T.; Claus, G.; Sand, W. Novel combination of atomic force microscopy and epifluorescence microscopy for visualization of leaching bacteria on pyrite. Appl. Environ. Microbiol. 2008, 74, 410–415. [Google Scholar] [CrossRef]
- Xi, C.; Marks, D.; Schlachter, S.; Luo, W.; Boppart, S.A. High-resolution three-dimensional imaging of biofilm development using optical coherence tomography. J. Biomed. Opt. 2006, 11, 034001–6. [Google Scholar] [CrossRef]
- Shen, Y.R. Surface properties probed by second-harmonic and sum-frequency generation. Nature 1989, 337, 519–525. [Google Scholar] [CrossRef]
- Morita, A.; Hynes, J.T. A theoretical analysis of the sum frequency generation spectrum of the water surface. Chem. Phys. 2000, 258, 371–390. [Google Scholar] [CrossRef]
- Shen, Y.R. A few selected applications of surface nonlinear optical spectroscopy. Appl. Phys. Sci. 1996, 96, 12104–12111. [Google Scholar]
- Sample Availability: Not available.
© 2009 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Vu, B.; Chen, M.; Crawford, R.J.; Ivanova, E.P. Bacterial Extracellular Polysaccharides Involved in Biofilm Formation. Molecules 2009, 14, 2535-2554. https://doi.org/10.3390/molecules14072535
Vu B, Chen M, Crawford RJ, Ivanova EP. Bacterial Extracellular Polysaccharides Involved in Biofilm Formation. Molecules. 2009; 14(7):2535-2554. https://doi.org/10.3390/molecules14072535
Chicago/Turabian StyleVu, Barbara, Miao Chen, Russell J. Crawford, and Elena P. Ivanova. 2009. "Bacterial Extracellular Polysaccharides Involved in Biofilm Formation" Molecules 14, no. 7: 2535-2554. https://doi.org/10.3390/molecules14072535
APA StyleVu, B., Chen, M., Crawford, R. J., & Ivanova, E. P. (2009). Bacterial Extracellular Polysaccharides Involved in Biofilm Formation. Molecules, 14(7), 2535-2554. https://doi.org/10.3390/molecules14072535