Using Permutations for Hierarchical Clustering of Time Series
Abstract
:1. Introduction and Main Definitions
- (c1)
- (c2)
- if , with .
| –type | –type | … | –type | ||
| –type | … | ||||
| –type | … | ||||
| … | … | … | … | … | … |
| –type | … | ||||
| … |
2. Synthetic Experiments
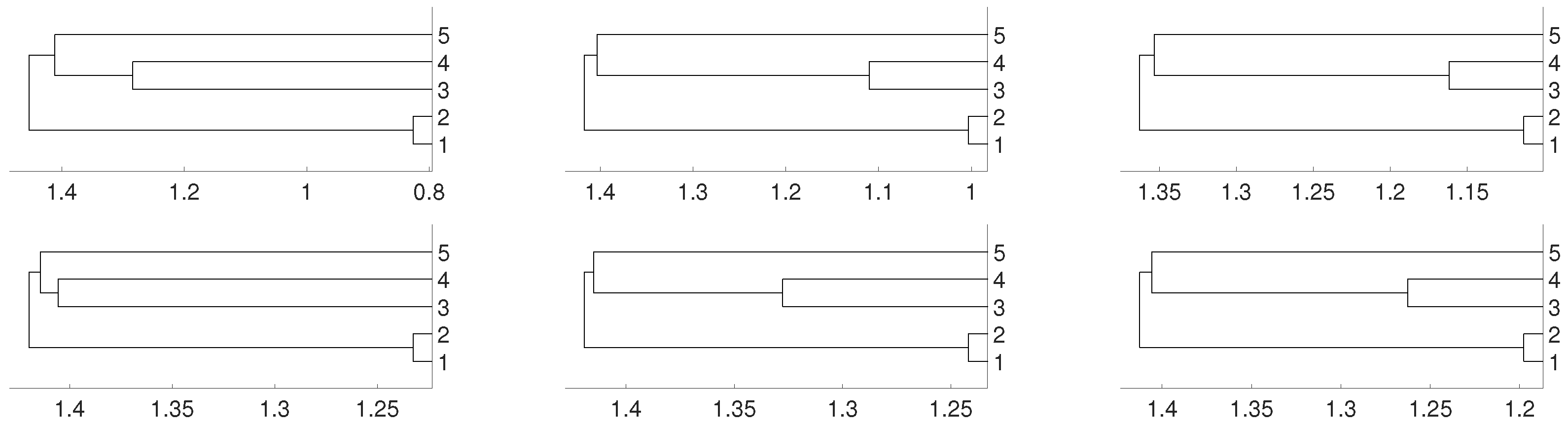
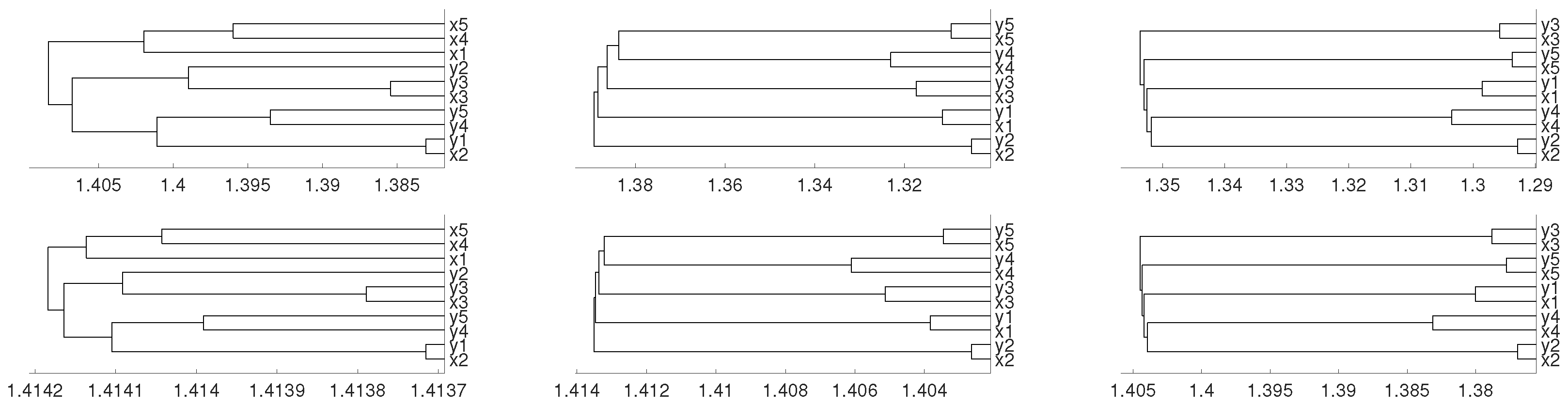
2.1. Linear Dependence
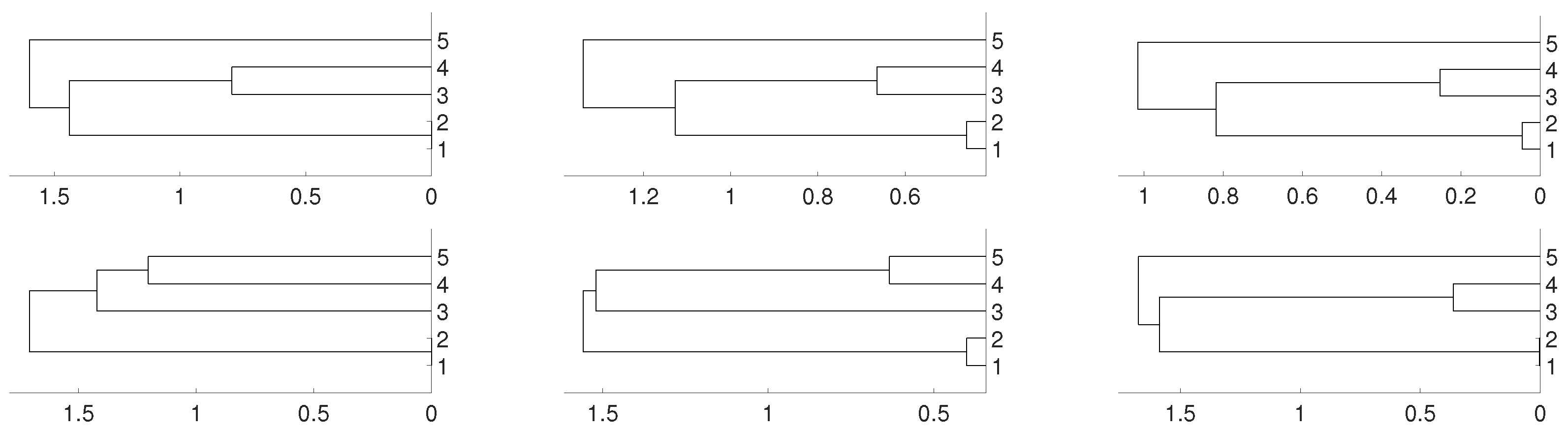
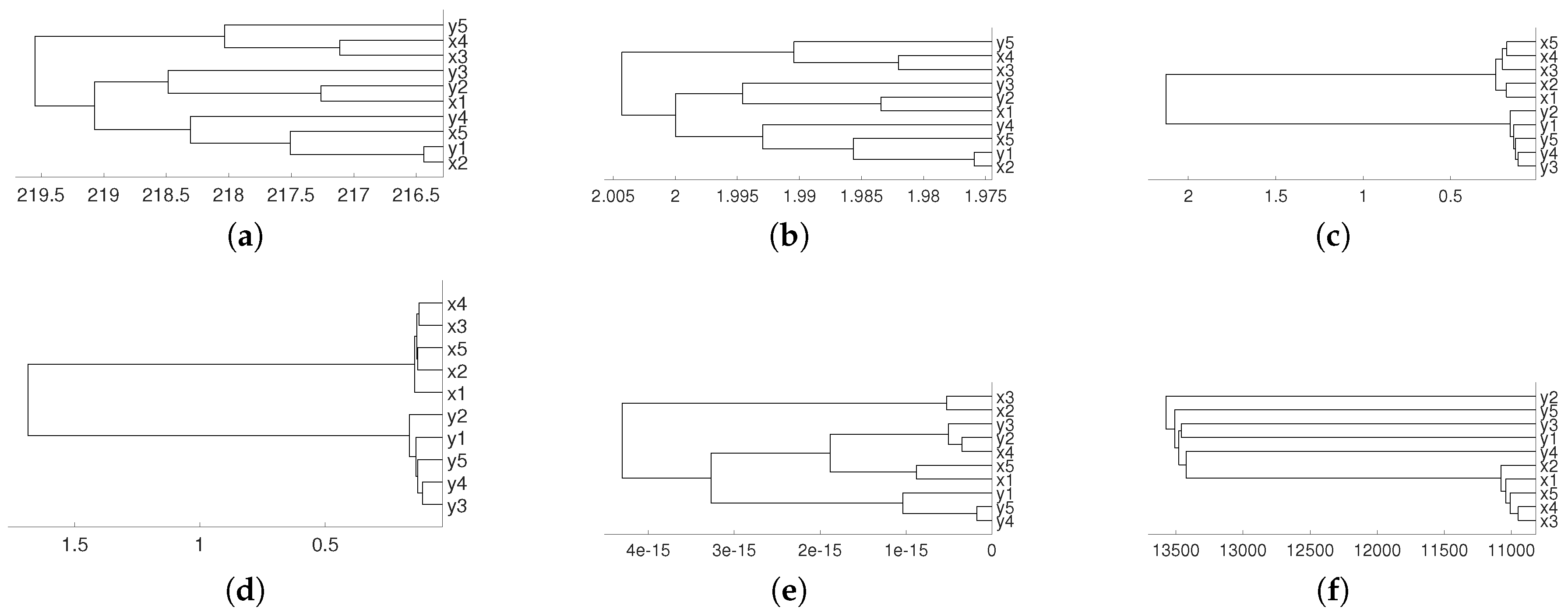
2.2. Non Linear Dependence: Deterministic Systems
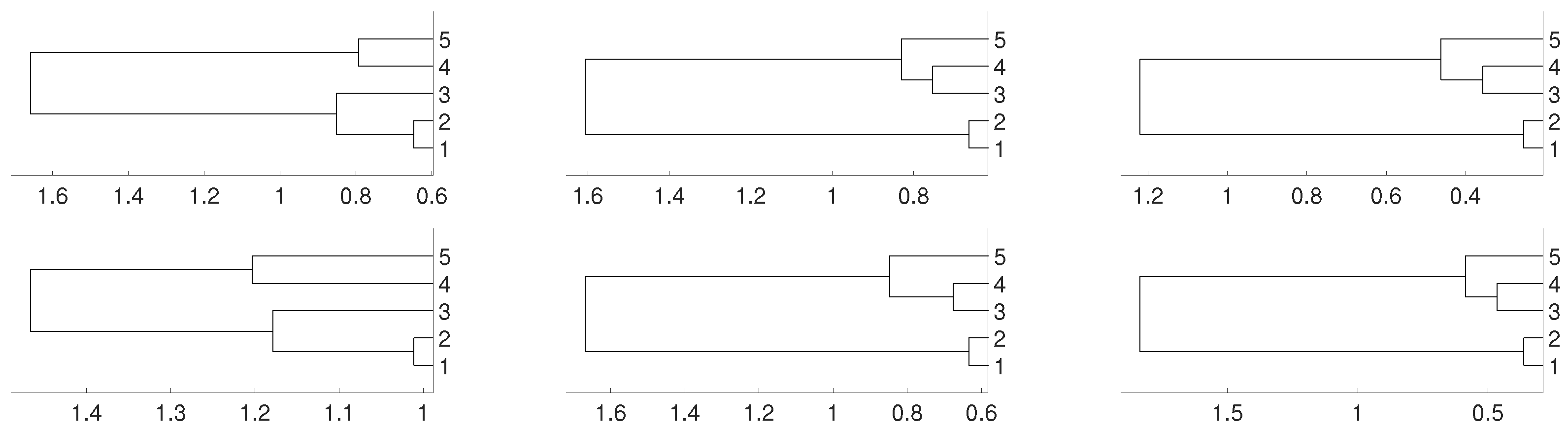
2.3. Non Linear Dependence: Non Deterministic Systems
3. Real Data Experiments
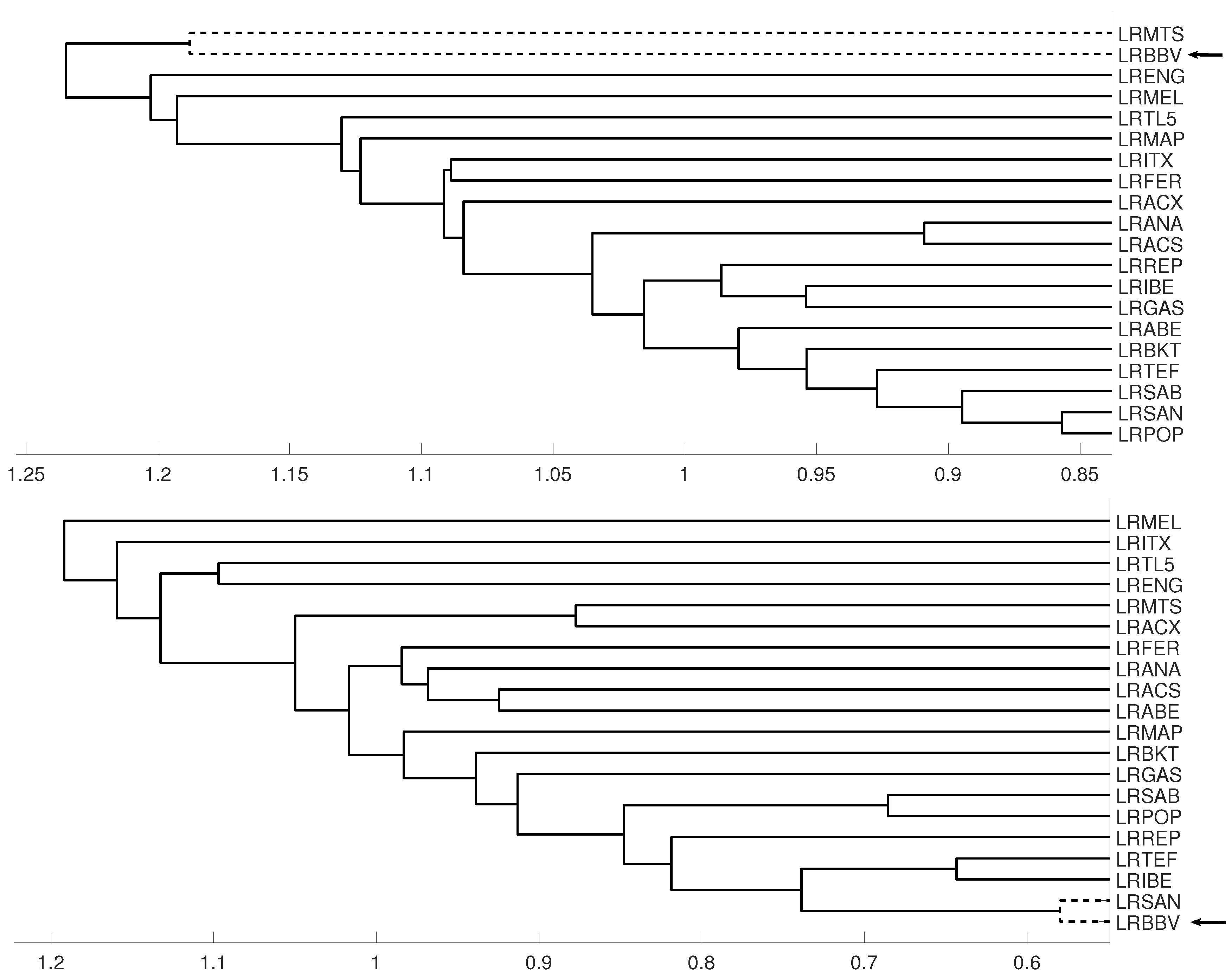
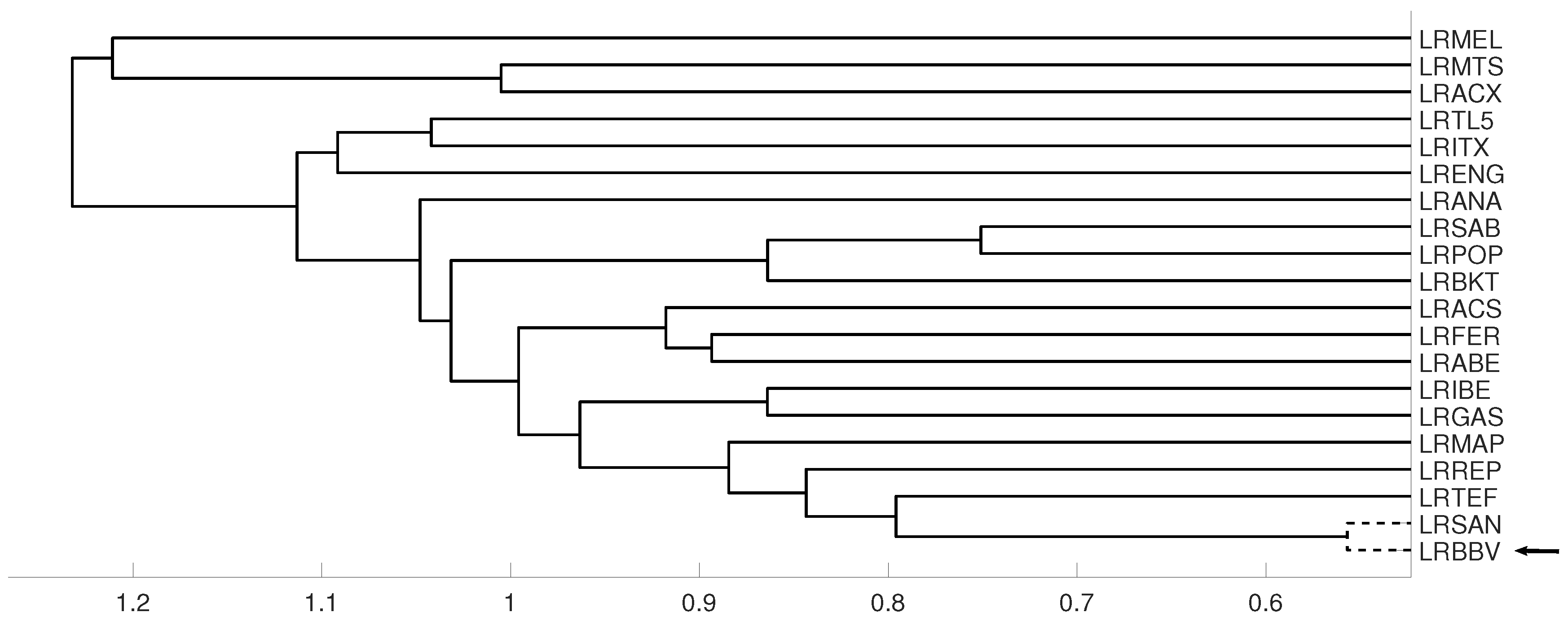
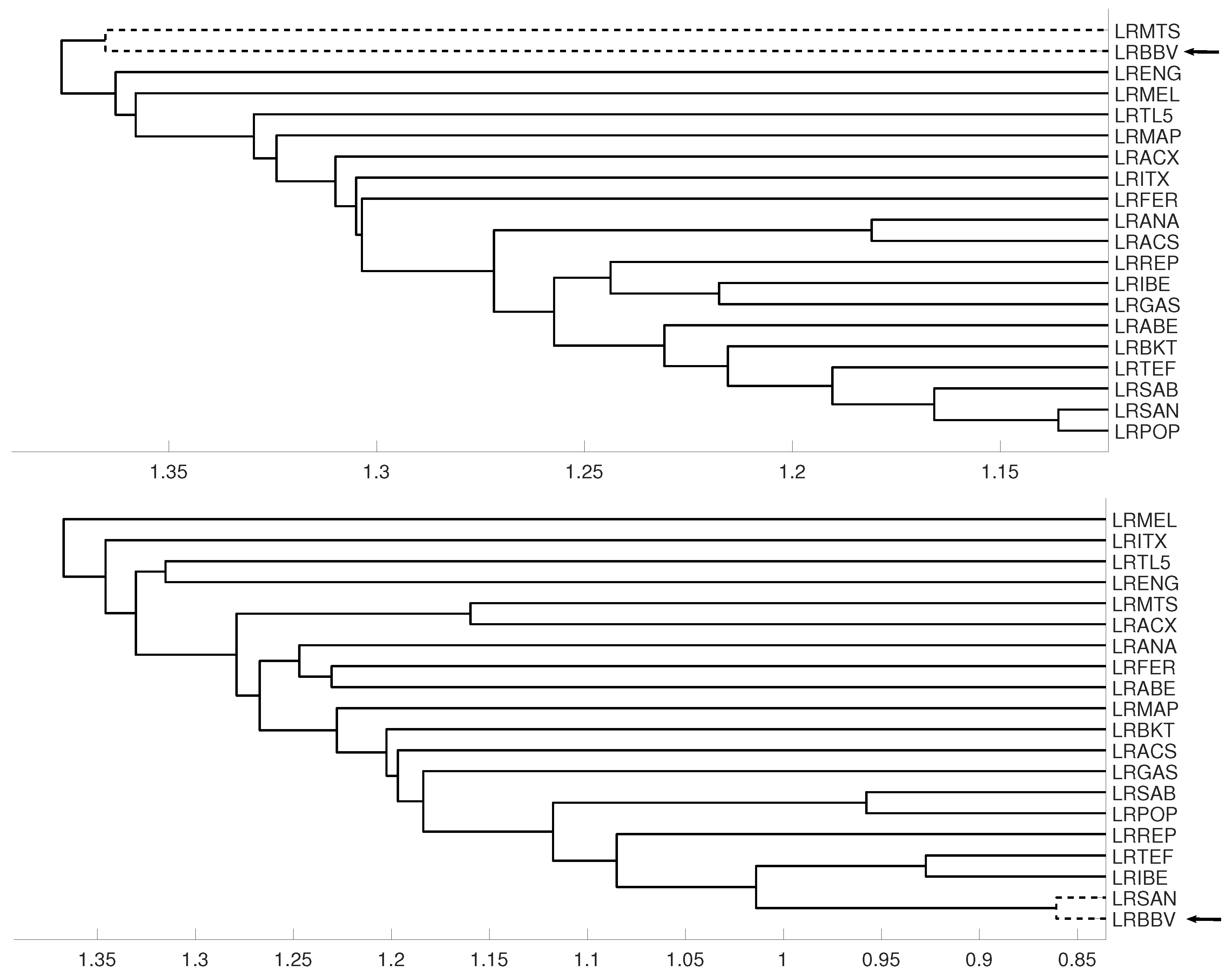
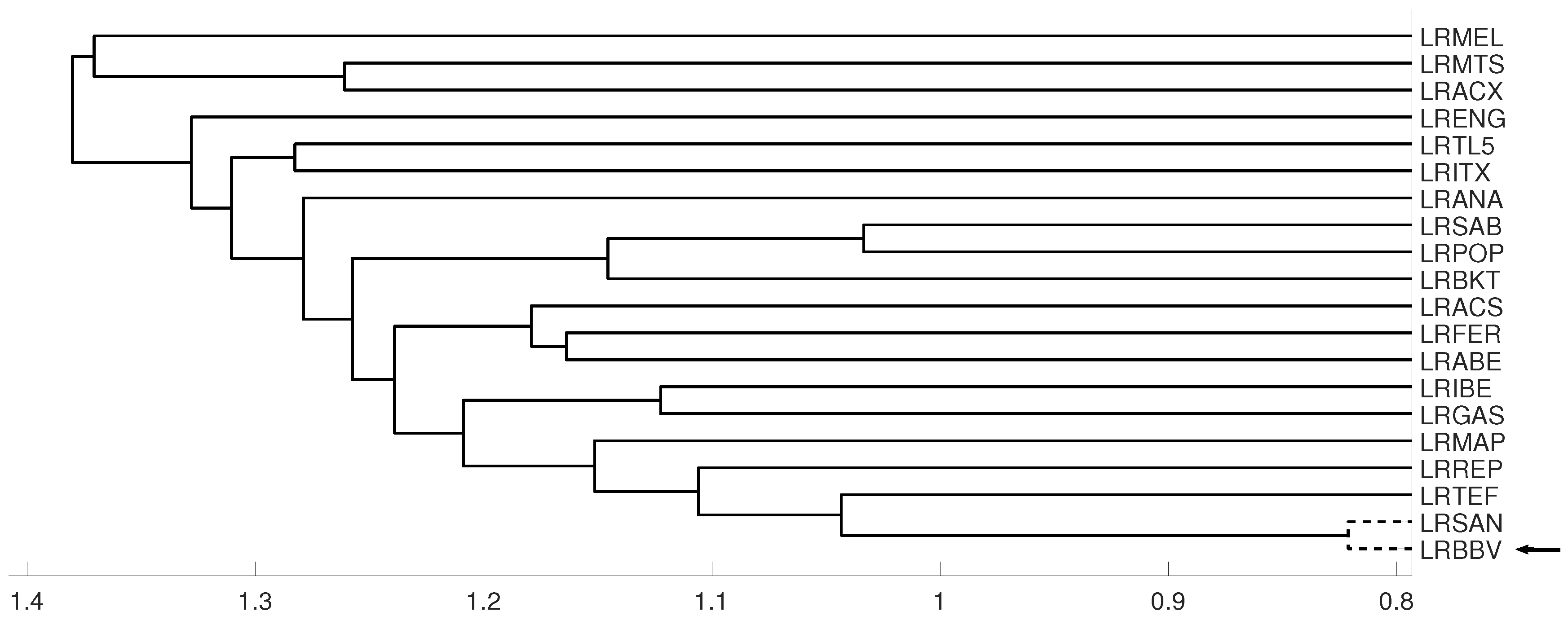
3.1. Latin American Exchange Rate Dependencies
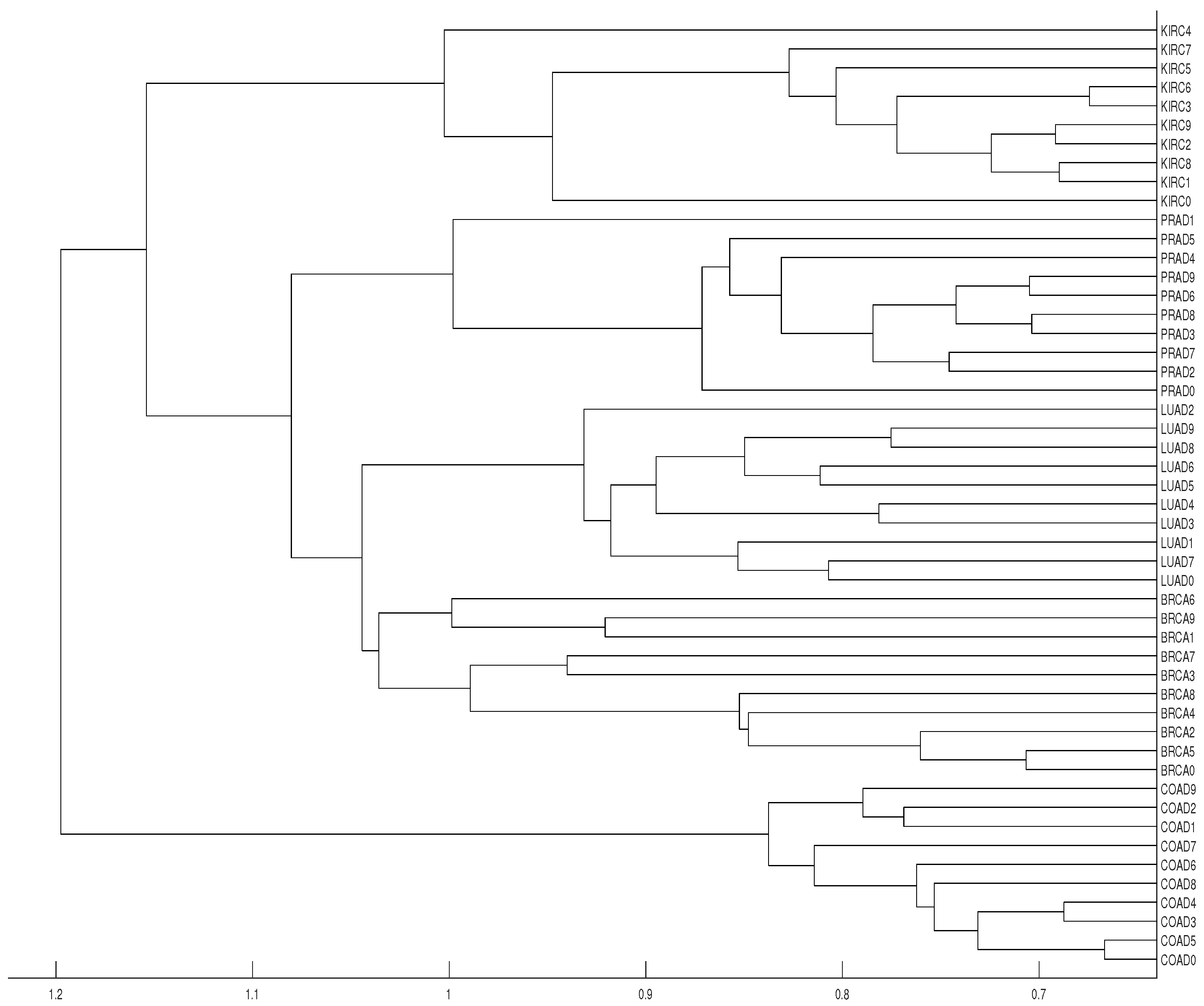
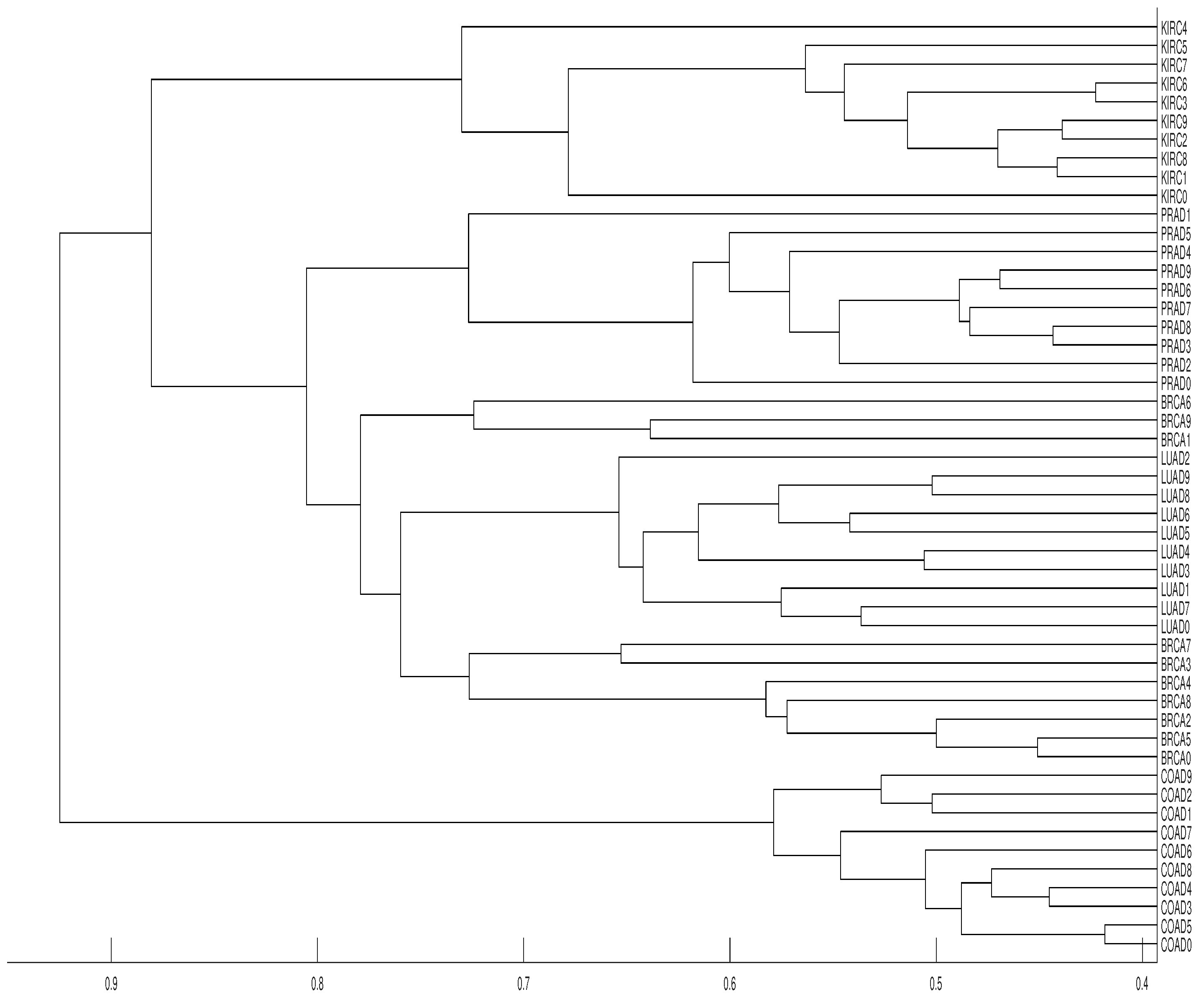
3.2. Tumor Clustering According to RNA Sequences
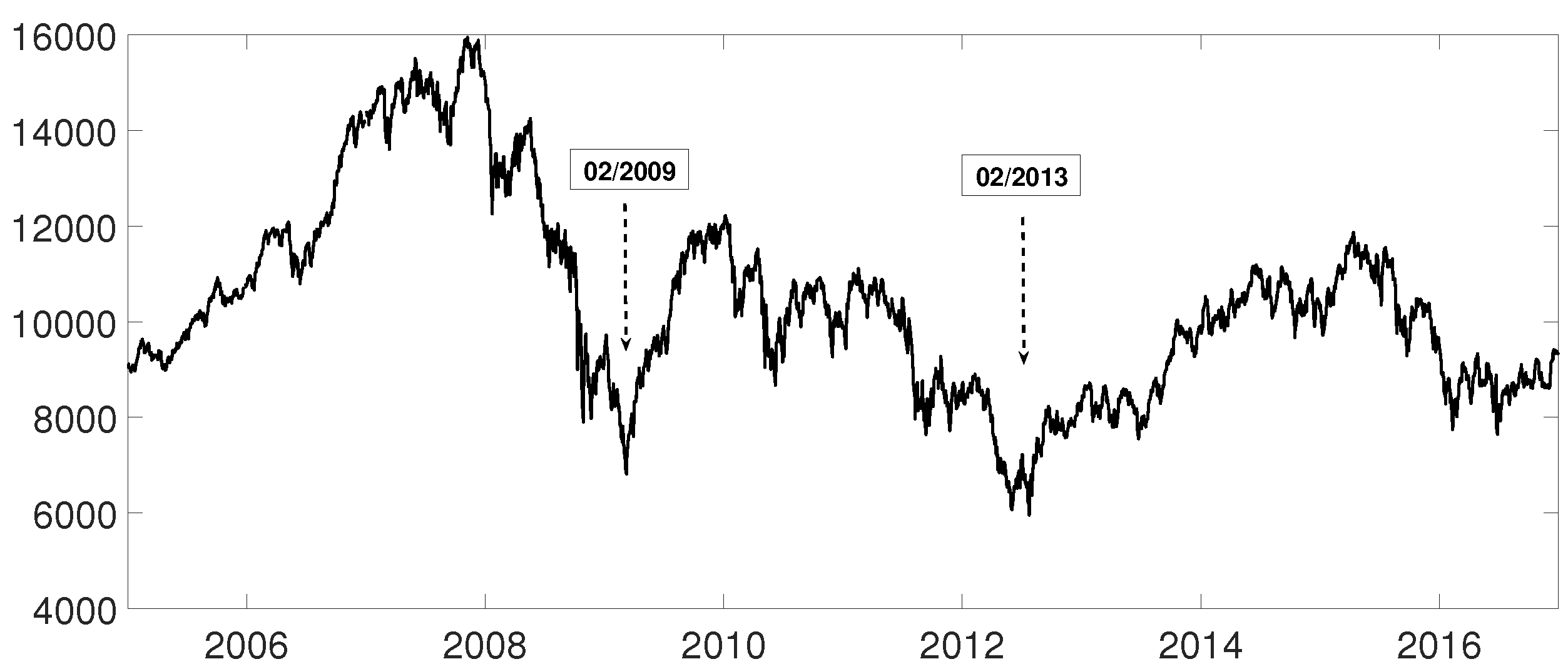
3.3. Evolution of Spanish IBEX35 Banks
4. Conclusions
- The proposed clustering approach is able to detect linear and non-linear dependencies among time series.
- In some cases, a very small embedding dimension like is not enough to detect dependencies among time series, thus a greater embedding dimension is required.
- The distance measure based on the mutual information has revealed a better performance than the distance measure based on the Crammer’s V statistic.
- There are not significant differences with respect to the selected linkage method.
Author Contributions
Funding
Conflicts of Interest
References
- Izakian, H.; Pedrycz, W.; Jamal, I. Fuzzy clustering of time series data using dynamic time warping distance. Eng. Appl. Artif. Intell. 2015, 39, 235–244. [Google Scholar] [CrossRef]
- Möller-Levet, C.S.; Klawonn, F.; Cho, K.H.; Wolkenhauer, O. Fuzzy clustering of short time-series and unevenly distributed sampling points. Adv. Intell. Data Anal. 2003, 330–340. [Google Scholar]
- Foster, E.D. State Space Time Series Clustering Using Discrepancies Based on the Kullback-Leibler Information and the Mahalanobis Distance. Ph.D. Thesis, University of Iowa, Iowa City, IA, USA, 2012. [Google Scholar]
- Zhang, B.; An, B. Clustering time series based on dependence structure. PLoS ONE 2018, 13, e0206753. [Google Scholar] [CrossRef]
- Aghabozorgi, S.; Shirkhorshidi, A.S.; Wah, T.Y. Time-series clustering—A decade review. Inf. Syst. 2105, 53, 16–38. [Google Scholar] [CrossRef]
- Liao, T.W. Clustering of time series data—A survey. Pattern Recognit. 2005, 38, 1857–1874. [Google Scholar] [CrossRef]
- Montero, P.; Vilar, J. TSclust: An R package for time series clustering. J. Stat. Softw. 2014, 62, 1–43. [Google Scholar] [CrossRef]
- Ando, T.; Bai, J. Panel data models with grouped factor structure under unknown group membership. J. Appl. Econ. 2016, 31, 163–191. [Google Scholar] [CrossRef]
- Ando, T.; Bai, J. Clustering huge number of financial time series: A panel data approach with high-dimensional predictor and factor structures. J. Am. Stat. Assoc. 2017, 112, 1182–1198. [Google Scholar] [CrossRef]
- Alonso, A.M.; Peña, D. Clustering time series by linear dependency. Stat. Comput. 2018. [Google Scholar] [CrossRef]
- McClellan, S.; Gibson, J. Spectral entropy: An alternative indicator for rate allocation? In Proceedings of the IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP’94), Adelaide, SA, Australia, 19–22 April 1994; pp. 201–204. [Google Scholar]
- Bruhn, J.; Ropcke, H.; Hoeft, A. Approximate entropy as an electroencephalographic measure of anesthetic drug effect during desflurane anesthesia. Anesthesiology 2000, 92, 715–726. [Google Scholar] [CrossRef]
- Bruzzo, A.A.; Gesierich, B.; Santi, M.; Tassinari, C.A.; Birbaumer, N.; Rubboli, G. Permutation entropy to detect vigilance changes and preictal states from scalp EEG in epileptic patients. A preliminary study. Neurol. Sci. 2008, 29, 3–9. [Google Scholar] [CrossRef] [PubMed]
- Olofsen, E.; Sleigh, J.W.; Dahan, A. Permutation entropy of the electroencephalogram: A measure of anaesthetic drug effect. Br. J. Anaesth. 2008, 6, 810–821. [Google Scholar] [CrossRef] [PubMed]
- Quian-Quiroga, R.; Blanco, S.; Rosso, O.A.; Garcıa, H.; Rabinowicz, A. Searching for hidden information with Gabor transform in generalized tonic–clonic seizures, Electroencephalography and Clinical. Neurophysiology 1997, 103, 434–439. [Google Scholar]
- Cánovas, J.S.; Garcia-Clemente, G.; Mu noz-Guillermo, M. Comparing permutation entropy functions to detect structural changes in time series. Phys. A 2018, 507, 153–174. [Google Scholar] [CrossRef]
- Parlitz, U.; Berg, S.; Luther, S.; Schirdewan, A.; Kurths, J.; Wessel, N. Classifying cardiac biosignals using ordinal pattern statistics and symbolic dynamics. Comput. Biol. Med. 2012, 42, 319–327. [Google Scholar] [CrossRef]
- Sinn, M.; Keller, K.; Chen, B. Segmentation and classification of time series using ordinal pattern distributions. Eur. Phys. J. Spec. Top. 2013, 222, 587–598. [Google Scholar] [CrossRef]
- Echegoyen, I.; Vera-Avila, V.; Sevilla-Escoboza, R.; Martinez, J.H.; Buldua, J.M. Ordinal synchronization: Using ordinal patterns to capture interdependencies between time series. Chaos Solitons Fractals 2019, 119, 8–18. [Google Scholar] [CrossRef]
- Ruiz-Abellón, M.C.; Guillamón, A.; Gabaldón, A. Dependency-aware clustering of time series and its application on Energy Markets. Energies 2016, 9, 809. [Google Scholar] [CrossRef]
- Bandt, C.; Pompe, B. Permutation entropy—A natural complexity measure for time series. Phys. Rev. Lett. 2002, 88, 174102. [Google Scholar] [CrossRef]
- Amigó, J.M.; Zambrano, S.; Sanjuán, M.A.F. True and false forbidden patterns in deterministic and random dynamics. Europhys. Lett. EPL 2007, 79, 50001. [Google Scholar] [CrossRef]
- Amigó, J.M.; Kennel, M.B. Topological permutation entropy. Phys. D Nonlinear Phenom. 2007, 231, 137–142. [Google Scholar] [CrossRef]
- Matilla, M.; Ruíz, M. A non–parametric independence test using permutation entropy. J. Econom. 2008, 144, 139–155. [Google Scholar] [CrossRef]
- Bandt, C.; Keller, G.; Pompe, B. Entropy of interval maps via permutations. Nonlinearity 2002, 15, 1595–1602. [Google Scholar] [CrossRef]
- Cánovas, J.S. Estimating topological entropy from individual orbits. Int. J. Comput. Math. 2009, 86, 1901–1906. [Google Scholar] [CrossRef]
- Cánovas, J.S.; Guillamón, A.; del Ruíz, M. Using permutations to detect dependence between time series. Phys. D Nonlinear Phenom. 2011, 240, 1199–1204. [Google Scholar] [CrossRef]
- Wallis, S. Measures of Association for Contingency Tables; University College London: London, UK, 2012. [Google Scholar]
- Kraskov, A.; Stögbauer, H.; Andrzejak, R.G.; Grassberger, P. Hierarchical Clustering Based on Mutual Information. arXiv, 2005; arXiv:q-bio/0311039. [Google Scholar]
- Walters, P. An Introduction to Ergodic Theory; Springer: New York, NY, USA, 1982. [Google Scholar]
- Herrera, A.R. Analysis of dispersal effects in metapopulation models. J. Math. Biol. 2016, 72, 683–698. [Google Scholar] [CrossRef] [PubMed]
- Gómez-González, J.E.; Melo-Velandia, L.F.; Maya, R.L. Latin American Exchange Rate Dependencies: A Regular Vine Copula Approach. Contemp. Econ. Policy 2015, 33, 535–549. [Google Scholar]

© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cánovas, J.S.; Guillamón, A.; Ruiz-Abellón, M.C. Using Permutations for Hierarchical Clustering of Time Series. Entropy 2019, 21, 306. https://doi.org/10.3390/e21030306
Cánovas JS, Guillamón A, Ruiz-Abellón MC. Using Permutations for Hierarchical Clustering of Time Series. Entropy. 2019; 21(3):306. https://doi.org/10.3390/e21030306
Chicago/Turabian StyleCánovas, Jose S., Antonio Guillamón, and María Carmen Ruiz-Abellón. 2019. "Using Permutations for Hierarchical Clustering of Time Series" Entropy 21, no. 3: 306. https://doi.org/10.3390/e21030306
APA StyleCánovas, J. S., Guillamón, A., & Ruiz-Abellón, M. C. (2019). Using Permutations for Hierarchical Clustering of Time Series. Entropy, 21(3), 306. https://doi.org/10.3390/e21030306




