LSTM-CRF for Drug-Named Entity Recognition
Abstract
:1. Introduction
2. LSTM-CRF Model
2.1. LSTM
2.2. CRF Model
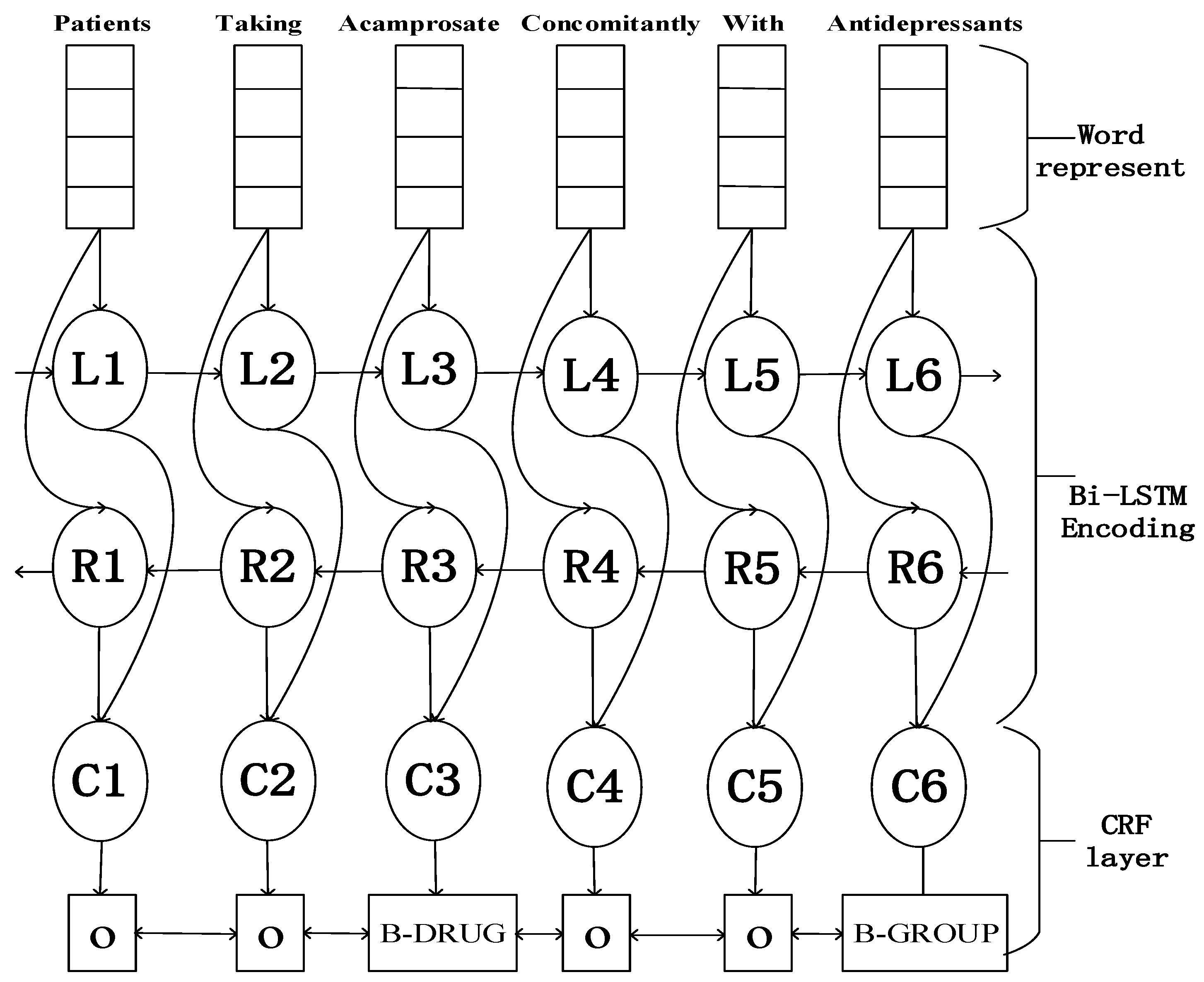
2.3. LSTM-CRF
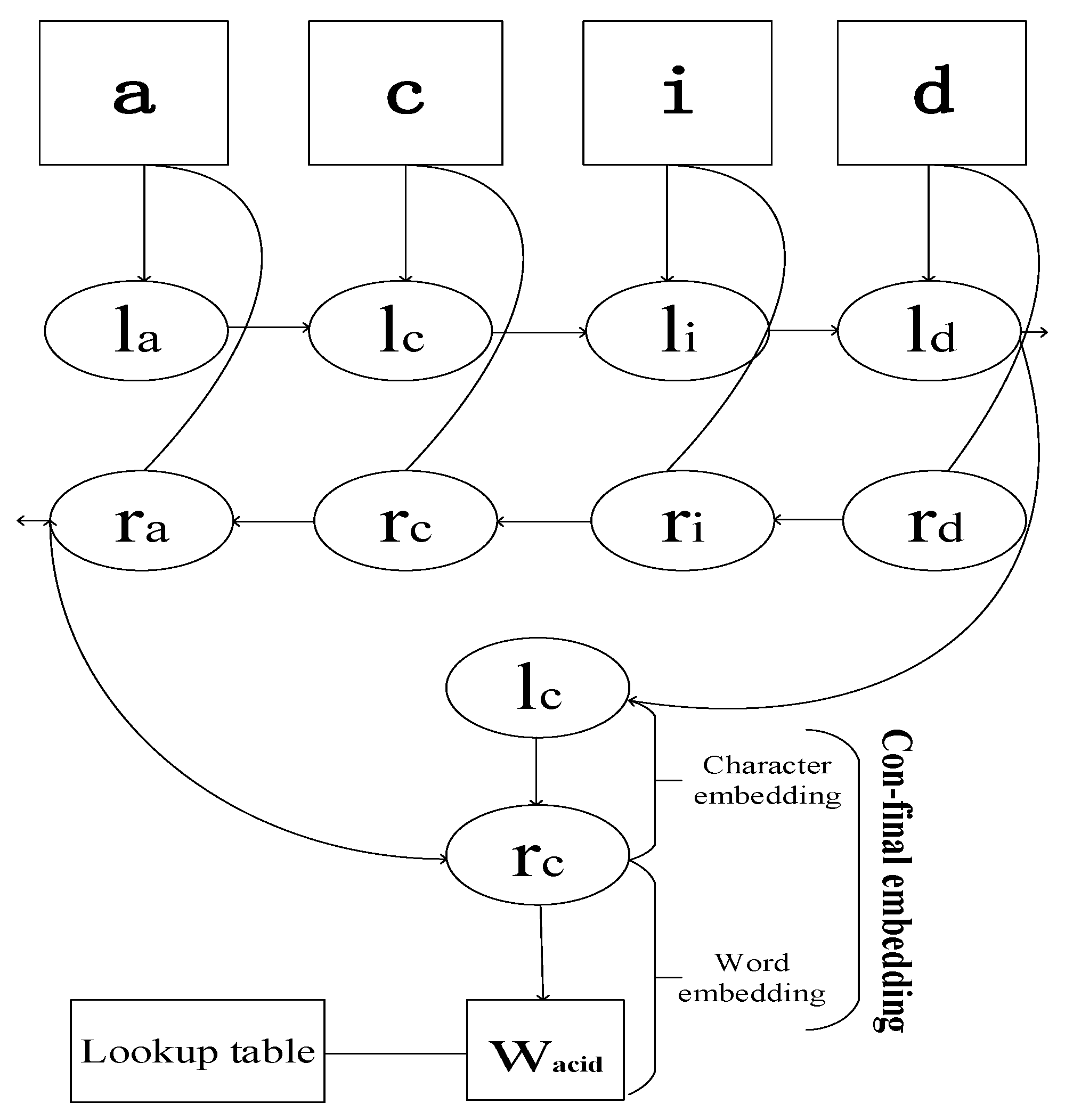
2.4. Embedding and Network Training
2.4.1. Parameters Initialization
2.4.2. Optimization Method
2.4.3. Parameters Adjustment
2.5. IOBES Tagging Scheme
3. Experiments
3.1. Data Sets
3.2. Evaluation Metrics
3.3. Results
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Segura-Bedmar, I.; Martínez, P.; Pablo-Sánchez, C. Using a shallow linguistic kernel for drug-drug interaction extraction. J. Biomed. Inform. 2011, 44, 789–804. [Google Scholar] [CrossRef]
- Warrer, P.; Hansen, W.; Juhl-Jensen, L.; Aagaard, L. Using text-mining techniques in electronic patient records to identify ADRs from medicine use. Br. J. Clin. Pharmacol. 2012, 73, 674–684. [Google Scholar] [CrossRef]
- Segura-Bedmar, I.; Suárez-Paniagua, V.; Martínez, P. Exploring word embedding for drug name recognition. In Proceedings of the International Workshop on Health Text Mining and Information Analysis, Lisbon, Portugal, 17 September 2015; pp. 64–72. [Google Scholar]
- Li, K.; Ai, W.; Tang, Z.; Zhang, F.; Jiang, L.; Li, K.; Hwang, K. Hadoop recognition of biomedical named entity using conditional random fields. IEEE Trans. Parallel Distrib. Syst. 2015, 26, 3040–3051. [Google Scholar] [CrossRef]
- Kazama, J.; Makino, T.; Ohta, Y.; Tsujii, J. Tuning support vector machines for biomedical named entity recognition. In Proceedings of the ACL Workshop on Natural Language Processing in the Biomedical Domain, Philadelphia, PA, USA, 11 July 2002; pp. 1–8. [Google Scholar]
- Saha, S.K.; Sarkar, S.; Mitra, P. Feature selection techniques for maximum entropy based biomedical named entity recognition. J. Biomed. Inform. 2009, 42, 905–911. [Google Scholar] [CrossRef]
- Lin, Y.; Tsai, T.; Chou, W.; Wu, K.; Sung, T.; Hsu, W. A maximum entropy approach to biomedical named entity recognition. In Proceedings of the 4th International Conference on Data Mining in Bioinformatics, Seattle, WA, USA, 28 June 2004; pp. 56–61. [Google Scholar]
- Finkel, J.R.; Grenager, T.; Manning, C.D. Incorporating non-local information into information extraction systems by Gibbs sampling. In Proceedings of the 43rd Annual Meeting on Association for Computational Linguistics, Ann Arbor, MI, USA, 25–30 June 2005; pp. 363–370. [Google Scholar]
- Tkachenko, M.; Simanovsky, A. Named entity recognition: Exploring features. In Proceedings of the 11th Conference on Natural Language Processing, Vienna, Austria, 19–21 September 2012; pp. 118–127. [Google Scholar]
- Zeng, D.; Sun, C.; Lin, L.; Liu, B. Enlarging drug dictionary with semi-supervised learning for Drug Entity Recognition. In Proceedings of the IEEE International Conference on Bioinformatics and Biomedicine, Shenzhen, China, 15–18 December 2016; pp. 1929–1931. [Google Scholar]
- Eriksson, R.; Jensen, P.B.; Frankild, S.; Jensen, L.J.; Brunak, S. Dictionary construction and identification of possible adverse drug events in Danish clinical narrative text. J. Am. Med. Inform. Assoc. 2013, 20, 947–953. [Google Scholar] [CrossRef]
- Chalapathy, R.; Borzeshi, E.Z.; Piccardi, M. An investigation of recurrent neural architectures for drug name recognition. arXiv, 2016; arXiv:1609.07585. [Google Scholar]
- Hochreiter, S.; Schmidhuber, J. Long short-term memory. Neural Comput. 1997, 9, 1735–1780. [Google Scholar] [CrossRef]
- Hammerton, J. Named entity recognition with long short-term memory. In Proceedings of the 7th conference on Natural language learning, Edmonton, AB, Canada, 31 May–1 June 2003; pp. 172–175. [Google Scholar]
- Huang, Z.; Xu, W.; Yu, K. Bidirectional LSTM-CRF models for sequence tagging. arXiv, 2015; arXiv:1508.01991. [Google Scholar]
- Dos Santos, C.N.; Zadrozny, B. Learning character-level representations for part-of-speech tagging. In Proceedings of the 31st International Conference on Machine Learning, Beijing, China, 21–26 June 2014; pp. 1818–1826. [Google Scholar]
- Rumelhart, D.; Hinton, G.; Williams, R. Learning representations by back-propagating errors. Nature 1986, 323, 533–536. [Google Scholar] [CrossRef]
- Elman, J. Finding structure in time. Cogn. Sci. 1990, 14, 179–211. [Google Scholar] [CrossRef]
- Werbos, P.J. Generalization of backpropagation with application to a recurrent gas market model. Neural Netw. 1998, 1, 339–356. [Google Scholar] [CrossRef]
- Bengio, Y.; Simard, P.; Frasconi, P. Learning long-term dependencies with gradient descent is difficult. IEEE Trans. Neural Netw. 1994, 5, 157–166. [Google Scholar] [CrossRef]
- Graves, A.; Schmidhuber, J. Framewise phoneme classification with bidirectional LSTM networks. In Proceedings of the 2005 International Joint Conference on Neural Networks, Montreal, QC, Canada, 31 July–4 Aug 2005; pp. 2047–2052. [Google Scholar]
- Bergstra, J.; Breuleux, O.; Bastien, F.; Lamblin, P.; Pascanu, R.; Desjardins, G.; Turian, J.; Bengio, Y. Theano: A CPU and GPU math expression compiler. In Proceedings of the Python for Scientific Computing Conference, Austin, TX, USA, 28 June–3 July 2010; pp. 3–10. [Google Scholar]
- Collobert, R.; Weston, J.; Bottou, L.; Karlen, M.; Kavukcuoglu, K.; Kuksa, P.P. Natural language processing (almost) from scratch. J. Mach. Learn. Res. 2011, 12, 2493–2537. [Google Scholar]
- Wikimedia Downloads. Available online: http://download.wikimedia.org (accessed on 11 June 2017).
- Ling, W.; Marujo, T.; Astudillo, F.; Amir, S.; Dyer, C.; Black, A.W.; Trancoso, I. Finding function in form: Compositional character models for open vocabulary word representation. In Proceedings of the Conference on Empirical Methods in Natural Language Processing, Lisbon, Portugal, 17–21 September 2015; pp. 1520–1530. [Google Scholar]
- Mikolov, T.; Chen, K.; Corrado, G.; Dean, J. Efficient estimation of word representations in vector space. arXiv, 2013; arXiv:1301.3781. [Google Scholar]
- Lample, G.; Ballesteros, M.; Subramanian, S.; Kawakami, K.; Dyer, C. Neural architectures for named entity recognition. arXiv, 2016; arXiv:1603.01360. [Google Scholar]
- Pascanu, R.; Mikolov, T.; Bengio, Y. On the difficulty of training recurrent neural networks. arXiv, 2012; arXiv:1211.5063. [Google Scholar]
- Zeiler, M.D. ADADELTA: An adaptive learning rate method. arXiv, 2015; arXiv:1212.5701. [Google Scholar]
- Kingma, D.; Ba, J. Adam: A method for stochastic optimization. arXiv, 2014; arXiv:1412.6980. [Google Scholar]
- Hinton, G.; Srivastava, N.; Swersky, K. RMSProp: Divide the gradient by a running average of its recent magnitude. Available online: http://www.cs.toronto.edu/~tijmen/csc321/slides/lecture_slides_lec6.pdf (accessed on 10 June 2017).
- Ma, X.; Hovy, E.H. End-to-end sequence labeling via bi-directional LSTM-CNNs-CRF. arXiv, 2016; arXiv:1603.01354. [Google Scholar]
- Srivastava, N.; Hinton, G.; Krizhevsky, A.; Sutskever, I.; Salakhutdinov, R. Dropout: A simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 2014, 15, 1929–1958. [Google Scholar]
- Dos Santos, C.N.; Guimarães, V. Boosting named entity recognition with neural character embeddings. arXiv, 2015; arXiv:1505.05008. [Google Scholar]
- Segura-Bedmar, I.; Martinez, P.; Sanchez-Cisneros, D. The 1st DDIExtraction-2011 challenge task: Extraction of drug-drug interactions from biomedical texts. In Proceedings of the 1st Challenge Task on Drug-Drug Interaction Extraction, Huelva, Spain, 7 September 2011; pp. 1–9. [Google Scholar]
- Segura-Bedmar, I.; Martínez, P.; Herrero-Zazo, M. Lessons learnt from the DDIExtraction-2013 shared task. J. Biomed. Inform. 2014, 51, 152–164. [Google Scholar] [CrossRef] [PubMed]
- He, L. Drug Name Recognition and Drug–Drug Interaction Extraction Based on Machine Learning. Master’s Thesis, Dalian University of Technology, Dalin, China, 2013. [Google Scholar]
- Segura-Bedmar, I.; Martínez, P.; Herrero-Zazo, M. Semeval-2013 task 9: Extraction of drug-drug interactions from biomedical texts. In Proceedings of the 7th International Workshop on Semantic Evaluation, Atlanta, GA, USA, 14–15 June 2013; pp. 341–350. [Google Scholar]
- Liu, S.; Tang, B.; Chen, Q.; Wang, X. Effects of semantic features on machine learning-based drug name recognition systems: Word embeddings vs. manually constructed dictionaries. Information 2015, 6, 848–865. [Google Scholar] [CrossRef]
- Grego, T.; Pinto, F.; Couto, F.M. LASIGE: Using conditional random fields and ChEBI ontology. In Proceedings of the 7th International Workshop on Semantic Evaluation, Atlanta, GA, USA, 14–15 June 2013; pp. 660–666. [Google Scholar]
- Collazo, A.; Ceballo, A.; Puig, D.; Gutiérrez, Y.; Abreu, J.; Pérez, R.; Orquín, A.; Montoyo, A.; Muñoz, R.; Camara, F. Semantic and lexical features for detection and classification drugs in biomedical texts. In Proceedings of the 7th International Workshop on Semantic Evaluation, Atlanta, GA, USA, 14–15 June 2013; pp. 636–643. [Google Scholar]
| Case Letters | a|b|c|d|e|f|g|h|i|j|k|l|m|n|o|p|q|r|s|t|u|v|w|x|y|z | 26 |
| Numbers | 0|1|2|3|4|5|6|7|8|9 | 10 |
| Punctuations | $|!|@|#|%|&|*|-|=|_|+|(|)|||[|]|;|’|:|"|,|.|/|<|>|?|`|’| |·|| | 33 |
| Hyper-Parameter | Values |
|---|---|
| initial state | 0.0 |
| dropout rate | 0.5 |
| initial learning rate | 0.01 |
| gradient clipping | 5.0 |
| word_dim | 100 |
| forward_char_dim | 25 |
| backward_char_dim | 25 |
| char_dim | 50 |
| con-final embedding | 150 |
| Set | Documents | Sentences | Drugs |
|---|---|---|---|
| Training | 435 | 4267 | 11,260 |
| Final Test | 144 | 1539 | 3689 |
| Total | 579 | 5806 | 14,949 |
| Type | DrugBank | MedLine | Total |
|---|---|---|---|
| Drug | 9901 (63%) | 1745 (63%) | 11,646 (63%) |
| Brand | 1824 (12%) | 42 (1.5%) | 1866 (10%) |
| Group | 3901 (25%) | 324 (12%) | 4225 (23%) |
| Drug_n | 130 (1%) | 635 (23%) | 765 (4%) |
| Total | 15,756 | 2746 | 18,502 |
| Type | DrugBank | MedLine | Total |
|---|---|---|---|
| Drug | 180 (59%) | 171 (44%) | 351 (51%) |
| Brand | 53 (18%) | 6 (2%) | 59 (8%) |
| Group | 65 (21%) | 90 (24%) | 155 (23%) |
| Drug_n | 6 (2%) | 115 (30%) | 121 (18%) |
| Total | 304 | 382 | 686 |
| System | Precision | Recall | F1 |
|---|---|---|---|
| Our System | 93.26% (%) | 91.11% (%) | 92.04% (%) |
| Best without Dic | 92.15% | 89.73% | 90.92% |
| Best with Dic | 94.75% | 90.44% | 92.54% |
| Type | Precision | Recall | F1 |
|---|---|---|---|
| Drug | 85.03% | 80.03% | 81.87% |
| Brand | 88.24% | 77.42% | 81.83% |
| Group | 86.01% | 88.59% | 86.86% |
| Drug_n | 78.39% | 57.26% | 62.83% |
| Micro-Average | 83.62% | 77.81% | 79.26% |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zeng, D.; Sun, C.; Lin, L.; Liu, B. LSTM-CRF for Drug-Named Entity Recognition. Entropy 2017, 19, 283. https://doi.org/10.3390/e19060283
Zeng D, Sun C, Lin L, Liu B. LSTM-CRF for Drug-Named Entity Recognition. Entropy. 2017; 19(6):283. https://doi.org/10.3390/e19060283
Chicago/Turabian StyleZeng, Donghuo, Chengjie Sun, Lei Lin, and Bingquan Liu. 2017. "LSTM-CRF for Drug-Named Entity Recognition" Entropy 19, no. 6: 283. https://doi.org/10.3390/e19060283
APA StyleZeng, D., Sun, C., Lin, L., & Liu, B. (2017). LSTM-CRF for Drug-Named Entity Recognition. Entropy, 19(6), 283. https://doi.org/10.3390/e19060283






