1. Background
Pathological brain detection (PBD) is of essential importance. It can help physicians make decisions, and to avoid wrong judgements on subjects’ condition. Magnetic resonance imaging (MRI) features high-resolution on soft tissues in the subjects’ brains, generating a mass dataset [
1]. At present, there are numerous works on the use of brain magnetic resonance (MR) images for solving PBD problems [
2,
3].
Due to the enormous volume of the imaging dataset from the human brain, traditional manual techniques are either tedious, or time-consuming, or costly. Therefore, it is necessary to develop a novel computer-aided diagnosis (CAD) system [
4] to help patients have enough time to receive treatment.
In the last decade, many methods from different countries were presented with the same goal of detecting pathological brains [
5,
6,
7,
8] (more references will be introduced in
Section 2). Most of them have two stages: (1) Feature extraction, to extract efficient features that can distinguish pathological brain from healthy brains; feature reduction can be skipped if the size of features dataset is reasonable; and (2) Classification, to construct a classifier using the extracted (and reduced) features.
For the first stage of feature extraction, the latest solutions transform the brain image by discrete wavelet transform (DWT) [
9], which is proven to be superior to the traditional Fourier transform. Nevertheless, DWT comes across a problem of choosing the best decomposition level and the optimal wavelet function.
For the final stage of classification, recent approaches like to use feed-forward neural network (FNN) or support vector machine (SVM), which are becoming popular in the fields of classification and detection [
10]. However, SVM has the limitation that its hyperplanes should be parallel. This parallelism restrains the classification perform.
To solve the above two problems, we propose two improvements: on the one hand, we propose a novel image feature—Fractional Fourier Entropy (FRFE)—which is based on two steps: (1) the use of a Fraction Fourier Transform (FRFT) to replace the traditional Fourier transform; and (2) Shannon entropy to extract features from the FRFT spectrums.
On the other hand, we suggest removing the hyperplane parallelism restraint [
11]. We introduce for this purpose two non-parallel SVMs, the generalized eigenvalue proximal SVM (GEPSVM) and twin support vector machine (TSVM).
In the remainder of this paper
Section 2 offers the state-of-the-art. Then,
Section 3 describes the materials used.
Section 4 presents the extracted features and how to select important features.
Section 5 offers the mechanisms of the standard support vector machine and non-parallel support vector machine.
Section 6 covers the experimental design.
Section 7 provides the results. Discussions are presented in
Section 8. Finally
Section 9 is devoted to our conclusions.
2. State-of-the-Art
Recent PBD methods are of two types. One treats a 3D dataset as a whole, and the other selects the most important slice from the 3D data. The former needs to scan the whole brain, which is expensive and time-consuming. The latter only needs to scan the focus related slice, which is cheap and rapid. In this study, we focus on the latter.
Chaplot
et al. [
12] were the first to apply “DWT” to PBD problems. Their classifiers are SVM and self-organizing map (SOM). El-Dahshan
et al. [
13] employed a 3-level discrete wavelet transform. Dong
et al. [
14] proposed the use of a novel scaled conjugate gradient (SCG) approach for PBD. Zhang and Wu [
15] proposed to employ kernel support vector machine (KSVM). Saritha
et al. [
16] combined wavelet transform with Shannon entropy, and they named the novel feature wavelet-entropy (WE). They harnessed spider-web plots (SWPs) with the aim of decreasing the number of WEs. They employed the probabilistic NN (PNN) for classification. Zhang
et al. [
17,
18] found spider-web plots had no effect on PBD. Das
et al. [
19] suggested using a Ripplet transform (RT) in PBD. Their classifier is a least squares support vector machine (LS-SVM). Zhang
et al. [
20] employed particle swarm optimization (PSO) to find the optimal parameters in a kernel support vector machine. El-Dahshan
et al. [
21] proposed the use of a feedback pulse-coupled neural network to segment brain images. Following Saritha’s work, Zhou
et al. [
22] employed WE, and Naive Bayes classifier (NBC) as the classifier. Zhang
et al. [
23] employed a discrete wavelet packet transform (DWPT) to replace DWT, and employed Tsallis entropy (TE) to replace Shannon entropy (SE). Yang
et al. [
24] employed wavelet-energy as the features. Damodharan and Raghavan [
25] used tissue segmentation to detect neoplasms in brains. Guang-Shuai
et al. [
26] employed both wavelet-energy and SVM. The overall accuracy was less than 83%. Wang
et al. [
27] employed genetic algorithm (GA) to solve the task of PBD. Nazir
et al. [
28] proposed performing image denoising first. Their overall accuracy was higher than 91%. Harikumar and Kumar [
29] used ANN with optimal performance achieved by use of a db4 wavelet and radial basis function (RBF) kernel. Wang
et al. [
30] suggested the use of a stationary wavelet transform (SWT). Zhang
et al. [
31] offered a new Hybridization of Biogeography-based optimization (BBO) and the Particle swarm optimization (PSO) method. Hence, they termed it HBP for short. Farzan
et al. [
32] used the longitudinal percentage of brain volume changes (PBVC) in a two-year follow up and its intermediate counterparts in early 6-month and late 18-month tests as features. Their experimental results showed SVM with RBF performed the best with an accuracy of 91.7%, higher than K-means at 83.3%, fuzzy c-means (FCM) at 83.3%, and linear SVM at 90%. Zhang
et al. [
33] used two types of features: one is Hu moment invariants (HMI); the other is wavelet entropy. Munteanu
et al. [
34] employed Proton Magnetic Resonance Spectroscopy (MRS) data, in order to identify mild cognitive impairment (MCI) and Alzheimer’s disease (AD) in healthy controls. Savio and Grana [
35] employed Regional Homogeneity to build a CAD for detecting schizophrenia based on resting-state function magnetic resonance imaging (fMRI). Zhang
et al. [
36] proposed employing a three dimensional discrete wavelet transform to extract features from structural MRI, with the aim of detecting Alzheimer’s disease and mild cognitive impairment.
The contribution of this paper is to use fractional Fourier entropy and non-parallel SVMs, with the aim of developing a novel PBD system which has superior classification performance than the above approaches.
3. Materials
At present, there are three benchmark datasets of different sizes, viz., D66, D160, and D255. They were all used for our tests. All datasets contain T2-weighted MR brain images, which were acquired along the axial axis with sizes of 256 × 256. The readers can download them from the Medical School of Harvard University website. The first two datasets consisted of examples from seven types of diseases (meningioma, AD, AD plus visual agnosia, Huntington’s disease, sarcoma, Pick’s disease, and glioma) along with normal brain images. The last dataset D255 contains all seven types of diseases as mentioned before, and four new diseases (multiple sclerosis, cerebral toxoplasmosis, chronic subdural hematoma, and herpes encephalitis).
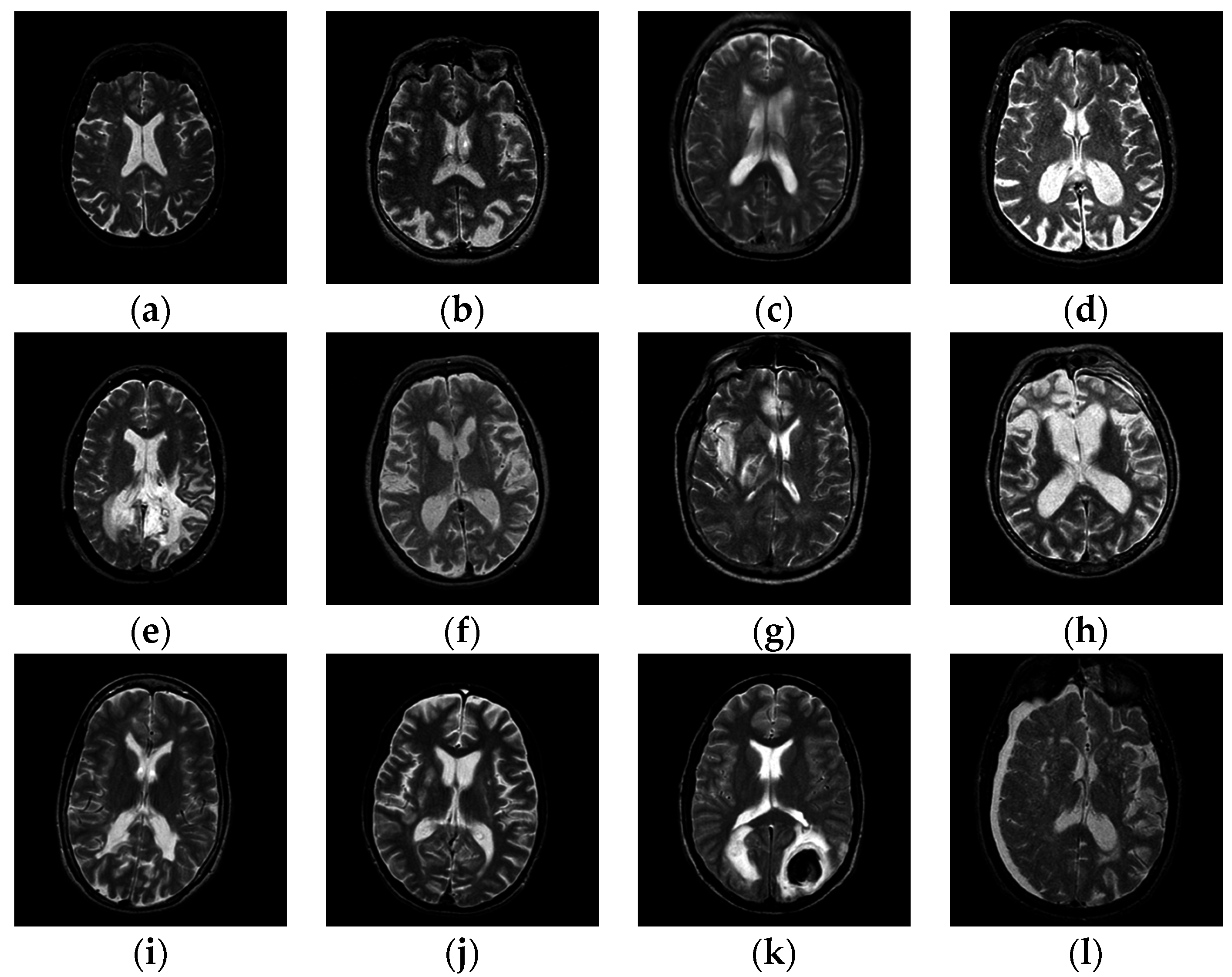
Figure 1 shows samples of the brain images.
Figure 1.
Samples of pathological brain images: (a) healthy brain; (b) AD with visual agnosia; (c) Meningioma; (d) AD; (e) Glioma; (f) Huntington’s disease; (g) Herpes encephalitis; (h) Pick’s disease; (i) Multiple sclerosis; (j) Cerebral toxoplasmosis; (k) Sarcoma; (l) Subdural hematoma.
Figure 1.
Samples of pathological brain images: (a) healthy brain; (b) AD with visual agnosia; (c) Meningioma; (d) AD; (e) Glioma; (f) Huntington’s disease; (g) Herpes encephalitis; (h) Pick’s disease; (i) Multiple sclerosis; (j) Cerebral toxoplasmosis; (k) Sarcoma; (l) Subdural hematoma.
The cost of predicting pathological images as normal is severe, as the treatmen s of patients may be deferred. On the other hand, the cost of misclassification of normal as abnormal is not serious, since other diagnosis means can remedy the error.
This cost-sensitivity (CS) problem can be solved by changing the class distribution at the beginning stage, since the original data is accessible. That means, we intentionally pick more abnormal brains than normal brains in the dataset, with the aim of making the classifier biased to pathological brains, with the aim of addressing the CS problem.
8. Discussion
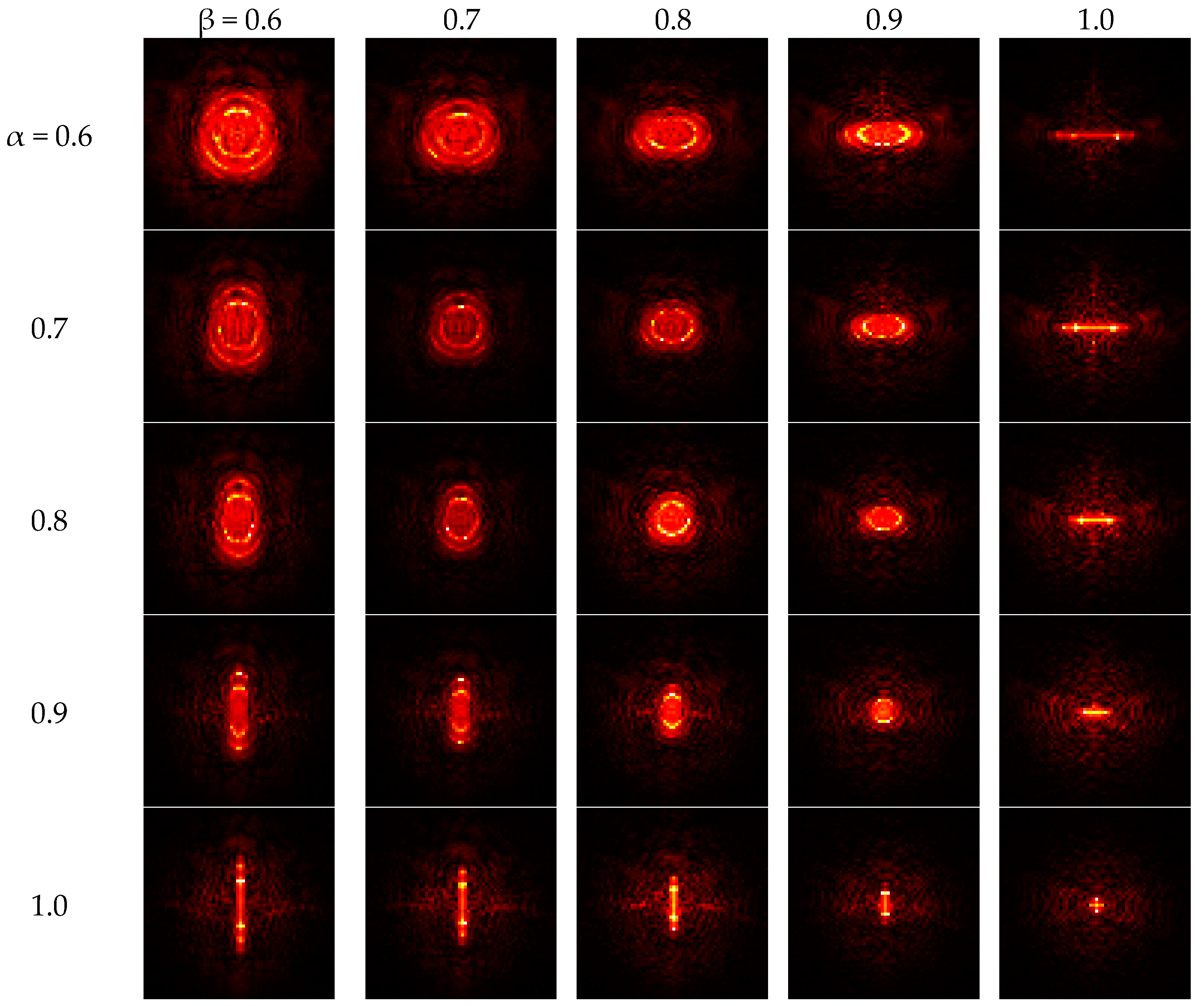
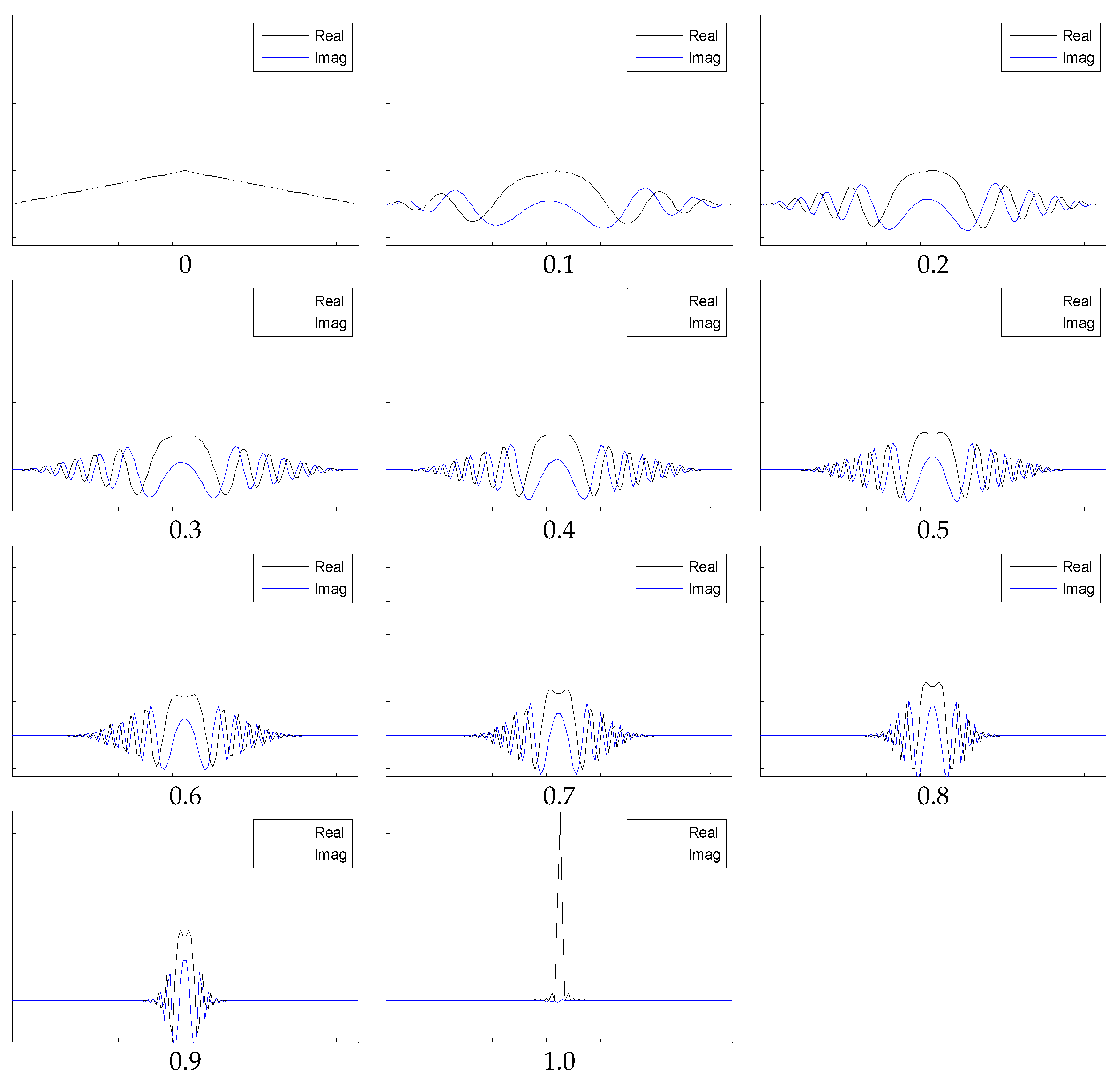
Figure 4 indicates that if both angles increase to one, the FRFT degrades to a standard FT. Contrarily, if both angles reduce to zero, the FRFT degrades to identity operator, which does not contain any frequency spectral information.
Table 4 shows the final selected twelve features are FRFE with parameters (α, β) are assigned with twelve different values of (0.6, 1.0), (0.7, 1.0), (0.8, 0.9), (0.8, 1.0), (0.9, 0.8), (0.9, 0.9), (0.9, 1.0), (1.0, 0.6), (1.0, 0.7), (1.0, 0.8), (1.0, 0.9), and (1.0, 1.0). Note that (α, β) = (1.0, 1.0) represents the standard Fourier Transform (SFT). Those selected features are all closer to the SFT point than those unselected features. It indicates that the SFT and those FRFT near to SFT are more efficient features than others. Note that ONLY (α, β) = (1.0, 1.0) corresponds the SFT. For example, the (0.6, 1.0) means a FRFT with angle of 0.6 along
x-axis and a SFT along
y-axis, so (0.6, 1.0) should be considered a 2D-FRFT other than SFT.
The proposed FRFE measures the information contents in the fractional Fourier domain (FRFD), which is extended from the standard Fourier domain. From another point of view, FRFE is a measure of diversity or unpredictability. A limitation is that FRFE value is not absolute. It depends on the model over FRFD.
Table 5 shows that for NBC, the wavelet entropy obtained accuracies of 92.58%, 91.87%, and 90.51 on D66, D160, and D255, respectively, while the FRFE + WTT + NBC obtained accuracies of 97.12%, 95.94%, and 95.69%, which are higher than the accuracies obtained by wavelet entropy. Therefore, we can conclude the FRFE performed better than wavelet entropy.
Table 6 shows the wavelet energy with SVM achieves accuracies of 82.58%, 80.13%, and 77.76% over three datasets, nevertheless, the FRFE + WTT + SVM yields accuracies of 100.00%, 99.69%, and 98.98% over three datasets. It suggests us FRFE is significantly better than wavelet energy. Comparing the “FRFE + WTT + SVM” in
Table 6 with “FRFE + WTT + NBC” in
Table 5, another finding is SVM is superior to NBC. The reason is SVM works well for large dimensional problems with relative few instances due to its regularization form [
58].
Results in
Table 7 indicate that GEPSVM is superior to standard SVM. Both obtain perfect detection for D66. For D160, the accuracy of GEPSVM is higher than that of SVM by 0.31%. For D255, the accuracy of GEPSVM is higher than that of SVM by 0.20%. Meanwhile, TSVM is superior to GEPSVM. The accuracy of TSVM is 0.39% higher than that of GEPSVM for D255.
The parallel hyperplane setting restrains standard SVM to generate complicated and flexible hyperplanes. NPSVMs discard this setting, so their performances are much better than SVM. TSVM has a resemblance to GEPSVM in spirit, since both drop parallelism. Their difference is that TSVM uses simpler formulation than GEPSVM, and the former can be solved by merely two QP problems. Our results align with the finding in Kumar and Gopal [
59], which says “
generalization performance of TSVM
is better than GEPSVM and conventional SVM”. Nevertheless, Ding
et al. [
60] claimed that TSVM has a lower generalization ability, so it is too early to make a decision about the classification performance of TSVM before more rigorous tests are implemented.
In total, our proposed FRFE + WTT + TSVM predicts 2539 success cases and 13 fail cases in 10 × 5-fold SCV for D255. Remember D255 contains 220 pathological brains and 35 healthy brains, so in total 2200 pathological and 350 healthy instances after 10 repetitions. For 2200 pathological instances, our method predicts 2191 cases successfully, and misclassifies nine pathological instances as healthy. For 350 healthy instances, our method predicts 348 instances successfully, and misclassifies two healthy instances as pathological. Therefore, the sensitivity of our method is 99.59%, specificity is 99.43%, and precision is 99.91% (See
Table 10).
Table 11 lists the comparison results. The first column lists the abbreviated name of the different algorithms. The second column lists the feature number used in each method. The third column lists the number of runs. Here all new algorithms were run 10 times, except some old algorithms which ran five times that were reported in literature [
19]. The last three columns list the classification accuracy over D66, D160, and D255, respectively.
Table 11.
Comparison with other Methods based on 10 × K-fold SCV (# stands for number).
Table 11.
Comparison with other Methods based on 10 × K-fold SCV (# stands for number).
| Existing Algorithms | Feature # | Run # | Accuracy |
|---|
| D66 | D160 | D255 |
|---|
| DWT+SOM [12] | 4761 | 5 | 94.00 | 93.17 | 91.65 |
| DWT+SVM [12] | 4761 | 5 | 96.15 | 95.38 | 94.05 |
| DWT + SVM + RBF [12] | 4761 | 5 | 98.00 | 97.33 | 96.18 |
| DWT + PCA + ANN [13] | 7 | 5 | 97.00 | 96.98 | 95.29 |
| DWT + PCA + KNN [13] | 7 | 5 | 98.00 | 97.54 | 96.79 |
| DWT + PCA + SCG-FNN [14] | 19 | 5 | 100.00 | 99.27 | 98.82 |
| DWT + PCA + SVM [15] | 19 | 5 | 96.01 | 95.00 | 94.29 |
| DWT + PCA + SVM + HPOL [15] | 19 | 5 | 98.34 | 96.88 | 95.61 |
| DWT + PCA + SVM + IPOL [15] | 19 | 5 | 100.00 | 98.12 | 97.73 |
| DWT + PCA + SVM + RBF [15] | 19 | 5 | 100.00 | 99.38 | 98.82 |
| WE + SWP + PNN [16] | 3 | 10 | 100.00 | 99.94 | 98.86 |
| RT + PCA + LS-SVM [19] | 9 | 5 | 100.00 | 100.00 | 99.39 |
| PCNN + DWT + PCA + BPNN [21] | 7 | 10 | 100.00 | 98.88 | 98.24 |
| DWPT + SE + GEPSVM [23] | 16 | 10 | 99.85 | 99.62 | 98.78 |
| DWPT + TE + GEPSVM [23] | 16 | 10 | 100.00 | 100.00 | 99.33 |
| SWT + PCA + IABAP-FNN [30] | 7 | 10 | 100.00 | 99.44 | 99.18 |
| SWT + PCA + ABC-SPSO-FNN [30] | 7 | 10 | 100.00 | 99.75 | 99.02 |
| SWT + PCA + HPA-FNN [30] | 7 | 10 | 100.00 | 100.00 | 99.45 |
| WE + HMI + GEPSVM [33] | 14 | 10 | 100.00 | 99.56 | 98.63 |
| WE + HMI + GEPSVM + RBF [33] | 14 | 10 | 100.00 | 100.00 | 99.45 |
| FRFE + WTT + TSVM (Proposed) | 12 | 10 | 100 | 100.00 | 99.57 |
Table 11 shows that D66 contains too few instances so that many algorithms achieve accuracy of 100%. For the D160, four algorithms achieve perfect classification. They are RT + PCA + LS-SVM [
19], DWPT + TE + GEPSVM [
23], SWT + PCA + HPA-FNN [
30], WE + HMI + GEPSVM + RBF [
33], and the proposed method of “FRFE + WTT + TSVM”. D255 is the most difficult one and no one yields a perfect classification. Among all algorithms, the proposed “FRFE + WTT + TSVM” yields the highest accuracy of 99.57%.
Comparing to “SWT + PCA + HPA-FNN [
30]” with accuracy of 99.45% over D255, our method increases this about 0.12%. Although the improvement is slight, it is obtained by 10 repetitions of 5-fold stratified cross validation, which means the improvement is robust and reliable. SWT + PCA + HPA-FNN [
30] used seven features and proved seven features is the best feature combination, and introducing new features will not improve their accuracy. It does not cost too much time for our method to use 14 features (double of that of SWT + PCA), since 14 features are not a burden to classifiers in current computers.
Our contributions are: (i) we are the first to propose a novel image feature called “Fractional Fourier Entropy (FRFE)”; (ii) WTT is employed to select important FRFEs; (iii) the proposed system “FRFE + WTT + TSVM” is superior to 20 state-of-the-art methods w.r.t. pathological brain detection.
9. Conclusions and Future Research
In this paper, we proposed a novel image feature—Fractional Fourier Entropy (FRFE)—and then use Welch’s t-test (WTT) to select important FRFEs for developing a PBD system. We finally tested four classifiers (NBC, SVM, GEPSVM, and TSVM). The simulation results showed that the proposed “FRFE + WTT + TSVM” yield better results than both other three proposed methods (FRFE + WTT + NBC, FRFE + WTT + SVM, and FRFE + WTT + GEPSVM) and 20 state-of-the-art approaches. Our PBD system may be further applied to brains with more complicated pathological conditions.
In the future research may be performed on the following points: (1) to develop evaluation methods to measure the influence from different values of α and β; (2) trying to consider the use of the least-squares technique to further improve the performance of SVM and NPSVMs; (3) application the FRFE to other pattern recognition problems, such as fruit classification [
61] and tea classification [
62]; (4) testing kernel methods [
63]; (4) mutual entropy [
64] will be introduced to test its performance in feature selection; (5) our method may be applied to X-ray [
65], AD images [
66], and CT images; (6) support vector data description (SVDD) [
67] is commonly used for detecting novel data or outliers, so we will get more medical image data, and test SVDD; (7) swarm intelligence approaches [
68] may be applied to help train classifiers.