The New Genetics and Natural versus Artificial Genetic Modification
Abstract
1. Introduction
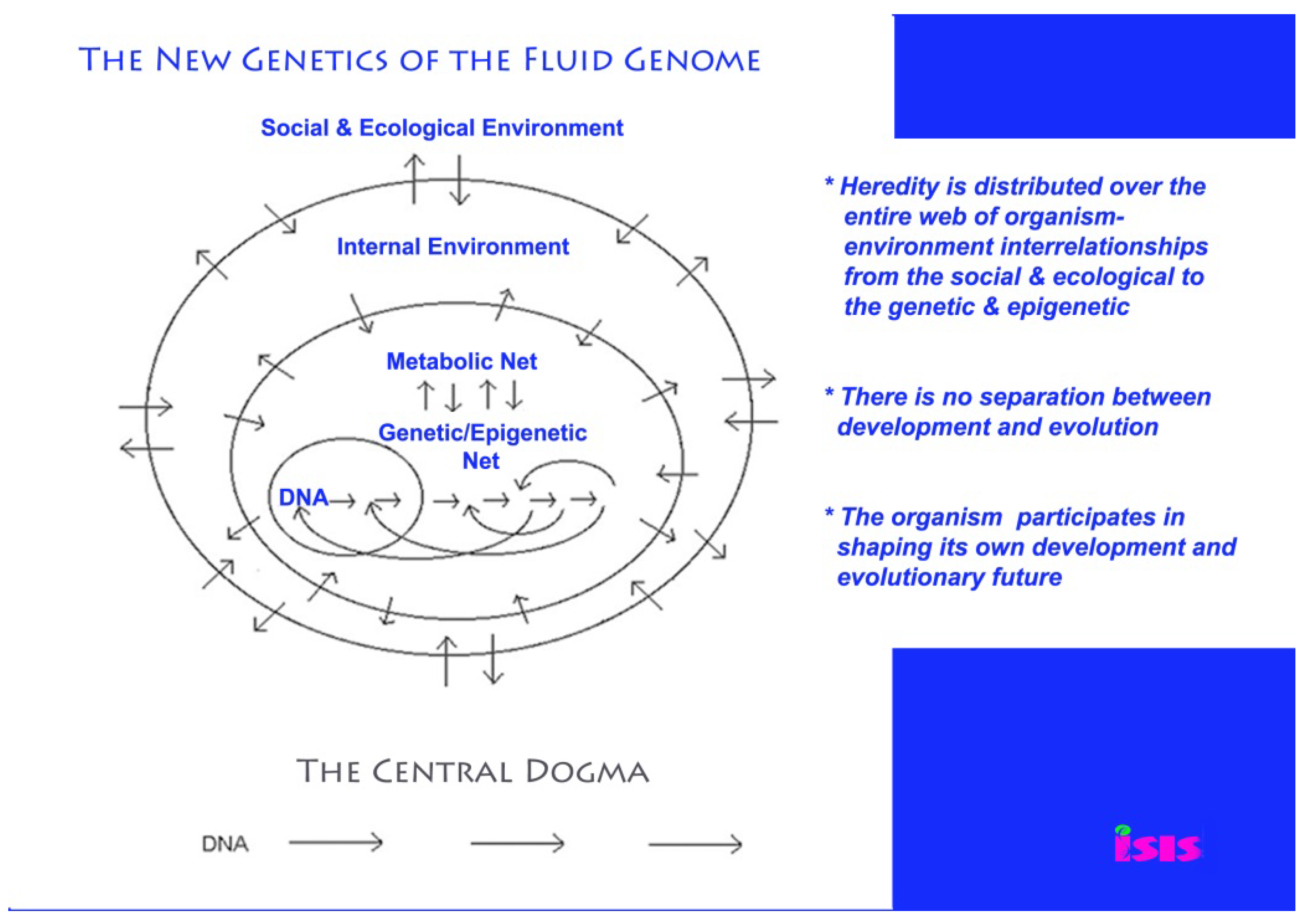
2. The Fluid Genome and Natural Genetic Engineering
3. Genetic Modification Inherently Hazardous
- A paper just published in 2013 reports significantly higher rates of severe stomach inflammation in pigs fed a mixed GM corn and soybean diet for 22.7 weeks compared with pigs fed an equivalent non-GM control diet: 32% compared to 12%. Female pigs on the GM diet also had uterus heavier by 25% on average [27].
- A 2-year lab feeding trial reported in 2012 found rats of both sexes exposed to Roundup and/or Roundup-tolerant maize not sprayed with herbicide were 2 to 3 times as likely to die as controls and to develop large tumours, of mammary glands in females and of kidney and skin in males [28] (see also [29,30]). In other words, the GMO without the herbicide was also harmful in every respect. Pituitary disease was up more than 2 fold in females and liver and kidney diseases up 1.5 to 2 fold in males on GM maize alone that was not sprayed with herbicide. These effects appeared after the 90 days period legally required for safety tests on GMOs, thereby exposing the inadequacy of the regulatory regime.
- A Danish farmer found excessive illnesses and deaths in his pigs fed GM soy meal including chronic diarrhoea, birth defects, reproductive problems, bloating, stomach ulcers, weak and smaller piglets, and reduced litter size. These were entirely reversed when he put them on a GM-free diet [31].
- A meta-analysis pooling all available data on 19 feeding trials carried out for 90 days on GM soybean and maize, both glyphosate tolerant and Bt crops representing 83% of commercialized GMOs—a period inadequate for detecting the most serious health impacts—nevertheless found significant disruption of liver and kidney functions [32,33].
- Between 2005 and 2006, senior scientist Irina Ermakova at the Russian Academy of Sciences reported that female rats fed glyphosate-tolerant GM soybeans produced excessive numbers of severely stunted pups and more than half of the litter dying within three weeks, while the surviving pups were completely sterile [36,37].
- Between 2004 and 2005, hundreds of farm workers and cotton handlers in Madhya Pradesh, India, reported allergy symptoms from exposure to Bt cotton containing Cry1Ac or both Cry1Ac and Cry1Ab proteins [38].
- Between 2005 and 2006, thousands of sheep died after grazing on Bt cotton crop residues in four villages in the Warangal district of Andhra Pradesh in India [39].
- In 2005, scientists at the Commonwealth Scientific and Industrial Research Organization in Canberra Australia tested a transgenic pea containing a normally harmless protein in bean (alpha-amylase inhibitor 1), and found it caused inflammation in the lungs of mice and provoked sensitivities to other proteins in the diet [40,41].
- In 2003, villagers in the south of the Philippines suffered mysterious illnesses when a Monsanto Bt maize hybrid containing Cry1Ab protein came into flower; antibodies to the Cry1Ab protein were found in the villagers, there have been at least five unexplained deaths and some remain ill to this day [47].
- In 2004, Monsanto’s research dossier, kept confidential for commercial reasons, showed that rats fed MON863 GM maize containing Cry3Bb protein developed serious kidney and blood abnormalities [48].
- Between 2001 and 2002, a dozen cows died in Hesse Germany after eating Syngenta GM maize Bt176 containing Cry1Ab/Cry1Ac plus glufosinate-tolerance; and more in the herd had to be slaughtered from illnesses [49]. In 2012, biotech giant Syngenta was criminally charged with denying knowledge it had since 1996, that its GM maize kills livestock, during a civil court case brought by the farmer that ended in 2007 [50].
- In 1998, senior scientist Arpad Pusztai and colleagues formerly of the Rowett Institute in Scotland reported damage in every organ system of young rats fed GM potatoes containing snowdrop lectin, including a stomach lining twice as thick as controls [51].
- Also in 1998, scientists in Egypt found similar effects in the gut of mice fed Bt potato containing a Cry1A protein [52].
- In 2002, Aventis company (later Bayer Cropscience) submitted data to UK regulators showing that chickens fed glufosinate-tolerant GM maize Chardon LL were twice as likely to die compared with controls [53].
4. Sources of Hazards from GMOs
|
|
|
|
5. Natural and Artificial Genetic Modification Are Completely Distinct
6. Transgene Instability and the Illegality of GMOs
7. Horizontal Gene Transfer from GMOs Does Happen
8. Hazards of the CaMV 35S Promoter
9. Hazards of the Agrobacterium Vector
Agrobacterium and Morgellons Disease
10. RNA Interference and Double-Stranded RNA
11. The Nucleic Acids Intercom
11.1. Darwin’s Pangenesis Is Precursor to Nucleic Acid Intercom
11.2. Nucleic Acids Trafficked between Cells
11.3. Homologous Recombination by Uptake of Circulating DNA
12. Conclusions
Acknowledgments
Conflicts of Interest
References
- Bruening, G.; Lyons, J.M. The case of the FLAVR SAVR tomato. Calif. Agric. 2000, 54, 6–7. [Google Scholar] [CrossRef]
- Crick, F.H.C. On protein synthesis. Symp. Soc. Exp. Biol. 1958, 12, 139–163. [Google Scholar]
- Crick, F. Central dogma of molecular biology. Nature 1970, 227, 561–563. [Google Scholar] [CrossRef] [PubMed]
- Watson, J.D.; Crick, F. A structure for deoxyribose nucleic acid. Nature 1953, 171, 737–738. [Google Scholar] [CrossRef] [PubMed]
- Oller, J.W., Jr. The antithesis of entropy: Biosemiotic communication from genetics to human language with special emphasis on the immune systems. Entropy 2010, 12, 631–705. [Google Scholar] [CrossRef]
- Gryder, B.; Nelson, C.; Shepard, S. Biosemiotic entropy of the genome: mutations and epigenetic imbalances resulting in cancer. Entropy 2013, 15, 234–261. [Google Scholar] [CrossRef]
- Samsel, A.; Seneff, S. Glyphosate’s suppression of cytochrome P450 enzymes and amino acid biosynthesis by the gut microbiome: Pathways to modern diseases. Entropy 2013, 15, 1416–1463. [Google Scholar] [CrossRef]
- Ho, M.W. Genetic Engineering Dream of Nightmare? The Brave New World of Bad Science and Big Business, 3rd ed.; Gateway books: New York, NY, USA, 2007. [Google Scholar]
- Dover, G.; Flavell, D. Genome Evolution; Oxford University Press: Oxford, UK, 1982. [Google Scholar]
- Ho, M.W. Evolution by process, not by consequence: Implicationsof the new molecular genetics for development and evolution. Int. J. Comp. Psychol. 1987, 1, 3–27. [Google Scholar]
- Pollard, J.W. The Fluid Genome and Evolution. In Evolutionary Processes and Metaphors; Ho, M.W., Fox, S., Eds.; Wiley: London, UK, 1988; pp. 63–84. [Google Scholar]
- Shapiro, J. Genome organization, natural genetic engineerging and adaptive mutation. Trends Genet. 1997, 13, 98–104. [Google Scholar] [CrossRef]
- Jablonka, E.; Lamb, M. Epigenetic Inheritance and Evolution, the Lamarckian Dimension; Oxford University Press: Oxford, UK, 1995. [Google Scholar]
- Ho, M.W. Death of the central dogma. Sci. Soc. 2004, 24, 4. [Google Scholar]
- Ho, M.W. Living with the Fluid Genome. Available online: http://www.i-sis.org.uk/fluidGenome.php (accessed on 2 August 2013).
- Mattick, J.S.; Mehler, M.F. RNA editing, DNA recoding and the evolution of human cognition. Trends Neurosci. 2008, 31, 227–233. [Google Scholar] [CrossRef] [PubMed]
- Cheng, C.C.; Johnson, T.L.; Hoffmann, A. Epigenetic control: Slow and global, nimble and local. Gene. Dev. 2008, 22, 1110–1114. [Google Scholar] [CrossRef] [PubMed]
- Dietert, R.; Dietert, J. The completed self: An immunological view of the human-microbiome superorganism and risk of chronic diseases. Entropy 2012, 14, 2036–2065. [Google Scholar] [CrossRef]
- Kosoy, M. Deepening the conception of functional information in the description of zoonotic infectious diseases. Entropy 2013, 15, 1929–1962. [Google Scholar] [CrossRef]
- Oller, J.W., Jr. Biosemiotic Entropy: Disorder, Disease, and Mortality. Available online: https://www.mdpi.com/journal/entropy/special_issues/biosemiotic_entropy (accessed on 2 August 2013).
- Tang, W.; Gei, Y.; Page, M. Biological significance of RNA editing in cells. Mol. Biotechnol. 2012, 52, 91–100. [Google Scholar] [CrossRef] [PubMed]
- Agrawal, N.; Dasaradhi, P.V.N.; Mohmmed, A.; Malhotra, P.; Bhatnager, R.K.; Mukherhee, S.K. RNA interference: Biology, mechanism and applications. Microbiol. Mol. Biol. Rev. 2003, 67, 657–685. [Google Scholar] [CrossRef] [PubMed]
- Derr, L.K.; Strathern, J.N. A role for reverse transcripts in gene conversion. Nature 1993, 361, 170–173. [Google Scholar] [CrossRef] [PubMed]
- Ho, M.W. Subverting the genetic text. Sci. Soc. 2004, 24, 6–8. [Google Scholar]
- Ho, M.W. Development and Evolution Revisited. In Handbook of Developmental Science, Behavior and Genetics; Hood, D., Halpern, C., Greenberg, G., Lerner, R., Eds.; Wiley: Chichester, UK, 2010; pp. 61–108. [Google Scholar]
- Ho, M.W.; Cummins, J.; Saunders, P.T. GM food nightmare unfolding in the regulatory sham. Microb. Ecol. Health Dis. 2007, 19, 66–77. [Google Scholar] [CrossRef]
- Carman, J.A.; Vlieger, H.R.; Ver Steeg, L.J.; Sneller, V.E.; Robinson, G.W.; Clinch-Jones, C.A.; Hayes, J.I.; Edwards, J.W. A long-term toxicology study on pigs fed a combined genetically modified (GM) soy and GM mazie diet. J. Org. Sys. 2013, 8, 39–54. [Google Scholar]
- Séralini, G.-E.; Clair, E.; Mesnage, R.; Gress, S.; Defarge, N.; Malatesta, M.; Hennequin, D.; de Vendômois, J.-S. Long term toxicity of a Roundup herbicide and a Roundup-tolerant genetically modified maize. Food Chem. Toxicol. 2012, 50, 4221–4231. [Google Scholar] [CrossRef] [PubMed]
- Saunders, P.T.; Ho, M.W. GM cancer warning can no longer be ignored. Sci. Soc. 2012, 56, 2–4. [Google Scholar]
- Saunders, P.T. Excess cancers and deaths with GM feed: The stats stand up. Sci. Soc. 2012, 56, 4–5. [Google Scholar]
- Sirinathsinghji, E. GM soy linked to illnesses in farm pigs. Sci. Soc. 2012, 55, 8–9. [Google Scholar]
- Séralini, G.-E.; Mesnage, R.; Clair, E.; Gress, S.; Spiroux de Vendôme, J.; Cellier, D. Genetically modified crops safety assessment: Present limits and possible improvements. Environ. Sci. Eur. 2011, 23. [Google Scholar] [CrossRef]
- Sirinathsinghji, E. GM feed toxic, meta-analysis reveals. Sci. Soc. 2011, 52, 30–32. [Google Scholar]
- Ho, M.W. Emergency! Pathogen new to science found in Roundup Ready GM crops. Sci. Soc. 2011, 50, 10–11. [Google Scholar]
- Ho, M.W. Scientist defends claim of new pathogen linked to GM crops. Sci. Soc. 2011, 50, 12–13. [Google Scholar]
- Ermakova, I.V. Genetically modified soy leads to the decrease of weight and high mortality of rat pups of the first generation. Preliminary studies. EcosInform 2006, 1, 4–9. [Google Scholar]
- Ho, M.W. GM soya-fed rats: Stunted, dead, or sterile. Sci. Soc. 2007, 33, 4–6. [Google Scholar]
- Ho, M.W. More illnesses linked to Bt crops. Sci. Soc. 2006, 30, 8–10. [Google Scholar]
- Ho, M.W. Mass deaths in sheep grazing on Bt cotton. Sci. Soc. 2006, 30, 12–13. [Google Scholar]
- Prescott, V.E.; Campbell, P.M.; Moore, A.; Mattes, J.; Rothenberg, M.E.; Foster, P.S.; Higgins, T.J.V.; Hogan, S.P. Transgenic expression of bean a-amylase inhibitor in peas results in altered structure and immunogenicity. J. Agric. Food Chem. 2005, 53, 9023–9030. [Google Scholar] [CrossRef] [PubMed]
- Ho, M.W. Transgenic pea that makes mice ill. Sci. Soc. 2006, 29, 28–29. [Google Scholar]
- Malatesta, M.; Caporaloni, C.; Rossi, L.; Battistelli, S.; Rocchi, M.B.L.; Tonucci, F.; Gazzanelli, G. Ultrastructural analysis of pancreatic acinar cells from mice fed on genetically modified soybean. J. Anat. 2002, 201, 409–415. [Google Scholar] [CrossRef] [PubMed]
- Malatesta, M.; Biggiogera, M.; Manuali, E.; Rochhi, M.B.L.; Baldelli, B.; Gazzanelli, G. Fine structural analyses of pancreatic acinar cell nuclei from mice fed on genetically modified soybean. Eur. J. Histochem. 2003, 47, 385–388. [Google Scholar] [PubMed]
- Malatesta, M.; Caporaloni, C.; Gavaudan, S.; Rocchi, M.B.L.; Serafini, S.; Tiberi, C.; Gazzanelli, G. Ultrastructural morphometrical and immunocytochemical analysis of hepatocyte nuclei from mice fed on genetically modified soybean. Cell Struct. Funct. 2002, 27, 175–180. [Google Scholar] [CrossRef]
- Malatesta, M.; Tiberi, C.; Baldelli, B.; Battistelli, S.; Manuali, E.; Biggiogera, M. Reversibility of hepatocyte nuclear modifications in mice fed on genetically modified soybean. Eur. J. Histochem. 2005, 49, 237–242. [Google Scholar] [PubMed]
- Vecchio, L.; Cisterna, B.; Malatesta, M.; Martin, T.E.; Biggiogera, M. Ultrastructural analysis of testes from mice fed on genetically modified soybean. Eur. J. Histochem. 2004, 48, 449–454. [Google Scholar] [CrossRef]
- Ho, M.W. GM ban long overdue. Sci. Soc. 2006, 29, 26–27. [Google Scholar]
- French experts very disturbed by health effects of Monsanto GM corn (translated from article in LeMonde). Available online: http://www.lobbywatch.org/archive2.asp?arcid=3308 (accessed on 15 October 2013).
- Ho, M.W.; Burcher, S. Cows ate GM maize and died. Sci. Soc. 2004, 21, 4–6. [Google Scholar]
- Sirinathsinghji, E. Syngenta charged for covering up livestock deaths from GM corn. Sci. Soc. 2012, 55, 4–5. [Google Scholar]
- Pusztai, A.; Bardocz, S.; Ewen, S.W.B. Genetically Modified Foods: Potential Human Health Effects. In Food Safety: Contaminants and Toxins; D’Mello, J.P.F., Ed.; Scottish Agricultural College: Edinburgh, UK, 2003; pp. 347–371. [Google Scholar]
- Fares, N.H.; El-Sayed, A.K. Fine structural changes in the ileum of mice fed on δ-endotoxin-treated potatoes and transgenic potatoes. Nat. Toxins 1998, 6, 219–233. [Google Scholar] [CrossRef]
- Novotny, E. Animals avoid GM food, for good reasons. Sci. Soc. 2004, 21, 9–11. [Google Scholar]
- Do seed companies control GM crop research? Available online: https://www.scientificamerican.com/article.cfm?id=do-seed-companies-control-gm-crop-research (accessed on 2 August 2013).
- Ho, M.W.; Lim, L.C. The Case for a GM-Free Sustainable World; Independent Science Panel Report; Institute of Science in Society and Third World Network: London, UK, Penang, Malaysia, 2003. [Google Scholar]
- Ho, M.W.; Lim, L.C. GM-Free, Exposing the Hazards of Biotechnology to Ensure the Integrity of Our Food Supply; Vitalhealth Publishing: Ridgefield, CT, USA, 2004. [Google Scholar]
- Sirinathsinghji, E.; Ho, M.W. Double Jeopardy of Glyphosate and Glyphosate Tolerant Crops. In Ban GMOs Now; ISIS special report; Ho, M.W., Ed.; The Institute of Science in Society: London, UK, 2013. [Google Scholar]
- Sirinathsinghji, E. Bt Crops Failing and Harmful to Health and the Environmen. In Ban GMOs Now; ISIS special report; Ho, M.W., Ed.; The Institute of Science in Society: London, UK, 2013. [Google Scholar]
- Selman, M.; Dankar, S.K.; Forbes, N.E.; Jia, J.-J.; Brown, E.G. Adaptive mutation in influenza A virus non-structural gene is linked to host switching and induces a novel protein by alternative splicing. Emerg. Microbe Infections 2012, 1, e53. [Google Scholar] [CrossRef] [PubMed]
- Ho, M.W. To mutate or not to mutate. Sci. Soc. 2004, 24, 9–10. [Google Scholar]
- Ho, M.W. The Rainbow and the Worm, the Physics of Organisms, 3rd ed.; World Scientific: Singapore, Singapore, 2008. [Google Scholar]
- Ho, M.W. Quantum computer? Is it alive? Available online: http://www.i-sis.org.uk/isisnews/i-sisnews11–24.php#quan (accessed on 2 August 2013).
- Ho, M.W. The real bioinformatics revolution—proteins and nucleic acids singing to one another? Sci. Soc. 2007, 33, 42–45. [Google Scholar]
- Ho, M.W. Non-random directed mutations confirmed. Sci. Soc. 2013, 60. in press. [Google Scholar]
- Wilson, A.; Latham, J.; Steinbrecher, R. Genome Scrambling Myth or Reality; Econexus Technical Report; Ecpmexis: Brighton, UK, 2004. [Google Scholar]
- Bergelson, J.; Purrington, C.B.; Wichmann, G. Promiscuity in transgenic plants. Nature 1998, 395. [Google Scholar] [CrossRef] [PubMed]
- Ho, M.W.; Traavik, T.; Olsvik, O.; Tappeser, B.; Howard, V.; von Weizsacker, C.; McGavin, G.C. Gene technology and gene ecology of infectious diseases. Microb. Ecol. Health Dis. 1998, 10, 33–39. [Google Scholar] [CrossRef] [Green Version]
- Ho, M.W. Horizontal gene transfer—the hidden hazards of genetic engineering. Available online: http://www.i-sis.org.uk/horizontal.php (accessed on 2 August 2013).
- Ho, M.W. GM DNA does jump species. Sci. Soc. 2010, 47, 30–33. [Google Scholar]
- Ho, M.W. Scientists discover new route for GM gene escape. Sci. Soc. 2011, 50, 14–15. [Google Scholar]
- Syvanen, M.; Kado, C.I. (Eds.) Horizontal Gene Transfer, 2nd ed.; Academic Press: Waltham, MA, USA, 2002.
- Eisen, M. GMOs: Gene transfer is neither unnatural nor dangerous. Available online: http://www.science20.com/profile/michael_eisen (accessed on 2 August 2013).
- Collonier, C.; Berthier, G.; Boyer, F.; Duplan, M.-N.; Fernandez, S.; Kebdani, N.; Kobilinsky, A.; Romanuk, M.; Bertheau, Y. Characterization of commercial GMO inserts: A source of useful material to study genome fluidity. Available online: http://www.crii-gen.org (accessed on 2 August 2013).
- Ho, M.W. Transgenic lines proven unstable. Sci. Soc. 2003, 20, 35–36. [Google Scholar]
- Ho, M.W. Unstable transgenic lines illegal. Sci. Soc. 2004, 21, 23. [Google Scholar]
- Windels, P.; Taverniers, I.; Depicker, A.; van Bockstaele, E.; de Loose, M. Characterization of the Roundup Ready soybean insert. Eur. Food Res. Technol. 2001, 213, 107–112. [Google Scholar] [CrossRef]
- Hernández, M.; Pla, M.; Esteve, T.; Prat, S.; Puigdomènech, P.; Ferrando, A. A specific real-time quantitative PCR detection system for event MON810 in maize YieldGard based on the 3’-transgene integration sequence. Transgenic Res. 2003, 12, 170–189. [Google Scholar] [CrossRef]
- Singh, C.K.; Ojka, A.; Kamle, S.; Kachru, D.N. Assessment of cry1Ab transgene cassette in commercial Bt corn MON810: Gene, event, construct & GMO specific concurrent characterization. Nat. Protoc. 2007. [Google Scholar] [CrossRef]
- Rosati, A.; Bogani, P.; Santarlasci, A.; Buiatti, M. Characterisation of 4’ transgene insertion site and derived mRNAs in MON810 YieldGard maize. Plant Mol. Biol. 2008, 67, 271–281. [Google Scholar] [CrossRef] [PubMed]
- Ho, M.W. MON 810 rearranged again. Sci. Soc. 2008, 38, 27. [Google Scholar]
- Ho, M.W. Transgenic lines unstable hence illegal and ineligible for protection. Sci. Soc. 2008, 38, 28–29. [Google Scholar]
- Yin, Z.; Plader, W.; Malepszy, S. Transgene inheritance in plants. J. Appl. Genet. 2004, 45, 127–144. [Google Scholar] [PubMed]
- Ho, M.W.; Cummins, J. Gene therapy risks exposed. Sci. Soc. 2003, 19, 48–50. [Google Scholar]
- Somers, D.A.; Makarevitch, I. Transgene integration in plants: Poking or patching holes in promiscuous genomes? Curr.Opin. Biotech. 2004, 15, 126–131. [Google Scholar] [CrossRef] [PubMed]
- Wu, B.; Sun, Y.H.; Wang, Y.W.; Wang, Y.P.; Zhu, Z.Y. Characterization of transgene integration pattern in F4hGH-transgenic common carp (Cyprinus carpio L.). Cell Res. 2005, 15, 447–454. [Google Scholar] [CrossRef] [PubMed]
- Reim, S.; Hanke, V. Investigation on stability of transgenes and their expression in transgenic apple plants (Malus x domestica Borkh). Acta. Hort. 2004, 663, 419–424. [Google Scholar]
- Flachowsky, J.; Riedel, M.; Reim, S.; Hanke, M.-V. Evaluation of the uniformity and stability of T-DNA integration and gene expression in transgenic apple plants. Electron. J. Biotech. 2008, 11. [Google Scholar] [CrossRef]
- Roman, E.; Soares, A.; Proite, K.; Neiva, S.; Grossi, M.; Faria, J.C.; Rech, E.L.; Aragão, F.J.L. Transgene elimination in genetically modified dry bean and soybean lines. Genet. Mol. Res. 2005, 4, 177–184. [Google Scholar]
- GMO and BIOHAZ Units. Consolidated presentation of the joint Scientific Opinions of the GMO and BIOHAZ panels on “Use of Antibiotic Resistance Genes as Marker Genes in Genetically Modified Plants” and the Scientific Opinion of the GMO Panel on “Consequences of the Opinion on the Use of Antibiotic Resistance Genes as Marker Genes in Genetically Modified Plants on Previous EFSA Assessments of Individual GM Plants” (Statement of EFSA. EFSA-Q-2009–00589 and EFSA-Q-2009–00593). EFSA J. 2009, 1108, 1–8. [Google Scholar]
- De Vries, J.; Herzfeld, T.; Wackernagel, W. Transfer of plastid DNA from tobacco to the soil bacterium Acinetobacter sp. By natural transformation. Mol. Microbiol. 2004, 53, 323–334. [Google Scholar] [CrossRef] [PubMed]
- De Vries, J.; Meier, P.; Wackernagel, W. Microbial horizontal gene transfer and the DNA release from transgenic crop plants. Plant Soil 2004, 266, 91–104. [Google Scholar] [CrossRef]
- Simpson, D.J.; Fry, J.C.; Rogers, H.J.; Day, M.J. Transformation of Acinetobacyer baylyi in non-sterile soil using recombinant plant nuclear DNA. Environ. Biosaf. Res. 2007, 6, 101–112. [Google Scholar] [CrossRef] [PubMed]
- Rizzi, A.; Pontiroli, A.; Brusetti, L.; Borin, S.; Solini, C.; Abruzzese, A.; Sacchi, G.A.; Vogel, T.M.; Simonet, P.; Bazzicalupo, M.; et al. Strategy for in situ detection of natural transformation-based horizontal gene transfer events. Appl. Environ. Microbiol. 2008, 74, 1250–1254. [Google Scholar] [CrossRef] [PubMed]
- Pontiroli, A.; Rizzi, A.; Simonet, P.; Daffonchio, D.; Vogel, T.M.; Monier, J.M. Visual evidence of horizontal gene transfer between plants and bacteria in the phytosphere of transplastomic tobacco. Appl. Environ. Microbiol. 2009, 75, 3314–3322. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Jin, M.; Qiu, Z.G.; Guo, C.; Chen, Z.L.; Shen, Z.Q.; Wang, X.W.; Li, J.W. A survey of drug resistance bla genes originating from synthetic plasmid vectors in six Chinese rivers. Environ. Sci. Tech. 2012, 46, 13448–13454. [Google Scholar] [CrossRef] [PubMed]
- Sirinathsinghji, E. GM antibiotic resistance in China’s rivers. Sci. Soc. 2013, 57, 6–7. [Google Scholar]
- Chowdhury, E.H.; Mikami, O.; Nakajima, Y.; Hino, A.; Kuribara, H.; Suga, K.; Hanazumi, M.; Yomemochi, C. Detection of genetically modified maize DNA fragments in the gastrointestinal contents of pigs fed StarLink CBH351. Vet. Hum. Toxicol. 2003, 45, 95–96. [Google Scholar] [PubMed]
- Reuter, T.; Aulrich, K. Investigations on genetically modified maize (Bt-maize) in pig nutrition: Fate of feed-ingested foreign DNA in pig bodies. Eur. Food Res. Technol. 2003, 216, 185–192. [Google Scholar]
- Duggan, P.S.; Chambers, P.A.; Heritage, J.; Forbes, J.M. Fate of genetically modified maize DNA in the oral cavity and rumen of sheep. Brit. J. Nutr. 2003, 89, 159–166. [Google Scholar] [CrossRef] [PubMed]
- Phipps, R.H.; Deaville, E.R.; Maddison, B.C. Detection of transgenic and endogenous plant DNA in rumen fluid, duodenal digesta, milk, blood and faeces of lactating dairy cows. J. Dairy Sci. 2003, 86, 4070–4078. [Google Scholar] [CrossRef]
- Ho, M.W. DNA in GM food & feed. Sci. Soc. 2004, 23, 34–36. [Google Scholar]
- Netherwood, T.; Martin-Orue, S.M.; O-Donnell, A.G.; Gockling, S.; Graham, J.; Mathers, J.C.; Gilbert, J.H. Assessing the survival of transgenic plant DNA in the human gastrointestinal tract. Nat. Biotechnol. 2004, 22, 204–209. [Google Scholar] [CrossRef] [PubMed]
- Netherwood, T.; Bowden, R.; Harrison, P.; O’Donnell, A.G.; Parker, D.S.; Gilbert, H.J. Gene transfer in the gastrointestinal tract. Appl. Environ. Microb. 1999, 65, 5139–5141. [Google Scholar]
- Kelly, B.G.; Vespermann, A.; Bolton, D.J. Gene transfer events and their occurrence in selected environments. Food Chem. Toxicol. 2009, 47, 978–983. [Google Scholar] [CrossRef] [PubMed]
- McCuddin, Z.; Carlson, S.A.; Rasmussen, M.A.; Franklin, S.K. Klebsiella to Salmonella gene transfer within rumen protozoa: Implications for antibiotic resistance and rumen defaunation. Vet. Microb. 2005, 114, 275–284. [Google Scholar] [CrossRef] [PubMed]
- Mercer, D.K.; Scott, K.P.; Bruce-Johnson, W.A.; Glover, L.A.; Flint, H.J. Fate of free DNA and transformation of the oral bacterium Streptococcus gordonii DL by plasmid DNA in human saliva. Appl. Environ. Microb. 1999, 65, 6–10. [Google Scholar]
- Ho, M.W.; Ryan, A.; Cummins, J.; Traavik, T. Slipping Through the Regulatory Net: Naked and Free Nucleic Acids; TWN: Penang, Malaysia, 2001. [Google Scholar]
- Podevin, N.; du Jardin, P. Possible consequences of the overlap between the CaMV 35S promoter regions in plant transformation vectors used and the viral gene VI in transgenic plants. GM Crop. Food 2012, 3, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Ho, M.W. Hazardous virus gene discovered in GM crops after 20 years. Sci. Soc. 2013, 57, 2–3. [Google Scholar]
- Latham, J.; Wilson, A. Potentially dangerous virus gene hidden in commercial GM crops. Sci. Soc. 2013, 57, 4–5. [Google Scholar]
- Ho, M.W.; Ryan, A.; Cummins, J. Cauliflower mosaic viral promoter—a recipe for disaster? Microb. Ecol. Health Dis. 1999, 11, 194–197. [Google Scholar] [CrossRef]
- Ho, M.W.; Ryan, A.; Cummins, J. Hazards of transgenic plants with the cauliflower mosaic viral promoter. Microb. Ecol. Health Dis. 2000, 12, 6–11. [Google Scholar] [CrossRef]
- Ho, M.W.; Ryan, A.; Cummins, J. CaMV35S promoter fragmentation hotspot confirmed and it is active in animals. Microb. Ecol. Health Dis. 2000, 12. [Google Scholar] [CrossRef]
- Ballas, N.; Broido, S.; Soreq, H.; Loyter, A. Efficient functioning of plant promoters and poly(A) sites in Xenopus oocytes. Nucl. Acids Res. 1989, 17, 7891–7903. [Google Scholar] [CrossRef] [PubMed]
- Burke, C.; Yu, X.B.; Marchitelli, L.; Davis, E.A.; Ackerman, S. Transcription factor IIA of wheat and human function similarly with plant and animal viral promoters. Nucl. Acids Res. 1990, 18, 3611–3620. [Google Scholar] [CrossRef] [PubMed]
- Cui, X.; Fan, B.; Scholz, J.; Chan, Z. Roles of Arabidopsis cyclin-dependent kinase C complexes in cauliflower mosaic virus infection, plant growth and development. Plant Cell 2007, 19, 1388–1402. [Google Scholar] [CrossRef] [PubMed]
- Ho, M.W.; Cummins, J. New evidence links CaMV 35S promoter to HIV transcription. Microb. Ecol. Health Dis. 2009, 21, 172–174. [Google Scholar] [CrossRef]
- Hazards of GMOS: Agrobacterium mediated transformation. Available online: http://www.bristol.ac.uk/news/2010/7279.html (accessed on 2 August 2013).
- Knight, C.J.; Bailey, A.M.; Foster, G.D. Investigating Agrobacterium-mediated transformation of Verticillium albo-atrum on plant surfaces. PLoS One 2010, 5, e13684. [Google Scholar] [CrossRef] [PubMed]
- McNicol, M.J.; Lyon, G.D.; Chen, M.Y.; Barrett, C.; Cobb, E. The Possibility of Agrobacterium as a Vehicle for Gene Escape; R&D and Surveillance Report, Contract No. RG 0202; Scottish Crop Research Institute: Scotland, UK, 1996. [Google Scholar]
- Barrett, C.; Cobb, E.; MacNiCol, R.; Lyon, G. A risk assessment study of plant genetic transformation using Agrobacterium and implication for analysis of transgenic plants. Plant Cell Tiss. Org. Cult. 1997, 19, 135–144. [Google Scholar] [CrossRef]
- Mogilner, N.; Zutra, D.; Gafny, R.; Bar-Joseph, M. the persistence of engineered Agrobacterium tumefaciens in agroinfected plants. Mol. Plant—Microb. Interact. 1993, 6, 673–675. [Google Scholar] [CrossRef]
- Soltani, J.; van Heusden, P.H.; Hooykaas, P.J.J. Agrobacterium-Mediated Transformation of Non-Plant Organisms. In Agrobacterium: From Biology to Biotechnology; Tzfira, T., Citovsky, V., Eds.; Springer: New York, NY, USA, 2008; pp. 649–674. [Google Scholar]
- Ho, M.W. Horizontal gene transfer from GMOs does happen. Sci. Soc. 2008, 38, 22–24. [Google Scholar]
- Ferguson, G.; Heinemann, J. Recent History of Trans-Kingdom Conjugation. In Horizontal Gene Transfer, 2nd ed.; Syvanen, M., Kado, C.I., Eds.; Academic Press: San Diego, CA, USA, 2002; pp. 3–17. [Google Scholar]
- Ho, M.W. Horizontal gene transfer. Heredity 2003, 90, 6–7. [Google Scholar] [CrossRef]
- Kado, C. Horiontal Transmission of Genes by Agrobacterium Species. In Horizontal Gene Transfer, 2nd ed.; Syvanen, M., Kado, C.I., Eds.; Academic Press: San Diego, CA, USA, 2002; pp. 45–50. [Google Scholar]
- Sengelov, G.; Kristensen, K.J.; Sorensen, A.H.; Kroer, N.; Sorensen, S.J. Effect of genomic location on horizontal transfer of a recombinant gene cassette between Pseudomonas strains in the rhizosphere and spermosphere of barley seedlings. Curr. Microb. 2001, 42, 160–167. [Google Scholar] [CrossRef]
- Kunik, T.; Tzfira, T.; Kapulnik, Y.; Gafni, Y.; Dingwall, C.; Citovsky, V. Genetic transformation of HeLa cells by Agrobacterium. Proc. Natl. Acad. Sci. USA 2001, 98, 1871–1876. [Google Scholar] [CrossRef] [PubMed]
- Cummins, J. Common plant vector injects genes into human cells. Available online: http://www.i-sis.org.uk/Agrobacterium.php (accessed on 2 August 2013).
- CDC to launch study on unexplained illness. Available online: http://www.cdc.gov/od/oc/media/transcripts/2008/t080116.htm#id=45169 (accessed on 2 August 2013).
- Unexplained Dermopathy (aka “Morgellons”). Available online http://www.cdc.gov/unexplaineddermopathy/general_info.html (accessed on 2 August 2013).
- Ho, M.W.; Cummins, J. Agrobacterium & Morgellons disease, a GM connection? Sci. Soc. 2008, 38, 33–36. [Google Scholar]
- Savely, V.R.; Leitao, M.M.; Stricker, R.B. The mystery of Morgellons disease, infection or delusion? Am. J. Clin. Dermatol. 2006, 7, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Stricker, R.B.; Savely, V.R.; Saltsman, A.; Citovsky, V. Contribution of Agrobacterium to Morgellons disease. J. Invest. Med. 2007, 55 (Suppl.), S123. [Google Scholar]
- Pearson, M.L.; Selby, J.V.; Katz, K.A.; Cantrell, V.; Braden, C.R.; Parise, M.E.; Paddock, C.D.; Lewin-Smith, M.R.; Kalasinsky, V.F.; Goldstein, F.C.; et al. Clinical, epidemiologic, histopathologic and molecular features of an unexplained dermopathy. PLoS One 2012, 7, e29908. [Google Scholar] [CrossRef] [PubMed]
- Heinemann, J.A.; Agapito-Tenfen, S.Z.; Carman, J.A. A comparative evaluation of the regulation of GM crops or products containing dsRNA and suggested improvements to risk assessment. Environ. Int. 2013, 55, 43–55. [Google Scholar] [CrossRef] [PubMed]
- Fire, A.; Xu, S.; Montgomery, M.K.; Kostas, S.A.; Diver, S.E.; Mello, C.C. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 1998, 391, 806–811. [Google Scholar] [CrossRef] [PubMed]
- Cogoni, C.; Macino, G. Post-transcriptional gene silencing across kingdoms. Curr. Opin. Genet. Dev. 2000, 10, 638–643. [Google Scholar] [CrossRef]
- Hawkins, P.G.; Sharon, S.S.; Adams, C.; Anest, V.; Morris, K.V. Promoter targeted small RNAs induce long-term transcriptional gene silencing in human cells. Nucl. Acids Res. 2009, 37. [Google Scholar] [CrossRef] [PubMed]
- Alder, M.N.; Dames, S.; Gaudet, J.; Mango, S.E. Gene silencing in Caernorhabditis elegans by transitive NA interference. RNA 2003, 9, 25–32. [Google Scholar] [CrossRef] [PubMed]
- Ho, M.W. Epigenetic inheritance, what genes remember. Sci. Soc. 2009, 41, 4–5. [Google Scholar]
- Ho, M.W. New GM nightmares with RNA. Sci. Soc. 2013, 58, 6–7. [Google Scholar]
- Zhang, L.; Hou, D.; Chen, X.; Li, D.; Zhu, L.; Zhang, Y.; Li, J.; Bian, Z.; Liang, X.; Cai, X.; et al. Exogenous plant MiR168a specifically targets mammalian LDIRAP1: Evidence of cross-kingdom regulation by microRNA. Cell Res. 2012, 22, 107–126. [Google Scholar] [CrossRef] [PubMed]
- Ho, M.W. How food affects genes. Sci. Soc. 2012, 53, 12–13. [Google Scholar]
- Zhang, Y.; Wiggins, E.; Lawrence, C.; Petrick, J.; Ivashuta, S.; Heck, G. Analysis of plant-derived miRNAs in animal small RNA datasets. BCM Genomics 2012, 13. [Google Scholar] [CrossRef] [PubMed]
- Thermo Scientific Tech Support. Off-target effects: disturbing the silence of RNA interference (RNAi). Available online: http://www.thermoscientificbio.com/uploadedFiles/Resources/off-target-tech-review.pdf (accessed on 2 August 2013).
- Helwak, A.; Kudla, G.; Dudnakova, T.; Tollervey, D. Mapping the human miRNA Interactome by CLASH reveals frequent noncanonical binding. Cell 2013, 153, 654–665. [Google Scholar] [CrossRef] [PubMed]
- Ho, M.W. RNA interference “complex and flexible” & beyond control. Sci. Soc. 2013, 59. in press. [Google Scholar]
- Grimm, D.; Streetz, K.L.; Jopling, C.L.; Storm, T.A.; Pandey, K.; Davis, C.R.; Marion, P.; Salazar, F.; Kay, M.A. Fatality in mice due to oversaturation of cellular microRNA/shorthairpin RNA pathways. Nature 2006, 441, 537–541. [Google Scholar] [CrossRef] [PubMed]
- Ho, M.W. Gene therapy nightmare for mice could human be next? Sci. Soc. 2006, 31, 25. [Google Scholar]
- Couzin, J. RNAi shows cracks in its armor. Science 2004, 306, 1124–1125. [Google Scholar] [CrossRef] [PubMed]
- Ho, M.W. Controversy over gene therapy breakthrough. Sci. Soc. 2005, 26, 38. [Google Scholar]
- Heinemann, J.A. Update on “Evaluation of risks from creation of novel RNA molecules in genetically engineered wheat plants and recommendations for risk assessment”. Personal communication, Rocky Mountain Laboratories: Hamilton, MT, USA, 21 March 2013. [Google Scholar]
- Ho, M.W. Human genome map spells death of genetic determinism. ISIS News. 2001, 7/8, pp. 1–2. Available online: http://www.i-sis.org.uk/isisnews/i-sisnews7-1.php (accessed on 2 August 2013).
- Ho, M.W. Ten years of the human genome. Sci. Soc. 2010, 48, 22–25. [Google Scholar]
- Zuk, O.; Hechter, E.; Sunyaev, S.R.; Lander, E.R. The mystery of missing heritability: Genetic interactions create phantom heritability. Available online: http://www.pnas.org/cgi/coi/10.1073/pnas.1119685109 (accessed on 2 August 2013).
- Ho, M.W. Mystery of missing heritability solved? Sci. Soc. 2012, 53, 26–31. [Google Scholar]
- Ho, M.W. No genes for intelligence. Sci. Soc. 2012, 53, 28–31. [Google Scholar]
- Ho, M.W. No genes for intelligence in the fluid genome. Adv. Child Dev. Behav. 2013, 45, 67–92. [Google Scholar] [PubMed]
- Ribozyme. Available online: http://en.wikipedia.org/wiki/Ribozyme (accessed on 2 August 2013).
- Wiedenheft, B.; Sternberg, S.H.; Doudna, J.A. RNA-guided genetic silencing systems in bacteria and archaea. Nature 2012, 482, 331–338. [Google Scholar] [CrossRef] [PubMed]
- Ho, M.W. Intercommunication via circulating nucleic acids. Sci. Soc. 2009, 42, 46–48. [Google Scholar]
- Liu, Y. A new perspective on Darwin’s Pangenesis. Biol. Rev. 2008, 83, 141–149. [Google Scholar] [CrossRef] [PubMed]
- Ho, M.W. Darwin’s pangenesis, the hidden history of genetics & the dangers of GMOs. Sci. Soc. 2009, 42, 42–45. [Google Scholar]
- Spadafora, C. Sperm-mediated “reverse” gene transfer: A role of reverse transcriptase in the generation of new genetic information. Hum. Reprod. 2008, 23, 735–740. [Google Scholar] [CrossRef] [PubMed]
- Ho, M.W. Epigenetic inheritance through sperm cells, the Lamarckian dimension in evolution. Sci. Soc. 2009, 42, 40–42. [Google Scholar]
- Rasoulzadegan, M.; Grandjean, V.; Gounon, P.; .Vicent, S.; Gillot, I.; Cuzin, F. RNA-mediated non-Mendelian inheritance of an epigenetic change in the mouse. Nature 2006, 441, 469–474. [Google Scholar] [CrossRef] [PubMed]
- Corrado, C.; Raimondo, S.; Chiesi, A.; Ciccia, F.; de Leo, G.; Alessandro, R. Exosomes as intercellular signaling organelles involved in health and disease: basic science and clinical applications. Int. J. Mol. Sci. 2013, 14, 5338–5366. [Google Scholar] [CrossRef] [PubMed]
- Sahoo, S.; Klychkod, E.; Thorne, T.; Misener, S.; Schultz, K.M.; Milay, M.; Ito, A.; Liu, T.; Kamide, C.; Agrawal, H.; et al. Exosomes from human CD34+ stem cells mediate the proangiogenic paracrine activity. Circ. Res. 2011, 109, 724–728. [Google Scholar] [CrossRef] [PubMed]
- Bergsmedh, A.; Szeles, A.; Henriksson, M.; Bratt, A.; Foldman, M.J.; Spetz, A.-L.; Holmgren, L. Horizontal transfer of oncogenes by uptake of apoptotic bodies. Proc. Natl. Acad. Sci. USA 2001, 98, 6407–6411. [Google Scholar] [CrossRef] [PubMed]
- Beling, M.; Wittrup, A. Nanotubes, exosomes, and nucleic acid-binding peptides provide novel mechanisms of intercellular communication in eukaryotic cells: implications in health and disease. J. Cell Biol. 2008, 183, 1187–1191. [Google Scholar] [CrossRef] [PubMed]
- Van der Vaart, M.; Pretorius, P.J. Circulating DNA, its origins and fluctuation. Ann. N. Y. Acad. Sci. 2008, 1137, 18–26. [Google Scholar] [CrossRef] [PubMed]
- Gahan, P.B.; Ander, P.; Stroun, M. Metabolic DNA as the origin of spontaneously released DNA? Ann. N. Y. Acad. Sci. 2008, 1137, 7–17. [Google Scholar] [CrossRef] [PubMed]
- Gahan, P.B.; Stroun, M. The Biology of CNAPS. In Extracellular Nucleic Acids; Rykova, E.Y., Kikuchi, Y., Eds.; Springer: Berlin, Germany, 2010. [Google Scholar]
- Beck, J.; Urnovitz, H.B.; Riggert, J.; Cierici, M.; Schütz, E. Profile of the circulating DNA in apparently healthy individuals. Clin. Chem. 2009, 55, 730–738. [Google Scholar] [CrossRef] [PubMed]
- Yakubov, L.A.; Petrova, N.A.; Popova, N.A.; Semenov, D.V.; Nikolin, V.P.; Os’kina, I.N. The role of extracellular DNA in the stability and variability of cell genomes. Dokl. Biochem. Biophys. 2002, 382, 31–34. [Google Scholar] [CrossRef] [PubMed]
- Yakubov, L.A.; Rogachev, V.A.; Likhacheva, A.C.; Bogachev, S.S.; Seeleva, T.E.; Shilov, A.G.; Baiborodin, S.I.; Petroa, N.; Mechelina, L.V.; Shurdov, M.A.; et al. Natural human gene correction by small extracellular genomic DNA fragments. Cell Cycle 2007, 6, 2293–2301. [Google Scholar] [CrossRef] [PubMed]
© 2013 by the author; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ho, M.-W. The New Genetics and Natural versus Artificial Genetic Modification. Entropy 2013, 15, 4748-4781. https://doi.org/10.3390/e15114748
Ho M-W. The New Genetics and Natural versus Artificial Genetic Modification. Entropy. 2013; 15(11):4748-4781. https://doi.org/10.3390/e15114748
Chicago/Turabian StyleHo, Mae-Wan. 2013. "The New Genetics and Natural versus Artificial Genetic Modification" Entropy 15, no. 11: 4748-4781. https://doi.org/10.3390/e15114748
APA StyleHo, M.-W. (2013). The New Genetics and Natural versus Artificial Genetic Modification. Entropy, 15(11), 4748-4781. https://doi.org/10.3390/e15114748




