Update on Rapid Diagnostics for COVID-19: A Systematic Review
Abstract
:1. Introduction
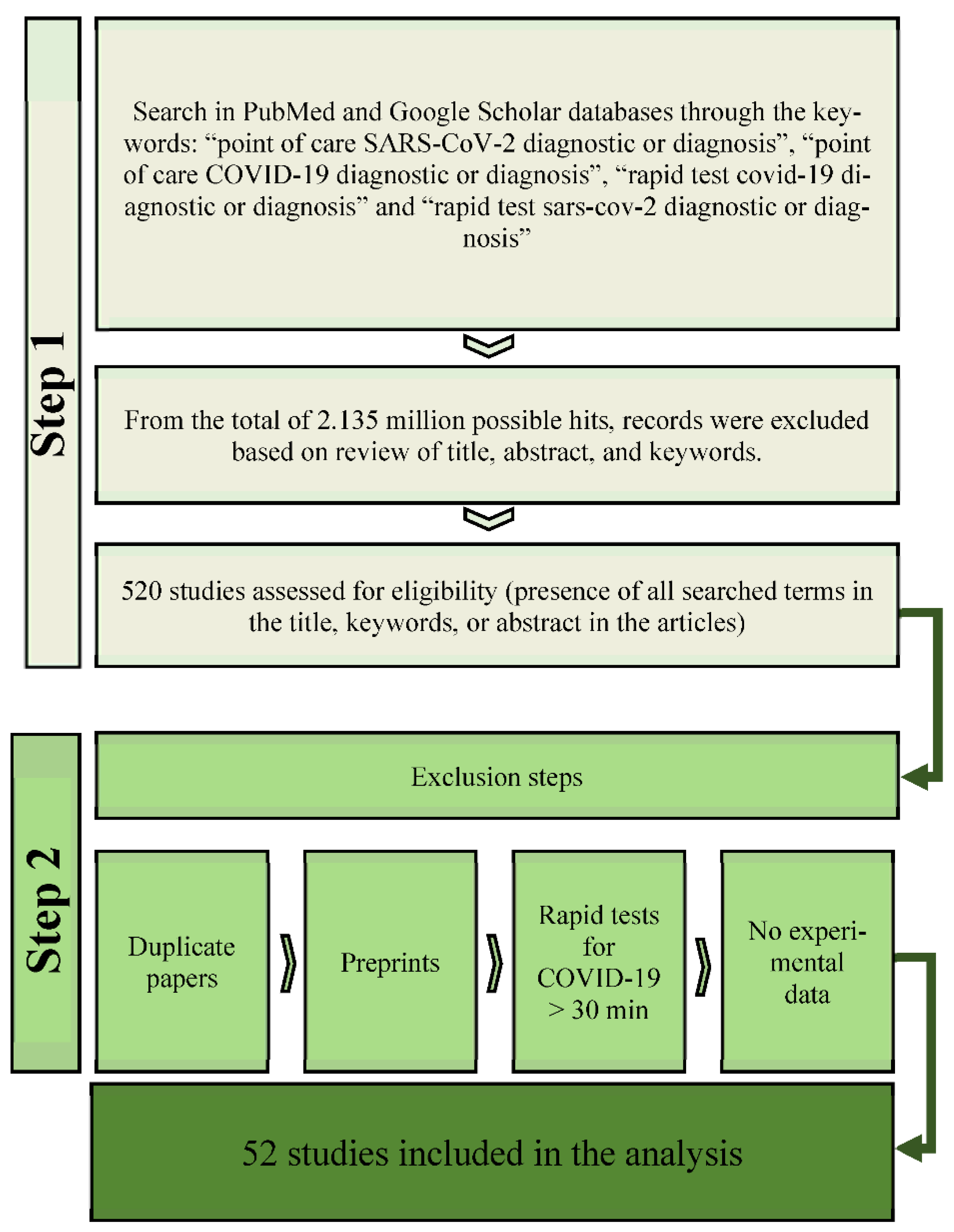
2. Methods
2.1. Literature Search
2.2. Selection Criteria
2.3. Data Extraction and Quality Assessment
3. Results
3.1. Direct Detection Tests
3.2. Immunoglobulin Detection Tests
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef] [Green Version]
- WHO. Director-General’s Opening Remarks at the Media Briefing on COVID-19. 11 March 2020. Available online: https://www.who.int/director-general/speeches/detail/who-director-general-s-opening-remarks-at-the-media-briefing-on-covid-19---11-march-2020 (accessed on 2 June 2022).
- WHO. Coronavirus (COVID-19) Dashboard. Available online: https://covid19.who.int/ (accessed on 2 June 2022).
- Jin, Y.; Wang, M.; Zuo, Z.; Fan, C.; Ye, F.; Cai, Z.; Wang, Y.; Cui, H.; Pan, K.; Xu, A. Diagnostic value and dynamic variance of serum antibody in coronavirus disease 2019. Int. J. Infect. Dis. 2020, 94, 49–52. [Google Scholar] [CrossRef] [PubMed]
- WHO. Diagnostic Testing for SARS-CoV-2. 2020. Available online: https://www.who.int/publications/i/item/diagnostic-testing-for-sars-cov-2 (accessed on 2 June 2022).
- Guo, L.; Ren, L.; Yang, S.; Xiao, M.; Chang, D.; Yang, F.; Dela Cruz, C.S.; Wang, Y.; Wu, C.; Xiao, Y.; et al. Profiling Early Humoral Response to Diagnose Novel Coronavirus Disease (COVID-19). Clin. Infect. Dis. 2020, 71, 778–785. [Google Scholar] [CrossRef] [Green Version]
- UNICEF. Supply Division, Most Affordable COVID-19 Rapid Diagnostic Test Now Available. Available online: https://www.unicef.org/supply/stories/most-affordable-covid-19-rapid-diagnostic-test-now-available (accessed on 2 June 2022).
- WHO. Antigen-Detection in the Diagnosis of SARS-CoV-2 Infection (2021). Available online: https://www.who.int/publications/i/item/antigen-detection-in-the-diagnosis-of-sars-cov-2infection-using-rapid-immunoassays (accessed on 2 June 2022).
- WHO. Global Partnership to Make Available 120 Million Affordable, Quality COVID-19 Rapid Tests for Low- and Middle-Income Countries. 2020. Available online: https://www.who.int/news/item/28-09-2020-global-partnership-to-make-available-120-million-affordable-quality-covid-19-rapid-tests-for-low--and-middle-income-countries (accessed on 2 June 2022).
- Liu, R.; He, L.; Hu, Y.; Luo, Z.; Zhang, J. A serological aptamer-assisted proximity ligation assay for COVID-19 diagnosis and seeking neutralizing aptamers. Chem. Sci. 2020, 11, 12157–12164. [Google Scholar] [CrossRef] [PubMed]
- Xiong, Y.; Zhang, J.; Yang, Z.; Mou, Q.; Ma, Y.; Xiong, Y.; Lu, Y. Functional DNA Regulated CRISPR-Cas12a Sensors for Point-of-Care Diagnostics of Non-Nucleic-Acid Targets. J. Am. Chem. Soc. 2020, 142, 207–213. [Google Scholar] [CrossRef] [PubMed]
- Liu, R.; Hu, Y.; He, Y.; Lan, T.; Zhang, J. Translating daily COVID-19 screening into a simple glucose test: A proof of concept study. Chem. Sci. 2021, 12, 9022–9030. [Google Scholar] [CrossRef]
- Zhang, W.; He, Y.; Feng, Z.; Zhang, J. Recent advances of functional nucleic acid-based sensors for point-of-care detection of SARS-CoV-2. Mikrochim. Acta 2022, 189, 128. [Google Scholar] [CrossRef]
- Maxmen, A. Untapped potential: More US labs could be providing tests for coronavirus. Nature 2020, 580, 312. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bassi, L.L.; Hwenda, L. COVID-19: Time to plan for prompt universal access to diagnostics and treatments. Lancet Glob. Health 2020, 8, e756–e757. [Google Scholar] [CrossRef]
- Borges, L.P.; Martins, A.F.; Silva, B.M.; Dias, B.P.; Goncalves, R.L.; Souza, D.R.V.; Oliveira, M.G.B.; Jesus, P.C.; Serafini, M.R.; Quintans, J.S.S.; et al. Rapid diagnosis of COVID-19 in the first year of the pandemic: A systematic review. Int. Immunopharmacol. 2021, 101, 108144. [Google Scholar] [CrossRef] [PubMed]
- Masia, M.; Fernandez-Gonzalez, M.; Sanchez, M.; Carvajal, M.; Garcia, J.A.; Gonzalo-Jimenez, N.; Ortiz de la Tabla, V.; Agullo, V.; Candela, I.; Guijarro, J.; et al. Nasopharyngeal Panbio COVID-19 Antigen Performed at Point-of-Care Has a High Sensitivity in Symptomatic and Asymptomatic Patients with Higher Risk for Transmission and Older Age. Open Forum Infect. Dis. 2021, 8, ofab059. [Google Scholar] [CrossRef] [PubMed]
- Kruger, L.J.; Gaeddert, M.; Tobian, F.; Lainati, F.; Gottschalk, C.; Klein, J.A.F.; Schnitzler, P.; Krausslich, H.G.; Nikolai, O.; Lindner, A.K.; et al. The Abbott PanBio WHO emergency use listed, rapid, antigen-detecting point-of-care diagnostic test for SARS-CoV-2-Evaluation of the accuracy and ease-of-use. PLoS ONE 2021, 16, e0247918. [Google Scholar] [CrossRef] [PubMed]
- Kolwijck, E.; Brouwers-Boers, M.; Broertjes, J.; van Heeswijk, K.; Runderkamp, N.; Meijer, A.; Hermans, M.H.A.; Leenders, A. Validation and implementation of the Panbio COVID-19 Ag rapid test for the diagnosis of SARS-CoV-2 infection in symptomatic hospital healthcare workers. Infect. Prev. Pract. 2021, 3, 100142. [Google Scholar] [CrossRef] [PubMed]
- Cortes Rubio, J.A.; Costa Zamora, M.P.; Canals Aracil, M.; Pulgar Feio, M.; Mata Martinez, A.; Carrasco Munera, A. Evaluation of the diagnostic test for rapid detection of COVID-19 antigen (Panbio Covid rapid test) in primary care. Semergen 2021, 47, 508–514. [Google Scholar] [CrossRef] [PubMed]
- L’Huillier, A.G.; Lacour, M.; Sadiku, D.; Gadiri, M.A.; De Siebenthal, L.; Schibler, M.; Eckerle, I.; Pinosch, S.; Kaiser, L.; Gervaix, A.; et al. Diagnostic Accuracy of SARS-CoV-2 Rapid Antigen Detection Testing in Symptomatic and Asymptomatic Children in the Clinical Setting. J. Clin. Microbiol. 2021, 59, e0099121. [Google Scholar] [CrossRef] [PubMed]
- Villaverde, S.; Dominguez-Rodriguez, S.; Sabrido, G.; Perez-Jorge, C.; Plata, M.; Romero, M.P.; Grasa, C.D.; Jimenez, A.B.; Heras, E.; Broncano, A.; et al. Diagnostic Accuracy of the Panbio Severe Acute Respiratory Syndrome Coronavirus 2 Antigen Rapid Test Compared with Reverse-Transcriptase Polymerase Chain Reaction Testing of Nasopharyngeal Samples in the Pediatric Population. J. Pediatrics 2021, 232, 287–289.e4. [Google Scholar] [CrossRef]
- Muhi, S.; Tayler, N.; Hoang, T.; Ballard, S.A.; Graham, M.; Rojek, A.; Kwong, J.C.; Trubiano, J.A.; Smibert, O.; Drewett, G.; et al. Multi-site assessment of rapid, point-of-care antigen testing for the diagnosis of SARS-CoV-2 infection in a low-prevalence setting: A validation and implementation study. Lancet Reg. Health-West. Pac. 2021, 9, 100115. [Google Scholar] [CrossRef] [PubMed]
- Merino, P.; Guinea, J.; Munoz-Gallego, I.; Gonzalez-Donapetry, P.; Galan, J.C.; Antona, N.; Cilla, G.; Hernaez-Crespo, S.; Diaz-de Tuesta, J.L.; Gual-de Torrella, A.; et al. Multicenter evaluation of the Panbio COVID-19 rapid antigen-detection test for the diagnosis of SARS-CoV-2 infection. Clin. Microbiol. Infect. 2021, 27, 758–761. [Google Scholar] [CrossRef]
- Albert, E.; Torres, I.; Bueno, F.; Huntley, D.; Molla, E.; Fernandez-Fuentes, M.A.; Martinez, M.; Poujois, S.; Forque, L.; Valdivia, A.; et al. Field evaluation of a rapid antigen test (Panbio COVID-19 Ag Rapid Test Device) for COVID-19 diagnosis in primary healthcare centres. Clin. Microbiol. Infect. 2021, 27, 472.e7–472.e10. [Google Scholar] [CrossRef] [PubMed]
- Carbonell-Sahuquillo, S.; Lazaro-Carreno, M.I.; Camacho, J.; Barres-Fernandez, A.; Albert, E.; Torres, I.; Breton-Martinez, J.R.; Martinez-Costa, C.; Navarro, D. Evaluation of a rapid antigen detection test (Panbio COVID-19 Ag Rapid Test Device) as a point-of-care diagnostic tool for COVID-19 in a pediatric emergency department. J. Med. Virol. 2021, 93, 6803–6807. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Lee, J.; Bal, J.; Seo, S.K.; Chong, C.K.; Lee, J.H.; Park, H. Development and Clinical Evaluation of an Immunochromatography-Based Rapid Antigen Test (GenBody COVAG025) for COVID-19 Diagnosis. Viruses 2021, 13, 796. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.; Gupta, P.; Mathuria, Y.P.; Kalita, D.; Prasad, A.; Panda, P.K.; Bahurupi, Y.; Sahoo, B.; Omar, B.J. Comparative Evaluation of Nasopharyngeal and Oropharyngeal Swab Based Rapid SARS-CoV-2 Antigen Detection and Real-Time RT-PCR for Diagnosis of COVID-19 in Tertiary Care Hospital. Cureus 2021, 13, e16785. [Google Scholar] [CrossRef]
- Mockel, M.; Corman, V.M.; Stegemann, M.S.; Hofmann, J.; Stein, A.; Jones, T.C.; Gastmeier, P.; Seybold, J.; Offermann, R.; Bachmann, U.; et al. SARS-CoV-2 antigen rapid immunoassay for diagnosis of COVID-19 in the emergency department. Biomarkers 2021, 26, 213–220. [Google Scholar] [CrossRef] [PubMed]
- Jegerlehner, S.; Suter-Riniker, F.; Jent, P.; Bittel, P.; Nagler, M. Diagnostic accuracy of a SARS-CoV-2 rapid antigen test in real-life clinical settings. Int. J. Infect. Dis. 2021, 109, 118–122. [Google Scholar] [CrossRef] [PubMed]
- Ciotti, M.; Maurici, M.; Pieri, M.; Andreoni, M.; Bernardini, S. Performance of a rapid antigen test in the diagnosis of SARS-CoV-2 infection. J. Med. Virol. 2021, 93, 2988–2991. [Google Scholar] [CrossRef]
- Kanaujia, R.; Ghosh, A.; Mohindra, R.; Singla, V.; Goyal, K.; Gudisa, R.; Sharma, V.; Mohan, L.; Kaur, N.; Mohi, G.K.; et al. Rapid antigen detection kit for the diagnosis of SARS-CoV-2—Are we missing asymptomatic patients? Indian J. Med. Microbiol. 2021, 39, 457–461. [Google Scholar] [CrossRef]
- Strömer, A.; Rose, R.; Schäfer, M.; Schön, F.; Vollersen, A.; Lorentz, T.; Fickenscher, H.; Krumbholz, A. Performance of a Point-of-Care Test for the Rapid Detection of SARS-CoV-2 Antigen. Microorganisms 2021, 9, 58. [Google Scholar] [CrossRef] [PubMed]
- Thakur, P.; Saxena, S.; Manchanda, V.; Rana, N.; Goel, R.; Arora, R. Utility of Antigen-Based Rapid Diagnostic Test for Detection of SARS-CoV-2 Virus in Routine Hospital Settings. Lab. Med. 2021, 52, e154–e158. [Google Scholar] [CrossRef]
- Chaimayo, C.; Kaewnaphan, B.; Tanlieng, N.; Athipanyasilp, N.; Sirijatuphat, R.; Chayakulkeeree, M.; Angkasekwinai, N.; Sutthent, R.; Puangpunngam, N.; Tharmviboonsri, T.; et al. Rapid SARS-CoV-2 antigen detection assay in comparison with real-time RT-PCR assay for laboratory diagnosis of COVID-19 in Thailand. Virol. J. 2020, 17, 177. [Google Scholar] [CrossRef]
- Amer, R.M.; Samir, M.; Gaber, O.A.; El-Deeb, N.A.; Abdelmoaty, A.A.; Ahmed, A.A.; Samy, W.; Atta, A.H.; Walaa, M.; Anis, R.H. Diagnostic performance of rapid antigen test for COVID-19 and the effect of viral load, sampling time, subject’s clinical and laboratory parameters on test accuracy. J. Infect. Public Health 2021, 14, 1446–1453. [Google Scholar] [CrossRef]
- Diao, B.; Wen, K.; Zhang, J.; Chen, J.; Han, C.; Chen, Y.; Wang, S.; Deng, G.; Zhou, H.; Wu, Y. Accuracy of a nucleocapsid protein antigen rapid test in the diagnosis of SARS-CoV-2 infection. Clin. Microbiol. Infect. 2021, 27, 289.e1–289.e4. [Google Scholar] [CrossRef] [PubMed]
- Mboumba Bouassa, R.S.; Veyer, D.; Pere, H.; Belec, L. Analytical performances of the point-of-care SIENNA COVID-19 Antigen Rapid Test for the detection of SARS-CoV-2 nucleocapsid protein in nasopharyngeal swabs: A prospective evaluation during the COVID-19 second wave in France. Int. J. Infect. Dis. 2021, 106, 8–12. [Google Scholar] [CrossRef]
- Tinker, S.C.; Szablewski, C.M.; Litvintseva, A.P.; Drenzek, C.; Voccio, G.E.; Hunter, M.A.; Briggs, S.; Heida, D.E.; Folster, J.; Shewmaker, P.L.; et al. Point-of-Care Antigen Test for SARS-CoV-2 in Asymptomatic College Students. Emerg. Infect. Dis. 2021, 27, 2662–2665. [Google Scholar] [CrossRef]
- Orsi, A.; Pennati, B.M.; Bruzzone, B.; Ricucci, V.; Ferone, D.; Barbera, P.; Arboscello, E.; Dentone, C.; Icardi, G. On-field evaluation of a ultra-rapid fluorescence immunoassay as a frontline test for SARS-CoV-2 diagnostic. J. Virol. Methods 2021, 295, 114201. [Google Scholar] [CrossRef]
- Cassuto, N.G.; Gravier, A.; Colin, M.; Theillay, A.; Pires-Roteira, D.; Pallay, S.; Serreau, R.; Hocqueloux, L.; Prazuck, T. Evaluation of a SARS-CoV-2 antigen-detecting rapid diagnostic test as a self-test: Diagnostic performance and usability. J. Med. Virol. 2021, 93, 6686–6692. [Google Scholar] [CrossRef]
- Rastawicki, W.; Gierczynski, R.; Juszczyk, G.; Mitura, K.; Henry, B.M. Evaluation of PCL rapid point of care antigen test for detection of SARS-CoV-2 in nasopharyngeal swabs. J. Med. Virol. 2021, 93, 1920–1922. [Google Scholar] [CrossRef]
- Bianco, G.; Boattini, M.; Barbui, A.M.; Scozzari, G.; Riccardini, F.; Coggiola, M.; Lupia, E.; Cavallo, R.; Costa, C. Evaluation of an antigen-based test for hospital point-of-care diagnosis of SARS-CoV-2 infection. J. Clin. Virol. 2021, 139, 104838. [Google Scholar] [CrossRef] [PubMed]
- Kruger, L.J.; Klein, J.A.F.; Tobian, F.; Gaeddert, M.; Lainati, F.; Klemm, S.; Schnitzler, P.; Bartenschlager, R.; Cerikan, B.; Neufeldt, C.J.; et al. Evaluation of accuracy, exclusivity, limit-of-detection and ease-of-use of LumiraDx: An antigen-detecting point-of-care device for SARS-CoV-2. Infection 2022, 50, 395–406. [Google Scholar] [CrossRef]
- Drain, P.K.; Ampajwala, M.; Chappel, C.; Gvozden, A.B.; Hoppers, M.; Wang, M.; Rosen, R.; Young, S.; Zissman, E.; Montano, M. A Rapid, High-Sensitivity SARS-CoV-2 Nucleocapsid Immunoassay to Aid Diagnosis of Acute COVID-19 at the Point of Care: A Clinical Performance Study. Infect. Dis. Ther. 2021, 10, 753–761. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.; Ju, C.; Han, C.; Shi, R.; Chen, X.; Duan, D.; Yan, J.; Yan, X. Nanozyme chemiluminescence paper test for rapid and sensitive detection of SARS-CoV-2 antigen. Biosens. Bioelectron. 2020, 173, 112817. [Google Scholar] [CrossRef] [PubMed]
- Egerer, R.; Edel, B.; Loffler, B.; Henke, A.; Rodel, J. Performance of the RT-LAMP-based eazyplex(R) SARS-CoV-2 as a novel rapid diagnostic test. J. Clin. Virol. 2021, 138, 104817. [Google Scholar] [CrossRef]
- Wang, Y.; Li, K.; Xu, G.; Chen, C.; Song, G.; Dong, Z.; Lin, L.; Wang, Y.; Xu, Z.; Yu, M.; et al. Low-Cost and Scalable Platform with Multiplexed Microwell Array Biochip for Rapid Diagnosis of COVID-19. Research 2021, 2021, 2813643. [Google Scholar] [CrossRef] [PubMed]
- Xun, G.; Lane, S.T.; Petrov, V.A.; Pepa, B.E.; Zhao, H. A rapid, accurate, scalable, and portable testing system for COVID-19 diagnosis. Nat. Commun. 2021, 12, 2905. [Google Scholar] [CrossRef] [PubMed]
- de Oliveira Coelho, B.; Sanchuki, H.B.S.; Zanette, D.L.; Nardin, J.M.; Morales, H.M.P.; Fornazari, B.; Aoki, M.N.; Blanes, L. Essential properties and pitfalls of colorimetric Reverse Transcription Loop-mediated Isothermal Amplification as a point-of-care test for SARS-CoV-2 diagnosis. Mol. Med. 2021, 27, 30. [Google Scholar] [CrossRef] [PubMed]
- Wei, S.; Suryawanshi, H.; Djandji, A.; Kohl, E.; Morgan, S.; Hod, E.A.; Whittier, S.; Roth, K.; Yeh, R.; Alejaldre, J.C.; et al. Field-deployable, rapid diagnostic testing of saliva for SARS-CoV-2. Sci. Rep. 2021, 11, 5448. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.Y.; Degani, I.; Cheong, J.; Lee, J.H.; Choi, H.J.; Cheon, J.; Lee, H. Fluorescence polarization system for rapid COVID-19 diagnosis. Biosens. Bioelectron. 2021, 178, 113049. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; He, P.; Xiong, D.; Lou, Y.; Pu, Q.; Zhang, H.; Zhang, H.; Yu, J. A Reverse Transcription Recombinase-Aided Amplification Method for Rapid and Point-of-Care Detection of SARS-CoV-2, including Variants. Viruses 2021, 13, 1875. [Google Scholar] [CrossRef]
- Margulis, M.; Erster, O.; Roth, S.; Mandelboim, M.; Danielli, A. A Magnetic Modulation Biosensing-Based Molecular Assay for Rapid and Highly Sensitive Clinical Diagnosis of Coronavirus Disease 2019 (COVID-19). J. Mol. Diagn. 2021, 23, 1680–1690. [Google Scholar] [CrossRef] [PubMed]
- Torres, M.D.T.; de Araujo, W.R.; de Lima, L.F.; Ferreira, A.L.; de la Fuente-Nunez, C. Low-cost biosensor for rapid detection of SARS-CoV-2 at the point of care. Matter 2021, 4, 2403–2416. [Google Scholar] [CrossRef] [PubMed]
- Kumar, V.; Ghosh, K.; Chandran, A.; Panwar, S.; Bhat, A.; Konaje, S.; Das, S.; Srikanta, S.; Jaganathan, L.; Prasad, S.; et al. Multicentric evaluation of a novel point of care electrochemical ELISA platform for SARS-CoV-2 specific IgG and IgM antibody assay. J. Virol. Methods 2021, 298, 114275. [Google Scholar] [CrossRef] [PubMed]
- De Munck, D.G.; Peeters, B.; Huyghe, E.; Goossens, H.; Ieven, M.; Matheeussen, V. Performance of the FREND COVID-19 IgG/IgM Duo point-of-care test for SARS-CoV-2 antibody detection. Acta Clin. Belg. 2022, 77, 647–652. [Google Scholar] [CrossRef] [PubMed]
- Peng, T.; Sui, Z.; Huang, Z.; Xie, J.; Wen, K.; Zhang, Y.; Huang, W.; Mi, W.; Peng, K.; Dai, X.; et al. Point-of-care test system for detection of immunoglobulin-G and -M against nucleocapsid protein and spike glycoprotein of SARS-CoV-2. Sens. Actuators B Chem. 2021, 331, 129415. [Google Scholar] [CrossRef] [PubMed]
- Kiziloglu, I.; Sener, A.; Siliv, N. Comparison of rapid antibody test and thorax computed tomography results in patients who underwent RT-PCR with the pre-diagnosis of COVID-19. Int. J. Clin. Pract. 2021, 75, e14524. [Google Scholar] [CrossRef]
- Plebani, M.; Parcina, M.; Bechri, I.; Zehender, G.; Terkes, V.; Abdel Hafith, B.; Antinori, S.; Pillet, S.; Gonzalo, S.; Hoerauf, A.; et al. Performance of the COVID19SEROSpeed IgM/IgG Rapid Test, an Immunochromatographic Assay for the Diagnosis of SARS-CoV-2 Infection: A Multicenter European Study. J. Clin. Microbiol. 2021, 59, e02240-20. [Google Scholar] [CrossRef] [PubMed]
- Villarreal, A.; Rangel, G.; Zhang, X.; Wong, D.; Britton, G.; Fernandez, P.L.; Perez, A.; Oviedo, D.; Restrepo, C.; Carreirra, M.B.; et al. Performance of a Point of Care Test for Detecting IgM and IgG Antibodies Against SARS-CoV-2 and Seroprevalence in Blood Donors and Health Care Workers in Panama. Front. Med. 2021, 8, 616106. [Google Scholar] [CrossRef]
- Scotta, M.C.; David, C.N.; Varela, F.H.; Sartor, I.T.S.; Polese-Bonatto, M.; Fernandes, I.R.; Zavaglia, G.O.; Ferreira, C.F.; Kern, L.B.; Santos, A.P.; et al. Low performance of a SARS-CoV-2 point-of-care lateral flow immunoassay in symptomatic children during the pandemic. J. Pediatr. 2022, 98, 136–141. [Google Scholar] [CrossRef]
- Fauziah, N.; Koesoemadinata, R.C.; Andriyoko, B.; Faridah, L.; Riswari, S.F.; Widyatmoko, L.; Prihatni, D.; Ekawardhani, S.; Fibriani, A.; Rachmawati, E.; et al. The performance of point-of-care antibody test for COVID-19 diagnosis in a tertiary hospital in Bandung, Indonesia. J. Infect. Dev. Ctries. 2021, 15, 237–241. [Google Scholar] [CrossRef]
- Pallett, S.J.C.; Denny, S.J.; Patel, A.; Charani, E.; Mughal, N.; Stebbing, J.; Davies, G.W.; Moore, L.S.P. Point-of-care SARS-CoV-2 serological assays for enhanced case finding in a UK inpatient population. Sci. Rep. 2021, 11, 5860. [Google Scholar] [CrossRef]
- Prendecki, M.; Clarke, C.; McKinnon, T.; Lightstone, L.; Pickering, M.C.; Thomas, D.C.; McAdoo, S.P.; Willicombe, M. SARS-CoV-2 Antibody Point-of-Care Testing in Dialysis and Kidney Transplant Patients With COVID-19. Kidney Med. 2021, 3, 54–59.e1. [Google Scholar] [CrossRef]
- Zhang, P.; Chen, L.; Hu, J.; Trick, A.Y.; Chen, F.E.; Hsieh, K.; Zhao, Y.; Coleman, B.; Kruczynski, K.; Pisanic, T.R., 2nd; et al. Magnetofluidic immuno-PCR for point-of-care COVID-19 serological testing. Biosens. Bioelectron. 2022, 195, 113656. [Google Scholar] [CrossRef]
- Elledge, S.K.; Zhou, X.X.; Byrnes, J.R.; Martinko, A.J.; Lui, I.; Pance, K.; Lim, S.A.; Glasgow, J.E.; Glasgow, A.A.; Turcios, K.; et al. Engineering luminescent biosensors for point-of-care SARS-CoV-2 antibody detection. Nat. Biotechnol. 2021, 39, 928–935. [Google Scholar] [CrossRef] [PubMed]
- Li, K.; Tong, C.; Ha, X.; Zeng, C.; Chen, X.; Xu, F.; Yang, J.; Du, H.; Chen, Y.; Cai, J.; et al. Development and clinical evaluation of a rapid antibody lateral flow assay for the diagnosis of SARS-CoV-2 infection. BMC Infect. Dis. 2021, 21, 860. [Google Scholar] [CrossRef] [PubMed]
- Gaythorpe, K.A.M.; Bhatia, S.; Mangal, T.; Unwin, H.J.T.; Imai, N.; Cuomo-Dannenburg, G.; Walters, C.E.; Jauneikaite, E.; Bayley, H.; Kont, M.D.; et al. Children’s role in the COVID-19 pandemic: A systematic review of early surveillance data on susceptibility, severity, and transmissibility. Sci. Rep. 2021, 11, 13903. [Google Scholar] [CrossRef] [PubMed]
- Cevik, M.; Tate, M.; Lloyd, O.; Maraolo, A.E.; Schafers, J.; Ho, A. SARS-CoV-2, SARS-CoV, and MERS-CoV viral load dynamics, duration of viral shedding, and infectiousness: A systematic review and meta-analysis. Lancet Microbe 2021, 2, e13–e22. [Google Scholar] [CrossRef]
- Centers for Disease Control and Prevention. Guidance for Antigen Testing for SARS-CoV-2 for Healthcare Providers Testing Individuals in the Community. 2022. Available online: https://www.cdc.gov/coronavirus/2019-ncov/lab/resources/antigen-tests-guidelines.html (accessed on 2 June 2022).
- Puhach, O.; Adea, K.; Hulo, N.; Sattonnet, P.; Genecand, C.; Iten, A.; Jacquerioz, F.; Kaiser, L.; Vetter, P.; Eckerle, I.; et al. Infectious viral load in unvaccinated and vaccinated individuals infected with ancestral, Delta or Omicron SARS-CoV-2. Nat. Med. 2022. online ahead of print. [Google Scholar] [CrossRef]
- Levine-Tiefenbrun, M.; Yelin, I.; Katz, R.; Herzel, E.; Golan, Z.; Schreiber, L.; Wolf, T.; Nadler, V.; Ben-Tov, A.; Kuint, J.; et al. Initial report of decreased SARS-CoV-2 viral load after inoculation with the BNT162b2 vaccine. Nat. Med. 2021, 27, 790–792. [Google Scholar] [CrossRef] [PubMed]
- McEllistrem, M.C.; Clancy, C.J.; Buehrle, D.J.; Lucas, A.; Decker, B.K. Single Dose of an mRNA Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-Cov-2) Vaccine Is Associated With Lower Nasopharyngeal Viral Load among Nursing Home Residents With Asymptomatic Coronavirus Disease 2019 (COVID-19). Clin. Infect. Dis. 2021, 73, e1365–e1367. [Google Scholar] [CrossRef]
- Long, Q.X.; Liu, B.Z.; Deng, H.J.; Wu, G.C.; Deng, K.; Chen, Y.K.; Liao, P.; Qiu, J.F.; Lin, Y.; Cai, X.F.; et al. Antibody responses to SARS-CoV-2 in patients with COVID-19. Nat. Med. 2020, 26, 845–848. [Google Scholar] [CrossRef]
- WHO. Advice on the Use of Point-of-Care Immunodiagnostic Tests for COVID-19. 2020. Available online: https://www.who.int/news-room/commentaries/detail/advice-on-the-use-of-point-of-care-immunodiagnostic-tests-for-covid-19 (accessed on 2 June 2022).
- De Carlo, A.; Lo Caputo, S.; Paolillo, C.; Rosa, A.M.; D’Orsi, U.; De Palma, M.; Reveglia, P.; Lacedonia, D.; Cinnella, G.; Foschino, M.P.; et al. SARS-COV-2 Serological Profile in Healthcare Professionals of a Southern Italy Hospital. Int. J. Environ. Res. Public Health 2020, 17, 9324. [Google Scholar] [CrossRef]
- Panbio™ COVID-19 Ag Rapid Test Device. Available online: https://dam.abbott.com/en-gb/panbio/120007883-v1-Panbio-COVID-19-Ag-Nasal-AsymptomaticSe.pdf (accessed on 2 June 2022).
- GenBody COVID-19 Ag. Available online: https://www.fda.gov/media/150788/download (accessed on 2 June 2022).
- Roche SARS-CoV-2 Rapid Antigen Test. Available online: https://diagnostics.roche.com/au/en_gb/c/covidqr-patients.html#package-insert (accessed on 2 June 2022).
- COVID-19 Ag Respi-Strip, Coris BioConcept. Available online: https://www.corisbio.com/pdf/Products/COVID-19-Respi-Strip_20201113.pdf (accessed on 2 June 2022).
- NADAL® COVID-19 Ag. Available online: https://www.nadal-test.com/en/antigen (accessed on 2 June 2022).
- STANDARDTM Q COVID-19 Ag Test. Available online: https://www.who.int/diagnostics_laboratory/eual/eul_0563_117_00_standard_q_covid19_ag_ifu.pdf (accessed on 26 November 2021).
- COVID19SEROSpeed-IgM-IgG. Available online: https://biospeedia.com/wp-content/uploads/2021/02/BioSpeedia-Covid-19-Antibody-BSD_501-IFU-English.pdf (accessed on 2 June 2022).
- Masyeni, S.; Santoso, M.S.; Widyaningsih, P.D.; Asmara, D.W.; Nainu, F.; Harapan, H.; Sasmono, R.T. Serological cross-reaction and coinfection of dengue and COVID-19 in Asia: Experience from Indonesia. Int. J. Infect. Dis. 2021, 102, 152–154. [Google Scholar] [CrossRef]
- Steinhardt, L.C.; Ige, F.; Iriemenam, N.C.; Greby, S.M.; Hamada, Y.; Uwandu, M.; Aniedobe, M.; Stafford, K.A.; Abimiku, A.; Mba, N.; et al. Cross-Reactivity of Two SARS-CoV-2 Serological Assays in a Setting Where Malaria Is Endemic. J. Clin. Microbiol. 2021, 59, e0051421. [Google Scholar] [CrossRef] [PubMed]
- Faccini-Martinez, A.A.; Rivero, R.; Garay, E.; Garcia, A.; Mattar, S.; Botero, Y.; Galeano, K.; Miranda, J.; Martinez, C.; Guzman, C.; et al. Serological cross-reactivity using a SARS-CoV-2 ELISA test in acute Zika virus infection, Colombia. Int. J. Infect. Dis. 2020, 101, 191–193. [Google Scholar] [CrossRef] [PubMed]
- Huber, T.; Steininger, P.; Irrgang, P.; Korn, K.; Tenbusch, M.; Diesch, K.; Achenbach, S.; Kremer, A.E.; Werblow, M.; Vetter, M.; et al. Diagnostic performance of four SARS-CoV-2 antibody assays in patients with COVID-19 or with bacterial and non-SARS-CoV-2 viral respiratory infections. Eur. J. Clin. Microbiol. Infect. Dis. 2021, 40, 1983–1997. [Google Scholar] [CrossRef]
- Lustig, Y.; Keler, S.; Kolodny, R.; Ben-Tal, N.; Atias-Varon, D.; Shlush, E.; Gerlic, M.; Munitz, A.; Doolman, R.; Asraf, K.; et al. Potential Antigenic Cross-reactivity Between Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) and Dengue Viruses. Clin. Infect. Dis. 2020, 73, e2444–e2449. [Google Scholar] [CrossRef] [PubMed]
- Lansbury, L.; Lim, B.; Baskaran, V.; Lim, W.S. Co-infections in people with COVID-19: A systematic review and meta-analysis. J. Infect. 2020, 81, 266–275. [Google Scholar] [CrossRef] [PubMed]
- Sardar, S.; Sharma, R.; Alyamani, T.Y.M.; Aboukamar, M. COVID-19 and Plasmodium vivax malaria co-infection. IDCases 2020, 21, e00879. [Google Scholar] [CrossRef]
- Fan, B.E.; Lim, K.G.E.; Chong, V.C.L.; Chan, S.S.W.; Ong, K.H.; Kuperan, P. COVID-19 and mycoplasma pneumoniae coinfection. Am. J. Hematol. 2020, 95, 723–724. [Google Scholar] [CrossRef]
- Wang, L.; Amin, A.K.; Khanna, P.; Aali, A.; McGregor, A.; Bassett, P.; Gopal Rao, G. An observational cohort study of bacterial co-infection and implications for empirical antibiotic therapy in patients presenting with COVID-19 to hospitals in North West London. J. Antimicrob. Chemother. 2021, 76, 796–803. [Google Scholar] [CrossRef]
- Ozaras, R.; Cirpin, R.; Duran, A.; Duman, H.; Arslan, O.; Bakcan, Y.; Kaya, M.; Mutlu, H.; Isayeva, L.; Kebanli, F.; et al. Influenza and COVID-19 coinfection: Report of six cases and review of the literature. J. Med. Virol. 2020, 92, 2657–2665. [Google Scholar] [CrossRef]
- Cuadrado-Payan, E.; Montagud-Marrahi, E.; Torres-Elorza, M.; Bodro, M.; Blasco, M.; Poch, E.; Soriano, A.; Pineiro, G.J. SARS-CoV-2 and influenza virus co-infection. Lancet 2020, 395, e84. [Google Scholar] [CrossRef]
- Konala, V.M.; Adapa, S.; Gayam, V.; Naramala, S.; Daggubati, S.R.; Kammari, C.B.; Chenna, A. Co-infection with Influenza A and COVID-19. Eur. J. Case Rep. Intern. Med. 2020, 7, 001656. [Google Scholar] [CrossRef] [PubMed]
- Verduyn, M.; Allou, N.; Gazaille, V.; Andre, M.; Desroche, T.; Jaffar, M.C.; Traversier, N.; Levin, C.; Lagrange-Xelot, M.; Moiton, M.P.; et al. Co-infection of dengue and COVID-19: A case report. PLoS Negl. Trop. Dis. 2020, 14, e0008476. [Google Scholar] [CrossRef] [PubMed]
- Parvu, V.; Gary, D.S.; Mann, J.; Lin, Y.C.; Mills, D.; Cooper, L.; Andrews, J.C.; Manabe, Y.C.; Pekosz, A.; Cooper, C.K. Factors that Influence the Reported Sensitivity of Rapid Antigen Testing for SARS-CoV-2. Front. Microbiol. 2021, 12, 714242. [Google Scholar] [CrossRef]
- Goncalves, R.L.; Leite, T.C.R.; Dias, B.P.; Caetano, C.; de Souza, A.C.G.; Batista, U.D.S.; Barbosa, C.C.; Reyes-Sandoval, A.; Coelho, L.F.L.; Silva, B.M. SARS-CoV-2 mutations and where to find them: An in silico perspective of structural changes and antigenicity of the spike protein. J. Biomol. Struct. Dyn. 2022, 40, 3336–3346. [Google Scholar] [CrossRef] [PubMed]
- Dejnirattisai, W.; Shaw, R.H.; Supasa, P.; Liu, C.; Stuart, A.S.; Pollard, A.J.; Liu, X.; Lambe, T.; Crook, D.; Stuart, D.I.; et al. Reduced neutralisation of SARS-CoV-2 omicron B.1.1.529 variant by post-immunisation serum. Lancet 2022, 399, 234–236. [Google Scholar] [CrossRef]
- U.S. Department of Health and Human Services Food and Drug Administration; Center for Devices and Radiological Health. Policy for Coronavirus Disease-2019 Tests during the Public Health Emergency (Revised): Guidance for Clinical Laboratories, Commercial Manufacturers, and Food and Drug Administration Staff. 2021. Available online: https://www.fda.gov/media/135659/download (accessed on 2 June 2022).



| Autor/Methods Used | Sensitivity | Specificity/Accuracy | N | Cross-Reaction | ||
|---|---|---|---|---|---|---|
| Day after Symptom Onset | ||||||
| 01–07 | 08–14 | 15–39 | ||||
| Masiá et al. [17]/Immunochromatographic test for rapid detection of SARS-CoV-2 nucleocapsid in nasopharyngeal swab (NPS), swab nasal and saliva samples, Panbio COVID-19 Ag RTD (Abbott). | NPS: 60.5% Saliva: 23.1% Nasal samples: 44.7% | ND | ND | 100%/ND | 913 patients included. 296 were asymptomatic, confirmed by RT-PCR assay. | ND |
| Krüger et al. [18]/Immunochromatographic test for rapid detection of SARS-CoV-2 nucleocapsid in nasopharyngeal swab, PanBio™ COVID-19 Ag Rapid Test Device (Abbott). | 90.8% overall | 61.5% overall | ND | 0–7 days: 99.6% overall/ND 08–14 days: 100% overall/ND | 1108 cases were tested in total. Information on symptom duration from the day of the test in days was available in 687 patients. | ND |
| Kolwijck, et al. [19]/Immunochromatographic test for rapid detection of SARS-CoV-2 nucleocapsid in nasopharyngeal sample, Panbio™ COVID-19 Ag RDT (Abbott). | 86.7% | ND | ND | 100%/ND | 433 participants, with 45 tested positive by RT-qPCR. | ND |
| Rubio et al. [20]/Immunochromatographic test for rapid detection of SARS-CoV-2 nucleocapsid in nasopharyngeal swab, Panbio™ COVID-19 Ag RDT (Abbott). | 72% | ND | ND | 100%/ND | 103 nasopharyngeal swabs were evaluated, confirmed by RT-PCR. | ND |
| L’Huillier et al. [21]/Immunochromatographic test for rapid detection of SARS-CoV-2 nucleocapsid in nasopharyngeal swab, Panbio™ COVID-19 Ag RDT (Abbott). | 66% overall | ND | ND | 100%/ND | 822 pediatric participants completed the study with RT-PCR positive. | ND |
| Villaverde et al. [22]/Immunochromatographic test for rapid detection of SARS-CoV-2 nucleocapsid in nasopharyngeal swab, Panbio™ COVID-19 Ag RDT (Abbott). | 45.4% | ND | ND | 99.8%/97.2% | 1620 pediatric patients aged 0–16 years with symptoms compatible with severe acute respiratory SARS-CoV-2 infection. | ND |
| Muhi et al. [23]/Immunochromatographic test for rapid detection of SARS-CoV-2 nucleocapsid in nasopharyngeal swab, Panbio™ COVID-19 Ag RDT (Abbott). | ND | ND | ND | 99.96%/ND | 2413 subjects tested in hospitals (or associated screening clinics), no subjects tested positive using RT-PCR. | ND |
| 100% | 100%/100% | 26 participants with COVID-19 (as notified to the Victorian Department of Health and Human Services) with the time from symptom onset ranged from 1 to 33 days. | ||||
| Merino et al. [24]/Immunochromatographic test for rapid detection of SARS-CoV-2 nucleocapsid in nasopharyngeal swab, Panbio™ COVID-19 Ag RDT (Abbott). | 90.5% | ND | ND | 98.8%/ND | 958 individuals who had at least one symptom compatible with COVID-19 (n = 830) or who had been in close contact with a diagnosed COVID-19 patient (n = 128) were included. RT-PCR was positive in 359 and negative in 599. | ND |
| Albert et al. [25]/Immunochromatographic test for rapid detection of SARS-CoV-2 nucleocapsid in nasopharyngeal swab, Panbio™ COVID-19 Ag RDT (Abbott). | Adults: 82.6% Pediatric: 62.5% Overall: 79.6% | ND | ND | 100% overall/ND | 412 patients with clinical suspicion of COVID-19, 327 were adults and 85 children. 43 were positive by RT-PCR and 358 were negative. | ND |
| Carbonell-Sahuquillo et al. [26]/Immunochromatographic test for rapid detection of SARS-CoV-2 nucleocapsid in nasopharyngeal swab, Panbio™ COVID-19 Ag RDT (Abbott). | 70.6% | ND | ND | 100%/95.6% | 357 patients (aged 0 to 14 years) with clinical suspicion of COVID-19 (≤5 days since symptom onset). 34 had a positive result by RT-PCR. | 7 out of the 10 specimens yielding discordant results (RT-PCR+/RAD-) were run in a multiplexed PCR assay targeting common respiratory viruses. Two of the 7 specimens tested positive for Rhinovirus/Enterovirus |
| Kim et al. [27]/Immunochromatographic lateral flow assay for rapid detection of SARS-CoV-2 nucleocapsid in nasopharyngeal swab, GenBody™ COVID-19 Ag test (COVAG025). | 90% | ND | ND | 98%/96.15% | 130 residual NPS swabs from individuals who either visited or were hospitalized at Yeungnam University Medical Centre. 30 were confirmed positive for COVID-19 and 100 were designated negative, based on the RT-PCR assay. | ND |
| 94% | 100%/97% | 200 symptomatic and asymptomatic NPS swabs, with 100 samples positive by Real-time PCR for COVID-19. | ||||
| Singh et al. [28]/Immunochromatographic lateral flow assay for rapid detection of SARS-CoV-2 nucleocapsid in nasopharyngeal and oropharyngeal swab, GenBody™ COVID-19 Ag test. | 35.29% | ND | ND | 100%/ND | 240 respiratory samples (nasopharyngeal and oropharyngeal smears) were collected from suspected cases of COVID-19. 102 samples were positive RT-PCR. | ND |
| Möckel et al. [29]/Immunoassay for the rapid determination of the presence of SARS-CoV-2 antigen in human oro-nasopharingeal swabs, Roche SARS-CoV-2 rapid antigen test (Penzberg, Germany). | Adult:75.3% Pediatric:72% | ND | ND | Adult: 100%/ND Pediatric: 99.4%/ND | 473 analyzed patients (n = 271 patients from the adult EDs and n = 202 from the pediatric ED). | ND |
| Jegerlehner et al. [30]/Immunoassay for the rapid determination of the presence of SARS-CoV-2 antigen in human nasopharyngeal swabs, Roche SARS-CoV-2 rapid antigen test (Penzberg, Germany). | 65.3% | ND | ND | 99.9%/ND | 1465 individuals have been tested, including individuals who were referred because of exposure to infected individuals. 141 individuals tested positive according to RT-PCR. | ND |
| Ciotti et al. [31]/Immunochromatographic test for rapid detection of SARS CoV-2 nucleoprotein in nasopharyngeal swabs, COVID-19 Ag Respi-Strip (Coris BioConcept). | 30.77% | ND | ND | 100%/86.15% | 50 sample have been tested, 39 confirmed by real-time RT-PCR. | ND |
| Kanaujia et al. [32]/Immunochromatographic test for rapid detection of SARS CoV-2 in nasopharyngeal swabs, COVID-19 Ag Respi-Strip (Coris BioConcept). | 71.96% | ND | ND | 99.32%/88.64% | 825 symptomatic patients and their contacts were included in the study; RT-PCR and antigen detection were performed simultaneously for 484 samples to determine the sensitivity and specificity of the test. | ND |
| Strömer et al. [33]/Lateral flow chromatographic immunoassay for rapid detection of SARS-CoV-2 nucleoprotein in nasopharyngeal swab, ADAL® COVID-19 Ag (nal von minden GmbH). | 73.1% overall | ND | ND | 99.3%/ND | 134 upper respiratory tract swab samples were used. 124 were positive samples, and 10 were negative samples confirmed by RT-PCR triplex of the N gene. | ND |
| Thakur et al. [34]/Immune-chromatographic lateral flow assay for the rapid determination of the SARS-Cov-2 antigen in nasopharyngeal swabs manufactured by PathoCatch/ACCUCARE, (Lab Care Diagnostics Private Ltd., Mumbai, India). | 34.5% | ND | ND | 99.8%/91.7% | 677 patients have been tested include asymptomatic patients. 55 specimens that tested positive by RT-qPCR. | ND |
| Chaimayo et al. [35]/Chromatographic immunoassay for rapid detection of SARS-CoV-2 nucleocapsid in respiratory samples (mainly nasopharyngeal and throat swabs), Standard™ Q COVID-19 Ag kit (SD Biosensor®). | 98.33% | ND | ND | 98.73%/ND | 454 respiratory samples suspected COVID-19 was evaluated. 60 were positive, and 394 were negative by real-time RT-PCR assay. | ND |
| Amer et al. [36]/Chromatographic immunoassay for rapid detection of SARS-CoV-2 nucleocapsid in oropharyngeal and nasopharyngeal swabs, Standard™ Q COVID-19 Ag kit (SD Biosensor®). | 78.2% | ND | ND | 64.2%/75.9% | 83 oropharyngeal and nasopharyngeal swabs were evaluated of participants should be suspected of having COVID-19 infection. 54 were positive by RT-qPCR. | ND |
| Diao et al. [37]/Fluorescence immunochromatographic (FIC) assay for rapid detection of SARS-CoV-2 nucleocapsid in nasopharyngeal swabs, manufactured by the authors. | 75.6% | ND | ND | 100%/80.5% | 251 participants with suspected COVID-19 symptoms. The Ct value 40 is the cutoff of RT-PCR testing. 201 had a Ct value of ≤40. | ND |
| Mboumba-Bouassa et al. [38]/Chromatographic immunoassay for rapid detection of SARS-CoV-2 nucleocapsid in nasopharyngeal swabs, Ag-RDT SIENNA™ COVID-19 Antigen Rapid Test Cassette (Nasopharyngeal Swab) (SIENNA™). | 90% | ND | ND | 100%/90% | 100 positive and 50 negative RNA swabs from SARS-CoV-2 by reference multiplex rtRT-PCR. | ND |
| Tinker et al. [39]/Lateral flow immunoassay for rapid detection of SARS-CoV-2 nucleocapsid in direct anterior nasal (nares) swabs, BinaxNOW COVID-19 Ag Card (BinaxNOW; Abbott Laboratories). | 20% | ND | ND | 100%/ND | 1540 asymptomatic cases were tested. 40 positives confirmed with RT PCR. | No specimens tested positive for Influenza A or B viruses. |
| Orsi et al. [40]/Qualitative fluorescence immunoassay (FIA) for rapid detection of SARS-CoV-2 nucleocapsid in nasopharyngeal swabs, FREND™ COVID-19 Ag assay (NanoEntek, South Korea). | 93.3% | 86.7% | ND | 100%/ND | 110 nasopharyngeal samples from patients with symptoms attributable to SARS-CoV-2 infection. 60of swabs tested positive by RT-qPCR. | ND |
| Cassuto et al. [41]/Lateral flow immunochromatographic assay for rapid detection of SARS-CoV-2 nucleocapsid in a nasal sample, COVID-VIRO® (AAZ-LMB). | 96.88% | ND | ND | 100%/ND | 234 patients with mild to moderate symptoms lasting less than 7 days and without the need for immediate hospitalization. RT-PCR confirmed 32 positive and 202 negative samples. | ND |
| Rastawicki, et al. [42]/Fluorescent immunoassay (FIA), PCL COVID-19 Ag (SD BIOSENSOR, Korea). | 38.9% | ND | ND | 83.3%/ND | 167 nasopharyngeal swabs. | ND |
| Bianco et al. [43]/Microfluidic immunofluorescence assay for detection of the nucleocapsid protein of SARS-CoV-2 in nasal, LumiraDx™ (LumiraDx GmbH, Colônia, Alemanha). | 90.3% overall | ND | ND | 92.1% overall/ND | 907 patients were evaluated, including participants asymptomatic. As a result, 298 participants were positive for SARS-CoV-2 confirmed by RT-PCR. | ND |
| Krüger et al. [44]/Microfluidic immunofluorescence assay to detect the nucleocapsid protein of SARS-CoV-2 in nasal mid-turbinate (NMT) self-swab manufactured by LumiraDx™ (LumiraDx™, London, UK). | 82.2% | ND | ND | 99.3%/ND | 761 samples were evaluated, of which 146 were RT-PCR positive and 615 negatives. | The respiratory swab samples contained four seasonal coronaviruses, Adenovirus, Bocavirus, Influenza virus, Metapneumovirus, Parainfluenza virus, Respiratory syncytial virus, Rhinovirus or Mycoplasma pneumoniae, Staphylococcus aureus and Streptococcus sp. No cross-reactivity was detected. |
| Drain et al. [45]/Microfluidic immunofluorescence assay to detect the nucleocapsid protein of SARS-CoV-2 in nasal and nasopharyngeal (NP) swab manufactured by LumiraDx™ (LumiraDx™, Reino Unido). | ND | Up to 12 days: 97.6% (nasal swab) 97.5% (NP swab) | ND | 96.6% (nasal swab)/ND 97.7% (NP swab)/ND | 512 participants, aged 0–90 years. | ND |
| Liu et al. [46]/Nanozyme chemiluminescence paper test for rapid and sensitive detection of SARS-CoV-2 antigen, manufactured by the authors. | ND | ND | ND | ND/ND | Viral samples were recombinant peak proteins (2 × 104 TCID50/mL titer) and 98% purity confirmed by SDS-PAGE. | Other human coronaviruses (SARS-CoV, MERS-CoV, HCoV-HKU1 and HCoV-OC43) were tested to validate the specificity. There was no cross-reaction with other coronaviruses or Influenza A subtypes. |
| Egerer et al. [47]/Reverse transcriptase-loop-mediated isothermal amplification (RT-LAMP) for rapid detection of SARS-CoV-2 N and ORF8 genes in oropharyngeal, eazyplex® SARS-CoV-2 (Amplex Diagnostics, Gars Bahnhof, Germany). | Ct ≤ 28: 97.4% | ND | ND | 100%/ND | 150 oropharyngeal and nasal swabs were evaluated and confirmed by RT-PCR. | ND |
| Wang et al. [48]/Multiplexed RT-LAMP microwell biochip for rapid detection of SARS-CoV-2 ORF1ab gene in throat swab, manufactured by the authors. | 95.4% | ND | ND | 95.35%/ND | 87 samples from PCR-positive and 43 PCR-negative patients. | ND |
| Xun et al. [49]/Scalable and Portable Testing (SPOT) assay comprise a one-pot RT-LAMP followed by PfAgo-based target sequence detection (detecting the N gene and E gene in a multiplexed reaction) in clinical saliva samples. | 93.3% | ND | ND | 98.6%/ND | 104 in total saliva samples, confirmed by qRT-PCR. | Samples of saliva were spiked with or without SARS-CoV-2, three other genomic RNA from human coronaviruses (OC43, 229E, and NL63), SARS, and MERS viruses (γ-irradiated), and Influenza. Among these samples, SARS-CoV-2 genes were only detected in the positive control. |
| de Oliveira Coelho et al. [50]/Reverse Transcription Loop-mediated Isothermal Amplification (RT-LAMP) for the rapid determination of SARS-CoV-2 copies in nasal and oral rayon-swabs samples, manufactured by the authors. | 93.8% | ND | ND | 90.4%/ND | 466 samples were evaluated. 250 were negative, and 216 were positive by RT-qPCR. | ND |
| Wei et al. [51]/High-Performance Loop-mediated isothermal Amplification (HP-LAMP) for the rapid determination of SARS-CoV-2 copies in saliva samples. | >96% | ND | ND | >96%/ND | 120 samples were evaluated. | Wet testing was performed to evaluate potential cross-reactivity of the assay with other organisms using ZeptoMetrix Corporation NATtrol Respiratory Verification Panel (ZeptoMetrix, NATRVP-IDI), including 19 respiratory pathogens, NATtrol Coronavirus-SARS Stock, NATtrol MERS-CoV Stock, and NATtrol SARS-Related Coronavirus 2 External Run Control. No cross-reactivity was detected. |
| Lee et al. [52]/CRISPR Optical Detection of Anisotropy for rapid detection of SARS-CoV-2 copies in a nasopharyngeal or oropharyngeal swabs or sputum, manufactured by the authors. | ND | ND | ND | ND/ND | 20 clinical specimens of nasopharyngeal and oropharyngeal smears and sputum from individuals suspected of COVID-19. 10 COVID-19 positive samples and 10 negative samples were confirmed by qRT-PCR. | ND |
| Li et al. [53]/reverse transcription recombinase-aided amplification (RT-RAA) for rapid detection of SARS-CoV-2 in throat swab. | Ct (22.1 to 32.8): 98% Ct (33.2 to 36.4): 33% | ND | ND | 100%/ND | 80 throat swab specimens were collected from the suspected SARS-CoV-2 infectious patients, confirmed by qRT-PCR. | Were tested 8 respiratory RNA viruses, including Influenza A viruses (H3N2, H7N9, H5N1, H1N1) and Influenza B viruses (Victoria and Yamagata lineages), which were isolated from humans or birds and adenoviruses (AdV3 of strain IVCAS16(A).00027 and AdV7 of strain IVCAS 16(A).00028). The non-targeted RNA genomes generated similar baselines as the negative control (water). |
| Margulis et al. [54]/Magnetic modulation biosensing (MMB) for rapid detection of SARS-CoV-2 in nasopharyngeal swabs. | 97.8% | ND | ND | 100%/ND | Were tested 309 clinical samples from SARS-CoV-2–positive and SARS-CoV-2–negative patients with a wide range of initial viral loads (CT ≤ 42). | 30 nasopharyngeal swab samples collected in 2019 from patients with different viral respiratory diseases, such as influenza A (13 samples), influenza B (10 samples), and respiratory syncytial virus (7 samples), were tested to evaluate the specificity of the assay further. All SARS-CoV-2 samples were identified as negative. |
| Torres et al. [55]/RAPID 1.0 (real-time accurate portable impedimetric detection prototype 1.0) in saliva, nasopharyngeal and oropharyngeal swabs, manufactured by the authors. | NP/OP: 83.5% Saliva: 100% | ND | ND | NP/OP: 100%/87.1% Saliva: 86.5%/90% | A total of 151 saliva and (NP/OP) swabs were evaluated, confirmed by RT-PCR | It performed specificity assays with three coronaviruses (MHV-Murine hepatitis virus, HCoV-OC43–human coronavirus OC43 and human coronavirus 229E) and four non-coronavirus viral strains (H1N1-A/California/2009, H3N2-A/Nicaragua, Influenza B-B/Colorado, HSV2-Herpes simplex virus-2). No cross-reactivity was detected. |
| Autor/Methods Used | Sensitivity | Specificity/Accuracy | N | Cross-Reaction | ||
|---|---|---|---|---|---|---|
| Day after Symptom Onset | ||||||
| 01–07 | 08–14 | 15–39 | ||||
| Kumar et al. [56]/Electrochemical ELISA platform detects both IgM and IgG antibodies against the SARS-CoV-2 Spike Glycoprotein (S1) in clinical whole blood and serum samples. anuPath™ Electrochemical ELISA Analyzer. | ND | ND | 100% | 100%/100% | 450 samples were evaluated, of which 252 were EDTA whole blood samples and 198 were sera samples. | ND |
| Munck et al. [57]/COVID-19 IgG/IgM Duo is a fluorescent lateral flow immunoassay detecting both IgM and IgG antibodies against the SARS-CoV-2 nucleocapsid (N) protein separately in serum samples. NanoEntec. | 46.2% | 93.8% | 100% | IgM: 87.5% IgG: 91.7% IgG/IgM: 95.8%/ND | 105 serum samples were evaluated, confirmed by PCR. | False-positive results were found in two samples with antibodies to other coronaviruses (NL63 and HKU1) and one sample with Epstein Barr viral capsid IgG. |
| Peng et al. [58]/Lateral Flow Immunoassay test for rapidly detecting of specific IgM and IgG antibodies against the nucleocapsid and S-RBD protein recombinant of SARS-CoV-2 in serum samples, manufactured by the authors. | Anti-N IgG: 96.6% Anti-N igM: 96.6% Anti-S-RBD-IgG: 95.9% Anti-S-RBD-IgM: 96.6% | ND | ND | Anti-N IgG: 94.1%/ND Anti-N igM: 100%/ND Anti-S-RBD-IgG: 96.1%/ND Anti-S-RBD-IgM: 100%/ND | 108 serum samples were tested, 82 positives of 83 confirmed with CLIA | ND |
| Kiziloglu et al. [59]/Gold Immunochromatography for the rapid detection of anti-SARS-CoV-2-IgG and IgM manufactured by Bioeasy COVID-19 Coronavirus IgG/IgM GICA Rapid Test Kit ©. | 57.5% | ND | ND | 85.5%/ND | 181 cases were tested: 71 positives were clinically confirmed (RT-PCR test). | ND |
| Plebani et al. [60]/Immunochromatographic assay that detects IgM and IgG antibodies against the spike protein SARS-CoV-2 (S1), and the nucleocapsid protein in a serum or plasma sample. COVID19SEROSpeed-IgM-IgG (BioSpeedia). | ND | ND | 92.5% | 98.1%/ND | 710 samples. For specificity analysis, 215 pre-pandemic serum samples were analyzed. For sensitivity analysis, the samples were positive by qRT-PCR. | Four false-positive samples tested positive for IgG in three of them and IgM in one of them; two of these four patients exhibited IgM targeted to Mycoplasma pneumoniae and Toxoplasma gondii, respectively. |
| Villarreal et al. [61]/Gold Immunochromatography for the rapid detection of specific IgM and IgG antibodies against SARS-CoV-2 nucleocapsid (N) protein in serum samples, manufactured by the group of Dr. Chong Li of the Institute of Biophysics of the Chinese Academy of Sciences. | 36.4% for IgM and IgG | 76.2% (IgM) and 71.4% (IgG) | 87.0% for IgM and IgG | 95.0%/ND | 810 serum samples, confirmed by qRT-PCR and a set of pre-pandemic panel samples. | Cross-reaction was tested on pre-pandemic samples from patients with Dengue, pulmonary tuberculosis, and latent tuberculosis infection. Only one patient who tested positive for Dengue showed a positive IgM. |
| Scotta et al. [62]/Lateral flow for the rapid detection of specific IgM and IgG antibodies against the spike protein SARS-CoV-2 in the blood sample, manufactured by Wondfo Biotech. | ND | 60.0% | 73.2% | 96.8% overall/ND | 175 whole blood samples of pediatric patients were evaluated, confirmed by qRT-PCR | ND |
| Fauziah et al. [63]/Lateral flow qualitative immunoassay for rapidly detecting specific IgM and IgG antibodies against SARS-CoV-2 in serum and capillary blood samples, manufactured by Guangzhou Wondfo Biotech Co., Ltd., Guangzhou, China | Serum: 63%. Capillary blood: 41.2% | ND | ND | Serum: 95 %/78.7% Capillary blood: 100%/50% | 47 sample, 27 patients had a positive rRT-PCR result. | Cross-reaction was tested on serum samples from a patient with Dengue or typhoid fever. No cross-reactivity was observed. |
| Pallett et al. [64]/Lateral flow immunoassay for the rapidly detecting specific IgM and IgG antibodies against SARS-CoV-2 in blood samples, OrientGene COVID-19 split IgG/IgM (OrientGene). | 74% | 86% | 100% | 96%/ND | 200 samples, 50 negatives and 130 positives of the 150 positives were confirmed with RT PCR. | ND |
| Prendecki et al. [65]/Lateral flow immunoassay for rapidly detecting specific IgG antibodies against the spike protein SARS-CoV-2 in blood, serum, and plasma sample, manufactured by Biomedomics Inc. | ND | ND | 96.6% | 97.7%/97.3% | Sixty samples were collected from maintenance hemodialysis patients and kidney transplant recipients. All patients had undergone RT-PCR testing. And 88 plasma samples were pre-pandemic. | ND |
| Zhang et al. [66]/Magnetofluidic immuno-PCR platform assay for anti-SARS-CoV-2 Immunoglobulin G (IgG) detection using a magnetofluidic instrument, manufactured by the authors. | ND | ND | 93.8% overall | 98.3%/ND | Were tested 108 samples, 34 of which were 34 convalescent plasma samples from patients with SARS-CoV-2 confirmed with the Roche ECLIA test (Elecsys® Anti-SARS-CoV-2), 14 convalescent serum samples confirmed by a custom serological test based on beads and 40 serum samples and 20 plasma samples as negative controls. | ND |
| Elledge et al. [67]/Split luciferase (spLUC) antibody sensor for the rapid detection of specific antibodies against the nucleocapsid and S-RBD protein recombinant of SARS-CoV-2 in serum, plasma, whole blood, and saliva samples, manufactured by the authors. | ND | ND | anti-S protein antibodies: 89% anti-N protein antibodies: 98% | anti-S protein antibodies: 100% anti-N protein antibodies: 99%/ND | Testing of over 150 patient serum/plasma samples. | They were tested for seasonal coronavirus patient samples and 20 pre- and post-vaccination influenza vaccine samples. These controls generated significantly lower luminescent signals than the COVID-19 patient sera samples. |
| Li et al. [68]/Lateral flow assay test for the rapid detection of specific IgA, IgM and IgG antibodies against the recombinant N protein and recombinant S1 protein of SARS-CoV-2 in serum and plasma samples manufactured by the authors. | ND | ND | 88.56% | 88.56%/ND | The kit was used to test the serum from 43 suspected COVID-19 patients, 97 COVID-19 patients, and 88 with general fever or pulmonary infection patients. | Cross-reactivity in patients infected with M. pneumoniae and respiratory tract infection. |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
de Paula Dias, B.; Gonçalves, R.L.; Ferreira, C.S.; Barbosa, C.C.; Arrieta, O.A.P.; dos Santos, S.M.S.A.; Malta, W.C.; Alves e Silva, M.; Gomes, M.L.M.D.; Guimarães, A.G.; et al. Update on Rapid Diagnostics for COVID-19: A Systematic Review. Int. J. Transl. Med. 2022, 2, 252-274. https://doi.org/10.3390/ijtm2020023
de Paula Dias B, Gonçalves RL, Ferreira CS, Barbosa CC, Arrieta OAP, dos Santos SMSA, Malta WC, Alves e Silva M, Gomes MLMD, Guimarães AG, et al. Update on Rapid Diagnostics for COVID-19: A Systematic Review. International Journal of Translational Medicine. 2022; 2(2):252-274. https://doi.org/10.3390/ijtm2020023
Chicago/Turabian Stylede Paula Dias, Bruna, Ricardo Lemes Gonçalves, Cyntia Silva Ferreira, Camila Cavadas Barbosa, Orlando Alfredo Pineda Arrieta, Samara Mayra Soares Alves dos Santos, Wellington Carvalho Malta, Mariela Alves e Silva, Maria Laura Maximiano Dias Gomes, Adriana Gibara Guimarães, and et al. 2022. "Update on Rapid Diagnostics for COVID-19: A Systematic Review" International Journal of Translational Medicine 2, no. 2: 252-274. https://doi.org/10.3390/ijtm2020023
APA Stylede Paula Dias, B., Gonçalves, R. L., Ferreira, C. S., Barbosa, C. C., Arrieta, O. A. P., dos Santos, S. M. S. A., Malta, W. C., Alves e Silva, M., Gomes, M. L. M. D., Guimarães, A. G., Borges, L. P., & de Mello Silva, B. (2022). Update on Rapid Diagnostics for COVID-19: A Systematic Review. International Journal of Translational Medicine, 2(2), 252-274. https://doi.org/10.3390/ijtm2020023








