Abstract
Microscopy slides are routinely created as part of sexual assault workflows for screening purposes and retained indefinitely with instances, such as cold cases, where they are the only remaining source of evidence. To date, no method has been developed to harvest the cells from these slides for differential extraction using the i-sep® DL column or Intimate extraction using the PrepFiler™ Express chemistry supplemented with 60 mM DTT. This study used mock sexual assault slides from 2010 to develop a potential cell harvesting method, then tested both the extraction methods on historic casework slides collected in the 1980s and 1990s. Key findings included the necessity to re-screen slides microscopically to assess current cellular loading and the utility of using phase contrast to enhance visualisation of spermatozoa on historic slides. Both extraction methods successfully recovered DNA and generated partial profiles from the 1990s slides, but 1980s slides were too degraded to provide informative profiles. Differential extraction provided additional contextual information by separating contributors into sperm and non-sperm fractions, resulting in cleaner sperm profiles for interpretation. The study successfully established methods for historic microscope slides to be used as viable sources of evidence, potentially aiding in the progression and resolution of sexual assault investigations.
1. Introduction
Advancements in technology have progressively allowed greater sensitivity and discrimination when processing exhibits for forensic DNA analysis. These method optimisations aim to facilitate maximum DNA recovery and generate profiles suitable for searching against databases for identification. This is especially advantageous for cold case investigations, where new leads of enquiry may prove crucial to resolving crimes which remain unsolved. DNA evidence can be crucial to the progression of a sexual assault case as eyewitnesses are often not available, the victim may be unable to recall the details of the offence and/or offender as well as potential issues with recall accuracy [1,2,3]. Sexual assault kits (SAK’s) are routinely collected during the investigation of alleged sexual offences and include the recovery of samples such as hairs, fibres, fingernail scrapings and bodily fluids including saliva, semen and blood [4]. Collection media can vary between jurisdictions but often includes gauzes and swabs, which can be examined in DNA analysis workflows. Due to the consumption of samples during testing, one of the fundamental limitations facing cold case investigations is obtaining sufficient evidentiary sample to undergo testing using novel or updated methodologies.
The differential extraction method to separate sperm and non-sperm cells into individual DNA extracts is commonly employed within forensic laboratories processing samples from sexual assault casework, with continuing development and optimization of the method [5,6,7,8,9]. This separation facilitates clearer resolution of individual contributor profiles from a mixed cell sample, increasing the likelihood of generating informative profiles [5,9]. Screening samples is an important aspect of SAK processing as the identification of sperm cells can provide contextual information relating to the case and direct the sample for the appropriate DNA analysis method [10]. A common screening method uses the SAK swabs to create smears on microscope slides which are then stained with traditional stains such as Christmas Tree, or newer fluorescence-based methods such as Sperm Hy-Liter™, which selectively stain sperm heads [11,12]. This simplifies screening by making sperm cells more distinguishable amongst heavy epithelial loading or cells with a similar visual appearance, such as yeast [12]. After identifying the presence of sperm cells, eligible swabs are then directed for the appropriate processing with slides entered into long-term storage as they were not the primary source of evidence.
Due to elapsed time and consumption of samples during previous testing, cold case sexual assault investigations often have no original exhibits available for additional analysis. In these cases microscopy slides created for screening may provide the only potential source of biological material, even though they were not originally intended for DNA processing. Cold case microscopy slides present a unique challenge for processing as dye stains can inconsistently fade over time, reducing both the visual identification of spermatozoa and the ability to target areas of the slide to recover sperm cells [11,13]. The alternative is to swab the entire slide surface in an attempt to collect all cells but the efficacy of cell harvesting in this manner is uncertain, with previous in-house work failing to yield quantifiable results.
The services provided by forensic laboratories have evolved from simple screening and presumptive testing of suspected semen stains to advanced DNA extraction and comparison of resulting profiles against both State and National DNA databases. A recent development was the use of the i-sep® DL spin column for differential separation and extraction of sexual assault samples [14]. Unlike conventional differential separation methods which rely on pelleting the sperm cells, the i-sep® method allows for direct lysis of the sample substrate which maximises the recovery of DNA in both sperm and non-sperm fractions. This increase in extraction efficiency has been shown to improve from ~50% for traditional pellet differential extraction to >99% sperm cell recovery using the i-sep® method [14]. Other laboratories employ direct whole sample lysis on intimate samples with no sperm cell specific lysis steps, instead performing mixture analysis on the final profiles [15]. Irrespective of method, the addition of dithiothreitol (DTT) is common in sexual assault workflows to facilitate cell lysis and DNA release from sperm cells [5,9].
Studies into the applicability of using microscope slides as a source of forensic evidence have been promising. A 1997 study demonstrated that DNA could be recovered from cytological smears, with the conclusion that this has value for sexual assault casework by providing new lines of enquiry for investigators [16]. In pathology laboratories, slides are often created from tissue using formalin-fixed, paraffin-embedded (FFPE), cytologic preparations or direct smears [17]. These are routinely processed by scraping the surface of the slide using a scalpel blade as a means of cell harvesting, before undergoing DNA analysis [17,18]. These laboratories often face a similar issue as cold case investigations where limited sample exists for analysis, prompting the development of methods to use pathology diagnostic slides as sources of DNA.
Applying the similar principles of slide processing undertaken in pathology laboratories to forensic workflows is not without precedent. The utility of this was demonstrated in a United States review of unsolved sexual assault homicides where microscopy smears underwent re-screening and extraction [19]. Six cases were identified to be unsolved and have visible sperm cells present, with profiles obtained from all six slides, resulting in both identification and exclusion of suspects. The processes used were simple and cost-effective, utilising an already implemented differential extraction method and simply swabbing of the slides for cell collection. While a promising study, no measure of recovery rates to ascertain the efficiency of cell collection and whether swabbing was sufficient to collect all cells on the slide surface was performed.
The differential separation and extraction method will remain a core process in sexual assault workflows, so developing a method for processing microscopy slides with the i-sep® differential separation (DL) would be advantageous. To date, no research has been published on the combination of scraping historic microscope slides for cell harvesting and processing using either the i-sep® DL or direct extraction methods undertaken in this study. Additionally, the use of multiple slides for scraping and pooling into one extraction has not been explored. Using a stepwise testing approach over 3 phases of increasing microscope slide sample age, this study aims to develop a method for maximising the collection of cells and evaluate the DNA recovery efficiency of direct extraction or i-sep® DL methods from archived microscope slide smears from sexual assault investigations.
2. Materials and Methods
2.1. Microscope Slide Samples
Both mock-samples and real casework samples were processed as part of this study which was separated into three distinct phases. Phase 1 used mock samples previously prepared for a 2010 project verifying the use of Christmas tree staining for sperm cell screening. These samples consisted of buccal cells from a known female donor and sperm cells from a known male donor were pipetted onto the surface of glass slides, dried on a drying rack before staining with either Crystal violet or Christmas tree stains [20,21]. Slides were then coverslipped with Depex mounting medium, dried in an oven overnight at 37 °C and stored at room temperature long term. Phase 2 and 3 comprised of historic Casework samples from the 1990s and 1980s, respectively. All slides used in Phases 2 and 3 were from solved cases and approved for use by the relevant authority. These historic slides were prepared from either swabs which were smeared across the slide surface or stains moistened with a drop of water and scraped across the slide surface. Slides were then dried on a hot plate followed by heat fixation by passing the slide face down through a Bunsen flame. Not all slides were coverslipped and/or stained with crystal violet before being placed into long term storage at room temperature.
2.2. Light Microscopy
Light microscopy was used to visualise any stained material, with methodical passes over the slide surface to ensure all areas were assessed. Microscope slides were re-screened using the Olympus BX53 light microscope and images captured using the cellSens Entry Imaging Software (Olympus, Tokyo, Japan) on 20× or 40× magnification, and scored using the criteria listed in Table 1.

Table 1.
Scoring criteria based on number of spermatozoa microscopically observed on either Christmas Tree or Crystal Violet stained slides at specified magnifications and the corresponding recorded result.
On historic casework slides which had lost their visible staining, microscope screening was performed on the same Olympus BX53 light microscope using phase contrast, or brightfield microscopy without the top lens to visualise any unstained cellular material deposited on the slide surface. Slides with spermatozoa visually confirmed were deemed candidates for processing with the remainder returned to storage.
2.3. Microscope Slide Preparation
The front and back of any cover slipped slides were wiped with 70% ethanol (Rowe Scientific, Wangara, Australia) before incubation in a Xylene (Rowe Scientific, Wangara, Australia) bath for three days to dissolve the Depex mounting media. Using a new sterile scalpel blade for each sample, the coverslip was carefully separated from the microscope slide, followed by dipping into an absolute ethanol (Rowe Scientific, Wangara, Australia) bath to remove any residual Depex and allowed to air dry. Non-coverslipped slides were wiped with ethanol on the back only, with care taken to ensure that the correct side was selected for wiping.
2.4. Microscope Slide Cell Harvest
Sixty microlitres of the appropriate lysis buffer (non-sperm lysis buffer (0.1 M NaCl, 10 mM Tris, 1 mM EDTA, pH 8.0, 0.8 mg/mL Proteinase K) for i-sep® samples, Prepfiler™ lysis buffer (Thermo Fisher Scientific, Waltham, MA, USA) supplemented with 60 mM DTT (Bio-Strategy, Campbellfield, Australia) for direct extraction samples) was pipetted onto the slide. Using a fresh sterile scalpel blade, the surface of the slide was scraped to dislodge any cellular material on the surface. An excised Copan rayon swab head (Copan Diagnostics Incorporated, Murrieta, CA, USA) was then used to collect all of the liquid with cellular material from the microscope slide and the surface of the scalpel blade and immediately placed into the appropriate tube (i-sep® DL column or Lysep column). A visual check of the slide and blade was made to ensure all liquid had been collected and the scalpel blade discarded. Once all samples had been prepared, the remainder of the appropriate buffer (to a final volume of 500 μL) was pipetted into the sample tubes. For Phase 3 testing where two different slides were combined into a single extraction, 30 μL of the appropriate lysis buffer was applied to each slide and half a cut swab head used to collect the liquid.
2.5. i-sep® Differential Separation (DS)
The i-sep® differential separation was performed as described in White et al. [14]. Samples were incubated using non-sperm lysis buffer for 1 h at 56 °C, on a thermal shaker set to 800 rpm. Samples were then centrifuged at 4500× g before the i-sep® column containing the substrate was transferred to a new collection tube, with the original tube containing the non-sperm fraction. The samples then underwent a second 1 h incubation using 500 µL of non-sperm lysis buffer with additional 2% SDS (PathWest Media, Nedlands, Australia), before three washes using non-sperm lysis buffer. Samples underwent a final incubation for sperm lysis using Prepfiler™ lysis buffer supplemented with 60 mM DTT for 40 min at 70 °C on a thermal shaker set to 750 rpm.
2.6. Direct Extraction Method
Samples undergoing direct extraction were incubated in 500 µL of PrepFiler™ lysis buffer supplemented with 60 mM DTT for 40 min at 70 °C, 750 rpm in a thermal shaker. After incubation, the samples were centrifuged at 10,000× g for 2 min to collect the lysate for DNA extraction.
2.7. DNA Extraction
The non-sperm fractions from the i-sep® differential method were concentrated to ~200 µL using the Microcon® DNA Fast Flow Centrifugal Filter Unit (Merck, Darmstadt, Germany), followed by DNA extraction using the PrepFiler Express™ Forensic DNA Extraction Kit and AutoMate Express™ System (Thermo Fisher Scientific, Waltham, MA, USA). For sperm cell lysis, sperm fractions were incubated in the i-sep® DL spin column with 500 µL of PrepFiler™ lysis buffer supplemented with 60 mM DTT for 40 min at 70 °C, 750 rpm in a thermal shaker. After incubation, the samples were centrifuged at 15,700× g for 5 min to collect the lysate, from which DNA was extracted on the AutoMate Express™ DNA extraction system.
2.8. DNA Quantification and STR Analysis
Samples were quantified using the Quantifiler™ Trio DNA Quantification Kit (Thermo Fisher Scientific, Waltham, MA, USA) on the Applied Biosystems 9700 Real-Time PCR System (Thermo Fisher Scientific, Waltham, MA, USA) and analyzed using HID real-time PCR analysis software v1.3 (Thermo Fisher Scientific, Waltham, MA, USA). Quantification metrics included the Short Autosomal (SA) which asses the total DNA within the sample, the male component (Y) as well as the Degradation Index (D.I). The D.I is presented as a numerical value, with greater degradation associated with increasing values. This degradation can impact profiling, as disproportionate reduction of allele amplification at loci associated with larger DNA fragments leads to a ski-slope morphological appearance of final electropherograms [22].
STR profiling was performed using the PowerPlex® 21 System (PP21) (Promega Corporation, Madison, WI, USA). DNA input was targeted to 0.5 ng, with samples being diluted according to an internally verified dilution table. For low-level samples unable to reach the target input a maximum loading of 15 μL of sample was added, to achieve a final PCR volume of 25 μL. All samples underwent 30 cycles of amplification on the ProFlex™ PCR System (Thermo Fisher Scientific, Waltham, MA, USA), followed by capillary electrophoresis using the Applied Biosystems 3500xL Genetic Analyser (Thermo Fisher Scientific, Waltham, MA, USA) at 1.2 kV for 24 s injection parameters. Profile analysis was performed using the GeneMapper ID-X software v1.6 (Thermo Fisher Scientific, Waltham, MA, USA), with internally validated limit of detection (LOD) threshold of 100 RFU and limit of quantification (LOQ) of 250 RFU. Homozygote peaks had equal or greater than 1200 RFU with heterozygote alleles having to balance ratio of 50% or greater to be considered passing. A profile was deemed to be informative if it contained passing loci at a minimum of six of the 21 loci within the PowerPlex® 21 kit.
3. Results
3.1. Phase 1—Evaluation of DNA Recovery and DNA Quality of Scraped Cellular Material from Cover Slipped Crystal Violet and Christmas Tree Stained Mock Sexual Assault Microscope Slide Smears of Decreasing Sperm Scores Prepared in 2010
All slides had visible purple (crystal violet) or green (Christmas tree) staining (Figure 1), indicating cellular loading was present on the surface.
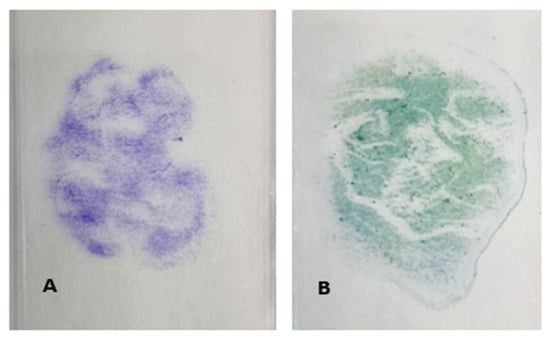
Figure 1.
Representative images of crystal violet (A) and Christmas tree (B) stained slides used in Phase 1 testing.
Microscopy screening confirmed the presence of sperm and non-sperm cells on the slides for all scored slides (Figure 2).
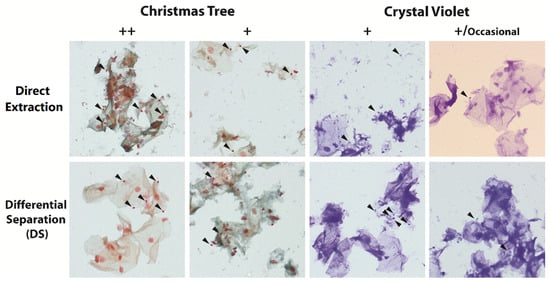
Figure 2.
Representative images of slides from displaying different microscopy scoring of 2010 stained with Christmas Tree or Crystal Violet and extracted using either the direct extraction or differential separation method. Black arrowheads indicate intact sperm heads.
Quantification results from harvested cellular material off the microscope slides indicated successful DNA recovery from all samples processed by both the direct extraction and i-sep® DS methods, irrespective of staining with crystal violet or Christmas tree stains (Table 2).

Table 2.
Comparison of DNA recovery, quality and PP21 passing loci from Secondary Phase 1 slides processed by either direct extraction or differential separation extraction methods for DNA recovery from crystal violet (CV) and Christmas tree (CT) stained slides from 2010.
Y-DNA was reported in both non-sperm and sperm fractions of the i-sep® samples, with increased Y to SA ratios as well as greater total Y values in the sperm fractions. DI values for all samples processed by the direct extraction method were higher than the corresponding i-sep® DS samples, most prominent with the crystal violet stained samples. All i-sep® DS sperm fractions demonstrated successful separation of the contributors with single source male profiles obtained for three samples, with the final sample a two-person mixture that contained a clear male major profile. All samples processed by the direct extraction method and non-sperm fractions from i-sep® DS had Y quantification results that were lower than the short autosomal concentration, indicating multiple contributors. This was reflected in the electropherograms as all contained 2-person major female/minor male profiles.
3.2. Phase 2—Evaluation of DNA Recovery and DNA Quality of Scraped Cellular Material from Historic Casework Sexual Assault Microscope Slide Smears from the 1990s
Results from 1990s slides were more variable than Phase 1, with sperm cells not observed on all slides. This included slides which had previously been scored ‘+++’ indicating large numbers of spermatozoa observed during the original examination, yet this was not replicated in contemporary screening (Figure 3).
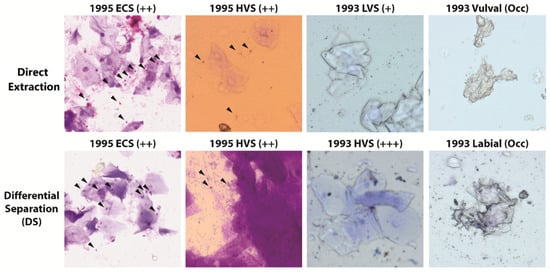
Figure 3.
Representative images of historic slides from sexual assault cases stained with crystal violet and extracted using the direct extraction or differential separation method. Black arrowheads indicate intact sperm heads. ECS = Endocervical Swab, HVS = High Vaginal Swab and LVS = Low Vaginal Swab.
Both ‘occasional’ scored slides provided no usable genetic information as no loci were detected when processed by either the direct extraction or i-sep® method (Table 3). A further direct extraction sample did not recover sufficient DNA to generate an informative profile. With the exception of a single sample, all i-sep® sperm fractions were able to generate informative profiles, with final profiles exhibiting ski-slope morphology of varying severity.

Table 3.
Comparison of DNA recovery, DNA quality and passing PP21 loci from cellular material harvested from crystal violet stained casework slides from the 1990s processed using direct extraction and differential separation extraction methods.
While the majority of male DNA was quantified in the sperm fraction, the non-sperm fraction all had Y-quantification results, indicating male DNA to be present in all samples. Final profiles did not necessarily reflect this, with sample 1993 high vaginal swab (HVS) appearing as a single source female profile for the non-sperm fraction and a two-person mixture in the sperm fraction due to female carryover. Both of the samples processed by the direct extraction method which had reportable DNA quantification resulted in mixed final profiles from two contributors.
3.3. Phase 3—Evaluation of DNA Recovery and DNA Quality of Scraped Cellular Material from Historic Casework Sexual Assault Microscope Slide Smears from the 1980s
Screening of the 1980s slides showed that the slides had very little to no visible cellular material clearly identifiable on the slide surface, which was reflected in the lack of material when viewed microscopically. Most of the structures seen were amorphous ‘jelly like’ or debris such as dust on the slide and not representative of any biological material. Utilising phase contrast or brightfield microscopy without the top lens, the morphology of spermatozoa (if present) was enhanced allowing for easier visual identification on historic casework slides (Figure 4).
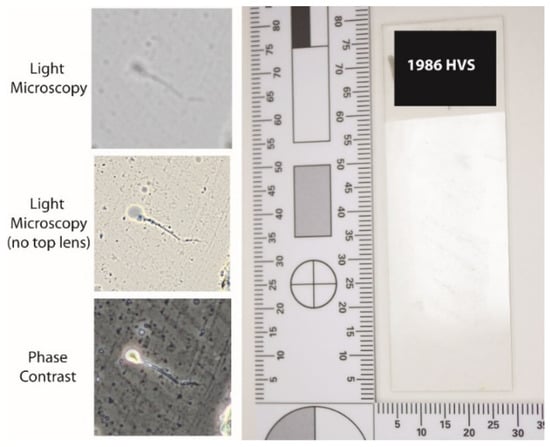
Figure 4.
Representative overview of historic casework slide from the 1980s as well as images showing the different visual appearance of the same spermatozoa with different microscopy conditions.
This increase in spermatozoa visualization was most pronounced on slides where epithelial cell loading was in excess to sperm cells and/or slides where the stain had bleached out (Figure 5). Overall, phase contrast provided the greatest enhancement to sperm cell identification as the contrast between the sperm head and background was more pronounced than just top lens removal (Figure 5).
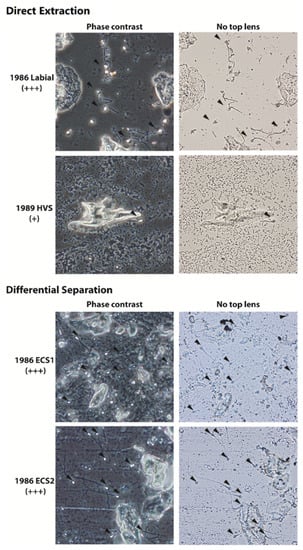
Figure 5.
Representative phase contrast and light microscope (captured without top lens) images of historic slides from 1980s sexual assault cases stained with crystal violet and extracted using the direct extraction or differential separation method. All slides had no visible stain remaining at the time of rescreening. Black arrowheads indicate intact sperm heads.
The comparison between individual slide extraction and pooling of slides into the same extraction failed to yield improvements in the recovery of DNA (Table 4). No samples from Phase 3 reported any Y quantification result, indicating no male DNA had been recovered from any slides (Table 4). Degradation Index value were not able to be established for any of the samples, indicating significant degradation beyond the limits of the quantification kit. Due to the lack of observed spermatozoa and poor recovery of DNA, STR profiling was not performed on these samples.

Table 4.
Summary of results from Casework slides from the 1980s processed using direct extraction and i-sep® extraction methods on quantification metrics.
4. Discussion
This study demonstrated that cellular material from microscope slides may be harvested by scraping using a sterile scalpel blade, with DNA recovery achieved by both the i-sep® differential separation and direct extraction methods. Microscopy became a critical part of the study, facilitating the visualisation of spermatozoa on the slides to ensure the appropriate samples were selected for processing. It is well established that DNA will degrade over time at varying rates due to internal and external factors on the sample such as environment pre and post-sampling, testing applied and storage [23,24]. Even the collection site can have a significant impact on the persistence of sperm cells, which can directly affect the number of spermatozoa that are able to be viewed microscopically [24].
The slides in this study were up to 36 years old and all were stored in cardboard boxes at room temperature. There is little doubt that age alone would be a significant contributor to the levels of DNA degradation observed across all three experimental phases. As such, the state of cells on the slides are not necessarily going to reflect their original microscopy scoring and it became apparent that the re-screen was required to ensure the most appropriate slides were being processed. As it is recognised that dye stains will fade over time, the observation of reduced evident staining on the historic slides was not unexpected [11,13]. This reduction of staining made it challenging to observe sperm cells on the historic slides using traditional microscopy, however several easily implemented variations proved beneficial, including the removal of the top lens which allowed greater depth, making sperm easier to locate. The use of phase-contrast microscopy was demonstrated to be advantageous in slide-screening as the greatly enhanced contrast facilitated more readily observable sperm cells on slides. The difference in visualisation was significant enough that sperm cells were located on slides which had previously been scored negative when using traditional light microscopy.
While the study successfully recovered DNA from the slides from 2010 and the 1990s, all slides from the 1980s failed to provide any sort of useable profile information, despite the sampling of two slides being combined into the same extraction. This indicates that these samples have experienced DNA loss and degradation to a level where the DNA was no longer suitable for processing using this slide method. Although age is likely to be a significant cause of degradation other factors, such as slide preparation methods, cannot be excluded contributing to this observation. No records exist that provide details on differences between processing of the 1980s or 1990s slides in this study, however that does not mean that differences did not exist. Furthermore, the relatively small sample size means that the findings of non-informative profiles are limited to this study and is not an absolute conclusion that all slides from the 1980s will have the same result. While the results of this study demonstrated difficulty in obtaining DNA from the 1980s slides, novel work in other fields has shown promise as alternative pathways for analysis. Theunissen et al. (2021) found the use of Whole Genome Sequencing (WGS) and cell picking to achieve STR typing from single sperm cells. The Authors describe potential limitations in regard to staining, but suggest that this sort of DNA profiling workflow may be well suited to historic cases where limitations exist of sample availability, specifically mentioning microscopy slides [25].
The quantifiable male DNA in the direct extraction samples demonstrated that it was able to successfully recover DNA from slides, but at a loss of the contextual information provided by separating the extract into sperm and non-sperm fractions. This could become problematic in instances where the non-sperm component of the mixture is in excess to the sperm component. In these cases, the minor sperm cell profile can be difficult to isolate using traditional autosomal STR analysis, meaning only Y-STR kits are suitable for profiling [25]. This limits the evidentiary potential by reducing the ability to generate a profile which is unique to the sperm contributor. Nevertheless, there is still investigative potential in the direct extraction method for slide extraction, especially in the context of Cold Cases, where every lead could be significant to the resolution of a case.
The use of laser microdissection (LMD) has been widely demonstrated to be an effective method to selectively recover sperm cells from microscope slides [15,26,27]. LMD employs a laser to cut out specifically stained cells from the slide surface, with the greater selectivity leading to reduced potential for mixtures in the resulting profiles [28]. Other novel techniques include the use of antibodies to selectively target sperm cells in a mixture and selective degradation kits (such as the Erase Sperm Isolation Kit) which specifically degrade non-sperm to leave a clean sperm DNA extract [26]. While these techniques show promise, all have their own limitations that limit their widespread implementation in forensic workflows. Despite the advantages of LMD, cost is a key consideration with significant initial investment and on-going consumables leading to greater per sample expenditure [15]. Sperm antibody separation still requires further research and validation before widespread implementation and, despite providing cleaner sperm fractions, the selective degradation kits have been shown to have significantly lower total DNA recovery from semen [29,30].
Considering the degradative effect that time has on the stability of DNA, it was unknown whether the i-sep® method and its multiple wash steps would lead to a loss of DNA when compared to the direct extraction method, which has the advantage of keeping all lysed DNA in a single tube. The results of this study demonstrate the efficiency of the i-sep® method in recovering DNA and the additional advantage of separating the sperm and non-sperm fractions. Overall, the sperm fractions from the i-sep® method represented either single-source or clear male major profiles, with little to no female carryover of female DNA. Equally, while there were traces of male DNA in the non-sperm fractions, this is not unexpected as there is non-sperm DNA present in ejaculate and the high degradation could mean some sperm cells could have prematurely lysed into the non-sperm fraction. This has been previously reported, with the emphasis that the measure of a successful extraction is reflected in the majority of male DNA being extracted into the sperm fraction [23]. For this study, all i-sep® samples had the clear majority of male DNA contained within the sperm fraction, supporting that the i-sep® is a suitable method for extracting DNA from historic slides.
Despite the development of new techniques for sperm cell screening, the differential extraction method remains the most utilised method for processing sexual assault samples [29]. In a review of current methods for examining sexual assault evidence, Clark et al. found the differential extraction to have its own limitations (such as time and manual complexity) but was still able to provide strong DNA recovery of sperm cells in mixtures, with more informative male profiles due to the separation of sperm and non-sperm fractions [29]. This aligns with the findings of this study where the i-sep® was able to provide more informative sperm fraction profiles when compared to the mixed profiles resulting from the direct extraction. With the differential extraction likely to remain a core process for sexual assault workflows, the ability to apply optimised versions to historic cases via microscopy slides is a clear advantage for investigators. From the results of this study, the recommendation would be to screen potential candidate slides to assess the current sperm cell loading, followed by extraction using the i-sep® method to maximise both DNA recovery and contextual information from the separation of the sperm fraction. The direct extraction method should be employed as appropriate, depending on the specific sample is the aim is to recover all male DNA into one final extraction. These are the primary workflows that have been implemented by the laboratory, with results from Casework samples having been presented and accepted as evidence in Court.
5. Conclusions
The ability to provide new investigative leads in all investigations is advantageous for any forensic laboratory with this especially true for cold cases, but Investigators often are limited by the sample available for testing. The results of the experiments in this study clearly demonstrated the ability of the i-sep® differential extraction and direct extraction methods to recover DNA from historic microscope slides up to 30 years old. While the method has limitations pertaining to age-related degradation of the cells on the slides, the added benefit of sperm and non-sperm fraction separation using the i-sep® method was retained. The study also highlighted the importance of ensuring that historic slides are thoroughly re-screened prior to analysis and selected in context to their original microscopy scopes to ensure the best potential candidates for processing. Overall, the study successfully developed a method for routine implementation that can provide new pathways of analysis and generate new information for investigators.
Author Contributions
Conceptualization, C.M.H., S.E.E. and J.W.T.; methodology, C.M.H., S.E.E. and J.W.T.; formal analysis, C.M.H. and J.W.T.; writing—original draft preparation, C.M.H. and J.W.T.; writing—review and editing, C.M.H., S.E.E. and J.W.T.; supervision, J.W.T. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
This study was exempt from review by the Sir Charles Gairdner Osborne Park Health Care Group (SCGOPHCG) Human Research Ethics Committee (HREC). This activity has been assessed as meeting the requirements of the National Statement on Ethical Conduct in Human Research and is endorsed.
Informed Consent Statement
Informed consent was obtained for slides used in Phase 1 of this study and slides examined in Phases in 2 and 3 were approved for use in this study by the Western Australia Police Force.
Data Availability Statement
Data sharing is not applicable to this article.
Acknowledgments
The Authors wish to acknowledge Dean Topping and Tracy Horner for their assistance in laboratory examination of slides as well as the Western Australia Police Force for approval of slides used in Phases 2 and 3 of this study.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Ramos, P.; Handt, O.; Taylor, D. Investigating the position and level of DNA transfer to undergarments during digital sexual assault. Forensic Sci. Int. Genet. 2020, 47, 102316. [Google Scholar] [CrossRef] [PubMed]
- Connell, M. Expert testimony in sexual assault cases: Alcohol intoxication and memory. Int. J. Law Psychiatry 2015, 42–43, 98–105. [Google Scholar] [CrossRef] [PubMed]
- Berkowitz, S.R.; Garrett, B.L.; Fenn, K.M.; Loftus, E. Convicting with confidence? Why we should not over-rely on eyewitness confidence. Memory 2022, 30, 10–15. [Google Scholar] [CrossRef] [PubMed]
- Davis, R.C.; Wells, W. DNA testing in sexual assault cases: When do the benefits outweigh the costs? Forensic Sci. Int. 2019, 299, 44–48. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, K.; Sekiguchi, K.; Mizuno, N.; Kasai, K.; Sakai, I.; Sato, H.; Seta, S. The modified method of two-step differential extraction of sperm and vaginal epithelial cell DNA from vaginal fluid mixed with semen. Forensic Sci. Int. 1995, 72, 25–33. [Google Scholar] [CrossRef]
- Lounsbury, J.A.; Nambiar, S.M.; Karlsson, A.; Cunniffe, H.; Norris, J.V.; Ferrance, J.P.; Landers, J.P. Enhanced recovery of spermatozoa and comprehensive lysis of epithelial cells from sexual assault samples having a low cell counts or aged up to one year. Forensic Sci. Int. Genet. 2014, 8, 84–89. [Google Scholar] [CrossRef]
- Timken, M.D.; Klein, S.B.; Kubala, S.; Scharnhorst, G.; Buoncristiani, M.R.; Miller, K.W.P. Automation of the standard DNA differential extraction on the Hamilton AutoLys STAR system: A proof-of-concept study. Forensic Sci. Int. Genet. 2019, 40, 96–104. [Google Scholar] [CrossRef]
- Sinha, S.K.; Brown, H.; Holt, H.; Khan, M.R.; Brown, R.; Sgueglia, J.B.; Loftus, A.; Murphy, G.; Montgomery, A. Development and validation of a novel method “SpermX” for high throughput differential extraction processing of sexual assault kits (SAKs) for DNA analysis. Forensic Sci. Int. Genet. 2022, 59, 102690. [Google Scholar] [CrossRef]
- Gill, P.; Jeffreys, A.J.; Werrett, D.J. Forensic application of DNA ‘fingerprints’. Nature 1985, 318, 577–579. [Google Scholar] [CrossRef]
- Virkler, K.; Lednev, I.K. Analysis of body fluids for forensic purposes: From laboratory testing to non-destructive rapid confirmatory identification at a crime scene. Forensic Sci. Int. 2009, 188, 1–17. [Google Scholar] [CrossRef]
- Golomingi, R.; Haas, C.; Dobay, A.; Kottner, S.; Ebert, L. Sperm hunting on optical microscope slides for forensic analysis with deep convolutional networks—A feasibility study. Forensic Sci. Int. Genet. 2022, 56, 102602. [Google Scholar] [CrossRef] [PubMed]
- Westring, C.G.; Wiuf, M.; Nielsen, S.J.; Fogleman, J.C.; Old, J.B.; Lenz, C.; Reich, K.A.; Morling, N. SPERM HY-LITER for the identification of spermatozoa from sexual assault evidence. Forensic Sci. Int. Genet. 2014, 12, 161–167. [Google Scholar] [CrossRef] [PubMed]
- Jones, E.L. Analysis of Body Fluids in Sexual Assault Cases; Forensic Science Handbook; CRC Press: Boca Raton, FL, USA, 2020; pp. 613–706. [Google Scholar]
- White, T.J.; Rye, M.S.; Tay, J.W. Developmental validation of an efficient differential separation method incorporating the i-sep(®) DL spin column with high sperm DNA recovery for the processing of sexual assault samples. J. Forensic Sci. 2022, 67, 1679–1689. [Google Scholar] [CrossRef] [PubMed]
- Costa, S.; Correia-de-Sa, P.; Porto, M.J.; Caine, L. The Use of Laser Microdissection in Forensic Sexual Assault Casework: Pros and Cons Compared to Standard Methods. J. Forensic Sci. 2017, 62, 998–1006. [Google Scholar] [CrossRef] [PubMed]
- Dimo-Simonin, N.; Grange, F.; Brandt-Casadevall, C. PCR-based forensic testing of DNA from stained cytological smears. J. Forensic Sci. 1997, 42, 506–509. [Google Scholar] [CrossRef] [PubMed]
- Roy-Chowdhuri, S.; Chow, C.W.; Kane, M.K.; Yao, H.; Wistuba, I.I.; Krishnamurthy, S.; Stewart, J.; Staerkel, G. Optimizing the DNA yield for molecular analysis from cytologic preparations. Cancer Cytopathol. 2016, 124, 254–260. [Google Scholar] [CrossRef]
- Dejmek, A.; Zendehrokh, N.; Tomaszewska, M.; Edsjö, A. Preparation of DNA from cytological material: Effects of fixation, staining, and mounting medium on DNA yield and quality. Cancer Cytopathol. 2013, 121, 344–353. [Google Scholar] [CrossRef]
- Clark, M.; Gill, J.; Sasinouski, K.; McGuire, A. Cold Case Homicides: DNA Testing of Retained Autopsy Sexual Assault Smears. J. Forensic Sci. 2019, 64, 1100–1104. [Google Scholar] [CrossRef]
- Oppitz, E. Eine neue Färbemethode zum Nachweis der Spermien bei Sittlichkeitsdelikten. J. Arch. Kriminol. 1969, 144, 145–148. [Google Scholar]
- Fischer, R. The Gram-staining behavior of spermatozoa. Experientia 1953, 9, 335–336. [Google Scholar] [CrossRef]
- Applied Biosystems Quantifiler™ HP and Trio DNA Quantification Kits Uer Guide: Publication Number 4485354 Revision G; Thermofisher Scientific: Waltham, MA, USA, 2017.
- Alaeddini, R.; Walsh, S.J.; Abbas, A. Forensic implications of genetic analyses from degraded DNA—A review. Forensic Sci. Int. Genet. 2010, 4, 148–157. [Google Scholar] [CrossRef] [PubMed]
- Casey, D.G.; Domijan, K.; MacNeill, S.; Rizet, D.; O’Connell, D.; Ryan, J. The Persistence of Sperm and the Development of Time Since Intercourse (TSI) Guidelines in Sexual Assault Cases at Forensic Science Ireland, Dublin, Ireland. J. Forensic Sci. 2017, 62, 585–592. [Google Scholar] [CrossRef] [PubMed]
- Theunissen, G.M.G.; Gibb, A.; Lin Paul, K.T.; Rolf, B.; Forat, S.; Jäger, R. DNA profiling of single sperm cells after whole genome amplification. Forensic Sci. Int. Rep. 2021, 4, 100240. [Google Scholar] [CrossRef]
- Elliott, K.; Hill, D.S.; Lambert, C.; Burroughes, T.R.; Gill, P. Use of laser microdissection greatly improves the recovery of DNA from sperm on microscope slides. Forensic Sci. Int. 2003, 137, 28–36. [Google Scholar] [CrossRef] [PubMed]
- Sanders, C.; Sanchez, N.; Ballantyne, J.; Peterson, D. Laser Microdissection Separation of Pure Spermatozoa from Epithelial Cells for Short Tandem Repeat Analysis*. J. Forensic Sci. 2006, 51, 748–757. [Google Scholar] [CrossRef]
- Meredith, M.; Bright, J.A.; Cockerton, S.; Vintiner, S. Development of a one-tube extraction and amplification method for DNA analysis of sperm and epithelial cells recovered from forensic samples by laser microdissection. Forensic Sci. Int. Genet. 2012, 6, 91–96. [Google Scholar] [CrossRef]
- Clark, C.; Turiello, R.; Cotton, R.; Landers, J.P. Analytical approaches to differential extraction for sexual assault evidence. Anal. Chim. Acta. 2021, 1141, 230–245. [Google Scholar] [CrossRef]
- Zhao, X.C.; Wang, L.; Sun, J.; Jiang, B.W.; Zhang, E.L.; Ye, J. Isolating Sperm from Cell Mixtures Using Magnetic Beads Coupled with an Anti-PH-20 Antibody for Forensic DNA Analysis. PLoS ONE 2016, 11, e0159401. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).