The Cancer Clock Is (Not) Ticking: Links between Circadian Rhythms and Cancer
Abstract
1. Introduction to Circadian Rhythms and Cancer
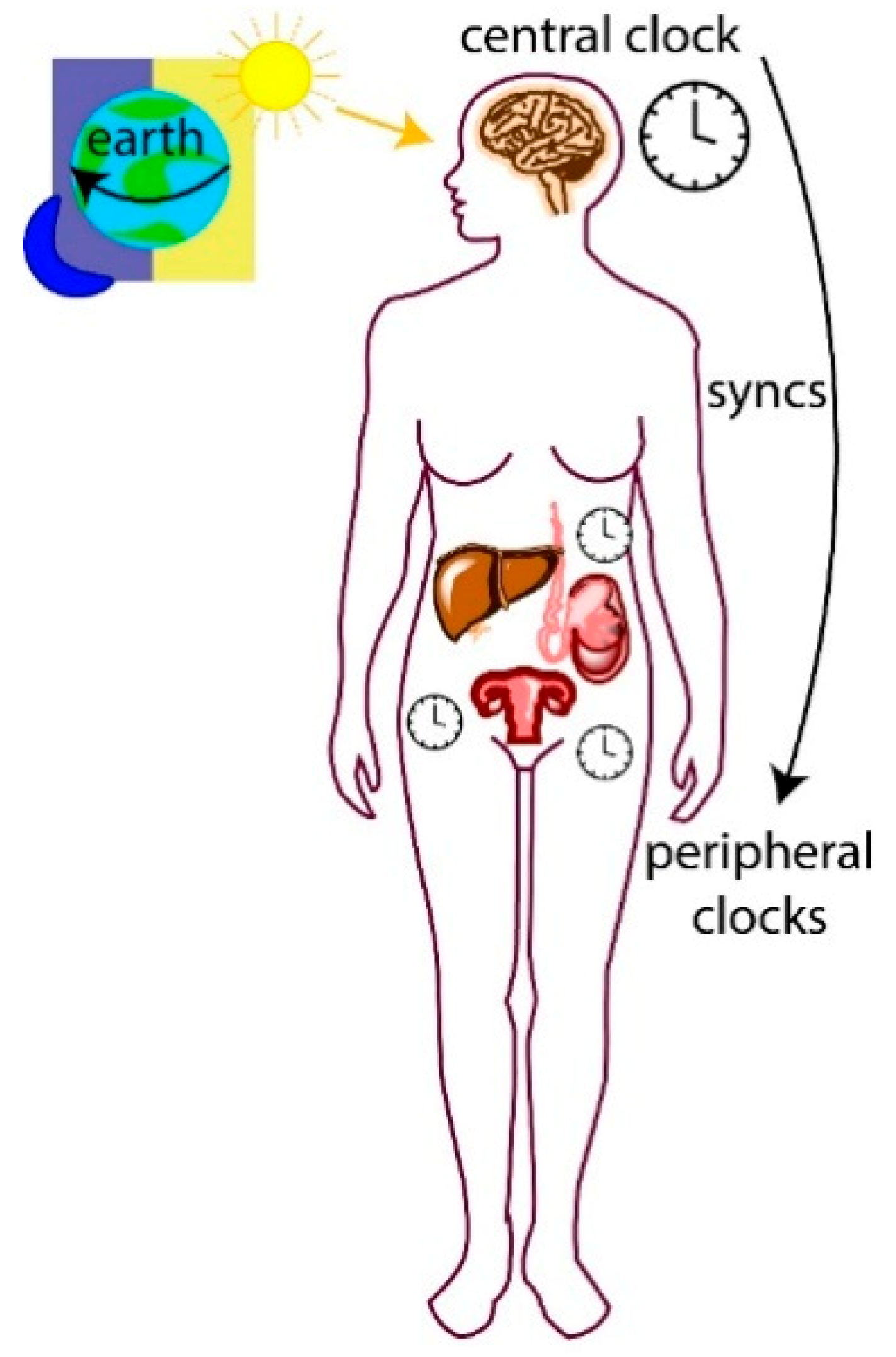
1.1. Central and Peripheral Clocks
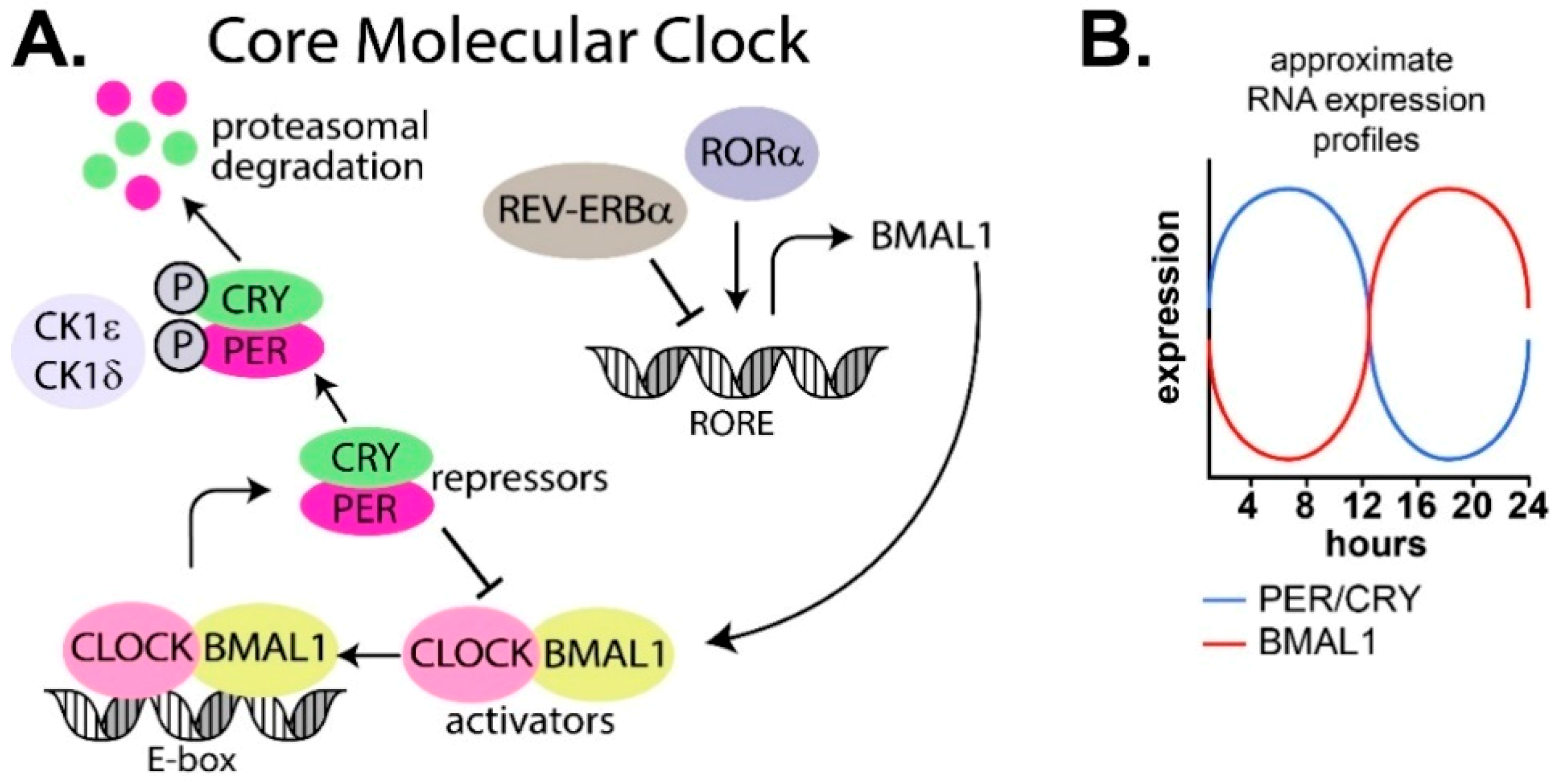
1.2. Molecular Regulation of the Circadian Clock
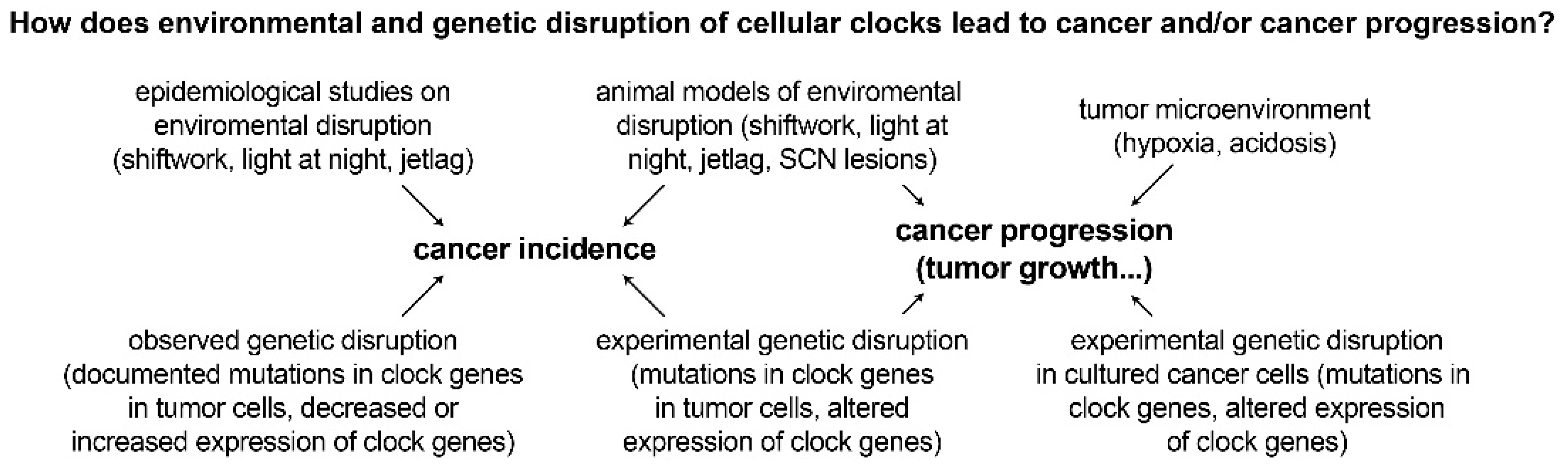
2. Environmental Disruption of Circadian Rhythms and Links to Cancer
2.1. Epidemiological Studies Link Shift Work and Disrupted Circadian Rhythms to Cancer Incidence
2.2. Animal Models and In Vitro Studies of Circadian Disruption Support the Epidemiological Studies and Demonstrate Increased Tumor Incidence and Faster Tumor Growth
3. Genetic Disruption of Circadian Rhythms and Links to Cancer
Mutations, Epigenetic Changes and Deregulated Expression of Clock-Related Genes are Common in Tumor Cells
4. Hallmarks and Characteristics of Cancer and their Interactions with Circadian Rhythms
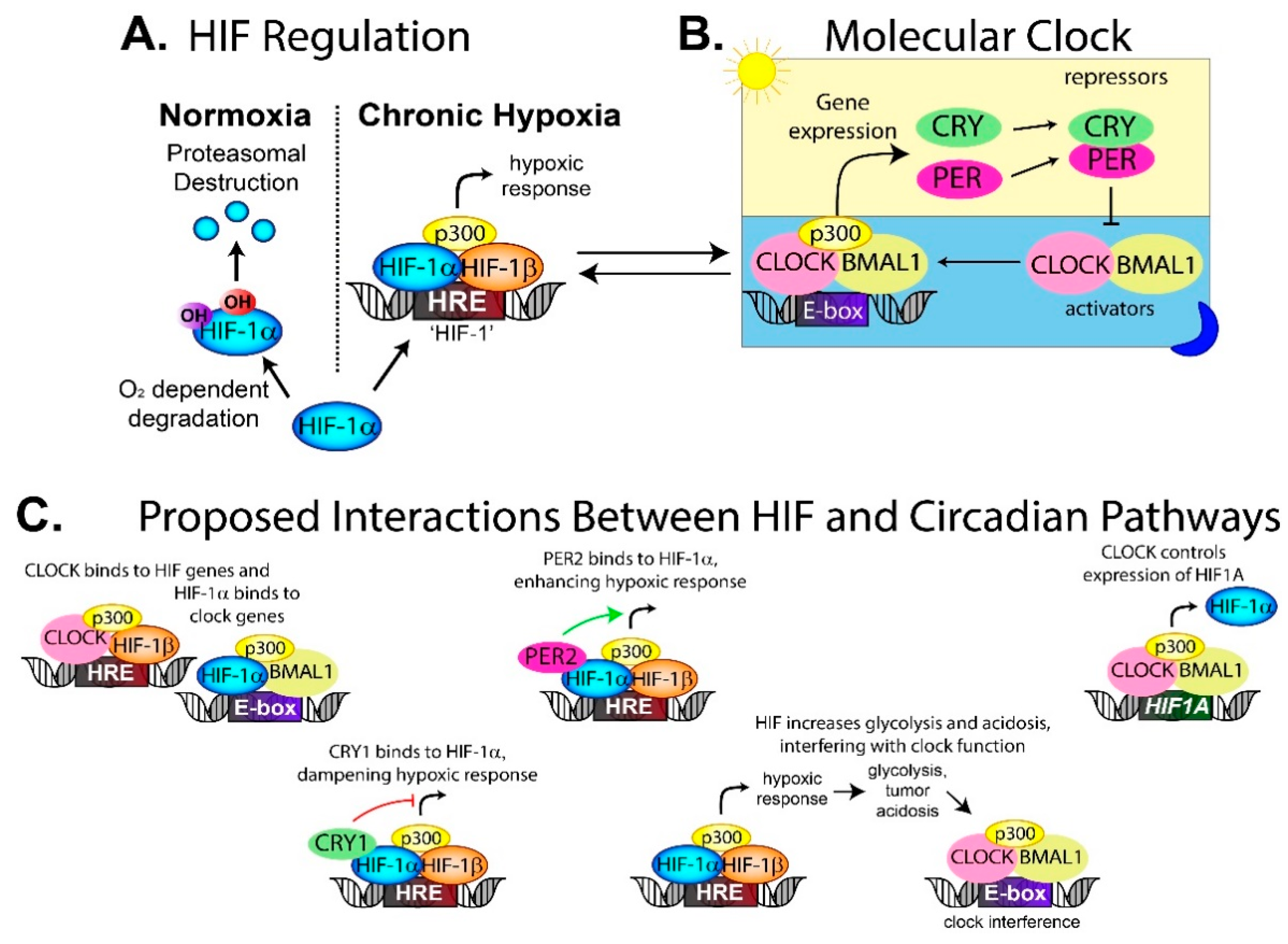
4.1. Disruption of Tumor Circadian Rhythms by Microenvironmental Factors Such as Hypoxia and Acidosis
4.2. Disrupted Cell Cycles Can Be Controlled by Circadian Rhythms
4.3. Circadian Control of Metabolism
4.4. Circadian Control of Other Cancer Characteristics
5. Restoring the Circadian Clock in Tumor Cells May Be an Effective Anti-Cancer Strategy
5.1. Circadian Anti-Cancer Drug Targets
5.2. Timing of Current Chemotherapy Protocols May Improve Efficacy through Circadian Mechanisms
5.3. Melatonin
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| 5-FU | 5-Fluorouracil |
| ARNTL2 | Aryl Hydrocarbon Receptor Nuclear Translocator Like 2 (BMAL2) |
| BMAL1 | Brain and Muscle ARNT-like protein-1 |
| Cdkn1a | Cyclin Dependent Kinase Inhibitor 1A |
| CDK4/6 | Cyclin-Dependent Kinase 4/6 |
| ChIP | Chromatin Immunoprecipitation |
| CML | Chronic Myeloid leukemia |
| CLOCK | Circadian Locomotor Output Cycles Kaput |
| CK1δ | Casein Kinase 1 Delta |
| CK1ε | Casein Kinase 1 Epsilon |
| CRY1/2 | Cryptochrome |
| DEX | Dexamethasone |
| GSK-3β | Glycogen Synthase Kinase -3 Beta |
| HIF | Hypoxia Inducible Factor |
| HIF-1α | Hypoxia-inducible factor 1-alpha |
| HRE | Hypoxia Response Element |
| IARC | International Agency for Research on Cancer |
| NR1D1 | Nuclear receptor subfamily 1, group D, member 1 (gene for REV-ERBα) |
| NPAS2 | Neuronal PAS Domain Protein 2 |
| PER1/2/3 | Period1, Period2, Period3 |
| PERK | Protein kinase RNA-like Endoplasmic Reticulum Kinase |
| ROR | Retinoic acid receptor-related orphan receptor |
| SCN | Suprachiasmatic nucleus |
| SNP | Single-nucleotide polymorphism |
References
- Partch, C.L.; Green, C.B.; Takahashi, J.S. Molecular architecture of the mammalian circadian clock. Trends Cell Biol. 2014, 24, 90–99. [Google Scholar] [CrossRef] [PubMed]
- Matsuo, T.; Yamaguchi, S.; Mitsui, S.; Emi, A.; Shimoda, F.; Okamura, H. Control Mechanism of the Circadian Clock for Timing of Cell Division in Vivo. Science 2003, 302, 255–259. [Google Scholar] [CrossRef] [PubMed]
- Zhang, R.; Lahens, N.F.; Balance, H.I.; Hughes, M.E.; Hogenesch, J.B. A circadian gene expression atlas in mammals: Implications for biology and medicine. Proc. Natl. Acad. Sci. USA 2014, 111, 16219–16224. [Google Scholar] [CrossRef] [PubMed]
- Asher, G.; Gatfield, D.; Stratmann, M.; Reinke, H.; Dibner, C.; Kreppel, F.; Mostoslavsky, R.; Alt, F.W.; Schibler, U. SIRT1 Regulates Circadian Clock Gene Expression through PER2 Deacetylation. Cell 2008, 134, 317–328. [Google Scholar] [CrossRef] [PubMed]
- Gréchez-Cassiau, A.; Rayet, B.; Guillaumond, F.; Teboul, M.; Delaunay, F. The Circadian Clock Component BMAL1 Is a Critical Regulator of p21WAF1/CIP1 Expression and Hepatocyte Proliferation. J. Biol. Chem. 2008, 283, 4535–4542. [Google Scholar] [CrossRef] [PubMed]
- Akle, V.; Stankiewicz, A.J.; Kharchenko, V.; Yu, L.; Kharchenko, P.V.; Zhdanova, I.V. Circadian Kinetics of Cell Cycle Progression in Adult Neurogenic Niches of a Diurnal Vertebrate. J. Neurosci. 2017, 37, 1900–1909. [Google Scholar] [CrossRef] [PubMed]
- Gery, S.; Komatsu, N.; Baldjyan, L.; Yu, A.; Koo, D.; Koeffler, H.P. The circadian gene per1 plays an important role in cell growth and dna damage control in human cancer cells. Mol. Cell 2006, 22, 375–382. [Google Scholar] [CrossRef]
- Koyanagi, S.; Kuramoto, Y.; Nakagawa, H.; Aramaki, H.; Ohdo, S.; Soeda, S.; Shimeno, H. A Molecular Mechanism Regulating Circadian Expression of Vascular Endothelial Growth Factor in Tumor Cells. Cancer Res. 2003, 63, 7277–7283. [Google Scholar] [PubMed]
- Lamia, K. Ticking time bombs: Connections between circadian clocks and cancer. F1000 Fac. Rev. 2017, 6. [Google Scholar] [CrossRef]
- Papagiannakopoulos, T.; Bauer, M.R.; Davidson, S.M.; Heimann, M.; Subbaraj, L.; Bhutkar, A.; Bartlebaugh, J.; Vander Heiden, M.G.; Jacks, T. Circadian Rhythm Disruption Promotes Lung Tumorigenesis. Cell Metab. 2016, 24, 324–331. [Google Scholar] [CrossRef]
- Ye, Y.; Xiang, Y.; Ozguc, F.M.; Kim, Y.; Liu, C.J.; Park, P.K.; Hu, Q.; Diao, L.; Lou, Y.; Lin, C.; et al. The Genomic Landscape and Pharmacogenomic Interactions of Clock Genes in Cancer Chronotherapy. Cell Syst. 2018, 6, 314.e2–328.e2. [Google Scholar] [CrossRef] [PubMed]
- Schernhammer, E.S.; Laden, F.; Speizer, F.E.; Willett, W.C.; Hunter, D.J.; Kawachi, I.; Colditz, G.A. Rotating Night Shifts and Risk of Breast Cancer in Women Participating in the Nurses’ Health Study. JNCI J. Nat. Cancer Inst. 2001, 93, 1563–1568. [Google Scholar] [CrossRef] [PubMed]
- Papantoniou, K.; Castaño-Vinyals, G.; Espinosa, A.; Aragonés, N.; Pérez-Gómez, B.; Burgos, J.; Gómez-Acebo, I.; Llorca, J.; Peiró, R.; Jimenez-Moleón, J.J.; et al. Night shift work, chronotype and prostate cancer risk in the MCC-Spain case-control study. Int. J. Cancer 2015, 137, 1147–1157. [Google Scholar] [CrossRef] [PubMed]
- Knutsson, A.; Alfredsson, L.; Karlsson, B.; Akerstedt, T.; Fransson, E.I.; Westerholm, P.; Westerlund, H. Breast cancer among shift workers: Results of the WOLF longitudinal cohort study. Scand J. Work Environ. Health 2013, 39, 170–177. [Google Scholar] [CrossRef] [PubMed]
- Åkerstedt, T.; Knutsson, A.; Narusyte, J.; Svedberg, P.; Kecklund, G.; Alexanderson, K. Night work and breast cancer in women: A Swedish cohort study. BMJ Open 2015, 5, e008127. [Google Scholar] [CrossRef] [PubMed]
- Alhopuro, P.; Björklund, M.; Sammalkorpi, H.; Turunen, M.; Tuupanen, S.; Biström, M.; Niittymäki, I.; Lehtonen, H.J.; Kivioja, T.; Launonen, V.; et al. Mutations in the Circadian Gene CLOCK in Colorectal Cancer. Mol. Cancer Res. 2010, 8, 952–960. [Google Scholar] [CrossRef]
- Walton, Z.E.; Patel, C.H.; Brooks, R.C.; Yu, Y.; Ibrahim-Hashim, A.; Riddle, M.; Porcu, A.; Jian, T.; Ecker, B.L.; Tameire, F.; et al. Acid Suspends the Circadian Clock in Hypoxia through Inhibition of mTOR. Cell 2018, 174, 72.e32–87.e32. [Google Scholar] [CrossRef]
- Reppert, S.M.; Weaver, D.R. Coordination of circadian timing in mammals. Nature 2002, 418, 935–941. [Google Scholar] [CrossRef]
- Akhtar, R.A.; Reddy, A.B.; Maywood, E.S.; Clayton, J.D.; King, V.M.; Smith, A.G.; Gant, T.W.; Hastings, M.H.; Kyriacou, C.P. Circadian Cycling of the Mouse Liver Transcriptome, as Revealed by cDNA Microarray, Is Driven by the Suprachiasmatic Nucleus. Curr. Biol. 2002, 12, 540–550. [Google Scholar] [CrossRef]
- Husse, J.; Eichele, G.; Oster, H. Synchronization of the mammalian circadian timing system: Light can control peripheral clocks independently of the SCN clock: Alternate routes of entrainment optimize the alignment of the body’s circadian clock network with external time. BioEssays 2015, 37, 1119–1128. [Google Scholar] [CrossRef]
- Albrecht, U. Timing to Perfection: The Biology of Central and Peripheral Circadian Clocks. Neuron 2012, 74, 246–260. [Google Scholar] [CrossRef] [PubMed]
- Mohawk, J.A.; Green, C.B.; Takahashi, J.S. Central and peripheral circadian clocks in mammals. Ann. Rev. Neurosci. 2012, 35, 445–462. [Google Scholar] [CrossRef] [PubMed]
- Bass, J.; Takahashi, J.S. Circadian Integration of Metabolism and Energetics. Science 2010, 330, 1349. [Google Scholar] [CrossRef]
- Damiola, F.; Le Minh, N.; Preitner, N.; Kornmann, B.; Fleury-Olela, F.; Schibler, U. Restricted feeding uncouples circadian oscillators in peripheral tissues from the central pacemaker in the suprachiasmatic nucleus. Genes Dev. 2000, 14, 2950–2961. [Google Scholar] [CrossRef] [PubMed]
- Welsh, D.K.; Yoo, S.H.; Liu, A.C.; Takahashi, J.S.; Kay, S.A. Bioluminescence Imaging of Individual Fibroblasts Reveals Persistent, Independently Phased Circadian Rhythms of Clock Gene Expression. Curr. Biol. 2004, 14, 2289–2295. [Google Scholar] [CrossRef] [PubMed]
- Nagoshi, E.; Saini, C.; Bauer, C.; Laroche, T.; Naef, F.; Schibler, U. Circadian Gene Expression in Individual Fibroblasts: Cell-Autonomous and Self-Sustained Oscillators Pass Time to Daughter Cells. Cell 2004, 119, 693–705. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, J.S. Transcriptional architecture of the mammalian circadian clock. Nat. Rev. Genet. 2016, 18, 164. [Google Scholar] [CrossRef]
- Savvidis, C.; Koutsilieris, M. Circadian rhythm disruption in cancer biology. Mol. Med. 2012, 18, 1249–1260. [Google Scholar] [CrossRef]
- Sahar, S.; Sassone-Corsi, P. Metabolism and cancer: The circadian clock connection. Nat. Rev. Cancer 2009, 9, 886. [Google Scholar] [CrossRef]
- Zhang, E.E.; Kay, S.A. Clocks not winding down: Unravelling circadian networks. Nat. Rev. Mol. Cell Biol. 2010, 11, 764. [Google Scholar] [CrossRef]
- Vitaterna, M.H.; King, D.P.; Chang, A.M.; Kornhauser, J.M.; Lowrey, P.L.; McDonald, J.D.; Dove, W.F.; Pinto, L.H.; Turek, F.W.; Takahashi, J.S. Mutagenesis and mapping of a mouse gene, Clock, essential for circadian behavior. Science 1994, 264, 719–725. [Google Scholar] [CrossRef] [PubMed]
- King, D.P.; Zhao, Y.; Sangoram, A.M.; Wilsbacher, L.D.; Tanaka, M.; Antoch, M.P.; Steeves, T.D.; Vitaterna, M.H.; Kornhauser, J.M.; Lowrey, P.L.; et al. Positional cloning of the mouse circadian clock gene. Cell 1997, 89, 641–653. [Google Scholar] [CrossRef]
- Zylka, M.J.; Shearman, L.P.; Weaver, D.R.; Reppert, S.M. Three period Homologs in Mammals: Differential Light Responses in the Suprachiasmatic Circadian Clock and Oscillating Transcripts Outside of Brain. Neuron 1998, 20, 1103–1110. [Google Scholar] [CrossRef]
- Tei, H.; Okamura, H.; Shigeyoshi, Y.; Fukuhara, C.; Ozawa, R.; Hirose, M.; Sakaki, Y. Circadian oscillation of a mammalian homologue of the Drosophila period gene. Nature 1997, 389, 512–516. [Google Scholar] [CrossRef] [PubMed]
- Kume, K.; Zylka, M.J.; Sriram, S.; Shearman, L.P.; Weaver, D.R.; Jin, X.; Maywood, E.S.; Hastings, M.H.; Reppert, S.M. mCRY1 and mCRY2 Are Essential Components of the Negative Limb of the Circadian Clock Feedback Loop. Cell 1999, 98, 193–205. [Google Scholar] [CrossRef]
- Ikeda, M.; Nomura, M. cDNA Cloning and Tissue-Specific Expression of a Novel Basic Helix–Loop–Helix/PAS Protein (BMAL1) and Identification of Alternatively Spliced Variants with Alternative Translation Initiation Site Usage. Biochem. Biophys. Res. Commun. 1997, 233, 258–264. [Google Scholar] [CrossRef]
- Eide, E.J.; Kang, H.; Crapo, S.; Gallego, M.; Virshup, D.M. Casein kinase I in the mammalian circadian clock. Methods Enzymol. 2005, 393, 408–418. [Google Scholar]
- Etchegaray, J.-P.; Machida, K.K.; Noton, E.; Constance, C.M.; Dallmann, R.; Di Napoli, M.N.; DeBruyne, J.P.; Lambert, C.M.; Yu, E.A.; Reppert, S.M.; et al. Casein Kinase 1 Delta Regulates the Pace of the Mammalian Circadian Clock. Mol. Cell. Biol. 2009, 29, 3853–3866. [Google Scholar] [CrossRef]
- Preitner, N.; Damiola, F.; Luis Lopez, M.; Zakany, J.; Duboule, D.; Albrecht, U.; Schibler, U. The Orphan Nuclear Receptor REV-ERBα Controls Circadian Transcription within the Positive Limb of the Mammalian Circadian Oscillator. Cell 2002, 110, 251–260. [Google Scholar] [CrossRef]
- Sato, T.K.; Panda, S.; Miraglia, L.J.; Reyes, T.M.; Rudic, R.D.; McNamara, P.; Naik, K.A.; FitzGerald, G.A.; Kay, S.A.; Hogenesch, J.B. A Functional Genomics Strategy Reveals Rora as a Component of the Mammalian Circadian Clock. Neuron 2004, 43, 527–537. [Google Scholar] [CrossRef]
- Ko, C.H.; Takahashi, J.S. Molecular components of the mammalian circadian clock. Human Mol. Genet. 2006, 15 (Suppl. 2), R271–R277. [Google Scholar] [CrossRef]
- Viswanathan, A.N.; Hankinson, S.E.; Schernhammer, E.S. Night shift work and the risk of endometrial cancer. Cancer Res. 2007, 67, 10618–10622. [Google Scholar] [CrossRef] [PubMed]
- Schernhammer, E.S.; Laden, F.; Speizer, F.E.; Willett, W.C.; Hunter, D.J.; Kawachi, I.; Fuchs, C.S.; Colditz, G.A. Night-Shift Work and Risk of Colorectal Cancer in the Nurses’ Health Study. JNCI J. Nat. Cancer Inst. 2003, 95, 825–828. [Google Scholar] [CrossRef] [PubMed]
- Papantoniou, K.; Devore, E.E.; Massa, J.; Strohmaier, S.; Vetter, C.; Yang, L.; Shi, Y.; Giovannucci, E.; Speizer, F.; Schernhammer, E.S. Rotating night shift work and colorectal cancer risk in the nurses’ health studies. Int. J. Cancer 2018, 143, 2709–2717. [Google Scholar] [CrossRef] [PubMed]
- Wegrzyn, L.R.; Tamimi, R.M.; Rosner, B.A.; Brown, S.B.; Stevens, R.G.; Eliassen, A.H.; Laden, F.; Willett, W.C.; Hankinson, S.E.; Schernhammer, E.S. Rotating Night-Shift Work and the Risk of Breast Cancer in the Nurses’ Health Studies. Am. J. Epidemiol. 2017, 186, 532–540. [Google Scholar] [CrossRef] [PubMed]
- Papantoniou, K.; Castano-Vinyals, G.; Espinosa, A.; Aragonés, N.; Perez-Gomez, B.; Ardanaz, E.; Miren Altzibar, J.; Martín, V.; Bsc, I.; Llorca, J.; et al. Breast cancer risk and night shift work in a case–control study in a Spanish population. Eur. J. Epidemiol. 2015, 31, 867–878. [Google Scholar] [CrossRef] [PubMed]
- Hansen, J. Increased Breast Cancer Risk among Women Who Work Predominantly at Night. Epidemiology 2001, 12, 74–77. [Google Scholar] [CrossRef]
- Schernhammer, E.S.; Kroenke, C.H.; Laden, F.; Hankinson, S.E. Night Work and Risk of Breast Cancer. Epidemiology 2006, 17, 108–111. [Google Scholar] [CrossRef]
- Megdal, S.P.; Kroenke, C.H.; Laden, F.; Pukkala, E.; Schernhammer, E.S. Night work and breast cancer risk: A systematic review and meta-analysis. Eur J. Cancer 2005, 41, 2023–2032. [Google Scholar] [CrossRef]
- Cordina-Duverger, E.; Menegaux, F.; Popa, A.; Rabstein, S.; Harth, V.; Pesch, B.; Brüning, T.; Fritschi, L.; Glass, D.C.; Heyworth, J.S.; et al. Night shift work and breast cancer: A pooled analysis of population-based case–control studies with complete work history. Eur. J. Epidemiol. 2018, 33, 369–379. [Google Scholar] [CrossRef]
- Pham, T.T.; Lee, E.-S.; Kong, S.Y.; Kim, J.; Kim, S.Y.; Joo, J.; Yoon, K.A.; Park, B. Night-shift work, circadian and melatonin pathway related genes and their interaction on breast cancer risk: Evidence from a case-control study in Korean women. Sci. Rep. 2019, 9, 10982. [Google Scholar] [CrossRef] [PubMed]
- Straif, K.; Baan, R.; Grosse, Y.; Secretan, B.; El Ghissassi, F.; Bouvard, V.; Altieri, A.; Benbrahim-Tallaa, L.; Cogliano, V. Carcinogenicity of shift-work, painting, and fire-fighting. Lancet Oncol. 2007, 8, 1065–1066. [Google Scholar] [CrossRef]
- Kubo, T.; Ozasa, K.; Mikami, K.; Wakai, K.; Fujino, Y.; Watanabe, Y.; Miki, T.; Nakao, M.; Hayashi, K.; Suzuki, K.; et al. Prospective Cohort Study of the Risk of Prostate Cancer among Rotating-Shift Workers: Findings from the Japan Collaborative Cohort Study. Am. J. Epidemiol. 2006, 164, 549–555. [Google Scholar] [CrossRef] [PubMed]
- Davis, S.; Mirick, D.K. Circadian disruption, shift work and the risk of cancer: A summary of the evidence and studies in Seattle. Cancer Causes Control 2006, 17, 539–545. [Google Scholar] [CrossRef] [PubMed]
- Tokumaru, O.; Haruki, K.; Bacal, K.; Katagiri, T.; Yamamoto, T.; Sakurai, Y. Incidence of cancer among female flight attendants: A meta-analysis. J. Trav. Med. 2006, 13, 127–132. [Google Scholar] [CrossRef]
- Pukkala, E.; Aspholm, R.; Auvinen, A.; Eliasch, H.; Gundestrup, M.; Haldorsen, T.; Hammar, N.; Hrafnkelsson, J.; Kyyrönen, P.; Linnersjö, A.; et al. Cancer Incidence Among 10,211 Airline Pilots: A Nordic Study. Aviat Space Environ. Med. 2003, 74, 699–706. [Google Scholar]
- Band, P.R.; Le, N.D.; Fang, R.; Deschamps, M.; Coldman, A.J.; Gallagher, R.P.; Moody, J. Cohort study of Air Canada pilots: Mortality, cancer incidence, and leukemia risk. Am. J. Epidemiol. 1996, 143, 137–143. [Google Scholar] [CrossRef]
- Schernhammer, E.S.; Razavi, P.; Li, T.Y.; Qureshi, A.A.; Han, J. Rotating Night Shifts and Risk of Skin Cancer in the Nurses’ Health Study. JNCI J. Natl. Cancer Inst. 2011, 103, 602–606. [Google Scholar] [CrossRef]
- Srour, B.; Plancoulaine, S.; Andreeva, V.A.; Fassier, P.; Julia, C.; Galan, P.; Hercberg, S.; Deschasaux, M.; Latino-Martel, P.; Touvier, M. Circadian nutritional behaviours and cancer risk: New insights from the NutriNet-santé prospective cohort study: Disclaimers. Int. J. Cancer 2018, 143, 2369–2379. [Google Scholar] [CrossRef]
- Kogevinas, M.; Espinosa, A.; Castelló, A.; Gómez-Acebo, I.; Guevara, M.; Martin, V.; Amiano, P.; Alguacil, J.; Peiro, R.; Moreno, V.; et al. Effect of mistimed eating patterns on breast and prostate cancer risk (MCC-Spain Study). Int. J. Cancer 2018, 143, 2380–2389. [Google Scholar] [CrossRef]
- Anisimov, V.N.; Baturin, D.A.; Popovich, I.G.; Zabezhinski, M.A.; Manton, K.G.; Semenchenko, A.V.; Yashin, A.I. Effect of exposure to light-at-night on life span and spontaneous carcinogenesis in female CBA mice. Int. J. Cancer 2004, 111, 475–479. [Google Scholar] [CrossRef]
- Van Dycke, K.C.; Rodenburg, W.; van Oostrom, C.T.; van Kerkhof, L.W.; Pennings, J.L.; Roenneberg, T.; van Steeg, H.; van der Horst, G.T. Chronically alternating light cycles increase breast cancer risk in mice. Curr. Biol. 2015, 25, 1932–1937. [Google Scholar] [CrossRef] [PubMed]
- Filipski, E.; King, V.M.; Li, X.; Granda, T.G.; Mormont, M.C.; Lui, X.; Claustrat, B.; Hastings, M.H.; Levi, F. Host circadian clock as a control point in tumor progression. J. Natl. Cancer Inst. 2002, 94, 690–697. [Google Scholar] [CrossRef] [PubMed]
- Filipski, E.; Delaunay, F.; King, V.M.; Wu, M.-W.; Claustrat, B.; Gréchez-Cassiau, A.; Guettier, C.; Hastings, M.H.; Francis, L. Effects of chronic jet lag on tumor progression in mice. Cancer Res. 2004, 64, 7879–7885. [Google Scholar] [CrossRef] [PubMed]
- Balsalobre, A.; Brown, S.A.; Marcacci, L.; Tronche, F.; Kellendonk, C.; Reichardt, H.M.; Schütz, G.; Schibler, U. A Serum Shock Induces Circadian Gene Expression in Mammalian Tissue Culture Cells. Cell 1998, 93, 929–937. [Google Scholar] [CrossRef]
- Tamaru, T.; Hattori, M.; Honda, K.; Benjamin, I.; Ozawa, T.; Takamatsu, K. Synchronization of Circadian Per2 Rhythms and HSF1-BMAL1: CLOCK Interaction in Mouse Fibroblasts after Short-Term Heat Shock Pulse. PLoS ONE 2011, 6, e24521. [Google Scholar] [CrossRef] [PubMed]
- Balsalobre, A.; Brown, S.A.; Marcacci, L.; Tronche, F.; Kellendonk, C.; Reichardt, H.M.; Schütz, G.; Schibler, U. Resetting of Circadian Time in Peripheral Tissues by Glucocorticoid Signaling. Science 2000, 289, 2344–2347. [Google Scholar] [CrossRef]
- Yagita, K.; Okamura, H. Forskolin induces circadian gene expression of rPer1, rPer2 and dbp in mammalian rat-1 fibroblasts. FEBS Lett. 2000, 465, 79–82. [Google Scholar] [CrossRef]
- Kiessling, S.; Beaulieu-Laroche, L.; Blum, I.D.; Landgraf, D.; Welsh, D.K.; Storch, K.-F.; Labrecque, N.; Cermakian, N. Enhancing circadian clock function in cancer cells inhibits tumor growth. BMC Biol. 2017, 15, 13. [Google Scholar] [CrossRef]
- Filipski, E.; Lévi, F. Circadian Disruption in Experimental Cancer Processes. Integr. Cancer Ther. 2009, 8, 298–302. [Google Scholar] [CrossRef]
- Fu, L.; Pelicano, H.; Liu, J.; Huang, P.; Lee, C.C. The Circadian Gene Period2 Plays an Important Role in Tumor Suppression and DNA Damage Response in vivo. Cell 2002, 111, 41–50. [Google Scholar] [CrossRef]
- Deng, F.; Yang, K. Current Status of Research on the Period Family of Clock Genes in the Occurrence and Development of Cancer. J. Cancer 2019, 10, 1117–1123. [Google Scholar] [CrossRef] [PubMed]
- Mteyrek, A.; Filipski, E.; Guettier, C.; Okyar, A.; Levi, F. Clock gene Per2 as a controller of liver carcinogenesis. Oncotarget 2016, 7, 85832–85847. [Google Scholar] [CrossRef] [PubMed]
- Miyazaki, K.; Wakabayashui, M.; Hara, Y.; Ishida, N. Tumor growth suppression in vivo by overexpression of the circadian component, PER2. Genes Cell. 2010, 15, 351–358. [Google Scholar] [CrossRef] [PubMed]
- Mocellin, S.; Tropea, S.; Benna, C.; Rossi, C.R. Circadian pathway genetic variation and cancer risk: Evidence from genome-wide association studies. BMC Med. 2018, 16, 20. [Google Scholar] [CrossRef]
- Fujioka, A.; Takashima, N.; Shigeyoshi, Y. Circadian rhythm generation in a glioma cell line. Biochem. Biophys. Res. Commun. 2006, 346, 169–174. [Google Scholar] [CrossRef] [PubMed]
- Xia, H.-C.; Niu, Z.-F.; Ma, H.; Cao, S.-Z.; Hao, S.-C.; Liu, Z.-T.; Wang, F. Deregulated Expression of the Per1 and Per2 in Human Gliomas. Can. J. Neurol. Sci. 2014, 37, 365–370. [Google Scholar] [CrossRef]
- Pogue-Geile, K.L.; Lyons-Weiler, J.; Whitcomb, D.C. Molecular overlap of fly circadian rhythms and human pancreatic cancer. Cancer Lett. 2006, 243, 55–57. [Google Scholar] [CrossRef]
- Chen, S.-T.; Choo, K.-B.; Hou, M.-F.; Yeh, K.-T.; Kuo, S.-J.; Chang, J.-G. Deregulated expression of the PER1, PER2 and PER3 genes in breast cancers. Carcinogenesis 2005, 26, 1241–1246. [Google Scholar] [CrossRef]
- Winter, S.L.; Bosnoyan-Collins, L.; Pinnaduwage, D.; Andrulis, I.L. Expression of the circadian clock genes Per1 and Per2 in sporadic and familial breast tumors. Neoplasia 2007, 9, 797–800. [Google Scholar] [CrossRef]
- Tokunaga, H.; Takebayashi, Y.; Utsunomiya, H.; Akahira, J.-I.; Higashimoto, M.; Mashiko, M.; Ito, K.; Niikura, H.; Takenoshita, S.-I.; Yaegashi, N. Clinicopathological significance of circadian rhythm-related gene expression levels in patients with epithelial ovarian cancer. Acta Obstet. Gynecol. Scand. 2008, 87, 1060–1070. [Google Scholar] [CrossRef]
- Yang, M.-Y.; Chang, J.-G.; Lin, P.-M.; Tang, K.-P.; Chen, Y.-H.; Lin, H.Y.-H.; Liu, T.-C.; Hsiao, H.-H.; Liu, Y.-C.; Lin, S.-F. Downregulation of circadian clock genes in chronic myeloid leukemia: Alternative methylation pattern of hPER3. Cancer Sci. 2006, 97, 1298–1307. [Google Scholar] [CrossRef] [PubMed]
- Taniguchi, H.; Fernández, A.F.; Setién, F.; Ropero, S.; Ballestar, E.; Villanueva, A.; Yamamoto, H.; Imai, K.; Shinomura, Y.; Esteller, M. Epigenetic Inactivation of the Circadian Clock Gene BMAL1 in Hematologic Malignancies. Cancer Res. 2009, 69, 8447. [Google Scholar] [CrossRef]
- Erdem, J.S.; Skare, O.; Petersen-Overleir, M.; Noto, H.O.; Lie, J.A.S.; Reszka, E.; Peplonska, B.; Zienolddiny, S. Mechanisms of Breast Cancer in Shift Workers: DNA Methylation in Five Core Circadian Genes in Nurses Working Night Shifts. J. Cancer 2017, 8, 2876–2884. [Google Scholar] [CrossRef]
- Hoffman, A.E.; Zheng, T.; Stevens, R.G.; Ba, Y.; Zhang, Y.; Leaderer, D.; Yi, C.; Holford, T.R.; Zhu, Y. Clock-Cancer Connection in Non–Hodgkins Lymphoma: A Genetic Association Study and Pathway Analysis of the Circadian Gene Cryptochrome 2. Cancer Res. 2009, 69, 3605. [Google Scholar] [CrossRef] [PubMed]
- De Assis, L.V.M.; Kinker, G.S.; Moraes, M.N.; Markus, R.P.; Fernandes, P.A.; Castrucci, A.M.D.L. Expression of the Circadian Clock Gene BMAL1 Positively Correlates with Antitumor Immunity and Patient Survival in Metastatic Melanoma. Front. Oncol. 2018, 8, 185. [Google Scholar] [CrossRef] [PubMed]
- Oshima, T.; Takenoshita, S.; Akaike, M.; Kunisaki, C.; Fujii, S.; Nozaki, A.; Numata, K.; Shiozawa, M.; Rino, Y.; Tanaka, K.; et al. Expression of circadian genes correlates with liver metastasis and outcomes in colorectal cancer. Oncol. Rep. 2011, 25, 1439–1446. [Google Scholar] [CrossRef] [PubMed]
- Eisele, L.; Prinz, R.; Klein-Hitpass, L.; Nückel, H.; Lowinski, K.; Thomale, J.; Moeller, L.C.; Dührsen, U.; Dürig, J. Combined PER2 and CRY1 expression predicts outcome in chronic lymphocytic leukemia. Eur. J. Haematol. 2009, 83, 320–327. [Google Scholar] [CrossRef] [PubMed]
- Abreu, M.; Basti, A.; Genov, N.; Mazzoccoli, G.; Relógio, A. The reciprocal interplay between TNFα and the circadian clock impacts on cell proliferation and migration in Hodgkin lymphoma cells. Sci. Rep. 2018, 8, 11474. [Google Scholar] [CrossRef] [PubMed]
- Preußner, M.; Wilhelmi, I.; Schultz, A.-S.; Finkernagel, F.; Michel, M.; Möröy, T.; Heyd, F. Rhythmic U2af26 Alternative Splicing Controls PERIOD1 Stability and the Circadian Clock in Mice. Mol. Cell 2014, 54, 651–662. [Google Scholar] [CrossRef] [PubMed]
- El-Athman, R.; Fuhr, L.; Relógio, A. A Systems-Level Analysis Reveals Circadian Regulation of Splicing in Colorectal Cancer. EBioMedicine 2018, 33, 68–81. [Google Scholar] [CrossRef] [PubMed]
- Genov, N.; Basti, A.; Abreu, M.; Astaburuaga, R.; Relógio, A. A bioinformatic analysis identifies circadian expression of splicing factors and time-dependent alternative splicing events in the HD-MY-Z cell line. Sci. Rep. 2019, 9, 11062. [Google Scholar] [CrossRef] [PubMed]
- Kajimoto, J.; Matsumura, R.; Node, K.; Akashi, M. Potential role of the pancreatic hormone insulin in resetting human peripheral clocks. Genes Cell. 2018, 23, 393–399. [Google Scholar] [CrossRef] [PubMed]
- Pevet, P.; Challet, E. Melatonin: Both master clock output and internal time-giver in the circadian clocks network. J. Physiol. Paris 2011, 105, 170–182. [Google Scholar] [CrossRef] [PubMed]
- Semenza, G.L. Targeting HIF-1 for cancer therapy. Nat. Rev. Cancer 2003, 3, 721. [Google Scholar] [CrossRef]
- Cook, K.M.; Schofield, C.J. Therapeutic strategies that target. The HIF system. In Angiogenesis; Figg, W.D., Folkman, J., Eds.; Springer US: New York, NY, USA, 2008; pp. 359–373. [Google Scholar]
- Hunyor, I.; Cook, K.M. Models of intermittent hypoxia and obstructive sleep apnea: Molecular pathways and their contribution to cancer. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2018, 315, R669–R687. [Google Scholar] [CrossRef] [PubMed]
- Harris, A.L. Hypoxia—A key regulatory factor in tumour growth. Nat. Rev. Cancer 2002, 2, 38–47. [Google Scholar] [CrossRef]
- Yu, C.; Yang, S.; Fang, X.; Jiang, J.-X.; Sun, C.-Y.; Huang, T. Hypoxia disrupts the expression levels of circadian rhythm genes in hepatocellular carcinoma. Mol. Med. Rep. 2015, 11, 4002–4008. [Google Scholar] [CrossRef]
- Dimova, E.Y.; Jakupovic, M.; Kubaichuk, K.; Mennerich, D.; Chi, T.F.; Tamanini, F.; Oklejewicz, M.; Hänig, J.; Byts, N.; Mäkelä, K.A.; et al. The Circadian Clock Protein CRY1 Is a Negative Regulator of HIF-1α. iScience 2019, 13, 284–304. [Google Scholar] [CrossRef]
- Kobayashi, M.; Morinibu, A.; Koyasu, S.; Goto, Y.; Hiraoka, M.; Harada, H. A circadian clock gene, PER2, activates HIF-1 as an effector molecule for recruitment of HIF-1alpha to promoter regions of its downstream genes. FEBS J. 2017, 284, 3804–3816. [Google Scholar] [CrossRef]
- Wu, Y.; Tang, D.; Liu, N.; Xiong, W.; Huang, H.; Li, Y.; Ma, Z.; Zhao, H.; Chen, P.; Qi, X.; et al. Reciprocal Regulation between the Circadian Clock and Hypoxia Signaling at the Genome Level in Mammals. Cell Metab. 2017, 25, 73–85. [Google Scholar] [CrossRef] [PubMed]
- Peek, C.B.; Levine, D.C.; Cedernaes, J.; Taguchi, A.; Kobayashi, Y.; Tsai, S.J.; Bonar, N.A.; McNulty, M.R.; Ramsey, K.M.; Bass, J. Circadian Clock Interaction with HIF1α Mediates Oxygenic Metabolism and Anaerobic Glycolysis in Skeletal Muscle. Cell Metab. 2017, 25, 86–92. [Google Scholar] [CrossRef] [PubMed]
- Adamovich, Y.; Ladeuix, B.; Golik, M.; Koeners, M.P.; Asher, G. Rhythmic Oxygen Levels Reset Circadian Clocks through HIF1α. Cell Metab. 2017, 25, 93–101. [Google Scholar] [CrossRef] [PubMed]
- Egg, M.; Köblitz, L.; Hirayama, J.; Schwerte, T.; Folterbauer, C.; Kurz, A.; Fiechtner, B.; Möst, M.; Salvenmoser, W.; Sassone-Corsi, P.; et al. Linking Oxygen to Time: The Bidirectional Interaction Between the Hypoxic Signaling Pathway and the Circadian Clock. Chronobiol. Int. 2013, 30, 510–529. [Google Scholar] [CrossRef] [PubMed]
- Egg, M.; Köblitz, L.; Hirayama, J.; Schwerte, T.; Folterbauer, C.; Kurz, A.; Fiechtner, B.; Möst, M.; Salvenmoser, W.; Sassone-Corsi, P.; et al. The basic-helix–loop–helix-PAS orphan MOP3 forms transcriptionally active complexes with circadian and hypoxia factors. Proc. Natl. Acad. Sci. USA 1998, 95, 5474–5479. [Google Scholar]
- Egg, M.; Köblitz, L.; Hirayama, J.; Schwerte, T.; Folterbauer, C.; Kurz, A.; Fiechtner, B.; Möst, M.; Salvenmoser, W.; Sassone-Corsi, P.; et al. Activation of TGF-β/activin signalling resets the circadian clock through rapid induction of Dec1 transcripts. Nat. Cell Biol. 2008, 10, 1463. [Google Scholar]
- Adamovich, Y.; Ladeuix, B.; Sobel, J.; Manella, G.; Neufeld-Cohen, A.; Assadi, M.H.; Golik, M.; Kuperman, Y.; Tarasiuk, A.; Koeners, M.P.; et al. Oxygen and Carbon Dioxide Rhythms Are Circadian Clock Controlled and Differentially Directed by Behavioral Signals. Cell Metab. 2019, 7, 1092. [Google Scholar] [CrossRef] [PubMed]
- Cummins, E.P.; Selfridge, A.C.; Sporn, P.H.; Sznajder, J.I.; Taylor, C.T. Carbon dioxide-sensing in organisms and its implications for human disease. Cell. Mol. Life Sci. 2014, 71, 831–845. [Google Scholar] [CrossRef] [PubMed]
- Martinez, C.; Kerr, B.; Jin, C.; Cistulli, P.; Cook, K. Obstructive Sleep Apnea Activates HIF-1 in a Hypoxia Dose-Dependent Manner in HCT116 Colorectal Carcinoma Cells. Int. J. Mol. Sci. 2019, 20, 445. [Google Scholar] [CrossRef] [PubMed]
- De Assis, L.V.M.; Moraes, M.N.; Magalhães-Marques, K.K.; Kinker, G.S.; Da Silveira Cruz-Machado, S.; De Lauro Castrucci, A.M. Non-Metastatic Cutaneous Melanoma Induces Chronodisruption in Central and Peripheral Circadian Clocks. Int. J. Mol. Sci. 2018, 19, 1065. [Google Scholar] [CrossRef] [PubMed]
- Fuhr, L.; Abreu, M.; Carbone, A.; El-Athman, R.; Bianchi, F.; Laukkanen, M.O.; Mazzoccoli, G.; Relógio, A. The Interplay between Colon Cancer Cells and Tumour-Associated Stromal Cells Impacts the Biological Clock and Enhances Malignant Phenotypes. Cancers 2019, 11, 988. [Google Scholar] [CrossRef] [PubMed]
- Hojo, H.; Enya, S.; Arai, M.; Suzuki, Y.; Nojiri, T.; Kangawa, K.; Koyama, S.; Kawaoka, S. Remote reprogramming of hepatic circadian transcriptome by breast cancer. Oncotarget 2017, 8, 34128–34140. [Google Scholar] [CrossRef] [PubMed]
- Masri, S.; Papagiannakopoulos, T.; Kinouchi, K.; Liu, Y.; Cervantes, M.; Baldi, P.; Jacks, T.; Sassone-Corsi, P. Lung Adenocarcinoma Distally Rewires Hepatic Circadian Homeostasis. Cell 2016, 165, 896–909. [Google Scholar] [CrossRef] [PubMed]
- Gaucher, J.; Montellier, E.; Sassone-Corsi, P. Molecular Cogs: Interplay between Circadian Clock and Cell Cycle. Trends Cell Biol. 2018, 28, 368–379. [Google Scholar] [CrossRef] [PubMed]
- Shostak, A. Circadian Clock, Cell Division, and Cancer: From Molecules to Organism. Int. J. Mol. Sci. 2017, 18, 873. [Google Scholar] [CrossRef]
- Mori, T.; Binder, B.; Johnson, C.H. Circadian gating of cell division in cyanobacteria growing with average doubling times of less than 24 h. Proc. Natl. Acad. Sci. USA 1996, 93, 10183–10188. [Google Scholar] [CrossRef] [PubMed]
- Mori, T.; Johnson, C.H. Independence of Circadian Timing from Cell Division in Cyanobacteria. J. Bacteriol. 2001, 183, 2439–2444. [Google Scholar] [CrossRef]
- Bolige, A.; Hagiwara, S.-Y.; Zhang, Y.; Goto, K. Circadian G2 Arrest as Related to Circadian Gating of Cell Population Growth in Euglena. Plant. Cell Physiol. 2005, 46, 931–936. [Google Scholar] [CrossRef][Green Version]
- Dong, G.; Yang, Q.; Wang, Q.; Kim, Y.I.; Wood, T.L.; Osteryoung, K.W.; van Oudenaarden, A.; Golden, S.S. Elevated ATPase activity of KaiC applies a circadian checkpoint on cell division in Synechococcus elongatus. Cell 2010, 140, 529–539. [Google Scholar] [CrossRef]
- Buchi, K.N.; Moore, J.G.; Hrushesky, W.J.M.; Sothern, R.B.; Rubin, N.H. Circadian rhythm of cellular proliferation in the human rectal mucosa. Gastroenterology 1991, 101, 410–415. [Google Scholar] [CrossRef]
- Scheving, L.E.; Burns, E.R.; Pauly, J.E.; Tsai, T.-H. Circadian variation in cell division of the mouse alimentary tract, bone marrow and corneal epithelium. Anat. Rec. 1978, 191, 479–486. [Google Scholar] [CrossRef] [PubMed]
- Smaaland, R.; Laerum, O.; Lote, K.; Sletvold, O.; Sothern, R.; Bjerknes, R. DNA synthesis in human bone marrow is circadian stage dependent. Blood 1991, 77, 2603–2611. [Google Scholar] [PubMed]
- Al-Nuaimi, Y.; Hardman, J.A.; Bíró, T.; Haslam, I.S.; Philpott, M.P.; Tóth, B.I.; Farjo, N.; Farjo, B.; Baier, G.; Watson, R.E.B.; et al. A Meeting of Two Chronobiological Systems: Circadian Proteins Period1 and BMAL1 Modulate the Human Hair Cycle Clock. J. Investig. Dermatol. 2014, 134, 610–619. [Google Scholar] [CrossRef] [PubMed]
- Bjarnason, G.A.; Jordan, R. Rhythms in human gastrointestinal mucosa and skin. Chronobiol. Int. 2002, 19, 129–140. [Google Scholar] [CrossRef] [PubMed]
- Spörl, F.; Korge, S.; Jürchott, K.; Wunderskirchner, M.; Schellenberg, K.; Heins, S.; Specht, A.; Stoll, C.; Klemz, R.; Maier, B.; et al. Krüppel-like factor 9 is a circadian transcription factor in human epidermis that controls proliferation of keratinocytes. Proc. Natl. Acad. Sci. USA 2012, 109, 10903–10908. [Google Scholar] [CrossRef]
- Janich, P.; Pascual, G.; Merlos-Suárez, A.; Batlle, E.; Ripperger, J.; Albrecht, U.; Cheng, H.-Y.M.; Obrietan, K.; Di Croce, L.; Benitah, S.A. The circadian molecular clock creates epidermal stem cell heterogeneity. Nature 2011, 480, 209. [Google Scholar] [CrossRef] [PubMed]
- Janich, P.; Toufighi, K.; Solanas, G.; Luis, N.M.; Minkwitz, S.; Serrano, L.; Lehner, B.; Benitah, S.A. Human Epidermal Stem Cell Function Is Regulated by Circadian Oscillations. Cell Stem Cell 2013, 13, 745–753. [Google Scholar] [CrossRef] [PubMed]
- Bouchard-Cannon, P.; Mendoza-Viveros, L.; Yuen, A.; Kærn, M.; Cheng, H.-Y.M. The Circadian Molecular Clock Regulates Adult Hippocampal Neurogenesis by Controlling the Timing of Cell-Cycle Entry and Exit. Cell Rep. 2013, 5, 961–973. [Google Scholar] [CrossRef] [PubMed]
- Méndez-Ferrer, S.; Lucas, D.; Battista, M.; Frenette, P.S. Haematopoietic stem cell release is regulated by circadian oscillations. Nature 2008, 452, 442. [Google Scholar] [CrossRef]
- Dekens, M.P.S.; Santoriello, C.; Vallone, D.; Grassi, G.; Whitmore, D.; Foulkes, N.S. Light Regulates the Cell Cycle in Zebrafish. Curr. Biol. 2003, 13, 2051–2057. [Google Scholar] [CrossRef]
- Laranjeiro, R.; Tamai, T.K.; Peyric, E.; Krusche, P.; Ott, S.; Whitmore, D. Cyclin-dependent kinase inhibitor p20 controls circadian cell-cycle timing. Proc. Natl. Acad. Sci. USA 2013, 110, 6835–6840. [Google Scholar] [CrossRef] [PubMed]
- Plikus, M.V.; Vollmers, C.; de la Cruz, D.; Chaix, A.; Ramos, R.; Panda, S.; Chuong, C.-M. Local circadian clock gates cell cycle progression of transient amplifying cells during regenerative hair cycling. Proc. Natl. Acad. Sci. USA 2013, 110, E2106–E2115. [Google Scholar] [CrossRef] [PubMed]
- Feillet, C.; van der Horst, G.T.J.; Levi, F.; Rand, D.A.; Delaunay, F. Coupling between the Circadian Clock and Cell Cycle Oscillators: Implication for Healthy Cells and Malignant Growth. Front. Neurol. 2015, 6, 96. [Google Scholar] [CrossRef] [PubMed]
- Farshadi, E.; Yan, J.; Leclere, P.; Goldbeter, A.; Chaves, I.; van der Horst, G.T.J. The positive circadian regulators CLOCK and BMAL1 control G2/M cell cycle transition through Cyclin B1. Cell Cycle 2019, 18, 16–33. [Google Scholar] [CrossRef] [PubMed]
- Kowalska, E.; Ripperger, J.A.; Hoegger, D.C.; Bruegger, P.; Buch, T.; Birchler, T.; Mueller, A.; Albrecht, U.; Contaldo, C.; Brown, S.A. NONO couples the circadian clock to the cell cycle. Proc. Natl. Acad. Sci. USA 2013, 110, 1592–1599. [Google Scholar] [CrossRef] [PubMed]
- Karpowicz, P.; Zhang, Y.; Hogenesch, J.B.; Emery, P.; Perrimon, N. The Circadian Clock Gates the Intestinal Stem Cell Regenerative State. Cell Rep. 2013, 3, 996–1004. [Google Scholar] [CrossRef] [PubMed]
- Yeom, M.; Pendergast, J.S.; Ohmiya, Y.; Yamazaki, S. Circadian-independent cell mitosis in immortalized fibroblasts. Proc. Natl. Acad. Sci. USA 2010, 107, 9665–9670. [Google Scholar] [CrossRef] [PubMed]
- Masri, S.; Cervantes, M.; Sassone-Corsi, P. The circadian clock and cell cycle: Interconnected biological circuits. Curr. Opin. Cell Biol. 2013, 25, 730–734. [Google Scholar] [CrossRef]
- Ou, J.; Li, H.; Qiu, P.; Li, Q.; Chang, H.-C.; Tang, Y.-C. CDK9 modulates circadian clock by attenuating REV-ERBα activity. Biochem. Biophys. Res. Commun. 2019, 513, 967–973. [Google Scholar] [CrossRef]
- Lowrey, P.L.; Takahashi, J.S. Genetics of circadian rhythms in mammalian model organisms. In Advances in Genetics; Brody, S., Ed.; Academic Press: Cambridge, MA, USA, 2011; Chapter 6; pp. 175–230. [Google Scholar]
- Sulli, G.; Lam, M.T.Y.; Panda, S. Interplay between Circadian Clock and Cancer: New Frontiers for Cancer Treatment. Trends Cancer 2019, 5, 475–494. [Google Scholar] [CrossRef]
- Kettner, N.M.; Voicu, H.; Finegold, M.J.; Coarfa, C.; Sreekumar, A.; Putluri, N.; Katchy, C.A.; Lee, C.; Moore, D.D.; Fu, L. Circadian Homeostasis of Liver Metabolism Suppresses Hepatocarcinogenesis. Cancer Cell 2016, 30, 909–924. [Google Scholar] [CrossRef] [PubMed]
- Guerrero-Vargas, N.N.; Navarro-Espíndola, R.; Guzmán-Ruíz, M.A.; Basualdo, M.d.C.; Espitia-Bautista, E.; López-Bago, A.; Lascurain, R.; Córdoba-Manilla, C.; Buijs, R.M.; Escobar, C. Circadian disruption promotes tumor growth by anabolic host metabolism; experimental evidence in a rat model. BMC Cancer 2017, 17, 625. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Song, L.; Liu, M.; Ge, R.; Zhou, Q.; Liu, W.; Li, R.; Qie, J.; Zhen, B.; Wang, Y.; et al. A proteomics landscape of circadian clock in mouse liver. Nat. Commun. 2018, 9, 1553. [Google Scholar] [CrossRef] [PubMed]
- Szabó, Á.; Papin, C.; Cornu, D.; Chélot, E.; Lipinszki, Z.; Udvardy, A.; Redeker, V.; Mayor, U.; Rouyer, F. Ubiquitylation Dynamics of the Clock Cell Proteome and TIMELESS during a Circadian Cycle. Cell Rep. 2018, 23, 2273–2282. [Google Scholar] [CrossRef] [PubMed]
- Zheng, X.; Sehgal, A. AKT and TOR Signaling Set the Pace of the Circadian Pacemaker. Curr. Biol. 2010, 20, 1203–1208. [Google Scholar] [CrossRef]
- Luciano, A.K.; Zhou, W.; Santana, J.M.; Kyriakides, C.; Velazquez, H.; Sessa, W.C. CLOCK phosphorylation by AKT regulates its nuclear accumulation and circadian gene expression in peripheral tissues. J. Biol. Chem. 2018, 293, 9126–9136. [Google Scholar] [CrossRef]
- Dallmann, R.; Viola, A.U.; Tarokh, L.; Cajochen, C.; Brown, S.A. The human circadian metabolome. Proc. Natl. Acad. Sci. USA 2012, 109, 2625–2629. [Google Scholar] [CrossRef]
- Eckel-Mahan, K.L.; Patel, V.R.; Mohney, R.P.; Vignola, K.S.; Baldi, P.; Sassone-Corsi, P. Coordination of the transcriptome and metabolome by the circadian clock. Proc. Natl. Acad. Sci. USA 2012, 109, 5541–5546. [Google Scholar] [CrossRef]
- Minami, Y.; Kasukawa, T.; Kakazu, Y.; Iigo, M.; Sugimoto, M.; Ikeda, S.; Yasui, A.; van der Horst, G.T.J.; Soga, T.; Ueda, H.R. Measurement of internal body time by blood metabolomics. Proc. Natl. Acad. Sci. USA 2009, 106, 9890–9895. [Google Scholar] [CrossRef]
- Verlande, A.; Masri, S. Circadian Clocks and Cancer: Timekeeping Governs Cellular Metabolism. Trends Endocrinol. Metab. 2019, 30, 445–458. [Google Scholar] [CrossRef] [PubMed]
- Masri, S.; Sassone-Corsi, P. The emerging link between cancer, metabolism, and circadian rhythms. Nat. Med. 2018, 24, 1795–1803. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Mauvoisin, D.; Martin, E.; Atger, F.; Galindo, A.N.; Dayon, L.; Sizzano, F.; Palini, A.; Kussmann, M.; Waridel, P.; et al. Nuclear Proteomics Uncovers Diurnal Regulatory Landscapes in Mouse Liver. Cell Metab. 2017, 25, 102–117. [Google Scholar] [CrossRef]
- Mure, L.S.; Le, H.D.; Benegiamo, G.; Chang, M.W.; Rios, L.; Jillani, N.; Ngotho, M.; Kariuki, T.; Dkhissi-Benyahya, O.; Cooper, H.M.; et al. Diurnal transcriptome atlas of a primate across major neural and peripheral tissues. Science 2018, 359, eaao0318. [Google Scholar] [CrossRef] [PubMed]
- Kang, T.-H.; Reardon, J.T.; Kemp, M.; Sancar, A. Circadian oscillation of nucleotide excision repair in mammalian brain. Proc. Natl. Acad. Sci. USA 2009, 106, 2864–2867. [Google Scholar] [CrossRef] [PubMed]
- Oklejewicz, M.; Destici, E.; Tamanini, F.; Hut, R.A.; Janssens, R.; van der Horst, G.T.J. Phase Resetting of the Mammalian Circadian Clock by DNA Damage. Curr. Biol. 2008, 18, 286–291. [Google Scholar] [CrossRef] [PubMed]
- Papp, S.J.; Huber, A.-L.; Jordan, S.D.; Kriebs, A.; Nguyen, M.; Moresco, J.J.; Yates, J.R., III; Lamia, K.A. DNA damage shifts circadian clock time via Hausp-dependent Cry1 stabilization. eLife 2015, 4, e04883. [Google Scholar] [CrossRef] [PubMed]
- Logan, R.W.; Zhang, C.; Murugan, S.; O’Connell, S.; Levitt, D.; Rosenwasser, A.M.; Sarkar, D.K. Chronic Shift-Lag Alters the Circadian Clock of NK Cells and Promotes Lung Cancer Growth in Rats. J. Immunol. 2012, 188, 2583–2591. [Google Scholar] [CrossRef]
- Curtis, A.M.; Fagundes, C.T.; Yang, G.; Palsson-McDermott, E.M.; Wochal, P.; McGettrick, A.F.; Foley, N.H.; Early, J.O.; Chen, L.; Zhang, H.; et al. Circadian control of innate immunity in macrophages by miR-155 targeting Bmal1. Proc. Natl. Acad. Sci. USA 2015, 112, 7231–7236. [Google Scholar] [CrossRef]
- Spengler, M.L.; Kuropatwinski, K.K.; Comas, M.; Gasparian, A.V.; Fedtsova, N.; Gleiberman, A.S.; Gitlin, I.I.; Artemicheva, N.M.; Deluca, K.A.; Gudkov, A.V.; et al. Core circadian protein CLOCK is a positive regulator of NF-κB–mediated transcription. Proc. Natl. Acad. Sci. USA 2012, 109, E2457–E2465. [Google Scholar] [CrossRef]
- Narasimamurthy, R.; Hatori, M.; Nayak, S.K.; Liu, F.; Panda, S.; Verma, I.M. Circadian clock protein cryptochrome regulates the expression of proinflammatory cytokines. Proc. Natl. Acad. Sci. USA 2012, 109, 12662–12667. [Google Scholar] [CrossRef] [PubMed]
- Scheiermann, C.; Gibbs, J.; Ince, L.; Loudon, A. Clocking in to immunity. Nat. Rev. Immunol. 2018, 18, 423–437. [Google Scholar] [CrossRef] [PubMed]
- Sulli, G.; Rommel, A.; Wang, X.; Kolar, M.J.; Puca, F.; Saghatelian, A.; Plikus, M.V.; Verma, I.M.; Panda, S. Pharmacological activation of REV-ERBs is lethal in cancer and oncogene-induced senescence. Nature 2018, 553, 351. [Google Scholar] [CrossRef] [PubMed]
- De Mei, C.; Ercolani, L.; Parodi, C.; Veronesi, M.; Lo Vecchio, C.; Bottegoni, G.; Torrente, E.; Scarpelli, R.; Marotta, R.; Ruffili, R.; et al. Dual inhibition of REV-ERBβ and autophagy as a novel pharmacological approach to induce cytotoxicity in cancer cells. Oncogene 2015, 34, 2597–2608. [Google Scholar] [CrossRef] [PubMed]
- De Mei, C.; Ercolani, L.; Parodi, C.; Veronesi, M.; Lo Vecchio, C.; Bottegoni, G.; Torrente, E.; Scarpelli, R.; Marotta, R.; Ruffili, R.; et al. A PERK-miR-211 axis suppresses circadian regulators and protein synthesis to promote cancer cell survival. Nat. Cell Biol. 2018, 20, 104–115. [Google Scholar]
- Chen, Z.; Yoo, S.-H.; Park, Y.-S.; Kim, K.-H.; Wei, S.; Buhr, E.; Ye, Z.-Y.; Pan, H.-L.; Takahashi, J.S. Identification of diverse modulators of central and peripheral circadian clocks by high-throughput chemical screening. Proc. Natl. Acad. Sci. USA 2012, 109, 101–106. [Google Scholar] [CrossRef]
- Isojima, Y.; Nakajima, M.; Ukai, H.; Fujishima, H.; Yamada, R.G.; Masumoto, K.-h.; Kiuchi, R.; Ishida, M.; Ukai-Tadenuma, M.; Minami, Y.; et al. CKIε/δ-dependent phosphorylation is a temperature-insensitive, period-determining process in the mammalian circadian clock. Proc. Natl. Acad. Sci. USA 2009, 106, 15744–15749. [Google Scholar] [CrossRef]
- Hirota, T.; Lewis, W.G.; Liu, A.C.; Lee, J.W.; Schultz, P.G.; Kay, S.A. A chemical biology approach reveals period shortening of the mammalian circadian clock by specific inhibition of GSK-3β. Proc. Natl. Acad. Sci. USA 2008, 105, 20746–20751. [Google Scholar] [CrossRef]
- Hirota, T.; Lee, J.W.; Lewis, W.G.; Zhang, E.E.; Breton, G.; Liu, X.; Garcia, M.; Peters, E.C.; Etchegaray, J.-P.; Traver, D.; et al. High-Throughput Chemical Screen Identifies a Novel Potent Modulator of Cellular Circadian Rhythms and Reveals CKIα as a Clock Regulatory Kinase. PLoS Biol. 2010, 8, e1000559. [Google Scholar] [CrossRef]
- Oshima, T.; Niwa, Y.; Kuwata, K.; Srivastava, A.; Hyoda, T.; Tsuchiya, Y.; Kumagai, M.; Tsuyuguchi, M.; Tamaru, T.; Sugiyama, A.; et al. Cell-based screen identifies a new potent and highly selective CK2 inhibitor for modulation of circadian rhythms and cancer cell growth. Sci Adv. 2019, 5, eaau9060. [Google Scholar] [CrossRef]
- Cook, K.M.; Hilton, S.T.; Mecinovic, J.; Motherwell, W.B.; Figg, W.D.; Schofield, C.J. Epidithiodiketopiperazines block the interaction between hypoxia-inducible factor-1alpha (HIF-1alpha) and p300 by a zinc ejection mechanism. J. Biol. Chem. 2009, 284, 26831–26838. [Google Scholar] [CrossRef] [PubMed]
- Goey, A.K.L.; Chau, C.H.; Sissung, T.M.; Cook, K.M.; Venzon, D.J.; Castro, A.; Ransom, T.R.; Henrich, C.J.; McKee, T.C.; McMahon, J.B.; et al. Screening and Biological Effects of Marine Pyrroloiminoquinone Alkaloids: Potential Inhibitors of the HIF-1α/p300 Interaction. J. Nat. Prod. 2016, 79, 1267–1275. [Google Scholar] [CrossRef] [PubMed]
- Chan, S.T.; Patel, P.R.; Ransom, T.R.; Henrich, C.J.; McKee, T.C.; Goey, A.K.; Cook, K.M.; Figg, W.D.; McMahon, J.B.; Schnermann, M.J.; et al. Structural Elucidation and Synthesis of Eudistidine A: An Unusual Polycyclic Marine Alkaloid that Blocks Interaction of the Protein Binding Domains of p300 and HIF-1alpha. J. Am. Chem. Soc. 2015, 137, 5569–5575. [Google Scholar] [CrossRef] [PubMed]
- Harris, E.M.; Strope, J.D.; Beedie, S.L.; Huang, P.A.; Goey, A.K.L.; Cook, K.M.; Schofield, C.J.; Chau, C.H.; Cadelis, M.M.; Copp, B.R.; et al. Preclinical Evaluation of Discorhabdins in Antiangiogenic and Antitumor Models. Mar. Drugs 2018, 16, 241. [Google Scholar] [CrossRef] [PubMed]
- Kung, A.L.; Zabludoff, S.D.; France, D.S.; Freedman, S.J.; Tanner, E.A.; Vieira, A.; Cornell-Kennon, S.; Lee, J.; Wang, B.; Wang, J.; et al. Small molecule blockade of transcriptional coactivation of the hypoxia-inducible factor pathway. Cancer Cell 2004, 6, 33–43. [Google Scholar] [CrossRef] [PubMed]
- Kung, A.L.; Zabludoff, S.D.; France, D.S.; Freedman, S.J.; Tanner, E.A.; Vieira, A.; Cornell-Kennon, S.; Lee, J.; Wang, B.; Wang, J.; et al. Inhibition of the HIF1alpha-p300 interaction by quinone- and indandione-mediated ejection of structural Zn(II). Eur. J. Med. Chem. 2015, 94, 509–516. [Google Scholar]
- Reece, K.M.; Richardson, E.D.; Cook, K.M.; Campbell, T.J.; Pisle, S.T.; Holly, A.J.; Venzon, D.J.; Liewehr, D.J.; Chau, C.H.; Price, D.K.; et al. Epidithiodiketopiperazines (ETPs) exhibit in vitro antiangiogenic and in vivo antitumor activity by disrupting the HIF-1alpha/p300 complex in a preclinical model of prostate cancer. Mol. Cancer 2014, 13, 91. [Google Scholar] [CrossRef] [PubMed]
- Miranda, E.; Nordgren, I.K.; Male, A.L.; Lawrence, C.E.; Hoakwie, F.; Cuda, F.; Court, W.; Fox, K.R.; Townsend, P.A.; Packham, G.K.; et al. A Cyclic Peptide Inhibitor of HIF-1 Heterodimerization That Inhibits Hypoxia Signaling in Cancer Cells. J. Am. Chem. Soc. 2013, 135, 10418–10425. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.; Zhang, H.; Qian, D.Z.; Rey, S.; Liu, J.O.; Semenza, G.L. Acriflavine inhibits HIF-1 dimerization, tumor growth, and vascularization. Proc. Natl. Acad. Sci. USA 2009, 106, 17910–17915. [Google Scholar] [CrossRef]
- Chen, W.; Hill, H.; Christie, A.; Kim, M.S.; Holloman, E.; Pavia-Jimenez, A.; Homayoun, F.; Ma, Y.; Patel, N.; Yell, P.; et al. Targeting renal cell carcinoma with a HIF-2 antagonist. Nature 2016, 539, 112–117. [Google Scholar] [CrossRef]
- Wallace, E.M.; Rizzi, J.P.; Han, G.; Wehn, P.M.; Cao, Z.; Du, X.; Cheng, T.; Czerwinski, R.M.; Dixon, D.D.; Goggin, B.S.; et al. A Small-Molecule Antagonist of HIF2α Is Efficacious in Preclinical Models of Renal Cell Carcinoma. Cancer Res. 2016, 76, 5491–5500. [Google Scholar] [CrossRef] [PubMed]
- Rogers, J.L.; Bayeh, L.; Scheuermann, T.H.; Longgood, J.; Key, J.; Naidoo, J.; Melito, L.; Shokri, C.; Frantz, D.E.; Bruick, R.K.; et al. Development of Inhibitors of the PAS-B Domain of the HIF-2α Transcription Factor. J. Med. Chem. 2013, 56, 1739–1747. [Google Scholar] [CrossRef] [PubMed]
- Scheuermann, T.H.; Li, Q.; Ma, H.-W.; Key, J.; Zhang, L.; Chen, R.; Garcia, J.A.; Naidoo, J.; Longgood, J.; Frantz, D.E.; et al. Allosteric inhibition of hypoxia inducible factor-2 with small molecules. Nat. Chem. Biol. 2013, 9, 271–276. [Google Scholar] [CrossRef] [PubMed]
- Mabjeesh, N.J.; Post, D.E.; Willard, M.T.; Kaur, B.; Van Meir, E.G.; Simons, J.W.; Zhong, H. Geldanamycin induces degradation of hypoxia-inducible factor 1alpha protein via the proteosome pathway in prostate cancer cells. Cancer Res. 2002, 62, 2478–2482. [Google Scholar]
- Kataria, N.; Martinez, C.A.; Kerr, B.; Zaiter, S.S.; Morgan, M.; McAlpine, S.R.; Cook, K.M. C-Terminal HSP90 Inhibitors Block the HIF-1 Hypoxic Response by Degrading HIF-1α through the Oxygen-Dependent Degradation Pathway. Cell. Physiol. Biochem. 2019, 53, 480–495. [Google Scholar]
- Kong, D.; Park, E.J.; Stephen, A.G.; Calvani, M.; Cardellina, J.H.; Monks, A.; Fisher, R.J.; Shoemaker, R.H.; Melillo, G. Echinomycin, a small-molecule inhibitor of hypoxia-inducible factor-1 DNA-binding activity. Cancer Res. 2005, 65, 9047–9055. [Google Scholar] [CrossRef]
- Hosoda, H.; Kato, K.; Asano, H.; Ito, M.; Kato, H.; Iwamoto, T.; Suzuki, A.; Masushige, S.; Kida, S. CBP/p300 is a cell type-specific modulator of CLOCK/BMAL1-mediated transcription. Mol. Brain 2009, 2, 34. [Google Scholar] [CrossRef]
- Dallmann, R.; Okyar, A.; Lévi, F. Dosing-time makes the poison: Circadian regulation and pharmacotherapy. Trends Mol. Med. 2016, 22, 430–445. [Google Scholar] [CrossRef]
- Lévi, F.; Zidani, R.; Misset, J.-L. Randomised multicentre trial of chronotherapy with oxaliplatin, fluorouracil, and folinic acid in metastatic colorectal cancer. Lancet 1997, 350, 681–686. [Google Scholar] [CrossRef]
- Bouchahda, M.; Adam, R.; Giacchetti, S.; Castaing, D.; Brezault-Bonnet, C.; Hauteville, D.; Innominato, P.F.; Focan, C.; Machover, D.; Lévi, F. Rescue chemotherapy using multidrug chronomodulated hepatic arterial infusion for patients with heavily pretreated metastatic colorectal cancer. Cancer 2009, 115, 4990–4999. [Google Scholar] [CrossRef]
- Lee, Y.; Lahens, N.F.; Zhang, S.; Bedont, J.; Field, J.M.; Sehgal, A. G1/S cell cycle regulators mediate effects of circadian dysregulation on tumor growth and provide targets for timed anticancer treatment. PLoS Biol. 2019, 17, e3000228. [Google Scholar] [CrossRef] [PubMed]
- Iurisci, I.; Filipski, E.; Reinhardt, J.; Bach, S.; Gianella-Borradori, A.; Iacobelli, S.; Meijer, L.; Lévi, F. Improved Tumor Control through Circadian Clock Induction by Seliciclib, a Cyclin-Dependent Kinase Inhibitor. Cancer Res. 2006, 66, 10720. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Giacchetti, S.; Parouchev, A.; Hadadi, E.; Li, X.; Dallmann, R.; Xandri-Monje, H.; Portier, L.; Adam, R.; Lévi, F.; et al. Dosing time dependent in vitro pharmacodynamics of Everolimus despite a defective circadian clock. Cell Cycle 2018, 17, 33–42. [Google Scholar] [CrossRef] [PubMed]
- Schernhammer, E.S.; Schulmeister, K. Melatonin and cancer risk: Does light at night compromise physiologic cancer protection by lowering serum melatonin levels? Br. J. Cancer 2004, 90, 941–943. [Google Scholar] [CrossRef] [PubMed]
- Haus, E.L.; Smolensky, M.H. Shift work and cancer risk: Potential mechanistic roles of circadian disruption, light at night, and sleep deprivation. Sleep Med. Rev. 2013, 17, 273–284. [Google Scholar] [CrossRef] [PubMed]
- Greene, M.W. Circadian rhythms and tumor growth. Cancer Lett. 2012, 318, 115–123. [Google Scholar] [CrossRef] [PubMed]
- Viswanathan, A.N.; Schernhammer, E.S. Circulating melatonin and the risk of breast and endometrial cancer in women. Cancer Lett. 2009, 281, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Cucina, A.; Proietti, S.; D’Anselmi, F.; Coluccia, P.; Dinicola, S.; Frati, L.; Bizzarri, M. Evidence for a biphasic apoptotic pathway induced by melatonin in MCF-7 breast cancer cells. J. Pineal Res. 2009, 46, 172–180. [Google Scholar] [CrossRef] [PubMed]
- Mediavilla, M.D.; Sanchez-Barcelo, E.J.; Tan, D.X.; Manchester, L.; Reiter, R.J. Basic mechanisms involved in the anti-cancer effects of melatonin. Curr. Med. Chem. 2010, 17, 4462–4481. [Google Scholar] [CrossRef]
- Blask, D.E.; Sauer, L.A.; Dauchy, R.T. Melatonin as a chronobiotic/anticancer agent: Cellular, biochemical, and molecular mechanisms of action and their implications for circadian-based cancer therapy. Curr. Top. Med. Chem. 2002, 2, 113–132. [Google Scholar] [CrossRef]
- Majidinia, M.; Sadeghpour, A.; Mehrzadi, S.; Reiter, R.J.; Khatami, N.; Yousefi, B. Melatonin: A pleiotropic molecule that modulates DNA damage response and repair pathways. J. Pineal Res. 2017, 63, e12416. [Google Scholar] [CrossRef]
- Jung-Hynes, B.; Huang, W.; Reiter, R.J.; Ahmad, N. Melatonin resynchronizes dysregulated circadian rhythm circuitry in human prostate cancer cells. J. Pineal Res. 2010, 49, 60–68. [Google Scholar] [CrossRef] [PubMed]
- Anisimov, V.N.; Alimova, I.N.; Baturin, D.A.; Popovich, I.G.; Zabezhinski, M.A.; Manton, K.G.; Semenchenko, A.V.; Yashin, A.I. The effect of melatonin treatment regimen on mammary adenocarcinoma development in HER-2/neu transgenic mice. Int. J. Cancer 2003, 103, 300–305. [Google Scholar] [CrossRef] [PubMed]
- Baturin, D.A.; Alimova, I.N.; Anisimov, V.N.; Popovich, I.G.; Zabezhinski, M.A.; Provinciali, M.; Mancini, R.; Franceschi, C. The effect of light regimen and melatonin on the development of spontaneous mammary tumors in HER-2/neu transgenic mice is related to a downregulation of HER-2/neu gene expression. Neuroendocrinol. Lett. 2001, 22, 441–447. [Google Scholar] [PubMed]
- Liu, Z.; Gan, L.; Luo, D.; Sun, C. Melatonin promotes circadian rhythm-induced proliferation through Clock/histone deacetylase 3/c-Myc interaction in mouse adipose tissue. J. Pineal Res. 2017, 62, e12383. [Google Scholar] [CrossRef] [PubMed]
- Scheiermann, C.; Kunisaki, Y.; Frenette, P.S. Circadian control of the immune system. Nat. Rev. Immunol. 2013, 13, 190. [Google Scholar] [CrossRef]
- Soták, M.; Sumová, A.; Pácha, J. Cross-talk between the circadian clock and the cell cycle in cancer. Ann. Med. 2014, 46, 221–232. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Morgan, M.N.; Dvuchbabny, S.; Martinez, C.-A.; Kerr, B.; Cistulli, P.A.; Cook, K.M. The Cancer Clock Is (Not) Ticking: Links between Circadian Rhythms and Cancer. Clocks & Sleep 2019, 1, 435-458. https://doi.org/10.3390/clockssleep1040034
Morgan MN, Dvuchbabny S, Martinez C-A, Kerr B, Cistulli PA, Cook KM. The Cancer Clock Is (Not) Ticking: Links between Circadian Rhythms and Cancer. Clocks & Sleep. 2019; 1(4):435-458. https://doi.org/10.3390/clockssleep1040034
Chicago/Turabian StyleMorgan, Monica N., Sapir Dvuchbabny, Chloe-Anne Martinez, Bernadette Kerr, Peter A. Cistulli, and Kristina M. Cook. 2019. "The Cancer Clock Is (Not) Ticking: Links between Circadian Rhythms and Cancer" Clocks & Sleep 1, no. 4: 435-458. https://doi.org/10.3390/clockssleep1040034
APA StyleMorgan, M. N., Dvuchbabny, S., Martinez, C.-A., Kerr, B., Cistulli, P. A., & Cook, K. M. (2019). The Cancer Clock Is (Not) Ticking: Links between Circadian Rhythms and Cancer. Clocks & Sleep, 1(4), 435-458. https://doi.org/10.3390/clockssleep1040034




