Soil Microbial Communities in Corn Fields Treated with Atoxigenic Aspergillus flavus
Abstract
1. Introduction
2. Materials and Methods
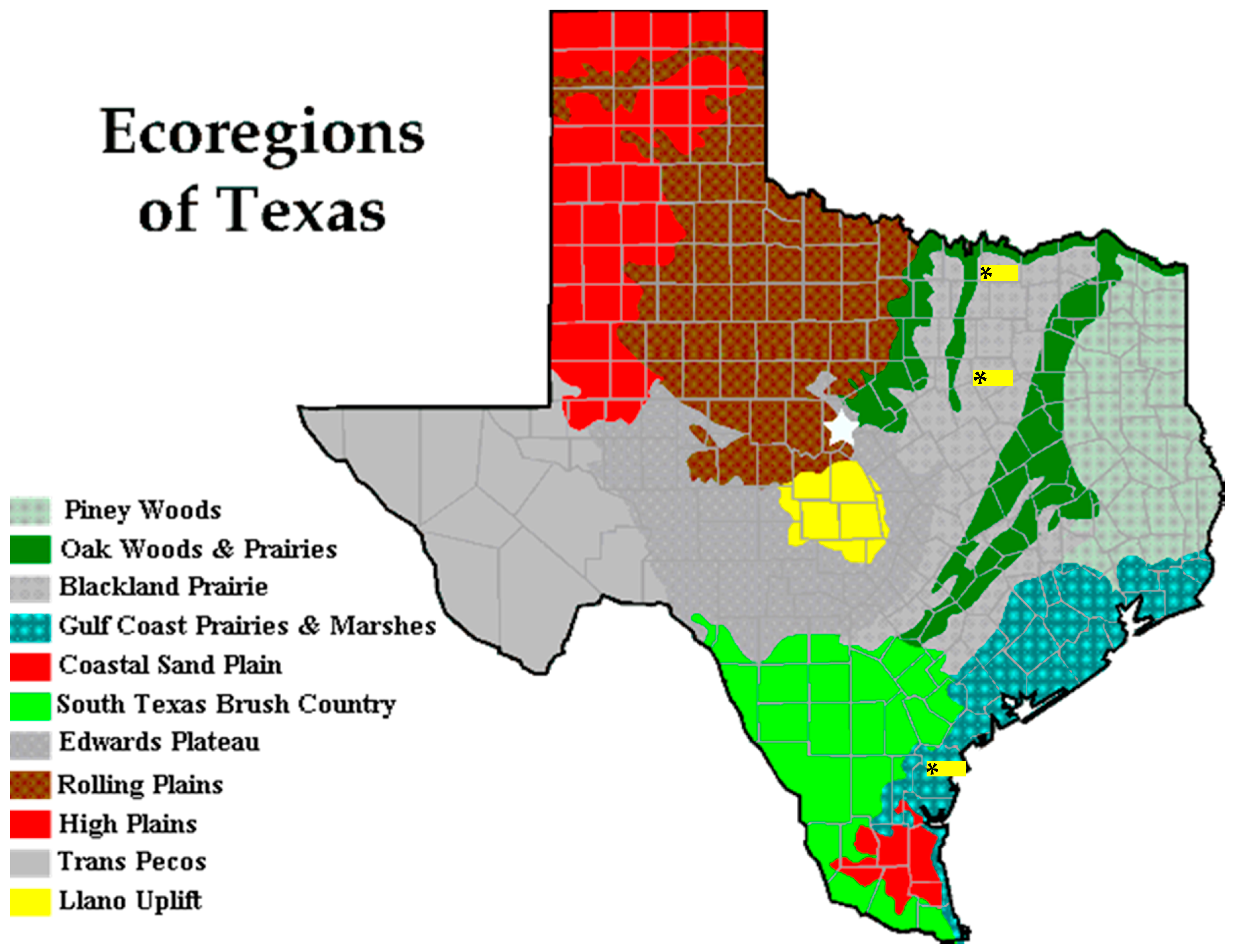
2.1. Description of Field Sites
2.2. Environmental Parameters
2.3. Soil Sampling
2.4. Soil Water Content Determination and EL-FAME Analysis for Soil Microbial Community Structure
2.5. Statistical Analyses
3. Results
3.1. Soil Water Content, Protozoal Abundances, and Total FAMEs
3.2. Fungal and Bacterial Abundances, and Their Ratios
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| AMF | arbuscular mycorrhizal fungi |
| EPA | Environmental Protection Agency |
| FAME | fatty acid methyl ester |
References
- Mehl, H.L.; Cotty, P.J. Variation in Competitive Ability among Isolates of Aspergillus flavus from Different Vegetative Compatibility Groups During Maize Infection. Phytopathology 2010, 100, 150–159. [Google Scholar] [CrossRef] [PubMed]
- Cotty, P.J. Influence of Field Application of an Atoxigenic Strain of Aspergillus flavus on the Populations of A. flavus Infecting Cotton Bolls and on the Aflatoxin Content of Cottonseed. Phytopathology 1994, 84, 1270. [Google Scholar] [CrossRef]
- Klich, M.A. Aspergillus flavus: The major producer of aflatoxin. Mol. Plant Pathol. 2007, 8, 713–722. [Google Scholar] [CrossRef] [PubMed]
- Bacon, C.W.; Nelson, P.E. Fumonisin Production in Corn by Toxigenic Strains of Fusarium moniliforme and Fusarium proliferatum. J. Food Prot. 1994, 57, 514–521. [Google Scholar] [CrossRef] [PubMed]
- Payne, G.A.; Widstrom, N.W. Aflatoxin in maize. Crit. Rev. Plant Sci. 1992, 10, 423–440. [Google Scholar] [CrossRef]
- Nishie, K.; Cole, R.J.; Domer, J.W. Toxicity and neurophamacology of cyclopiazonic acid. Food Chem. Toxicol. 1985, 9, 831–839. [Google Scholar] [CrossRef]
- Jensen, A.B.; Aronstein, K.; Flores, J.M.; Vojvodic, S.; Palacio, M.A.; Spivak, M. Standard methods for fungal brood disease research. J. Apic. Res. 2013, 52, 1–20. [Google Scholar] [CrossRef] [PubMed]
- Bayman, P.; Cotty, P.J. Vegetative compatibility and genetic variation in the Aspergillus flavus population of a single field. Can. J. Bot. 1991, 69, 1707–1711. [Google Scholar] [CrossRef]
- Ehrlich, K.C. Non-aflatoxigenic Aspergillus flavus to prevent aflatoxin contamination in crops: Advantages and limitations. Front. Microbiol. 2014, 5. [Google Scholar] [CrossRef] [PubMed]
- US EPA. Ecological Risk Assessment for Experimental Testing of the Microbial Pesticide End-Use Product. In FourSure. M (A.I.: Aspergillus flavus Strains TCI 6F, TC35C, TC38B and) to Control Aflatoxin-Producing Toxigenic Aspergillus flavus; US EPA: Washington, DC, USA, 2016. [Google Scholar]
- Tabatabai, M.A. Soil enzymes. In Methods of Soil Analysis, Part 2: Microbiological and Biochemical Properties; Weaver, R.W., Angle, S., Bottomley, P., Bezdicek, D., Smith, S., Tabatabai, A., Eds.; SSSA Book Series 5; SSSA: Madison, WI, USA, 1994; pp. 775–833. [Google Scholar]
- Lehman, R.M.; Cambardella, C.A.; Stott, D.; Acosta-Martinez, V.; Manter, D.K.; Buyer, J.; Maul, J.E.; Smith, J.L.; Collins, H.; Halvorson, J.; et al. Understanding and Enhancing Soil Biological Health: The Solution for Reversing Soil Degradation. Sustainability 2015, 7, 988–1027. [Google Scholar] [CrossRef]
- Acosta-Martínez, V.; Zobeck, T.M.; Allen, V. Soil Microbial, Chemical and Physical Properties in Continuous Cotton and Integrated Crop-Livestock Systems. Soil Sci. Soc. Am. J. 2004, 68, 1875–1884. [Google Scholar] [CrossRef]
- Bhandari, K.B.; Longing, S.; West, C.P. Bees Occurring in Corn Production Fields Treated with Atoxigenic Aspergillus flavus (Texas, USA). Agronomy 2020, 10, 571. [Google Scholar] [CrossRef]
- Schutter, M.E.; Dick, R.P. Comparison of Fatty Acid Methyl Ester (FAME) Methods for Characterizing Microbial Communities. Soil Sci. Soc. Am. J. 2000, 64, 1659–1668. [Google Scholar] [CrossRef]
- Bhandari, K.B.; West, C.P.; Acosta-Martinez, V.; Cotton, J.; Cano, A.M. Soil health indicators as affected by diverse forage species and mixtures in semi-arid pastures. Appl. Soil Ecol. 2018, 132, 179–186. [Google Scholar] [CrossRef]
- Acosta-Martínez, V.; Bell, C.; Morris, B.; Zak, J.; Allen, V. Long-term soil microbial community and enzyme activity responses to an integrated cropping-livestock system in a semi-arid region. Agric. Ecosyst. Environ. 2010, 137, 231–240. [Google Scholar] [CrossRef]
- Liebig, M.; Carpenter-Boggs, L.; Johnson, J.; Wright, S.; Barbour, N. Cropping system effects on soil biological characteristics in the Great Plains. Renew. Agric. Food Syst. 2006, 21, 36–48. [Google Scholar] [CrossRef]
- Zelles, L. Phospholipid fatty acid profiles in selected members of soil microbial communities. Chemosphere 1997, 35, 275–294. [Google Scholar] [CrossRef]
- Frostegard, A.; Baath, E. The use of phospholipid fatty acid analysis to estimate bacterial and fungal biomass in soil. Biol. Fert. Soils 1996, 22, 59–65. [Google Scholar] [CrossRef]
- Olsson, P.A.; Bååth, E.; Jakobsen, I.; Söderström, B. The use of phospholipid and neutral lipid fatty acids to estimate biomass of arbuscular mycorrhizal fungi in soil. Mycol. Res. 1995, 99, 623–629. [Google Scholar] [CrossRef]
- Zelles, L. Fatty acid patterns of phospholipids and lipopolysaccharides in the characterization of microbial communities in soil: A review. Biol. Fertil. Soils 1999, 29, 111–129. [Google Scholar] [CrossRef]
- Littell, R.C.; Milliken, G.A.; Stroup, W.W.; Wilfinger, R.D.; Schabenberger, O. SAS for Mixed Models, 2nd ed.; SAS Inst.: Cary, NC, USA, 2006. [Google Scholar]
| Variables | Treatment | County | Mean | ||
|---|---|---|---|---|---|
| San Patricio | Ellis | Grayson | |||
| Soil water (g g−1) | FourSure™ | 0.17 | 0.24 | 0.25 | 0.22 |
| Control | 0.13 | 0.25 | 0.23 | 0.20 | |
| Treatment effect | p = 0.09 | p = 0.75 | p = 0.66 | p = 0.46 | |
| County effect | p < 0.001 | ||||
| Protozoa (nmol g−1) | FourSure™ | 1.1 | 1.7 | 1.6 | 1.5 |
| Control | 1.4 | 1.3 | 1.3 | 1.3 | |
| Treatment effect | p = 0.48 | p = 0.10 | p = 0.53 | p = 0.46 | |
| County effect | p = 0.59 | ||||
| Total FAME (nmol g−1) | FourSure™ | 116 | 192 | 220 | 176 |
| Control | 155 | 161 | 192 | 169 | |
| Treatment effect | p = 0.55 | p = 0.31 | p = 0.64 | p = 0.82 | |
| County effect | p = 0.17 | ||||
| Variables | Treatment | County | Mean | ||
|---|---|---|---|---|---|
| San Patricio | Ellis | Grayson | |||
| (nmol g−1) | |||||
| AMF fungi | FourSure™ | 3.5 | 7.4 | 9.6 | 6.8 |
| Control | 4.4 | 5.5 | 3.6 | 4.5 | |
| Treatment effect | p = 0.50 | p = 0.59 | p = 0.23 | p = 0.13 | |
| County effect | p = 0.30 | ||||
| Saprophytic fungi | FourSure™ | 29.6 | 44.1 | 50.7 | 41.5 |
| Control | 41.7 | 35.5 | 39.3 | 38.8 | |
| Treatment effect | p = 0.58 | p = 0.42 | p = 0.45 | p = 0.73 | |
| County effect | p = 0.61 | ||||
| Total fungi | FourSure™ | 33.2 | 51.5 | 60.3 | 48.3 |
| Control | 46.1 | 40.9 | 42.8 | 43.3 | |
| Treatment effect | p = 0.57 | p = 0.45 | p = 0.38 | p = 0.57 | |
| County effect | p = 0.54 | ||||
| Gram+ bacteria | FourSure™ | 15.1 | 27.9 | 29.2 | 24.1 |
| Control | 20.9 | 24.9 | 29.2 | 25.0 | |
| Treatment effect | p = 0.48 | p = 0.40 | p = 0.99 | p = 0.81 | |
| County effect | p = 0.08 | ||||
| Gram− bacteria | FourSure™ | 6.5 | 10.2 | 12.8 | 9.8 |
| Control | 6.7 | 8.6 | 9.5 | 8.3 | |
| Treatment effect | p = 0.82 | p = 0.34 | p = 0.17 | p = 0.14 | |
| County effect | p < 0.01 | ||||
| Actinobacteria | FourSure™ | 11.3 | 21.0 | 22.5 | 18.3 |
| Control | 13.9 | 19.6 | 18.7 | 17.4 | |
| Treatment effect | p = 0.49 | p = 0.69 | p = 0.36 | p = 0.72 | |
| County effect | p = 0.047 | ||||
| Total bacteria | FourSure™ | 32.9 | 59.1 | 64.5 | 52.2 |
| Control | 41.5 | 53.1 | 57.4 | 50.7 | |
| Treatment effect | p = 0.50 | p = 0.49 | p = 0.54 | p = 0.84 | |
| County effect | p = 0.047 | ||||
| Fungi–bacteria ratio | FourSure™ | 1.0 | 0.9 | 0.9 | 0.9 |
| Control | 1.1 | 0.8 | 0.8 | 0.9 | |
| Treatment effect | p = 0.81 | p = 0.46 | p = 0.19 | p = 0.28 | |
| County effect | p = 0.05 | ||||
| Total FAMEs † | AMF † | Saprophytic Fungi | Total Fungi | Gram+ Bacteria | Gram−Bacteria | Actino-Bacteria | Total Bacteria | |
|---|---|---|---|---|---|---|---|---|
| Soil Water Content | 0.32 | 0.33 | 0.22 | 0.25 | 0.41 | 0.48 | 0.49 | 0.46 |
| p = 0.20 | p = 0.18 | p = 0.38 | p = 0.31 | p = 0.09 | p = 0.04 | p = 0.04 | p = 0.05 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bhandari, K.B.; Longing, S.D.; West, C.P. Soil Microbial Communities in Corn Fields Treated with Atoxigenic Aspergillus flavus. Soil Syst. 2020, 4, 35. https://doi.org/10.3390/soilsystems4020035
Bhandari KB, Longing SD, West CP. Soil Microbial Communities in Corn Fields Treated with Atoxigenic Aspergillus flavus. Soil Systems. 2020; 4(2):35. https://doi.org/10.3390/soilsystems4020035
Chicago/Turabian StyleBhandari, Krishna B., Scott D. Longing, and Charles P. West. 2020. "Soil Microbial Communities in Corn Fields Treated with Atoxigenic Aspergillus flavus" Soil Systems 4, no. 2: 35. https://doi.org/10.3390/soilsystems4020035
APA StyleBhandari, K. B., Longing, S. D., & West, C. P. (2020). Soil Microbial Communities in Corn Fields Treated with Atoxigenic Aspergillus flavus. Soil Systems, 4(2), 35. https://doi.org/10.3390/soilsystems4020035





