SARS-CoV-2 Detection for Diagnosis Purposes in the Setting of a Molecular Biology Research Lab
Abstract
:1. Introduction
1.1. Overview of the Technique
1.2. Applications and Target Audience
1.3. Advantages
1.4. Comparison with Other Methods
1.5. Expertise Needed to Implement the Protocol and Limitations
2. Materials
2.1. Sample Reception, Deconditioning and Labeling (Pre-Analytical Step)
2.1.1. Reagents
2.1.2. Equipment
2.1.3. Human Resources
2.2. Inactivation of Viral Infectivity (Pre-Analytical Step)
2.2.1. Reagents
- Guanidinium thiocyanate solution (TRI Reagent from Sigma-Aldrich or TRIzol from Invitrogen or QIAzol from QIAGEN). CAUTION: Vapors are toxic and the solution must be handled under a chemical hood.
- Internal Control (IC):
- 3.
- Umonium Medical Spray (Huckert’s International).
2.2.2. Equipment
2.2.3. Human Resources
2.3. RNA Extraction (Analytical Step)
2.3.1. Reagents
- Chloroform (Sigma-Aldrich). CAUTION: Vapors are toxic and the solution must be handled under a chemical hood.
- GlycoBlue (Thermofisher Scientifc).Alternatively, GlycoBlue can be substituted by Glycogen, RNA grade (Thermo scientific) to serve as RNA carrier according to the manufacturer instructions.
- Isopropanol (Sigma-Aldrich).
- Ethanol 75% (Sigma-Aldrich). In a falcon 50 mL tube, add 12.5 mL of RNAse free water to 37.5 mL of ethanol 96–100% (Sigma-Aldrich).
- RNAse free water.
2.3.2. Equipment
- Vortex apparatus.
- Four to eight high-speed 1.5 mL tube refrigerated centrifuges.
2.3.3. Human Resources
2.4. Taqman RT-qPCR for SARS-CoV-2 (Analytical Step)
2.4.1. Reagents
- 5X Master Mix containing DNA polymerase, MgCl2 (5.5 mM final concentration) and dNTPs (Eurogentec Takyon One-Step No Rox Probe 5X MasterMix dTTP). Important note: Use a low ROX or high ROX PCR Master Mix if your PCR machine requires ROX normalization.
- Euroscript II RT (50 u/µL) and RNAse inhibitor (20 u/µL) (Euroscript II Reverse Transcriptase/RNAse inhibitor provided in the Eurogentec Takyon One-Step No Rox Probe 5X MasterMix dTTP).
- Additive (provided in the Eurogentec Takyon One-Step No Rox Probe 5X MasterMix dTTP).
- RNase free water.
- Primers and Probe Mixes:
- E_Sarbeco_Fw: 5′- ACAGGTACGTTAATAGTTAATAGCGT-3′
- E_Sarbeco_Rev: 5′- ATATTGCAGCAGTACGCACACA-3′
- E_Sarbeco_Probe: 5′-(FAM)ACACTAGCCATCCTTACTGCGCTTCG(BHQ1)-3′
- IC_Fw: 5′-TTGCCGTTTGATTTTGAAGTTGTG-3′
- IC_Rev: 5′-TCAGGGATCGCAAATTAAAGAACC-3′
- IC_Probe: 5′-(FAM)TCATCCGTGCTGACCCTCTGCGAG(BHQ1)-3′
- Primers and Probe Mix for SARS-CoV-2
- 10 µL of E_Sarbeco_Probe (100 µM)
- +20 µL of E_Sarbeco_Fw (100 µM)
- +20 µL of E_Sarbeco_Rev (100 µM)
- +950 µL of RNAse free water
- +10 µL of IC_Probe (100 µM)
- +20 µL of IC_Fw (100 µM)
- +20 µL of IC_Rev (100 µM)
- +950 µL of RNAse free water
- 6.
- Positive Controls (PC):
- 7.
- Internal Control (IC), as described in Section 2.2. Inactivation of Viral Infectivity.
- 8.
- Negative Control (NC):
2.4.2. Equipment
2.4.3. Human Resources
2.5. Data Analysis and Validation (Post-Analytical Step)
Human Resources
3. Procedure
3.1. Sample Reception, Deconditioning and Labelling (Pre-Analytical Step)
- The 1st label is stuck to the original sample;
- The 2nd label is stuck to the sample’s sheet (medical prescription form);
- The 3rd label is stuck to the first 1.5 mL tube for inactivation of viral infectivity;
- The 4th label is stuck to the second 1.5 mL tube for RNA precipitation and resuspension.
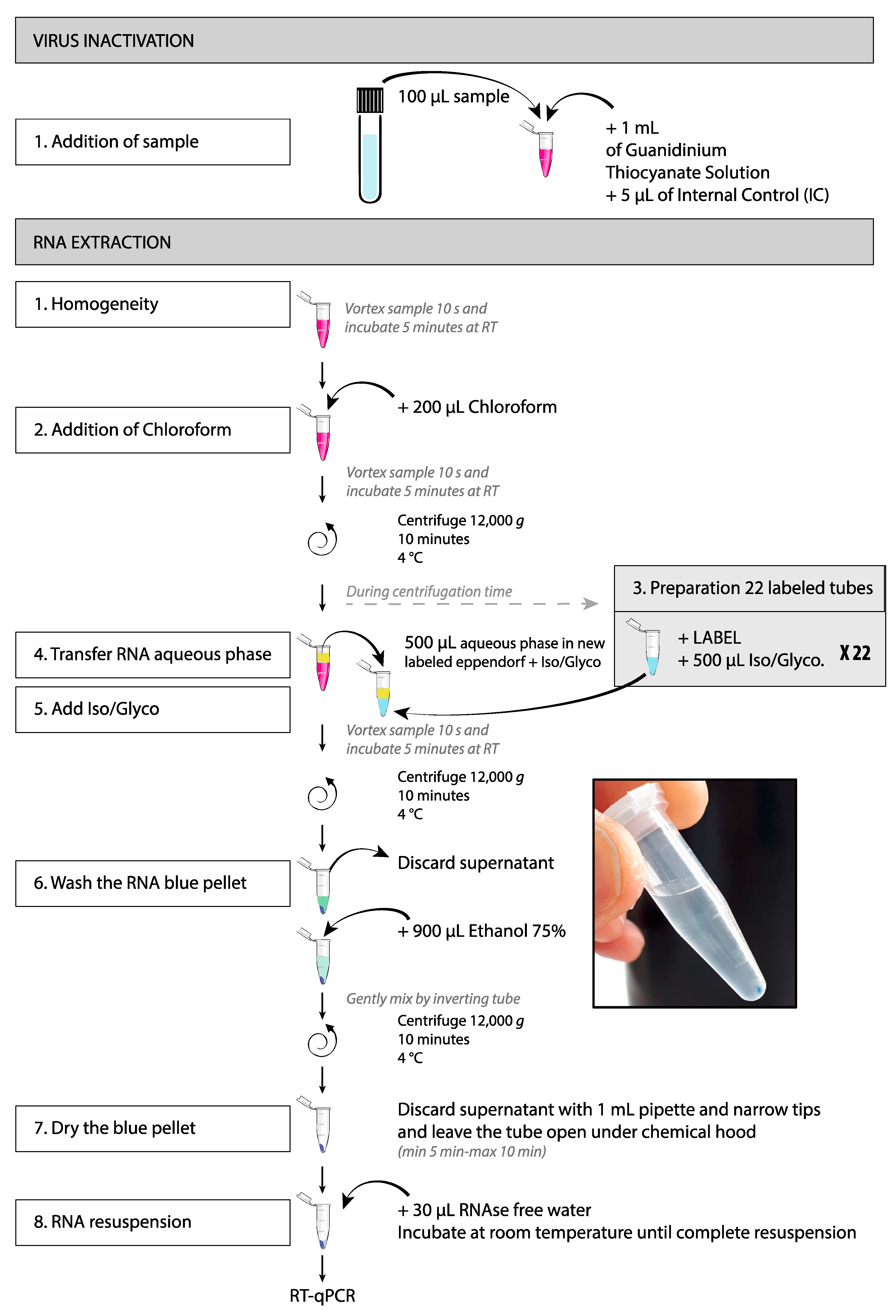
3.2. Inactivation of Viral Infectivity (Pre-Analytical Step)
- Vortex nasopharyngeal swabs, aspirates or BAL (BronchoAlveolar Lavage) fluids briefly;
- Spin the clinical specimen collection tube at 200 g for 1 min to make sure the sample (transport media inside the clinical sample collection tube) is at the bottom of the tube. Use a centrifuge with sealed buckets. Clean the centrifuge with Umonium Medical Spray or equivalent antiviral disinfectant;
- Under a BSL2 hood, homogenize the sample by up and down pipetting and transfer 100 µL of the sample in the corresponding labeled 1.5 mL tube containing 1 mL of guanidinium thiocyanate solution (Trizol or equivalent) supplemented with 5 µL of Internal Control. Mix by inverting the tube. This procedure instantly inactivates viral infectivity;
- The 1.5 mL tubes containing 1 mL of guanidinium thiocyanate solution supplemented with 5 µL of Internal Control must be prepared ahead under a chemical hood (vapors of guanidinium thiocyanate solution are toxic). These aliquots can be stored at −80 °C if necessary;
- Before the samples can get out of the biosafety cabinet, spray the tubes with quaternary ammonium solution (Umonium Medical Spray or equivalent). Wipe the tubes;
- Samples are now handled without any biosafety issue. Organize the inactivated samples by a series of 21 samples +1 tube called Extraction Control or EC. The EC tube contains 1 mL of guanidinium thiocyanate solution supplemented with 100 µL of PBS and 5 µL of Internal Control ONLY, i.e., without clinical sample. RNA extraction must be performed by a batch of 22 tubes (21 samples +1 EC);
- If necessary, samples can be stored at −80 °C for further processing. Because we usually receive the clinical specimens by the afternoon, the deconditioning and viral inactivation steps occur at day zero and the inactivated samples are stored at −80 °C overnight. RNA extraction, RT-qPCR, and data validation and results communication occur at day +1.
3.3. RNA Extraction (Analytical Step)
- Vortex the samples 10 s and incubate 5 min at room temperature;
- Add 200 µL of chloroform and vortex for 10 s CAUTION: Vapors are toxic and the solution must be handled under a chemical hood;
- Vortex the samples 10 s and incubate 5 min at room temperature;
- Centrifuge at 12,000 g for 10 min at 4 °C;
- Transfer 500 µL of the colorless upper aqueous phase (containing the RNA) in the second 1.5 mL tube, avoid contact with the ring or the lower organic phase (pink).
- 6.
- Add 500 µL of the mix isopropanol-GlycoBlue;
- 7.
- Vortex for 10 s and incubate 5 min at room temperature;
- 8.
- Centrifuge at 12,000 to 14,000 g for 10 min at 4 °C.
- 9.
- Discard the supernatant;
- 10.
- Add 900 µL of 75% ethanol;
- 11.
- Gently mix by inverting the tubes;
- 12.
- Centrifuge at 12,000 to 14,000 g for 10 min at 4 °C;
- 13.
- Aspirate slowly the supernatant with a 1 mL pipette by avoiding contact with the blue pellet (slow pipetting allows the ethanol to drain along the tube wall);
- 14.
- Use a narrow tip (e.g., gel loading tip) to remove residual ethanol;
- 15.
- To dry the pellet, leave the tube open under the chemical hood until complete ethanol evaporation. This step lasts about 5 min. Do not over-dry the RNA by letting the sample dries more than 10 min;
- 16.
- Resuspend the pellet in 30 µL of RNAse free water;
- 17.
- Incubate at room temperature until complete resuspension of the blue pellet;
- 18.
- If necessary, RNA can be stored at −80 °C for further processing.
3.4. Taqman RT-qPCR for SARS-CoV-2 (Analytical Step)
- S stands for Sample, 42 samples per plate (2 batches of 21 samples);
- NC for Negative Control (water);
- PC1 for Positive Control 1, PC2 for Positive Control 2 (RNA from SARS-CoV-2-infected cells);
- EC1 for Extraction Control of the first extraction batch, EC2 for Extraction Control of the second extraction batch.
- A hard copy of the PCR plate is kept for the validation process (Figure 3).
- First deposit 16 µL of the PCR Mix per well;
- Next, add 4 µL of sample per well (or 4 µL of water for NC, or 4 µL of PC, or 4 µL of EC);
- Stick the adhesive film;
- Spin the PCR plate to collect the PCR mix at the bottom of the well, 200 g, 1 min.
- 48 °C 10 min;
- 95 °C 3 min;
- 45 cycles: 95 °C 15 s, 58 °C 30 s.
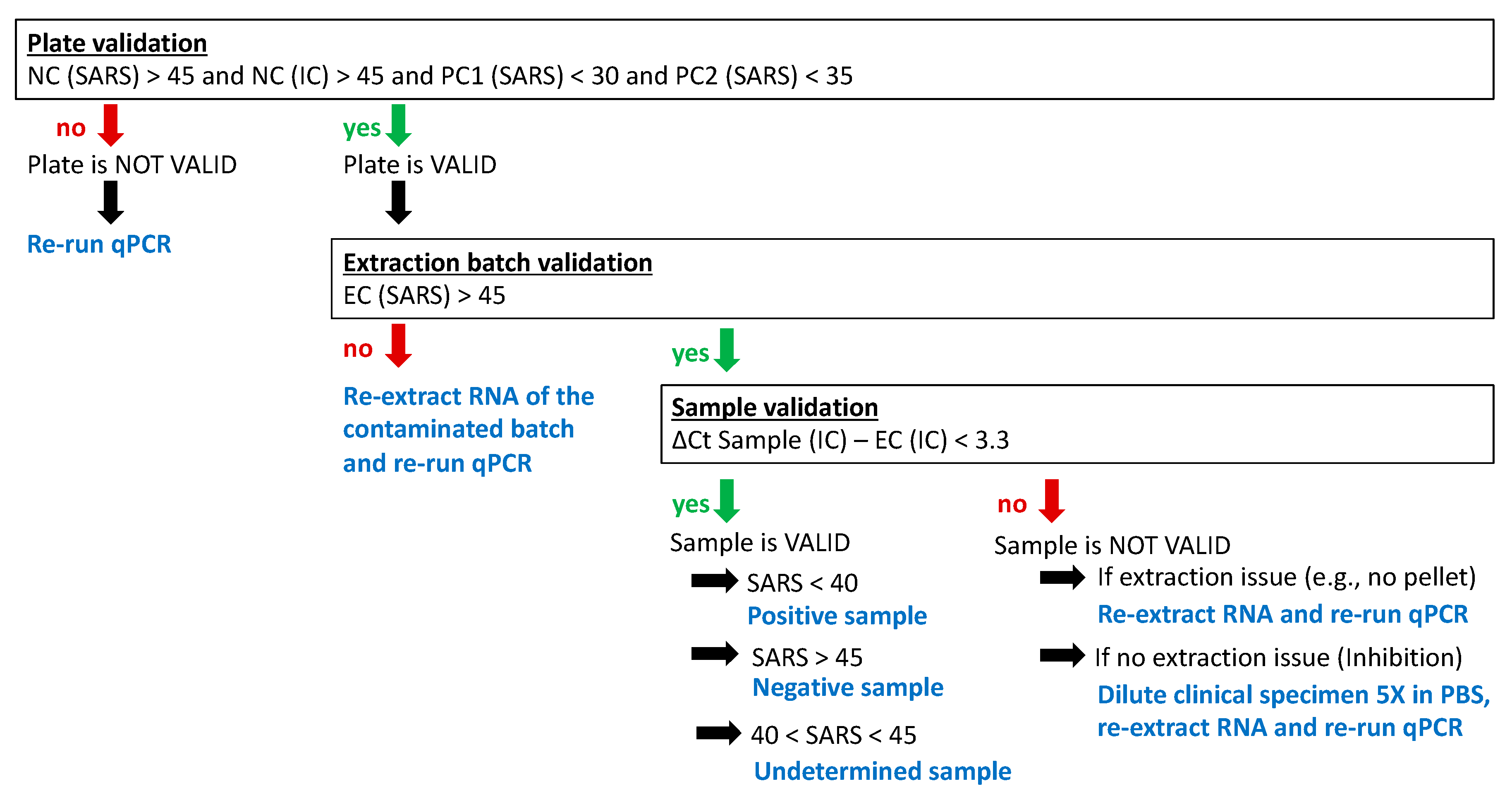
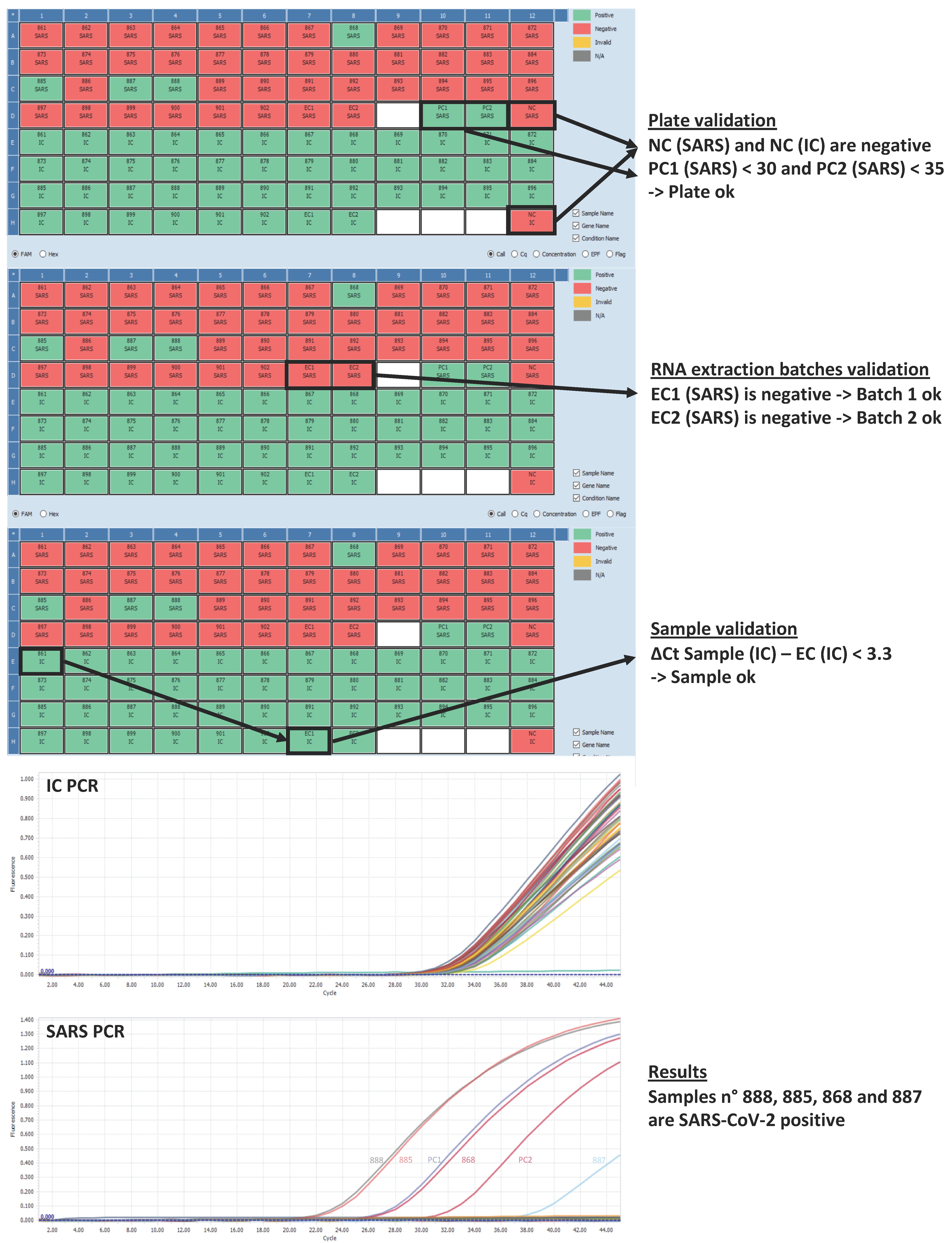
3.5. Data Analysis and Validation (Post-Analytical Step)
- We consider the plate as VALID only if Ct values for
- NC (SARS) > 45 and NC (IC) > 45
- and PC1 (SARS) < 30 and PC2 (SARS) < 35.
- Otherwise, the plate must be re-run.
4. Timing
5. Anticipated Results
6. Notes
- We do not recommend thermal inactivation of the clinical sample prior to mixing with the solution of guanidinium thiocyanate, as it might lead to false-negative results [4].
- The present protocol has been tested in dual-color using a FAM probe for the SARS-CoV-2 PCR and a HEX or Cy5 probe for the Internal Control PCR. Unfortunately, the results were not satisfactory, with a strong inhibition of the IC PCR. Therefore, the two PCRs (SARS-CoV-2 and IC) must be done in two separate wells to perform as expected.
- The primers and Taqman probe targeting the E gene fragment [2] have been preferred to those targeting the RdRp gene. We found the PCR on the E gene to be more sensitive with earlier Ct values on several clinical specimens (data not shown). This might be explained partly by the higher abundance of subgenomic RNAs compared to the genomic ones. The primers and probe binding sequences are conserved among the SARS-related CoV [2]. However, a point mutation on the sequence recognized by the E_Sarbeco_Probe (position 26,340) has been recently reported in some viral isolates, impeding detection [5]. It will be critical to follow the evolution of such strains (https://www.gisaid.org/) and, if necessary, to adapt the protocol using a degenerated probe or targeting a second viral region.
- In principle, the RNA spike-in used as internal control can be substituted by any known non-human exogenous single stranded RNA.
- Another alternative to the RNA spike-in control is the titration of house-keeping gene RNA (e.g., Glyceraldehyde-3-Phosphate Dehydrogenase GAPDH mRNA). The advantage of this approach is that it allows viral load comparison between samples. The disadvantage is that it cannot directly identify the samples whose RT-PCR is inhibited and is largely impacted by the type of sample and the sampling technique. Normalization using GAPDH mRNA has been reported for SARS-CoV-1 quantification [6] using the following primers and probes:
- GAPDH-Fw 5′-GAAGGTGAAGGTCGGAGT-3′
- GAPDH-Rv 5′-GAAGATGGTGATGGGATTTC-3′
- GAPDH-Probe 5′-(FAM)CAAGCTTCCCGTTCTCAGCC(BHQ1)-3′
- Because some samples must be redone and others must be fast-tracked, we dedicated an experienced researcher (the trouble-shooter) for that work. They take care of the whole process independently of the other workers to deliver the results in time.
- Potentially contaminated material (disposable gloves, gown, pipette tips, absorbing paper, FFP2 masks) are autoclaved at 121 °C for 20 min and then incinerated (double-neutralization of highly contagious material).
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bai, Y.; Yao, L.; Wei, T.; Tian, F.; Jin, D.Y.; Chen, L.; Wang, M. Presumed Asymptomatic Carrier Transmission of COVID-19. J. Am. Med. Assoc. 2020, 323, 1406–1407. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Corman, V.M.; Landt, O.; Kaiser, M.; Molenkamp, R.; Meijer, A.; Chu, D.K.; Bleicker, T.; Brünink, S.; Schneider, J.; Schmidt, M.L. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Eurosurveillance 2020, 25, 2000045. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fischer, M.; Schirrmeier, H.; Wernike, K.; Wegelt, A.; Beer, M.; Hoffmann, B. Development of a pan-Simbu real-time reverse transcriptase PCR for the detection of Simbu serogroup viruses and comparison with SBV diagnostic PCR systems. Virol. J. 2013, 10, 327. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pan, Y.; Long, L.; Zhang, D.; Yuan, T.; Cui, S.; Yang, P.; Wang, Q.; Ren, S. Potential False-Negative Nucleic Acid Testing Results for Severe Acute Respiratory Syndrome Coronavirus 2 from Thermal Inactivation of Samples with Low Viral Loads. Clin. Chem. 2020, 66, 794–801. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Artesi, M.; Long, L.; Zhang, D.; Yuan, T.; Cui, S.; Yang, P.; Wang, Q.; Ren, S. A recurrent mutation at position 26,340 of SARS-CoV-2 is associated with failure of the E-gene qRT-PCR utilized in a commercial dual-target diagnostic assay. J. Clin. Microbiol. 2020, 66, 794–801. [Google Scholar] [CrossRef] [PubMed]
- Wong, S.C.C.; Chan, J.K.C.; Lee, K.C.; Lo, E.S.F.; Tsang, D.N.C. Development of a quantitative assay for SARS coronavirus and correlation of GAPDH mRNA with SARS coronavirus in clinical specimens. J. Clin. Pathol. 2005, 58, 276–280. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kozera, B.; Rapacz, M. Reference genes in real-time PCR. J. Appl. Genet. 2013, 54, 391–406. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Radonić, A.; Thulke, S.; Mackay, I.M.; Landt, O.; Siegert, W.; Nitsche, A. Guideline to reference gene selection for quantitative real-time PCR. Biochem. Biophys. Res. Commun. 2004, 313, 856–862. [Google Scholar] [CrossRef] [PubMed]
- Radonić, A.; Thulke, S.; Bae, H.G.; Müller, M.A.; Siegert, W.; Nitsche, A. Reference gene selection for quantitative real-time PCR analysis in virus infected cells: SARS corona virus, Yellow fever virus, Human Herpesvirus-6, Camelpox virus and Cytomegalovirus infections. Virol. J. 2005, 2, 7. [Google Scholar] [CrossRef] [PubMed] [Green Version]



| Tracking Form | ||||||||
|---|---|---|---|---|---|---|---|---|
| Samples processed: | Date of clinical specimens’ reception: | |||||||
| UNPACKING and LABELING | Logistician: | |||||||
| INACTIVATION | Date: | Researcher: | ||||||
| Comments: | ||||||||
| EXTRACTION | Date: | Researcher: | ||||||
| chemical hood used n° | hood cleaning: | |||||||
| Comments: | ||||||||
| RT-qPCR | Date: | Researcher: | ||||||
| PCR bench n° | bench cleaning: | |||||||
| PCR machine n° | ||||||||
| Comments | ||||||||
| DATA VALIDATION | Date: | Researchers: | ||||||
| Comments | ||||||||
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | ||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| SARS | A | S1 | S2 | S3 | S4 | S5 | S6 | S7 | S8 | S9 | S10 | S11 | S12 |
| B | S13 | S14 | S15 | S16 | S17 | S18 | S19 | S20 | S21 | S22 | S23 | S24 | |
| C | S25 | S26 | S27 | S28 | S29 | S30 | S31 | S32 | S33 | S34 | S35 | S36 | |
| D | S37 | S38 | S39 | S40 | S41 | S42 | EC1 | EC2 | PC1 | PC2 | NC water | ||
| IC | E | S1 | S2 | S3 | S4 | S5 | S6 | S7 | S8 | S9 | S10 | S11 | S12 |
| F | S13 | S14 | S15 | S16 | S17 | S18 | S19 | S20 | S21 | S22 | S23 | S24 | |
| G | S25 | S26 | S27 | S28 | S29 | S30 | S31 | S32 | S33 | S34 | S35 | S36 | |
| H | S37 | S38 | S39 | S40 | S41 | S42 | EC1 | EC2 | NC water |
| Volume in µL | Per Reaction | 1 Plate | 2 Plates | 3 Plates | 4 Plates | 5 Plates |
|---|---|---|---|---|---|---|
| 5X Master Mix | 4 | 200 | 400 | 600 | 800 | 1000 |
| Euroscript II (RT) and RNAse inhibitor | 0.2 | 10 | 20 | 30 | 40 | 50 |
| Primers and Probes Mix SARS-CoV-2 | 4 | 200 | 400 | 600 | 800 | 1000 |
| RT Additive | 0.2 | 10 | 20 | 30 | 40 | 50 |
| RNAse free water | 7.6 | 380 | 760 | 1140 | 1520 | 1900 |
| Sample or PC or NC or EC | 4 |
| Volume in µL | Per Reaction | 1 Plate | 2 Plates | 3 Plates | 4 Plates | 5 Plates |
|---|---|---|---|---|---|---|
| 5X Master Mix | 4 | 200 | 400 | 600 | 800 | 1000 |
| Euroscript II (RT) and RNAse inhibitor | 0.2 | 10 | 20 | 30 | 40 | 50 |
| Primers and Probes Mix IC (SBV) | 4 | 200 | 400 | 600 | 800 | 1000 |
| RT Additive | 0.2 | 10 | 20 | 30 | 40 | 50 |
| RNAse free water | 7.6 | 380 | 760 | 1140 | 1520 | 1900 |
| Sample or PC or NC or EC | 4 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Coupeau, D.; Burton, N.; Lejeune, N.; Loret, S.; Petit, A.; Pejakovic, S.; Poulain, F.; Bonil, L.; Trozzi, G.; Wiggers, L.; et al. SARS-CoV-2 Detection for Diagnosis Purposes in the Setting of a Molecular Biology Research Lab. Methods Protoc. 2020, 3, 59. https://doi.org/10.3390/mps3030059
Coupeau D, Burton N, Lejeune N, Loret S, Petit A, Pejakovic S, Poulain F, Bonil L, Trozzi G, Wiggers L, et al. SARS-CoV-2 Detection for Diagnosis Purposes in the Setting of a Molecular Biology Research Lab. Methods and Protocols. 2020; 3(3):59. https://doi.org/10.3390/mps3030059
Chicago/Turabian StyleCoupeau, Damien, Nicolas Burton, Noémie Lejeune, Suzanne Loret, Astrid Petit, Srdan Pejakovic, Florian Poulain, Laura Bonil, Gabrielle Trozzi, Laetitia Wiggers, and et al. 2020. "SARS-CoV-2 Detection for Diagnosis Purposes in the Setting of a Molecular Biology Research Lab" Methods and Protocols 3, no. 3: 59. https://doi.org/10.3390/mps3030059
APA StyleCoupeau, D., Burton, N., Lejeune, N., Loret, S., Petit, A., Pejakovic, S., Poulain, F., Bonil, L., Trozzi, G., Wiggers, L., Willemart, K., André, E., Laenen, L., Cuypers, L., Van Ranst, M., Bogaerts, P., Muylkens, B., & Gillet, N. A. (2020). SARS-CoV-2 Detection for Diagnosis Purposes in the Setting of a Molecular Biology Research Lab. Methods and Protocols, 3(3), 59. https://doi.org/10.3390/mps3030059





