The Key Lnc (RNA)s in Cardiac and Skeletal Muscle Development, Regeneration, and Disease
Abstract
:1. Introduction
2. Long Non-Coding RNA in Cardiac Muscle
2.1. Cardiac Development
2.2. Cardiac Muscle Regeneration
2.3. Cardiac Disease
2.3.1. Cardiac Hypertrophy
2.3.2. Myocardial Infarction
3. Long Non-Coding RNA in Skeletal Muscle
3.1. Skeletal Muscle Development
3.2. Skeletal Muscle Differentiation and Regeneration
3.2.1. Positive Regulators
3.2.2. Negative Regulators
3.3. Skeletal Muscle Disease
3.4. LncRNAs That Function as ncRNAs and Micropeptides
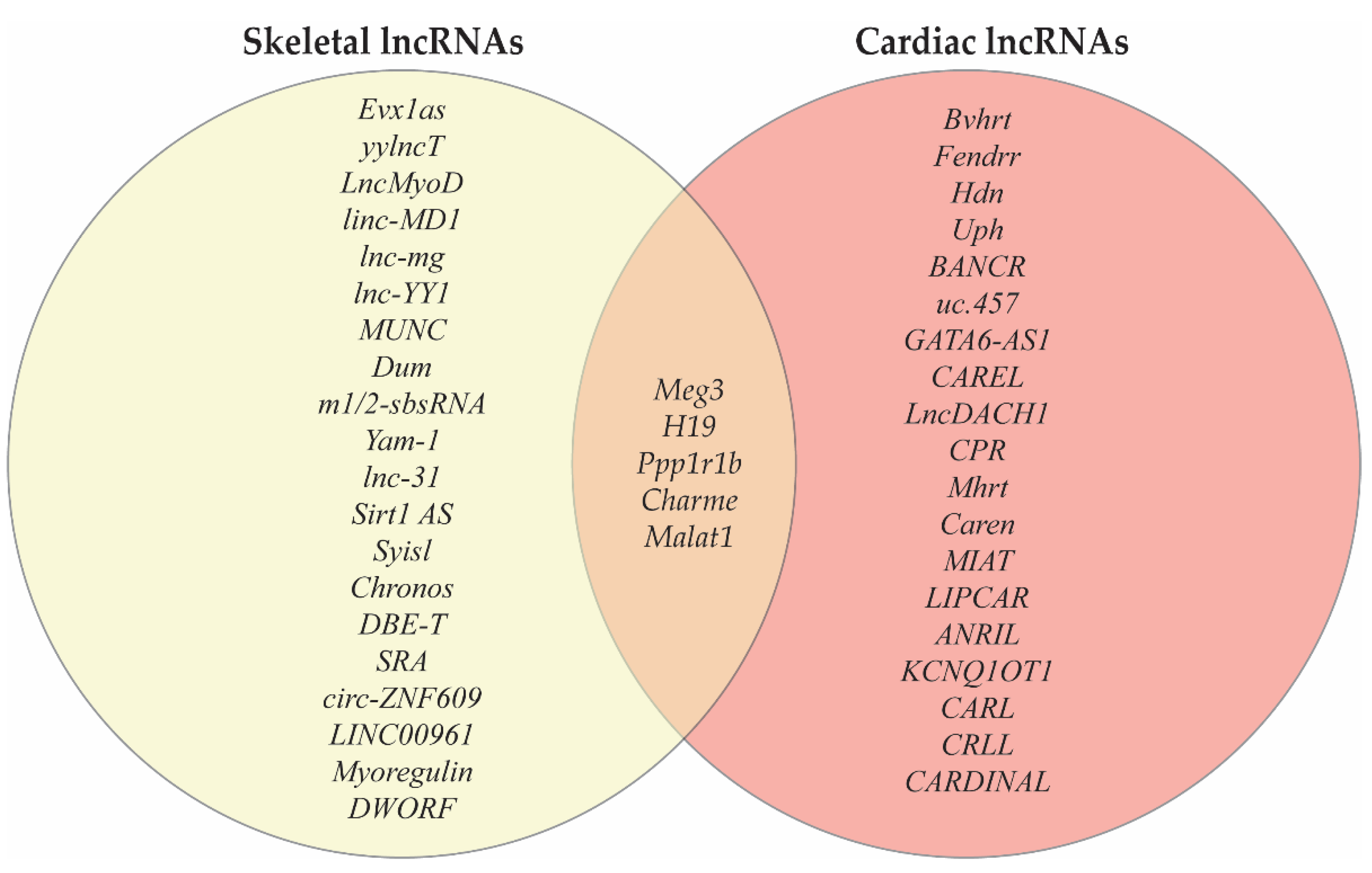
4. LncRNAs Co-Expressed in Cardiac and Skeletal Muscle
5. LncRNAs and Genomic Imprinting
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Howe, K.L.; Achuthan, P.; Allen, J.; Allen, J.; Alvarez-Jarreta, J.; Amode, M.R.; Armean, I.M.; Azov, A.G.; Bennett, R.; Bhai, J.; et al. Ensembl 2021. Nucleic Acids Res. 2021, 49. [Google Scholar] [CrossRef]
- International Human Genome Sequencing Consortium Initial Sequencing and Analysis of the Human Genome. Nature 2001, 409, 860–921. [CrossRef] [Green Version]
- Panni, S.; Lovering, R.C.; Porras, P.; Orchard, S. Non-Coding RNA Regulatory Networks. Biochim. Biophys. Acta Gene Regul. Mech. 2020, 1863, 194417. [Google Scholar] [CrossRef] [PubMed]
- Esteller, M. Non-Coding RNAs in Human Disease. Nat. Rev. Genet. 2011, 12, 861–874. [Google Scholar] [CrossRef]
- Martone, J.; Mariani, D.; Desideri, F.; Ballarino, M. Non-Coding RNAs Shaping Muscle. Front. Cell Dev. Biol. 2020, 7, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Peschansky, V.J.; Wahlestedt, C. Non-Coding RNAs as Direct and Indirect Modulators of Epigenetic Regulation. Epigenetics 2014, 3, 3–12. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, P.; Wu, W.; Chen, Q.; Chen, M. Non-Coding RNAs and Their Integrated Networks. J. Integr. Bioinform. 2019, 16, 1–12. [Google Scholar] [CrossRef]
- O’Brien, J.; Hayder, H.; Zayed, Y.; Peng, C. Overview of MicroRNA Biogenesis, Mechanisms of Actions, and Circulation. Front. Endocrinol. 2018, 9, 402. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dinger, M.E.; Amaral, P.P.; Mercer, T.R.; Pang, K.C.; Bruce, S.J.; Gardiner, B.B.; Askarian-Amiri, M.E.; Ru, K.; Solda, G.; Simons, C.; et al. Long Noncoding RNAs in Mouse Embryonic Stem Cell Pluripotency and Differentiation. Genome Res. 2008, 18, 1433–1445. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, M.; Kraus, W.L. From Discovery to Function: The Expanding Roles of Long NonCoding RNAs in Physiology and Disease. Endocr. Rev. 2015, 36, 25–64. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hon, C.-C.; Ramilowski, J.A.; Harshbarger, J.; Bertin, N.; Rackham, O.J.L.; Gough, J.; Denisenko, E.; Schmeier, S.; Poulsen, T.M.; Severin, J.; et al. An Atlas of Human Long Non-Coding RNAs with Accurate 5′ Ends. Nature 2017, 543, 199–204. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Iyer, M.K.; Niknafs, Y.S.; Malik, R.; Singhal, U.; Sahu, A.; Hosono, Y.; Barrette, T.R.; Prensner, J.R.; Evans, J.R.; Zhao, S.; et al. The Landscape of Long Noncoding RNAs in the Human Transcriptome. Nat. Genet. 2015, 47, 199–208. [Google Scholar] [CrossRef]
- Slavoff, S.A.; Mitchell, A.J.; Schwaid, A.G.; Cabili, M.N.; Ma, J.; Levin, J.Z.; Karger, A.D.; Budnik, B.A.; Rinn, J.L.; Saghatelian, A. Peptidomic Discovery of Short Open Reading Frame-Encoded Peptides in Human Cells. Nat. Chem. Biol. 2013, 9, 59–64. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Anderson, D.M.; Anderson, K.M.; Chang, C.L.; Makarewich, C.A.; Nelson, B.R.; McAnally, J.R.; Kasaragod, P.; Shelton, J.M.; Liou, J.; Bassel-Duby, R.; et al. A Micropeptide Encoded by a Putative Long Noncoding RNA Regulates Muscle Performance. Cell 2015, 160, 595–606. [Google Scholar] [CrossRef] [Green Version]
- Rion, N.; Rüegg, M.A. LncRNA-Encoded Peptides: More than Translational Noise? Cell Res. 2017, 27, 604–605. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, L.; Fan, J.; Han, L.; Qi, H.; Wang, Y.; Wang, H.; Chen, S.; Du, L.; Li, S.; Zhang, Y.; et al. The Micropeptide LEMP Plays an Evolutionarily Conserved Role in Myogenesis. Cell Death Dis. 2020, 11. [Google Scholar] [CrossRef]
- McMullen, J.R.; Drew, B.G. Long Non-Coding RNAs (LncRNAs) in Skeletal and Cardiac Muscle: Potential Therapeutic and Diagnostic Targets? Clin. Sci. 2016, 130, 2245–2256. [Google Scholar] [CrossRef] [PubMed]
- Devaux, Y.; Zangrando, J.; Schroen, B.; Creemers, E.E.; Pedrazzini, T.; Chang, C.P.; Dorn, G.W.; Thum, T.; Heymans, S. Long Noncoding RNAs in Cardiac Development and Ageing. Nat. Rev. Cardiol. 2015, 12, 415–425. [Google Scholar]
- Zhou, H.; Wang, B.; Yang, Y.-X.; Jia, Q.-J.; Zhang, A.; Qi, Z.-W.; Zhang, J.-P. Long Noncoding RNAs in Pathological Cardiac Remodeling: A Review of the Update Literature. BioMed Res. Int. 2019, 2019, 7159592. [Google Scholar] [CrossRef] [PubMed]
- Chaumeil, J.; Le Baccon, P.; Wutz, A.; Heard, E. A Novel Role for Xist RNA in the Formation of a Repressive Nuclear Compartment into Which Genes Are Recruited When Silenced. Genes Dev. 2006, 20, 2223–2237. [Google Scholar] [CrossRef] [Green Version]
- Rinn, J.L.; Kertesz, M.; Wang, J.K.; Squazzo, S.L.; Xu, X.; Brugmann, S.A.; Goodnough, L.H.; Helms, J.A.; Farnham, P.J.; Segal, E.; et al. Functional Demarcation of Active and Silent Chromatin Domains in Human HOX Loci by Noncoding RNAs. Cell 2007, 129, 1311–1323. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kazimierczyk, M.; Kasprowicz, M.K.; Kasprzyk, M.E.; Wrzesinski, J. Human Long Noncoding RNA Interactome: Detection, Characterization and Function. Int. J. Mol. Sci. 2020, 21, 1027. [Google Scholar] [CrossRef] [Green Version]
- Moore, I.V.; Joseph, B.; Uchida, S. Functional Characterization of Long Non-Coding RNAs Joseph. Curr. Opin. Cardiol. 2020, 35, 199–206. [Google Scholar] [CrossRef] [PubMed]
- Sohi, G.; Dilworth, F.J. Noncoding RNAs as Epigenetic Mediators of Skeletal Muscle Regeneration. FEBS J. 2015, 282, 1630–1646. [Google Scholar] [CrossRef] [Green Version]
- Ulitsky, I.; Shkumatava, A.; Jan, C.H.; Sive, H.; Bartel, D.P. Conserved Function of LincRNAs in Vertebrate Embryonic Development despite Rapid Sequence Evolution. Cell 2011, 147, 1537–1550. [Google Scholar] [CrossRef] [Green Version]
- Sarropoulos, I.; Marin, R.; Cardoso-Moreira, M.; Kaessmann, H. Developmental Dynamics of LncRNAs across Mammalian Organs and Species. Nature 2019, 571, 510–514. [Google Scholar] [CrossRef] [PubMed]
- Wojciechowska, A.; Braniewska, A.; Kozar-Kamińska, K. MicroRNA in Cardiovascular Biology and Disease. Adv. Clin. Exp. Med. Off. Organ Wroclaw Med. Univ. 2017, 26, 865–874. [Google Scholar] [CrossRef] [Green Version]
- Zaffran, S.; Frasch, M. Early Signals in Cardiac Development. Circ. Res. 2002, 91, 457–469. [Google Scholar] [CrossRef] [Green Version]
- Paige, S.L.; Plonowska, K.; Xu, A.; Wu, S.M. Molecular Regulation of Cardiomyocyte Differentiation. Circ. Res. 2015, 116, 341–353. [Google Scholar] [CrossRef] [Green Version]
- Doevendans, P.A.; Van Bilsen, M. Transcription Factors and the Cardiac Gene Programme. Int. J. Biochem. Cell Biol. 1996, 28, 387–403. [Google Scholar] [CrossRef]
- García-Padilla, C.; Domínguez, J.N.; Aránega, A.E.; Franco, D. Differential Chamber-Specific Expression and Regulation of Long Non-Coding RNAs during Cardiac Development. Biochim. Biophys. Acta BBA Gene Regul. Mech. 2019, 1862, 194435. [Google Scholar] [CrossRef]
- Saga, Y.; Hata, N.; Kobayashi, S.; Magnuson, T.; Seldin, M.F.; Taketo, M.M. MesP1: A Novel Basic Helix-Loop-Helix Protein Expressed in the Nascent Mesodermal Cells during Mouse Gastrulation. Development 1996, 122, 2769–2778. [Google Scholar] [CrossRef]
- Lescroart, F.; Wang, X.; Lin, X.; Swedlund, B.; Gargouri, S.; Sànchez-Dànes, A.; Moignard, V.; Dubois, C.; Paulissen, C.; Kinston, S.; et al. Defining the Earliest Step of Cardiovascular Lineage Segregation by Single-Cell RNA-Seq. Science 2018, 359, 1177–1181. [Google Scholar] [CrossRef] [Green Version]
- Klattenhoff, C.; Scheuermann, J.C.; Surface, L.E.; Robert, K.; Fields, P.A.; Steinhauser, M.L.; Ding, H.; Butty, V.L.; Torrey, L.; Haas, S.; et al. Braveheart, a Long Non-Coding RNA Required for Cardiovascular Lineage Commitment. Cell 2014, 152, 570–583. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grote, P.; Wittler, L.; Hendrix, D.; Koch, F.; Währisch, S.; Beisaw, A.; Macura, K.; Bläss, G.; Kellis, M.; Werber, M.; et al. The Tissue-Specific LncRNA Fendrr Is an Essential Regulator of Heart and Body Wall Development in the Mouse. Dev. Cell 2013, 24, 206–214. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ritter, N.; Ali, T.; Kopitchinski, N.; Dimmeler, S.; Grote, P.; Ritter, N.; Ali, T.; Kopitchinski, N.; Schuster, P.; Beisaw, A.; et al. The LncRNA Locus Handsdown Regulates Cardiac Gene Programs and Is Essential for Early Mouse Article The LncRNA Locus Handsdown Regulates Cardiac Gene Programs and Is Essential for Early Mouse Development. Dev. Cell 2019, 50, 644–657. [Google Scholar] [CrossRef] [PubMed]
- Anderson, K.M.; Anderson, D.M.; Mcanally, J.R.; Shelton, J.M.; Bassel-Duby, R.; Olson, E.N. Transcription of the Non-Coding RNA Upperhand Controls Hand2 Expression and Heart Development. Nature 2016, 539, 433–436. [Google Scholar] [CrossRef] [Green Version]
- Wilson, K.; Ameen, M.; Guo, H.; Abilez, O.J.; Tian, L.; Mumbach, M.R.; Guo, H.; Karakikes, I.; Chang, H.Y.; Wu, J.C. Article Endogenous Retrovirus-Derived LncRNA BANCR Promotes Cardiomyocyte Migration in Humans and Non-Human Primates Ll Article Endogenous Retrovirus-Derived LncRNA BANCR Promotes Cardiomyocyte Migration in Humans and Non-Human Primates. Dev. Cell 2020, 54, 694–709.e9. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Cheng, Z.; Yu, Z.; Zhu, C.; Qian, L. Role of LncRNA Uc.457 in the Differentiation and Maturation of Cardiomyocytes. Mol. Med. Rep. 2019, 19, 4927–4934. [Google Scholar] [CrossRef]
- Jha, R.; Li, D.; Wu, Q.; Ferguson, K.E.; Forghani, P.; Gibson, G.C.; Xu, C. A Long Non-Coding RNA GATA6-AS1 Adjacent to GATA6 Is Required for Cardiomyocyte Differentiation from Human Pluripotent Stem Cells. FASEB J. 2020, 34, 14336–14352. [Google Scholar] [CrossRef]
- Kang, X.; Zhao, Y.; Van Arsdell, G.; Nelson, S.F.; Touma, M. Ppp1r1b-LncRNA Inhibits PRC2 at Myogenic Regulatory Genes to Promote Cardiac and Skeletal Muscle Development in Mouse and Human. RNA 2020, 26, 481–491. [Google Scholar] [CrossRef] [Green Version]
- Ballarino, M.; Cipriano, A.; Tita, R.; Santini, T.; Desideri, F.; Morlando, M.; Colantoni, A.; Carrieri, C.; Nicoletti, C.; Musarò, A.; et al. Deficiency in the Nuclear Long Noncoding RNA Charme Causes Myogenic Defects and Heart Remodeling in Mice. EMBO J. 2018, 37, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Porrello, E.R.; Olson, E.N. A Neonatal Blueprint for Cardiac Regeneration. Stem Cell Res. 2014, 13, 556–570. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Porrello, E.R.; Mahmoud, A.I.; Simpson, E.; Hill, J.A.; Richardson, J.A.; Olson, E.N.; Sadek, H.A. Transient Regenerative Potential of the Neonatal Mouse Heart. Science 2011, 331, 1078–1080. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, Y.-M.; Li, H.; Fan, Y.; Zhang, Q.-J.; Li, X.; Wu, L.-J.; Chen, Z.-J.; Zhu, C.; Qian, L.-M. Identification of Differentially Expressed LncRNAs Involved in Transient Regeneration of the Neonatal C57BL/6J Mouse Heart by next-Generation High-Throughput RNA Sequencing. Oncotarget 2017, 8, 28052. [Google Scholar] [CrossRef]
- Adamowicz, M.; Morgan, C.C.; Haubner, B.J.; Noseda, M.; Collins, M.J.; Paiva, M.A.; Srivastava, P.K.; Gellert, P.; Razzaghi, B.; O’Gara, P.; et al. Functionally Conserved Noncoding Regulators of Cardiomyocyte Proliferation and Regeneration in Mouse and Human. Circ. Genomic Precis. Med. 2018, 11. [Google Scholar] [CrossRef] [Green Version]
- Cai, B.; Ma, W.; Ding, F.; Zhang, L.; Huang, Q.; Wang, X.; Hua, B.; Xu, J.; Li, J.; Bi, C.; et al. The Long Noncoding RNA CAREL Controls Cardiac Regeneration. J. Am. Coll. Cardiol. 2018, 72, 534–550. [Google Scholar] [CrossRef]
- Cai, B.; Ma, W.; Wang, X.; Sukhareva, N.; Hua, B.; Zhang, L.; Xu, J.; Li, X.; Li, S.; Liu, S.; et al. Targeting LncDACH1 Promotes Cardiac Repair and Regeneration after Myocardium Infarction. Cell Death Differ. 2020, 27, 2158–2175. [Google Scholar] [CrossRef]
- Ponnusamy, M.; Liu, F.; Zhang, Y.H.; Li, R.B.; Zhai, M.; Liu, F.; Zhou, L.Y.; Liu, C.Y.; Yan, K.W.; Dong, Y.H.; et al. Long Noncoding RNA CPR (Cardiomyocyte Proliferation Regulator) Regulates Cardiomyocyte Proliferation and Cardiac Repair. Circulation 2019, 139, 2668–2684. [Google Scholar] [CrossRef]
- Han, P.; Li, W.; Lin, C.H.; Yang, J.; Shang, C.; Nurnberg, S.T.; Jin, K.K.; Xu, W.; Lin, C.Y.; Lin, C.J.; et al. A Long Noncoding RNA Protects the Heart from Pathological Hypertrophy. Nature 2014, 514, 102–106. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sato, M.; Kadomatsu, T.; Miyata, K.; Warren, J.S.; Tian, Z.; Zhu, S.; Horiguchi, H.; Makaju, A.; Bakhtina, A.; Morinaga, J.; et al. The LncRNA Caren Antagonizes Heart Failure by Inactivating DNA Damage Response and Activating Mitochondrial Biogenesis. Nat. Commun. 2021, 12, 1–21. [Google Scholar] [CrossRef]
- Viereck, J.; Bührke, A.; Foinquinos, A.; Chatterjee, S.; Kleeberger, J.A.; Xiao, K.; Janssen-Peters, H.; Batkai, S.; Ramanujam, D.; Kraft, T.; et al. Targeting Muscle-Enriched Long Non-Coding RNA H19 Reverses Pathological Cardiac Hypertrophy. Eur. Heart J. 2020, 41, 3462–3474. [Google Scholar] [CrossRef]
- Piccoli, M.T.; Gupta, S.K.; Viereck, J.; Foinquinos, A.; Samolovac, S.; Kramer, F.L.; Garg, A.; Remke, J.; Zimmer, K.; Batkai, S.; et al. Inhibition of the Cardiac Fibroblast-Enriched LncRNA Meg3 Prevents Cardiac Fibrosis and Diastolic Dysfunction. Circ. Res. 2017, 121, 575–583. [Google Scholar] [CrossRef] [PubMed]
- Ishii, N.; Ozaki, K.; Sato, H.; Mizuno, H.; Susumu, S.; Takahashi, A.; Miyamoto, Y.; Ikegawa, S.; Kamatani, N.; Hori, M.; et al. Identification of a Novel Non-Coding RNA, MIAT, That Confers Risk of Myocardial Infarction. J. Hum. Genet. 2006, 51, 1087–1099. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kumarswamy, R.; Bauters, C.; Volkmann, I.; Maury, F.; Fetisch, J.; Holzmann, A.; Lemesle, G.; De Groote, P.; Pinet, F.; Thum, T. Circulating Long Noncoding RNA, LIPCAR, Predicts Survival in Patients with Heart Failure. Circ. Res. 2014, 114, 1569–1575. [Google Scholar] [CrossRef] [Green Version]
- Vausort, M.; Wagner, D.R.; Devaux, Y. Long Noncoding RNAs in Patients with Acute Myocardial Infarction. Circ. Res. 2014, 115, 668–677. [Google Scholar] [CrossRef] [Green Version]
- Wang, K.; Long, B.; Zhou, L.Y.; Liu, F.; Zhou, Q.Y.; Liu, C.Y.; Fan, Y.Y.; Li, P.F. CARL LncRNA Inhibits Anoxia-Induced Mitochondrial Fission and Apoptosis in Cardiomyocytes by Impairing MiR-539-Dependent PHB2 Downregulation. Nat. Commun. 2014, 5. [Google Scholar] [CrossRef] [Green Version]
- Chen, G.; Li, H.; Li, X.; Li, B.; Zhong, L.; Huang, S.; Zheng, H.; Li, M.; Jin, G.; Liao, W.; et al. Loss of Long Non-Coding RNA CRRL Promotes Cardiomyocyte Regeneration and Improves Cardiac Repair by Functioning as a Competing Endogenous RNA. J. Mol. Cell. Cardiol. 2018, 122, 152–164. [Google Scholar] [CrossRef] [Green Version]
- Gong, L.; Xu, H.; Chang, H.; Tong, Y.; Zhang, T.; Guo, G. Knockdown of Long Non-Coding RNA MEG3 Protects H9c2 Cells from Hypoxia-Induced Injury by Targeting MicroRNA-183. J. Cell. Biochem. 2018, 119, 1429–1440. [Google Scholar] [CrossRef] [PubMed]
- Anderson, D.M.; Anderson, K.M.; Nelson, B.R.; McAnally, J.R.; Bezprozvannaya, S.; Shelton, J.M.; Bassel-Duby, R.; Olson, E.N. A Myocardin-Adjacent LncRNA Balances SRF-Dependent Gene Transcription in the Heart. Genes Dev. 2021, 35, 835–840. [Google Scholar] [CrossRef]
- Nakaya, Y.; Sheng, G. Epithelial to Mesenchymal Transition during Gastrulation: An Embryological View. Dev. Growth Differ. 2008, 50, 755–766. [Google Scholar] [CrossRef] [PubMed]
- Lepper, C.; Partridge, T.A.; Fan, C.M. An Absolute Requirement for Pax7-Positive Satellite Cells in Acute Injury-Induced Skeletal Muscle Regeneration. Development 2011, 138, 3639–3646. [Google Scholar] [CrossRef] [Green Version]
- Chargé, S.B.P.; Rudnicki, M.A. Cellular and Molecular Regulation of Muscle Regeneration. Physiol. Rev. 2004, 84, 209–238. [Google Scholar] [CrossRef] [PubMed]
- Wosczyna, M.N.; Rando, T.A. A Muscle Stem Cell Support Group: Coordinated Cellular Responses in Muscle Regeneration. Dev. Cell 2018, 46, 135–143. [Google Scholar] [CrossRef] [Green Version]
- Yin, H.; Price, F.; Rudnicki, M.A. Satellite Cells and the Muscle Stem Cell Niche. Physiol. Rev. 2013, 93, 23–67. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Relaix, F.; Rocancourt, D.; Mansouri, A.; Buckingham, M. A Pax3/Pax7-Dependent Population of Skeletal Muscle Progenitor Cells. Nature 2005, 435, 948–953. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Desjardins, C.A.; Naya, F.J. The Function of the MEF2 Family of Transcription Factors in Cardiac Development, Cardiogenomics, and Direct Reprogramming. J. Cardiovasc. Dev. Dis. 2016, 3, 26. [Google Scholar] [CrossRef] [Green Version]
- Bonnet, S.; Boucherat, O.; Paulin, R.; Wu, D.; Hindmarch, C.C.T.; Archer, S.L.; Song, R.; Moore, J.B.; Provencher, S.; Zhang, L.; et al. Clinical Value of Non-Coding RNAs in Cardiovascular, Pulmonary, and Muscle Diseases. Am. J. Physiol.-Cell Physiol. 2020, 318, C1–C28. [Google Scholar] [CrossRef]
- Luo, S.; Lu, J.Y.; Liu, L.; Yin, Y.; Chen, C.; Han, X.; Wu, B.; Xu, R.; Liu, W.; Yan, P.; et al. Divergent LncRNAs Regulate Gene Expression and Lineage Differentiation in Pluripotent Cells. Cell Stem Cell 2016, 18, 637–652. [Google Scholar] [CrossRef]
- Frank, S.; Ahuja, G.; Bartsch, D.; Russ, N.; Yao, W.; Kuo, J.C.C.; Derks, J.P.; Akhade, V.S.; Kargapolova, Y.; Georgomanolis, T.; et al. YylncT Defines a Class of Divergently Transcribed LncRNAs and Safeguards the T-Mediated Mesodermal Commitment of Human PSCs. Cell Stem Cell 2019, 24, 318–327.e8. [Google Scholar] [CrossRef] [Green Version]
- Gonçalves, T.J.M.; Armand, A.S. Non-Coding RNAs in Skeletal Muscle Regeneration. Non-Coding RNA Res. 2017, 2, 56–67. [Google Scholar] [CrossRef]
- Boudoukha, S.; Cuvellier, S.; Polesskaya, A. Role of the RNA-Binding Protein IMP-2 in Muscle Cell Motility. Mol. Cell. Biol. 2010, 30, 5710–5725. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gong, C.; Li, Z.; Ramanujan, K.; Clay, I.; Zhang, Y.; Lemire-Brachat, S.; Glass, D.J. A Long Non-Coding RNA, LncMyoD, Regulates Skeletal Muscle Differentiation by Blocking IMP2-Mediated MRNA Translation. Dev. Cell 2015, 34, 181–191. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cesana, M.; Cacchiarelli, D.; Legnini, I.; Santini, T.; Sthandier, O.; Chinappi, M.; Tramontano, A.; Bozzoni, I. A Long Noncoding RNA Controls Muscle Differentiation by Functioning as a Competing Endogenous RNA. Cell 2011, 147, 358–369. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Von Roretz, C.; Beauchamp, P.; Di Marco, S.; Gallouzi, I.E. HuR and Myogenesis: Being in the Right Place at the Right Time. Biochim. Biophys. Acta Mol. Cell Res. 2011, 1813, 1663–1667. [Google Scholar] [CrossRef] [Green Version]
- Legnini, I.; Morlando, M.; Mangiavacchi, A.; Fatica, A.; Bozzoni, I. A Feedforward Regulatory Loop between HuR and the Long Noncoding RNA Linc-MD1 Controls Early Phases of Myogenesis. Mol. Cell 2014, 53, 506–514. [Google Scholar] [CrossRef] [Green Version]
- Zhu, M.; Liu, J.; Xiao, J.; Yang, L.; Cai, M.; Shen, H.; Chen, X.; Ma, Y.; Hu, S.; Wang, Z.; et al. Lnc-Mg Is a Long Non-Coding RNA That Promotes Myogenesis. Nat. Commun. 2017, 8, 1–11. [Google Scholar] [CrossRef] [Green Version]
- Zhou, L.; Sun, K.; Zhao, Y.; Zhang, S.; Wang, X.; Li, Y.; Lu, L.; Chen, X.; Chen, F.; Bao, X.; et al. Linc-YY1 Promotes Myogenic Differentiation and Muscle Regeneration through an Interaction with the Transcription Factor YY1. Nat. Commun. 2015, 6. [Google Scholar] [CrossRef] [Green Version]
- Mueller, A.C.; Cichewicz, M.A.; Dey, B.K.; Layer, R.; Reon, B.J.; Gagan, J.R.; Dutta, A. MUNC, a Long Noncoding RNA That Facilitates the Function of MyoD in Skeletal Myogenesis. Mol. Cell. Biol. 2015, 35, 498–513. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.; Zhao, Y.; Bao, X.; Zhu, X.; Kwok, Y.K.Y.; Sun, K.; Chen, X.; Huang, Y.; Jauch, R.; Esteban, M.A.; et al. LncRNA Dum Interacts with Dnmts to Regulate Dppa2 Expression during Myogenic Differentiation and Muscle Regeneration. Cell Res. 2015, 25, 335–350. [Google Scholar] [CrossRef] [Green Version]
- Watts, R.; Johnsen, V.L.; Shearer, J.; Hittel, D.S. Myostatin-Induced Inhibition of the Long Noncoding RNA Malat1 Is Associated with Decreased Myogenesis. Am. J. Physiol. Cell Physiol. 2013, 304. [Google Scholar] [CrossRef] [Green Version]
- Chen, X.; He, L.; Zhao, Y.; Li, Y.; Zhang, S.; Sun, K.; So, K.; Chen, F.; Zhou, L.; Lu, L.; et al. Malat1 Regulates Myogenic Differentiation and Muscle Regeneration through Modulating MyoD Transcriptional Activity. Cell Discov. 2017, 3, 1–23. [Google Scholar] [CrossRef]
- Dey, B.K.; Pfeifer, K.; Dutta, A. The H19 Long Noncoding RNA Gives Rise to MicroRNAs MiR-675-3p and MiR-675-5p to Promote Skeletal Muscle Differentiation and Regeneration. Genes Dev. 2014, 28, 491–501. [Google Scholar] [CrossRef] [Green Version]
- Martinet, C.; Monnier, P.; Louault, Y.; Benard, M.; Gabory, A.; Dandolo, L. H19 Controls Reactivation of the Imprinted Gene Network during Muscle Regeneration. Development 2016, 143, 962–971. [Google Scholar] [CrossRef] [Green Version]
- Dill, T.L.; Carroll, A.; Pinheiro, A.; Gao, J.; Naya, F.J. The Long Noncoding RNA Meg3 Regulates Myoblast Plasticity and Muscle Regeneration through Epithelial-Mesenchymal Transition. Dev. Camb. Engl. 2021, 148. [Google Scholar] [CrossRef]
- Wang, J.; Gong, C.; Maquat, L.E. Control of Myogenesis by Rodent SINE-Containing LncRNAs. Genes Dev. 2013, 27, 793–804. [Google Scholar] [CrossRef] [Green Version]
- Lu, L.; Sun, K.; Chen, X.; Zhao, Y.; Wang, L.; Zhou, L.; Sun, H.; Wang, H. Genome-Wide Survey by ChIP-Seq Reveals YY1 Regulation of LincRNAs in Skeletal Myogenesis. EMBO J. 2013, 32, 2575–2588. [Google Scholar] [CrossRef] [Green Version]
- Dimartino, D.; Colantoni, A.; Ballarino, M.; Martone, J.; Mariani, D.; Danner, J.; Bruckmann, A.; Meister, G.; Morlando, M.; Bozzoni, I. The Long Non-Coding RNA Lnc-31 Interacts with Rock1 MRNA and Mediates Its YB-1-Dependent Translation. Cell Rep. 2018, 23, 733–740. [Google Scholar] [CrossRef] [Green Version]
- Ballarino, M.; Cazzella, V.; D’Andrea, D.; Grassi, L.; Bisceglie, L.; Cipriano, A.; Santini, T.; Pinnarò, C.; Morlando, M.; Tramontano, A.; et al. Novel Long Noncoding RNAs (LncRNAs) in Myogenesis: A MiR-31 Overlapping LncRNA Transcript Controls Myoblast Differentiation. Mol. Cell. Biol. 2015, 35, 728–736. [Google Scholar] [CrossRef] [Green Version]
- Wang, G.Q.; Wang, Y.; Xiong, Y.; Chen, X.C.; Ma, M.L.; Cai, R.; Gao, Y.; Sun, Y.M.; Yang, G.S.; Pang, W.J. Sirt1 AS LncRNA Interacts with Its MRNA to Inhibit Muscle Formation by Attenuating Function of MiR-34a. Sci. Rep. 2016, 6, 1–13. [Google Scholar] [CrossRef]
- Jin, J.J.; Lv, W.; Xia, P.; Xu, Z.Y.; Zheng, A.D.; Wang, X.J.; Wang, S.S.; Zeng, R.; Luo, H.M.; Li, G.L.; et al. Long Noncoding RNA SYISL Regulates Myogenesis by Interacting with Polycomb Repressive Complex 2. Proc. Natl. Acad. Sci. USA 2018, 115, E9802–E9811. [Google Scholar] [CrossRef] [Green Version]
- Neppl, R.L.; Wu, C.L.; Walsh, K. IncRNA Chronos Is an Aging-Induced Inhibitor of Muscle Hypertrophy. J. Cell Biol. 2017, 216, 3497–3507. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Y.; Li, Y.; Hu, Q.; Xi, Y.; Xing, Z.; Zhang, Z.; Wu, J.; Liang, K.; Nguyen, T.K.; Egranov, S.D.; et al. LncRNA H19 Alleviates Muscular Dystrophy Through Stabilizing Dystrophin. Nat. Cell Biol. 2021, 22, 1332–1345. [Google Scholar] [CrossRef]
- Cabianca, D.S.; Casa, V.; Bodega, B.; Xynos, A.; Ginelli, E.; Tanaka, Y.; Gabellini, D. A Long NcRNA Links Copy Number Variation to a Polycomb/Trithorax Epigenetic Switch in FSHD Muscular Dystrophy. Cell 2012, 149, 819–831. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Matsumoto, A.; Pasut, A.; Matsumoto, M.; Yamashita, R.; Fung, J.; Monteleone, E.; Saghatelian, A.; Nakayama, K.I.; Clohessy, J.G.; Pandolfi, P.P. MTORC1 and Muscle Regeneration Are Regulated by the LINC00961-Encoded SPAR Polypeptide. Nature 2017, 541, 228–232. [Google Scholar] [CrossRef]
- Lanz, R.B.; McKenna, N.J.; Onate, S.A.; Albrecht, U.; Wong, J.; Tsai, S.Y.; Tsai, M.J.; O’Malley, B.W. A Steroid Receptor Coactivator, SRA, Functions as an RNA and Is Present in an SRC-1 Complex. Cell 1999, 97, 17–27. [Google Scholar] [CrossRef] [Green Version]
- Caretti, G.; Louis Schiltz, R.; Dilworth, F.J.; Di Padova, M.; Zhao, P.; Ogryzko, V.; Fuller-Pace, F.V.; Hoffman, E.P.; Tapscott, S.J.; Sartorelli, V. The RNA Helicases P68/P72 and the Noncoding RNA SRA Are Coregulators of MyoD and Skeletal Muscle Differentiation. Dev. Cell 2006, 11, 547–560. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hubé, F.; Velasco, G.; Rollin, J.; Furling, D.; Francastel, C.; Lanz, R.B.; McKenna, N.J.; Onate, S.A.; Albrecht, U.; Wong, J.; et al. Steroid Receptor RNA Activator Protein Binds to and Counteracts SRA RNA-Mediated Activation of MyoD and Muscle Differentiation. Nucleic Acids Res. 2011, 39, 513–525. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Legnini, I.; Di Timoteo, G.; Rossi, F.; Morlando, M.; Briganti, F.; Sthandier, O.; Fatica, A.; Santini, T.; Andronache, A.; Wade, M.; et al. Circ-ZNF609 Is a Circular RNA That Can Be Translated and Functions in Myogenesis. Mol. Cell 2017, 66, 22–37. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rutering, J.; Ilmer, M.; Recio, A.; Coleman, M.; Vykoukal, J.; Alt, E.; Orleans, N. A Peptide Encoded by a Transcript Annotated as Long Noncoding RNA Enhances SERCA Activity in Muscle. Science 2016, 351, 271–275. [Google Scholar] [CrossRef] [Green Version]
- Li, E.; Beard, C.; Jaenisch, R. Role for DNA Methylation in Genomic Imprinting. Nature 1993, 366, 362–365. [Google Scholar] [CrossRef]
- Moresi, V.; Marroncelli, N.; Coletti, D.; Adamo, S. Regulation of Skeletal Muscle Development and Homeostasis by Gene Imprinting, Histone Acetylation and MicroRNA. Biochim. Biophys. Acta Gene Regul. Mech. 2015, 1849, 309–316. [Google Scholar] [CrossRef] [Green Version]
- Moore, T.; Haig, D. Genomic Imprinting in Mammalian Development: A Parental Tug-of-War. Trends Genet. 1991, 7, 45–49. [Google Scholar] [CrossRef]
- Bartolomei, M.S.; Ferguson-Smith, A.C. Mammalian Genomic Imprinting. Cold Spring Harb. Perspect. Biol. 2011, 3, 1–17. [Google Scholar] [CrossRef] [Green Version]
- Davis, R.L.; Weintraub, H.; Lassar, A.B. Expression of a Single Transfected CDNA Converts Fibmblasts to Myoblasts. Cell 1987, 51, 987–1000. [Google Scholar] [CrossRef]
- Tierling, S.; Dalbert, S.; Schoppenhorst, S.; Tsai, C.E.; Oliger, S.; Ferguson-Smith, A.C.; Paulsen, M.; Walter, J. High-Resolution Map and Imprinting Analysis of the Gtl2-Dnchc1 Domain on Mouse Chromosome 12. Genomics 2006, 87, 225–235. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Clark, A.L.; Naya, F.J. MicroRNAs in the Myocyte Enhancer Factor 2 (MEF2)-Regulated Gtl2-Dio3 Noncoding RNA Locus Promote Cardiomyocyte Proliferation by Targeting the Transcriptional Coactivator Cited2. J. Biol. Chem. 2015, 290, 23162–23172. [Google Scholar] [CrossRef] [Green Version]
- Snyder, C.M.; Rice, A.L.; Estrella, N.L.; Held, A.; Kandarian, S.C.; Naya, F.J. MEF2A Regulates the Gtl2-Dio3 MicroRNA Mega-Cluster to Modulate WNT Signaling in Skeletal Muscle Regeneration. Dev. Camb. 2013, 140, 31–42. [Google Scholar] [CrossRef] [Green Version]
- Dentice, M.; Salvatore, D. Local Impact of Thyroid Hormone Inactivation. J. Endocrinol. 2011, 209, 273–282. [Google Scholar] [CrossRef]
- Falix, F.A.; Aronson, D.C.; Lamers, W.H.; Gaemers, I.C. Possible Roles of DLK1 in the Notch Pathway during Development and Disease. Biochim. Biophys. Acta Mol. Basis Dis. 2012, 1822, 988–995. [Google Scholar] [CrossRef] [Green Version]
- Hagan, M.; Zhou, M.; Ashraf, M.; Kim, I.; Su, H.; Neal, L.; Tang, Y.; Hospital, R. Long Noncoding RNAs and Their Roles in Skeletal Muscle Fate Determination. Noncoding RNA Investig. 2018, 1–6. [Google Scholar] [CrossRef] [Green Version]
- Takahashi, N.; Okamoto, A.; Kobayashi, R.; Shirai, M.; Obata, Y.; Ogawa, H.; Sotomaru, Y.; Kono, T. Deletion of Gtl2, Imprinted Non-Coding RNA, with Its Differentially Methylated Region Induces Lethal Parent-Origin-Dependent Defects in Mice. Hum. Mol. Genet. 2009, 18, 1879–1888. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, Y.; Cheunsuchon, P.; Nakayama, Y.; Lawlor, M.W.; Zhong, Y.; Rice, K.A.; Zhang, L.; Zhang, X.; Gordon, F.E.; Lidov, H.G.W.; et al. Activation of Paternally Expressed Genes and Perinatal Death Caused by Deletion of the Gtl2 Gene. Dev. Stem Cells 2010, 137, 10. [Google Scholar] [CrossRef] [Green Version]
- Mademtzoglou, D.; Asakura, Y.; Borok, M.J.; Alonso-Martin, S.; Mourikis, P.; Kodaka, Y.; Mohan, A.; Asakura, A.; Relaix, F. Cellular Localization of the Cell Cycle Inhibitor Cdkn1c Controls Growth Arrest of Adult Skeletal Muscle Stem Cells. eLife 2018, 7, 1–25. [Google Scholar] [CrossRef]

| Name | Species | Function | Mechanism | Citation |
|---|---|---|---|---|
| Hdn * | m | Cardiac development; Negative regulation of Hand2 | Transcriptional regulation of Hand2 | [36] |
| Uph * | m | Ventricle development; Hand2 activation | Transcriptional regulation of Hand2 | [37] |
| Bvht * | m | Activation of cardiovascular gene network; Epigenetic regulation | SUZ12 interaction | [34] |
| Fendrr * | m | Chamber development | Binds to PRC2 and TrxG/MLL complexes | [31,35] |
| BANCR * | p | Cardiomyocyte migration | Downstream effector of TEAD/YAP | [38] |
| uc.457 *^ | m | Cardiomyocyte proliferation | Transcription factor interactions | [39] |
| GATA6-AS1 * | h | Cardiomyocyte differentiation | Regulation of gene expression | [40] |
| Ppp1r1b * | h, m | Striated muscle differentiation | PRC2 interaction | [41] |
| Charme * | h, m | Striated muscle differentiation | Chromatin interaction | [42] |
| MIAT ^ | h | SNP increases MI risk | ? | [54] |
| Mhrt ^ | m | Cardioprotective | Modulation of chromatin | [50] |
| LIPCAR ^ | h | Cardiac remodeling | ? | [55] |
| ANRIL ^ | h | Cardiac event biomarker | ? | [56] |
| KCNQ1OT1 ^ | h | Cardiac event biomarker | ? | [56] |
| Malat1 ^ | h | Cardiac event biomarker | ? | [56] |
| CARL ^ | m | Suppress mitochondrial fission | miR-539 sponge | [57] |
| CRP | m | Cardiomyocyte proliferation | Promotes methylation | [49] |
| CRLL | r | Cardiomyocyte proliferation | miR-199a3 sponge | [58] |
| H19 *^ | h, m | Cardiac development | Cardiomyocyte differentiation and hypertrophy | [31,52] |
| Meg3 ^ | h, m | Cardiac fibrosis | P53 interaction | [53,59] |
| Caren ^ | m | Cardioprotective | Decrease Hint1 translation | [51] |
| LncDACH1 ^ | h, m | Cardiomyocyte proliferation | PP1A inhibitor | [48] |
| CAREL | h, m | Cardiomyocyte proliferation | ceRNA for miR-296 | [47] |
| CARDINAL ^ | h, m | Cardioprotective | Inhibits SRF/TCF complex | [60] |
| Name | Species | Function | Mechanism | Citation |
|---|---|---|---|---|
| Evx1as * | m | Skeletal muscle differentiation | Chromatin binding | [69] |
| yylncT * | h | Mesodermal lineage commitment | Binds to DNMT3B | [70] |
| Sirt1 AS * | m | Promotes proliferation and represses differentiation | Sirt1 mRNA binding | [90] |
| Syisl * | m | Repress differentiation | Recruits Ezh2 | [91] |
| LncMyoD | h, m | Promotes differentiation | Blocks IMP mediate translation | [72,73] |
| linc-MD ^ | h, m | Promotes muscle specific transcription factors and differentiation | Sponges miR-133 and miR-133 | [74,76] |
| lnc-mg | m | Promotes differentiation | Sponges miR-125b | [77] |
| lnc-YY1 | h, m | Promotes differentiation | Binds YY1 | [78] |
| MUNC | m | Gene expression regulation | MyoD binding | [79] |
| Malat1 | h, m | Proliferation and differentiation | Recruitment of Suv39h | [81,82] |
| Dum | m | Promyogenic | Silences Duppa | [80] |
| Meg3 | h, m | Myoblast differentiation; Regulates EMT | PRC2 interaction | [85] |
| m1/2-sbsRNA | m | Promote cell activation | mRNA degradation | [71,86] |
| Yam-1 | m | Repress differentiation | Activates miR-715 | [87] |
| lnc-31 ^ | h, m | Promote proliferation and inhibits differentiation | ? | [89] |
| Chronos | m | Inhibition of hypertrophic growth | Represses Bmp7 | [92] |
| SRA | h, m | Regulation of gene expression | Ribonuclear complexes | [96] |
| DBE-T ^ | h | De-repression of 4q35 genes | Interaction with Ash1L | [94] |
| circ-ZNF609 | h, m | Proliferation | ? | [99] |
| LINC00961 | h, m | Micropeptide | mTOR mediated regeneration | [95] |
| Myoregulin | h, m | Micropeptide | Impedes calcium reuptake | [14] |
| DWORF | m | Micropeptide | Enhances SERCA activity | [100] |
| H19 ^ | h, m | Promotes myoblast differentiation/ Stabilizes dystrophin | Mediated via miR-675-3p and miR-675-3b; Association with dystrophin | [83,84,93] |
| Ppp1r1b * | h, m | Striated muscle differentiation | PRC2 interaction | [41] |
| Charme * | h, m | Striated muscle differentiation | Chromatin interaction | [42] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pinheiro, A.; Naya, F.J. The Key Lnc (RNA)s in Cardiac and Skeletal Muscle Development, Regeneration, and Disease. J. Cardiovasc. Dev. Dis. 2021, 8, 84. https://doi.org/10.3390/jcdd8080084
Pinheiro A, Naya FJ. The Key Lnc (RNA)s in Cardiac and Skeletal Muscle Development, Regeneration, and Disease. Journal of Cardiovascular Development and Disease. 2021; 8(8):84. https://doi.org/10.3390/jcdd8080084
Chicago/Turabian StylePinheiro, Amanda, and Francisco J. Naya. 2021. "The Key Lnc (RNA)s in Cardiac and Skeletal Muscle Development, Regeneration, and Disease" Journal of Cardiovascular Development and Disease 8, no. 8: 84. https://doi.org/10.3390/jcdd8080084
APA StylePinheiro, A., & Naya, F. J. (2021). The Key Lnc (RNA)s in Cardiac and Skeletal Muscle Development, Regeneration, and Disease. Journal of Cardiovascular Development and Disease, 8(8), 84. https://doi.org/10.3390/jcdd8080084






