Design and Pilot Evaluation of an IoT-Based Blood Pressure Monitoring System for Rabbits
Abstract
1. Introduction
2. Materials and Methods
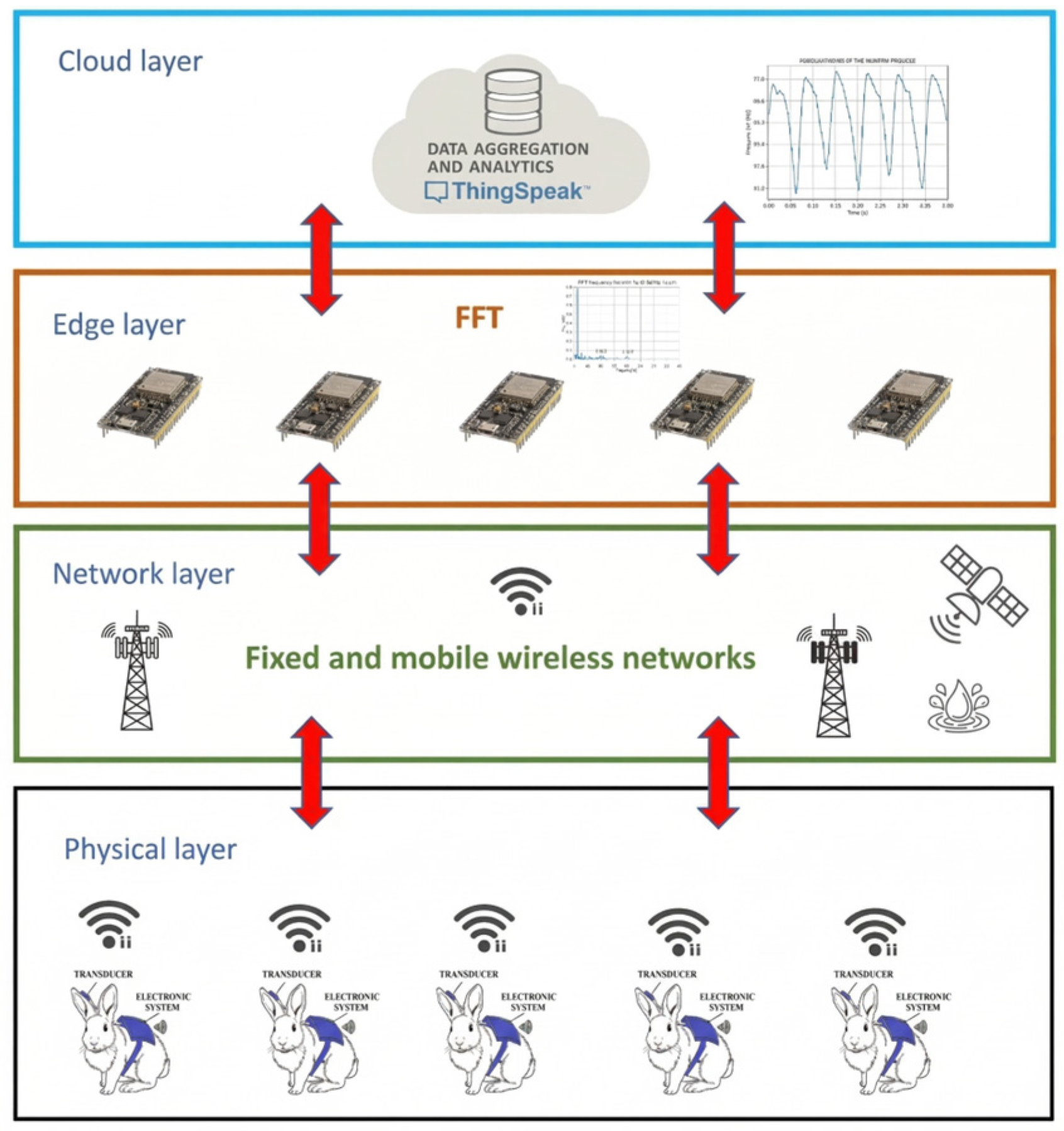
2.1. System Architecture
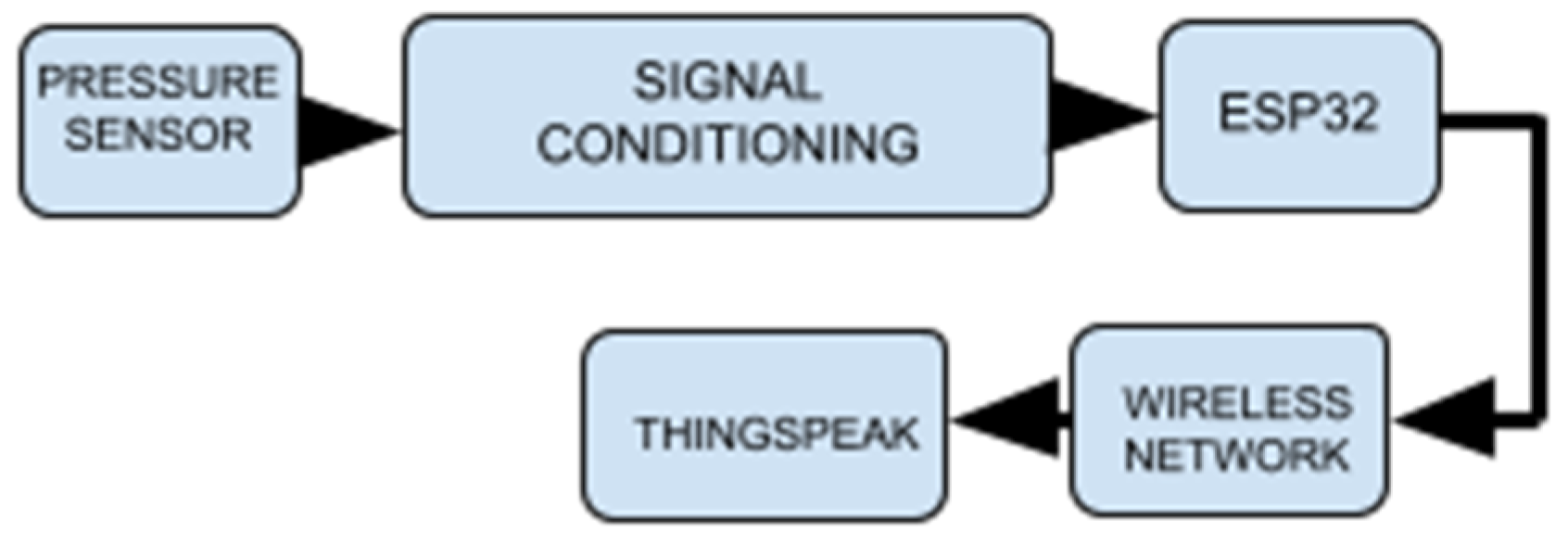
2.2. System Development
2.3. Rabbit Preparation
2.4. Experimental Procedures
3. Results
3.1. System Calibration
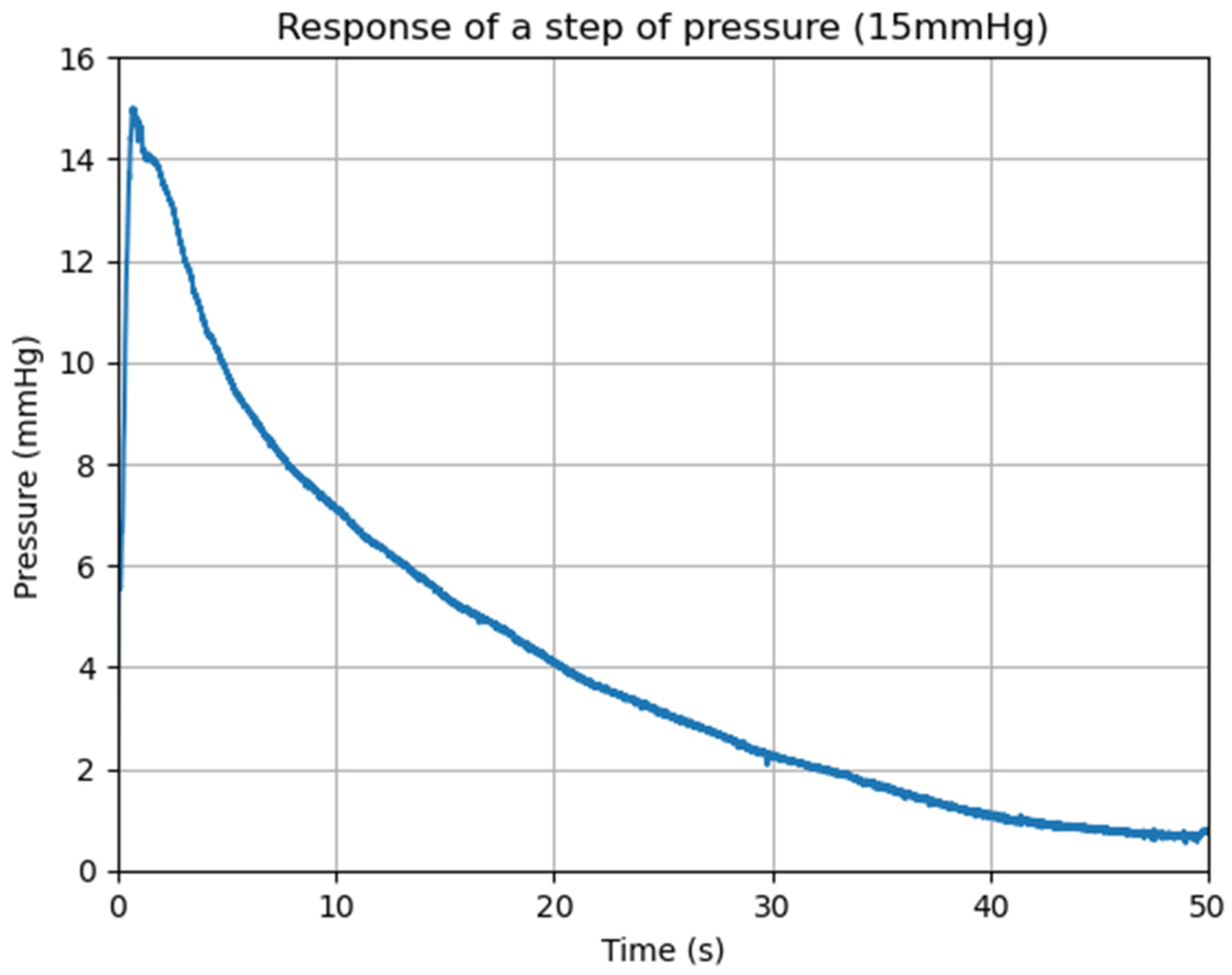
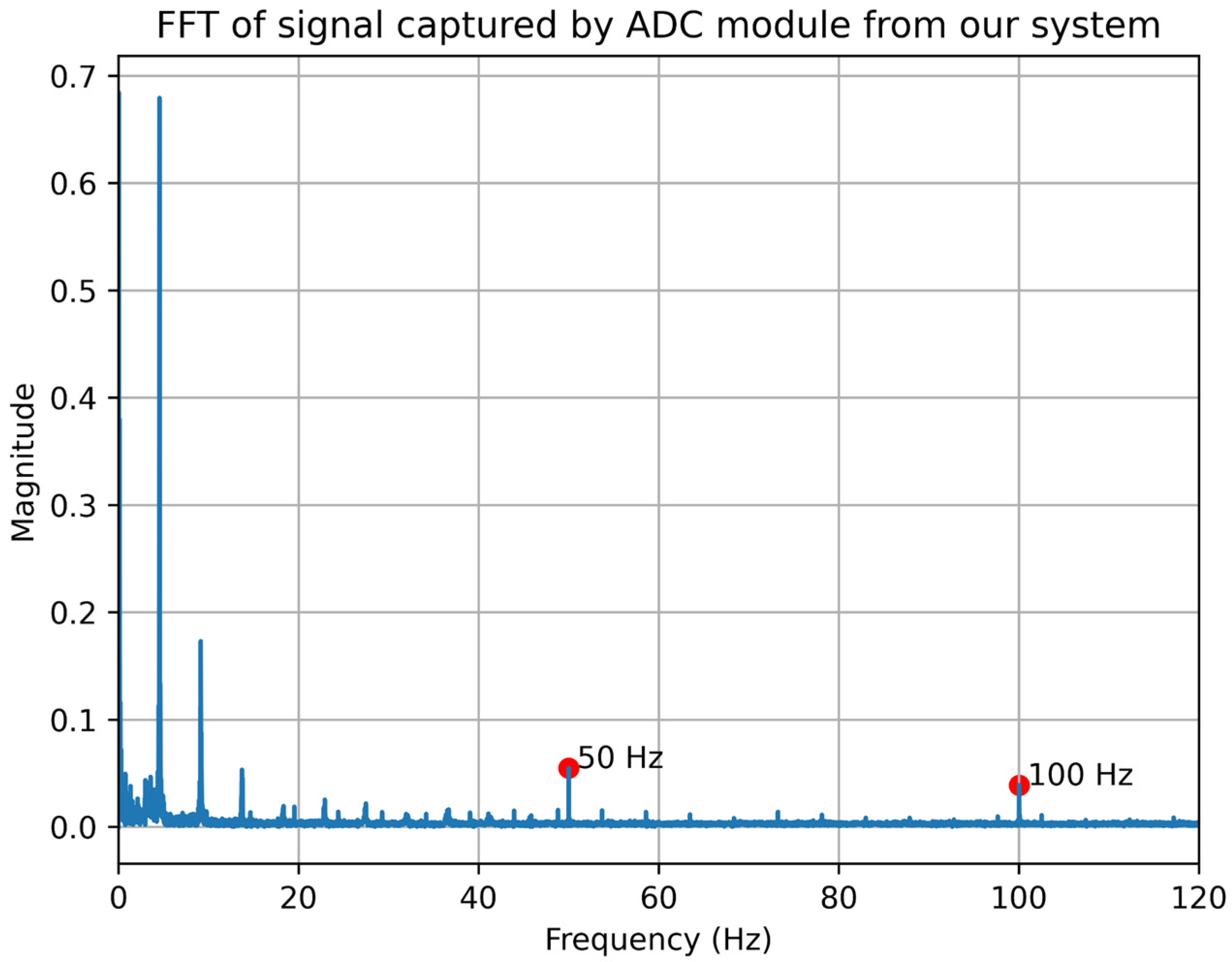
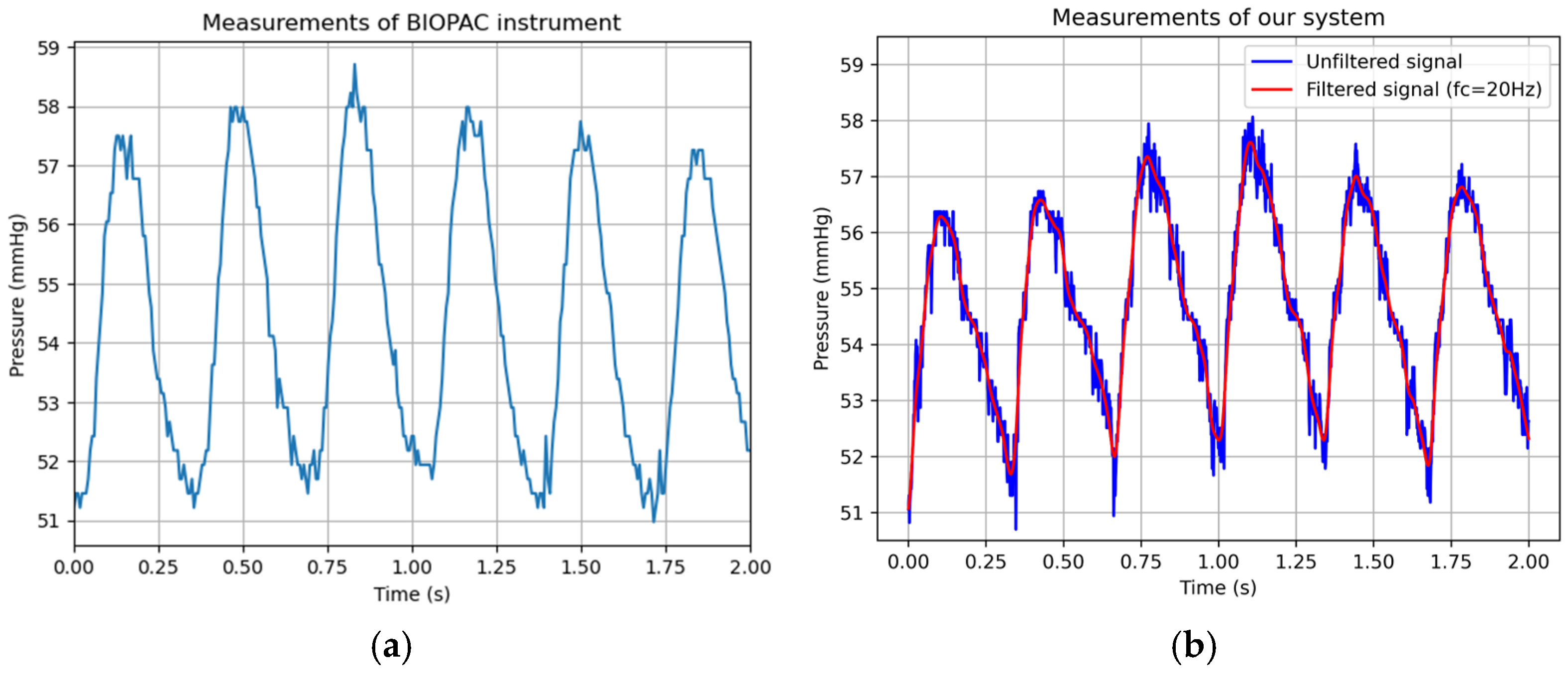
3.2. Data Acquisition and Processing
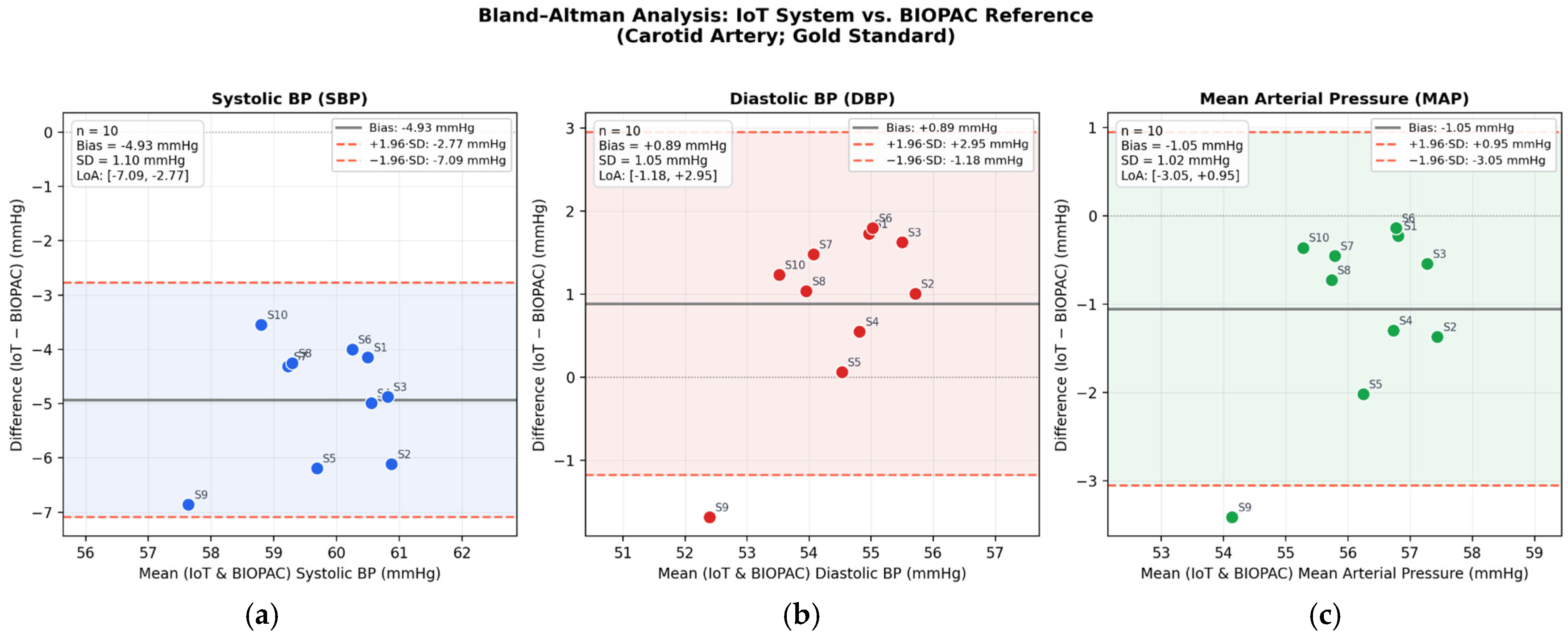
3.3. Performance Assessment in the Non-Terminal Group (Group 1)
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Han, Y.; Wei, T.; Chen, Z.; Wang, H.; Wang, L.; Li, C.; Mei, X.; Kuang, L.; Gong, J. LIVEMOS-G: A High Throughput Gantry Monitoring System with Multi-Source Imaging and Environmental Sensing for Large-Scale Commercial Rabbit Farming. Animals 2025, 15, 3177. [Google Scholar] [CrossRef]
- Ashton, K. That ‘Internet of Things’ Thing. RFID J. 2009, 22, 97–114. [Google Scholar]
- Vailshery, L.S. Number of Internet of Things (IoT) Connected Devices Worldwide from 2019 to 2021, with Forecasts from 2022 to 2034; Statista: Hamburg, Germany, 2022; Available online: https://www.statista.com/statistics/1183457/iot-connected-devices-worldwide/ (accessed on 15 February 2026).
- Yan, H.; Xu, L.D.; Bi, Z.; Pang, Z.; Zhang, J.; Chen, Y. An Emerging Technology—Wearable Wireless Sensor Networks with Applications in Human Health Condition Monitoring. J. Manag. Anal. 2015, 2, 121–137. [Google Scholar] [CrossRef]
- Sadowski, S.; Spachos, P. Wireless Technologies for Smart Agricultural Monitoring Using Internet of Things Devices with Energy Harvesting Capabilities. Comput. Electron. Agric. 2020, 172, 105338. [Google Scholar] [CrossRef]
- Zhang, M.; Wang, X.; Feng, H.; Huang, Q.; Xiao, X.; Zhang, X. Wearable Internet of Things Enabled Precision Livestock Farming in Smart Farms: A Review of Technical Solutions for Precise Perception, Biocompatibility, and Sustainability Monitoring. J. Clean. Prod. 2021, 312, 127712. [Google Scholar] [CrossRef]
- Li, C.; Wang, J.; Wang, S.; Zhang, Y. A Review of IoT Applications in Healthcare. Neurocomputing 2024, 565, 127017. [Google Scholar] [CrossRef]
- Karthick, G.S.; Sridhar, M.; Pankajavalli, P.B. Internet of Things in Animal Healthcare (IoTAH): Review of Recent Advancements in Architecture, Sensing Technologies and Real-Time Monitoring. SN Comput. Sci. 2020, 1, 301. [Google Scholar] [CrossRef]
- Bello, S.A.; Passaglia, C.L. A Wireless Pressure Sensor for Continuous Monitoring of Intraocular Pressure in Conscious Animals. Ann. Biomed. Eng. 2017, 45, 2592–2604. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Niimi, M.; Zhang, L.; Tang, X.; Lu, J.; Fan, J. A Simple Telemetry Sensor System for Monitoring Body Temperature in Rabbits—A Brief Report. Animals 2023, 13, 1677. [Google Scholar] [CrossRef]
- Pariaut, R. Cardiovascular Physiology and Diseases of the Rabbit. Vet. Clin. North Am. Exot. Anim. Pract. 2009, 12, 135–144. [Google Scholar] [CrossRef]
- Williams, P.B.; Schapiro, H.; Yeiser, P.E. Noninvasive Blood Pressure Determination in the Rabbit with a Doppler Ultrasound Probe. Proc. Soc. Exp. Biol. Med. 1979, 161, 417–420. [Google Scholar] [CrossRef]
- Jekl, V.; Agudelo, C.F.; Hauptman, K. Cardiology in Rodents, Rabbits, and Small Exotic Mammals—Diagnostic Workup. Vet. Clin. Exot. Anim. Pract. 2022, 25, 503–524. [Google Scholar] [CrossRef] [PubMed]
- Calero Rodriguez, A.; Van Zeeland, Y.R.; Schoemaker, N.J.; De Grauw, J.C. Agreement between Invasive and Oscillometric Arterial Blood Pressure Measurement Using a High-Definition Oscillometric Device in Normotensive New Zealand White Rabbits Using Two Different Anaesthetic Protocols. Vet. Anaesth. Analg. 2021, 48, 679–687. [Google Scholar] [CrossRef]
- Barter, L.S.; Epstein, S.E. Comparison of Doppler, Oscillometric, Auricular and Carotid Arterial Blood Pressure Measurements in Isoflurane Anesthetized New Zealand White Rabbits. Vet. Anaesth. Analg. 2014, 41, 393–397. [Google Scholar] [CrossRef] [PubMed]
- Data Sciences International. Pressure Sensing Technologies. Available online: https://www.datasci.com/solutions/cardiovascular/pressure-sensing-technologies (accessed on 16 March 2026).
- emka TECHNOLOGIES. easyTEL+ Digital Telemetry. Available online: https://www.emkatech.com/product/implanted-telemetry/ (accessed on 16 March 2026).
- Akhigbe, B.I.; Munir, K.; Akinade, O.; Akanbi, L.; Oyedele, L.O. IoT Technologies for Livestock Management: A Review of Present Status, Opportunities, and Future Trends. Big Data Cogn. Comput. 2021, 5, 10. [Google Scholar] [CrossRef]
- The MathWorks, Inc. Learn More About ThingSpeak; The MathWorks, Inc.: Natick, MA, USA, 2026; Available online: https://thingspeak.mathworks.com/pages/learn_more (accessed on 16 February 2026).
- OpenFog Consortium. Openfog Reference Architecture for Fog Computing. 2017. Available online: https://www.iiconsortium.org/pdf/OpenFog_Reference_Architecture_2_09_17.pdf (accessed on 23 March 2026).
- Markakis, E.; Mastorakis, G.; Mavromoustakis, C.X.; Pallis, E. Cloud and Fog Computing in 5G Mobile Networks: Emerging Advances and Applications; Institution of Engineering and Technology: Stevenage, UK, 2017; ISBN 1-78561-083-X. [Google Scholar]
- Rahmani, A.M.; Gia, T.N.; Negash, B.; Anzanpour, A.; Azimi, I.; Jiang, M.; Liljeberg, P. Exploiting Smart E-Health Gateways at the Edge of Healthcare Internet-of-Things: A Fog Computing Approach. Future Gener. Comput. Syst. 2018, 78, 641–658. [Google Scholar] [CrossRef]
- Texas Instruments. INA122 Datasheet. Available online: https://www.ti.com/lit/ds/symlink/ina122.pdf (accessed on 16 February 2026).
- Espressif Systems. ADC API Reference (ESP-IDF v4.4). Available online: https://docs.espressif.com/projects/esp-idf/en/v4.4/esp32/api-reference/peripherals/adc.html (accessed on 16 February 2026).
- Harrenstien, K. Time Server; RFC 738; Internet Engineering Task Force: Fremont, CA, USA, 1977; Available online: https://www.rfc-editor.org/rfc/rfc738.html (accessed on 16 February 2026).
- Mills, D.; Martin, J.; Burbank, J.; Kasch, W. Network Time Protocol Version 4: Protocol and Algorithms Specification; RFC 5905; Internet Engineering Task Force: Fremont, CA, USA, 2010; Available online: https://www.rfc-editor.org/rfc/rfc5905 (accessed on 16 February 2026).
- Espressif Systems. Sleep Modes API Reference. Available online: https://docs.espressif.com/projects/esp-idf/en/stable/esp32/api-reference/system/sleep_modes.html (accessed on 16 February 2026).
- INSIBIO. Apoyo a la Investigación—Conejos. Available online: http://insibio.org.ar/apoyo-a-la-investigacion/conejos/ (accessed on 16 February 2026).
- Comolli, J.; d’Ovidio, D.; Adami, C.; Schnellbacher, R. Technological Advances in Exotic Pet Anesthesia and Analgesia. Vet. Clin. Exot. Anim. Pract. 2019, 22, 419–439. [Google Scholar] [CrossRef] [PubMed]
- BIOPAC Systems, Inc. AcqKnowledge Software Guide. Available online: https://www.biopac.com/wp-content/uploads/acqknowledge_software_guide.pdf (accessed on 16 February 2026).
- Limaye, H.; Deshmukh, V.V. ECG Noise Sources and Various Noise Removal Techniques: A Survey. Int. J. Appl. Innov. Eng. Manag. 2016, 5, 86–92. [Google Scholar]
- Mewett, D.T.; Nazeran, H.; Reynolds, K.J. Removing Power Line Noise from Recorded EMG. In Proceedings of the 2001 Conference Proceedings of the 23rd Annual International Conference of the IEEE Engineering in Medicine and Biology Society, Istanbul, Turkey, 25–28 October 2001; IEEE: Piscataway, NJ, USA, 2001; Volume 3, pp. 2190–2193. [Google Scholar]
- Tompkins, W.J. Biomedical Digital Signal Processing; Prentice Hall: Saddle River, NJ, USA, 1993; Volume 237. [Google Scholar]
- Lyons, R.G. Understanding Digital Signal Processing, 3rd ed.; Prentice Hall: Saddle River, NJ, USA, 2011. [Google Scholar]
- Gustafsson, F. Determining the Initial States in Forward-Backward Filtering. IEEE Trans. Signal Process. 2002, 44, 988–992. [Google Scholar] [CrossRef]
- De Oliveira, M.A.; Da Rocha, A.M.; Puntel, F.E.; Cavalheiro, G.G.H. Time Series Compression for IoT: A Systematic Literature Review. Wirel. Commun. Mob. Comput. 2023, 2023, 5025255. [Google Scholar] [CrossRef]
- Smith, S.W. The Scientist and Engineer’s Guide to Digital Signal Processing, 2nd ed.; California Technical Pub.: San Diego, CA, USA, 1999. [Google Scholar]
- Chiarot, G.; Silvestri, C. Time Series Compression Survey. ACM Comput. Surv. 2023, 55, 198. [Google Scholar] [CrossRef]
- Bland, J.M.; Altman, D.G. Statistical methods for assessing agreement between two methods of clinical measurement. Lancet 1986, 327, 307–310. [Google Scholar] [CrossRef]
- O’Rourke, M.F.; Hartley, C.; McDonald, D.A. McDonald’s Blood Flow in Arteries: Theoretic, Experimental, and Clinical Principles; Arnold: London, UK, 1998. [Google Scholar]
- Magder, S. The highs and lows of blood pressure: Toward meaningful clinical targets in patients with shock. Crit. Care Med. 2014, 42, 1241–1251. [Google Scholar] [CrossRef]
- Waterfall, J.F. A review of the preclinical cardiovascular pharmacology of cilazapril, a new angiotensin converting enzyme inhibitor. Br. J. Clin. Pharmacol. 1989, 27, 139S–150S. [Google Scholar] [CrossRef] [PubMed]










| Parameter | Mean ± SD | Range | CV (%) | Unit |
|---|---|---|---|---|
| SBP | 52.79 ± 0.47 | 51.55–53.91 | 0.9 | mmHg |
| DBP | 51.26 ± 0.57 | 49.91–52.46 | 1.1 | mmHg |
| MAP | 51.77 ± 0.52 | 50.52–52.88 | 1.0 | mmHg |
| Step Pressure | 1.53 ± 0.23 | 0.93–2.22 | 15.0 | mmHg |
| Heart Rate | 274.5 ± 8.9 | 239–313 | 3.3 | bpm |
| Window | SBP (mmHg) | DBP (mmHg) | MAP (mmHg) | HR (bpm) |
|---|---|---|---|---|
| 1 | 52.63 | 51.04 | 51.57 | 275.9 |
| 2 | 52.60 | 51.06 | 51.57 | 274.5 |
| 3 | 52.35 | 50.68 | 51.24 | 274.7 |
| 4 | 53.01 | 51.58 | 52.05 | 274.2 |
| 5 | 53.37 | 51.96 | 52.43 | 273.0 |
| Parameter | Mean Bias (mmHg) | SD of Differences (mmHg) | Lower LoA (mmHg) | Upper LoA (mmHg) | Measurements Within LoA (%) | r (Pearson) | p-Value |
|---|---|---|---|---|---|---|---|
| SBP | −4.93 | 1.10 | −7.09 | −2.77 | 100% | 0.976 | <0.05 |
| DBP | +0.89 | 1.05 | −1.18 | +2.95 | 90% | 0.982 | <0.05 |
| MAP | −1.05 | 1.02 | −3.05 | +0.95 | 90% | 0.979 | <0.05 |
| Metric | BIOPAC PA (Mean ± SD) | Proposed System (All Beats, Mean ± SD) | Relative Error (%) | Proposed System (Session Mean ± SD, N = 10) | Range (Min–Max) |
|---|---|---|---|---|---|
| Detected beats (n) | 230 | 230 | — | — | — |
| SBP (mmHg) | 62.24 ± 1.46 | 56.90 ± 2.07 | −8.58 | 56.71 ± 1.74 | 54.14–58.41 |
| DBP (mmHg) | 54.00 ± 1.04 | 55.31 ± 2.21 | +2.43 | 55.10 ± 1.78 | 52.00–56.72 |
| MAP (mmHg) | 56.75 ± 1.17 | 55.84 ± 2.11 | −1.60 | 55.64 ± 1.72 | 52.72–57.28 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Garay, C.E.; Mansilla, G.N.; Madrid, R.E.; González Colombres, A.; Jerez, S.J. Design and Pilot Evaluation of an IoT-Based Blood Pressure Monitoring System for Rabbits. Bioengineering 2026, 13, 384. https://doi.org/10.3390/bioengineering13040384
Garay CE, Mansilla GN, Madrid RE, González Colombres A, Jerez SJ. Design and Pilot Evaluation of an IoT-Based Blood Pressure Monitoring System for Rabbits. Bioengineering. 2026; 13(4):384. https://doi.org/10.3390/bioengineering13040384
Chicago/Turabian StyleGaray, Carlos Exequiel, Gonzalo Nicolás Mansilla, Rossana Elena Madrid, Agustina González Colombres, and Susana Josefina Jerez. 2026. "Design and Pilot Evaluation of an IoT-Based Blood Pressure Monitoring System for Rabbits" Bioengineering 13, no. 4: 384. https://doi.org/10.3390/bioengineering13040384
APA StyleGaray, C. E., Mansilla, G. N., Madrid, R. E., González Colombres, A., & Jerez, S. J. (2026). Design and Pilot Evaluation of an IoT-Based Blood Pressure Monitoring System for Rabbits. Bioengineering, 13(4), 384. https://doi.org/10.3390/bioengineering13040384









