M5GP: Parallel Multidimensional Genetic Programming with Multidimensional Populations for Symbolic Regression
Abstract
1. Introduction
- Representation: the way in which solutions are represented (trees, graphs, arrays or stacks);
- Objectives: the number of objectives (single- or multi-objective), or the type of objectives used (error, correlation or novelty);
- Search process: how the method searches for or constructs a feasible model, including how the population is managed (for example, the use of single or multiple populations) or the type of search operators used (syntactic operators, semantic operators or hybrid techniques with local search methods);
- Search space: this is related to the set of feasible models that the technique can produce; i.e., the structure or the type of model than can be generated by the search process.
2. Background
2.1. M2GP: Multidimensional Multiclass Genetic Programming
2.2. M3GP: M2GP with Multidimensional Populations
2.3. M4GP: Incorporating a Stack-Based Representation
2.4. Related Work
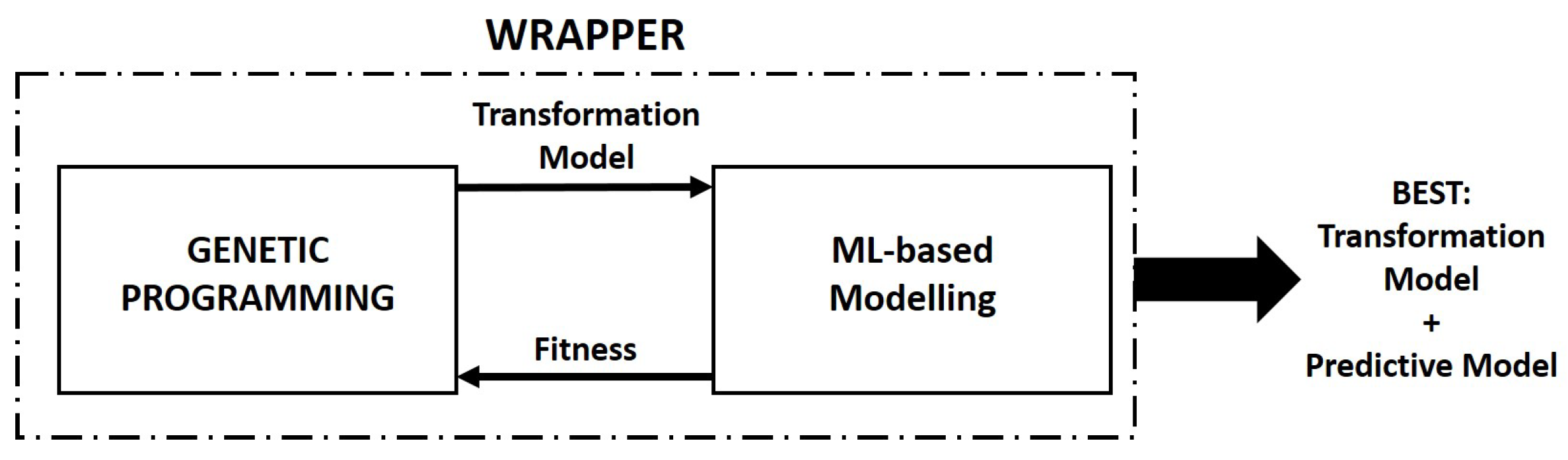
3. GPU-Based Parallel Implementation of M5GP
- The main evolutionary loop is controlled using Python code executed on the CPU but each of the main evolutionary processes are executed on the GPU, including population initialization, program evaluation done by an interpreter, selection and mutation;
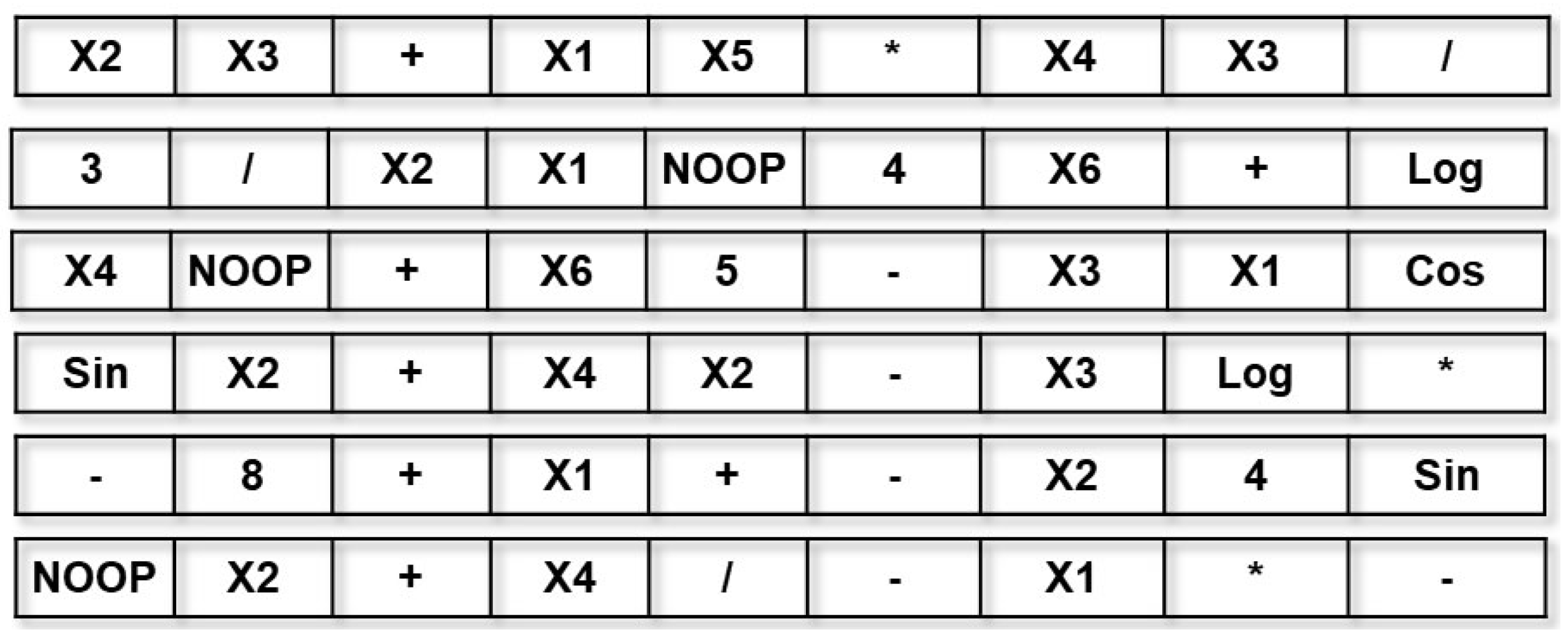
- Individuals are encoded using a linear representation, basically initialized as a list of randomly ordered primitives (terminals and functions). The individuals are considered to be transformation functions of the input feature space of a problem;
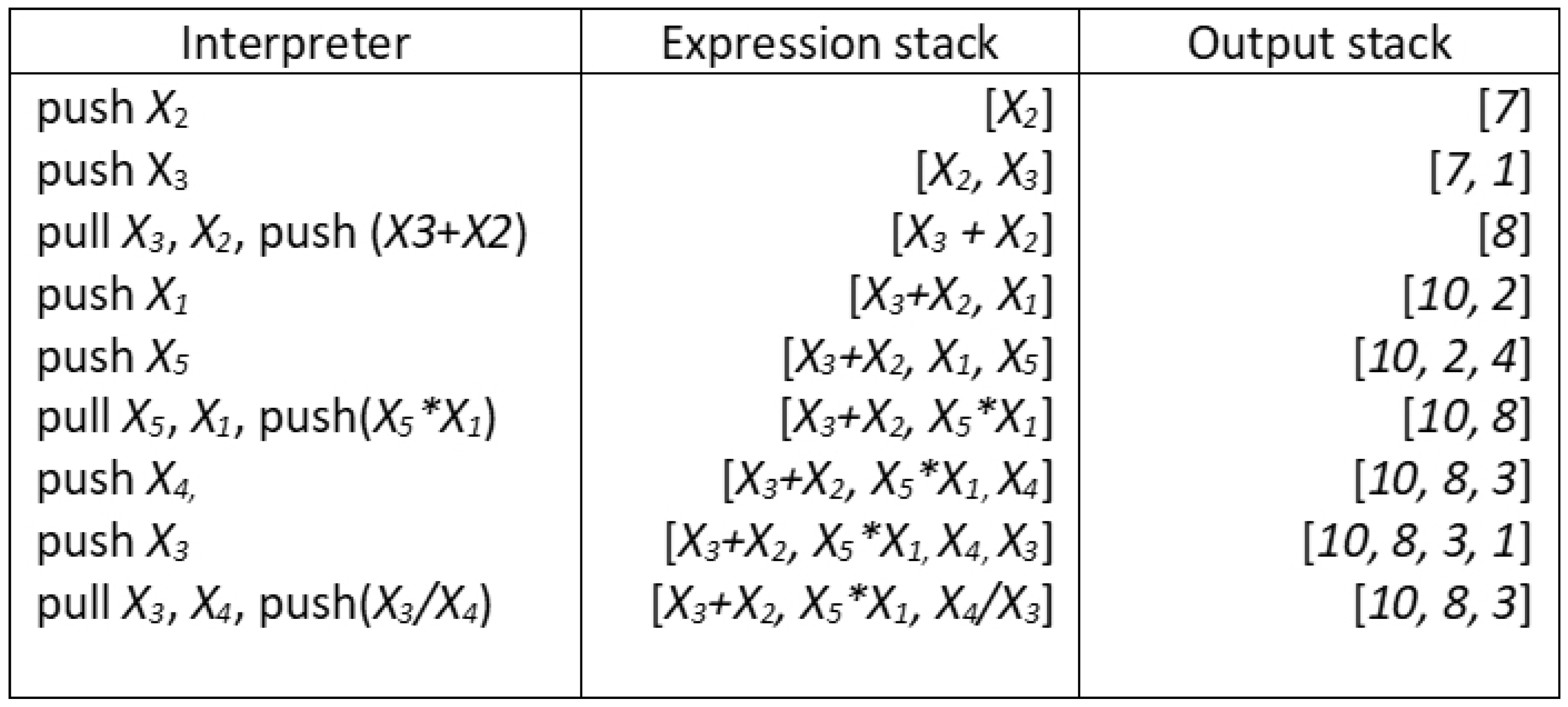
- Individuals are interpreted using a stack. Since most individuals will encode a set of sub-expressions, they tend to produce stacks of different sizes, the transformed feature space of the training data;
- The mutation operator used by M5GP is based on the Uniform Mutation by Addition and Deletion operator (UMAD) [25] and selection is performed using tournaments. The former was chosen because it has been shown to produce strong results with stack-based GP, while the latter was chosen instead of more advanced methods (such as lexicase selection [44] used by M4GP) for simplicity and efficiency purposes;
- All of the evolutionary processes in M5GP are implemented as GPU kernels using Numba, except for parameter estimation which is carried out using linear regression with cuML;
- Linear models are constructed using the output stacks of each individual using cuML, and the Root Mean Squared Error (RMSE) of the model is returned as fitness;
- M5GP returns the best feature transformation model and the corresponding linear model fitted using the cuML library;
- For ease of use, M5GP implements the scikit-learn API, allowing it to be evaluated on SRBench [4].
3.1. Individual Representation, Interpretation and Parallel Processing
3.2. Fitness Evaluation Using CUDA-Based Machine Learning
3.3. Search Operators
4. Experiments
4.1. Experimental Setup
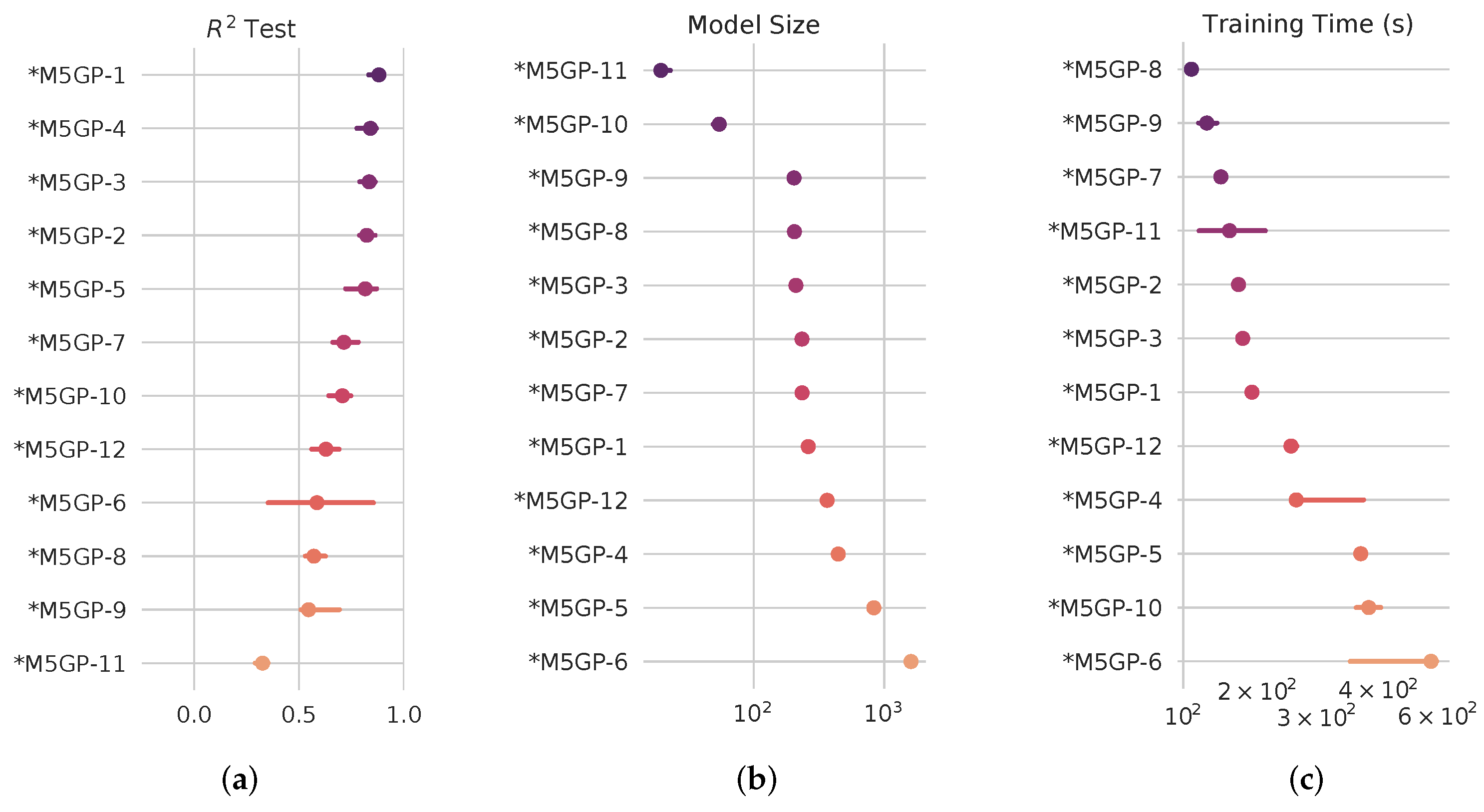
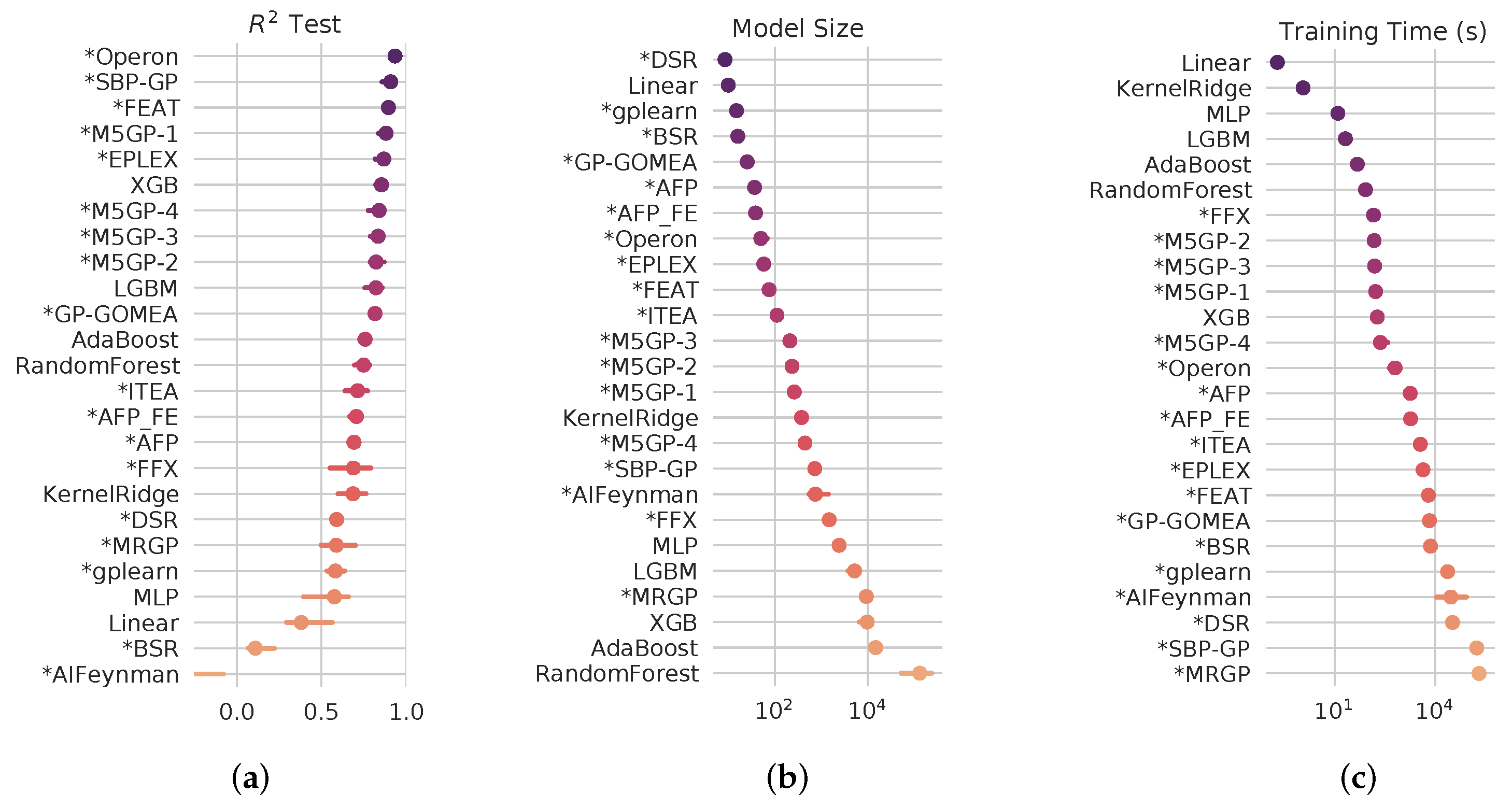
4.2. SRBench Results
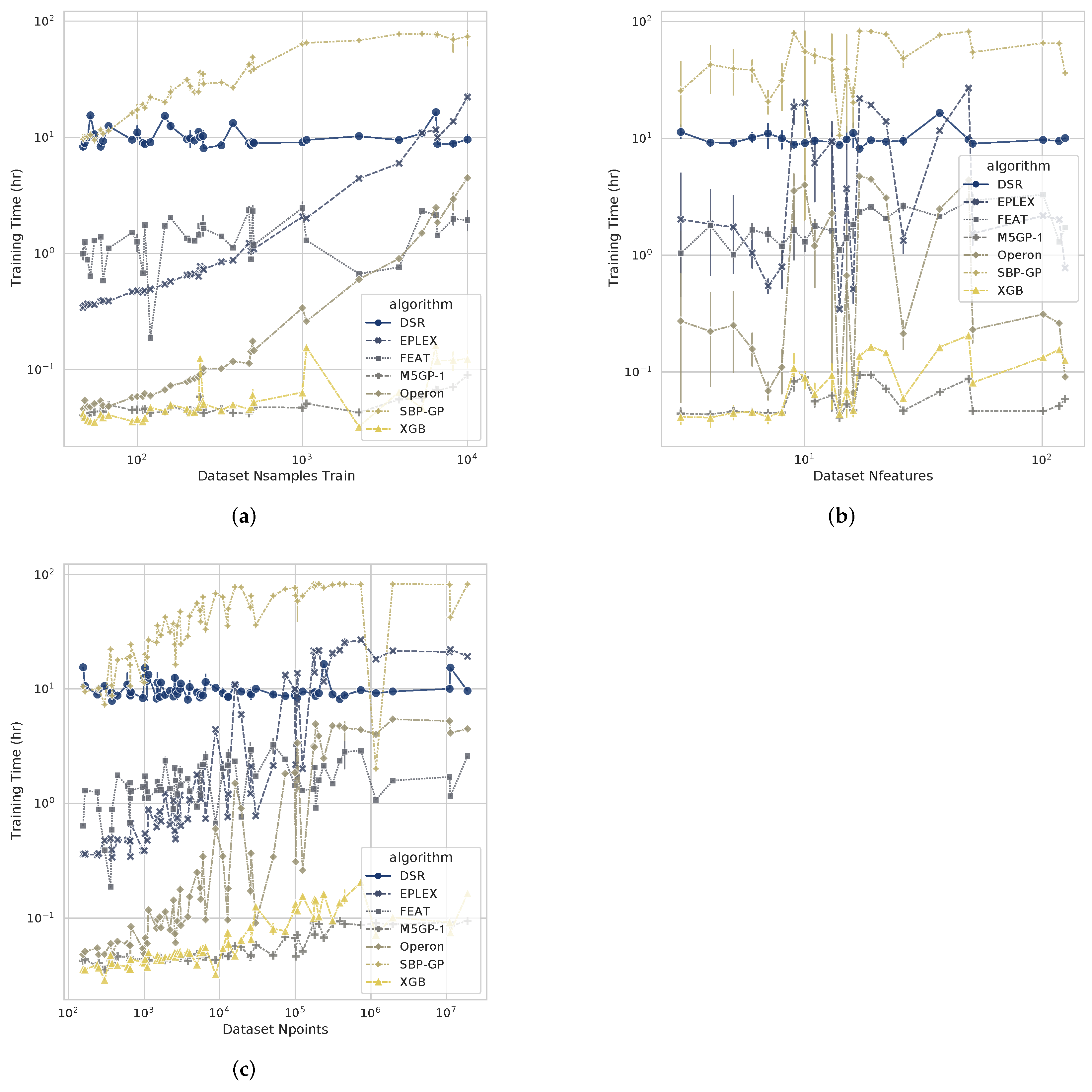
4.3. Computational Cost
5. Conclusions and Future Work
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Koza, J.R. Genetic Programming; Complex Adaptive Systems, Bradford Books: Cambridge, MA, USA, 1992. [Google Scholar]
- Koza, J.R. Human-competitive results produced by genetic programming. Genet. Program. Evolvable Mach. 2010, 11, 251–284. [Google Scholar] [CrossRef]
- Orzechowski, P.; La Cava, W.; Moore, J.H. Where Are We Now? A Large Benchmark Study of Recent Symbolic Regression Methods. In Proceedings of the GECCO ’18: Genetic and Evolutionary Computation Conference, Kyoto, Japan, 15–19 July 2018; Association for Computing Machinery: New York, NY, USA, 2018; pp. 1183–1190. [Google Scholar]
- La Cava, W.; Orzechowski, P.; Burlacu, B.; de Franca, F.; Virgolin, M.; Jin, Y.; Kommenda, M.; Moore, J. Contemporary Symbolic Regression Methods and Their Relative Performance. arXiv 2021, arXiv:2107.14351. [Google Scholar]
- Rudin, C. Stop explaining black box machine learning models for high stakes decisions and use interpretable models instead. Nat. Mach. Intell. 2019, 1, 206–215. [Google Scholar] [CrossRef] [PubMed]
- Spector, L.; Robinson, A. Genetic Programming and Autoconstructive Evolution with the Push Programming Language. Genet. Program. Evolvable Mach. 2002, 3, 7–40. [Google Scholar] [CrossRef]
- Burlacu, B.; Kronberger, G.; Kommenda, M. Operon C++: An Efficient Genetic Programming Framework for Symbolic Regression. In Proceedings of the GECCO ’20: 2020 Genetic and Evolutionary Computation Conference Companion, Cancún, Mexico, 8–12 July 2020; Association for Computing Machinery: New York, NY, USA, 2020; pp. 1562–1570. [Google Scholar]
- Arnaldo, I.; Krawiec, K.; O’Reilly, U.M. Multiple Regression Genetic Programming. In Proceedings of the GECCO ’14: 2014 Annual Conference on Genetic and Evolutionary Computation, Vancouver, BC, Canada, 12–16 July 2014; Association for Computing Machinery: New York, NY, USA, 2014; pp. 879–886. [Google Scholar]
- Muñoz, L.; Silva, S.; Trujillo, L. M3GP—Multiclass Classification with GP. In Lecture Notes in Computer Science; Springer International Publishing: Berlin/Heidelberg, Germany, 2015; pp. 78–91. [Google Scholar]
- Moraglio, A.; Krawiec, K.; Johnson, C.G. Geometric Semantic Genetic Programming. In Proceedings of the PPSN’12: 12th International Conference on Parallel Problem Solving from Nature—Volume Part I, Taormina, Italy, 1–5 September 2012; pp. 21–31. [Google Scholar]
- Muñoz, L.; Trujillo, L.; Silva, S. Transfer learning in constructive induction with Genetic Programming. Genet. Program. Evolvable Mach. 2019, 21, 529–569. [Google Scholar] [CrossRef]
- Montgomery, D.C.; Peck, E.A.; Vining, G.G. Introduction to Linear Regression Analysis, 6th ed.; Wiley Series in Probability and Statistics; John Wiley & Sons: Nashville, TN, USA, 2021. [Google Scholar]
- Rudin, C. Do Simpler Models Exist and How Can We Find Them? In Proceedings of the KDD ’19: 25th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, Anchorage, AK, USA, 4–8 August 2019; pp. 1–2. [Google Scholar]
- Tian, Y.; Zhang, Y. A comprehensive survey on regularization strategies in machine learning. Inf. Fusion 2022, 80, 146–166. [Google Scholar] [CrossRef]
- Trujillo, L.; Z-Flores, E.; Juárez-Smith, P.S.; Legrand, P.; Silva, S.; Castelli, M.; Vanneschi, L.; Schütze, O.; Muñoz, L. Local Search is Underused in Genetic Programming. In Genetic and Evolutionary Computation; Springer International Publishing: Berlin/Heidelberg, Germany, 2018; pp. 119–137. [Google Scholar]
- Iba, H. Inference of differential equation models by genetic programming. Inf. Sci. 2008, 178, 4453–4468. [Google Scholar] [CrossRef]
- Pan, I.; Das, S. When Darwin meets Lorenz: Evolving new chaotic attractors through genetic programming. Chaos Solitons Fractals 2015, 76, 141–155. [Google Scholar] [CrossRef]
- Falco, I.; Cioppa, A.; Tarantino, E. A Genetic Programming System for Time Series Prediction and Its Application to El Niño Forecast. In Advances in Soft Computing; Springer-Verlag: Berlin/Heidelberg, Germany, 1999; pp. 151–162. [Google Scholar]
- Arfken, G.B.; Weber, H.J.; Harris, F.E. Mathematical Methods for Physicists, 6th ed.; Academic Press: San Diego, CA, USA, 2005. [Google Scholar]
- McConaghy, T. FFX: Fast, Scalable, Deterministic Symbolic Regression Technology. In Genetic and Evolutionary Computation; Springer: New York, NY, USA, 2011; pp. 235–260. [Google Scholar]
- Cranmer, M. Interpretable Machine Learning for Science with PySR and SymbolicRegression.jl. arXiv 2023, arXiv:2305.01582. [Google Scholar]
- Ingalalli, V.; Silva, S.; Castelli, M.; Vanneschi, L. A Multi-dimensional Genetic Programming Approach for Multi-class Classification Problems. In Lecture Notes in Computer Science; Springer: Berlin/Heidelberg, Germany, 2014; pp. 48–60. [Google Scholar]
- Cava, W.L.; Silva, S.; Danai, K.; Spector, L.; Vanneschi, L.; Moore, J.H. Multidimensional genetic programming for multiclass classification. Swarm Evol. Comput. 2019, 44, 260–272. [Google Scholar] [CrossRef]
- Muñoz, L.; Trujillo, L.; Silva, S.; Castelli, M.; Vanneschi, L. Evolving multidimensional transformations for symbolic regression with M3GP. Memetic Comput. 2018, 11, 111–126. [Google Scholar] [CrossRef]
- Helmuth, T.; McPhee, N.F.; Spector, L. Program Synthesis Using Uniform Mutation by Addition and Deletion. In Proceedings of the GECCO ’18: Genetic and Evolutionary Computation Conference, Kyoto, Japan, 15–19 July 2018; Association for Computing Machinery: New York, NY, USA, 2018; pp. 1127–1134. [Google Scholar]
- Trujillo, L.; Muñoz Contreras, J.M.; Hernandez, D.E.; Castelli, M.; Tapia, J.J. GSGP-CUDA—A CUDA framework for Geometric Semantic Genetic Programming. SoftwareX 2022, 18, 101085. [Google Scholar] [CrossRef]
- Raschka, S.; Patterson, J.; Nolet, C. Machine learning in Python: Main developments and technology trends in data science, machine learning, and artificial intelligence. Information 2020, 11, 193. [Google Scholar] [CrossRef]
- Buitinck, L.; Louppe, G.; Blondel, M.; Pedregosa, F.; Mueller, A.; Grisel, O.; Niculae, V.; Prettenhofer, P.; Gramfort, A.; Grobler, J.; et al. API design for machine learning software: Experiences from the scikit-learn project. arXiv 2013, arXiv:1309.0238. [Google Scholar]
- Chandrashekar, G.; Sahin, F. A Survey on Feature Selection Methods. Comput. Electr. Eng. 2014, 40, 16–28. [Google Scholar] [CrossRef]
- Cava, W.L.; Singh, T.R.; Taggart, J.; Suri, S.; Moore, J.H. Learning Concise Representations for Regression by Evolving Networks of Trees. arXiv 2019, arXiv:1807.00981. [Google Scholar]
- Altarabichi, M.G.; Nowaczyk, S.; Pashami, S.; Mashhadi, P.S. Fast Genetic Algorithm for feature selection—A qualitative approximation approach. Expert Syst. Appl. 2023, 211, 118528. [Google Scholar] [CrossRef]
- Liao, L.; Pindur, A.K.; Iba, H. Genetic Programming with Random Binary Decomposition for Multi-Class Classification Problems. In Proceedings of the 2021 IEEE Congress on Evolutionary Computation (CEC), Krakow, Poland, 28 June–1 July 2021; IEEE: New York, NY, USA, 2021. [Google Scholar]
- Viegas, F.; Rocha, L.; Gonçalves, M.; Mourão, F.; Sá, G.; Salles, T.; Andrade, G.; Sandin, I. A Genetic Programming approach for feature selection in highly dimensional skewed data. Neurocomputing 2018, 273, 554–569. [Google Scholar] [CrossRef]
- Espejo, P.; Ventura, S.; Herrera, F. A Survey on the Application of Genetic Programming to Classification. IEEE Trans. Syst. Man Cybern. C Appl. Rev. 2010, 40, 121–144. [Google Scholar] [CrossRef]
- Z-Flores, E.; Trujillo, L.; Schütze, O.; Legrand, P. A Local Search Approach to Genetic Programming for Binary Classification. In Proceedings of the GECCO ’15: 2015 Annual Conference on Genetic and Evolutionary Computation, Madrid, Spain, 11–15 July 2015; Association for Computing Machinery: New York, NY, USA, 2015; pp. 1151–1158. [Google Scholar]
- Langdon, W.B. Failed Disruption Propagation in Integer Genetic Programming. In Proceedings of the GECCO ’22: Genetic and Evolutionary Computation Conference Companion, Boston, MA, USA, 9–13 July 2022; Association for Computing Machinery: New York, NY, USA, 2022; pp. 574–577. [Google Scholar]
- Batista, J.E.; Silva, S. Comparative study of classifier performance using automatic feature construction by M3GP. In Proceedings of the 2022 IEEE Congress on Evolutionary Computation (CEC), Padua, Italiy, 18–23 July 2022; IEEE: New York, NY, USA, 2022. [Google Scholar]
- Chen, T.; Guestrin, C. XGBoost: A Scalable Tree Boosting System. In Proceedings of the KDD ’16: 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco, CA, USA, 13–17 August 2016; Association for Computing Machinery: New York, NY, USA, 2016; pp. 785–794. [Google Scholar]
- Batista, J.E.; Cabral, A.I.R.; Vasconcelos, M.J.P.; Vanneschi, L.; Silva, S. Improving Land Cover Classification Using Genetic Programming for Feature Construction. Remote Sens. 2021, 13, 1623. [Google Scholar] [CrossRef]
- Yang, Y.; Wang, X.; Zhao, X.; Huang, M.; Zhu, Q. M3GPSpectra: A novel approach integrating variable selection/construction and MLR modeling for quantitative spectral analysis. Anal. Chim. Acta 2021, 1160, 338453. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; Yang, Y.; Zhang, G.; Xu, L.; Wang, M. EBM3GP: A novel evolutionary bi-objective genetic programming for dimensionality reduction in classification of hyperspectral data. Infrared Phys. Technol. 2023, 129, 104577. [Google Scholar] [CrossRef]
- Langdon, W.B. Graphics processing units and genetic programming: An overview. Soft Comput. 2011, 15, 1657–1669. [Google Scholar] [CrossRef]
- Chitty, D.M. Faster GPU-based genetic programming using a two-dimensional stack. Soft Comput. 2016, 21, 3859–3878. [Google Scholar] [CrossRef][Green Version]
- Spector, L. Assessment of Problem Modality by Differential Performance of Lexicase Selection in Genetic Programming: A Preliminary Report. In Proceedings of the GECCO ’12: 14th Annual Conference Companion on Genetic and Evolutionary Computation, Philadelphia, PA, USA, 7– 11 July 2012; Association for Computing Machinery: New York, NY, USA, 2012; pp. 401–408. [Google Scholar]
- Schmidt, M.D.; Lipson, H. Age-Fitness Pareto Optimization. In Proceedings of the GECCO ’10: 12th Annual Conference on Genetic and Evolutionary Computation, Cancun, Mexico, 8–12 July 2020; Association for Computing Machinery: New York, NY, USA, 2010; pp. 543–544. [Google Scholar]
- Olson, R.S.; Bartley, N.; Urbanowicz, R.J.; Moore, J.H. Evaluation of a Tree-based Pipeline Optimization Tool for Automating Data Science. In Proceedings of the GECCO ’16: Genetic and Evolutionary Computation Conference 2016, Denver, CO, USA, 20–24 July 2016; Association for Computing Machinery: New York, NY, USA, 2016; pp. 485–492. [Google Scholar]
- McDermott, J.; White, D.R.; Luke, S.; Manzoni, L.; Castelli, M.; Vanneschi, L.; Jaskowski, W.; Krawiec, K.; Harper, R.; De Jong, K.; et al. Genetic Programming Needs Better Benchmarks. In Proceedings of the GECCO ’12: 14th Annual Conference on Genetic and Evolutionary Computation, Philadelphia, PA, USA, 7–11 July 2012; Association for Computing Machinery: New York, NY, USA, 2012; pp. 791–798. [Google Scholar]
- McDermott, J.; Kronberger, G.; Orzechowski, P.; Vanneschi, L.; Manzoni, L.; Kalkreuth, R.; Castelli, M. Genetic Programming Benchmarks: Looking Back and Looking Forward. SIGEVOlution 2022, 15, 1–19. [Google Scholar] [CrossRef]
- Crary, C.; Piard, W.; Stitt, G.; Bean, C.; Hicks, B. Using FPGA Devices to Accelerate Tree-Based Genetic Programming: A Preliminary Exploration with Recent Technologies. In Lecture Notes in Computer Science; Springer Nature: Cham, Switzerland, 2023; pp. 182–197. [Google Scholar]
- Virgolin, M.; Alderliesten, T.; Bosman, P.A.N. Linear Scaling with and within Semantic Backpropagation-Based Genetic Programming for Symbolic Regression. In Proceedings of the GECCO ’19: Genetic and Evolutionary Computation Conference, Prague, Czech Republic, 13–17 July 2019; Association for Computing Machinery: New York, NY, USA; pp. 1084–1092. [Google Scholar]
- Melo, V.V.D. Kaizen programming. In Proceedings of the 2014 Annual Conference on Genetic and Evolutionary Computation, Vancouver, BC, Canada, 12–16 July 2014; ACM: New York, NY, USA, 2014. [Google Scholar]
- de Franca, F.O.; Aldeia, G.S.I. Interaction–Transformation Evolutionary Algorithm for Symbolic Regression. Evol. Comput. 2021, 29, 367–390. [Google Scholar] [CrossRef] [PubMed]
- Lam, S.K.; Pitrou, A.; Seibert, S. Numba: A llvm-based python jit compiler. In Proceedings of the Second Workshop on the LLVM Compiler Infrastructure in HPC, Austin, TX, USA, 15 November 2015; pp. 1–6. [Google Scholar]
- Ni, J.; Drieberg, R.H.; Rockett, P.I. The Use of an Analytic Quotient Operator in Genetic Programming. IEEE Trans. Evol. Comput. 2012, 17, 146–152. [Google Scholar] [CrossRef]
- Okuta, R.; Unno, Y.; Nishino, D.; Hido, S.; Loomis, C. CuPy: A NumPy-Compatible Library for NVIDIA GPU Calculations. In Proceedings of the 31st Conference on Neural Information Processing Systems (NIPS 2017), Long Beach, CA, USA, 4–9 December 2017. [Google Scholar]
- Olson, R.S.; La Cava, W.; Orzechowski, P.; Urbanowicz, R.J.; Moore, J.H. PMLB: A large benchmark suite for machine learning evaluation and comparison. BioData Min. 2017, 10, 36. [Google Scholar] [CrossRef]
- Schapire, R.E. Explaining adaboost. In Empirical Inference; Springer: Berlin/Heidelberg, Germany, 2013; pp. 37–52. [Google Scholar]
- Ho, T.K. Random decision forests. In Proceedings of the 3rd International Conference on Document Analysis and Recognition, Montreal, QC, Canada, 14–15 August 1995; IEEE: New York, NY, USA, 1995; Volume 1, pp. 278–282. [Google Scholar]
- Virgolin, M.; Alderliesten, T.; Witteveen, C.; Bosman, P.A.N. Scalable Genetic Programming by Gene-Pool Optimal Mixing and Input-Space Entropy-Based Building-Block Learning. In Proceedings of the GECCO ’17: Genetic and Evolutionary Computation Conference, Berlin, Germany, 15–19 July 2017; Association for Computing Machinery: New York, NY, USA, 2017; pp. 1041–1048. [Google Scholar]
- Petersen, B.K.; Landajuela, M.; Mundhenk, T.N.; Santiago, C.P.; Kim, S.K.; Kim, J.T. Deep symbolic regression: Recovering mathematical expressions from data via risk-seeking policy gradients. In Proceedings of the International Conference on Learning Representations, Virtual Only Conference, 3–7 May 2021. [Google Scholar]
- Sipper, M.; Fu, W.; Ahuja, K.; Moore, J.H. Investigating the parameter space of evolutionary algorithms. BioData Min. 2018, 11, 2. [Google Scholar] [CrossRef]
- Trujillo, L.; Álvarez González, E.; Galván, E.; Tapia, J.J.; Ponsich, A. On the Analysis of Hyper-Parameter Space for a Genetic Programming System with Iterated F-Race. Soft Comput. 2020, 24, 14757–14770. [Google Scholar] [CrossRef]
- Brookhouse, J.; Otero, F.E.; Kampouridis, M. Working with OpenCL to Speed up a Genetic Programming Financial Forecasting Algorithm: Initial Results. In Proceedings of the GECCO Comp’14: Companion Publication of the 2014 Annual Conference on Genetic and Evolutionary Computation, Vancouver, BC, Canada, 12–16 July 2014; Association for Computing Machinery: New York, NY, USA, 2014; pp. 1117–1124. [Google Scholar]
- Kamienny, P.-A.; d’Ascoli, S.; Lample, G.; Charton, F. End-to-end Symbolic Regression with Transformers. In Advances in Neural Information Processing Systems; Koyejo, S., Mohamed, S., Agarwal, A., Belgrave, D., Cho, K., Oh, A., Eds.; Curran Associates, Inc.: New York, NY, USA, 2022; Volume 35, pp. 10269–10281. [Google Scholar]
- Zhang, R.; Lensen, A.; Sun, Y. Speeding up Genetic Programming Based Symbolic Regression Using GPUs. In Proceedings of the PRICAI 2022: Trends in Artificial Intelligence, Shanghai, China, 10–13 November 2022; Khanna, S., Cao, J., Bai, Q., Xu, G., Eds.; Springer Nature: Cham, Switzerland, 2022; pp. 519–533. [Google Scholar]
- Holt, S.; Qian, Z.; van der Schaar, M. Deep Generative Symbolic Regression. In Proceedings of the The Eleventh International Conference on Learning Representations, Kigali, Rwanda, 1–5 May 2023. [Google Scholar]





| Identifier | Regression | Tuning | Population | Genes | Fitness |
|---|---|---|---|---|---|
| M5GP-1 | LR | Yes | 256 | 128 | RMSE |
| M5GP-2 | LR | No | 256 | 128 | RMSE |
| M5GP-3 | LR-n | No | 256 | 128 | RMSE |
| M5GP-4 | LR | No | 256 | 256 | RMSE |
| M5GP-5 | LR | No | 256 | 512 | RMSE |
| M5GP-6 | LR | No | 512 | 256 | RMSE |
| M5GP-7 | LR | No | 256 | 128 | |
| M5GP-8 | LR-m | No | 256 | 128 | RMSE |
| M5GP-9 | LASSO | No | 256 | 128 | RMSE |
| M5GP-10 | LASSO | No | 512 | 256 | RMSE |
| M5GP-11 | EN | No | 256 | 128 | RMSE |
| M5GP-12 | EN | No | 512 | 256 | RMSE |
| Generations | Population | Genes | Tournament Size |
|---|---|---|---|
| 30 | 128 | 128 | 0.25 |
| 30 | 128 | 128 | 0.15 |
| 30 | 256 | 128 | 0.1 |
| 30 | 256 | 256 | 0.1 |
| 50 | 128 | 128 | 0.25 |
| 50 | 128 | 128 | 0.15 |
| Hyperparameter | Value | Description |
|---|---|---|
| Generations | 30 | Iterations in the evolutionary loop |
| Function set | Applied at the gene level | |
| Terminal set | constants and features | Constants are in the range of |
| Replacement rate | 0.1 | Applied at the gene level |
| Delete rate | 0.01 | Applied at the gene level |
| Tournament size | 0.15 | Percentage of the population that will be included in the tournament |
| Gene function probability | 0.50 | Probability to choose a function during mutation or population initialization |
| Gene feature probability | 0.39 | Probability to choose a feature during mutation or population initialization |
| Gene constant probability | 0.1 | Probability to choose a constant during mutation or population initialization |
| Gene NOOP probability | 0.01 | Probability to choose a NOOP during mutation or population initialization |
| Identifier | Model Size | Training Time (s) | |
|---|---|---|---|
| M5GP-1 | 0.880 | 261.75 | 166.77 |
| M5GP-2 | 0.822 | 234 | 150.89 |
| M5GP-3 | 0.834 | 210.5 | 155.66 |
| M5GP-4 | 0.840 | 444 | 231.81 |
| M5GP-5 | 0.815 | 831 | 374.51 |
| M5GP-6 | 0.584 | 1600 | 631.66 |
| M5GP-7 | 0.713 | 234.5 | 132.32 |
| M5GP-8 | 0.570 | 205 | 106.28 |
| M5GP-9 | 0.544 | 203.5 | 119.18 |
| M5GP-10 | 0.706 | 54.5 | 397.40 |
| M5GP-11 | 0.325 | 19.5 | 141.09 |
| M5GP-12 | 0.627 | 365 | 222.92 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cárdenas Florido, L.; Trujillo, L.; Hernandez, D.E.; Muñoz Contreras, J.M. M5GP: Parallel Multidimensional Genetic Programming with Multidimensional Populations for Symbolic Regression. Math. Comput. Appl. 2024, 29, 25. https://doi.org/10.3390/mca29020025
Cárdenas Florido L, Trujillo L, Hernandez DE, Muñoz Contreras JM. M5GP: Parallel Multidimensional Genetic Programming with Multidimensional Populations for Symbolic Regression. Mathematical and Computational Applications. 2024; 29(2):25. https://doi.org/10.3390/mca29020025
Chicago/Turabian StyleCárdenas Florido, Luis, Leonardo Trujillo, Daniel E. Hernandez, and Jose Manuel Muñoz Contreras. 2024. "M5GP: Parallel Multidimensional Genetic Programming with Multidimensional Populations for Symbolic Regression" Mathematical and Computational Applications 29, no. 2: 25. https://doi.org/10.3390/mca29020025
APA StyleCárdenas Florido, L., Trujillo, L., Hernandez, D. E., & Muñoz Contreras, J. M. (2024). M5GP: Parallel Multidimensional Genetic Programming with Multidimensional Populations for Symbolic Regression. Mathematical and Computational Applications, 29(2), 25. https://doi.org/10.3390/mca29020025







