Salivary miR-30c-5p as Potential Biomarker for Detection of Oral Squamous Cell Carcinoma
Abstract
:1. Introduction
2. Materials and Methods
2.1. Patient Cohort and Saliva Collection
2.2. Total RNA Extraction and miRNA TaqMan Assay
2.3. HPV Detection in Tumor Specimens
2.4. Discrimination Power Analysis
2.5. TCGA Dataset of OSCC
2.6. miR-30c-5p Target Prediction and Pathway Enrichment Analysis
2.7. Survival Analyses
3. Results
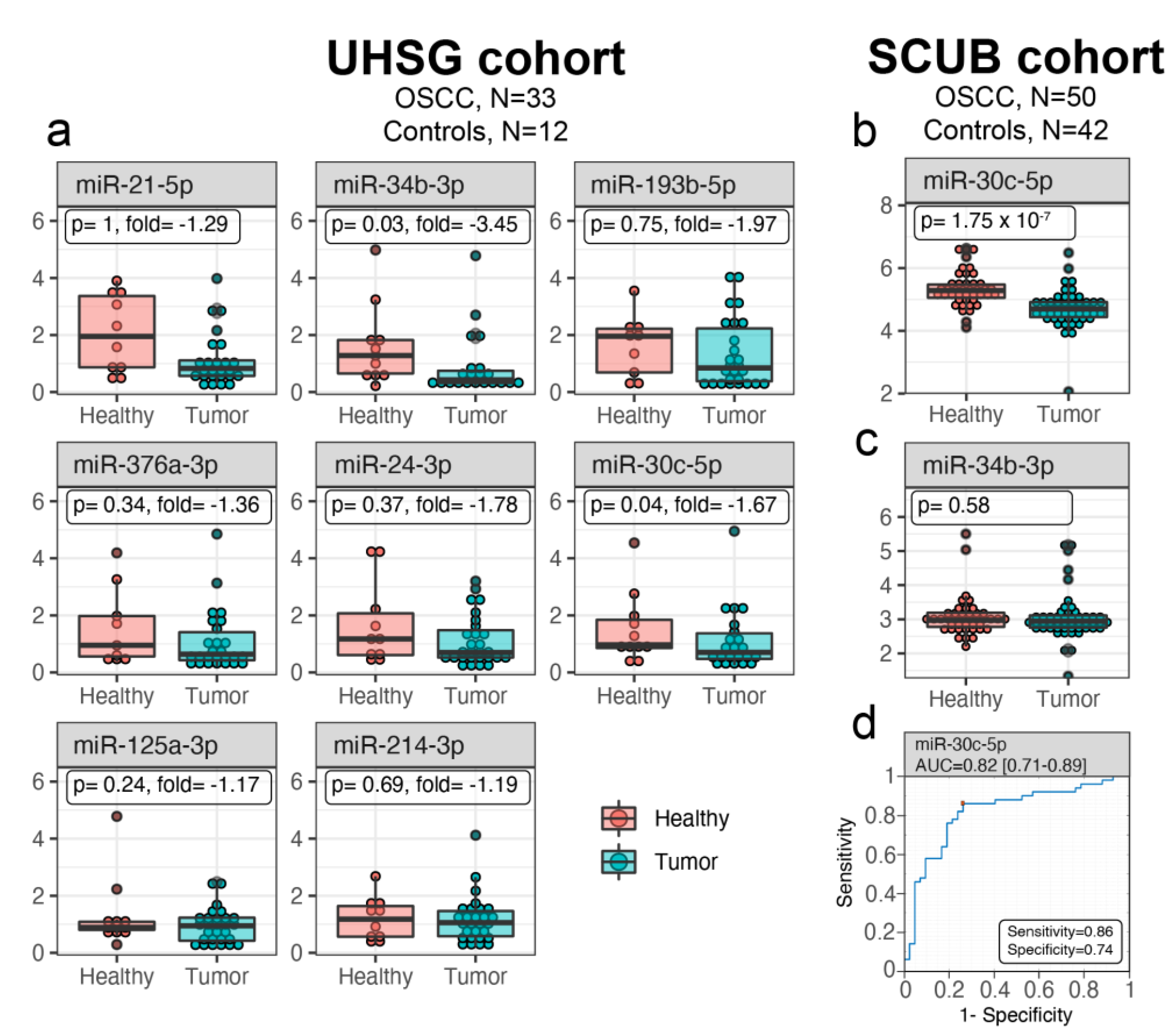
3.1. miR-30c-5p Is Weakly Expressed in OSCC Saliva
3.2. Salivary miR-30c-5p Has the Potential to Discriminate OSCC Patients from Healthy Controls
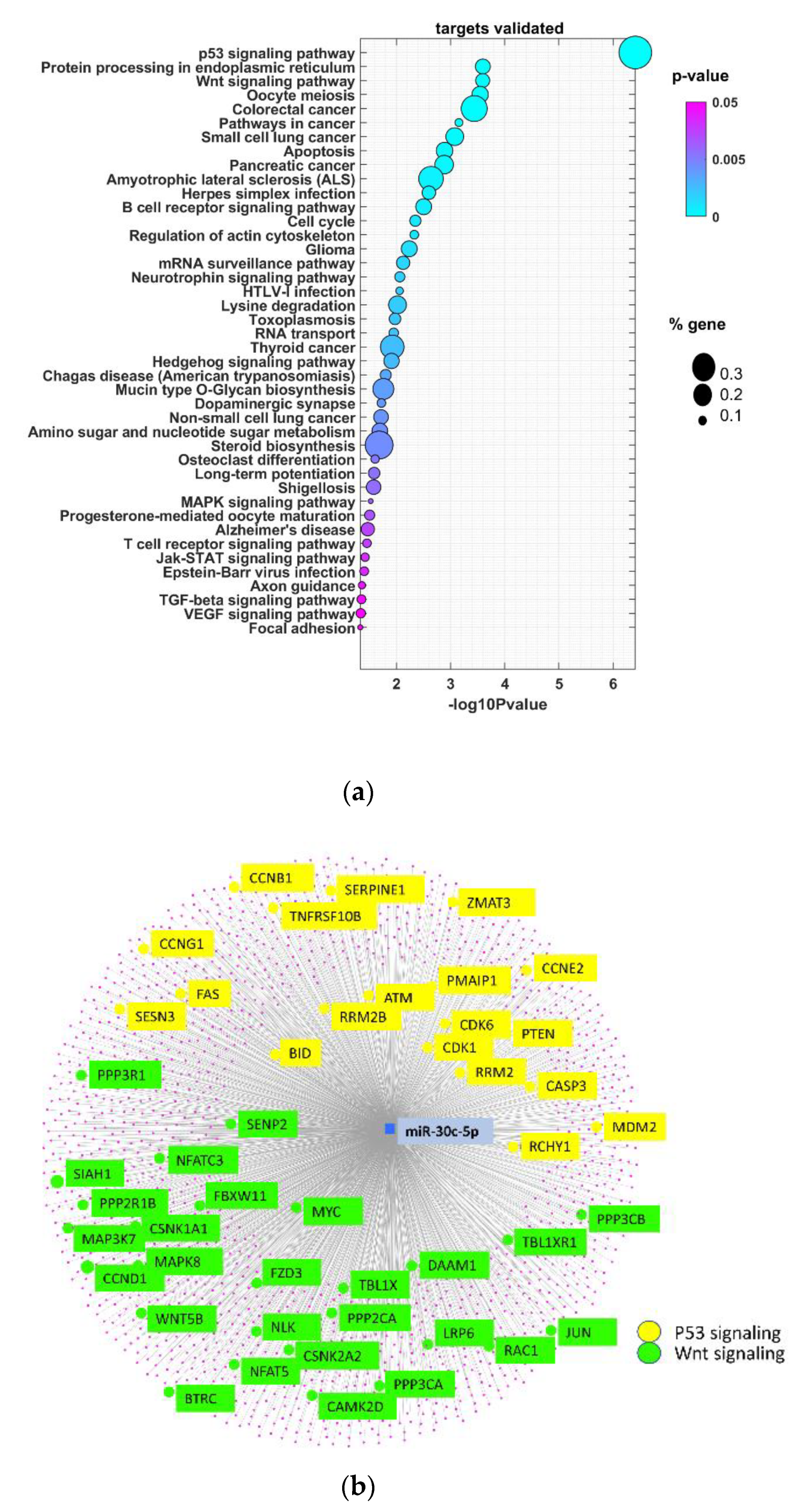
3.3. miR-30c-5p Target Genes Are Involved in p53 and Wnt Signaling Pathways in OSCC
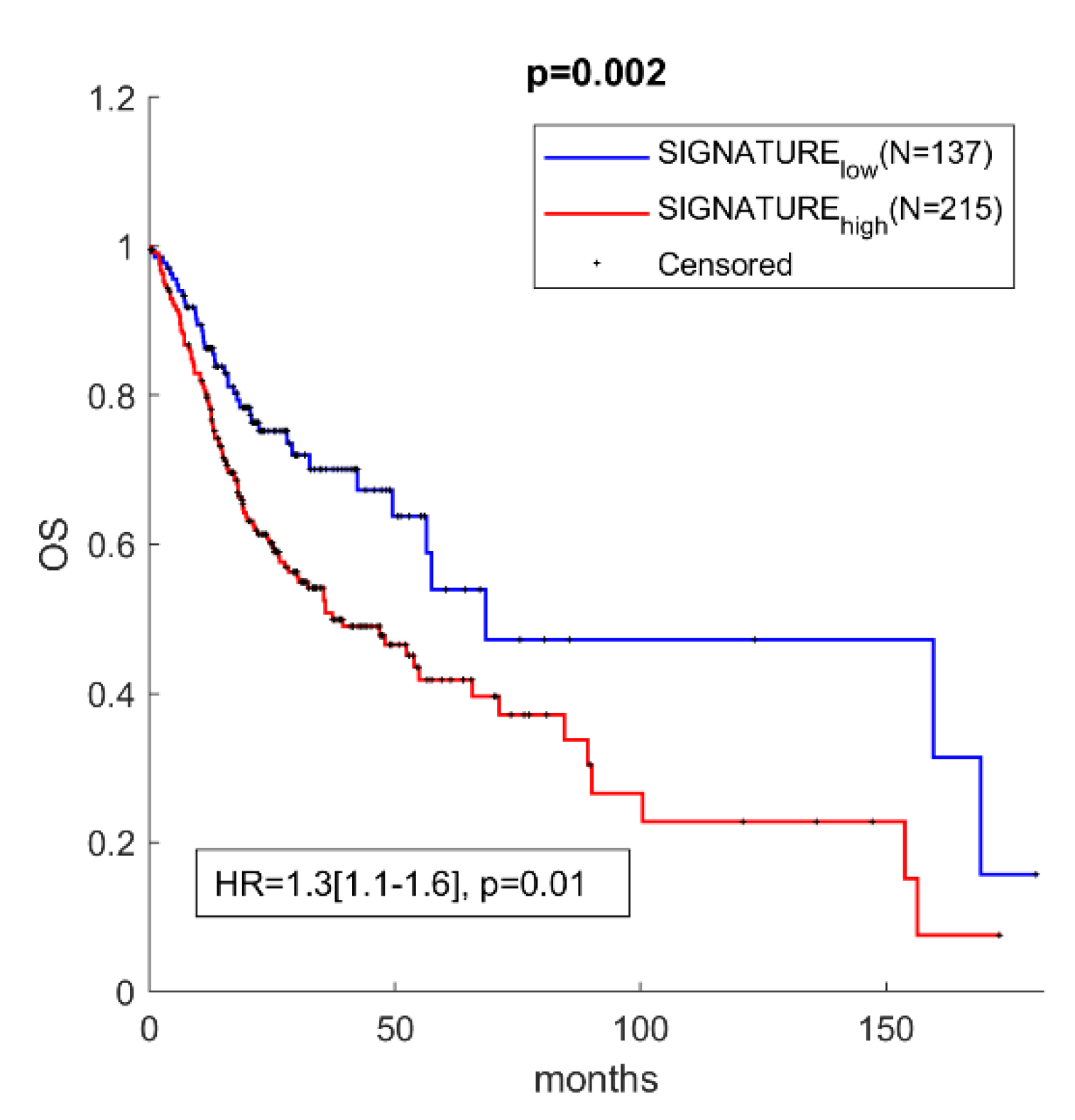
3.4. miR-30c-5p Targets Have Prognostic Value in OSCC
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ferlay, J.; Colombet, M.; Soerjomataram, I.; Mathers, C.; Parkin, D.M.; Pineros, M.; Znaor, A.; Bray, F. Estimating the global cancer incidence and mortality in 2018: GLOBOCAN sources and methods. Int. J. Cancer 2019, 144, 1941–1953. [Google Scholar] [CrossRef] [Green Version]
- Funk, G.F.; Karnell, L.H.; Robinson, R.A.; Zhen, W.K.; Trask, D.K.; Hoffman, H.T. Presentation, treatment, and outcome of oral cavity cancer: A National Cancer Data Base report. Head Neck 2002, 24, 165–180. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Zhou, X.; St John, M.A.; Wong, D.T.W. RNA profiling of cell-free saliva using microarray technology. J. Dent. Res. 2004, 83, 199–203. [Google Scholar] [CrossRef]
- St John, M.A.R.; Li, Y.; Zhou, X.; Denny, P.; Ho, C.M.; Montemagno, C.; Shi, W.; Qi, F.; Wu, B.; Sinha, U.; et al. Interleukin 6 and interleukin 8 as potential biomarkers for oral cavity and oropharyngeal squamous cell carcinoma. Arch. Otolaryngol Head Neck Surg. 2004, 130, 929–935. [Google Scholar] [CrossRef] [Green Version]
- Zanoni, D.K.; Montero, P.H.; Migliacci, J.C.; Shah, J.P.; Wong, R.J.; Ganly, I.; Patel, S.G. Survival outcomes after treatment of cancer of the oral cavity (1985–2015). Oral Oncol. 2019, 90, 115–121. [Google Scholar] [CrossRef]
- Panta, P.; Venna, V.R. Salivary RNA signatures in oral cancer detection. Anal. Cell Pathol. 2014, 2014, 450629. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zahran, F.; Ghalwash, D.; Shaker, O.; Al-Johani, K.; Scully, C. Salivary microRNAs in oral cancer. Oral Dis. 2015, 21, 739–747. [Google Scholar] [CrossRef]
- Yoshizawa, J.M.; Wong, D.T.W. Salivary microRNAs and oral cancer detection. Methods Mol. Biol. 2013, 936, 313–324. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shah, K.; Patel, S.; Modi, B.; Shah, F.; Rawal, R. Uncovering the potential of CD44v/SYNE1/miR34a axis in salivary fluids of oral cancer patients. J. Oral Pathol. Med. 2018, 47, 345–352. [Google Scholar] [CrossRef] [PubMed]
- Gai, C.; Camussi, F.; Broccoletti, R.; Gambino, A.; Cabras, M.; Molinaro, L.; Carossa, S.; Camussi, G.; Arduino, P.G. Salivary extracellular vesicle-associated miRNAs as potential biomarkers in oral squamous cell carcinoma. BMC Cancer 2018, 18, 1–11. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Korostoff, A.; Reder, L.; Masood, R.; Sinha, U.K. The role of salivary cytokine biomarkers in tongue cancer invasion and mortality. Oral Oncol. 2011, 47, 282–287. [Google Scholar] [CrossRef] [PubMed]
- Ogawa, Y.; Taketomi, Y.; Murakami, M.; Tsujimoto, M.; Yanoshita, R. Small RNA transcriptomes of two types of exosomes in human whole saliva determined by next generation sequencing. Biol. Pharm. Bull. 2013, 36, 66–75. [Google Scholar] [CrossRef] [Green Version]
- Wagenblast, E.; Araujo, J.; Gan, O.I.; Cutting, S.K.; Murison, A.; Krivdova, G.; Azkanaz, M.; McLeod, J.L.; Smith, S.A.; Gratton, B.A.; et al. Mapping the cellular origin and early evolution of leukemia in Down syndrome. Science 2021, 373, eabf6202. [Google Scholar] [CrossRef]
- Mishra, S.; Yadav, T.; Rani, V. Exploring miRNA based approaches in cancer diagnostics and therapeutics. Crit. Rev. Oncol. Hematol. 2016, 98, 12–23. [Google Scholar] [CrossRef]
- Okada, R.; Koshizuka, K.; Yamada, Y.; Moriya, S.; Kikkawa, N.; Kinoshita, T.; Hanazawa, T.; Seki, N. Regulation of oncogenic targets by miR-99a-3p (Passenger Strand of miR-99a-Duplex) in head and neck squamous cell carcinoma. Cells 2019, 8, 1535. [Google Scholar] [CrossRef] [Green Version]
- Fehlmann, T.; Backes, C.; Alles, J.; Fischer, U.; Hart, M.; Kern, F.; Langseth, H.; Rounge, T.; Umu, S.U.; Kahraman, M.; et al. A high-resolution map of the human small non-coding transcriptome. Bioinformatics 2018, 34, 1621–1628. [Google Scholar] [CrossRef] [Green Version]
- Di Leva, G.; Croce, C.M. MicroRNAs. Clin. Biochem. 2013, 46, 840–841. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Ba, Y.; Ma, L.; Cai, X.; Yin, Y.; Wang, K.; Guo, J.; Zhang, Y.; Chen, J.; Guo, X.; et al. Characterization of microRNAs in serum: A novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res. 2008, 18, 997–1006. [Google Scholar] [CrossRef] [Green Version]
- Chiabotto, G.; Gai, C.; Deregibus, M.C.; Camussi, G. Salivary extracellular vesicle-associated exRNA as cancer biomarker. Cancers Basel 2019, 11, 891. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Machida, T.; Tomofuji, T.; Maruyama, T.; Yoneda, T.; Ekuni, D.; Azuma, T.; Miyai, H.; Mizuno, H.; Kato, H.; Tsutsumi, K.; et al. miR1246 and miR4644 in salivary exosome as potential biomarkers for pancreatobiliary tract cancer. Oncol. Rep. 2016, 36, 2375–2381. [Google Scholar] [CrossRef] [Green Version]
- Michael, A.; Bajracharya, S.D.; Yuen, P.S.T.; Zhou, H.; Star, R.A.; Illei, G.G.; Alevizos, I. Exosomes from human saliva as a source of microRNA biomarkers. Oral Dis. 2010, 16, 34–38. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Byun, J.-S.; Hong, S.-H.; Choi, J.-K.; Jung, J.-K.; Lee, H.-J. Diagnostic profiling of salivary exosomal microRNAs in oral lichen planus patients. Oral Dis. 2015, 21, 987–993. [Google Scholar] [CrossRef]
- Li, Y.; St John, M.A.R.; Zhou, X.; Kim, Y.; Sinha, U.; Jordan, R.C.; Eisele, D.; Abemayor, E.; Elashoff, D.; Park, N.H.; et al. Salivary transcriptome diagnostics for oral cancer detection. Clin. Cancer Res. 2004, 10, 8442–8450. [Google Scholar] [CrossRef] [Green Version]
- Romani, C.; Salviato, E.; Paderno, A.; Zanotti, L.; Ravaggi, A.; Deganello, A.; Berretti, G.; Gualtieri, T.; Marchini, S.; D’Incalci, M.; et al. Genome-wide study of salivary miRNAs identifies miR-423-5p as promising diagnostic and prognostic biomarker in oral squamous cell carcinoma. Theranostics 2021, 11, 2987–2999. [Google Scholar] [CrossRef]
- The Cancer Genome Atlas Network. Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature 2015, 517, 576–582. [Google Scholar] [CrossRef] [Green Version]
- Paluszczak, J. The Significance of the Dysregulation of Canonical Wnt Signaling in Head and Neck Squamous Cell Carcinomas. Cells 2020, 9, 723. [Google Scholar] [CrossRef] [Green Version]
- Pentangelo, G.; Nistico, S.P.; Provenzano, E.; Cisale, G.Y.; Bennardo, L. Topical 5% imiquimod sequential to surgery for HPV-related squamous cell carcinoma of the lip. Med. Kaunas 2021, 57, 563. [Google Scholar] [CrossRef]
- Bennardo, L.; Bennardo, F.; Giudice, A.; Passante, M.; Dastoli, S.; Morrone, P.; Provenzano, E.; Patruno, C.; Nistico, S.P. Local chemotherapy as an adjuvant treatment in unresectable squamous cell carcinoma: What do we know so far? Curr. Oncol. 2021, 28, 2317–2325. [Google Scholar] [CrossRef]
- Yoshizawa, J.M.; Schafer, C.A.; Schafer, J.J.; Farrell, J.J.; Paster, B.J.; Wong, D.T. Salivary biomarkers: Toward future clinical and diagnostic utilities. Clin. Microbiol. Rev. 2013, 26, 781–791. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bahn, J.H.; Zhang, Q.; Li, F.; Chan, T.M.; Lin, X.; Kim, Y.; Wong, D.T.; Xiao, X. The landscape of microRNA, Piwi-interacting RNA, and circular RNA in human saliva. Clin. Chem. 2015, 61, 221–230. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mishra, S.; Saadat, D.; Kwon, O.; Lee, Y.; Choi, W.-S.; Kim, J.-H.; Yeo, W.H. Recent advances in salivary cancer diagnostics enabled by biosensors and bioelectronics. Biosens. Bioelectron. 2016, 81, 181–197. [Google Scholar] [CrossRef]
- Herichova, I.; Reis, R.; Hasakova, K.; Vician, M. Downregulation of miR-30c-5p expression in colorectal cancer tissue is sex-dependent. Physiol. Res. 2020, 69, S479–S487. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.; Guo, Y.; Chen, C.; Fan, D.; Wu, X.; Zhao, L.; Shao, B.; Sun, Z.; Ji, Z. Circ3823 contributes to growth, metastasis and angiogenesis of colorectal cancer: Involvement of miR-30c-5p/TCF7 axis. Mol. Cancer 2021, 20, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Shi, H.; Du, Y.; Zhao, G.; Wang, X.; Li, Q.; Liu, J.; Ye, L.; Shen, Z.; Guo, Y.; et al. lncRNA DLEU2 modulates cell proliferation and invasion of non-small cell lung cancer by regulating miR-30c-5p/SOX9 axis. Aging Albany N. Y. 2019, 11, 7386–7401. [Google Scholar] [CrossRef]
- MacDonagh, L.; Gallagher, M.F.; Ffrench, B.; Gasch, C.; Gray, S.G.; Reidy, M.; Nicholson, S.; Leonard, N.; Ryan, R.; Young, V.; et al. MicroRNA expression profiling and biomarker validation in treatment-naive and drug resistant non-small cell lung cancer. Transl. Lung Cancer Res. 2021, 10, 1773–1791. [Google Scholar] [CrossRef] [PubMed]
- Cao, J.-M.; Li, G.-Z.; Han, M.; Xu, H.-L.; Huang, K.-M. MiR-30c-5p suppresses migration, invasion and epithelial to mesenchymal transition of gastric cancer via targeting MTA1. Biomed. Pharmacother. 2017, 93, 554–560. [Google Scholar] [CrossRef]
- Tanaka, T.; Okada, R.; Hozaka, Y.; Wada, M.; Moriya, S.; Satake, S.; Idichi, T.; Kurahara, H.; Ohtsuka, T.; Seki, N. Molecular pathogenesis of pancreatic ductal adenocarcinoma: Impact of miR-30c-5p and miR-30c-2-3p regulation on oncogenic genes. Cancers 2020, 12, 2731. [Google Scholar] [CrossRef]
- Onyshchenko, K.V.; Voitsitskyi, T.V.; Grygorenko, V.M.; Saidakova, N.O.; Pereta, L.V.; Onyschuk, A.P.; Skrypkina, I.Y. Expression of micro-RNA hsa-miR-30c-5p and hsa-miR-138-1 in renal cell carcinoma. Exp. Oncol. 2020, 42, 115–119. [Google Scholar] [CrossRef]
- Song, S.; Long, M.; Yu, G.; Cheng, Y.; Yang, Q.; Liu, J.; Wang, Y.; Sheng, J.; Wang, L.; Wang, Z.; et al. Urinary exosome miR-30c-5p as a biomarker of clear cell renal cell carcinoma that inhibits progression by targeting HSPA5. J. Cell. Mol. Med. 2019, 23, 6755–6765. [Google Scholar] [CrossRef] [PubMed]
- Oksuz, Z.; Serin, M.S.; Kaplan, E.; Dogen, A.; Tezcan, S.; Aslan, G.; Emekdas, G.; Sezgin, O.; Altintas, E.; Tiftik, E.N. Serum microRNAs; miR-30c-5p, miR-223-3p, miR-302c-3p and miR-17-5p could be used as novel non-invasive biomarkers for HCV-positive cirrhosis and hepatocellular carcinoma. Mol. Biol. Rep. 2015, 42, 713–720. [Google Scholar] [CrossRef]
- Sheng, L.-Q.; Li, J.-R.; Qin, H.; Liu, L.; Zhang, D.-D.; Zhang, Q.; Huang, M.-L.; Li, X.-L.; Xu, X.-Y.; Wei, Y.-N.; et al. Blood exosomal micro ribonucleic acid profiling reveals the complexity of hepatocellular carcinoma and identifies potential biomarkers for differential diagnosis. World J. Gastrointest. Oncol. 2020, 12, 1195–1208. [Google Scholar] [CrossRef]
- Wong, V.C.-L.; Wong, M.-I.; Lam, C.-T.; Lung, M.L.; Lam, K.-O.; Lee, V.H.-F. Hallmark microRNA signature in liquid biopsy identifies hepatocellular carcinoma and differentiates it from liver metastasis. J. Cancer 2021, 12, 4585–4594. [Google Scholar] [CrossRef]
- Cai, W.; Ji, J.; Wu, B.; Hao, K.; Ren, P.; Jin, Y.; Yang, L.; Tong, Q.; Shen, Z. Characterization of the small RNA transcriptomes of cell protrusions and cell bodies of highly metastatic hepatocellular carcinoma cells via RNA sequencing. Oncol. Lett. 2021, 22, 1–15. [Google Scholar] [CrossRef]
- Pei, B.; Li, T.; Qian, Q.; Fan, W.; He, X.; Zhu, Y.; Xu, L. Downregulation of microRNA-30c-5p was responsible for cell migration and tumor metastasis via COTL1-mediated microfilament arrangement in breast cancer. Gland. Surg. 2020, 9, 747–758. [Google Scholar] [CrossRef]
- Chang, J.T.-H.; Wang, F.; Chapin, W.; Huang, R.S. Identification of MicroRNAs as breast cancer prognosis markers through the cancer genome atlas. PLoS ONE 2016, 11, e0168284. [Google Scholar] [CrossRef] [Green Version]
- Yen, M.-C.; Shih, Y.-C.; Hsu, Y.-L.; Lin, E.-S.; Lin, Y.-S.; Tsai, E.-M.; Ho, Y.-W.; Hou, M.-F.; Kuo, P.-L. Isolinderalactone enhances the inhibition of SOCS3 on STAT3 activity by decreasing miR-30c in breast cancer. Oncol. Rep. 2016, 35, 1356–1364. [Google Scholar] [CrossRef] [Green Version]
- Turashvili, G.; Lightbody, E.D.; Tyryshkin, K.; SenGupta, S.K.; Elliott, B.E.; Madarnas, Y.; Ghaffari, A.; Day, A.; Nicol, C.J.B. Novel prognostic and predictive microRNA targets for triple-negative breast cancer. FASEB J. 2018, 32, 5937–5954. [Google Scholar] [CrossRef] [PubMed]
- Amorim, M.; Lobo, J.; Fontes-Sousa, M.; Estevao-Pereira, H.; Salta, S.; Lopes, P.; Coimbra, N.; Antunes, L.; De Sousa, S.P.; Henrique, R.; et al. Predictive and prognostic value of selected MicroRNAs in luminal breast cancer. Front. Genet. 2019, 10, 815. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, X.; Xu, F.; Meng, Q.; Gong, N.; Teng, Z.; Xu, R.; Zhao, M.; Xia, M. Long noncoding RNA DLEU2 predicts a poor prognosis and enhances malignant properties in laryngeal squamous cell carcinoma through the miR-30c-5p/PIK3CD/Akt axis. Cell Death Dis. 2020, 11, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Ganci, F.; Sacconi, A.; Ben-Moshe, N.B.; Manciocco, V.; Sperduti, I.; Strigari, L.; Covello, R.; Benevolo, M.; Pescarmona, E.; Domany, E.; et al. Expression of TP53 mutation-associated microRNAs predicts clinical outcome in head and neck squamous cell carcinoma patients. Ann. Oncol. 2013, 24, 3082–3088. [Google Scholar] [CrossRef] [PubMed]
- Lei, Y.; Chen, L.; Zhang, G.; Shan, A.; Ye, C.; Liang, B.; Sun, J.; Liao, X.; Zhu, C.; Chen, Y.; et al. MicroRNAs target the Wnt/betacatenin signaling pathway to regulate epithelialmesenchymal transition in cancer (Review). Oncol. Rep. 2020, 44, 1299–1313. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Jiang, H.; Zhao, J.; Wen, H. MiRNA-16 inhibited oral squamous carcinoma tumor growth in vitro and in vivo via suppressing Wnt/beta-catenin signaling pathway. Onco. Targets Ther. 2018, 11, 5111–5119. [Google Scholar] [CrossRef] [Green Version]
- Zhao, C.; Liu, Z. MicroRNA 617 Targeting SERPINE1 inhibited the progression of oral squamous cell carcinoma. Mol. Cell. Biol. 2021, 41, e0056520. [Google Scholar] [CrossRef] [PubMed]
- Behera, A.K.; Kumar, M.; Shanmugam, M.K.; Bhattacharya, A.; Rao, V.J.; Bhat, A.; Vasudevan, M.; Gopinath, K.S.; Mohiyuddin, A.; Chatterjee, A.; et al. Functional interplay between YY1 and CARM1 promotes oral carcinogenesis. Oncotarget 2019, 10, 3709–3724. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, L.; Li, R.; Qiu, J.-Z.; Yu, J.-B.; Cao, Y.; Yuan, R.-T. YY1-mediated PTEN dephosphorylation antagonizes IR-induced DNA repair contributing to tongue squamous cell carcinoma radiosensitization. Mol. Cell. Probes 2020, 53, 101577. [Google Scholar] [CrossRef]
- Nakamura, A.; Kakihara, Y.; Funayama, A.; Haga, K.; Mikami, T.; Kobayashi, D.; Yoshida, Y.; Izumi, K.; Kobayashi, T.; Saeki, M. HEATR1, a novel interactor of Pontin/Reptin, stabilizes Pontin/Reptin and promotes cell proliferation of oral squamous cell carcinoma. Biochem. Biophys. Res. Commun. 2021, 557, 294–301. [Google Scholar] [CrossRef]

| Characteristics | OSCC Patients | Controls |
|---|---|---|
| Number | 33 | 12 |
| Gender | M 27 (81.8%)/F 6 (18.2%) | M 7 (58.3%)/F 5 (41.7%) |
| Age | 61.61 | 54.17 |
| Range | 47–80 | 40–69 |
| Smokers | 29 (87.9%) | 10 (83.3%) |
| HPV positivity | No | N/A |
| Histology | SSC | N/A |
| Location | Tongue 7 (21%) | |
| Floor of mouth 15 (45.5%) | N/A | |
| Retromolar triangle 1 (3%) | ||
| Gingiva 10 (30.3%) | ||
| Size (mm) | 36,1 | N/A |
| Range (mm) | 10–70 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mehterov, N.; Vladimirov, B.; Sacconi, A.; Pulito, C.; Rucinski, M.; Blandino, G.; Sarafian, V. Salivary miR-30c-5p as Potential Biomarker for Detection of Oral Squamous Cell Carcinoma. Biomedicines 2021, 9, 1079. https://doi.org/10.3390/biomedicines9091079
Mehterov N, Vladimirov B, Sacconi A, Pulito C, Rucinski M, Blandino G, Sarafian V. Salivary miR-30c-5p as Potential Biomarker for Detection of Oral Squamous Cell Carcinoma. Biomedicines. 2021; 9(9):1079. https://doi.org/10.3390/biomedicines9091079
Chicago/Turabian StyleMehterov, Nikolay, Boyan Vladimirov, Andrea Sacconi, Claudio Pulito, Marcin Rucinski, Giovanni Blandino, and Victoria Sarafian. 2021. "Salivary miR-30c-5p as Potential Biomarker for Detection of Oral Squamous Cell Carcinoma" Biomedicines 9, no. 9: 1079. https://doi.org/10.3390/biomedicines9091079
APA StyleMehterov, N., Vladimirov, B., Sacconi, A., Pulito, C., Rucinski, M., Blandino, G., & Sarafian, V. (2021). Salivary miR-30c-5p as Potential Biomarker for Detection of Oral Squamous Cell Carcinoma. Biomedicines, 9(9), 1079. https://doi.org/10.3390/biomedicines9091079