Matrix Stiffness Modulates Mechanical Interactions and Promotes Contact between Motile Cells
Abstract
1. Introduction
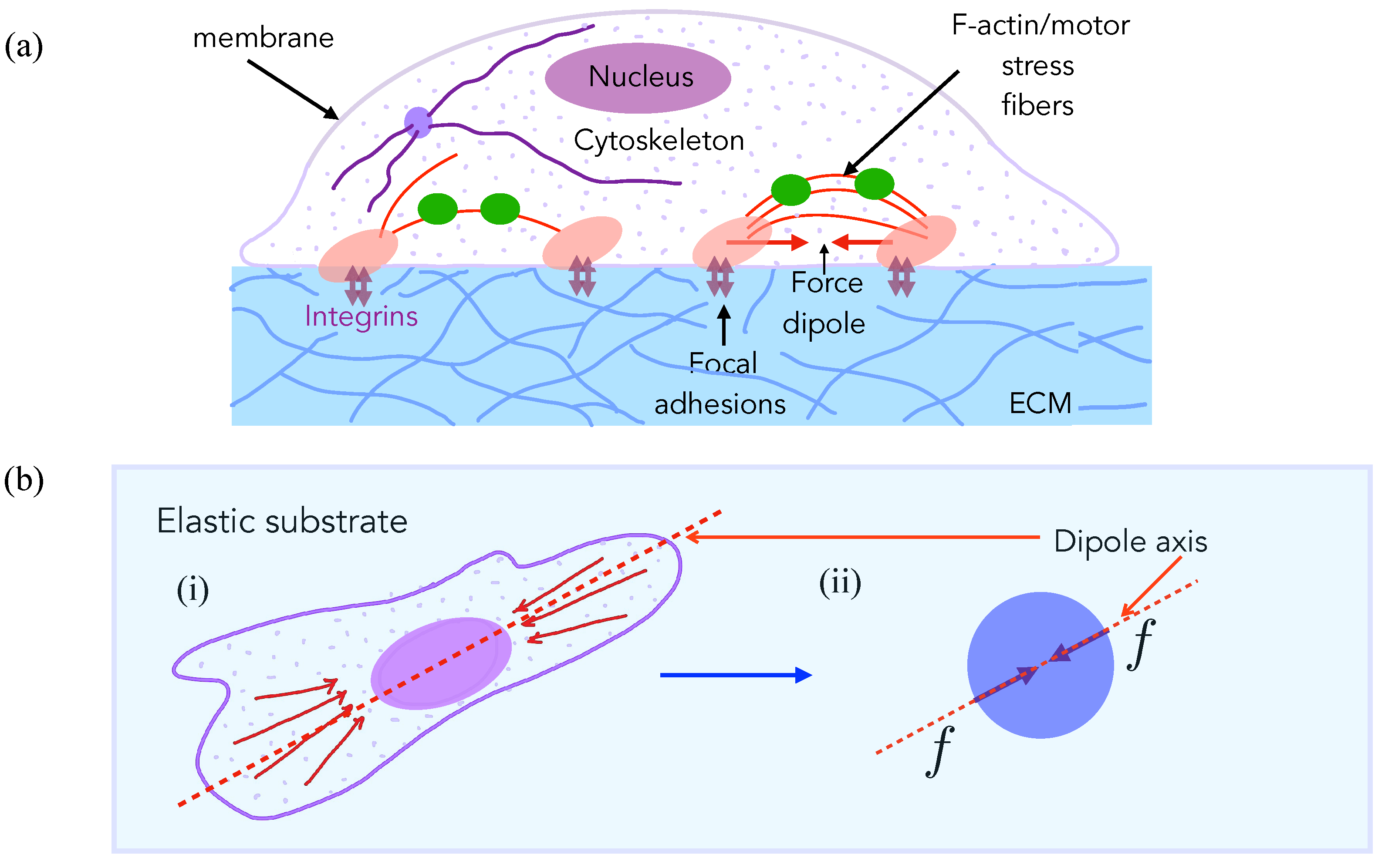
2. Experimental Observations Motivate Model for Cell Elastic Interactions
3. Materials and Methods
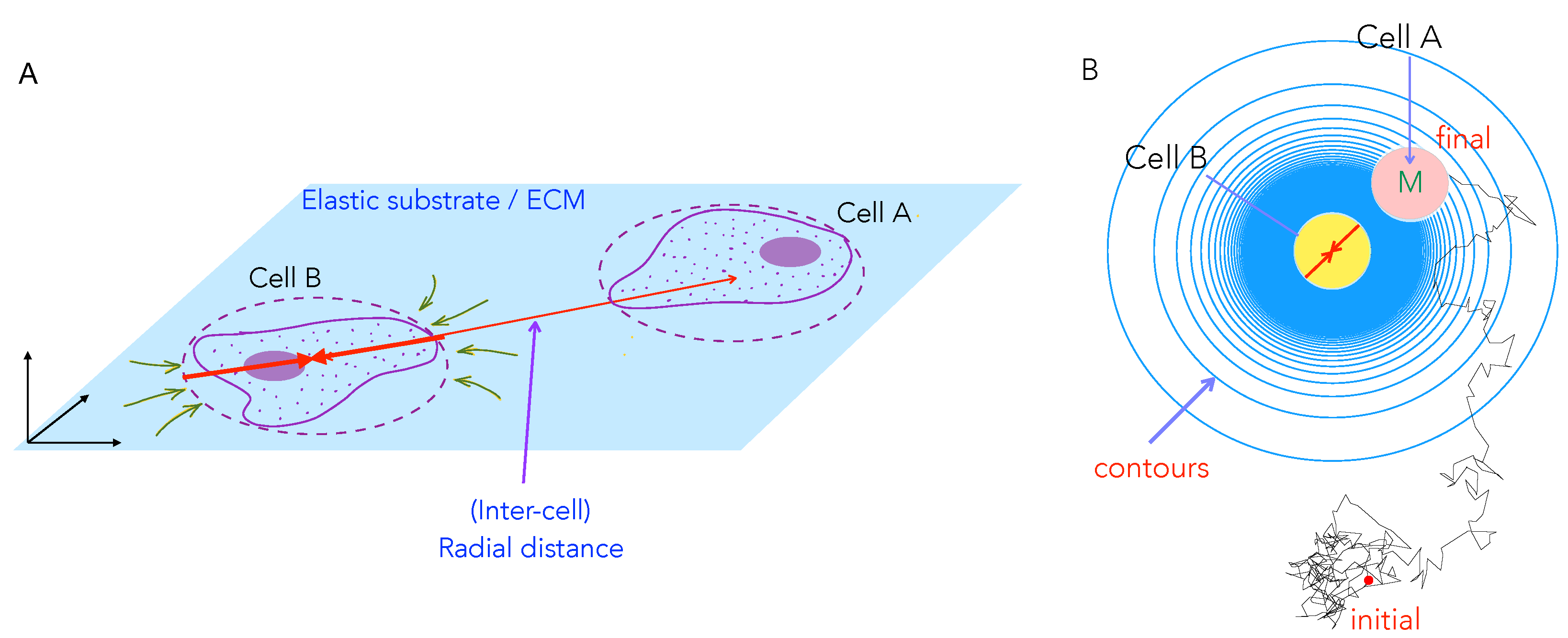
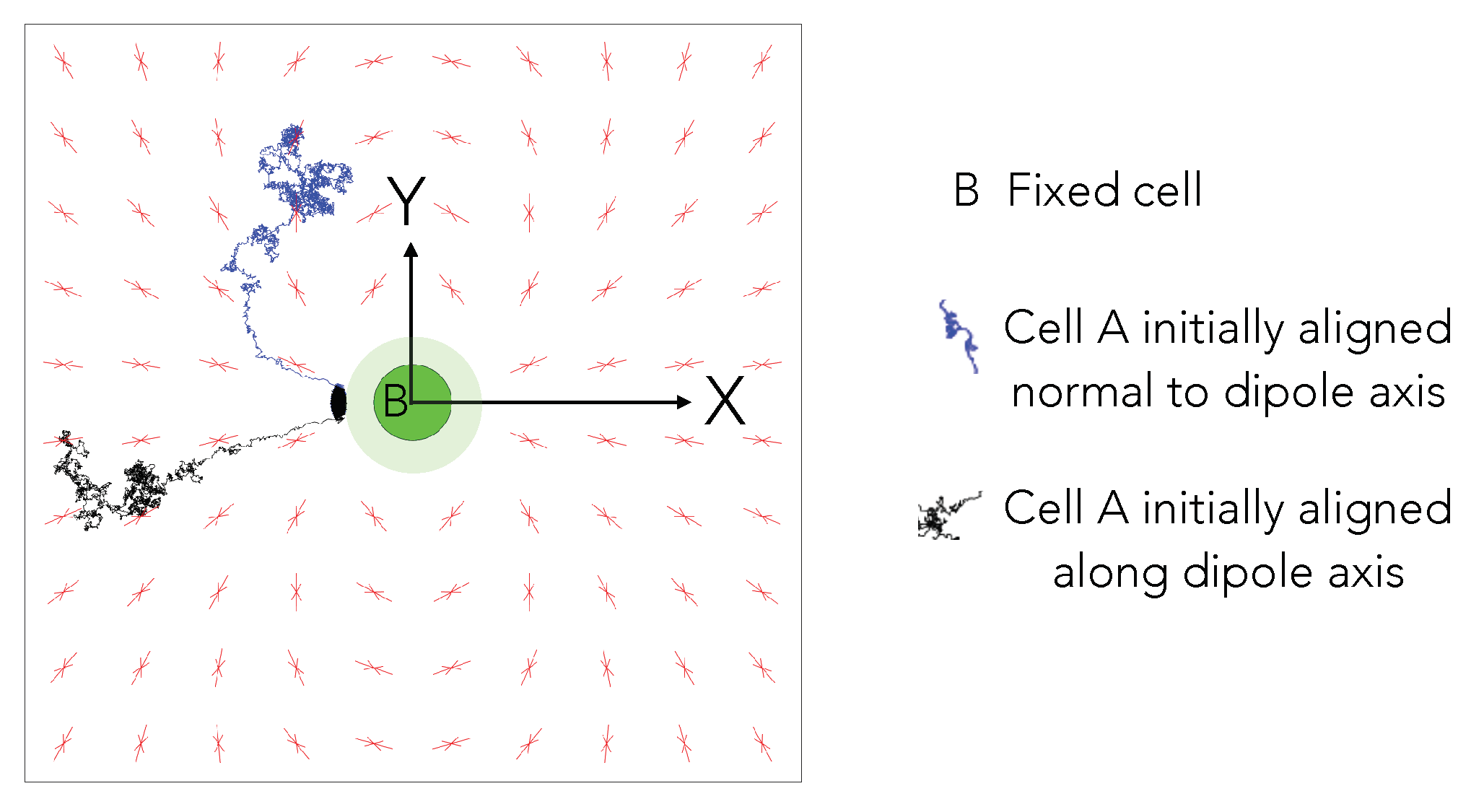
3.1. Model for Two-Cell Interactions
3.2. Dimensionless Parameters Quantifying Cell Motion and Interactions
3.3. Numerical Solution and Tracking Cell Trajectories
4. Results
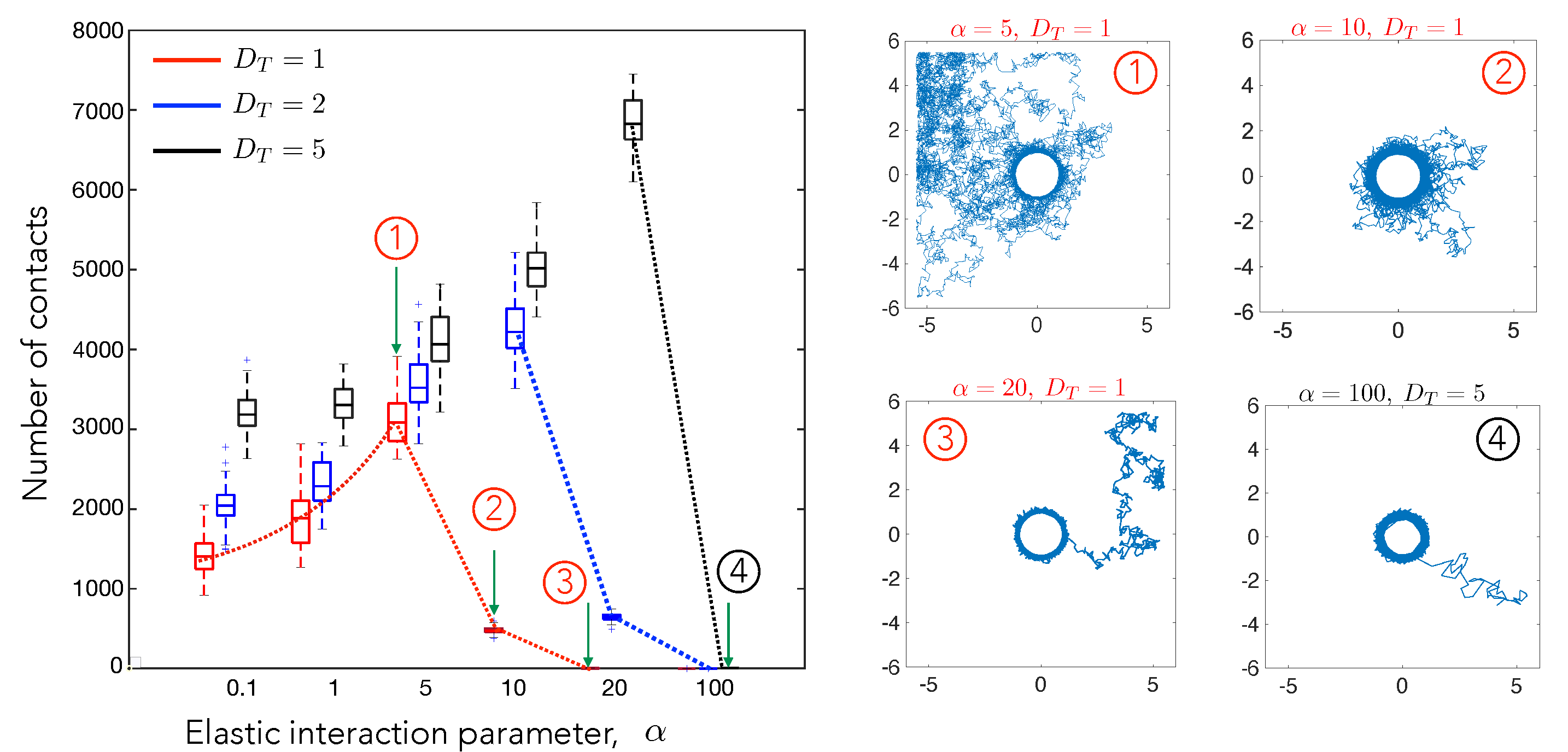
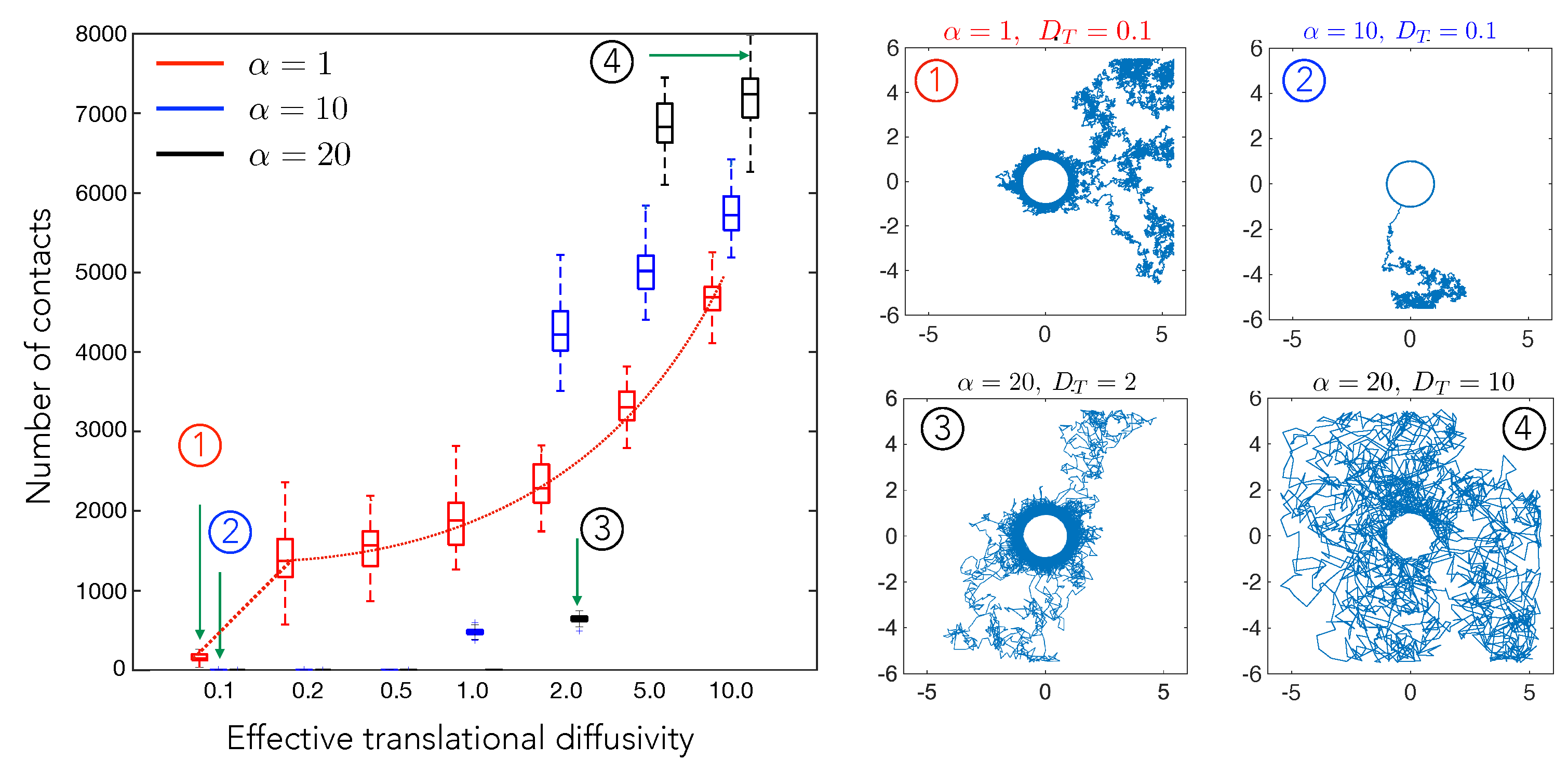
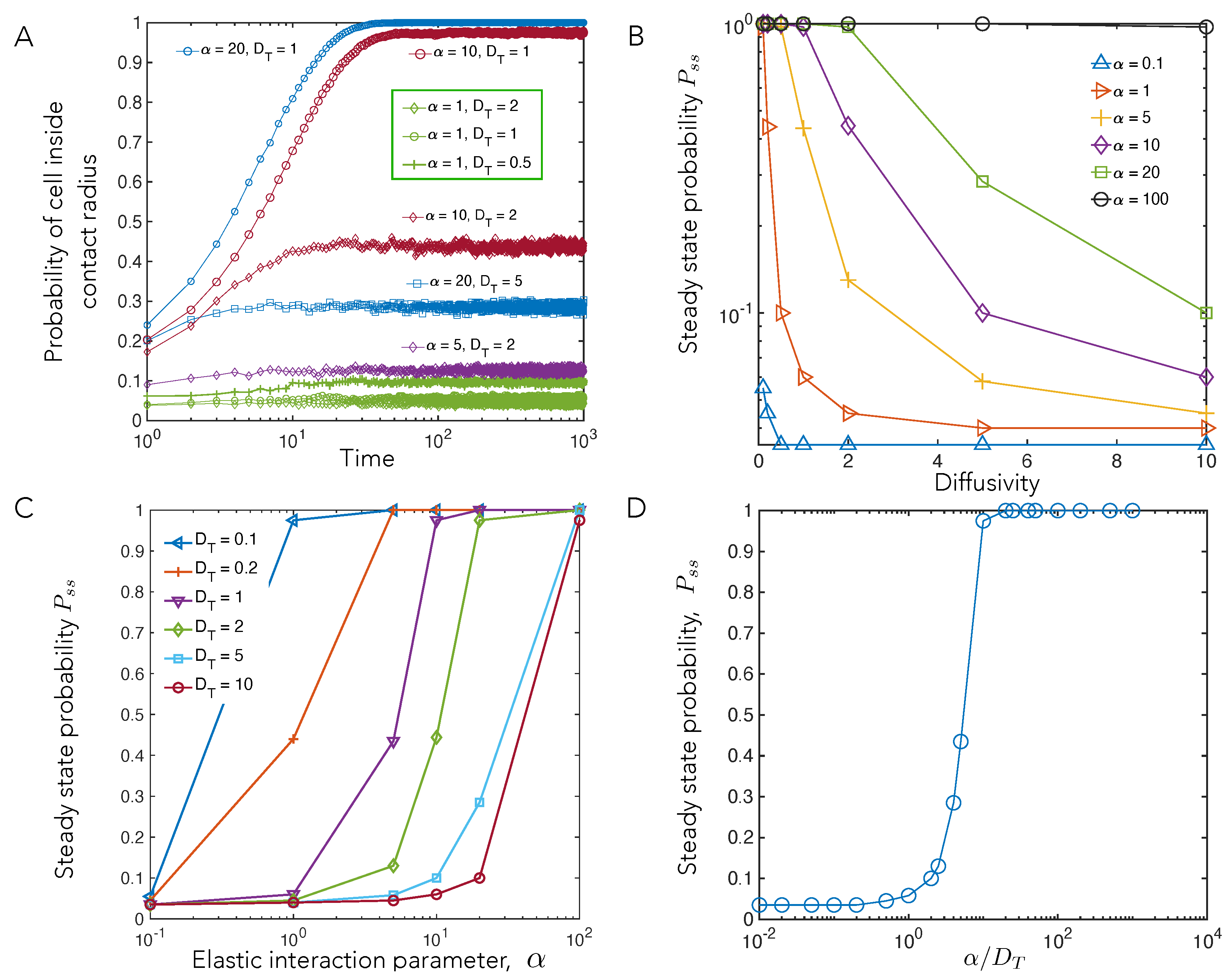
4.1. Cell–Cell Contact Frequency Is Controlled by Matrix Elastic Interactions
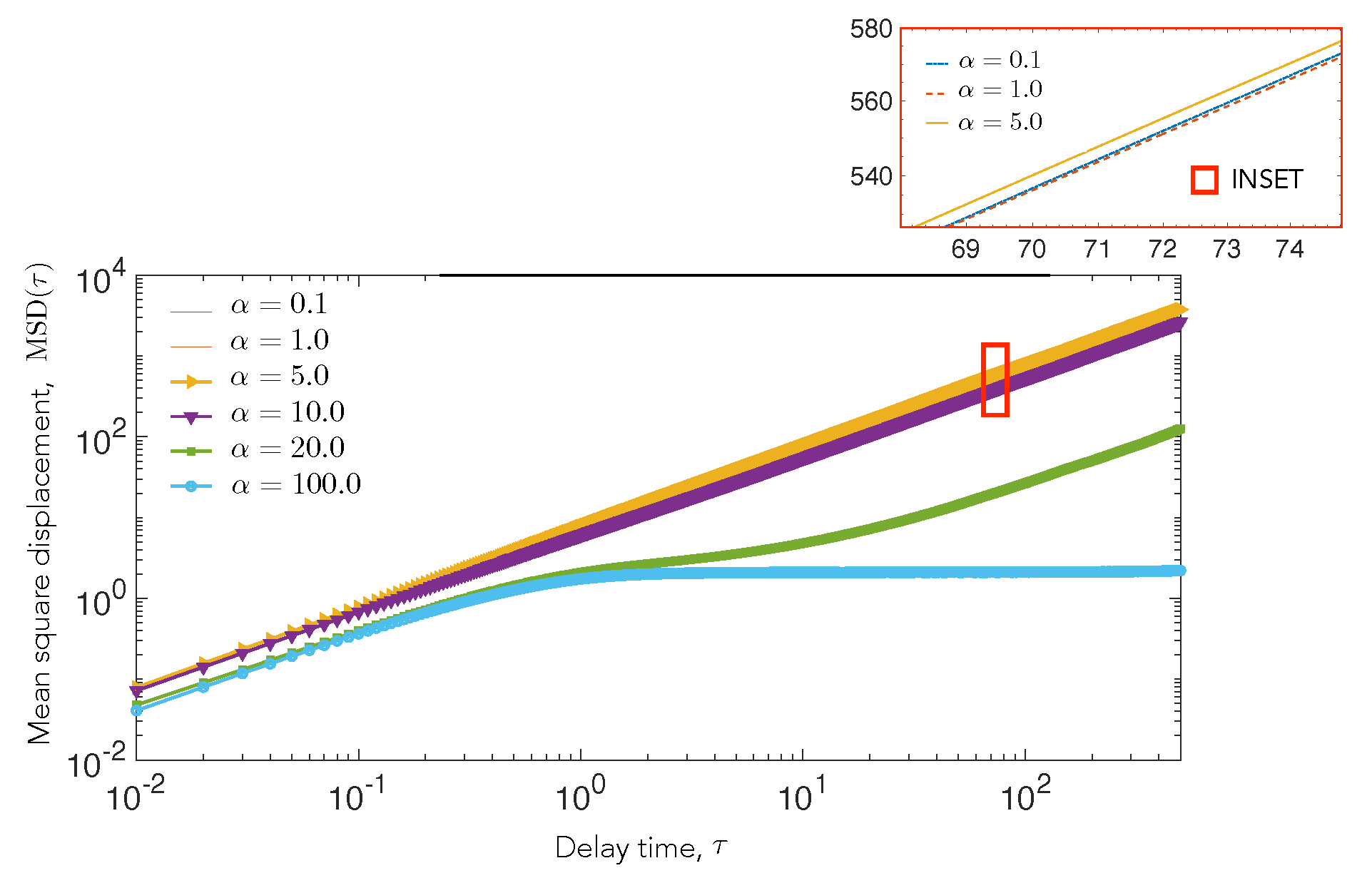
4.2. Cell Motility Characteristics Depend on Elastic Interactions
4.3. Elastic Interactions Lead to Effective Capture of Motile Cell
5. Discussion and Future Extensions to Other Forms of Interactions
5.1. Anisotropic Cell-Cell Elastic Interactions
5.2. Extensions to Near-Contact Biochemical or Bond Interactions
6. Summary
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| MSD | Mean Square Displacement |
Appendix A Model for a Moving Cell Interacting with a Stationary Cell Via Substrate Elasticity
References
- Ladoux, B.; Mège, R.M. Mechanobiology of collective cell behaviours. Nat. Rev. Mol. Cell Biol. 2017, 18, 743–757. [Google Scholar] [CrossRef]
- Alert, R.; Trepat, X. Physical Models of Collective Cell Migration. Annu. Rev. Condens. Matter Phys. 2020, 11, 77–101. [Google Scholar] [CrossRef]
- Reinhart-King, C.A.; Dembo, M.; Hammer, D.A. Cell-Cell Mechanical Communication through Compliant Substrates. Biophys. J. 2008, 95, 6044–6051. [Google Scholar] [CrossRef] [PubMed]
- Winer, J.P.; Oake, S.; Janmey, J.P. Non-Linear Elasticity of Extracellular Matrices Enables Contractile Cells to Communicate Local Position and Orientation. PLoS ONE 2009, 4, e6382. [Google Scholar] [CrossRef]
- Discher, D.E.; Janmey, P.; Wang, Y.L. Tissue Cells Feel and Respond to the Stiffness of Their Substrate. Science 2005, 310, 1139–1143. [Google Scholar] [CrossRef] [PubMed]
- Vaziri, A.; Gopinath, A. Cell and biomolecular mechanics in silico. Nat. Mater. 2008, 7, 15–23. [Google Scholar] [CrossRef] [PubMed]
- Engler, A.J.; Sen, S.; Sweeney, H.L.; Discher, D.E. Matrix Elasticity Directs Stem Cell Lineage Specification. Cell 2006, 126, 677–689. [Google Scholar] [CrossRef] [PubMed]
- Eyckmans, J.; Boudou, T.; Yu, X.; Chen, C. A Hitchhiker’s Guide to Mechanobiology. Dev. Cell 2011, 21, 35–47. [Google Scholar] [CrossRef]
- Barriga, E.H.; Franze, K.; Charras, G.; Mayor, R. Tissue stiffening coordinates morphogenesis by triggering collective cell migration in vivo. Nature 2018, 554, 523–527. [Google Scholar] [CrossRef]
- Tee, S.Y.; Fu, J.; Chen, C.S.; Janmey, P.A. Cell Shape and Substrate Rigidity Both Regulate Cell Stiffness. Biophys. J. 2011, 100, L25–L27. [Google Scholar] [CrossRef] [PubMed]
- Chopra, A.; Tabdanov, E.; Patel, H.; Janmey, P.A.; Kresh, J.Y. Cardiac myocyte remodeling mediated by N-cadherin-dependent mechanosensing. Am. J. Physiol. 2011, 300, H1252–H1266. [Google Scholar] [CrossRef] [PubMed]
- Vichare, S.; Sen, S.; Inamdar, M.M. Cellular mechanoadaptation to substrate mechanical properties: Contributions of substrate stiffness and thickness to cell stiffness measurements using AFM. Soft Matter 2014, 10, 1174–1181. [Google Scholar] [CrossRef] [PubMed]
- Staunton, J.R.; Doss, B.L.; Lindsay, S.; Ros, R. Correlating confocal microscopy and atomic force indentation reveals metastatic cancer cells stiffen during invasion into collagen I matrices. Sci. Rep. 2016, 6, 19686. [Google Scholar] [CrossRef] [PubMed]
- Cross, S.E.; Jin, Y.S.; Rao, J.; Gimzewski, J.K. Nanomechanical analysis of cells from cancer patients. Nat. Nanotechnol. 2007, 2, 780–783. [Google Scholar] [CrossRef]
- Han, S.J.; Bielawski, K.S.; Ting, L.H.; Rodriguez, M.L.; Sniadecki, N.J. Decoupling substrate stiffness, spread area, and micropost density: A close spatial relationship between traction forces and focal adhesions. Biophys. J. 2012, 103, 640–648. [Google Scholar] [CrossRef]
- Rheinlaender, J.; Dimitracopoulos, A.; Wallmeyer, B.; Kronenberg, N.M.; Chalut, K.J.; Gather, M.C.; Betz, T.; Charras, G.; Franze, K. Cortical cell stiffness is independent of substrate mechanics. Nat. Mater. 2020, 19, 1019–1025. [Google Scholar] [CrossRef]
- Balaban, N.Q.; Schwarz, U.S.; Riveline, D.; Goichberg, P.; Tzur, G.; Sabanay, I.; Mahalu, D.; Safran, S.; Bershadsky, A.; Addadi, L.; et al. Force and focal adhesion assembly: A close relationship studied using elastic micropatterned substrates. Nat. Cell Biol. 2001, 3, 466. [Google Scholar] [CrossRef]
- Guo, C.L.; Ouyang, M.; Yu, J.Y.; Maslov, J.; Price, A.; Shen, C.Y. Long-range mechanical force enables self-assembly of epithelial tubular patterns. Proc. Natl. Acad. Sci. USA 2012, 109, 5576–5582. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Abhilash, A.; Chen, C.S.; Wells, R.G.; Shenoy, V.B. Long-Range Force Transmission in Fibrous Matrices Enabled by Tension-Driven Alignment of Fibers. Biophys. J. 2014, 107, 2592–2603. [Google Scholar] [CrossRef]
- Natan, S.; Koren, Y.; Shelah, O.; Goren, S.; Lesman, A. Long-range mechanical coupling of cells in 3D fibrin gels. Mol. Biol. Cell 2020, 31, 1474–1485. [Google Scholar] [CrossRef] [PubMed]
- Pakshir, P.; Alizadehgiashi, M.; Wong, B.; Coelho, N.M.; Chen, X.; Gong, Z.; Shenoy, V.B.; McCulloch, C.A.; Hinz, B. Dynamic fibroblast contractions attract remote macrophages in fibrillar collagen matrix. Nat. Commun. 2019, 10, 1850. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Kim, T.H.; Thauland, T.J.; Li, H.; Majedi, F.S.; Ly, C.; Gu, Z.; Butte, M.J.; Rowat, A.C.; Li, S. Unraveling the mechanobiology of immune cells. Curr. Opin. Biotechnol. 2020, 66, 236–245. [Google Scholar] [CrossRef] [PubMed]
- Kwon, T.G.; Yang, T.D.; Lee, K.J. Enhancement of Chemotactic Cell Aggregation by Haptotactic Cell-To-Cell Interaction. PLoS ONE 2016, 11, e0154717. [Google Scholar] [CrossRef]
- Yang, T.D.; Kwon, T.G.; Park, J.S.; Lee, K.J. Trail networks formed by populations of immune cells. New J. Phys. 2014, 16, 023017. [Google Scholar] [CrossRef]
- Nasu-Tada, K.; Koizumi, S.; Inoue, K. Involvement of β1 integrin in microglial chemotaxis and proliferation on fibronectin: Different regulations by ADP through PKA. Glia 2005, 52, 98–107. [Google Scholar] [CrossRef]
- Fetler, L.; Amigorena, S. Brain under surveillance: The microglia patrol. Neuroscience 2005, 309, 329–393. [Google Scholar]
- Vincent, C.; Siddiqui, T.A.; Schlichter, L.C. Podosomes in migrating microglia: Components and matrix degradation. J. Neuroinflammation 2012, 9, 190. [Google Scholar] [CrossRef]
- Théry, C.; Ostrowski, M.; Segura, E. Membrane vesicles as conveyors of immune responses. Nat. Rev. Immunol. 2009, 9, 581–593. [Google Scholar] [CrossRef] [PubMed]
- Theoharides, T.C.; Tsilioni, I.; Bawazeer, M. Mast cells, neuroinflammation and pain in fibromyalgia syndrome. Front. Cell. Neurosci. 2019, 13, 353. [Google Scholar] [CrossRef] [PubMed]
- Demopoulos, C.; Antonopoulou, S.; Theoharides, T.C. COVID-19, microthromboses, and platelet activating factor. Biofactors 2020, 46, 927–933. [Google Scholar] [CrossRef] [PubMed]
- Yuval, J.; Safran, S.A. Dynamics of elastic interactions in soft and biological matter. Phys. Rev. E 2013, 87, 042703. [Google Scholar] [CrossRef]
- Tang, X.; Bajaj, P.; Bashir, R.; Saif, T.A. How far cardiac cells can see each other mechanically. Soft Matter 2011, 7, 6151–6158. [Google Scholar] [CrossRef]
- Nitsan, I.; Drori, S.; Lewis, Y.E.; Cohen, S.; Tzlil, S. Mechanical communication in cardiac cell synchronized beating. Nat. Phys. 2016, 12, 472–477. [Google Scholar] [CrossRef]
- Dasbiswas, K.; Majkut, S.; Discher, D.E.; Safran, S.A. Substrate stiffness-modulated registry phase correlations in cardiomyocytes map structural order to coherent beating. Nat. Commun. 2015, 6, 6085. [Google Scholar] [CrossRef] [PubMed]
- Tomasek, J.J.; Gabbiani, G.; Hinz, B.; Chaponnier, C.; Brown, R.A. Myofibroblasts and mechano-regulation of connective tissue remodelling. Nat. Rev. Mol. Cell Biol. 2002, 3, 349–363. [Google Scholar] [CrossRef] [PubMed]
- Korff, T.; Augustin, H. Tensional forces in fibrillar extracellular matrices control directional capillary sprouting. J. Cell Sci. 1999, 112, 3249–3258. [Google Scholar]
- Liu, Z.; Woo, S.; Weiner, O.D. Nodal signaling has dual roles in fate specification and directed migration during germ layer segregation in zebrafish. Development 2018, 145. [Google Scholar] [CrossRef]
- Pelham, R.J.; Wang, Y.L. Cell locomotion and focal adhesions are regulated by substrate flexibility. Proc. Natl. Acad. Sci. USA 1997, 94, 13661–13665. [Google Scholar] [CrossRef]
- Bischofs, I.B.; Schwarz, U.S. Cell organization in soft media due to active mechanosensing. Proc. Natl. Acad. Sci. USA 2003, 100, 9274–9279. [Google Scholar] [CrossRef] [PubMed]
- van Oers, R.; Rens, E.; LaValley, D.; Reinhart-King, C.; Merks, R. Mechanical Cell-Matrix Feedback Explains Pairwise and Collective Endothelial Cell Behavior In Vitro. PLoS Comput. Biol. 2014, 10, e1003774. [Google Scholar] [CrossRef]
- Schwarz, U.S.; Safran, S.A. Physics of adherent cells. Rev. Mod. Phys. 2013, 85, 1327–1381. [Google Scholar] [CrossRef]
- Yang, J.; Arratia, P.E.; Patteson, A.E.; Gopinath, A. Quenching active swarms: Effects of light exposure on collective motility in swarming Serratia marcescens. J. R. Soc. Interface 2019, 16, 1620180960. [Google Scholar] [CrossRef]
- Mandal, K.; Wang, I.; Vitiello, E.; Orellana, L.A.C.; Balland, M. Cell dipole behaviour revealed by ECM sub-cellular geometry. Nat. Commun. 2014, 5, 5749. [Google Scholar] [CrossRef] [PubMed]
- Dasbiswas, K.; Hu, S.; Schnorrer, F.; Safran, S.A.; Bershadsky, A.D. Ordering of myosin II filaments driven by mechanical forces: Experiments and theory. Philos. Trans. R. Soc. B Biol. Sci. 2018, 373, 20170114. [Google Scholar] [CrossRef] [PubMed]
- Mura, T. Micromechanics of Defects in Solids; Martnius Nijhoff: TheHague, The Netherlands, 1991. [Google Scholar]
- Ben-Yaakov, D.; Golkov, R.; Shokef, Y.; Safran, S.A. Response of adherent cells to mechanical perturbations of the surrounding matrix. Soft Matter 2015, 11, 1412–1424. [Google Scholar] [CrossRef]
- Golkov, R.; Shokef, Y. Shape regulation generates elastic interaction between living cells. New J. Phys. 2017, 19, 063011. [Google Scholar] [CrossRef]
- Fily, Y.; Subramanian, P.; Schneider, T.M.; Chelakkot, R.; Gopinath, A. Buckling instabilities and spatio-temporal dynamics of active elastic filaments. J. R. Soc. Interface 2019, 17, 20190794. [Google Scholar] [CrossRef] [PubMed]
- Notbohm, J.; Lesman, A.; Rosakis, P.; Tirrell, D.A.; Ravichandran, G. Microbuckling of fibrin provides a mechanism for cell mechanosensing. J. R. Soc. Interface 2015, 12, 20150320. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Safran, S.A. Nonlinearities of biopolymer gels increase the range of force transmission. Phys. Rev. E 2015, 92, 032728. [Google Scholar] [CrossRef] [PubMed]
- Golkov, R.; Shokef, Y. Elastic interactions between anisotropically contracting circular cells. Phys. Rev. E 2019, 99, 032418. [Google Scholar] [CrossRef]
- Bischofs, I.B.; Safran, S.A.; Schwarz, U.S. Elastic interactions of active cells with soft materials. Phys. Rev. E 2004, 69, 021911. [Google Scholar] [CrossRef]
- Phillips, R.; Kondev, J.; Theriot, J. Physical Biology of the Cell; Garland Science, Taylor & Francis Group: New York, NY, USA, 2008. [Google Scholar]
- Pei, D.; Shu, X.; Gassama-Diagne, A.; Thiery, J.P. Mesenchymal–epithelial transition in development and reprogramming. Nat. Cell Biol. 2019, 21, 44–53. [Google Scholar] [CrossRef]
- Bischofs, I.B.; Schwarz, U.S. Effect of Poisson Ratio on Cellular Structure Formation. Phys. Rev. Lett. 2005, 95, 068102. [Google Scholar] [CrossRef]
- Hong, J.; Ge, C.; Jothikumar, P.; Yuan, Z.; Liu, B.; Bai, K.; Li, K.; Rittase, W.; Shinzawa, M.; Zhang, Y.E.A. A TCR mechanotransduction signaling loop induces negative selection in the thymus. Nat. Immunol. 2018, 19, 1379–1390. [Google Scholar] [CrossRef]
- Kumari, S.; Mak, M.; Poh, Y.C.; Tohme, M.; Watson, N.; Melo, M.; Janssen, E.; Dustin, M.; Geha, R.; Irvine, D.J. Cytoskeletal tension actively sustains the migratory T-cell synaptic contact. EMBO J. 2020, 39, e102783. [Google Scholar] [CrossRef]
- Iskratsch, T.; Wolfenson, H.; Sheetz, M.P. Appreciating force and shape—The rise of mechanotransduction in cell biology. Nat. Rev. Mol. Cell Biol. 2014, 15, 825–833. [Google Scholar] [CrossRef] [PubMed]
- Huse, M. Mechanical forces in the immune system. Nat. Rev. Immunol. 2017, 17, 679–690. [Google Scholar] [CrossRef] [PubMed]
- Zhu, C.; Chen, W.; Lou, J.; Rittase, W.; Li, K. Mechanosensing through immunoreceptors. Nat. Immunol. 2019, 20, 1269–1278. [Google Scholar] [CrossRef] [PubMed]
- Huse, M. Integrins and their role in immune cell adhesion. Cell 2019, 177, 499–501. [Google Scholar]
- Mani, M.; Gopinath, A.; Mahadevan, L. How things get stuck: Kinetics, elastohydrodynamics, and soft adhesion. Phys. Rev. Lett. 2012, 108, 226104. [Google Scholar] [CrossRef] [PubMed]
- Sunyer, R.; Conte, V.; Escribano, J.; Elosegui-Artola, A.; Labernadie, A.; Valon, L.; Navajas, D.; García-Aznar, J.M.; Muñoz, J.J.; Roca-Cusachs, P.; et al. Collective cell durotaxis emerges from long-range intercellular force transmission. Science 2016, 353, 1157–1161. [Google Scholar] [CrossRef]
- Novikova, E.A.; Raab, M.; Discher, D.E.; Storm, C. Persistence-Driven Durotaxis: Generic, Directed Motility in Rigidity Gradients. Phys. Rev. Lett. 2017, 118, 078103. [Google Scholar] [CrossRef]
- Califano, J.P.; Reinhart-King, C.A. A Balance of Substrate Mechanics and Matrix Chemistry Regulates Endothelial Cell Network Assembly. Cell. Mol. Bioeng. 2008, 1, 122. [Google Scholar] [CrossRef]
- McKee, C.T.; Raghunathan, V.K.; Nealey, P.F.; Russell, P.; Murphy, C.J. Topographic Modulation of the Orientation and Shape of Cell Nuclei and Their Influence on the Measured Elastic Modulus of Epithelial Cells. Biophys. J. 2011, 9, 2139–2146. [Google Scholar] [CrossRef] [PubMed]
- Zhou, D.W.; Lee, T.T.; Weng, S.; Fuc, J.; García, A.J. Effects of substrate stiffness and actomyosin contractility on coupling between force transmission and vinculin–paxillin recruitment at single focal adhesions. Mol. Biol. Cell 2017, 28, 1901–1911. [Google Scholar] [CrossRef] [PubMed]
- Vargas, D.A.; Heck, T.; Smeets, B.; Ramon, H.; Parameswaran, H.; Van Oosterwyck, H. Intercellular adhesion stiffness moderates cell decoupling on stiff substrates. bioRxiv 2019. [Google Scholar] [CrossRef]
- Landau, L.D.; Lifshitz, E.M. Theory of Elasticity; Course of Theoretical Physics; Pergamon Press: London, UK, 1959; Volume 7. [Google Scholar]
| Quantity | Interpretation | Experimental Values |
|---|---|---|
| Cell size | 10–100 m | |
| T | Temperature | 25 °C |
| Thermal Diffusivity | 25 m2/min | |
| Effective Diffusivity | 3–50 m2/min | |
| E | Young’s modulus | 0.5–33 kPa |
| Poisson ratio | 0.3–0.5 | |
| P | Contractility | Nm |
| Parameter | Interpretation | Definition | Simulation Values |
|---|---|---|---|
| Diffusivity | 0.1–10 | ||
| Cell-cell interaction | 0.1–100 | ||
| Self-avoidance | – |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bose, S.; Dasbiswas, K.; Gopinath, A. Matrix Stiffness Modulates Mechanical Interactions and Promotes Contact between Motile Cells. Biomedicines 2021, 9, 428. https://doi.org/10.3390/biomedicines9040428
Bose S, Dasbiswas K, Gopinath A. Matrix Stiffness Modulates Mechanical Interactions and Promotes Contact between Motile Cells. Biomedicines. 2021; 9(4):428. https://doi.org/10.3390/biomedicines9040428
Chicago/Turabian StyleBose, Subhaya, Kinjal Dasbiswas, and Arvind Gopinath. 2021. "Matrix Stiffness Modulates Mechanical Interactions and Promotes Contact between Motile Cells" Biomedicines 9, no. 4: 428. https://doi.org/10.3390/biomedicines9040428
APA StyleBose, S., Dasbiswas, K., & Gopinath, A. (2021). Matrix Stiffness Modulates Mechanical Interactions and Promotes Contact between Motile Cells. Biomedicines, 9(4), 428. https://doi.org/10.3390/biomedicines9040428






