Natural Compounds as Target Biomolecules in Cellular Adhesion and Migration: From Biomolecular Stimulation to Label-Free Discovery and Bioactivity-Based Isolation
Abstract
1. Introduction
2. Relationship between Adhesion, Movement and Inflammation
3. Mechanisms of Action of Natural Compounds
3.1. Prestimulation with Cytokines
3.2. Inhibition of CAMs by Suppression of Their Expression (Downregulation)
3.3. Mechanism of Downregulation
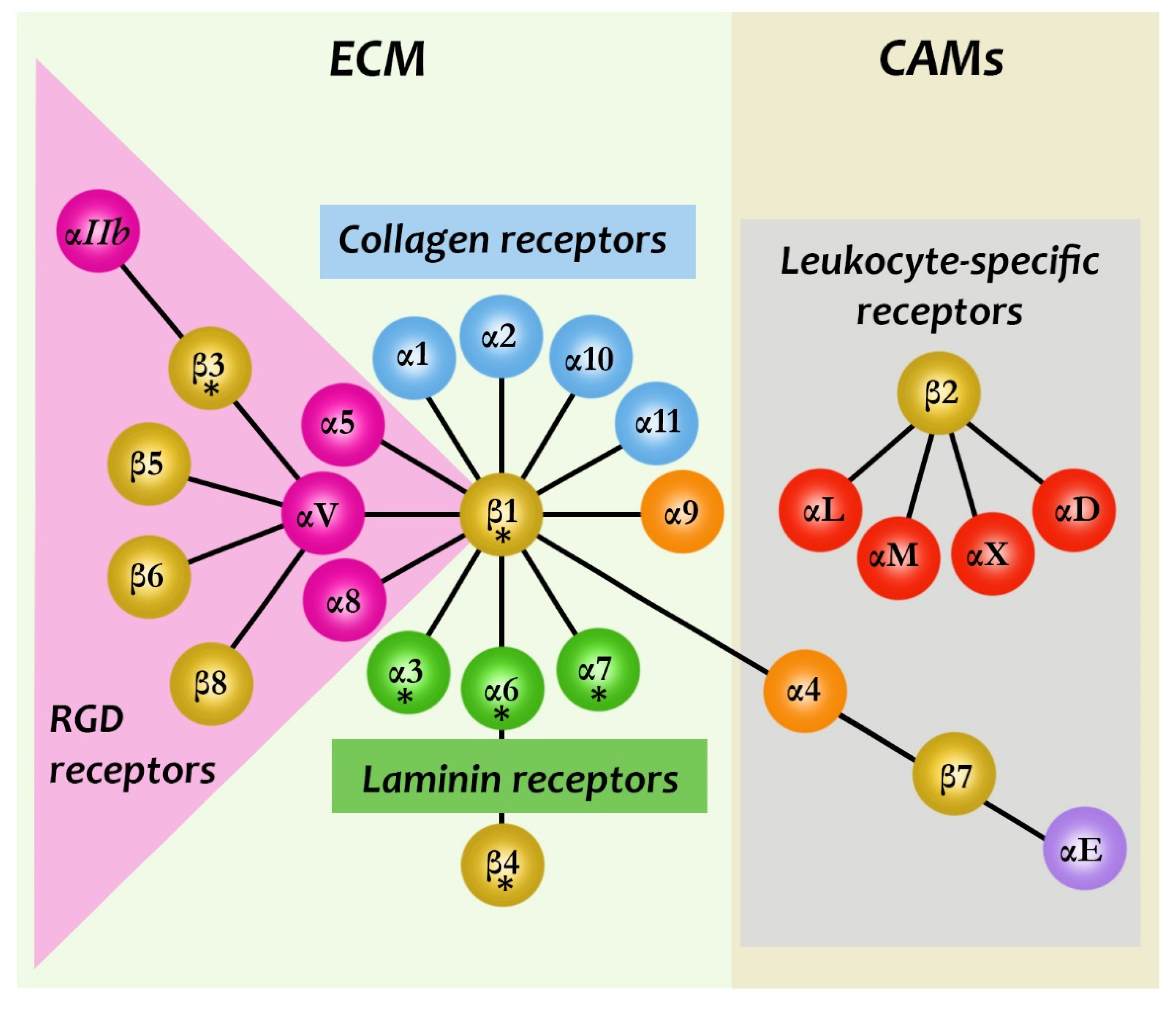
3.4. Intervention at the ECM
3.5. Inhibition of CAM Binding by Blocking Specific Cell–Surface Receptor Sites
4. Measurement Techniques for Monitoring Cellular Functions, Adhesion and Viability
4.1. Classical Techniques for Measuring Cell Viability
4.2. Limitations and Considerations When Using Membrane Integrity or Metabolic Assays
4.3. Measurement Techniques for Monitoring Cellular Movement and Adhesion
4.3.1. Label-Free Biosensors
4.3.2. Living Cell Movements—Holographic Microscopy
5. Preparation of Natural Compounds
6. Natural Compounds of Plant, Animal and Fungal Origin
| Active Substance | IUPACName (CAS Number) | Source | Cellular Effect | Effect Type | Molecular Mechanisms | Tested Cell Line 1, Animal | Method | Ref. |
|---|---|---|---|---|---|---|---|---|
| Ganoderma lucidum polysaccharide | Ganoderma lucidum | Inhibition of tumour cell adhesion | P | Ateration in β1-integrin expression | MT-1 | Coomassie blue staining, Western blot | Wu et al. 2006 [47] | |
| Rhodostomin peptide  | Pichia pastoris | Inhibition of cell adhesion, platelet aggregation and the binding of fibrinogen to platelet by ARGDWN mutants | B | Prevention of integrin αIIbβ3 interaction | CHO K562 | Mass spectrometry, Fibrinogen binding assay, Flow cytometry, Platelet aggregation assay, Nuclear magnetic resonance spectroscopy, Molecular docking | Chang et al. 2017 [56] | |
Cyclopeptolide HUN-7293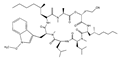 | 3-((2R,5S,8S,11S,14S,17R,20S)-8,11-diisobutyl-14-((1-methoxy-1H-indol-3-yl)methyl)-7,13,19,20-tetramethyl-18-methylene-5-((R)-2-methylheptyl)-17-((R)-2-methylhexyl)-3,6,9,12,15,21-hexaoxo-1-oxa-4,7,10,13,16,19-hexaazacyclohenicosan-2-yl)propanenitrile (129893-84-1) | Bartalinia robillardoides | Inhibition of VCAM-1 and ICAM-1 | P | Inhibition of the expression of VCAM-1 and ICAM-1 | HMEC-1 | ELISA | Schreiner et al. 2004 [242] |
Cytochalasin-E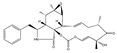 | (1E,4S,6R,7E,11aS,14S,14aS,15S,15aR,16aR,16bS)-14-benzyl-6-hydroxy-4,6,15,15a-tetramethyl-3,13,14,14a,15,15a,16a,16b-octahydro-[1,3]dioxacyclotridecino[4,5-d]oxireno[2,3-f]isoindole-5,10,12(4H,6H)-trione (36011-19-5) | Mycotypha sp. UMF-006 (soil) | Inhibition of cell aggregation and adhesion | No data | No data | HL-60 CHO | Cell aggregation assay, Adhesion assay (CFSE-labelling | Takamatsu et al.n 2002 [243] |
Cordyceptin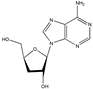 | (2R,5S)-2-(6-amino-9H-purin-9-yl)-5-(hydroxymethyl)tetrahydrofuran-3-ol (73-03-0) | Cordiceps militaris | Inhibition of motility, invasion and migration | P | Inhibition of PI3K/Akt pathway and expression of claudin family proteins. Downregulation of MMP activity | LNCaP | Wound healing migration assay, MTT assay, Boyden chamber, Matrigel invasion assay, Measurement of TER (EVOM voltohmmeter) | Jeong et al. 2012 [244] |
Fusarisetin A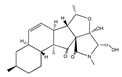 | (3S,3aR,5S,5aS,5bS,7aS,9R,11aR,11bS,12aR)-3a-hydroxy-3-(hydroxymethyl)-2,5,9,11b-tetramethyl-3,3a,5,5a,5b,7a,8,9,10,11,11a,11b-dodecahydro-12H-benzo[4′,5′]indeno[2′,1′:3,4]furo[2,3-c]pyrrole-1,12(2H)-dione (1300041-53-5) | Fusarium sp. FN080326 | Inhibition of migration | P | Pathway inhibition | MDA-MB-231 | Scratch wound assay, Boyden chamber transwell assay | Xu et al. 2012 [245] |
Terrein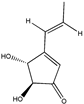 | (4R,5S)-4,5-dihydroxy-3-((E)-prop-1-en-1-yl)cyclopent-2-en-1-one (582-46-7) | Aspergillus terreus CRI301 | Inhibition of adhesion and migration | P | Downregulation of MMP-2 and MMP-9 transcription. Reduction of protein levels for the Rho GTPases | MDA-MB-231 MCF-7 | Cell adhesion assay (crystal violet staining), Wound healing assay, Matrigel cel migration assay | Kasorn et al. 2018 [246] |
Ophiobolin A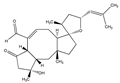 | (3S,3aR,3′S,5′R,6aS,9R,9aS,10aR,E)-9-hydroxy-3′,9,10a-trimethyl-5′-(2-methylprop-1-en-1-yl)-7-oxo-1,3a,4,4′,5′,6a,7,8,9,9a,10,10a-dodecahydro-2H,3′H-spiro[dicyclopenta[a,d][8]annulene-3,2′furan]-6-carbaldehyde (4611-05-6) | Drechslera gigantea | Decreased the 2D-migration potential | No data | No data | U373 MG Glioblastoma multiforme (GBM) | MTT assay, Quantitative video microscopy, Immunofluorescence/pseudo-confocal microscopy, Flow cytometry | Bury et al. 2013 [247] |
Fusicoccin A | (2S)-2-((1S,4R,5S,6S,9S,10aR,E)-4-(((2S,3R,4S,5R,6R)-4-acetoxy-3,5-dihydroxy-6-(((2-methylbut-3-en-2-yl)oxy)methyl)tetrahydro-2H-pyran-2-yl)oxy)-6-formyl-1,5-dihydroxy-9-(methoxymethyl)-6,10a-dimethyl-1,2,4,5,6,6a,7,8,9,10a-decahydrodicyclopenta[a,d][8]annulen-3-yl)propyl acetate (20108-30-9) | Fusicoccum amygdali | Decreased the 2D and 3D-migration potential. Decreased adhesion | No data | No data | U373-MG Glioblastoma multiforme (GBM) | Quantitative video microscopy, Boyden chamber assay, In vitro adhesion assay (hematoxylin staining) | Bury et al. 2013 [248] |
Altersolanol A | (1R,2S,3R,4S)-1,2,3,4,5-pentahydroxy-7-methoxy-2-methyl-1,2,3,4-tetrahydroanthracene-9,10-dione (22268-16-2) | Stemphylium globuliferum | Inhibition of migration | P | Inhibition of TNFα-activated NF-κB signalling pathway. Inhibition of NF-κB-mediated MMP expression | A549 | Incucyte Live-Cell imaging system, Videomicroscopy analysis, Scratch wound assay, flow cytometry, Western blot | Teiten et al. 2013 [249] |
Sphaeropsidin A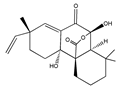 | (2R,4aR,4bR,8aS,9S)-4a,9-dihydroxy-2,8,8-trimethyl-2-vinyl-2,3,4,4a,5,6,7,8,8a,9-decahydro-10H-9,4b-(epoxymethano)phenanthrene-10,12-dione (38991-80-9) | Smardea sp. AZ0432 | Inhibition of migration | No data | No data | MDA-MB-231 | Wound healing assay | Wang et al. 2011 [250] |
Gliotoxin | (3R,5aS,6S,10aR)-6-hydroxy-3-(hydroxymethyl)-2-methyl-2,3,5a,6-tetrahydro-10H-3,10a-epidithiopyrazino[1,2-a]indole-1,4-dione (67-99-2) | Unidentified fungal strains Y90086/Y80805 | Inhibition of migration | No data | No data | HUVEC | Wound migration assay | Lee et al. 2001 [251] |
Methylthiogliotoxin | (3R,5aS,6S,10aR)-6-hydroxy-3-(hydroxymethyl)-2-methyl-3,10a-bis(methylthio)-2,3,5a,6,10,10a-hexahydropyrazino[1,2-a]indole-1,4-dione (74149-38-5) | |||||||
Cytochalasin D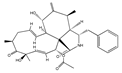 | (3S,3aR,4S,6S,6aR,7E,10S,12R,13E,15R,15aS)-3-benzyl-6,12-dihydroxy-4,10,12-trimethyl-5-methylene-1,11-dioxo-2,3,3a,4,5,6,6a,9,10,11,12,15-dodecahydro-1H-cycloundeca[d]isoindol-15-yl acetate (22144-77-0) | Not mentioned (but many fungi produce, e.g., Zygosporium mansonii) | Inhibition of actin polymerization, decreased cell motility and colonization. Inhibition of actin polymerization | No data | No data | MFC-7 A549 | Quantitative videomicroscopy, Scratch wound assay | Hayot et al. 2006 [252] |
FTY720 (fingolimod)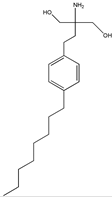 | 2-amino-2-(4-octylphenethyl)propane-1,3-diol (162359-55-9) | Isaria sinclairii | Inhibition of cell migration, motility and invasion | P | Inhibition of RhoA-GTPase expression | DU145 PC3 | Wound closure assay, 3D collagen colony forming assay | Zhou et al. 2006 [253] |
| Staitins | ||||||||
Lovastatin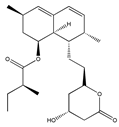 * (Fluvastatin (syn))  | (1S,3R,7S,8S,8aR)-8-(2-((2S,4R)-4-hydroxy-6-oxotetrahydro-2H-pyran-2-yl)ethyl)-3,7-dimethyl-1,2,3,7,8,8a-hexahydronaphthalen-1-yl (S)-2-methylbutanoate (75330-75-5) | Pleurotus ostreatus ** | Inhibition of EGF-induced migration and invasiveness | P | Inhibition of translocation of RhoA | PANC-1 | Matrigel invasion assay | Kusama et al. 2001 [254] |
| (3S,5R,E)-7-(3-(4-fluorophenyl)-1-isopropyl-1H-indol-2-yl)-3,5-dihydroxyhept-6-enoic acid (93957-54-1) | Inhibition of tumor cell attachement and migration | No data | No data | F3II | Adhesion assay, Migration assay (in vitro wound assay) | Alonso et al., 1998 [255] | ||
| Suppressed the EGF-induced cell adhesion, actin filament reorganization and transmigration | P | Suppression of Rho/ROCK and FAK/paxillin signaling | ARO | Matrigel invasion assay, Cell adhesion assay | Zhong et al., 2005 [256] | |||
| Inhibition of adhesion. Reduced tumor cell migration, attachement and motility. Changed the tumor cell shape | P | Reduced membrane localization of Rho protein | F3II | Adhesion assay, Migration assay (in vitro wound assay) | Farina et al., 2002 [257] | |||
| Reduced migration and invasion | P | Diminished ERK signaling. Impaired the regulation of the mevalonate- and the Ras-Raf-MEK-ERK pathway. Affected the post-translational modification of H-Ras and Rac1 | U87 U343 | Migration assay, Matrigel invasion assay | Afshordel et al., 2014 [258] | |||
Lovastatin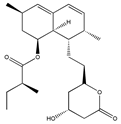 | (1S,7S,8S,8aR)-8-(2-((2R,4R)-4-hydroxy-6-oxotetrahydro-2H-pyran-2-yl)ethyl)-7-methyl-1,2,3,7,8,8a-hexahydronaphthalen-1-yl (S)-2-methylbutanoate (73573-88-3) | Pleurotus ostreatus ** | Inhibition of invasion and migration | P | Alteration in expression of matrix-metalloproteases | M14 | Matrigel invasion assay, Integrin-mediated binding assays | Glynn et al., 2008 [259] |
Mevastatin | (1S,7S,8S,8aR)-8-(2-((2R,4R)-4-hydroxy-6-oxotetrahydro-2H-pyran-2-yl)ethyl)-7-methyl-1,2,3,7,8,8a-hexahydronaphthalen-1-yl (S)-2-methylbutanoate (73573-88-3) | Penicillium citrinum ** | HT144 | |||||
Simvastain (syn)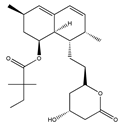 | (1S,3R,7S,8S,8aR)-8-(2-((2S,4R)-4-hydroxy-6-oxotetrahydro-2H-pyran-2-yl)ethyl)-3,7-dimethyl-1,2,3,7,8,8a-hexahydronaphthalen-1-yl 2,2-dimethylbutanoate (79902-63-9) | Aspergillus terreus | SK-MEL-28 | |||||
| Macrosphelides (MSs) | ||||||||
| Macrosphelide A 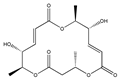 | (4S,7E,9R,10S,13E,15R,16S)-9,15-dihydroxy-4,10,16-trimethyl-1,5,11-trioxacyclohexadeca-7,13-diene-2,6,12-trione (172923-77-2) | Microsphaeropsis sp. FO-5050 (soil) | Inhibition of HL-60 cell adhesion to HUVEC | B | Blocking the binding of SLex to ELAM-1 | HL-60 | Cell adhesion assay | Hayashi et al., 1995 [260] |
B | (4S,7E,9R,10S,13E,16S)-9-hydroxy-4,10,16-trimethyl-1,5,11-trioxacyclohexadeca-7,13-diene-2,6,12,15-tetraone (172923-78-3) | HUVEC | ||||||
| Macrosphelide C 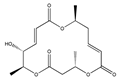 | (4S,7E,10S,13E,15R,16S)-15-hydroxy-4,10,16-trimethyl-1,5,11-trioxacyclohexadeca-7,13-diene-2,6,12-trione (199731-56-1) | Microsphaeropsis sp. FO-5050 (soil) | Inhibition of adhesion | No data | Not discussed | HL-60 | Cell adhesion assay | Takamatsu et al., 1997 [261] |
D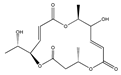 | (2S,6R,7E,11S,13E)-12-hydroxy-6-((S)-1-hydroxyethyl)-2,11-dimethyl-1,5,10-trioxacyclopentadeca-7,13-diene-4,9,15-trione (199731-57-2) | HUVEC | ||||||
Macrosphelide J | (4S,9R,10S,13S,16S,E)-9-hydroxy-13-methoxy-4,10,16-trimethyl-1,5,11-trioxacyclohexadec-7-ene-2,6,12,15-tetraone (239094-07-6) | Microsphaeropsis sp. FO-5050 (soil) | No effect | No effect | No effect | HL-60 HUVEC | Cell adhesion assay | Fukami et al., 1999 [262] |
K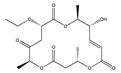 | (4S,9R,10S,13S,16S,E)-13-ethoxy-9-hydroxy-4,10,16-trimethyl-1,5,11-trioxacyclohexadec-7-ene-2,6,12,15-tetraone | |||||||
epi-5’-hydroxymycosporulone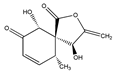 | (4S,5R,6R,10R)-4,6-dihydroxy-10-methyl-3-methylene-2-oxaspiro[4.5]dec-8-ene-1,7-dione (238735-98-3) | |||||||
Macroshelide A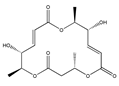 and structurally related compounds of Macroshelide A (Macroshelide C, E, F, G, H, I, L and M) | (4S,7E,9R,10S,13E,15R,16S)-9,15-dihydroxy-4,10,16-trimethyl-1,5,11-trioxacyclohexadeca-7,13-diene-2,6,12-trione (172923-77-2) | Periconia byssoides OUPS-N133 | Inhibition of HL-60 cell adhesion to HUVEC | No data | No data | HL-60 HUVEC | Cell adhesion assay with MTT staining (“Miki’s method” [263]) | Yamada et al., 2001, 2002, 2007 [264,265,266] |
Macroshelide B | (4S,7E,9R,10S,13E,16S)-9-hydroxy-4,10,16-trimethyl-1,5,11-trioxacyclohexadeca-7,13-diene-2,6,12,15-tetraone (172923-78-3) | Microsphaeropsis sp. FO-5050 (soil) | Inhibition of cell binding to HUVEC | B, P | Blocking the binding of sLex to E-selectin. Suppressed the expression of adhesion molecules. | B16/BL6 L5178Y-ML HUVEC | Adhesion assay (CFSE-labelling), flow cytometry analysis | Fukami et al., 2002 [267] |
| Peribysins | ||||||||
Peribysin A and structurally related compounds of Peribysin A (Peribysin B, C, D, E, F, G, H, I and J ) | (1aR,2S,3aR,7S,7aR,7bS)-1a-(3-hydroxyprop-1-en-2-yl)-7,7a-dimethyldecahydronaphtho[1,2-b]oxiren-2-ol | Periconia byssoides OUPS-N133 | Inhibition of HL-60 cell adhesion to HUVEC | No data | No data | HL-60 HUVEC | Cell adhesion assay with MTT staining (“Miki’s method” [263]) | Yamada et al., 2004, 2005, 2006, 2007 [266,268,269,270] |
7. SARS-CoV-2 and Possible Treatments with Herbal Extracts
8. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Peter, B.; Bosze, S.; Horvath, R. Biophysical characteristics of living cells exposed to the green tea polyphenol epigallocatecin-3-gallate (EGCg): Review of recent advances from molecular mechanisms to clinical trials. Eur. Biophys. J. 2016, 46, 1–35. [Google Scholar] [CrossRef]
- Ling, S.; Nheu, L.; Dai, A.; Guo, Z.; Komesaroff, P. Effects of four medicinal herbs on human vascular endothelial cells in culture. Int. J. Cardiol. 2008, 128, 350–358. [Google Scholar] [CrossRef] [PubMed]
- Kinghorn, A.D.; De Blanco, E.J.C.; Chai, H.; Orjala, J.; Farnsworth, N.R.; Soejarto, D.D.; Oberlies, N.H.; Wani, M.C.; Kroll, D.J.; Pearce, C.J.; et al. Discovery of anticancer agents of diverse natural origin. Pure Appl. Chem. 2009, 81, 1051–1063. [Google Scholar] [CrossRef]
- Gumbiner, B.M. Cell adhesion: The molecular basis of tissue architecture and morphogenesis. Cell 1996, 84, 345–357. [Google Scholar] [CrossRef]
- Spelman, K.; Aldag, R.; Hamman, A.; Kwasnik, E.M.; Mahendra, M.A.; Obasi, T.M.; Morse, J.; Williams, E.J. Traditional herbal remedies that influence cell adhesion molecule activity. Phytother. Res. 2011, 25, 473–483. [Google Scholar] [CrossRef]
- Goldstein, I.M.; Malmsten, C.L.; Samuelsson, B.; Weissmann, G. Prostaglandins, thromboxanes, and polymorphonuclear leukocytes. Inflammation 1977, 2, 309–317. [Google Scholar] [CrossRef] [PubMed]
- Chang, D.-M.; Kuo, S.-Y.; Lai, J.-H.; Chang, M.-L. Effects of anti-rheumatic herbal medicines on cellular adhesion molecules. Ann. Rheum. Dis. 1999, 58, 366–371. [Google Scholar] [CrossRef] [PubMed]
- Bevilacqua, M.P.; Pober, J.S.; Wheeler, M.E.; Cotran, R.S.; Gimbrone, M.A. Interleukin 1 acts on cultured human vascular endothelium to increase the adhesion of polymorphonuclear leukocytes, monocytes, and related leukocyte cell lines. J. Clin. Investig. 1985, 76, 2003–2011. [Google Scholar] [CrossRef]
- Cronstein, B.N.; Weissmann, G. The adhesion molecules of inflammation. Arthritis Rheum. 1993, 36, 147–157. [Google Scholar] [CrossRef]
- Hynes, R.O. Integrins: Bidirectional, Allosteric Signaling Machines. Cell 2002, 110, 673–687. [Google Scholar] [CrossRef]
- Bevilacqua, M.P. Endothelial-leukocyte cellular adhesion molecules. Annu. Rev. Immunol. 1993, 11, 767–804. [Google Scholar] [CrossRef]
- Grober, J.S.; Bowen, B.L.; Ebling, H.; Athey, B.; Thompson, C.B.; Fox, D.A.; Stoolman, L.M. Monocyte-endothelial adhesion in chronic rheumatoid arthritis: In situ detection of selectin and integrin-dependent interactions. J. Clin. Investig. 1993, 91, 2609–2619. [Google Scholar] [CrossRef]
- Kawasaki, K.; Muroyama, K.; Yamamoto, N.; Murosaki, S. A hot water extract of Curcuma longa inhibits adhesion molecule protein expression and monocyte adhesion to TNF-alpha-stimulated human endothelial cells. Biosci. Biotechnol. Biochem. 2015, 79, 1654–1659. [Google Scholar] [CrossRef]
- Ramadori, G.; Moriconi, F.; Malik, I.; Dudas, J. Physiology and pathophysiology of liver inflammation, damage and repair. J. Physiol. Pharmacol. 2008, 59, 107–117. [Google Scholar]
- Parsons, J.T.; Horwitz, A.R.; Schwartz, M.A. Cell adhesion: Integrating cytoskeletal dynamics and cellular tension. Nat. Rev. Mol. Cell Biol. 2010, 11, 633–643. [Google Scholar] [CrossRef]
- Mezu-Ndubuisi, O.J.; Maheshwari, A. The role of integrins in inflammation and angiogenesis. Pediatr. Res. 2020, 89, 1619–1626. [Google Scholar] [CrossRef]
- Yue, J.; Zhang, K.; Chen, J.F. Role of integrins in regulating proteases to mediate extracellular matrix remodeling. Cancer Microenviron. 2012, 5, 275–283. [Google Scholar] [CrossRef]
- Papoutsi, Z.; Kassi, E.; Chinou, I.; Halabalaki, M.; Skaltsounis, L.A.; Moutsatsou, P. Walnut extract (Juglans regia L.) and its component ellagic acid exhibit anti-inflammatory activity in human aorta endothelial cells and osteoblastic activity in the cell line KS483. Br. J. Nutr. 2008, 99, 715–722. [Google Scholar] [CrossRef] [PubMed]
- Tang, P.; Hung, M.C.; Klostergaard, J. Human pro-tumor necrosis factor is a homotrimer. Biochemistry 1996, 35, 8216–8225. [Google Scholar] [CrossRef]
- Spelman, K.; Burns, J.J.; Nichols, D.; Winters, N.; Ottersberg, S.; Tenborg, M. Modulation of Cytokine Expression by Traditional Medicines: A Review of Herbal Immunomodulators. Altern. Med. Rev. 2006, 11, 128–150. [Google Scholar]
- Carluccio, M.A.; Siculella, L.; Ancora, M.A.; Massaro, M.; Scoditti, E.; Storelli, C.; Visioli, F.; Distante, A.; De Caterina, R. Olive oil and red wine antioxidant polyphenols inhibit endothelial activation: Antiatherogenic properties of Mediterranean diet phytochemicals. Arterioscler. Thromb. Vasc. Biol. 2003, 23, 622–629. [Google Scholar] [CrossRef]
- Zhang, W.-J.; Hufnag, P.; Binder, B.R.; Wojta, J. Antiinflammatory activity of astragaloside IV is mediated by inhibition of NF-κB activation and adhesion molecule expression. Thromb. Haemost. 2003, 90, 904–914. [Google Scholar] [CrossRef]
- Tao, X.; Lipsky, P.E. The Chinese anti-inflammatory and immunosuppressive herbal remedy Tripterygium Wilfordii Hook F. Rheum. Dis. Clin. N. Am. 2000, 26, 29–50. [Google Scholar] [CrossRef]
- Iademarco, M.F.; McQuillan, J.J.; Rosen, G.D.; Dean, D.C. Characterization of the promoter for vascular cell adhesion molecule-1 (VCAM-1). J. Biol. Chem. 1992, 267, 16323–16329. [Google Scholar] [CrossRef]
- Schindler, U.; Baichwal, V.R. Three NF-kappa B binding sites in the human E-selectin gene required for maximal tumor necrosis factor alpha-induced expression. Mol. Cell. Biol. 1994, 14, 5820–5831. [Google Scholar] [CrossRef]
- Collins, T.; Read, M.A.; Neish, A.S.; Whitley, M.Z.; Thanos, D.; Maniatis, T. Transcriptional regulation of endothelial cell adhe- sion molecules: NF-kappa B and cytokine-inducible enhancers. FASEB J. 1995, 10, 899–909. [Google Scholar] [CrossRef]
- Voraberger, G.; Schäfer, R.; Stratowa, C. Cloning of the human gene for intercellular adhesion molecule 1 and analysis of its 5′- regulatory region. Induction by cytokines and phorbol ester. J. Immunol. 1991, 147, 2777–2786. [Google Scholar]
- Tak, P.P.; Firestein, G.S. NF-κB: A key role in inflammatory diseases. J. Clin. Investig. 2001, 107, 7–11. [Google Scholar] [CrossRef] [PubMed]
- Baldwin, A.S., Jr. The NF-κB and I kappa B proteins: New discoveries and insights. Annu. Rev. Immunol. 1996, 14, 649–683. [Google Scholar] [CrossRef]
- Kurucz, I.; Peter, B.; Prosz, A.; Szekacs, I.; Erdei, A.; Horvath, R. Label-free optical biosensor for on-line monitoring the integrated response of human B cells upon the engagement of stimulatory and inhibitory immune receptors. Sens. Actuators B 2017, 240, 528–535. [Google Scholar] [CrossRef]
- Erdei, A.; Sándor, N.; Szilvia, B.M.; Kremlitzka, M.; Bajtay, Z. The versatile functions of complement C3-derived ligands. Immunol. Rev. 2016, 274, 127–140. [Google Scholar] [CrossRef]
- Gorfu, G.; Rivera-Nieves, J.; Ley, K. Role of β7 integrins in intestinal lymphocyte homing and retention. Curr. Mol. Med. 2009, 9, 836–850. [Google Scholar] [CrossRef]
- Caswell, P.T.; Vadrevu, S.; Norman, J.C. Integrins: Masters and slaves of endocytic transport. Nat. Rev. Mol. Cell Biol. 2009, 10, 843–853. [Google Scholar] [CrossRef]
- Liu, X.; Wang, W.; Song, G.; Wei, X.; Zeng, Y.; Han, P.; Wang, D.; Shao, M.; Wu, J.; Sun, H.; et al. Astragaloside IV ameliorates diabetic nephropathy by modulating the mitochondrial quality control network. PLoS ONE 2017, 12, e0182558. [Google Scholar] [CrossRef]
- Date, R. Research review on the pharmacological effects of Astragaloside IV. Fundam. Clin. Pharmacol. 2017, 31, 17–36. [Google Scholar] [CrossRef]
- Gui, D.; Huang, J.; Guo, Y.; Chen, J.; Chen, Y.; Xiao, W.; Liu, X.; Wang, N. Cytokine Astragaloside IV ameliorates renal injury in streptozotocin-induced diabetic rats through inhibiting NF- j B-mediated inflammatory genes expression. Cytokine 2013, 61, 970–977. [Google Scholar] [CrossRef]
- Wang, Z.S.; Xiong, F.; Xie, X.H.; Chen, D.; Pan, J.H.; Cheng, L. Astragaloside IV attenuates proteinuria in streptozotocin-induced diabetic nephropathy via the inhibition of endoplasmic reticulum stress. Nephrology 2015, 16, 44. [Google Scholar] [CrossRef]
- Chen, J.; Chen, Y.; Luo, Y.; Gui, D.; Huang, J.; He, D. Astragaloside IV ameliorates diabetic nephropathy involving protection of podocytes in streptozotocin induced diabetic rats. Eur. J. Pharmacol. 2014, 736, 86–94. [Google Scholar] [CrossRef]
- Cheng, S.; Wu, Y.; Huang, W.; Pang, J.S. Cytokine Anti-in fl ammatory property of quercetin through downregulation of ICAM-1 and MMP-9 in TNF- α -activated retinal pigment epithelial cells. Cytokine 2019, 116, 48–60. [Google Scholar] [CrossRef]
- Lee, J.; Zhou, H.Y.; Cho, S.Y.; Kim, Y.S.; Lee, Y.S.; Jeong, C.S. Anti-inflammatory Mechanisms of Apigenin: Inhibition of Cyclooxygenase-2 Expression, Adhesion of Monocytes to Human Umbilical Vein Endothelial Cells, and Expression of Cellular Adhesion Molecules. Arch. Pharm. Res. 2007, 30, 1318–1327. [Google Scholar] [CrossRef] [PubMed]
- Journal, A.I.; Song, Y.; Tian, X.; Wang, X.; Feng, H. Vascular protection of salicin on IL-1 β -induced endothelial inflammatory response and damages in retinal endothelial cells. Artif. Cells Nanomed. Biotechnol. 2019, 47, 1995–2002. [Google Scholar] [CrossRef]
- Rho, M.; Kwon, O.E.; Kim, K.; Lee, S.W.; Chung, M.Y.; Kim, Y.H.; Hayashi, M.; Lee, H.S.; Kim, Y. Inhibitory Effects of Manassantin A and B Isolated from the Roots of Saururus chinensis on PMA-Induced ICAM-1 Expression. Planta Med. 2003, 69, 1147–1149. [Google Scholar]
- Choi, Y.; Jin, H.; Park, S.; Chung, J.; Lee, H.; Oh, S.; Kim, B.; Kim, J.; Chung, H.; Yu, B.; et al. Inhibition of endothelial cell adhesion by the new anti-inflammatory agent α -iso-cubebene. Vasc. Pharmacol. 2009, 51, 215–224. [Google Scholar] [CrossRef]
- Jiang, C.; Li, J.; Liu, F.; Wu, T.A.O.; Yu, M.; Xu, H. Andrographolide Inhibits the Adhesion of Gastric Cancer Cells to Endothelial Cells by Blocking E-selectin Expression. Anticancer Res. 2007, 2448, 2439–2447. [Google Scholar]
- Wu, S.; Xu, H.; Peng, J.; Wang, C.; Jin, Y.; Liu, K. Biochimie Potent anti-inflammatory effect of dioscin mediated by suppression of TNF- a -induced VCAM-1, ICAM-1and EL expression via the NF- k B pathway. Biochimie 2015, 110, 62–72. [Google Scholar] [CrossRef]
- Kumar, S.; Arya, P.; Mukherjee, C.; Singh, B.K.; Singh, N.; Parmar, V.S.; Prasad, A.K.; Ghosh, B. Novel aromatic ester from Piper longum and its analogues inhibit expression of cell adhesion molecules on endothelial cells. Biochemistry 2005, 44, 15944–15952. [Google Scholar] [CrossRef]
- Wu, Q.P.; Xie, Y.Z.; Li, S.Z.; La Pierre, D.P.; Deng, Z.; Chen, Q.; Li, C.; Zhang, Z.; Guo, J.; Wong, C.K.A.; et al. Tumour cell adhesion and integrin expression affected by Ganoderma lucidum. Enzyme Microb. Technol. 2006, 40, 32–41. [Google Scholar] [CrossRef]
- Wang, L.; Ling, Y.; Chen, Y.; Li, C.L.; Feng, F.; You, Q.D.; Lu, N.; Guo, Q.L. Flavonoid baicalein suppresses adhesion, migration and invasion of MDA-MB-231 human breast cancer cells. Cancer Lett. 2010, 297, 42–48. [Google Scholar] [CrossRef]
- Jang, J.H.; Yang, E.S.; Min, K.-J.; Kwon, T.K. Inhibitory effect of butein on tumor necrosis factor-α-induced expression of cell adhesion molecules in human lung epithelial cells via inhibition of reactive oxygen species generation, NF-κB activation and Akt phosphorylation. Int. J. Mol. Med. 2012, 30, 1357–1364. [Google Scholar] [CrossRef]
- Peter, B.; Farkas, E.; Forgacs, E.; Saftics, A.; Kovacs, B.; Kurunczi, S.; Szekacs, I.; Csampai, A.; Bosze, S.; Horvath, R. Green tea polyphenol tailors cell adhesivity of RGD displaying surfaces: Multicomponent models monitored optically. Nat. Publ. Gr. 2017, 7, 42220. [Google Scholar] [CrossRef]
- Szekacs, I.; Orgovan, N.; Peter, B.; Kovacs, B.; Horvath, R. Receptor specific adhesion assay for the quantification of integrin–ligand interactions in intact cells using a microplate based, label-free optical biosensor. Sens. Actuators B Chem. 2018, 256, 729–734. [Google Scholar] [CrossRef]
- Cox, D.; Brennan, M.; Moran, N. Integrins as therapeutic targets: Lessons and opportunities. Nat. Rev. Drug Discov. 2010, 9, 804–820. [Google Scholar] [CrossRef] [PubMed]
- Ley, K.; Rivera-Nieves, J.; Sandborn, W.J.; Shattil, S. Integrin-based therapeutics: Biological basis, clinical use and new drugs. Nat. Rev. Drug Discov. 2016, 15, 173–183. [Google Scholar] [CrossRef]
- McLane, M.A.; Sanchez, E.E.; Wong, A.; Paquette-Straub, C.; Perez, J.C. Disintegrins. Curr. Drug Targets Cardiovasc. Haematol. Disord. 2004, 4, 327–355. [Google Scholar] [CrossRef] [PubMed]
- Wu, W.-B.; Peng, H.-C.; Huang, T.-F. Disintegrin causes proteolysis of β-catenin and apoptosis of endothelial cells: Involvement of cell—cell and cell—ECM interactions in regulating cell viability. Exp. Cell Res. 2003, 286, 115–127. [Google Scholar] [CrossRef]
- Chang, Y.; Shiu, J.; Huang, C.; Chen, Y.; Chen, C.; Chang, Y.; Chuang, W. Effects of the RGD loop and C-terminus of rhodostomin on regulating integrin α IIb β 3 recognition. PLoS ONE 2017, 12, e0175321. [Google Scholar] [CrossRef]
- McColl, J.; Horvath, R.; Yakubov, G.E.; Ramsden, J.J. Surface rearrangement of adsorbed EGCG–mucin complexes on hydrophilic surfaces. Int. J. Biol. Macromol. 2017, 95, 704–712. [Google Scholar] [CrossRef]
- Habtemariam, S. Cistifolin, an Integrin-Dependent Cell Adhesion Blocker from the Anti- Rheumatic Herbal Drug, Gravel Root (Rhizome of Eupatorium purpureum). Planta Med. 1998, 64, 683–685. [Google Scholar] [CrossRef]
- Peter, B.; Ungai-Salanki, R.; Szabó, B.; Nagy, A.G.; Szekacs, I.; Bösze, S.; Horvath, R. High-Resolution Adhesion Kinetics of EGCG-Exposed Tumor Cells on Biomimetic Interfaces: Comparative Monitoring of Cell Viability Using Label-Free Biosensor and Classic End-Point Assays. ACS Omega 2018, 3, 3882–3891. [Google Scholar] [CrossRef]
- Cheung, K.C.; Di Berardino, M.; Schade-Kampmann, G.; Hebeisen, M.; Pierzchalski, A.; Bocsi, J.; Mittag, A.; Tárnok, A. Microfluidic impedance-based flow cytometry. Cytom. Part A 2010, 77, 648–666. [Google Scholar] [CrossRef]
- Wang, P.; Henning, S.M.; Heber, D. Limitations of MTT and MTS-based assays for measurement of antiproliferative activity of green tea polyphenols. PLoS ONE 2010, 5, e10202. [Google Scholar] [CrossRef] [PubMed]
- Niles, A.L.; Moravec, R.A.; Riss, T.L. In vitro viability and cytotoxicity testing and same-well multi-parametric combinations for high throughput screening. Curr. Chem. Genom. 2009, 3, 33–41. [Google Scholar] [CrossRef]
- Mahto, S.K.; Chandra, P.; Rhee, S.W. In vitro models, endpoints and assessment methods for the measurement of cytotoxicity. Toxicol. Environ. Health Sci. 2010, 2, 87–93. [Google Scholar] [CrossRef]
- Menyhárt, O.; Harami-Papp, H.; Sukumar, S.; Schäfer, R.; Magnani, L.; de Barrios, O.; Győrffy, B. Guidelines for the selection of functional assays to evaluate the hallmarks of cancer. Biochim. Biophys. Acta Rev. Cancer 2016, 1866, 300–319. [Google Scholar] [CrossRef]
- Markossian, S.; Grossman, A.; Brimacombe, K.; Arkin, M.; Auld, D.; Austin, C.P.; Baell, J.; Chung, T.D.Y.; Coussens, N.P.; Dahlin, J.L.; et al. (Eds.) Assay Guidance Manual; Eli Lilly & Company and the National Center for Advancing Translational Sciences: Bethesda, MD, USA, 2004; p. 540958. Available online: https://www.ncbi.nlm.nih.gov/books/NBK540958/ (accessed on 23 November 2021).
- Feoktistova, M.; Geserick, P.; Leverkus, M. Crystal violet assay for determining viability of cultured cells. Cold Spring Harb. Protoc. 2016, 2016, 343–346. [Google Scholar] [CrossRef]
- Lee, Y.; Chen, M.; Lee, J.D.; Zhang, J.; Lin, S.; Fu, M.; Chen, H.; Ishikawa, T.; Chiang, S.; Katon, J.; et al. HHS Public Access through inhibition of a MYC-WWP1 inhibitory pathway. Science 2020, 364, 1–40. [Google Scholar] [CrossRef]
- Vasan, N.; Razavi, P.; Johnson, J.L.; Shao, H.; Shah, H.; Antoine, A.; Ladewig, E.; Gorelick, A.; Lin, T.Y.; Toska, E.; et al. Double PIK3CA mutations in cis increase oncogenicity and sensitivity to PI3Ka inhibitors. Science 2019, 366, 714–723. [Google Scholar] [CrossRef] [PubMed]
- Johnston, G. Automated handheld instrument improves counting precision across multiple cell lines. Biotechniques 2010, 48, 325–327. [Google Scholar] [CrossRef]
- Strober, W. Trypan Blue Exclusion Test of Cell Viability. Curr. Protoc. Immunol. 2001, 21, A-3B. [Google Scholar] [CrossRef]
- Aslantürk, S.; Çelik, T.A. Antioxidant activity and anticancer effect of Vitex agnus-castus L. (Verbenaceae) seed extracts on MCF–7 breast cancer cells. Caryologia 2013, 66, 257–267. [Google Scholar] [CrossRef]
- Stone, V.; Johnston, H.; Schins, R.P.F. Development ofin vitrosystems for nanotoxicology: Methodological considerations. Crit. Rev. Toxicol. 2009, 39, 613–626. [Google Scholar] [CrossRef]
- Yip, D.K.; Auersperg, N. The dye-exclusion test for cell viability: Persistence of differential staining following fixation. In Vitro J. Tissue Cult. Assoc. 1972, 7, 323–329. [Google Scholar] [CrossRef]
- Ruben, R.L. Cell culture for testing anticancer compounds. In Advances in Cell Culture; Marqmarosh, K., Sato, G.H., Eds.; Academic Press: San Diego, CA, USA, 1988; Volume 6, pp. 161–188. [Google Scholar]
- Kim, S.I.; Kim, H.J.; Lee, H.J.; Lee, K.; Hong, D.; Lim, H.; Cho, K.; Jung, N.; Yi, Y.W. Application of a non-hazardous vital dye for cell counting with automated cell counters. Anal. Biochem. 2016, 492, 8–12. [Google Scholar] [CrossRef]
- Marmion, D.M. Handbook of U.S. Colortants for Foods, Drugs, and Cosmetics; Wiley Interscience: New York, NY, USA, 1979. [Google Scholar]
- Aslantürk, Ö.; Çelik, T.; Karabey, B.; Karabey, F. Active Phytochemical Detecting, Antioxidant, Cytotoxic, Apoptotic Activities of Ethyl Acetate and Methanol Extracts of Galium aparine L. Br. J. Pharm. Res. 2017, 15, 1–16. [Google Scholar] [CrossRef]
- Bopp, S.K.; Lettieri, T. Comparison of four different colorimetric and fluorometric cytotoxicity assays in a zebrafish liver cell line. BMC Pharmacol. 2008, 8, 8. [Google Scholar] [CrossRef] [PubMed]
- Mosmann, T. Rapid colorimetric assay for cellular growth and survival: Application to proliferation and cytotoxicity assays. J. Immunol. Methods 1983, 65, 55–63. [Google Scholar] [CrossRef]
- Berg, K.; Zhai, L.; Chen, M.; Kharazmi, A.; Owen, T.C. The use of a water-soluble formazan complex to quantitate the cell number and mitochondrial function of Leishmania major promastigotes. Parasitol. Res. 1994, 80, 235–239. [Google Scholar] [CrossRef]
- Tominaga, H.; Ishiyama, M.; Ohseto, F.; Sasamoto, K.; Hamamoto, T.; Suzuki, K.; Watanabe, M. A water-soluble tetrazolium salt useful for colorimetric cell viability assay. Anal. Commun. 1999, 36, 47–50. [Google Scholar] [CrossRef]
- Rotter, B.A.; Thompson, B.K.; Clarkin, S.; Owen, T.C. Rapid colorimetric bioassay for screening of fusarium mycotoxins. Nat. Toxins 1993, 1, 303–307. [Google Scholar] [CrossRef] [PubMed]
- Buttke, T.M.; McCubrey, J.A.; Owen, T.C. Use of an aqueous soluble tetrazolium/formazan assay to measure viability and proliferation of lymphokine-dependent cell lines. J. Immunol. Methods 1993, 157, 233–240. [Google Scholar] [CrossRef]
- Promega Corparation Technical Bulletin. CellTitre 96 AQueous Non-Radioactive Cell Proliferation Assay. 2006. Available online: https://worldwide.promega.com/resources/protocols/technical-bulletins/0/celltiter-96-aqueous-nonradioactive-cell-proliferation-assay-protocol/ (accessed on 23 November 2021).
- Cory, A.H.; Owen, T.C.; Barltrop, J.A.; Cory, J.G. Use of an aqueous soluble tetrazolium/formazan assay for cell growth assays in culture. Cancer Comm. 1991, 3, 207–212. [Google Scholar] [CrossRef]
- Riss, T.L.; Moravec, R.A. Comparison of MTT, XTT, and a novel tetrazolium compound MTS for in vitro proliferation and chemosensitivity assays. Mol. Biol. Cell 1992, 3, A184. [Google Scholar]
- Scudiero, D.A.; Shoemaker, R.H.; Paull, K.D.; Monks, A.; Tierney, S.; Nofziger, T.H.; Currens, M.J.; Seniff, D.; Boyd, M.R. Evaluation of a Soluble Tetrazolium/Formazan Assay for Cell Growth and Drug Sensitivity in Culture Using Human and Other Tumor Cell Lines. Cancer Res. 1988, 48, 4827–4833. [Google Scholar] [PubMed]
- Ishiyama, M.; Shiga, M.; Sasamoto, K.; Mizoguchi, M.; He, P. A new sulfonated tetrazolium salt that produces a high water-soluble formazan dye. Chem. Pharm. Bull. 1993, 41, 1118–1122. [Google Scholar] [CrossRef]
- Decker, T.; Lohmann-Matthes, M.L. A quick and simple method for the quantitation of lactate dehydrogenase release in measurements of cellular cytotoxicity and tumor necrosis factor (TNF) activity. J. Immunol. Methods 1988, 115, 61–69. [Google Scholar] [CrossRef]
- Schins, R.P.F.; Duffin, R.; Höhr, D.; Knaapen, A.M.; Shi, T.; Weishaupt, C.; Stone, V.; Donaldson, K.; Borm, P.J.A. Surface modification of quartz inhibits toxicity, particle uptake, and oxidative DNA damage in human lung epithelial cells. Chem. Res. Toxicol. 2002, 15, 1166–1173. [Google Scholar] [CrossRef]
- Fotakis, G.; Timbrell, J.A. In vitro cytotoxicity assays: Comparison of LDH, neutral red, MTT and protein assay in hepatoma cell lines following exposure to cadmium chloride. Toxicol. Lett. 2006, 160, 171–177. [Google Scholar] [CrossRef] [PubMed]
- Lappalanien, K.; Jaaskelainen, L.; Syrjanen, K.; Urtti, A.; Syrjanen, S. Comparison of cell proliferation and toxicity assays using two catronic liposomes. Pharm. Res. 1994, 11, 1127–1131. [Google Scholar] [CrossRef]
- Skehan, P.; Storeng, R.; Scudiero, D.; Monks, A.; Mcmahon, J.; Vistica, D.; Warren, J.T.; Bokesch, H.; Kenney, S.; Boyd, M.R. New colorimetric cytotoxicity assay for anticancer-drug screening. J. Natl. Cancer Inst. 1990, 82, 1107–1112. [Google Scholar] [CrossRef]
- Borenfreund, E.; Puerner, J.A. A simple quantitative procedure using monolayer cultures for cytotoxicity assays (HTD/NR-90). J. Tissue Cult. Methods 1985, 9, 7–9. [Google Scholar] [CrossRef]
- Repetto, G.; del Peso, A.; Zurita, J.L. Neutral red uptake assay for the estimation of cell viability/ cytotoxicity. Nat. Protoc. 2008, 3, 1125–1131. [Google Scholar] [CrossRef]
- Ringwood, A.H.; Conners, D.E.; Hoguet, J. Effects of natural and anthropogenic stressors on lysosomal destabilization in oysters Crassostrea virginica. Mar. Ecol. Prog. Ser. 1998, 166, 163–171. [Google Scholar] [CrossRef]
- Geserick, P.; Hupe, M.; Moulin, M.; Wong, W.W.L.; Feoktistova, M.; Kellert, B.; Gollnick, H.; Silke, J.; Leverkus, M. Cellular IAPs inhibit a cryptic CD95-induced cell death by limiting RIP1 kinase recruitment. J. Cell Biol. 2009, 187, 1037–1054. [Google Scholar] [CrossRef]
- Degterev, A.; Hitomi, J.; Germscheid, M.; Ch’en, I.L.; Korkina, O.; Teng, X.; Abbott, D.; Cuny, G.D.; Yuan, C.; Wagner, G.; et al. Identification of RIP1 kinase as a specific cellular target of necrostatins. Nat. Chem. Biol. 2008, 4, 313–321. [Google Scholar] [CrossRef]
- Sun, L.; Wang, H.; Wang, Z.; He, S.; Chen, S.; Liao, D.; Wang, L.; Yan, J.; Liu, W.; Lei, X.; et al. Mixed lineage kinase domain-like protein mediates necrosis signaling downstream of RIP3 kinase. Cell 2012, 148, 213–227. [Google Scholar] [CrossRef]
- Feoktistova, M.; Geserick, P.; Kellert, B.; Dimitrova, D.P.; Langlais, C.; Hupe, M.; Cain, K.; MacFarlane, M.; Häcker, G.; Leverkus, M. CIAPs Block Ripoptosome Formation, a RIP1/Caspase-8 Containing Intracellular Cell Death Complex Differentially Regulated by cFLIP Isoforms. Mol. Cell 2011, 43, 449–463. [Google Scholar] [CrossRef]
- O’Brien, J.; Wilson, I.; Orton, T.; Pognan, F. Investigation of the Alamar Blue (resazurin) fluorescent dye for the assessment of mammalian cell cytotoxicity. Eur. J. Biochem. 2000, 267, 5421–5426. [Google Scholar] [CrossRef]
- Ansar Ahmed, S.; Gogal, R.M.; Walsh, J.E. A new rapid and simple non-radioactive assay to monitor and determine the proliferation of lymphocytes: An alternative to [3H]thymidine incorporation assay. J. Immunol. Methods 1994, 170, 211–224. [Google Scholar] [CrossRef]
- Page, B.; Page, M.; Noel, C. A new fluorometric assay for cytotoxicity measurements in vitro. Int. J. Oncol. 1993, 3, 473–476. [Google Scholar] [CrossRef]
- Pace, R.T.; Burg, K.J.L. Toxic effects of resazurin on cell cultures. Cytotechnology 2015, 67, 13–17. [Google Scholar] [CrossRef]
- Schreer, A.; Tinson, C.; Sherry, J.P.; Schirmer, K. Application of Alamar blue/5-carboxyfluorescein diacetate acetoxymethyl ester as a noninvasive cell viability assay in primary hepatocytes from rainbow trout. Anal. Biochem. 2005, 344, 76–85. [Google Scholar] [CrossRef]
- Ganassi, R.C.; Schirmer, K.; Bols, N.C. Cell and tissue culture. In The Laboratory Fish; Ostrander, G.K., Ed.; Academic Press: San Diego, CA, USA, 2000; pp. 631–651. [Google Scholar]
- Sidman, R.L.; Miale, I.L.; Feder, N. Cell proliferation and migration in the primitive ependymal zone; An autoradiographic study of histogenesis in the nervous system. Exp. Neurol. 1959, 1, 322–333. [Google Scholar] [CrossRef]
- Miller, M.W.; Nowakowski, R.S. Use of bromodeoxyuridine-immunohistochemistry to examine the proliferation, migration and time of origin of cells in the central nervous system. Brain Res. 1988, 457, 44–52. [Google Scholar] [CrossRef]
- Salic, A.; Mitchison, T.J. A chemical method for fast and sensitive detection of DNA synthesis in vivo. Proc. Natl. Acad. Sci. USA 2008, 105, 2415–2420. [Google Scholar] [CrossRef]
- Nowakowski, R.S.; Lewin, S.B.; Miller, M.W. Bromodeoxyuridine immunohistochemical determination of the lengths of the cell cycle and the DNA-synthetic phase for an anatomically defined population. J. Neurocytol. 1989, 18, 311–318. [Google Scholar] [CrossRef]
- Taupin, P. BrdU immunohistochemistry for studying adult neurogenesis: Paradigms, pitfalls, limitations, and validation. Brain Res. Rev. 2007, 53, 198–214. [Google Scholar] [CrossRef]
- Duque, A.; Rakic, P. Different effects of BrdU and (3)H-thymidine incorporation into DNA on cell proliferation, position and fate. J. Neurosci. Off. J. Soc. Neurosci. 2001, 31, 15205–15217. [Google Scholar] [CrossRef]
- Maehara, Y.; Anai, H.; Tamada, R.; Sugimachi, K. The ATP assay is more sensitive than the succinate dehydrogenase inhibition test for predicting cell viability. Eur. J. Cancer Clin. Oncol. 1987, 23, 273–276. [Google Scholar] [CrossRef]
- García, O.; Massieu, L. Glutamate Uptake Inhibitor L-Trans-Pyrrolidine 2,4-Dicarboxylate Becomes Neurotoxic in the Presence of Subthreshold Concentrations of Mitochondrial Toxin 3-Nitropropionate: Involvement of Mitochondrial Reducing Activity and ATP Production. J. Neurosci. Res. 2003, 74, 956–966. [Google Scholar] [CrossRef]
- Andreotti, P.E.; Hartmann, D.M.; Linder, D.; Harel, G.; Gleiberman, I.; Caruso, P.A.; Ricks, S.H.; Cree, I.A.; Kurbacher, C.M.; Untch, M.; et al. Chemosensitivity Testing of Human Tumors Using a Microplate Adenosine Triphosphate Luminescence Assay: Clinical Correlation for Cisplatin Resistance of Ovarian Carcinoma. Cancer Res. 1995, 55, 5276–5282. [Google Scholar]
- Duellman, S.J.; Zhou, W.; Meisenheimer, P.; Vidugiris, G.; Cali, J.J.; Gautam, P.; Wennerberg, K.; Vidugiriene, J. Bioluminescent, Nonlytic, Real-Time Cell Viability Assay and Use in Inhibitor Screening. Assay Drug Dev. Technol. 2015, 13, 456–465. [Google Scholar] [CrossRef]
- Suzuki, T.; Fujikura, K.; Higashiyama, T.; Takata, K. DNA staining for fluorescence and laser confocal microscopy. J. Histochem. Cytochem. 1997, 45, 49–53. [Google Scholar] [CrossRef]
- Liu, X.; Chen, H.; Patel, D.J. Solution structure of actinomycin-DNA complexes: Drug intercalation at isolated G-C sites. J. Biomol. NMR 1991, 1, 323–347. [Google Scholar] [CrossRef]
- Latt, S.A. Fluorescent probes of chromosome structure and replication. Can. J. Genet. Cytol. 1977, 19, 603–623. [Google Scholar] [CrossRef]
- Liu, W.; Liang, Y.; Chan, Q.; Jiang, L.; Dong, J. CX3CL1 promotes lung cancer cell migration and invasion via the Src/focal adhesion kinase signaling pathway. Oncol. Rep. 2019, 41, 1911–1917. [Google Scholar] [CrossRef]
- Réu, P.; Svedberg, G.; Hässler, L.; Möller, B.; Svahn, H.A.; Gantelius, J. A 61% lighter cell culture dish to reduce plastic waste. PLoS ONE 2019, 14, e0216251. [Google Scholar] [CrossRef]
- Avlasevich, S.L.; Bryce, S.M.; Cairns, S.E.; Dertinger, S.D. In vitro micronucleus scoring by flow cytometry: Differential staining of micronuclei versus apoptotic and necrotic chromatin enhances assay reliability. Environ. Mol. Mutagen. 2006, 47, 56–66. [Google Scholar] [CrossRef]
- Yan, X.; Habbersett, R.C.; Cordek, J.M.; Nolan, J.P.; Yoshida, T.M.; Jett, J.H.; Marrone, B.L. Development of a mechanism-based, DNA staining protocol using SYTOX orange nucleic acid stain and DNA fragment sizing flow cytometry. Anal. Biochem. 2000, 286, 138–148. [Google Scholar] [CrossRef]
- Bryce, S.M.; Bemis, J.C.; Avlasevich, S.L.; Dertinger, S.D. In vitro micronucleus assay scored by flow cytometry provides a comprehensive evaluation of cytogenetic damage and cytotoxicity. Mutat. Res. 2007, 630, 78–91. [Google Scholar] [CrossRef]
- Mukhopadhyay, P.; Rajesh, M.; Haskó, G.; Hawkins, B.J.; Madesh, M.; Pacher, P. Simultaneous detection of apoptosis and mitochondrial superoxide production in live cells by flow cytometry and confocal microscopy. Nat. Protoc. 2007, 2, 2295–2301. [Google Scholar] [CrossRef]
- Kerscher, B.; Barlow, J.L.; Rana, B.M.; Jolin, H.E.; Gogoi, M.; Bartholomew, M.A.; Jhamb, D.; Pandey, A.; Tough, D.F.; Van Oosterhout, A.J.M.; et al. BET bromodomain inhibitor IBET151 impedes human ILC2 activation and prevents experimental allergic lung inflammation. Front. Immunol. 2019, 10, 678. [Google Scholar] [CrossRef]
- Paivandy, A.; Eriksson, J.; Melo, F.R.; Sellin, M.E.; Pejler, G. Lysosomotropic challenge of mast cells causes intra-granular reactive oxygen species production. Cell Death Discov. 2019, 5, 95. [Google Scholar] [CrossRef]
- Vig, S.; Buitinga, M.; Rondas, D.; Crèvecoeur, I.; van Zandvoort, M.; Waelkens, E.; Eizirik, D.L.; Gysemans, C.; Baatsen, P.; Mathieu, C.; et al. Cytokine-induced translocation of GRP78 to the plasma membrane triggers a pro-apoptotic feedback loop in pancreatic beta cells. Cell Death Dis. 2019, 10, 309. [Google Scholar] [CrossRef]
- Akagi, J.; Kordon, M.; Zhao, H.; Matuszek, A.; Dobrucki, J.; Errington, R.; Smith, P.J.; Takeda, K.; Darzynkiewicz, Z.; Wlodkowic, D. Real-time cell viability assays using a new anthracycline derivative DRAQ7®. Cytom. Part A 2013, 83 A, 227–234. [Google Scholar] [CrossRef]
- Severini, A.; Richard Morgan, A. An assay for proteinases and their inhibitors based on DNA/ethidium bromide fluorescence. Anal. Biochem. 1991, 193, 83–89. [Google Scholar] [CrossRef]
- Zipper, H.; Brunner, H.; Bernhagen, J.; Vitzthum, F. Investigations on DNA intercalation and surface binding by SYBR Green I, its structure determination and methodological implications. Nucleic Acids Res. 2004, 32, e103. [Google Scholar] [CrossRef]
- Singer, V.L.; Lawlor, T.E.; Yue, S. Comparison of SYBR® Green I nucleic acid gel stain mutagenicity and ethidium bromide mutagenicity in the Salmonella/mammalian microsome reverse mutation assay (Ames test). Mutat. Res. Genet. Toxicol. Environ. Mutagen. 1999, 439, 37–47. [Google Scholar] [CrossRef]
- Mirrett, S. Acridin orange stain. Infect. Control. 1982, 3, 250–252. [Google Scholar] [CrossRef]
- Kumar, R.; Kaur, M.; Kumari, M. Acridine: A versatile heterocyclic nucleus. Acta Pol. Pharm. Drug Res. 2012, 69, 3–9. [Google Scholar]
- Darzynkiewicz, Z.; Juan, G.; Srour, E.F. Differential staining of DNA and RNA. Curr. Protoc. Cytom. 2004, 30, 7.3.1–7.3.16. [Google Scholar] [CrossRef]
- Lekishvili, T.; Campbell, J.J. Rapid comparative immunophenotyping of human mesenchymal stromal cells by a modified fluorescent cell barcoding flow cytometric assay. Cytom. Part A 2018, 93, 905–915. [Google Scholar] [CrossRef] [PubMed]
- BD Biosciences. BD Biosciences Brochure. Available online: https://www.bdbiosciences.com/en-eu/applications/research-applications/multicolor-flow-cytometry (accessed on 23 November 2021).
- Prado-Garcia, H.; Romero-Garcia, S.; Rumbo-Nava, U.; Lopez-Gonzalez, J.S. Predominance of Th17 over regulatory T-cells in pleural effusions of patients with lung cancer implicates a proinflammatory profile. Anticancer Res. 2015, 35, 1529–1536. [Google Scholar]
- McMaster, S.R.; Gabbard, J.D.; Koutsonanos, D.G.; Compans, R.W.; Tripp, R.A.; Tompkins, S.M.; Kohlmeier, J.E. Memory T cells generated by prior exposure to influenza cross react with the novel H7N9 influenza virus and confer protective heterosubtypic immunity. PLoS ONE 2015, 10, e0115725. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Rodríguez, N.; Apostolidis, S.A.; Penaloza-MacMaster, P.; Martín Villa, J.M.; Barouch, D.H.; Tsokos, G.C.; Crispín, J.C. Programmed Cell Death 1 and Helios Distinguish TCR-αβ + Double-Negative (CD4 − CD8 −) T Cells That Derive from Self-Reactive CD8 T Cells. J. Immunol. 2015, 194, 4207–4214. [Google Scholar] [CrossRef]
- Flies, D.B.; Higuchi, T.; Chen, L. Mechanistic Assessment of PD-1H Coinhibitory Receptor–Induced T Cell Tolerance to Allogeneic Antigens. J. Immunol. 2015, 194, 5294–5304. [Google Scholar] [CrossRef] [PubMed]
- Akabane, S.; Matsuzaki, K.; Yamashita, S.I.; Arai, K.; Okatsu, K.; Kanki, T.; Matsuda, N.; Oka, T. Constitutive activation of PINK1 protein leads to proteasome-mediated and non-apoptotic cell death independently of mitochondrial autophagy. J. Biol. Chem. 2016, 291, 16162–16174. [Google Scholar] [CrossRef]
- Iraolagoitia, X.L.R.; Spallanzani, R.G.; Torres, N.I.; Araya, R.E.; Ziblat, A.; Domaica, C.I.; Sierra, J.M.; Nuñez, S.Y.; Secchiari, F.; Gajewski, T.F.; et al. NK Cells Restrain Spontaneous Antitumor CD8 + T Cell Priming through PD-1/PD-L1 Interactions with Dendritic Cells. J. Immunol. 2016, 197, 953–961. [Google Scholar] [CrossRef]
- Mercer, F.; Diala, F.G.I.; Chen, Y.P.; Molgora, B.M.; Ng, S.H.; Johnson, P.J. Leukocyte Lysis and Cytokine Induction by the Human Sexually Transmitted Parasite Trichomonas vaginalis. PLoS Negl. Trop. Dis. 2016, 10, e0004913. [Google Scholar] [CrossRef]
- Souza-Fonseca-Guimaraes, F.; Young, A.; Mittal, D.; Martinet, L.; Bruedigam, C.; Takeda, K.; Andoniou, C.E.; Degli-Esposti, M.A.; Hill, G.R.; Smyth, M.J. NK cells require IL-28R for optimal in vivo activity. Proc. Natl. Acad. Sci. USA 2015, 112, E2376–E2384. [Google Scholar] [CrossRef]
- Matsui, K.; Adelsberger, J.W.; Kemp, T.J.; Baseler, M.W.; Ledgerwood, J.E.; Pinto, L.A. Circulating CXCR5+CD4+ T follicular-like helper cell and memory B cell responses to human papillomavirus vaccines. PLoS ONE 2015, 10, e0137195. [Google Scholar] [CrossRef]
- Jones, A.; Opejin, A.; Henderson, J.G.; Gross, C.; Jain, R.; Epstein, J.A.; Flavell, R.A.; Hawiger, D. Peripherally Induced Tolerance Depends on Peripheral Regulatory T Cells That Require Hopx to Inhibit Intrinsic IL-2 Expression. J. Immunol. 2015, 195, 1489–1497. [Google Scholar] [CrossRef]
- Nath, A.; Li, I.; Roberts, L.R.; Chan, C. Elevated free fatty acid uptake via CD36 promotes epithelial-mesenchymal transition in hepatocellular carcinoma. Sci. Rep. 2015, 5, 14752. [Google Scholar] [CrossRef]
- Kanemaru, K.; Noguchi, E.; Tokunaga, T.; Nagai, K.; Hiroyama, T.; Nakamura, Y.; Tahara-Hanaoka, S.; Shibuya, A. Tie2 Signaling Enhances Mast Cell Progenitor Adhesion to Vascular Cell Adhesion Molecule-1 (VCAM-1) through α4β1 Integrin. PLoS ONE 2015, 10, e0144436. [Google Scholar] [CrossRef]
- Talabot-Ayer, D.; Martin, P.; Vesin, C.; Seemayer, C.A.; Vigne, S.; Gabay, C.; Palmer, G. Severe Neutrophil-Dominated Inflammation and Enhanced Myelopoiesis in IL-33–Overexpressing CMV/IL33 Mice. J. Immunol. 2015, 194, 750–760. [Google Scholar] [CrossRef]
- Keppel, M.P.; Saucier, N.; Mah, A.Y.; Vogel, T.P.; Cooper, M.A. Activation-Specific Metabolic Requirements for NK Cell IFN-γ Production. J. Immunol. 2015, 194, 1954–1962. [Google Scholar] [CrossRef]
- Shade, K.T.C.; Platzer, B.; Washburn, N.; Mani, V.; Bartsch, Y.C.; Conroy, M.; Pagan, J.D.; Bosques, C.; Mempel, T.R.; Fiebiger, E.; et al. A single glycan on IgE is indispensable for initiation of anaphylaxis. J. Exp. Med. 2015, 212, 457–467. [Google Scholar] [CrossRef]
- Weiser, J.N.; Roche, A.M.; Hergott, C.B.; LaRose, M.I.; Connolly, T.; Jorgensen, W.L.; Leng, L.; Bucala, R.; Das, R. Macrophage Migration Inhibitory Factor Is Detrimental in Pneumococcal Pneumonia and a Target for Therapeutic Immunomodulation. J. Infect. Dis. 2015, 212, 1677–1682. [Google Scholar] [CrossRef]
- Allen, T.M.; Cleland, L.G. Serum-induced leakage of liposome contents. BBA Biomembr. 1980, 597, 418–426. [Google Scholar] [CrossRef]
- Patel, H.; Tscheka, C.; Heerklotz, H. Characterizing vesicle leakage by fluorescence lifetime measurements. Soft Matter 2009, 5, 2849–2851. [Google Scholar] [CrossRef]
- Szakács, G.; Váradi, A.; Özvegy-Laczka, C.; Sarkadi, B. The role of ABC transporters in drug absorption, distribution, metabolism, excretion and toxicity (ADME-Tox). Drug Discov. Today 2008, 13, 379–393. [Google Scholar] [CrossRef]
- Tóvári, J.; Futosi, K.; Bartal, A.; Tátrai, E.; Gacs, A.; Kenessey, I.; Paku, S. Boyden chamber-based method for characterizing the distribution of adhesions and cytoskeletal structure in HT1080 fibrosarcoma cells. Cell Adhes. Migr. 2014, 8, 509–516. [Google Scholar] [CrossRef][Green Version]
- Khalili, A.A.; Ahmad, M.R. A Review of cell adhesion studies for biomedical and biological applications. Int. J. Mol. Sci. 2015, 16, 18149–18184. [Google Scholar] [CrossRef]
- Zepeda-Moreno, A.; Taubert, I.; Hellwig, I.; Hoang, V.; Pietsch, L.; Lakshmanan, V.K.; Wagner, W.; Ho, A.D. Innovative method for quantification of cell-cell adhesion in 96-well plates. Cell Adhes. Migr. 2011, 5, 215–219. [Google Scholar] [CrossRef]
- Kashef, J.; Franz, C.M. Quantitative methods for analyzing cell-cell adhesion in development. Dev. Biol. 2015, 401, 165–174. [Google Scholar] [CrossRef]
- Zhou, D.W.; García, A.J. Measurement Systems for Cell Adhesive Forces. J. Biomech. Eng. 2015, 137, 020908. [Google Scholar] [CrossRef]
- Ungai-Salánki, R.; Peter, B.; Gerecsei, T.; Orgovan, N.; Horvath, R.; Szabó, B. A practical review on the measurement tools for cellular adhesion force. Adv. Colloid Interface Sci. 2019, 269, 309–333. [Google Scholar] [CrossRef]
- Kramer, N.; Walzl, A.; Unger, C.; Rosner, M.; Krupitza, G.; Hengstschläger, M.; Dolznig, H. In vitro cell migration and invasion assays. Mutat. Res. Rev. Mutat. Res. 2013, 752, 10–24. [Google Scholar] [CrossRef]
- Boyden, S. The chemotactic effect of mixtures of antibody and antigen on polymorphonuclear leucocytes. J. Exp. Med. 1962, 115, 453–466. [Google Scholar] [CrossRef]
- Restouin, A.; Aresta, S.; Prébet, T.; Borg, J.P.; Badache, A.; Collette, Y. A simplified, 96-well-adapted, ATP luminescence-based motility assay. Biotechniques 2009, 47, 871–875. [Google Scholar] [CrossRef]
- Marshall, J. Transwell((R)) invasion assays. Methods Mol. Biol. 2011, 769, 97–110. [Google Scholar]
- Liang, C.C.; Park, A.Y.; Guan, J.L. In vitro scratch assay: A convenient and inexpensive method for analysis of cell migration in vitro. Nat. Protoc. 2007, 2, 329–333. [Google Scholar] [CrossRef]
- Gyorffy, B.; Stelniec-Klotz, I.; Sigler, C.; Kasack, K.; Redmer, T.; Qian, Y.; Schäfer, R. Effects of RAL signal transduction in KRAS-and BRAF-mutated cells and prognostic potential of the RAL signature in colorectal cancer. Oncotarget 2015, 6, 13334–13346. [Google Scholar] [CrossRef]
- Gorshkova, I.; He, D.; Berdyshev, E.; Usatuyk, P.; Burns, M.; Kalari, S.; Zhao, Y.; Pendyala, S.; Garcia, J.G.N.; Pyne, N.J.; et al. Protein kinase C-ε regulates sphingosine 1-phosphate-mediated migration of human lung endothelial cells through activation of phospholipase D2, protein kinase C-ζ, and Rac1. J. Biol. Chem. 2008, 283, 11794–11806. [Google Scholar] [CrossRef]
- Lo, C.M.; Keese, C.R.; Giaever, I. Impedance analysis of MDCK cells measured by electric cell-substrate impedance sensing. Biophys. J. 1995, 69, 2800–2807. [Google Scholar] [CrossRef]
- Tamada, M.; Perez, T.D.; Nelson, W.J.; Sheetz, M.P. Two distinct modes of myosin assembly and dynamics during epithelial wound closure. J. Cell Biol. 2007, 176, 27–33. [Google Scholar] [CrossRef]
- Zordan, M.D.; Mill, C.P.; Riese, D.J.; Leary, J.F. A high throughput, interactive imaging, bright-field wound healing assay. Cytom. Part A 2011, 79 A, 227–232. [Google Scholar] [CrossRef]
- Poujade, M.; Grasland-Mongrain, E.; Hertzog, A.; Jouanneau, J.; Chavrier, P.; Ladoux, B.; Buguin, A.; Silberzan, P. Collective migration of an epithelial monolayer in response to a model wound. Proc. Natl. Acad. Sci. USA 2007, 104, 15988–15993. [Google Scholar] [CrossRef]
- Simpson, K.J.; Selfors, L.M.; Bui, J.; Reynolds, A.; Leake, D.; Khvorova, A.; Brugge, J.S. Identification of genes that regulate epithelial cell migration using an siRNA screening approach. Nat. Cell Biol. 2008, 10, 1027–1038. [Google Scholar] [CrossRef]
- Gough, W.; Hulkower, K.I.; Lynch, R.; McGlynn, P.; Uhlik, M.; Yan, L.; Lee, J.A. A quantitative, facile, and high-throughput image-based cell migration method is a robust alternative to the scratch assay. J. Biomol. Screen. 2011, 16, 155–163. [Google Scholar] [CrossRef]
- Leto, T.L.; Pratt, B.M.; Madri, J.A. Mechanisms of cytoskeletal regulation: Modulation of aortic endothelial cell protein band 4.1 by the extracellular matrix. J. Cell. Physiol. 1986, 127, 423–431. [Google Scholar] [CrossRef]
- Varani, J.; Orr, W.; Ward, P.A. A comparison of the migration patterns of normal and malignant cells in two assay systems. Am. J. Pathol. 1978, 90, 159–172. [Google Scholar]
- Kroening, S.; Goppelt-Struebe, M. Analysis of matrix-dependent cell migration with a barrier migration assay. Sci. Signal. 2010, 3, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Rosen, E.M.; Meromsky, L.; Setter, E.; Vinter, D.W.; Goldberg, I.D. Quantitation of cytokine-stimulated migration of endothelium and epithelium by a new assay using microcarrier beads. Exp. Cell Res. 1990, 186, 22–31. [Google Scholar] [CrossRef]
- Konduri, S.D.; Tasiou, A.; Chandrasekar, N.; Rao, J.S. Overexpression of tissue factor pathway inhibitor-2 (TFPI-2), decreases the invasiveness of prostate cancer cells in vitro. Int. J. Oncol. 2001, 18, 127–131. [Google Scholar] [CrossRef] [PubMed]
- Chaubey, S.; Ridley, A.J.; Wells, C.M. Using the Dunn Chemotaxis Chamber to Analyze Primary Cell Migration in Real Time, in Cell Migration: Developmental Methods and Protocols. Methods Mol. Biol. 2011, 769, 41–51. [Google Scholar] [CrossRef]
- Echeverria, V.; Meyvantsson, I.; Skoien, A.; Worzella, T.; Lamers, C.; Hayes, S. An automated high-content assay for tumor cell migration through 3-Dimensional matrices. J. Biomol. Screen. 2010, 15, 1144–1151. [Google Scholar] [CrossRef] [PubMed]
- Miura, K. Tracking movement in cell biology. Adv. Biochem. Eng. Biotechnol. 2005, 95, 267–295. [Google Scholar] [CrossRef] [PubMed]
- Gu, W.; Pellegrino, T.; Parak, W.J.; Boudreau, R.; Le Gros, M.A.; Alivisatos, A.P.; Larabell, C.A. Measuring cell motility using quantum dot probes. Methods Mol. Biol. 2007, 374, 125–131. [Google Scholar] [CrossRef]
- Lin, M.; DiVito, M.M.; Merajver, S.D.; Boyanapalli, M.; van Golen, K.L. Regulation of pancreatic cancer cell migration and invasion by RhoC GTPase and caveolin-1. Mol. Cancer 2005, 4, 21. [Google Scholar] [CrossRef]
- Niinaka, Y.; Haga, A.; Raz, A. Quantification of cell motility. Metastasis Res. Protoc. 2001, 58, 55–60. [Google Scholar]
- Swaminathan, V.; Mythreye, K.; Tim O’Brien, E.; Berchuck, A.; Blobe, G.C.; Superfine, R. Mechanical Stiffness grades metastatic potential in patient tumor cells and in cancer cell lines. Cancer Res. 2011, 71, 5075–5080. [Google Scholar] [CrossRef]
- Cribb, J.A.; Osborne, L.D.; Beicker, K.; Psioda, M.; Chen, J.; O’Brien, E.T.; Taylor, R.M.; Vicci, L.; Hsiao, J.P.L.; Shao, C.; et al. An Automated High-throughput Array Microscope for Cancer Cell Mechanics. Sci. Rep. 2016, 6, 27371. [Google Scholar] [CrossRef]
- Hillman, H. Procedures used in modern cell biology ignore natural laws. J. Biol. Phys. Chem. 2009, 9, 130–135. [Google Scholar] [CrossRef]
- Ramsden, J.J. Towards zero-perturbation methods for investigating biomolecular interactions. Colloids Surf. A Physicochem. Eng. Asp. 1998, 141, 287–294. [Google Scholar] [CrossRef]
- Ramsden, J.J. Experimental methods for investigating protein adsorption kinetics at surfaces. Q. Rev. Biophys. 1993, 27, 41–105. [Google Scholar] [CrossRef]
- Ramsden, J.J. Optical biosensors. J. Mol. Recog. 1997, 10, 109–120. [Google Scholar] [CrossRef]
- Song, H.P.; Wang, H.; Zhao, X.; He, L.; Zhong, H.; Wu, S.Q.; Li, P.; Yang, H. Label-free pharmacological profiling based on dynamic mass redistribution for characterization and authentication of hazardous natural products. J. Hazard. Mater. 2017, 333, 265–274. [Google Scholar] [CrossRef]
- Zhang, X.; Deng, H.; Xiao, Y.; Xue, X.; Ferrie, A.M.; Tran, E.; Liang, X.; Fang, Y. Label-free cell phenotypic profiling identifies pharmacologically active compounds in two traditional Chinese medicinal plants. RSC Adv. 2014, 4, 26368–26377. [Google Scholar] [CrossRef]
- Peter, B.; Nador, J.; Juhasz, K.; Dobos, A.; Korosi, L.; Székács, I.; Patko, D.; Horvath, R. Incubator proof miniaturized Holomonitor to in situ monitor cancer cells exposed to green tea polyphenol and preosteoblast cells adhering on nanostructured titanate surfaces: Validity of the measured parameters and their corrections. J. Biomed. Opt. 2015, 20, 067002. [Google Scholar] [CrossRef]
- McColl, J.; Horvath, R.; Aref, A.; Larcombe, L.; Chianella, I.; Morgan, S.; Yakubov, G.E.; Ramsden, J.J. Polyphenol control of cell spreading on glycoprotein substrata. J. Biomater. Sci. Polym. Ed. 2009, 20, 841–851. [Google Scholar] [CrossRef]
- Orgovan, N.; Peter, B.; Bősze, S.; Ramsden, J.J.; Szabó, B.; Horvath, R. Dependence of cancer cell adhesion kinetics on integrin ligand surface density measured by a high-throughput label-free resonant waveguide grating biosensor. Sci. Rep. 2014, 4, 4034. [Google Scholar] [CrossRef]
- Aref, A.; Horvath, R.; Ramsden, J.J. Spreading kinetics for quantifying cell state during stem cell differentiation. J. Biol. Phys. Chem. 2010, 10, 145–151. [Google Scholar] [CrossRef]
- Ramsden, J.J. Biospecific interaction analysis using integrated optics techniques. In Quantitative Analysis of Biospecific Interactions; Lundahl, P., Lundqvist, A., Greijer, E., Eds.; Harwood: Amsterdam, The Netherlands, 1998; pp. 149–162. [Google Scholar]
- Michielin, O.; Vergères, G.; Ramsden, J.J. Myristoylation-induced compaction of a membrane-binding protein. J. Am. Chem. Soc. 1999, 121, 6523–6526. [Google Scholar] [CrossRef]
- Shanshiashvili, L.V.; Suknidze, N.C.; Machaidze, G.G.; Mikeladze, D.G.; Ramsden, J.J. Adhesion and clustering of charge isomers of myelin basic protein at model myelin membranes. Arch. Biochem. Biophys. 2003, 419, 170–177. [Google Scholar] [CrossRef] [PubMed]
- Vergères, G. and Ramsden, J.J. Regulation of the binding of MARCKS-related protein to lipid bilayer membranes by calmodulin. Arch. Biochem. Biophys. 2000, 378, 45–50. [Google Scholar] [CrossRef]
- Ramsden, J.J. OWLS–A versatile technique for drug discovery. Front. Drug Des. Discov. 2006, 2, 211–223. [Google Scholar]
- Nikolelis, D.P.; Nikoleli, G.P. Biosensors for Security and Bioterrorism Applications; Springer International Publishing: Cham, Switzerland, 2016. [Google Scholar] [CrossRef]
- Ramsden, J.J. From Kinetics to Structure: High resolution molecular microscopy. In Proteins at Solid-Liquid Interfaces; Dejardin, P., Ed.; Springer: Berlin/Heidelberg, Germany, 2006; pp. 23–49. [Google Scholar]
- Horvath, R.; McColl, J.; Yakubov, G.E.; Ramsden, J.J. Structural hysteresis and hierarchy in adsorbed glycoproteins. J. Chem. Phys. 2008, 129, 08B401. [Google Scholar] [CrossRef]
- Saftics, A.; Kurunczi, S.; Peter, B.; Szekacs, I.; Ramsden, J.J.; Horvath, R. Data evaluation for surface-sensitive label-free methods to obtain real-time kinetic and structural information of thin films: A practical review with related software packages. Adv. Colloid Interface Sci. 2021, 294, 102431. [Google Scholar] [CrossRef] [PubMed]
- Jankovics, H.; Kovacs, B.; Saftics, A.; Gerecsei, T.; Tóth, É.; Szekacs, I.; Vonderviszt, F.; Horvath, R. Grating-coupled interferometry reveals binding kinetics and affinities of Ni ions to genetically engineered protein layers. Sci. Rep. 2020, 10, 22253. [Google Scholar] [CrossRef]
- Kliment, K.; Szekacs, I.; Peter, B.; Erdei, A.; Kurucz, I.; Horvath, R. Biosensors and Bioelectronics Label-free real-time monitoring of the BCR-triggered activation of primary human B cells modulated by the simultaneous engagement of inhibitory receptors. Biosens. Bioelectron. 2021, 191, 113469. [Google Scholar] [CrossRef]
- Sztilkovics, M.; Gerecsei, T.; Peter, B.; Saftics, A.; Kurunczi, S.; Szekacs, I.; Szabo, B.; Horvath, R. Single-cell adhesion force kinetics of cell populations from combined label-free optical biosensor and robotic fluidic force microscopy. Sci. Rep. 2020, 10, 61. [Google Scholar] [CrossRef]
- Ramsden, J.J.; Li, S.-Y.; Heinzle, E.; Prenosil, J.E. Optical method for measurement of number and shape of attached cells in real time. Cytometry 1995, 1, 97–102. [Google Scholar] [CrossRef]
- Wang, R.; Wang, J.; Liu, Y.; Zhang, X.; Liang, X. Resonant waveguide grating based assays for colloidal aggregate detection and promiscuity characterization in natural products. RSC Adv. 2019, 9, 38055–38064. [Google Scholar] [CrossRef]
- Persson, J.; Mölder, A.; Pettersson, S.; Alm, K. Cell motility studies using digital holographic microscopy. Education 2010, 1063–1072. [Google Scholar]
- El-Schish, Z.; Mölder, A.; Sebesta, M.; Gisselsson, L.; Alm, K.; Gjörloff, A.; Wingren, A. Digital holographic microscopy–Innovative and non-destructive analysis of living cells. Microsc. Sci. Technol. Appl. Educ. 2010, 4, 1055–1062. [Google Scholar]
- Murray, E.M.; Allen, C.F.; Handy, T.E.; Hu, C.A.; Craig, W.R.; Seaton, S.C.; Wolfe, A.L. Development of a Robust and Quantitative High-Throughput Screening Method for Antibiotic Production in Bacterial Libraries. ACS Omega 2019, 4, 15414–15420. [Google Scholar] [CrossRef]
- Cordell, G.A.; Shin, Y.G. Finding the needle in the haystack. The dereplication of natural product extracts. Pure Appl. Chem. 1999, 71, 1089–1094. [Google Scholar] [CrossRef]
- Dias, D.A.; Urban, S.; Roessner, U. A Historical Overview of Natural Products in Drug Discovery. Metabolites 2012, 303–336. [Google Scholar] [CrossRef] [PubMed]
- Newman, D.J.; Cragg, G.M. Natural Products as Sources of New Drugs over the Nearly Four Decades from 01/1981 to 09/2019. J. Nat. Prod. 2020, 83, 770–803. [Google Scholar] [CrossRef] [PubMed]
- Atanasov, A.G.; Zotchev, S.B.; Dirsch, V.M.; Orhan, I.E.; Banach, M.; Rollinger, J.M.; Barreca, D.; Weckwerth, W.; Bauer, R.; Bayer, E.A.; et al. Natural products in drug discovery: Advances and opportunities. Nat. Rev. Drug Discov. 2021, 20, 200–216. [Google Scholar] [CrossRef]
- Thomford, N.E.; Senthebane, D.A.; Rowe, A.; Munro, D.; Seele, P.; Maroyi, A.; Dzobo, K. Natural products for drug discovery in the 21st century: Innovations for novel drug discovery. Int. J. Mol. Sci. 2018, 19, 1578. [Google Scholar] [CrossRef]
- Walker, J.M. Natural products isolation. In Methods in Biotechnology; Sarker, S.D., Latif, Z., Gray, A.I., Eds.; Humana Press Inc: Totowa, NJ, USA, 2005. [Google Scholar]
- Sarker, S.D.; Nahar, L. An Introduction to Natural Products Isolation. Methods Mol. Biol. 2012, 864, 1–25. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.W.; Lin, L.G.; Ye, W.C. Techniques for extraction and isolation of natural products: A comprehensive review. Chin. Med. 2018, 13, 20. [Google Scholar] [CrossRef]
- Cavalcanti, R.N. Uses and Applications of Extracts from Natural Sources. In RSC Green Chemistry No. 21 Natural Product Extraction: Principles and Applications; Rostagno, M.A., Prado, J.M., Eds.; The Royal Society of Chemistry: London, UK, 2013. [Google Scholar]
- Li, J.; Hu, S.; Jian, W.; Xie, C.; Yang, X. Plant antimicrobial peptides: Structures, functions, and applications. Bot. Stud. 2021, 62, 5. [Google Scholar] [CrossRef] [PubMed]
- Dos Santos-Silva, C.A.; Zupin, L.; Oliveira-Lima, M.; Vilela, L.M.B.; Bezerra-Neto, J.P.; Ferreira-Neto, J.R.; Ferreira, J.D.C.; de Oliveira-Silva, R.L.; de J. Pires, C.; Aburjaile, F.F.; et al. Plant Antimicrobial Peptides: State of the Art, In Silico Prediction and Perspectives in the Omics Era. Bioinform. Biol. Insights 2020, 14, 1177932220952739. [Google Scholar] [CrossRef]
- Hegedüs, N.; Marx, F. Antifungal proteins: More than antimicrobials? Fungal Biol. Rev. 2013, 26, 132–145. [Google Scholar] [CrossRef]
- Shwaiki, L.N.; Lynch, K.M.; Arendt, E.K. Future of antimicrobial peptides derived from plants in food application–A focus on synthetic peptides. Trends Food Sci. Technol. 2021, 112, 312–324. [Google Scholar] [CrossRef]
- Xiang, L.; Chi, T.; Tang, Q.; Yang, X.; Ou, M.; Chen, X.; Yu, X.; Chen, J.; Ho, R.J.Y.; Shao, J.; et al. A pentacyclic triterpene natural product, ursolic acid and its prodrug US597 inhibit targets within cell adhesion pathway and prevent cancer metastasis. Oncotarget 2015, 6, 9295–9312. [Google Scholar] [CrossRef]
- Rahimi-Esboei, B.; Ebrahimzadeh, M.A.; Gholami, S.H.; Falah-Omrani, V. Anti-giardial activity of Sambucus ebulus. Eur. Rev. Med. Pharmacol. Sci. 2013, 17, 2047–2050. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, Y.; Isemura, M. Inhibitory effect of epigallocatechin gallate on adhesion of murine melanoma cells to laminin. Cancer Lett. 2001, 173, 15–20. [Google Scholar] [CrossRef]
- De Campos, P.S.; Matte, B.F.; Diel, L.F.; Jesus, L.H.; Bernardi, L.; Alves, A.M.; Rados, P.V.; Lamers, M.L. Low Doses of Curcuma longa Modulates Cell Migration and Cell-Cell Adhesion. Phyther. Res. 2017, 31, 1433–1440. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.D.; Liu, L.; Guo, W.; Meydani, M. Chemical structure of flavonols in relation to modulation of angiogenesis and immune-endothelial cell adhesion. J. Nutr. Biochem. 2006, 17, 165–176. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.M.; Bignon, J.; Haroun-Bouhedja, F.; Bittoun, P.; Vassy, J.; Fermandjian, S.; Wdzieczak-Bakala, J.; Boisson-Vidal, C. Inhibitory effect of fucoidan on the adhesion of adenocarcinoma cells to fibronectin. Anticancer Res. 2005, 25, 2129–2133. [Google Scholar]
- Di Mattia, C.; Battista, N.; Sacchetti, G.; Serafini, M. Antioxidant Activities in vitro of Water and Liposoluble Extracts Obtained by Different Species of Edible Insects and Invertebrates. Front. Nutr. 2019, 6, 106. [Google Scholar] [CrossRef]
- Arancibia-avila, P.; Fuentes, O.; Alarco, J.; Zamorano-ponce, E.; Herna, M. Production and Purification of Statins from Pleurotus ostreatus (Basidiomycetes ) Strains. Z. Naturforsch. C J. Biosci. 2003, 58, 62–64. [Google Scholar]
- Tamura, C.; Terahara, A. ML-236A, ML-236B, and ML-236C, new inhibitors of cholesterogenesis produced by Penicillium citrinium. J. Antibiot. 1976, 29, 1346–1348. [Google Scholar]
- Altwairgi, A.K. Statins are potential anticancerous agents. Oncol. Rep. 2015, 33, 1019–1039. [Google Scholar] [CrossRef]
- Gazzerro, P.; Proto, M.C.; Gangemi, G.; Malfitano, A.M.; Ciaglia, E.; Pisanti, S.; Santoro, A. Pharmacological Actions of Statins: A Critical Appraisal in the Management of Cancer. Pharmacol. Rev. 2012, 64, 102–146. [Google Scholar] [CrossRef]
- Iannelli, F.; Lombardi, R.; Milone, M.R.; Pucci, B.; Rienzo, S. De Targeting Mevalonate Pathway in Cancer Treatment: Repurposing of Statins. Recent Pat. Anticancer Drug Discov. 2018, 13, 184–200. [Google Scholar] [CrossRef]
- Takamatsu, S. Naturally occurring cell adhesion inhibitors. J. Nat. Med. 2018, 72, 817–835. [Google Scholar] [CrossRef] [PubMed]
- Schreiner, E.P.; Kern, M.; Steck, A.; Foster, C.A. Synthesis of ether analogues derived from HUN-7293 and evaluation as inhibitors of VCAM-1 expression. Bioorgan. Med. Chem. Lett. 2004, 14, 5003–5006. [Google Scholar] [CrossRef]
- Takamatsu, S.; Zhang, Q.; Schrader, K.K.; ElSohly, H.N.; Walker, L.A. Characterization of Mycotypha metabolites found ro be inhibitors of cell adhesion molecules. J. Antibiot. 2002, 55, 585–592. [Google Scholar] [CrossRef]
- Jeong, J.; Jin, C.; Park, C.; Han, M.I.N.H.O.; Kim, G.; Moon, S.; Kim, C.G.I.L.; Jeong, Y.K.E.E.; Kim, W.; Lee, J.; et al. Inhibition of migration and invasion of LNCaP human prostate carcinoma cells by cordycepin through inactivation of Akt. Int. J. Oncol. 2012, 40, 1697–1704. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Caro-diaz, E.J.E.; Lacoske, M.H.; Hung, C.; Theodorakis, E.A. EDGE ARTICLE Fusarisetin A: Scalable total synthesis and related studies. Chem. Sci. 2012, 12, 3378–3386. [Google Scholar] [CrossRef]
- Kasorn, A.; Loison, F.; Kangsamaksin, T. Terrein inhibits migration of human breast cancer cells via inhibition of the Rho and Rac signaling pathways. Oncol. Rep. 2018, 39, 1378–1386. [Google Scholar] [CrossRef]
- Mathieu, V.; Berger, W.; Evidente, A.; Kornienko, A.; Gailly, P.; Vandier, C. Ophiobolin A induces paraptosis-like cell death in human glioblastoma cells by decreasing BKCa channel activity. Cell Death Dis. 2013, 4, e561. [Google Scholar] [CrossRef]
- Bury, M.; Andolfi, A.; Rogister, B.; Cimmino, A.; Mégalizzi, V.; Mathieu, V.; Feron, O.; Evidente, A.; Kiss, R. Fusicoccin A, a phytotoxic carbotricyclic diterpene glucoside of fungal Origin, reduces proliferation and invasion of glioblastoma cells by targeting multiple tyrosine kinases. Transl. Oncol. 2013, 6, 112–123. [Google Scholar] [CrossRef]
- Teiten, M.; Mack, F.; Debbab, A.; Aly, A.H.; Dicato, M.; Proksch, P.; Diederich, M. Bioorganic & Medicinal Chemistry Anticancer effect of altersolanol A, a metabolite produced by the endophytic fungus Stemphylium globuliferum, mediated by its pro-apoptotic and anti-invasive potential via the inhibition of NF- j B activity. Bioorgan. Med. Chem. 2013, 21, 3850–3858. [Google Scholar] [CrossRef]
- Wang, X.; Bashyal, B.P.; Wijeratne, E.M.K.; Ren, J.M.U.; Liu, M.X.; Gunatilaka, M.K.; Arnold, A.E.; Gunatilaka, A.A.L. Smardaesidins A–G, Isopimarane and 20- nor -Isopimarane Diterpenoids from Smardaea sp., a Fungal Endophyte of the Moss Ceratodon purpureus 1. J. Nat. Prod. 2011, 74, 2052–2061. [Google Scholar] [CrossRef]
- Lee, H.J.; Lee, J.H.; Hwang, B.Y.; Kim, H.S.; Lee, J.J. Anti-Angiogenic Activities of Gliotoxin and Its Methylthio- Derivative, Fungal Metabolites. Arch. Pharm. Res. 2001, 24, 397–401. [Google Scholar] [CrossRef]
- Hayot, C.; Debeir, O.; Van Ham, P.; Van Damme, M.; Kiss, R.; Decaestecker, C. Characterization of the activities of actin-affecting drugs on tumor cell migration. Toxicol. Appl. Pharmacol. 2006, 211, 30–40. [Google Scholar] [CrossRef] [PubMed]
- Zhou, C.; Ling, M.; Lee, T.K.; Man, K.; Wang, X.; Wong, Y. FTY720, a fungus metabolite, inhibits invasion ability of androgen-independent prostate cancer cells through. Cancer Lett. 2006, 233, 36–47. [Google Scholar] [CrossRef] [PubMed]
- Kusama, T.; Mukai, M.; Iwasaki, T.; Tatsuta, M.; Matsumoto, Y.; Akedo, H. Inhibition of Epidermal Growth Factor-induced RhoA Translocation and Invasion of Human Pancreatic Cancer Cells by 3-Hydroxy-3-methylglutaryl-coenzyme A. Cancer Res. 2001, 61, 4885–4891. [Google Scholar]
- Alonso, D.F.; Farina, H.G.; Skilton, G.; Gabri, M.R.; de Lorenzo, M.S.; Gomez, D.E. Reduction of mouse mammary tumor formation and metastasis by lovastatin, an inhibitor of the mevalonate pathway ofcholesterol synthesis. Breast Cancer Res. Treat. 1998, 50, 83–93. [Google Scholar] [CrossRef]
- Zhong, W.; Liang, Y.; Wang, C.; Chang, T.; Lee, W. Lovastatin suppresses invasiveness of anaplastic thyroid cancer cells by inhibiting Rho geranylgeranylation and RhoA / ROCK signaling. Endocr. Relat. Cancer 2005, 3, 615–630. [Google Scholar] [CrossRef]
- Bublik, R.; Alonso, D.F.; Gomez, D.E. Lovastatin alters cytoskeleton organization and inhibits experimental metastasis of mammary carcinoma cells. Clin. Exp. Metastasis 2002, 19, 551–559. [Google Scholar]
- Afshordel, S.; Kern, B.; Clasohm, J.; König, H.; Priester, M.; Weissenberger, J.; Kögel, D.; Eckert, G.P. Lovastatin and perillyl alcohol inhibit glioma cell invasion, migration, and proliferation–Impact of Ras-/Rho-prenylation. Pharmacol. Res. 2015, 91, 69–77. [Google Scholar] [CrossRef]
- Glynn, S.A.; Sullivan, D.O.; Eustace, A.J.; Clynes, M.; Donovan, N.O. The 3-hydroxy-3-methylglutaryl-coenzyme A reductase inhibitors, invasion of melanoma cells. BMC Cancer 2008, 8, 9. [Google Scholar] [CrossRef] [PubMed]
- Hayashi, M.; Kim, Y.P.; Hiraoka, H.; Natori, M.; Takamatsu, S.; Kawakubo, T.; Masuma, R.; Komiyama, K.; Somura, S. Macrosphelide, a novel inhibitor of cell-cell adhesion molecule. I. Taxonomy, fermentation, isolation and biological activities. J. Antibiot. 1995, 48, 1435–1439. [Google Scholar] [CrossRef]
- Takamatsu, S.; Hiraoka, H.; Kim, Y.; Hayashi, M.; Natori, M.; Komiyama, K.; Omura, S. Macrosphelides C and D, Novel Inhibitors of Cell Adhesion. J. Antibiot. 1997, 50, 878–880. [Google Scholar] [CrossRef] [PubMed]
- Fukami, A.; Taniguchi, Y.; Nakamura, T.; Rho, M.; Kawaguchi, K.; Hayashi, M.; Komiyama, K.; Omura, S. New Membersof the Macrosphelides from Microsphaeropsis sp. FO-5050 IV. J. Antibiot. 1999, 52, 501–504. [Google Scholar] [CrossRef]
- Miki, I.; Ishihara, N.; Otoshi, M.; Kase, H. Simple colorimetric cell-cell adhesion assay using MTT-stained leukemia cells. J. Immunol. Methods 1993, 164, 255–261. [Google Scholar] [CrossRef]
- Yamada, T.; Iritani, M.; Doi, M.; Minoura, K.; Ito, T.; Numata, A. Absolute stereostructures of cell-adhesion inhibitors, macrosphelides C, E–G and I, produced by a sea hare-derived Periconia sp. J. Antibiot. 2001, 3046–3053. [Google Scholar] [CrossRef]
- Yamada, T.; Iritani, M.; Minoura, K.; Numata, A. Absolute Stereostructures of Cell Adhesion Inhibitors, Macrosphelides H and L, from Periconia byssoides OUPS-N133. J. Antibiot. 2002, 55, 147–154. [Google Scholar] [CrossRef] [PubMed]
- Yamada, T.; Minoura, K.; Tanaka, R.; Numata, A. Cell-adhesion Inhibitors Produced by a Sea Hare-derived Periconia sp. J. Antibiot. 2007, 60, 370–375. [Google Scholar] [CrossRef]
- Fukami, A.; Iijima, K.; Hayashi, M.; Komiyama, K.; Omura, S. Macrosphelide B Suppressed Metastasis through Inhibition of Adhesion of sLe x / E-Selectin Molecules. Biochem. Biophys. Res. Commun. 2002, 1070, 1065–1070. [Google Scholar] [CrossRef]
- Yamada, T.; Iritani, M.; Minoura, K.; Kawai, K.; Numata, A. Peribysins A–D, potent cell-adhesion inhibitors from a sea hare-derived culture of Periconia species. Org. Biomol. Chem. 2004, 2, 2131–2135. [Google Scholar] [CrossRef]
- Yamada, T.; Doi, M.; Miura, A.; Harada, W.; Hiramura, M.; Minoura, K.; Tanaka, R.; Numata, A. Absolute Stereostructures of Cell-adhesion Inhibitors, Peribysins A, E, F and G, Produced by a Sea Hare-derived Periconia sp. J. Antibiot. 2005, 58, 185–191. [Google Scholar] [CrossRef]
- Yamada, T.; Minoura, K.; Tanaka, R.; Numata, A. Cell-adhesion Inhibitors Produced by a Sea Hare-derived Periconia sp. II. J. Antibiot. 2006, 59, 345–350. [Google Scholar] [CrossRef]
- Sigrist, C.J.A.; Bridge, A.; Le Mercier, P. A potential role for integrins in host cell entry by SARS-CoV-2. Antiviral Res. 2020, 177, 104759. [Google Scholar] [CrossRef] [PubMed]
- Kiss, B.; Kis, Z.; Pályi, B.; Kellermayer, M.S.Z. Topography, Spike Dynamics, and Nanomechanics of Individual Native SARS-CoV-2 Virions. Nano Lett. 2021, 21, 2675–2680. [Google Scholar] [CrossRef] [PubMed]
- Kanyo, N.; Kovacs, K.D.; Saftics, A.; Szekacs, I.; Peter, B.; Santa-Maria, A.R.; Walter, F.R.; Dér, A.; Deli, M.A.; Horvath, R. Glycocalyx regulates the strength and kinetics of cancer cell adhesion revealed by biophysical models based on high resolution label-free optical data. Sci. Rep. 2020, 10, 22422. [Google Scholar] [CrossRef]
- Dakal, T.C. SARS-CoV-2 attachment to host cells is possibly mediated via RGD-integrin interaction in a calcium-dependent manner and suggests pulmonary EDTA chelation therapy as a novel treatment for COVID 19. Immunobiology 2021, 226, 152021. [Google Scholar] [CrossRef] [PubMed]
- Yang, Q.; Hughes, T.A.; Kelkar, A.; Yu, X.; Cheng, K.; Park, S.J.; Huang, W.C.; Lovell, J.F.; Neelamegham, S. Inhibition of SARS-CoV-2 viral entry in vitro upon blocking N- and O-glycan elaboration. bioRxiv 2020, 1–19. [Google Scholar] [CrossRef]
- Bhuiyan, F.R.; Howlader, S.; Raihan, T.; Hasan, M. Plants Metabolites: Possibility of Natural Therapeutics Against the COVID-19 Pandemic. Front. Med. 2020, 7, 444. [Google Scholar] [CrossRef]
- Antonelli, M.; Donelli, D.; Maggini, V.; Firenzuoli, F. Phytotherapic compounds against coronaviruses: Possible streams for future research. Phyther. Res. 2020, 34, 1469–1470. [Google Scholar] [CrossRef]
- Seifert, G.; Jeitler, M.; Stange, R.; Michalsen, A.; Cramer, H.; Brinkhaus, B.; Esch, T.; Kerckhoff, A.; Paul, A.; Teut, M.; et al. The Relevance of Complementary and Integrative Medicine in the COVID-19 Pandemic: A Qualitative Review of the Literature. Front. Med. 2020, 7, 946. [Google Scholar] [CrossRef]
- Subbaiyan, A.; Ravichandran, K.; Singh, S.V.; Sankar, M.; Thomas, P.; Dhama, K.; Malik, Y.S.; Singh, R.K.; Chaudhuri, P. In silico molecular docking analysis targeting SARS-CoV-2 Spike protein and selected herbal constituents. J. Pure Appl. Microbiol. 2020, 14, 989–998. [Google Scholar] [CrossRef]
- Liu, J.; Bodnar, B.H.; Meng, F.; Khan, A.I.; Wang, X.; Saribas, S.; Wang, T.; Lohani, S.C.; Wang, P.; Wei, Z.; et al. Epigallocatechin gallate from green tea effectively blocks infection of SARS-CoV-2 and new variants by inhibiting spike binding to ACE2 receptor. Cell Biosci. 2021, 11, 1–15. [Google Scholar] [CrossRef]
- Kalus, U.; Grigorov, A.; Kadecki, O.; Jansen, J.P.; Kiesewetter, H.; Radtke, H. Cistus incanus (CYSTUS052) for treating patients with infection of the upper respiratory tract. A prospective, randomised, placebo-controlled clinical study. Antiviral Res. 2009, 84, 267–271. [Google Scholar] [CrossRef] [PubMed]
- Bramer, W.M.; De Jonge, G.B.; Rethlefsen, M.L.; Mast, F.; Kleijnen, J. A systematic approach to searching: An efficient and complete method to develop literature searches. J. Med. Libr. Assoc. 2018, 106, 531–541. [Google Scholar] [CrossRef] [PubMed]



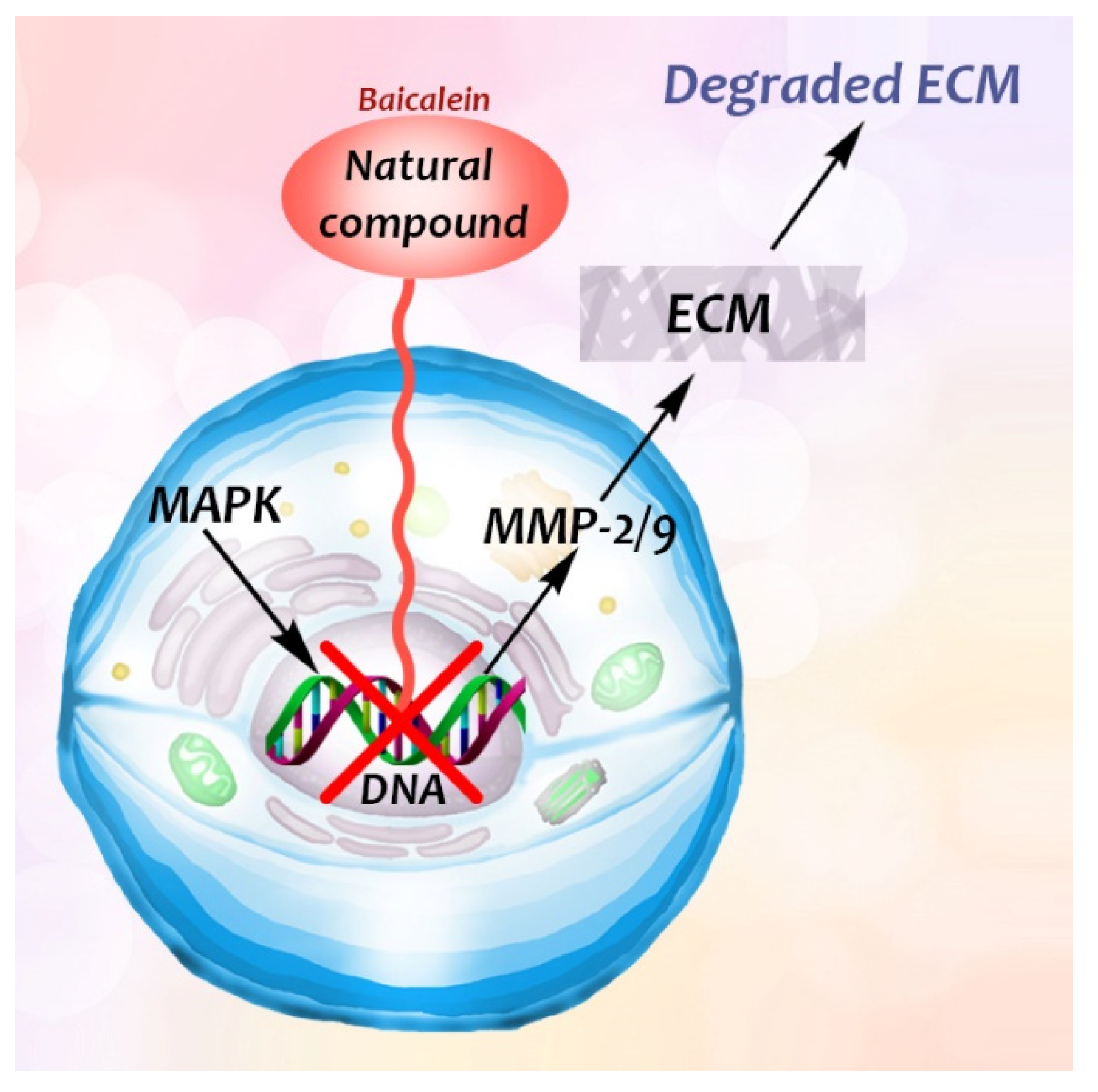
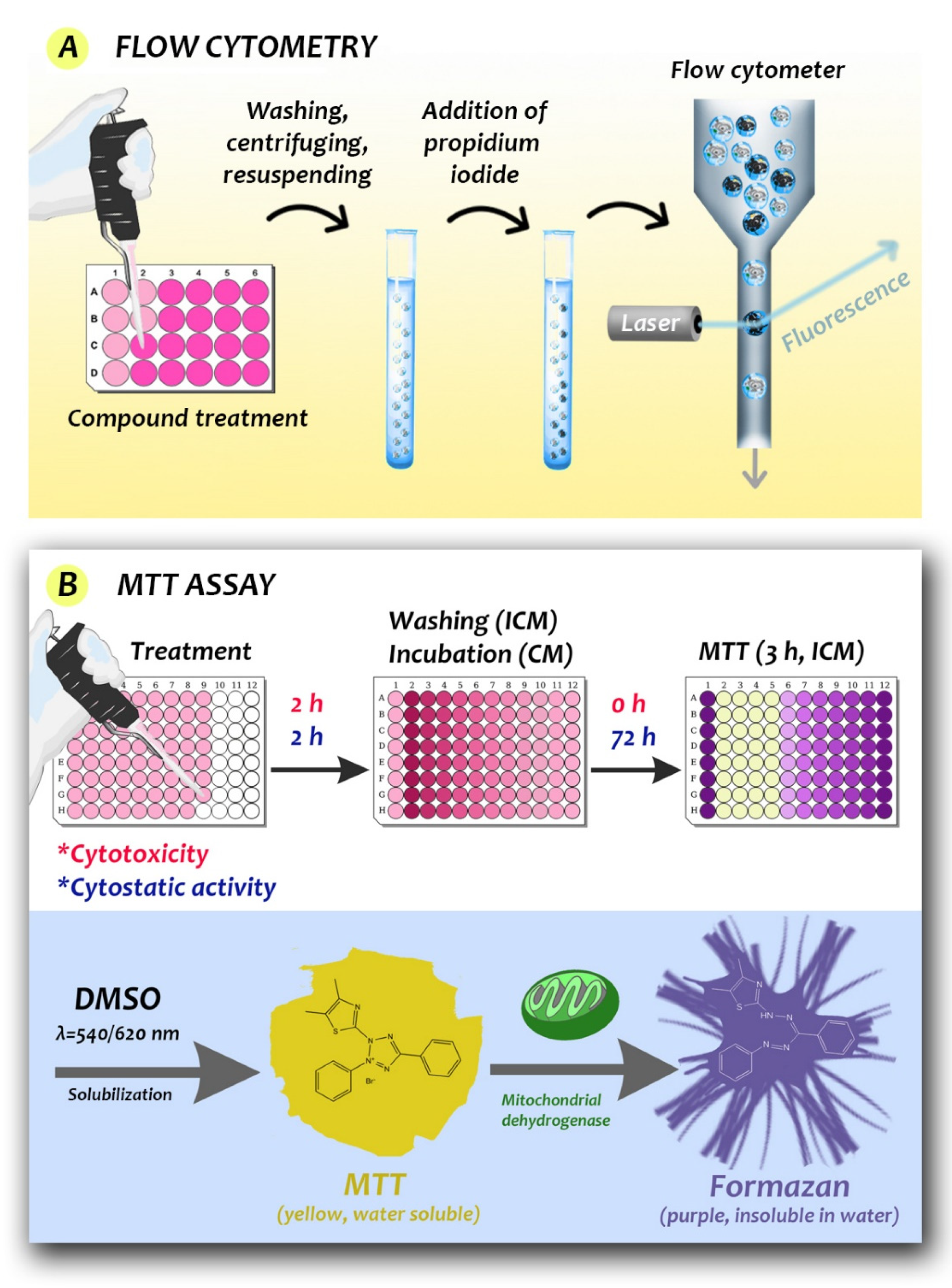
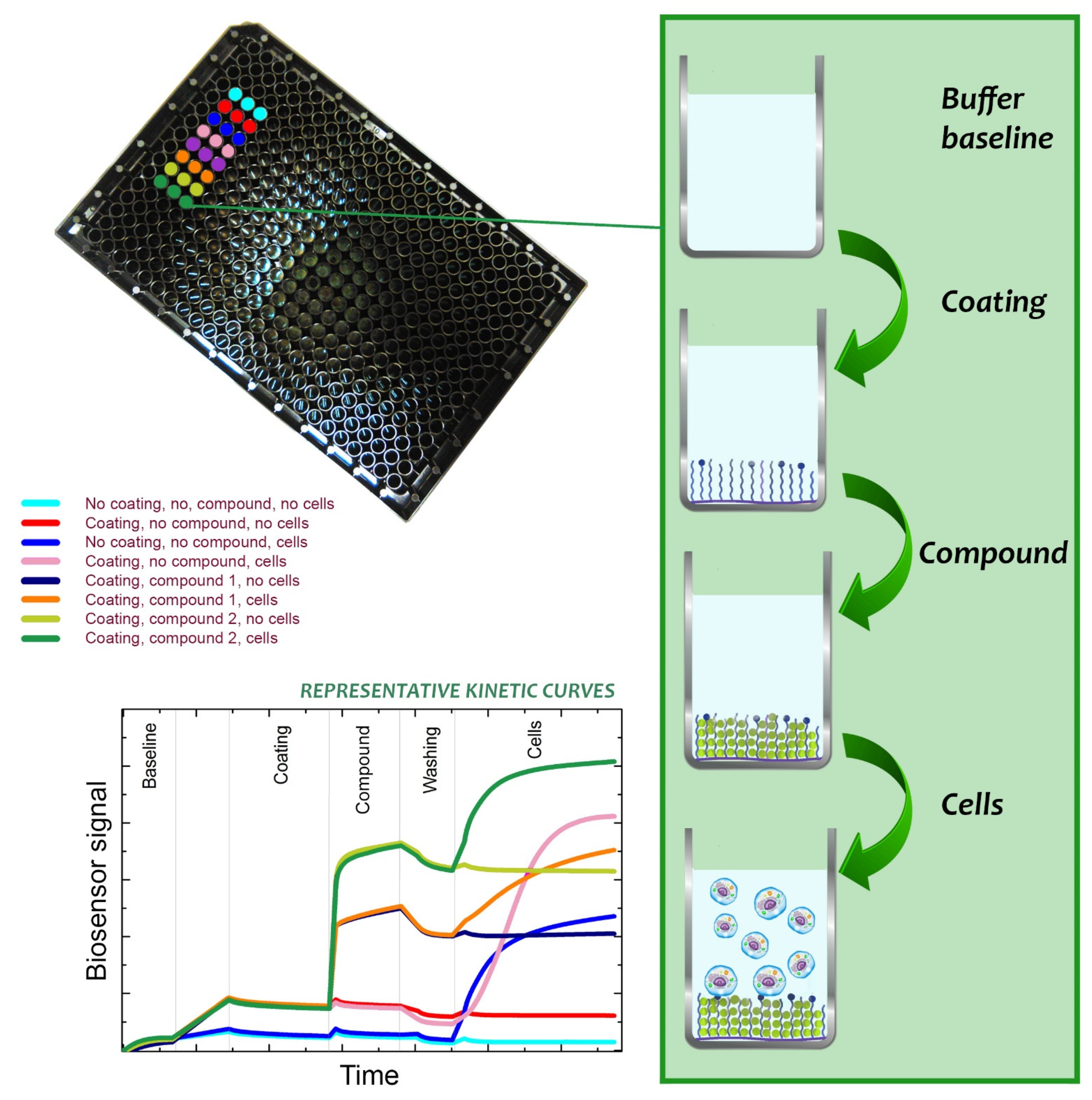
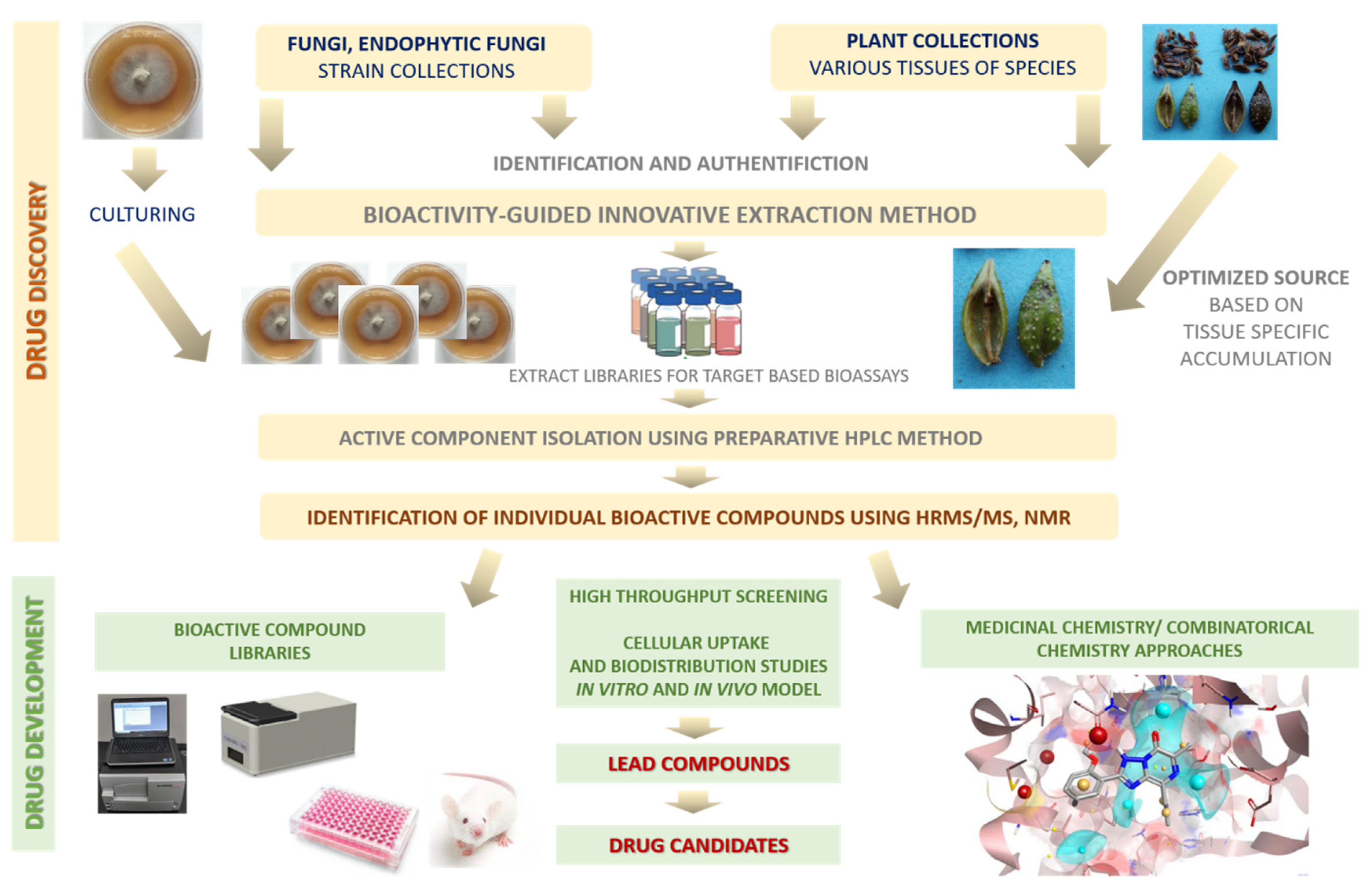

| Integrin | Ligands | Functions |
|---|---|---|
| β1 | RGD, VCAM-1, E-cadherin | Adhesion |
| β2 | iC3b, fibrinogen, ICAM-1 | Adhesion, phagocytosis, apoptotic cell clearance |
| β6 | RGD, fibronectin | Adhesion, endocytosis, inflammation |
| β7 | RGD, VCAM-1 | Adhesion, inflammation |
| Mode | Action |
|---|---|
| P1 | Inhibiting the dephosphorylation of I κB and, hence, the activation of NF-κB |
| P2 | Inhibiting translocation of activated NF-κB into the nucleus |
| P3 | Inhibition of binding of activated NF-κB to promoter sites for CAM expression |
| Mechanism | Action |
|---|---|
| B1 | Blocking adhesion receptors on the surface of mobile cells (e.g., leukocytes) |
| B2 | Blocking adhesion receptors on the surface of tissue cells (e.g., of the endothelium) |
| B3 | Blocking adhesion motifs in the extracellular matrix (ECM) |
| Cell Viability and Cytotoxicity Assays Absorbance-Based or Colorimetric | |||||
|---|---|---|---|---|---|
| Principal | Advantage | Disadvantage | References | ||
| Dye exclusion assays | Trypan blue | membrane integrity | simple, easy to use, economic, fast | hemocytometer required, counting error, difficult to process large number of samples, cannot be distinguished healthy cells and cell function lost alive cells, toxic on mammalian cells | Jonston 2010 [69], Strober 2001 [70], Aslantürk and Celik 2013 [71], Stone et al., 2009 [72], Yip and Auersperg 1972 [73], Ruben 1988 [74] |
| Eosin (Erythrosin B) | membrane integrity | economic, versatility, biosafety | time-consuming, labor-intensive, sensitive for the contamination, filling rate, and inter-user variation | Kim et al., 2016 [75], Marmion 1979 [76] | |
| Colorymetric assays | MTT | enzyme (mitochondrial) activity | easy to use, safe, high reproductibility, widly used | organic solvents required, significant well-to-well error, interaction with compounds results false positive, or false negative data | Stone et al., 2009 [72], Aslantürk et al. [77], Bopp and Lettieri 2008 [78], Mosmann 1983 [79] |
| MTS | easy to use, rapid, sensitive, economic, specific, inexpensive | measured absorbance level is influenced by incubation time, cell type, and cell number; optimal incubation time: 1–3 h | Berg et al., 1994 [80], Tominaga et al., 1999 [81], Rotter et al., 1993 [82], Buttke et al., 1993 [83], Promega Technical Bulletin [84], Cory et al., 1991 [85], Riss et al., 1992 [86] | ||
| XTT | results water soluble crystals, quick, sensitive, easy-to-use, safe; highly sensitive and accurate | strongly depends on reductive capacity of viable cells due to pH, cellular ion concentration, cell cycle variation, environmental factors | Scudiero et al.1988 [87] | ||
| WST-1 | easy to use, high reproducibility, wildly used, no interference with phenol red, water soluble dye, no additional incubation time | relatively long incubation time (2 h) | Ishiyama et al., 1993 [88] | ||
| WST-8 | not cell permeable; cells can be used after the assay; water soluble formazan | intracellular metabolic activity can influence the reduction of WST-8 | Tominaga et al., 1999 [81], Strober 2001 [70] | ||
| LDH (lactate dehydrogenase) | enzyme (lactate dehydrogenase) activity | reliability; quick, simple evaluation, indicator of cell death | serum incompetence | Decker and Lohmann-Matthes 1988 [89], Schins et al., 2002 [90], Fotakis et al., 2006 [91], Lappalanien et al., 1994 [92] | |
| SRB (sulforhodamine B) | SRB-protein interaction | simple, fast, sensitive, good linearity with cell number, metabolism independent, high reproducibility | homogenous cell suspension is required; cellular clumps and aggregates interfere with SRB | Skehan et al., 1990 [93] | |
| NRU (neutral red uptake) | neutral red uptake and lysosomal accumulation | good marker for lysosomal damage, fast and simple evaluation | influenced by pollutants | Borenfreund and Puerner 1984 [94], Repetto et al., 2008 [95], Ringwood et al.1998 [96] | |
| Crystal violet | binding for proteins and DNA of viable cells | quick; chemical inhibtors can be incorporated into the assay | metabolism affected compounds can not be tested, not able to measure cell proliferation rate | Feoktisova et al., 2016 [66], Geserick et al., 2009 [97], Degterev et al., 2008 [98], Sun et al., 2012 [99], Feoktisova et al., 2011 [100] | |
| Fluorometric Assay | ||||
|---|---|---|---|---|
| Principal | Advantage | Disadvantage | References | |
| AlamarBlue (resazurin reduction assay) | enzyme (mitochondrial and other (e.g., Diaphorases)) activity | relatively inexpensive, sensitive, multiplexed with other methods (e.g., Caspase activity) | fluorescence interference with tested compound; direct cytotoxic effect can be occured (depending on incubation time) | O’Brien et al., 2000 [101], Ahmed et al., 1994 [102], Page et al., 1993 [103], Markossian et al., 2004 [65], Pace et al., 2013 [104] |
| CFDA-AM (5-carboxyfluorescein diacetate, acetoxymethyl ester) | plasma membrane integrity | can be used parallel with alamarBlue on the same set of cells | fluorescence interference with tested compound | Bopp et al., 2006 [103], Schreer et al., 2005 [105], Ganassi 2000 [106] |
| protease viability marker assay; GF-AFC assay (glycylphenyl-alaninyl)-aminofluoro-cumarine) | enzyme (aminopeptidase) activity | relatively nontoxic; multiplex with other assays, short incubation time (30–60 min) | fluorescence interference with tested compound | Niles et al., 2009 [62] |
| BrdU (bromoeoxyuridin) and EdU (5-ethynyl-2 deoxyuridine) | DNA synthesis | rapid, highly selective, results in the most reliable and direct index of proliferation, in contrast to 3H-thymidine incorporation assay, which requires a scintillation beta-counter, BrdU and EdU can be detected by antibodies, allowing analysis by flow cytometry or by immuno-histochemistry | BrdU is toxic and mutagenic, alters cell cycle, required DNA denaturation | Sidman et al., 1959 [107] Miller and Nowakowski 1988 [108] Salic and Mitchison 2008 [109] Nowakowski et al. 1989 [110] Taupin 2007 [111] P.L Duque and Rakic 2011 [112] |
| Luminometric Assay | ||||
|---|---|---|---|---|
| Principal | Advantage | Disadvantage | References | |
| ATP assay | membrane integrity | the fastest and the most sensitive assay to use; no artefacts; no plate handling step | sensitivity depends on reproducibility of pipetting | Maehara et al., 1987 [113], García et al., 2003 [114], Andreotti et al., [115], Markossian et al., 2004 [65] |
| Real-time viability assay | metabolic activity | real-time measurement; multiplex | incubation time is cell type and seeding density dependent | Duellman et al., 2015 [116], Markossian et al., 2004 [65] |
| Cell Viability Assays for Flow Cytometry and Microscopic Imaging | |||||
|---|---|---|---|---|---|
| λex | Advantage | Disadvantage | References | ||
| Nucleic acid dies | Propidium iodide | 488 and 561 nm | can be added directly to the samples | not membrane permeable to live cells, not possible to use on fixed cells, can also bind to RNA not only to DNA, toxigenic and mutagenic | Suzuki et al., 1997 [117] |
| 7-amino-actinomycin-D (7-AAD) | 488 and 561 nm | can be added directly to the samples, can be used in combination with formaldehyde fixation | not membrane permeable to live cells, potential carcinogen | Liu et al., 1991 [118] Latt 1977 [119] | |
| Hoechst 33342 | 350 and 454 nm | not membrane permeable to live cells | Liu et al., 2019 [120], Réu et al., 2019 [121] | ||
| Helix NP™ Green, NIR and Blue and SYTOX® based dyes | wide range from 444–640 nm | it can also be used for viability in microscopy on live cells or as a nuclear counterstain on fixed and permeabilized cells and tissue sensitive nucleic acid stain, in combination with resuzarine can be used, works with mammalian and Gram-positive and Gram-negative bacteria, can be incorporated SYTOX® stains into a number of assays for apoptosis, cell viability, and metabolism, easy-to-use. | non cell-permeable to live cells | Avlasevich et al., 2006 [122], Yan et al., 2000 [123], Bryce et al., 2007 [124], Mukhopadhyay et al., 2007 [125] | |
| DRAQ7™ far-red fluorescent DNA dye | 633 nm | can be combined with FITC, PE, and other UV or violet excitable dyes for multicolor analysis, non toxic, can be used for siRNA studies and other dynamic viablity assays | not membrane-permeable to live cells | Kerscher et al., 2019 [126] Paivandy et al., 2019 [127] Vig et al., 2019 [128] Akagi et al., 2013 [129] | |
| Ethidium bromide | 518 nm | can be used on fixed cells, economic | not membrane-permeable, intercalates double-stranded DNA and RNA, mutagen, carcinogen, | Severini and Morgan 1991 [130] | |
| SYBR® Green | 494 nm | highly selective for DNA, can be used in combination with propidium iodide | carcinogen, less mutagenic than ethidium bromide | Zipper et al., 2004 [131], Singer et al., 1999 [132] | |
| Acridine orange | 500 nm | cell permeable, low-cost, sensitive, rapid, intercalate to DNA, and electrostatically interact with RNA, sensitive for pH, acidic organells can be also detected, compatible for ethidium bromide and propidium iodide | nonfixable | Mirrett 1982 [133], Kumar et al., 2012 [134], Darzynkiewicz et al., 2004 [135] | |
| Protein binding dies | eFluor fixable dyes | wide range from 401–645 nm | traditional organic fluorescence dies, multiple application, fully compatible with most convential dies | Lekishvili et al., 2018 [136] | |
| BD Horizon Brilliant dyes | polymer dyes with brighter fluorescence signal | BD Biosciences products [137] | |||
| Biolegend Zombie dyes | wide range from 360–633 nm | Pardo-Garcia et al., 2015 [138], McMaster et al., 2015 [139], Rodríguez- Rodríguez et al., 2015 [140], Files et al., 2015 [141], Akabane et al., 2016 [142], Iraolagoitia et al., 2016 [143], Mercer et al., 2016 [144], Souza-Fonseca-Guimaraes et al., 2015 [145], Matsui et al., 2015 [146], Jones et al., 2015 [147], Nath et al., 2015 [148], Kanemaru et al., 2015 [149], Tabalot-Ayer et al., 2015 [150], Keppel et al., 2015 [151], Shade et al., 2015 [152], Weiser et al., 2015 [153] | |||
| Calcein AM | 496 | cell permeant, indicator of lipid vesicle leakage, neutral substrate for MDR efflux transporters, selective for live cells; suitable for proliferating and non-proliferating cells; ideal for suspension and adherent cells, rapid, ideal for high-throughput assays; commonly used for cell tracing and in studies of endocytosis, cell migration, and gap junctions; adaptable to a wide variety of techniques, including: microplate assays, immunocytochemistry, flow cytometry, and in vivo cell tracking | Allen and Cleland 1980 [154], Patel et al., 2009 [155], Glavinas et al., 2004 [156] | ||
| Migration Assays | ||||
|---|---|---|---|---|
| Principal | Advantage | Disadvantage | References | |
| Transwell migration and invasion assay (Boyden chamber) | chemotaxis, migration, invasion | easy setup, most frequently used method, can be visualized by cytological dyes, stained fluorescent, or lysed and assessed by a plate reader; invasive index can be calculated | endpoint measurement, only adherent cells can be used, only vertical movement can be detected, must be optimized for each cell type | Menyhárt et al., 2016 [64], Kramer et al., 2013 [163], Boyden 1962 [164], Restouin et al., 2009 [165], Marshall 2011 [166] |
| Scratch (wound healing) assay | two dimensional (2D) cell migration in confluent, monolayer cell cultures | simple, easy setup, low cost, kinetic measurement, can be combined with other techniques (e.g., gene transfections), allows high -throughput screening (HTS) | only adherent cells can be used, only horizontal movement can be detected, require relatively large cell and reagent quantities, is not ideal for chemotaxis studies | Menyhárt et al., 2016 [64], Liang et al., 2007 [167], Győrffy et al., 2015 [168], Gorshkova et al., 2008 [169], Lo et al., 1995 [170], Tamada 2007 [171], Zordan et al., 2011 [172], Poujade et al., 2007 [173], Simpson et al., 2008 [174] |
| Cell exclusion zone assays | migration | Several kinds of barrier can be used (e.g., glass, silicone, metal, Teflon, microfabricated soft and elastic “stencils” or agarose gels), cells are visualized several manner (e.g., photomicrography or labeled with fluorescence and measured with a microplate reader), migratory capacity and interaction between two different populations can be compared, kinetic measurement, this is currently the only method that allows investigation of the effects of ECM proteins on cell motility, allows high -throughput screening (HTS) | only adherent cells can be used, only horizontal movement can be detected | Menyhárt et al., 2016 [64], Poujade et al., 2007 [173], Gough et al., 2011 [175], Pratt et al., 1984 [176] Varani et al., 1978 [177] Kroening 2010 [178] |
| Microcarrier bead and spheroid migration assays | migration | adherent cells and spheroids can be used, vertical and horizontal movement, plastic surface and beads can be used, microscope, time-lapse microscope and fluorescent staining can be used to count the cells, endpoint and kinetic measurement can be performed | Menyhárt et al., 2016 [64], Rosen et al., 1990 [179], Konduri et al., 2001 [180] | |
| The capillary chamber migration assay | migration and morphology | adherent and suspension cells can be used, migratory behavior and morphological responses can be visualized in real time by time-lapse microscopy, endpoint and kinetic measurement can be performed, suitable for rare cell types and expensive compounds due to its small cell and volume require, liquid handling and image processing can be fully automated, allows high- throughput imaging system (HTIS) | only horizontal movement can be detected | Menyhárt et al., 2016 [64], Chaubey et al., 2011 [181], Echeverria et al., 2010 [182] |
| Motility of individual cells | migration | adherent and suspension cells can be used, time-lapse video microscopy, 3D tracking is also possible, analysis of invasive properties of single cells as well as of populations can be performed, kinetic measurement | only horizontal movement can be detected, requires specialized microscopes and image analyzing software | Menyhárt et al., 2016 [64], Miura 2005 [183], Gu et al., 2007 [184], Lin et al., 2005 [185], Niinaka et al., 2001 [186] |
| Mechanical properties | migration and invasion in 3D structures | 3D force microscope and coated magnetic beads can be also used, high-speed video camera, video spot tracker can be applied | only adherent cells can be tested, only horizontal movement can be detected, endpoint assay | Menyhárt el al. 2016 [64], Swaminathan et al., 2011 [187], Cribb et al., 2016 [188] |
| Active Substance | IUPAC Name (CAS Number) | Source | Cellular Effect | Effect Type | Molecular Mechanisms | Tested Cell Line 1, Animal | Method | Ref. |
|---|---|---|---|---|---|---|---|---|
Cistifolin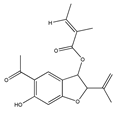 | 5-acetyl-6-hydroxy-2-(prop-1-en-2-yl)-2,3-dihydrobenzofuran-3-yl (E)-2-methylbut-2-enoate (31685-99-1) | Eupatorium purpureum (rhizome) | Inhibition of cell adhesion | No data | Modulation of integrin-dependent cell-cell and cell-protein interaction | U937 EA.hy926 | Chromatography, Homotypic cell aggregation assay, Cell attachment assay, Carrageenan oedema test | Habtemariam. 1998 [58] |
Ursolic acid (UA)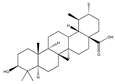 | (3β)-3-Hydroxyurs-12-en-28-oic acid (77-52-1) | Berries and wax-like coatings of apples, pears, etc. | Inhibition of cancer cell adhesion, invasion and migration. Preventing cancer metastasis. Inhibition on proliferation. Inhibition of cell invasion | P | ICAM-1 downregulation | MHCC-97H MHCC-97L HepG2 M619 MDA-MB-231v MCF-7 HT29 SW620 B16-F10 Sprague Dawley rats (males) | MTT cell viability assay, Adhesion assay (fluorescence microscope photographed method), Invasion assay (matrigel coated transwell inserts), Wound healing assay, Flow cytometry, Western blot, Microarray assay, Real-time PCR, In vivo tumor growth and metastasis assay, ICAM-1 immunohistochemistry UPLC-MS/MS | Xiang et al., 2015 [229] |
Ellagic acid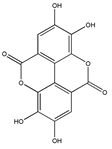 | 2,3,7,8-tetrahydroxychromeno[5,4,3-cde]chromene-5,10-dione (476-66-4) | Juglans regia | Antiinflammatory activity and osteoblastic activity | P | Inhibition of the effect on TNF-α-induced VCAM-1 and ICAM-1 expression | HAEC KS483 | HPLC, Cell-enzyme-linked immunosorbent assay (cell-ELISA), MTT assay | Papoutsi et al., 2007 [18] |
Eugenol | 4-allyl-2-methoxyphenol (97-53-0) | Syzygium aromaticum | Inhibition of trophozoite adhesion. Anti-Giardia activity | No data | No data | Gardia lamblia (WB strain [ATCC 30957]) | GC, GC/MS, Light microscope, Neubauer cell-counter chamber, Morphological assay, Transmission and scanning electron microscopy, MTT assay | Machado et al., 2011 [230] |
Elenolic acid,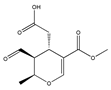 | 2-[(2S,3S,4S)-3-formyl-5-methoxycarbonyl-2-methyl-3,4-dihydro-2H-pyran-4-yl]acetic acid (34422-12-3) | Olea europaea and Vitis vinifera | Decreased monocytoid cell adhesion to HUVECs. Anti-inflammatory activity | P | VCAM-1 down-expression | HUEVEC BAEC U937 | Enzyme immunoassay (EIA), Cell number and viability assessment (Trypan blue,) Adhesion assay, Northern analysis, Transfection assays (calcium phosphate precipitation method), Electrophoretic mobility shift assay (EMSA) | Carluccio et al., 2003 [21] |
Hydroxytyrosol,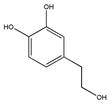 | 4-(2-hydroxyethyl)benzene-1,2-diol (10597-60-1) | |||||||
Oleuropein, | (2S,3E,4S)-3-ethylidene-2-(β-D-glucopyranosyloxy)-3,4-dihydro-5-(methoxycarbonyl)-2H-pyran-4-acetic acid, 2-(3,4-dihydroxyphenyl)ethyl ester (32619-42-4) | |||||||
Resveratrol | 5-[(E)-2-(4-hydroxyphenyl)ethenyl]benzene-1,3-diol (501-36-0) | |||||||
Tyrosol, | 4-(2-hydroxyethyl)phenol (501-94-0) | |||||||
Epigallocatechin-gallate (EGCG) | Camellia sinensis | Inhibition of cancer cell adhesion | B | EGCG forms multilayers in the PLL-g-PEG–RGD chains, bocking the RGD motifs | HeLa | Epic BT (RWG label-free biosensor) | Peter et al., 2017 [50] | |
| (2R,3R)-5,7-dihydroxy-2-(3,4,5-trihydroxyphenyl)chroman-3-yl 3,4,5-trihydroxybenzoate (989-51-5) | Inhibition of cancer cell movement (motility, average motility speed and migration) | No data | No data | HeLa | HoloMonitor M4 | Peter et al., 2015 [195] | ||
| Inhibition of cancer cell adhesion to laminin | B | Binding of EGCG to laminin | B16 | Trypan blue dye exclusion assay Affinity chromatography | Suzuki and Isemura, 2001 [231] | |||
Curcumin | (1E,6E)-1,7-bis(4-hydroxy-3-methoxyphenyl)hepta-1,6-diene-3,5-dione (458-37-7) | Curcuma longa | Suppressed monocyte and HUVEC adhesion | P | Down-expression of adhesion molecules. Blocks the NF- κB signaling pathway | HUVEC THP-1 | WST-1 colorimetry assay, ELISA, Western Blot | Kawasaki et al., 2015 [13] |
| Dose dependent decrease of cell-cell adhesion, especially on tumor-derived spheroids. Decreased cell proliferation in cell lines with mesenchymal characteristics. Decreased the migration speed of highly migratory cells. Decreased tumor growth and aggressiveness. | No data | No data | HACAT NIH-3T3 SCC25 CAL27 HCPA.BALB/c (nude mice) | CyQUANT NF cell proliferation assay kit, Fluorometer, Spheroid assay, Time-lapse analysis (phase microscopy, inverted microscope), Flow cytometry, Xenograft model | Santos de Campos et al., 2017 [232] | |||
Ethyl 3’,4’,5’-trimethoxycinamate (1), | (1) ethyl 3,4,5-trimethoxybenzoate (6178-44-5) | Piper longum | Anti-inflammatory effect. (1) Significantly but reversibly blocked the adhesion of neutrophils to endothelium | P | Down-expression of ICAM-1 | Primary endothelial cells from human umblical cord | Trypan blue exclusion test, MTT assay, ELISA, Flow cytometry, NMR, TLC | Kumar et al., 2005 [46] |
piperine (2) | (2) (2E,4E)-5-(benzo[d][1,3]dioxol-5-yl)-1-(piperidin-1-yl)penta-2,4-dien-1-one (94-62-2) | Neutrophil cells from peripheral blood of healthy individuals | ||||||
Saponin astagaloside IV (AS-IV) | (3β,6α,9β,16β,20R,24S)-16,25-dihydroxy-3-(β-D-xylopyranosyloxy)-20,24-epoxy-9,19-cyclolanostan-6-yl β-D-glucopyranoside (84687-43-4) | Astagalus membranaceus | Reduced adhesion of LPS-stimulated HUVECs for PMNs and THP-1 cells | P | Decreased the LPS-induced expression of E-selectin and VCAM-1 on the surface of HUVECs. Decreased the LPS- and TNFα-induced specific mRNA levels for E-selectin and VCAM-1. Abolished LPS- and TNFα-induced nuclear translocation of NF- κB and NF- κB DNA binding activity in endothelial cells | HUVEC THP-1 PMN | ELISA, Northern blot, Electrophoretic mobility shift assay (EMSA), Immunofluorescence staining | Zhang et al., 2003 [22] |
Galangin (G) | (G) 3,5,7-trihydroxy-2-phenyl-4H-chromen-4-one (548-83-4) | Alpinia officinarium (G) | Dose-dependent suppression of U937 cell adhesion to HUVEC monolayer | P | M and Q reduced the IL-1β stimulated expression of VCAM-1, ICAM-1 and E-selectin | HUVEC U937 | Trypan blue exclusion test, Ferric-reducing ability assay of plasma method (FRAP), Cytofluor fluorescence multiwell plate reader (dichlorofluorescein), Fluoresent labeling (BCECF-AM), ELISA | Kim et al., 2006 [233] |
Kaempferol (K) | (K) 3,5,7-trihydroxy-2-(4-hydroxyphenyl)-4H-chromen-4-one (520-18-3) | Gingko biloba, fruits and vegetables (K) | ||||||
Myricetin (M) | (M) 3,5,7-trihydroxy-2-(3,4,5-trihydroxyphenyl)-4H-chromen-4-one (529-44-2) | Berries, Camellia sinensis, etc. (M) | ||||||
Quercetin (Q) | (Q) 2-(3,4-dihydroxyphenyl)-3,5,7-trihydroxy-4H-chromen-4-one (117-39-5) | Vitis vinifera, Camellia sinensis,fruits and vegetables (Q) | ||||||
Baicalein | 5,6,7-trihydroxy-2-phenyl-4H-chromen-4-one (491-67-8) | Scutelaria baicalensis | Suppressed MDA-MB-231 cell adhesion to fibronectin-coated substrate | P | Down-regulated the expression of MMP-2/9 involved MAPK signaling pathway | MDA-MB-231 | MTT assay, Wound healing assay, Invasion assay, Gelatin zymography, Western blot | Wang et al., 2010 [48] |
Fucoidan | (2S,3S,4S,5S,6R)-4,5-dihydroxy-2,6-dimethyltetrahydro-2H-pyran-3-yl hydrogen sulfate (9072-19-9) | Ascophyllum nodosum | Inhibits cell adhesion to fibronectin | B | Binds to fibronectin with high affinity. Binds to MDA-MB 231 cell membrane and is internalized. Compromises α5 subunit distribution on MDA-MB-231 cell membrane | MDA-MB-231 | Western blot, Flow cytometer, Scatchard plot analysis, Fluorescence microscopy, Immunofluorescence and confocal laser imaging, Lab-tek chamber | Liu et al., 2005 [234] |
Butein | (E)-1-(2,4-dihydroxyphenyl)-3-(3,4-dihydroxyphenyl)prop-2-en-1-one (487-52-5) | Semecarpus anacardium, Dalbergia odorifera,  Caragana jubata,  Rhus verniciflua  | Significantly decreased TNF-α -induced U937 cell adhesion to lung epithelial cells in a dose dependent manner. Anti-inflammatory | P | Inhibition of TNF-α-induced ICAM-1 and VCAM-1 expression by inhibiting the NF- κB/ MAPK/Akt signaling pathway. Inhibited expression of ICAM-1 and VCAM-1 | A549 U937 | Western blot, Fluorescent labeling (BCECF-AM), RT-PCR, Luciferase reporter gene assay, Fluorescence microscopy, Fluocytometer (H2DCFDA) | Jang et al., 2012 [49] |
Tripterygium wilfordii extract (TWH-f), and tetrandrine | (1β)-6,6’,7,12-tetramethoxy-2,2’-dimethylberbaman (518-34-3) | Tripterygium wilfordii | Affects the secretion and expression of adhesion molecules on cells | P | TWH-f at high concentration has significant inhibitory effect on expression and secretion of E-selectin, ICAM-1, VCAM-1. Tetrandrine did not demonstrate the same effects. | Neutrophils HUVEC HFB | Flow cytometry, ELISA | Chang et al., 1999 [7] |
| „Danshen” (DS) | Salvia miltiorrhiza (DS) | SZ and HSW significantly inhibited apoptosis in HUVEC undergoing serum deprivation and TNF-α stimulation | P | Down-regulation of caspase-3-gene expression. DS and SQ significantly attenuated TNF-α-induced expression of VCAM-1 and ICAM-1 | HUVEC | Annexin-V staining, Western blot, Flow cytometry, [3H]-thymidine incorporation assay | Ling et al., 2007 [2] | |
| „Sanchi” (SQ) | Panax notoginseng (SQ) | |||||||
| „Shanzai” (SZ) | Crataegus (SZ) | |||||||
| „Heshouwu”(HSW) | Polygonum multiflorum (HSW) |
| Active Substance | Source | Cellular Effect | Effect Type | Molecular Mechanisms | Tested Cell Line 1, Animal | Method | Ref. |
|---|---|---|---|---|---|---|---|
Rhodostomin Disintegrin  | Calloselasma rhodostoma (venom) | Cell detachment. Actin cytoskeleton perturbed | B, P | Prevention of integrin αvβ3 interaction. Decreased pp125FAK phosphorylation | HUVEC | Cell detachment assay (Trypan blue), MTT assay, Flow cytometry, DNA fragmentation assay, Precipitation of cell extracts and immunoprecipitation, Western blot, Immunofluorescence microscopy | Wu et al., 2002 [55] |
Echistatin | Echis carinatus (venom)  | Decreased cell adhesion | B | Prevention of αIIbβ3, αvβ3 and α5β1 integrin interactions | HeLa | Epic BT (RWG label-free biosensor) | Szekacs et al., 2018 [51] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Péter, B.; Boldizsár, I.; Kovács, G.M.; Erdei, A.; Bajtay, Z.; Vörös, A.; Ramsden, J.J.; Szabó, I.; Bősze, S.; Horvath, R. Natural Compounds as Target Biomolecules in Cellular Adhesion and Migration: From Biomolecular Stimulation to Label-Free Discovery and Bioactivity-Based Isolation. Biomedicines 2021, 9, 1781. https://doi.org/10.3390/biomedicines9121781
Péter B, Boldizsár I, Kovács GM, Erdei A, Bajtay Z, Vörös A, Ramsden JJ, Szabó I, Bősze S, Horvath R. Natural Compounds as Target Biomolecules in Cellular Adhesion and Migration: From Biomolecular Stimulation to Label-Free Discovery and Bioactivity-Based Isolation. Biomedicines. 2021; 9(12):1781. https://doi.org/10.3390/biomedicines9121781
Chicago/Turabian StylePéter, Beatrix, Imre Boldizsár, Gábor M. Kovács, Anna Erdei, Zsuzsa Bajtay, Alexandra Vörös, Jeremy J. Ramsden, Ildikó Szabó, Szilvia Bősze, and Robert Horvath. 2021. "Natural Compounds as Target Biomolecules in Cellular Adhesion and Migration: From Biomolecular Stimulation to Label-Free Discovery and Bioactivity-Based Isolation" Biomedicines 9, no. 12: 1781. https://doi.org/10.3390/biomedicines9121781
APA StylePéter, B., Boldizsár, I., Kovács, G. M., Erdei, A., Bajtay, Z., Vörös, A., Ramsden, J. J., Szabó, I., Bősze, S., & Horvath, R. (2021). Natural Compounds as Target Biomolecules in Cellular Adhesion and Migration: From Biomolecular Stimulation to Label-Free Discovery and Bioactivity-Based Isolation. Biomedicines, 9(12), 1781. https://doi.org/10.3390/biomedicines9121781







