Proteome of Glioblastoma-Derived Exosomes as a Source of Biomarkers
Abstract
1. Introduction
2. Results
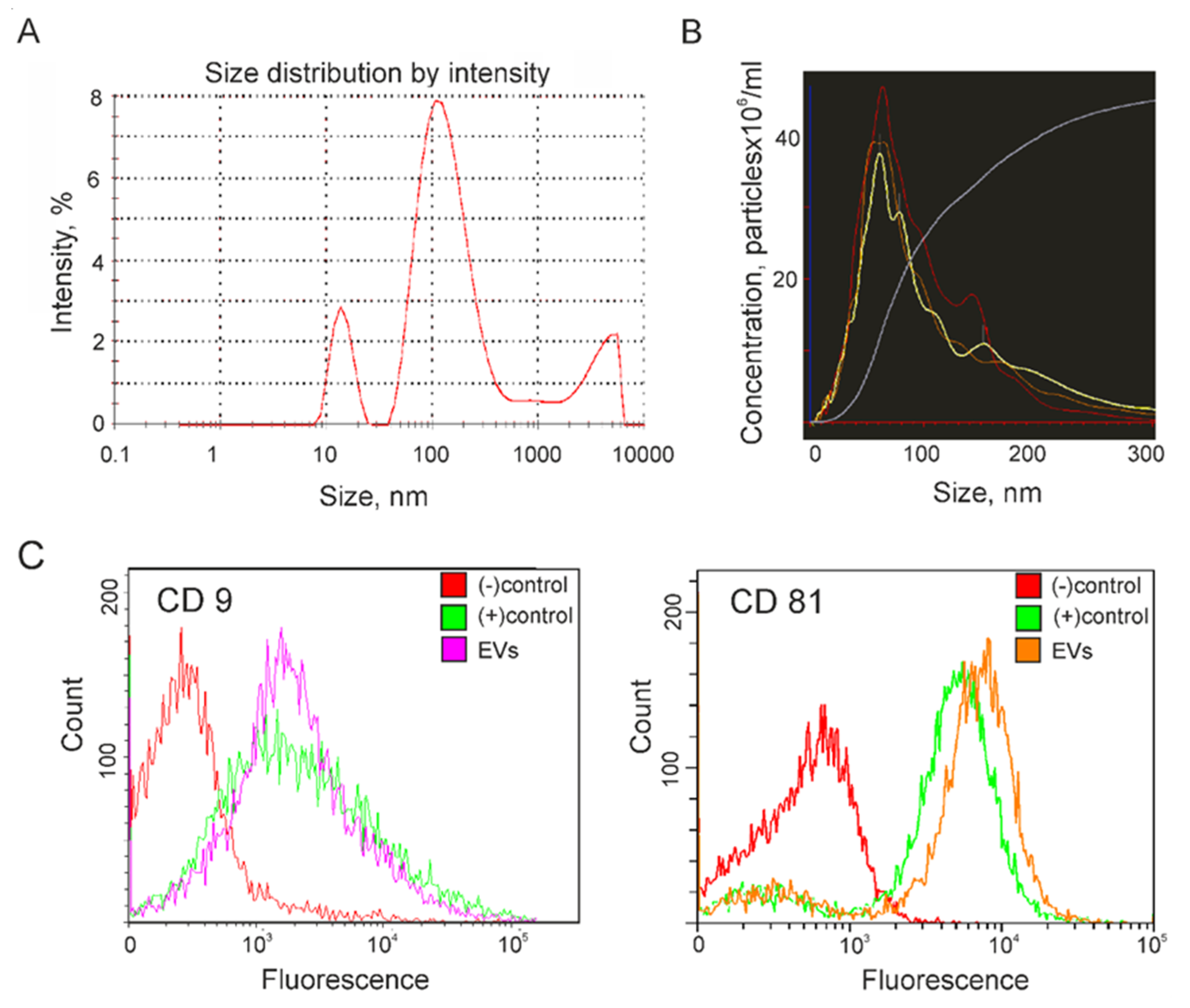
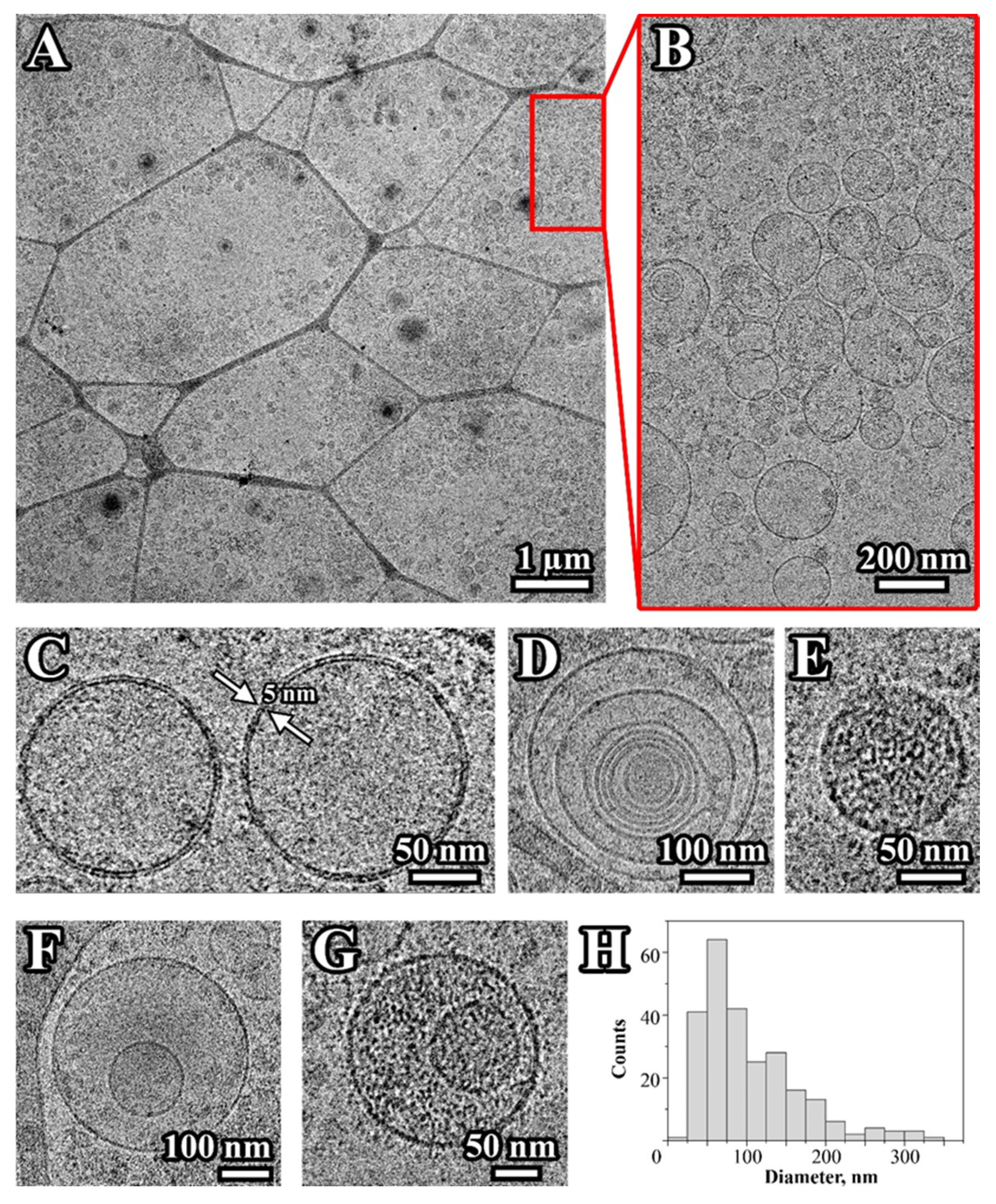
2.1. Characterization of Extracellular Vesicles Isolated from Conditioned Culture Medium of Glioma Cell Lines
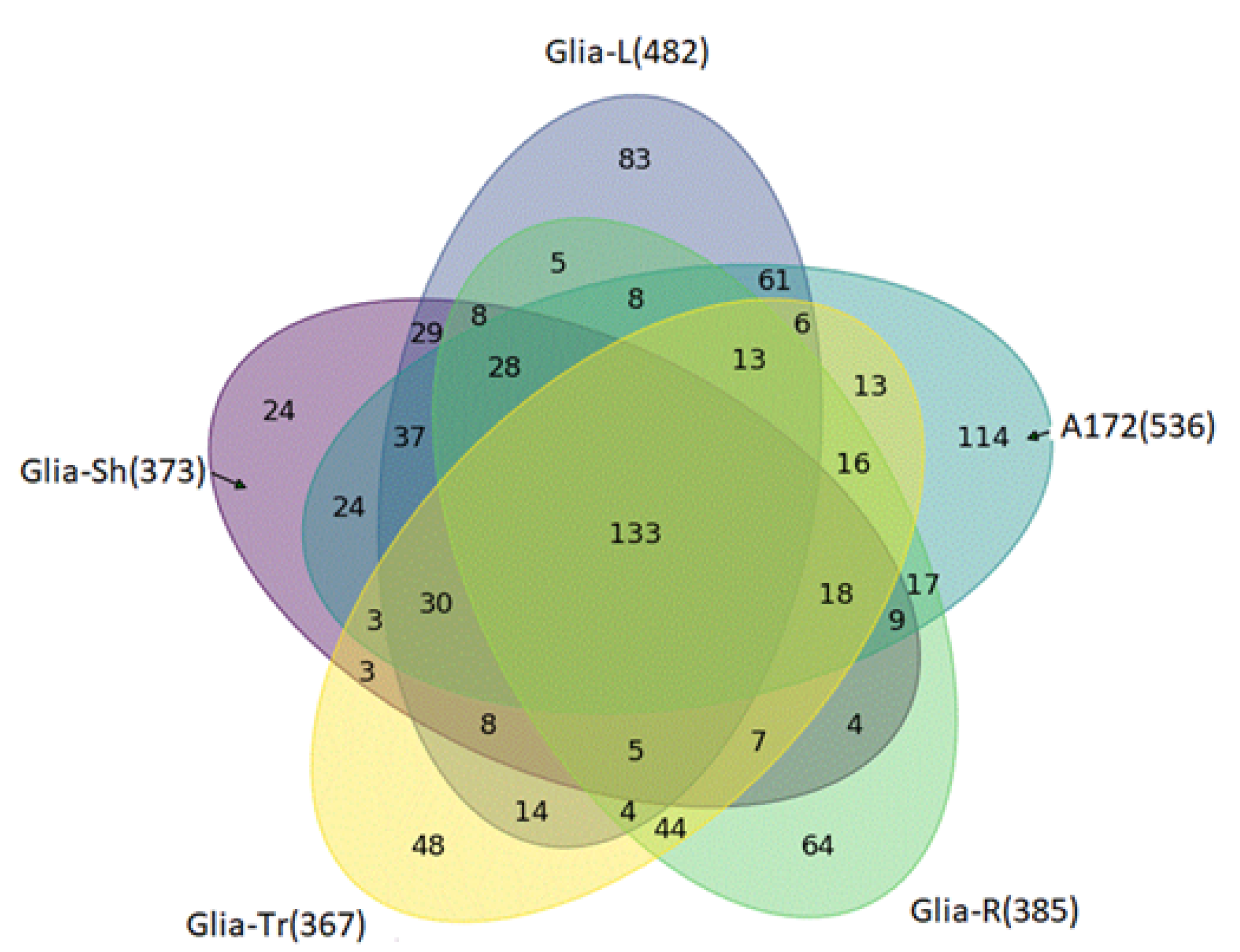
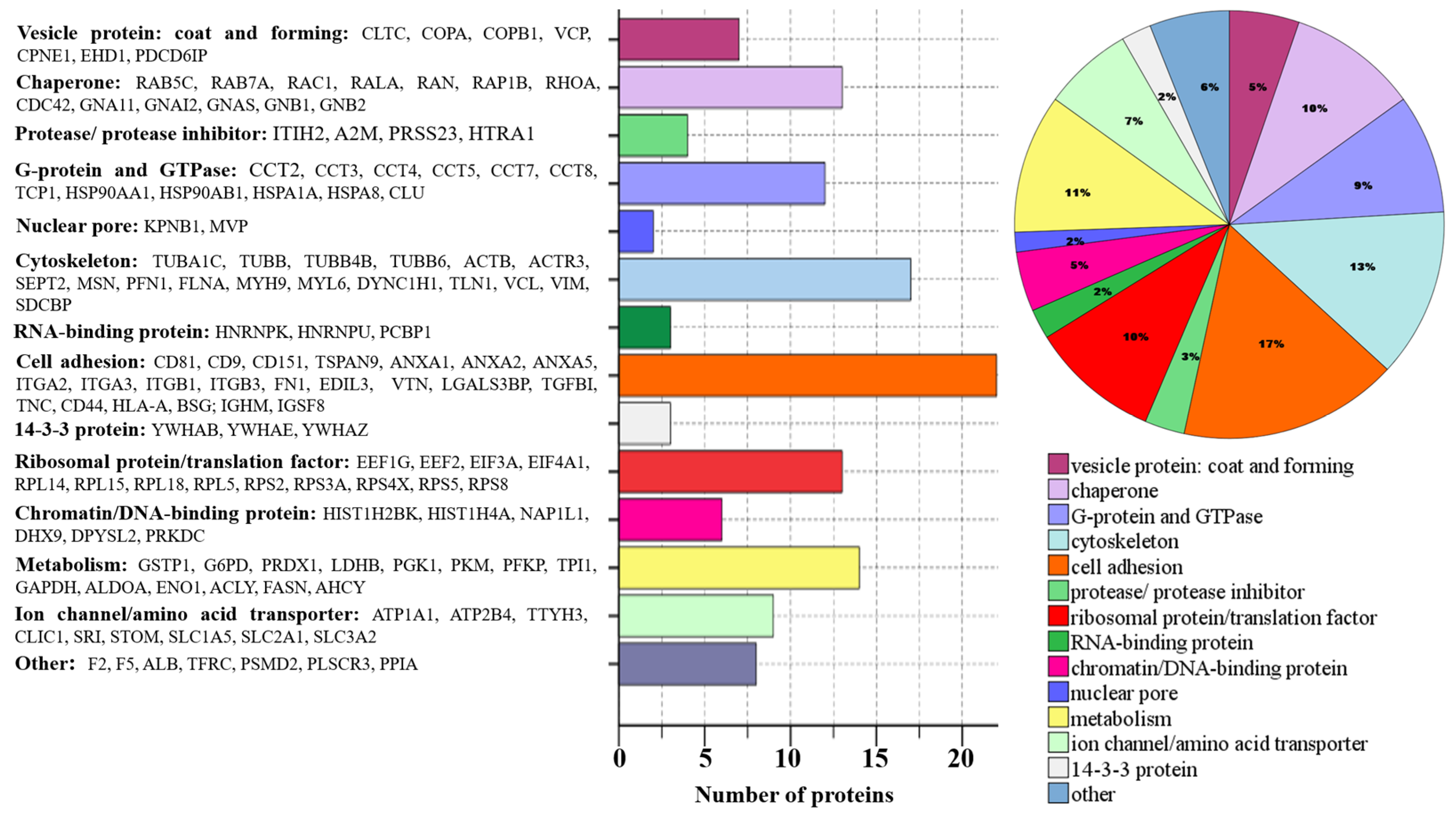
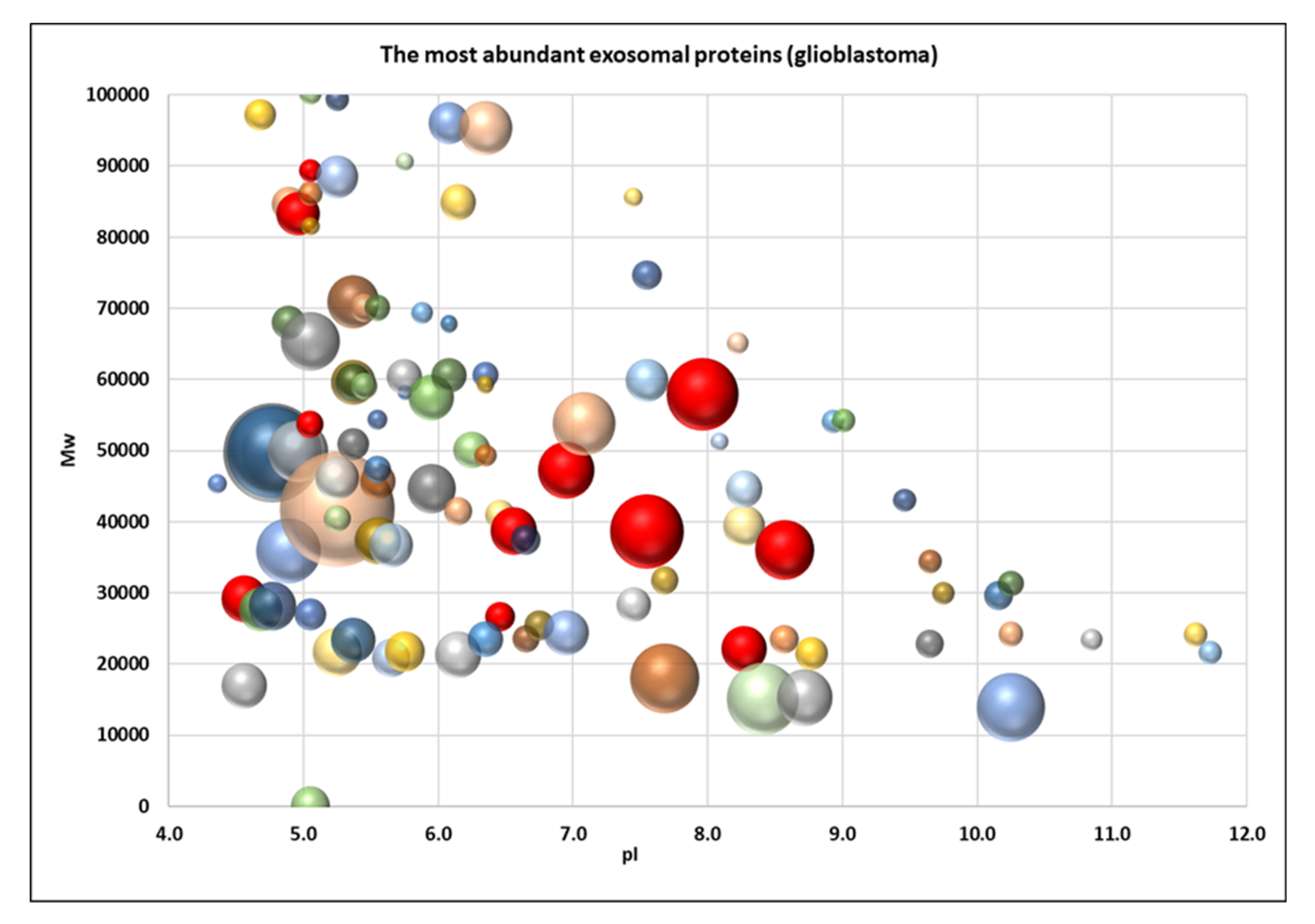
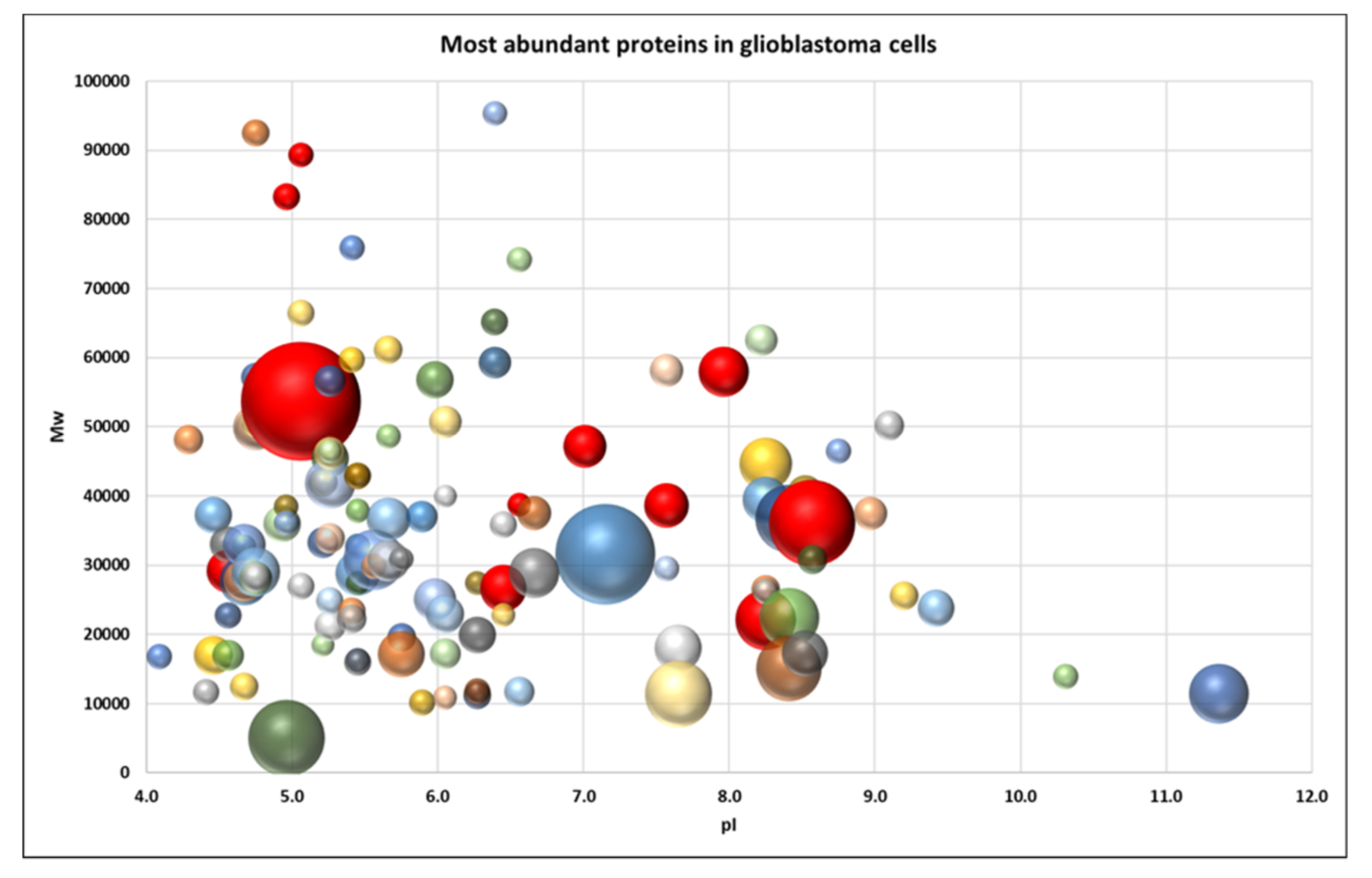
2.2. Proteome Profiling of Glioblastoma-Derived Extracellular Vesicles
3. Discussion
4. Materials and Methods
4.1. Glioma Cell Lines and Cultivation Conditions
4.2. Isolation of Exosomes from the Conditioned Culture Medium (CCM)
4.3. Exosome Analysis
4.4. Mass Spectrometry (MS)
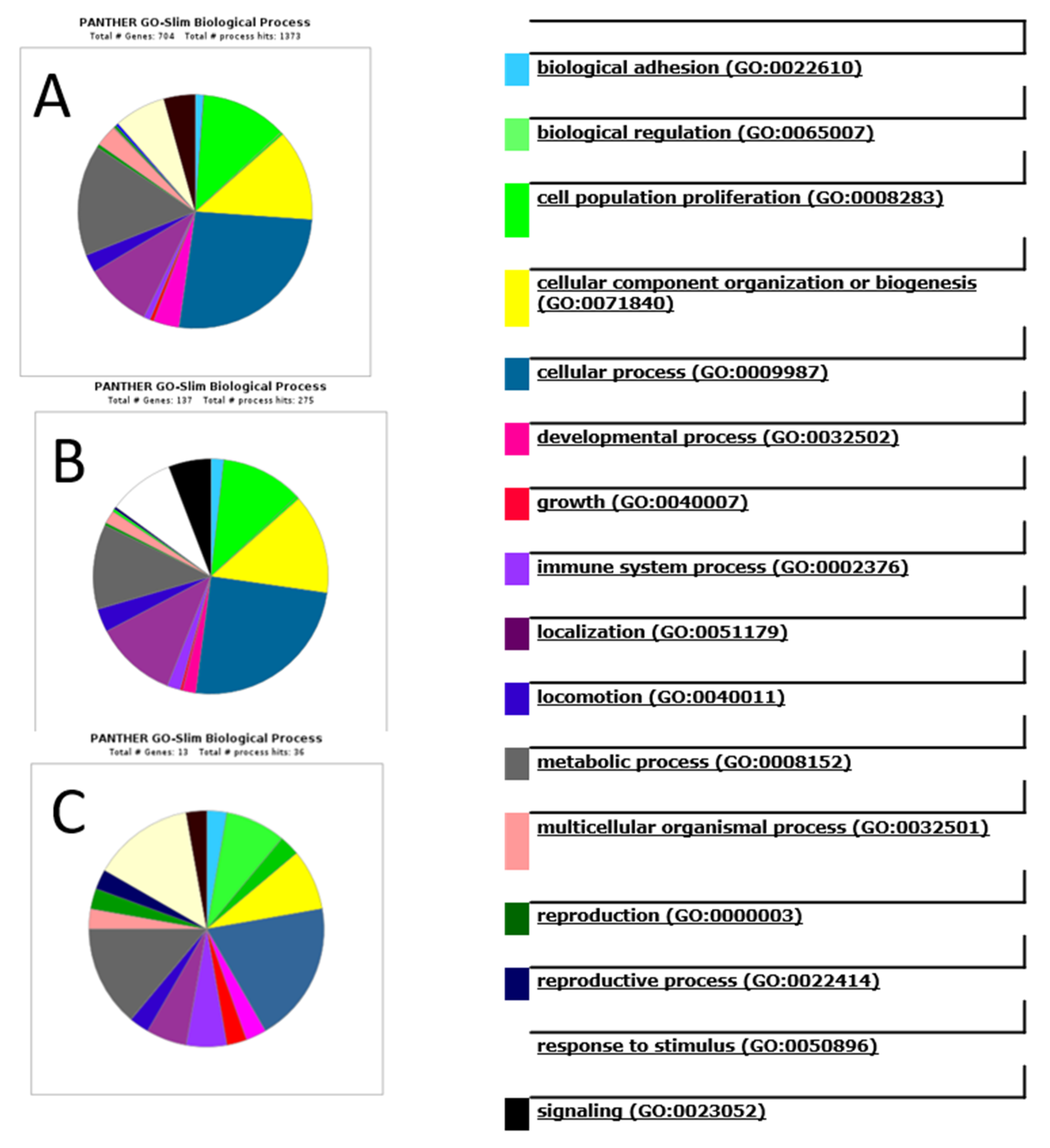
4.5. Gene Ontology Functional Annotation and Pathway Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| EV | Extracellular vesicles |
| MS | Mass spectrometry |
| ESI LC-MS/MS | Liquid chromatography-electrospray ionization-tandem mass spectrometry |
| GBM | The glioblastoma multiforme |
| 2DE | Two-dimensional electrophoresis |
| NTA | Nanoparticle tracking analysis |
| AFM | Atomic force microscopy |
| DLS | Dynamic light scattering |
| CCM | Conditioned culture medium |
| emPAI | Exponential modified form of protein abundance index |
| FASP | Filter-aided sample preparation |
| cryo-EM | Cryo-electron microscopy |
| HPA | Human protein atlas |
References
- Louis, D.N.; Perry, A.; Reifenberger, G.; von Deimling, A.; Figarella-Branger, D.; Cavenee, W.K.; Ohgaki, H.; Wiestler, O.D.; Kleihues, P.; Ellison, D.W. The 2016 World Health Organization Classification of Tumors of the Central Nervous System: A summary. Acta Neuropathol. 2016, 131, 803–820. [Google Scholar] [CrossRef]
- Stupp, R.; Mason, W.P.; van den Bent, M.J.; Weller, M.; Fisher, B.; Taphoorn, M.J.B.; Belanger, K.; Brandes, A.A.; Marosi, C.; Bogdahn, U.; et al. Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N. Engl. J. Med. 2005, 352, 987–996. [Google Scholar] [CrossRef] [PubMed]
- Mallawaaratchy, D.M.; Hallal, S.; Russell, B.; Ly, L.; Ebrahimkhani, S.; Wei, H.; Christopherson, R.I.; Buckland, M.E.; Kaufman, K.L. Comprehensive proteome profiling of glioblastoma-derived extracellular vesicles identifies markers for more aggressive disease. J. Neurooncol. 2017, 131, 233–244. [Google Scholar] [CrossRef]
- Kalluri, R. The biology and function of exosomes in cancer. J. Clin. Invest. 2016, 126, 1208–1215. [Google Scholar] [CrossRef]
- Raposo, G.; Stoorvogel, W. Extracellular vesicles: Exosomes, microvesicles, and friends. J. Cell Biol. 2013, 200, 373–383. [Google Scholar] [CrossRef]
- Théry, C.; Ostrowski, M.; Segura, E. Membrane vesicles as conveyors of immune responses. Nat. Rev. Immunol. 2009, 9, 581–593. [Google Scholar] [CrossRef] [PubMed]
- Skog, J.; Würdinger, T.; van Rijn, S.; Meijer, D.H.; Gainche, L.; Curry, W.T.; Carter, B.S.; Krichevsky, A.M.; Breakefield, X.O. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat. Cell Biol. 2008, 10, 1470–1476. [Google Scholar] [CrossRef] [PubMed]
- Li, C.C.Y.; Eaton, S.A.; Young, P.E.; Lee, M.; Shuttleworth, R.; Humphreys, D.T.; Grau, G.E.; Combes, V.; Bebawy, M.; Gong, J.; et al. Glioma microvesicles carry selectively packaged coding and noncoding RNAs which alter gene expression in recipient cells. RNA Biol. 2013, 10, 1333–1344. [Google Scholar] [CrossRef] [PubMed]
- Barile, L.; Vassalli, G. Exosomes: Therapy delivery tools and biomarkers of diseases. Pharmacol. Ther. 2017, 174, 63–78. [Google Scholar] [CrossRef]
- Graner, M.W.; Alzate, O.; Dechkovskaia, A.M.; Keene, J.D.; Sampson, J.H.; Mitchell, D.A.; Bigner, D.D. Proteomic and immunologic analyses of brain tumor exosomes. FASEB J. 2009, 23, 1541–1557. [Google Scholar] [CrossRef]
- Schey, K.L.; Luther, J.M.; Rose, K.L. Proteomics characterization of exosome cargo. Methods 2015, 87, 75–82. [Google Scholar] [CrossRef] [PubMed]
- Shtam, T.A.; Naryzhny, S.N.; Landa, S.B.; Burdakov, V.S.; Artamonova, T.O.; Filatov, M.V. Purification and in vitro analysis of exosomes secreted by malignantly transformed human cells. Cell Tissue biol. 2012, 6, 317–325. [Google Scholar] [CrossRef]
- Naryzhny, S.N.; Ronzhina, N.L.; Mainskova, M.A.; Belyakova, N.V.; Pantina, R.A.; Filatov, M.V. Development of barcode and proteome profiling of glioblastoma. Biochem. Suppl. Ser. B Biomed. Chem. 2014, 8, 243–251. [Google Scholar]
- Naryzhny, S.; Klopov, N.; Ronzhina, N.; Zorina, E.; Zgoda, V.; Kleyst, O.; Belyakova, N.; Legina, O. A database for inventory of proteoform profiles: “2DE-pattern”. Electrophoresis 2020, 41, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Emelyanov, A.; Shtam, T.; Kamyshinsky, R.; Garaeva, L.; Verlov, N.; Miliukhina, I.; Kudrevatykh, A.; Gavrilov, G.; Zabrodskaya, Y.; Pchelina, S.; et al. Cryo-electron microscopy of extracellular vesicles from cerebrospinal fluid. PLoS ONE 2020, 15, e0227949. [Google Scholar] [CrossRef] [PubMed]
- Höög, J.L.; Lötvall, J. Diversity of extracellular vesicles in human ejaculates revealed by cryo-electron microscopy. J. Extracell. Vesicles 2015, 4, 28680. [Google Scholar] [CrossRef] [PubMed]
- Yuana, Y.; Koning, R.I.; Kuil, M.E.; Rensen, P.C.N.; Koster, A.J.; Bertina, R.M.; Osanto, S. Cryo-electron microscopy of extracellular vesicles in fresh plasma. J. Extracell. Vesicles 2013, 2, 21494. [Google Scholar] [CrossRef]
- Naryzhny, S.N.; Maynskova, M.A.; Zgoda, V.G.; Ronzhina, N.L.; Novikova, S.E.; Belyakova, N.V.; Kleyst, O.A.; Legina, O.K.; Pantina, R.A. FM Proteomic profiling of high-grade glioblastoma using virtual-experimental 2DE. J. Proteom. Bioinform. 2016, 9, 158–165. [Google Scholar] [CrossRef]
- Burdakov, V.; Shtam, T.; Naryzhny, S.; Landa, S.; Ronzina, N.; Filatov, M. Proteomic analysis of exosomes secreted by human glioblastoma cells hold promise for identifying markers of brain cancer: SW06. S25–22. FEBS J. 2013, 280, 3–617. [Google Scholar]
- Salomon, C.; Ryan, J.; Sobrevia, L.; Kobayashi, M.; Ashman, K.; Mitchell, M.; Rice, G.E. Exosomal signaling during hypoxia mediates microvascular endothelial cell migration and vasculogenesis. PLoS ONE 2013, 8, e68451. [Google Scholar] [CrossRef]
- Indira Chandran, V.; Welinder, C.; Gonçalves de Oliveira, K.; Cerezo-Magaña, M.; Månsson, A.S.; Johansson, M.C.; Marko-Varga, G.; Belting, M. Global extracellular vesicle proteomic signature defines U87-MG glioma cell hypoxic status with potential implications for non-invasive diagnostics. J. Neurooncol. 2019, 144, 477–488. [Google Scholar] [CrossRef]
- Jayaseelan, V.P. Emerging role of exosomes as promising diagnostic tool for cancer. Cancer Gene Ther. 2019, 27, 395–398. [Google Scholar] [CrossRef]
- Domenis, R.; Cesselli, D.; Toffoletto, B.; Bourkoula, E.; Caponnetto, F.; Manini, I.; Beltrami, A.P.; Ius, T.; Skrap, M.; di Loreto, C.; et al. Systemic T cells immunosuppression of glioma stem cell-derived exosomes is mediated by monocytic myeloid-derived suppressor cells. PLoS ONE 2017, 12, e0169932. [Google Scholar] [CrossRef]
- Luhtala, N.; Aslanian, A.; Yates, J.R.; Hunter, T. Secreted glioblastoma nanovesicles contain intracellular signaling proteins and active ras incorporated in a farnesylation-dependent manner. J. Biol. Chem. 2017, 292, 611–628. [Google Scholar] [CrossRef]
- Ma, Z.; Cui, X.; Lu, L.; Chen, G.; Yang, Y.; Hu, Y.; Lu, Y.; Cao, Z.; Wang, Y.; Wang, X. Exosomes from glioma cells induce a tumor-like phenotype in mesenchymal stem cells by activating glycolysis. Stem. Cell Res. Ther. 2019, 10. [Google Scholar] [CrossRef]
- Hochberg, F.H.; Atai, N.A.; Gonda, D.; Hughes, M.S.; Mawejje, B.; Balaj, L.; Carter, R.S. Glioma diagnostics and biomarkers: An ongoing challenge in the field of medicine and science. Expert Rev. Mol. Diagn. 2014, 14, 439–452. [Google Scholar] [CrossRef]
- Théry, C.; Witwer, K.W.; Aikawa, E.; Alcaraz, M.J.; Anderson, J.D.; Andriantsitohaina, R.; Antoniou, A.; Arab, T.; Archer, F.; Atkin-Smith, G.K.; et al. Minimal information for studies of extracellular vesicles 2018 (MISEV2018): A position statement of the International Society for Extracellular Vesicles and update of the MISEV2014 guidelines. J. Extracell. Vesicles 2018, 7. [Google Scholar] [CrossRef]
- Liu, C.Y.; Lin, H.H.; Tang, M.J.; Wang, Y.K. Vimentin contributes to epithelial-mesenchymal transition ancer cell mechanics by mediating cytoskeletal organization and focal adhesion maturation. Oncotarget 2015, 6, 15966–15983. [Google Scholar] [CrossRef]
- Baietti, M.F.; Zhang, Z.; Mortier, E.; Melchior, A.; Degeest, G.; Geeraerts, A.; Ivarsson, Y.; Depoortere, F.; Coomans, C.; Vermeiren, E.; et al. Syndecan-syntenin-ALIX regulates the biogenesis of exosomes. Nat. Cell Biol. 2012, 14, 677–685. [Google Scholar] [CrossRef]
- Chatellard-Causse, C.; Blot, B.; Cristina, N.; Torch, S.; Missotten, M.; Sadoul, R. Alix (ALG-2-interacting protein X), a protein involved in apoptosis, binds to endophilins and induces cytoplasmic vacuolization. J. Biol. Chem. 2002, 277, 29108–29115. [Google Scholar] [CrossRef]
- Li, Z.; Mbah, N.E.; Maltese, W.A. Vacuole-inducing compounds that disrupt endolysosomal trafficking stimulate production of exosomes by glioblastoma cells. Mol. Cell. Biochem. 2018, 439, 1–9. [Google Scholar] [CrossRef]
- Zhai, H.; Acharya, S.; Gravanis, I.; Mehmood, S.; Seidman, R.J.; Shroyer, K.R.; Hajjar, K.A.; Tsirka, S.E. Annexin A2 promotes glioma cell invasion and tumor progression. J. Neurosci. 2011, 31, 14346–14360. [Google Scholar] [CrossRef] [PubMed]
- Pietras, A.; Katz, A.M.; Ekström, E.J.; Wee, B.; Halliday, J.J.; Pitter, K.L.; Werbeck, J.L.; Amankulor, N.M.; Huse, J.T.; Holland, E.C. Osteopontin-CD44 signaling in the glioma perivascular niche enhances cancer stem cell phenotypes and promotes aggressive tumor growth. Cell Stem Cell 2014, 14, 357–369. [Google Scholar] [CrossRef]
- Leins, A.; Riva, P.; Lindstedt, R.; Davidoff, M.S.; Mehraein, P.; Weis, S. Expression of tenascin-C in various human brain tumors and its relevance for survival in patients with astrocytoma. Cancer 2003, 98, 2430–2439. [Google Scholar] [CrossRef] [PubMed]
- Volnitskiy, A.V.; Semenova, E.V.; Shtam, T.A.; Kovalev, R.A.; Filatov, M.V. Aberrant expression of the sox2 gene in malignant gliomas. Cell Tissue Biol. 2014, 8, 368–373. [Google Scholar] [CrossRef]
- Volnitskiy, A.; Shtam, T.; Burdakov, V.; Kovalev, R.; Konev, A.; Filatov, M. Abnormal activity of transcription factors gli in high-grade gliomas. PLoS ONE 2019, 14, e0211980. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Jiang, Y.; Meisenhelder, J.; Yang, W.; Hawke, D.H.; Zheng, Y.; Xia, Y.; Aldape, K.; He, J.; Hunter, T.; et al. Mitochondria-translocated PGK1 functions as a protein kinase to coordinate glycolysis and the TCA cycle in tumorigenesis. Mol. Cell 2016, 61, 705–719. [Google Scholar] [CrossRef]
- Daido, S.; Yamamoto, A.; Fujiwara, K.; Sawaya, R.; Kondo, S.; Kondo, Y. Inhibition of the DNA-dependent protein kinase catalytic subunit radiosensitizes malignant glioma cells by inducing autophagy. Cancer Res. 2005, 65, 4368–4375. [Google Scholar] [CrossRef]
- Jiang, N.; Shen, Y.; Fei, X.; Sheng, K.; Sun, P.; Qiu, Y.; Larner, J.; Cao, L.; Kong, X.; Mi, J. Valosin-containing protein regulates the proteasome-mediated degradation of DNA-PKcs in glioma cells. Cell Death Dis. 2013, 4. [Google Scholar] [CrossRef]
- Lara, P.C.; Pruschy, M.; Zimmermann, M.; Henríquez-Hernández, L.A. MVP and vaults: A role in the radiation response. Radiat. Oncol. 2011, 6, 149. [Google Scholar] [CrossRef]
- Stavrovskaya, A.A.; Shushanov, S.S.; Rybalkina, E.Y. Problems of glioblastoma multiforme drug resistance. Biochemistry 2016, 81, 91–100. [Google Scholar] [CrossRef] [PubMed]
- Cui, D.; Xu, Q.; Wang, K.; Che, X. Gli1 is a potential target for alleviating multidrug resistance of gliomas. J. Neurol. Sci. 2010, 288, 156–166. [Google Scholar] [CrossRef]
- Théry, C.; Amigorena, S.; Raposo, G.; Clayton, A. Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Curr. Protoc. Cell Biol. 2006, 30, 3–22. [Google Scholar] [CrossRef]
- Shtam, T.A.; Samsonov, R.B.; Volnitskiy, A.V.; Kamyshinsky, R.A.; Verlov, N.A.; Kniazeva, M.S.; Korobkina, E.A.; Orehov, A.S.; Vasiliev, A.L.; Konevega, A.L.; et al. Isolation of extracellular microvesicles from cell culture medium: Comparative evaluation of methods. Biochem. Suppl. Ser. B Biomed. Chem. 2018, 12, 167–175. [Google Scholar] [CrossRef]
- Nečas, D.; Klapetek, P. Gwyddion: An open-source software for SPM data analysis. Cent. Eur. J. Phys. 2012, 10, 181–188. [Google Scholar] [CrossRef]
- Naryzhny, S.; Zgoda, V.; Kopylov, A.; Petrenko, E.; Kleist, O.; Archakov, A. Variety and dynamics of proteoforms in the human proteome: Aspects of markers for hepatocellular carcinoma. Proteomes 2017, 5, 33. [Google Scholar] [CrossRef]
- Naryzhny, S.N.; Zgoda, V.G.; Maynskova, M.A.; Novikova, S.E.; Ronzhina, N.L.; Vakhrushev, I.V.; Khryapova, E.V.; Lisitsa, A.V.; Tikhonova, O.V.; Ponomarenko, E.A.; et al. Combination of virtual and experimental 2DE together with ESI LC-MS/MS gives a clearer view about proteomes of human cells and plasma. Electrophoresis 2016, 37, 302–309. [Google Scholar] [CrossRef]
- Naryzhny, S.; Maynskova, M.; Zgoda, V.; Archakov, A. Zipf’s law in proteomics. J. Proteomics Bioinform. 2017, 10, 79–84. [Google Scholar] [CrossRef]
- Vaudel, M.; Barsnes, H.; Berven, F.S.; Sickmann, A.; Martens, L. SearchGUI: An open-source graphical user interface for simultaneous OMSSA and X!Tandem searches. Proteomics 2011, 11, 996–999. [Google Scholar] [CrossRef]
- Ge, S.X.; Jung, D. ShinyGO: A graphical enrichment tool for animals and plants. bioRxiv 2018, 2, 315150. [Google Scholar]
- Huang, D.W.; Sherman, B.T.; Tan, Q.; Collins, J.R.; Alvord, W.G.; Roayaei, J.; Stephens, R.; Baseler, M.W.; Lane, H.C.; Lempicki, R.A. The DAVID gene functional classification tool: A novel biological module-centric algorithm to functionally analyze large gene lists. Genome Biol. 2007, 8, R183. [Google Scholar] [CrossRef]
- Nakaya, A.; Katayama, T.; Itoh, M.; Hiranuka, K.; Kawashima, S.; Moriya, Y.; Okuda, S.; Tanaka, M.; Tokimatsu, T.; Yamanishi, Y.; et al. KEGG OC: A large-scale automatic construction of taxonomy-based ortholog clusters. Nucleic Acids Res. 2013, 41, D1. [Google Scholar] [CrossRef] [PubMed]
- Schaefer, C.F.; Anthony, K.; Krupa, S.; Buchoff, J.; Day, M.; Hannay, T.; Buetow, K.H. PID: The pathway interaction database. Nucleic Acids Res. 2009, 37, D674–D679. [Google Scholar] [CrossRef] [PubMed]
- Fabregat, A.; Jupe, S.; Matthews, L.; Sidiropoulos, K.; Gillespie, M.; Garapati, P.; Haw, R.; Jassal, B.; Korninger, F.; May, B.; et al. The reactome pathway knowledgebase. Nucleic Acids Res. 2018, 46, D649–D655. [Google Scholar] [CrossRef]

© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Naryzhny, S.; Volnitskiy, A.; Kopylov, A.; Zorina, E.; Kamyshinsky, R.; Bairamukov, V.; Garaeva, L.; Shlikht, A.; Shtam, T. Proteome of Glioblastoma-Derived Exosomes as a Source of Biomarkers. Biomedicines 2020, 8, 216. https://doi.org/10.3390/biomedicines8070216
Naryzhny S, Volnitskiy A, Kopylov A, Zorina E, Kamyshinsky R, Bairamukov V, Garaeva L, Shlikht A, Shtam T. Proteome of Glioblastoma-Derived Exosomes as a Source of Biomarkers. Biomedicines. 2020; 8(7):216. https://doi.org/10.3390/biomedicines8070216
Chicago/Turabian StyleNaryzhny, Stanislav, Andrey Volnitskiy, Arthur Kopylov, Elena Zorina, Roman Kamyshinsky, Viktor Bairamukov, Luiza Garaeva, Anatoly Shlikht, and Tatiana Shtam. 2020. "Proteome of Glioblastoma-Derived Exosomes as a Source of Biomarkers" Biomedicines 8, no. 7: 216. https://doi.org/10.3390/biomedicines8070216
APA StyleNaryzhny, S., Volnitskiy, A., Kopylov, A., Zorina, E., Kamyshinsky, R., Bairamukov, V., Garaeva, L., Shlikht, A., & Shtam, T. (2020). Proteome of Glioblastoma-Derived Exosomes as a Source of Biomarkers. Biomedicines, 8(7), 216. https://doi.org/10.3390/biomedicines8070216