Genotype-Specific Distribution of High-Risk Human Papillomavirus (HPV) and Microbial Co-Detections in HPV-Positive Women from Southern Croatia
Abstract
1. Introduction
2. Materials and Methods
Data Analysis
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| AIC | Akaike Information Criterion |
| CE-IVD | Conformité Européenne—In Vitro Diagnostic |
| CIN 2 | Cervical Intraepithelial Neoplasia Grade 2 |
| CIN 3 | Cervical Intraepithelial Neoplasia Grade 3 |
| CT | Chlamydia trachomatis |
| DALY | Disability-Adjusted Life Year |
| GV | Gardnerella vaginalis |
| HPV | Human Papillomavirus |
| HR | High Risk |
| IQR | Interquartile Range |
| MG | Mycoplasma genitalium |
| MH | Mycoplasma hominis |
| NG | Neisseria gonorrhoeae |
| OR | Odds Ratio |
| Pap | Papanicolaou |
| PCR | Polymerase Chain Reaction |
| SE | Standard Error |
| STI | Sexually Transmitted Infection |
| UU | Ureaplasma urealyticum |
| TV | Trichomonas vaginalis |
References
- Ghosh, S.K.; Choudhury, B.; Hansa, J.; Mondal, R.; Singh, M.; Singh, M.; Duttagupta, S.; Das, A.; Kumar, R.; Laskar, R.S.; et al. Human papillomavirus testing for suspected cervical cancer patients from Southern Assam by fast-PCR. Asian Pac. J. Cancer Prev. 2011, 12, 749–751. [Google Scholar]
- Okunade, K.S. Human papillomavirus and cervical cancer. J. Obstet. Gynaecol. 2020, 40, 602–608. [Google Scholar] [CrossRef]
- Naghavi, M.; Meštrović, T.; Gray, A.; Gershberg Hayoon, A.; Swetschinski, L.R.; Robles Aguilar, G.; Davis Weaver, N.; Ikuta, K.S.; Chung, E.; Wool, E.E.; et al. Global burden associated with 85 pathogens in 2019: A systematic analysis for the Global Burden of Disease Study 2019. Lancet Infect. Dis. 2024, 24, 868–895. [Google Scholar] [CrossRef] [PubMed]
- Su, P.; Ma, J.; Yu, L.; Tang, S.; Sun, P. Clinical significance of extended high-risk human papillomavirus genotyping and viral load in cervical cancer and precancerous lesions. Gynaecol. Obstet. Clin. Med. 2023, 3, 22–29. [Google Scholar] [CrossRef]
- Baba, S.K.; Alblooshi, S.S.E.; Yaqoob, R.; Behl, S.; Al Saleem, M.; Rakha, E.A.; Malik, F.; Singh, M.; Macha, M.A.; Akhtar, M.K.; et al. Human papilloma virus (HPV) mediated cancers: An insightful update. J. Transl. Med. 2025, 23, 483. [Google Scholar] [CrossRef] [PubMed]
- Van Den Borst, E.; Bell, M.; Op De Beeck, K.; Van Camp, G.; Van Keer, S.; Vorsters, A. Lineages and sublineages of high-risk HPV types associated with cervical cancer and precancer: A systematic review and meta-analysis. J. Natl. Cancer Inst. 2025; ahead of print. [Google Scholar] [CrossRef]
- Zang, L.; Hu, Y. Risk factors associated with HPV persistence after conization in high-grade squamous intraepithelial lesion. Arch. Gynecol. Obstet. 2021, 304, 1409–1416. [Google Scholar] [CrossRef]
- Ntuli, L.; Mtshali, A.; Mzobe, G.; Liebenberg, L.J.; Ngcapu, S. Role of Immunity and Vaginal Microbiome in Clearance and Persistence of Human Papillomavirus Infection. Front. Cell. Infect. Microbiol. 2022, 12, 927131. [Google Scholar] [CrossRef] [PubMed]
- Lin, S.; Gao, K.; Gu, S.; You, L.; Qian, S.; Tang, M.; Wang, J.; Chen, K.; Jin, M. Worldwide trends in cervical cancer incidence and mortality, with predictions for the next 15 years. Cancer 2021, 127, 4030–4039. [Google Scholar] [CrossRef]
- Du, J.; Ahrlund-Richter, A.; Nasman, A.; Dalianis, T. Human papilloma virus (HPV) prevalence upon HPV vaccination in Swedish youth: A review based on our findings 2008–2018, and perspectives on cancer prevention. Arch. Gynecol. Obstet. 2021, 303, 329–335. [Google Scholar] [CrossRef] [PubMed]
- Oyervides-Munoz, M.A.; Perez-Maya, A.A.; Sanchez-Dominguez, C.N.; Berlanga-Garza, A.; Antonio-Macedo, M.; Valdez-Chapa, L.D.; Cerda-Flores, R.M.; Trevino, V.; Barrera-Saldana, H.A.; Garza-Rodriguez, M.L. Multiple HPV Infections and Viral Load Association in Persistent Cervical Lesions in Mexican Women. Viruses 2020, 12, 380. [Google Scholar] [CrossRef]
- Tao, X.; Zhang, H.; Zhang, H.; Xiao, Y.; Zhong, F.; Zhou, X.; Cong, Q.; Sui, L.; Zhao, C. The clinical utility of extended high-risk HPV genotyping in risk-stratifying women with L-SIL cytology: A retrospective study of 8726 cases. Cancer Cytopathol. 2022, 130, 542–550. [Google Scholar] [CrossRef]
- De Marco, L.; Bisanzi, S.; Ronco, G.; Mancuso, P.; Carozzi, F.; Allia, E.; Rizzolo, R.; Gustinucci, D.; Frayle, H.; Viti, J.; et al. Extended HPV genotyping by the BD Onclarity assay: Concordance with screening HPV-DNA assays, triage biomarkers, and histopathology in women from the NTCC2 study. Microbiol. Spectr. 2025, 13, e0089724. [Google Scholar] [CrossRef] [PubMed]
- Ye, Y.; Jones, T.; Wang, T.; Zeng, X.; Liu, Y.; Zhao, C. Comprehensive overview of genotype distribution and prevalence of human papillomavirus in cervical lesions. Gynecol. Obstet. Clin. Med. 2024, 4, e000005. [Google Scholar] [CrossRef]
- Lei, J.; Cuschieri, K.; Patel, H.; Lawrence, A.; Deats, K.; Sasieni, P.; Lim, A.W.W. Human papillomavirus genotype and cycle threshold value from self-samples and risk of high-grade cervical lesions: A post hoc analysis of a modified stepped-wedge implementation feasibility trial. PLoS Med. 2024, 21, e1004494. [Google Scholar] [CrossRef]
- Massad, L.S.; Clarke, M.A.; Perkins, R.B.; Garcia, F.; Chelmow, D.; Cheung, L.C.; Darragh, T.M.; Egemen, D.; Lorey, T.S.; Nayar, R.; et al. Applying Results of Extended Genotyping to Management of Positive Cervicovaginal Human Papillomavirus Test Results: Enduring Guidelines. J. Low. Genit. Tract. Dis. 2025, 29, 134–143. [Google Scholar] [CrossRef]
- Anton, L.; Ferguson, B.; Friedman, E.S.; Gerson, K.D.; Brown, A.G.; Elovitz, M.A. Gardnerella vaginalis alters cervicovaginal epithelial cell function through microbe-specific immune responses. Microbiome 2022, 10, 119. [Google Scholar] [CrossRef]
- Gargiulo Isacco, C.; Balzanelli, M.G.; Garzone, S.; Lorusso, M.; Inchingolo, F.; Nguyen, K.C.D.; Santacroce, L.; Mosca, A.; Del Prete, R. Alterations of Vaginal Microbiota and Chlamydia trachomatis as Crucial Co-Causative Factors in Cervical Cancer Genesis Procured by HPV. Microorganisms 2023, 11, 662. [Google Scholar] [CrossRef]
- Cui, M.; Wu, Y.; Liu, Z.; Liu, Y.; Fan, L. Advances in the interrelated nature of vaginal microecology, HPV infection, and cervical lesions. Front. Cell. Infect. Microbiol. 2025, 15, 1608195. [Google Scholar] [CrossRef]
- Kaliterna, V.; Kaliterna, P.; Pejkovic, L.; Vulic, R.; Zanchi, L.; Cerskov, K. Prevalence of Human Papillomavirus (HPV) among Females in the General Population of the Split and Dalmatia County and Its Association with Genital Microbiota and Infections: A Prospective Study. Viruses 2023, 15, 443. [Google Scholar] [CrossRef] [PubMed]
- Demarco, M.; Hyun, N.; Carter-Pokras, O.; Raine-Bennett, T.R.; Cheung, L.; Chen, X.; Hammer, A.; Campos, N.; Kinney, W.; Gage, J.C.; et al. A study of type-specific HPV natural history and implications for contemporary cervical cancer screening programs. EClinicalMedicine 2020, 22, 100293. [Google Scholar] [CrossRef] [PubMed]
- de Sanjosé, S.; Serrano, B.; Tous, S.; Alejo, M.; Lloveras, B.; Quirós, B.; Clavero, O.; Vidal, A.; Ferrándiz-Pulido, C.; Pavón, M.Á.; et al. Burden of Human Papillomavirus (HPV)-Related Cancers Attributable to HPVs 6/11/16/18/31/33/45/52 and 58. JNCI Cancer Spectr. 2019, 2, pky045. [Google Scholar] [CrossRef]
- Bouvard, V.; Wentzensen, N.; Mackie, A.; Berkhof, J.; Brotherton, J.; Giorgi-Rossi, P.; Kupets, R.; Smith, R.; Arrossi, S.; Bendahhou, K.; et al. The IARC Perspective on Cervical Cancer Screening. N. Engl. J. Med. 2021, 385, 1908–1918. [Google Scholar] [CrossRef] [PubMed]
- Suba, E.J.; Ortega, R.E.; Mutch, D.G. The IARC Perspective on Cervical Cancer Screening. N. Engl. J. Med. 2022, 386, 607. [Google Scholar]
- R Core Team. R: A language and environment for statistical computing. In R Foundation for Statistical Computing; R Core Team: Vienna, Austria, 2020; Available online: https://www.R-project.org/ (accessed on 15 August 2025).
- The jamovi Project. jamovi (Version 2.6) [Computer Software]. 2025. Available online: https://www.jamovi.org (accessed on 15 August 2025).
- WHO/ICO Information Centre on HPV and Cervical Cancer. HPV and cervical cancer in the 2007 report. Vaccine 2007, 25 (Suppl. S3), C1–C230. [Google Scholar]
- Poljak, M.; Seme, K.; Maver, P.J.; Kocjan, B.J.; Cuschieri, K.S.; Rogovskaya, S.I.; Arbyn, M.; Syrjänen, S. Human Papillomavirus Prevalence and Type-Distribution, Cervical Cancer Screening Practices and Current Status of Vaccination Implementation in Central and Eastern Europe. Vaccine 2013, 31, H59–H70. [Google Scholar] [CrossRef] [PubMed]
- Tsakogiannis, D.; Zografos, E.; Tzioga, L.; Zografos, C.G.; Zagouri, F.; Bletsa, G. Prevalence and Genotype Distribution of High-Risk HPV Genotypes Among Women in Greece: A Retrospective Analysis of 3500 Women. Cancers 2025, 17, 1267. [Google Scholar] [CrossRef] [PubMed]
- Kang, L.N.; Castle, P.E.; Zhao, F.H.; Jeronimo, J.; Chen, F.; Bansil, P.; Li, J.; Chen, W.; Zhang, X.; Qiao, Y.L. A prospective study of age trends of high-risk human papillomavirus infection in rural China. BMC Infect. Dis. 2014, 14, 96. [Google Scholar] [CrossRef]
- Burger, E.A.; Kim, J.J.; Sy, S.; Castle, P.E. Age of Acquiring Causal Human Papillomavirus (HPV) Infections: Leveraging Simulation Models to Explore the Natural History of HPV-induced Cervical Cancer. Clin. Infect. Dis. 2017, 65, 893–899. [Google Scholar] [CrossRef]
- Liu, S.; Mei, B.; Ouyang, Y.; Li, C. Prevalence and genotype distribution of human papillomavirus infection among women in Jingzhou, China: A population-based study of 51,720 women. Virol. J. 2023, 20, 297. [Google Scholar] [CrossRef]
- Hu, J.P.; Wang, J.L.; Li, Y.; Feng, Y.; Tian, C.Q.; Zhang, G.H.; Chen, X.Q.; Liu, H.X.; Yang, J.S.; Fang, Z.W.; et al. Prevalence and genotype distribution of human papillomavirus infection among 66000 women from 2014 to 2023 in the plateau region of Southwest China. Virol. J. 2024, 21, 176. [Google Scholar] [CrossRef]
- Berza, N.; Zodzika, J.; Kivite-Urtane, A.; Baltzer, N.; Curkste, A.; Pole, I.; Nygard, M.; Parna, K.; Stankunas, M.; Tisler, A.; et al. Understanding the high-risk human papillomavirus prevalence and associated factors in the European country with a high incidence of cervical cancer. Eur. J. Public Health 2024, 34, 826–832. [Google Scholar] [CrossRef]
- Clarke, M.A.; Risley, C.; Stewart, M.W.; Geisinger, K.R.; Hiser, L.M.; Morgan, J.C.; Owens, K.J.; Ayyalasomayajula, K.; Rives, R.M.; Jannela, A.; et al. Age-specific prevalence of human papillomavirus and abnormal cytology at baseline in a diverse statewide prospective cohort of individuals undergoing cervical cancer screening in Mississippi. Cancer Med. 2021, 10, 8641–8650. [Google Scholar] [CrossRef]
- Wang, Y.; Hou, H.; Dong, G.; Zhang, H.; Zhang, X.; Zhou, Y.; Xue, M.; Wang, Z.; Geng, J.; Liu, L. Age-related distribution of human papillomavirus genotypes in women with cervical squamous cell carcinoma from Linyi, China, 2015–2023. Virol. J. 2025, 22, 157. [Google Scholar] [CrossRef]
- Koshiol, J.; Lindsay, L.; Pimenta, J.M.; Poole, C.; Jenkins, D.; Smith, J.S. Persistent human papillomavirus infection and cervical neoplasia: A systematic review and meta-analysis. Am. J. Epidemiol. 2008, 168, 123–137. [Google Scholar] [CrossRef] [PubMed]
- Bruno, M.T.; Valenti, G.; Ruggeri, Z.; Incognito, G.G.; Coretti, P.; Montana, G.D.; Panella, M.M.; Mereu, L. Correlation of the HPV 16 Genotype Persistence in Women Undergoing LEEP for CIN3 with the Risk of CIN2+ Relapses in the First 18 Months of Follow-Up: A Multicenter Retrospective Study. Diagnostics 2024, 14, 509. [Google Scholar] [CrossRef] [PubMed]
- Agwu, A. Sexuality, Sexual Health, and Sexually Transmitted Infections in Adolescents and Young Adults. Top. Antivir. Med. 2020, 28, 459–462. [Google Scholar] [PubMed]
- Gupta, K.; Harrison, S.A.; Davis, N.A.; Culp, M.L.; Hand, S.C.; Simpson, T.; Van Der Pol, B.; Galbraith, J.W.; Van Wagoner, N.J.; Morrison, S.G.; et al. Prevalence of Chlamydia trachomatis Infection in Young Women and Associated Predictors. Sex. Transm. Dis. 2021, 48, 529–535. [Google Scholar] [CrossRef]
- Li, Y.; Liao, Z.; Wang, Q.; He, W.; Deng, Y.; Liu, C. Prevalence of Chlamydia trachomatis, Neisseria gonorrhoeae, and Ureaplasma urealyticum Infections in Males and Females of Childbearing Age in Chengdu, China. Front. Cell. Infect. Microbiol. 2025, 15, 1566163. [Google Scholar] [CrossRef]
- Martins, E.R.; Melo-Cristino, J.; Ramirez, M.; Portuguese Group for the Study of Streptococcal Infections. Dominance of Serotype Ia among Group B Streptococci Causing Invasive Infections in Nonpregnant Adults in Portugal. J. Clin. Microbiol. 2012, 50, 1219–1227. [Google Scholar] [CrossRef]
- Kaliterna, V.; Kaliterna, M.; Pejković, L.; Hofman, I.D.; Andelinović, S. Prevalence and genotyping of the human papillomavirus in the cervical specimens among women of Southern Croatia (Dalmatia County). Cent. Eur. J. Public Health 2013, 21, 26–29. [Google Scholar] [CrossRef]
- Sabol, I.; Milutin Gašperov, N.; Matovina, M.; Božinović, K.; Grubišić, G.; Fistonić, I.; Belci, D.; Alemany, L.; Džebro, S.; Dominis, M.; et al. Cervical HPV type-specific pre-vaccination prevalence and age distribution in Croatia. PLoS ONE 2017, 12, e0180480. [Google Scholar] [CrossRef] [PubMed]
- Sousa, H.; Tavares, A.; Campos, C.; Marinho-Dias, J.; Brito, M.; Medeiros, R.; Baldaque, I.; Lobo, C.; Leča, L.; Monteiro, P.; et al. High-Risk human papillomavirus genotype distribution in the Northern region of Portugal: Data from regional cervical cancer screening program. Papillomavirus Res. 2019, 8, 100179. [Google Scholar] [CrossRef]
- Pešut, E.; Šimić, I.; Fureš, R.; Milutin Gašperov, N.; Lež, C.; Feratović, F.; Kukina Žvigač, T.; Grce, M.; Erceg Ivkošić, I.; Sabol, I. Monitoring HPV Prevalence and Risk Cofactors for Abnormal Cytology in the Post-Vaccination Period among Croatian Women. Viruses 2024, 16, 642. [Google Scholar] [CrossRef] [PubMed]
- Reuschenbach, M.; Valente, S.; Takyar, J.; Dhawan, A.; Hall, A.; Agrawal, N.; Ghelardi, A.; Del Pino, M.; Nowakowski, A.; Sabale, U. Treatment characteristics, HPV genotype distribution and risk of subsequent disease among women with high-grade cervical intraepithelial neoplasia in Europe: A systematic literature review. Eur. J. Obstet. Gynecol. Reprod. Biol. 2024, 300, 129–140. [Google Scholar] [CrossRef] [PubMed]
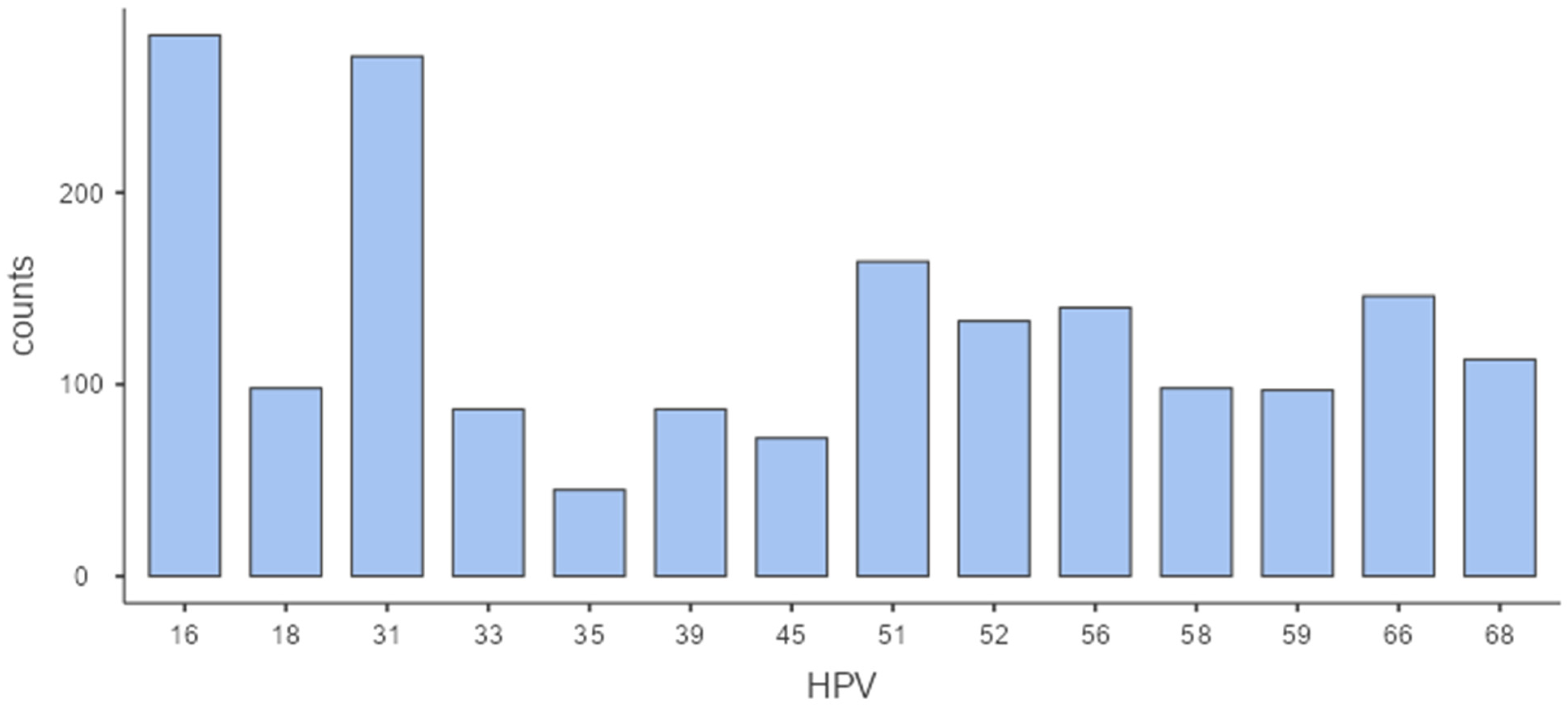
| HPV Genotype | Counts | % of Total High-Risk HPV-Positive Women (N = 1211) | % of Total Genotypes (N = 1833) |
|---|---|---|---|
| 16 | 282 | 23.3% | 15.4% |
| 18 | 98 | 8.1% | 5.3% |
| 31 | 271 | 22.4% | 14.8% |
| 33 | 87 | 7.2% | 4.7% |
| 35 | 45 | 3.7% | 2.5% |
| 39 | 87 | 7.2% | 4.7% |
| 45 | 72 | 5.9% | 3.9% |
| 51 | 164 | 13.5% | 8.9% |
| 52 | 133 | 11.0% | 7.3% |
| 56 | 140 | 11.6% | 7.6% |
| 58 | 98 | 8.1% | 5.3% |
| 59 | 97 | 8.0% | 5.3% |
| 66 | 146 | 12.1% | 8.0% |
| 68 | 113 | 9.3% | 6.2% |
| High-Risk HPV Group | |||||
|---|---|---|---|---|---|
| Age Group | 18/45 | 16 High Risk | 16-Related | Lower Risk | Total |
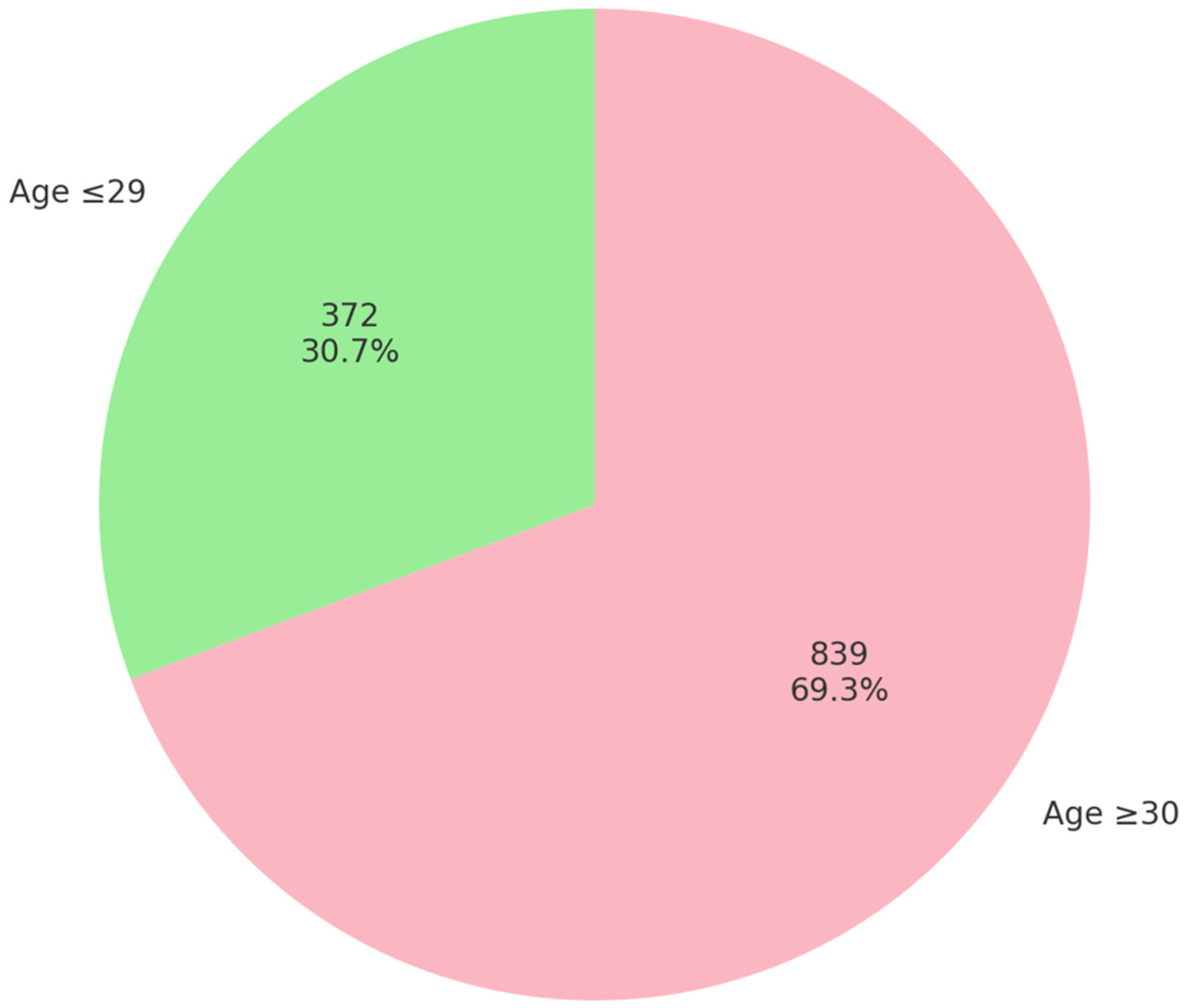
| 29 or under | 109 | 183 | 393 | 467 | 1152 |
| 30 or over | 61 | 99 | 241 | 280 | 681 |
| Total | 170 | 282 | 634 | 747 | 1833 |
| p-value | 0.835 | ||||
| HPV Groups | Predictor | Estimate | SE | Z | p-Value | OR |
|---|---|---|---|---|---|---|
| 16 high-risk—18/45 | Intercept | 0.5626 | 0.169 | 3.336 | <0.001 | 1.755 |
| CT: | ||||||
| P—N | −0.6578 | 0.531 | −1.239 | 0.215 | 0.518 | |
| MG: | ||||||
| P—N | −0.3040 | 0.946 | −0.321 | 0.748 | 0.738 | |
| Mh: | ||||||
| P—N | −0.1740 | 0.471 | −0.369 | 0.712 | 0.840 | |
| Uu: | ||||||
| P—N | −0.0786 | 0.407 | −0.193 | 0.847 | 0.924 | |
| GV: | ||||||
| P—N | 0.8512 | 0.475 | 1.791 | 0.073 | 2.343 | |
| 16-related—18/45 | Intercept | 1.3876 | 0.151 | 9.195 | <0.001 | 4.005 |
| CT: | ||||||
| P—N | −0.9583 | 0.479 | −2.000 | 0.045 | 0.384 | |
| MG: | ||||||
| P—N | −1.0822 | 0.941 | −1.150 | 0.250 | 0.339 | |
| Mh: | ||||||
| P—N | −0.1778 | 0.425 | −0.418 | 0.676 | 0.837 | |
| Uu: | ||||||
| P—N | −0.2572 | 0.371 | −0.694 | 0.488 | 0.773 | |
| GV: | ||||||
| P—N | 0.9619 | 0.441 | 2.180 | 0.029 | 2.617 | |
| Lower risk—18/45 | Intercept | 1.6403 | 0.147 | 11.134 | <0.001 | 5.157 |
| CT: | ||||||
| P—N | −1.2722 | 0.474 | −2.684 | 0.007 | 0.280 | |
| MG: | ||||||
| P—N | −0.4983 | 0.822 | −0.606 | 0.544 | 0.608 | |
| Mh: | ||||||
| P—N | 0.1108 | 0.408 | 0.271 | 0.786 | 1.117 | |
| Uu: | ||||||
| P—N | −0.1185 | 0.357 | −0.332 | 0.740 | 0.888 | |
| GV: | ||||||
| P—N | 0.8316 | 0.434 | 1.916 | 0.055 | 2.297 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kaliterna, V.; Meštrović, T.; Čorić-Mesarić, M.; Božičević, I. Genotype-Specific Distribution of High-Risk Human Papillomavirus (HPV) and Microbial Co-Detections in HPV-Positive Women from Southern Croatia. Biomedicines 2025, 13, 2100. https://doi.org/10.3390/biomedicines13092100
Kaliterna V, Meštrović T, Čorić-Mesarić M, Božičević I. Genotype-Specific Distribution of High-Risk Human Papillomavirus (HPV) and Microbial Co-Detections in HPV-Positive Women from Southern Croatia. Biomedicines. 2025; 13(9):2100. https://doi.org/10.3390/biomedicines13092100
Chicago/Turabian StyleKaliterna, Vanja, Tomislav Meštrović, Mirjana Čorić-Mesarić, and Ivana Božičević. 2025. "Genotype-Specific Distribution of High-Risk Human Papillomavirus (HPV) and Microbial Co-Detections in HPV-Positive Women from Southern Croatia" Biomedicines 13, no. 9: 2100. https://doi.org/10.3390/biomedicines13092100
APA StyleKaliterna, V., Meštrović, T., Čorić-Mesarić, M., & Božičević, I. (2025). Genotype-Specific Distribution of High-Risk Human Papillomavirus (HPV) and Microbial Co-Detections in HPV-Positive Women from Southern Croatia. Biomedicines, 13(9), 2100. https://doi.org/10.3390/biomedicines13092100






