Characterization of T Helper 1 and 2 Cytokine Profiles in Newborns of Mothers with COVID-19
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethical Aspects and Hospital Units
2.2. Target Population
2.3. Collection and Processing of Samples
2.4. Profile Characterization of Cytokines by Flow Cytometry
2.5. Statistical Analysis
3. Results
3.1. Clinical Aspects
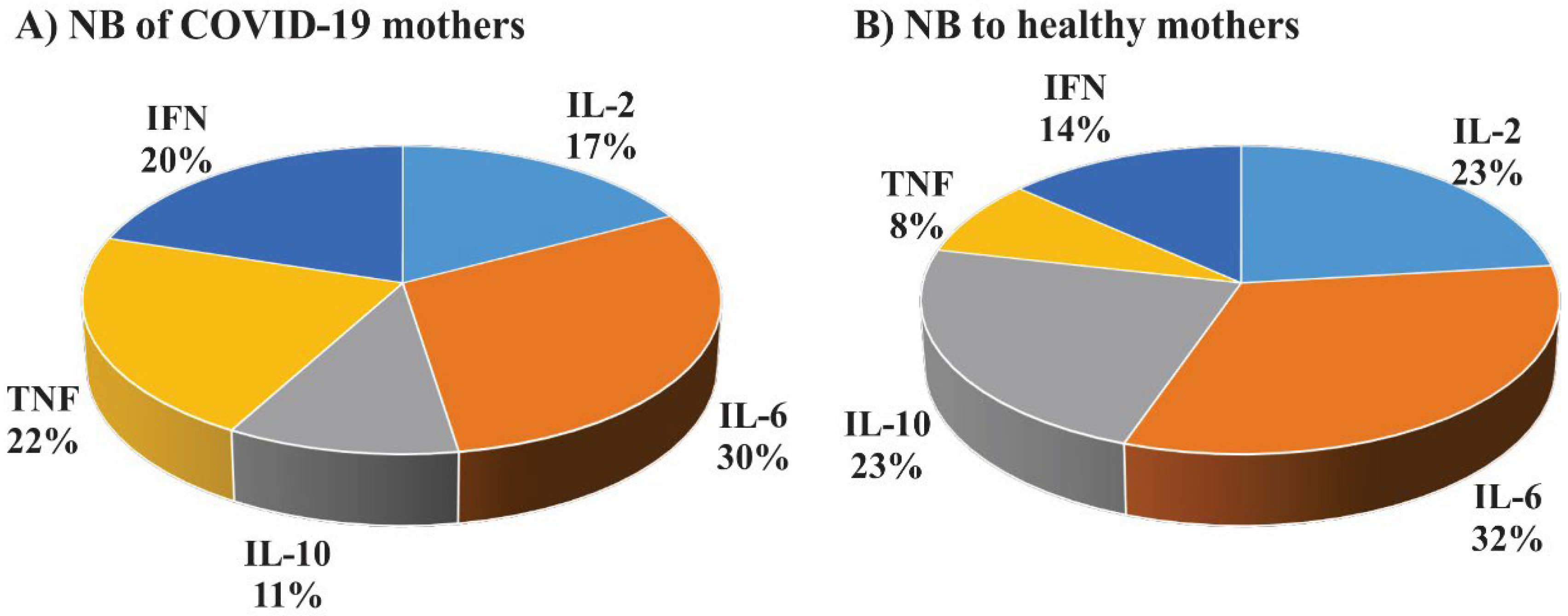
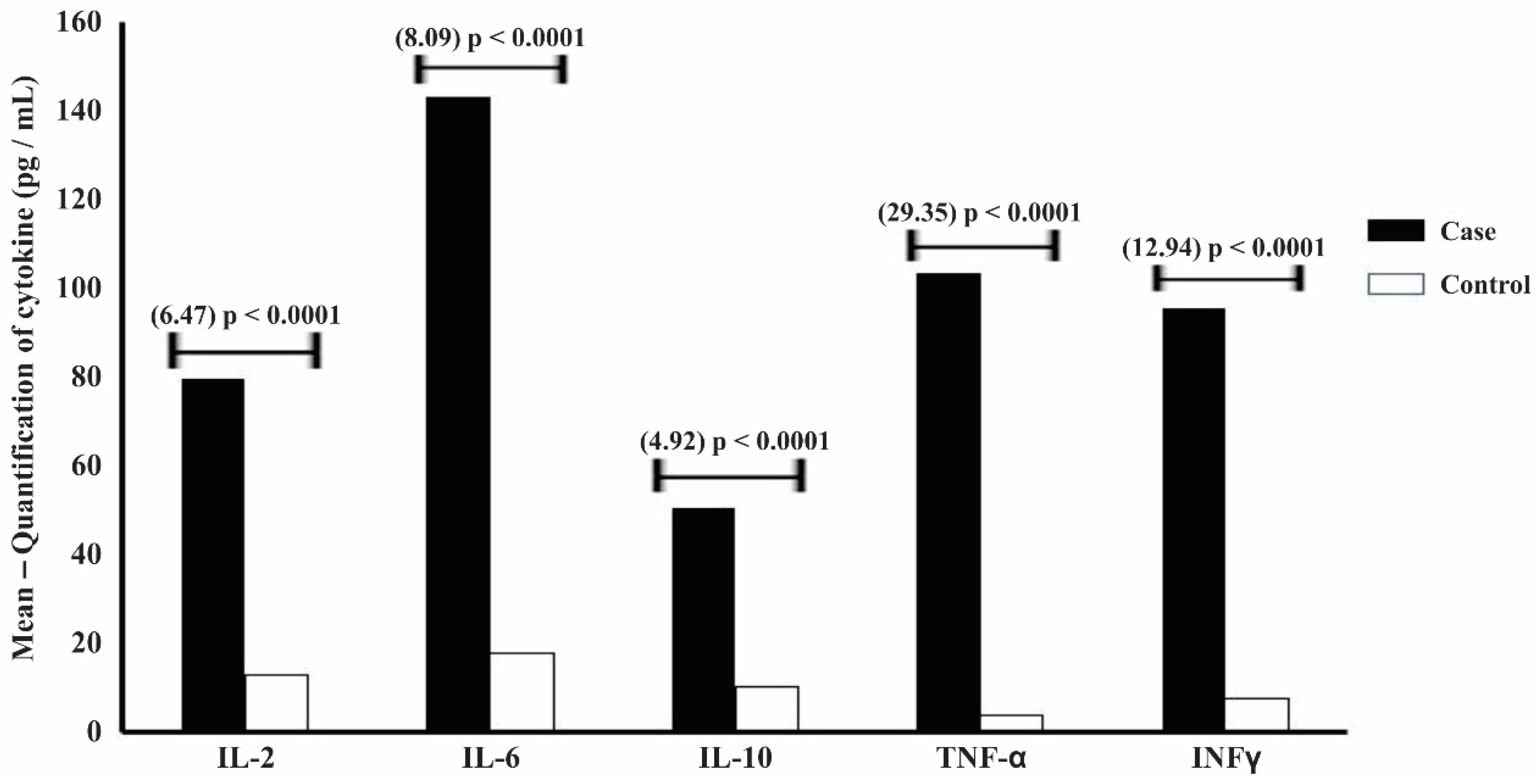
3.2. Immunophenotypic Analysis of Cytokines Using Flow Cytometry
3.3. Comparative Analysis of Immune Humoral and Cellular Responses in Newborns of Mothers with COVID-19
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Zhou, P.; Yang, X.L.; Wang, X.G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.R.; Zhu, Y.; Li, B.; Huang, C.L.; et al. Addendum: A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 588, E6. [Google Scholar] [CrossRef] [PubMed]
- Berman, S. Epidemiology of acute respiratory infections in children of developing countries. Rev. Infect. Dis. 1991, 13, S454–S462. [Google Scholar] [CrossRef] [PubMed]
- WHO. Overview of Sentinel Systems for Hospitalized Severe Acute Respiratory Infections (SARI) Represented in the Weekly EuroFlu Surveillance Bulletin (as of 10 February 2013) 2013 18 July 2018; WHO: Geneva, Switzerland, 2018.
- Li, H.; Wei, Q.; Tan, A.; Wang, L. Epidemiological analysis of respiratory viral etiology for influenza-like illness during 2010 in Zhuhai, China. Virol. J. 2013, 10, 143. [Google Scholar] [CrossRef]
- Parveen, S.; Sullender, W.M.; Fowler, K.; Lefkowitz, E.J.; Kapoor, S.K.; Broor, S. Genetic variability in the G protein gene of group A and B respiratory syncytial viruses from India. J. Clin. Microbiol. 2006, 44, 3055–3064. [Google Scholar] [CrossRef] [PubMed]
- Williams, B.G.; Gouws, E.; Boschi-Pinto, C.; Bryce, J.; Dye, C. Estimates of world-wide distribution of child deaths from acute respiratory infections. Lancet Infect. Dis. 2002, 2, 25–32. [Google Scholar] [CrossRef]
- Bharaj, P.; Sullender, W.M.; Kabra, S.K.; Mani, K.; Cherian, J.; Tyagi, V.; Chahar, H.S.; Kaushik, S.; Dar, L.; Broor, S. Respiratory viral infections detected by multiplex PCR among pediatric patients with lower respiratory tract infections seen at an urban hospital in Delhi from 2005 to 2007. Virol. J. 2009, 6, 89. [Google Scholar] [CrossRef]
- Collins, S.A.; Surmala, P.; Osborne, G.; Greenberg, C.; Bathory, L.W.; Edmunds-Potvin, S.; Arbour, L. Causes and risk factors for infant mortality in Nunavut, Canada 1999–2011. BMC Pediatr. 2012, 12, 190. [Google Scholar] [CrossRef]
- Nair, H.; Simões, E.A.; Rudan, I.; Gessner, B.D.; Azziz-Baumgartner, E.; Zhang, J.S.F.; Feikin, D.R.; Mackenzie, G.A.; Moiïsi, J.C.; Roca, A. Global and regional burden of hospital admissions for severe acute lower respiratory infections in young children in 2010: A systematic analysis. Lancet 2013, 381, 1380–1390. [Google Scholar] [CrossRef]
- Jamieson, D.J.; Rasmussen, S.A. An update on COVID-19 and pregnancy. Am. J. Obstet. Gynecol. 2021, 226, 177–186. [Google Scholar] [CrossRef]
- Argueta, L.B.; Lacko, L.A.; Bram, Y.; Tada, T.; Carrau, L.; Zhang, T.; Uhl, S.; Lubor, B.C.; Chandar, V.; Gil, C. SARS-CoV-2 infects syncytiotrophoblast and activates inflammatory responses in the placenta. BioRxiv 2021. [Google Scholar] [CrossRef]
- Cribiù, F.M.; Erra, R.; Pugni, L.; Rubio-Perez, C.; Alonso, L.; Simonetti, S.; Croci, G.A.; Serna, G.; Ronchi, A.; Pietrasanta, C. Severe SARS-CoV-2 placenta infection can impact neonatal outcome in the absence of vertical transmission. J. Clin. Investig. 2021, 131, e145427. [Google Scholar] [CrossRef] [PubMed]
- Bahadur, G.; Bhat, M.; Acharya, S.; Janga, D.; Cambell, B.; Huirne, J.; Yoong, W.; Govind, A.; Pardo, J.; Homburg, R. Retrospective observational RT-PCR analyses on 688 babies born to 843 SARS-CoV-2 positive mothers, placental analyses and diagnostic analyses limitations suggest vertical transmission is possible. Facts Views Vis. ObGyn 2021, 13, 53. [Google Scholar] [CrossRef] [PubMed]
- Jin, J.C.; Ananthanarayanan, A.; Brown, J.A.; Rager, S.L.; Bram, Y.; Sanidad, K.Z.; Amir, M.; Baergen, R.N.; Stuhlmann, H.; Schwartz, R.E. SARS CoV-2 detected in neonatal stool remote from maternal COVID-19 during pregnancy. Pediatr. Res. 2022, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Civardi, E.; Tzialla, C.; Baldanti, F.; Strocchio, L.; Manzoni, P.; Stronati, M. Viral outbreaks in neonatal intensive care units: What we do not know. Am. J. Infect. Control 2013, 41, 854–856. [Google Scholar] [CrossRef]
- Gelber, S.E.; Ratner, A.J. Hospital-acquired viral pathogens in the neonatal intensive care unit. In Seminars in Perinatology; WB Saunders: Philadelphia, PA, USA, 2002; pp. 346–356. [Google Scholar] [CrossRef]
- Faden, H.; Wynn, R.J.; Campagna, L.; Ryan, R.M. Outbreak of adenovirus type 30 in a neonatal intensive care unit. J. Pediatr. 2005, 146, 523–527. [Google Scholar] [CrossRef]
- Gagneur, A.; Vallet, S.; Talbot, P.J.; Legrand-Quillien, M.-C.; Picard, B.; Payan, C.; Sizun, J. Outbreaks of human coronavirus in a paediatric and neonatal intensive care unit. Eur. J. Pediatr. 2008, 167, 1427–1434. [Google Scholar] [CrossRef]
- Nair, H.; Nokes, D.J.; Gessner, B.D.; Dherani, M.; Madhi, S.A.; Singleton, R.J.; O’Brien, K.L.; Roca, A.; Wright, P.F.; Bruce, N. Global burden of acute lower respiratory infections due to respiratory syncytial virus in young children: A systematic review and meta-analysis. Lancet 2010, 375, 1545–1555. [Google Scholar] [CrossRef]
- Kusel, M.M.; de Klerk, N.H.; Holt, P.G.; Kebadze, T.; Johnston, S.L.; Sly, P.D. Role of respiratory viruses in acute upper and lower respiratory tract illness in the first year of life: A birth cohort study. Pediatr. Infect. Dis. J. 2006, 25, 680–686. [Google Scholar] [CrossRef]
- Straliotto, S.M.; Siqueira, M.M.; Machado, V.; Maia, T.M. Respiratory viruses in the pediatric intensive care unit: Prevalence and clinical aspects. Memórias Inst. Oswaldo Cruz 2004, 99, 883–887. [Google Scholar] [CrossRef]
- Gupta, N.; Richter, R.; Robert, S.; Kong, M. Viral sepsis in children. Front. Pediatr. 2018, 6, 252. [Google Scholar] [CrossRef]
- Lin, G.-L.; McGinley, J.P.; Drysdale, S.B.; Pollard, A.J. Epidemiology and immune pathogenesis of viral sepsis. Front. Immunol. 2018, 9, 2147. [Google Scholar] [CrossRef]
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef]
- Abbas, A.K.; Lichtman, A.H.; Pillai, S. Cellular and Molecular Immunology; WB Saunders: Philadelphia, PA, USA, 2017. [Google Scholar]
- Pedersen, S.F.; Ho, Y.-C. SARS-CoV-2: A storm is raging. J. Clin. Investig. 2020, 130, 2202–2205. [Google Scholar] [CrossRef] [PubMed]
- Mehta, P.; McAuley, D.F.; Brown, M.; Sanchez, E.; Tattersall, R.S.; Manson, J.J. COVID-19: Consider cytokine storm syndromes and immunosuppression. Lancet 2020, 395, 1033–1034. [Google Scholar] [CrossRef] [PubMed]
- Jose, R.J.; Manuel, A. COVID-19 cytokine storm: The interplay between inflammation and coagulation. Lancet Respir. Med. 2020, 8, e46–e47. [Google Scholar] [CrossRef] [PubMed]
- Saunders-Hastings, P.R.; Krewski, D. Reviewing the history of pandemic influenza: Understanding patterns of emergence and transmission. Pathogens 2016, 5, 66. [Google Scholar] [CrossRef]
- Knight, M.; Bunch, K.; Vousden, N.; Morris, E.; Simpson, N.; Gale, C.; O’Brien, P.; Quigley, M.; Brocklehurst, P.; Kurinczuk, J.J. Characteristics and outcomes of pregnant women hospitalised with confirmed SARS-CoV-2 infection in the UK: A national cohort study using the UK Obstetric Surveillance System (UKOSS). Medrxiv 2020. [Google Scholar] [CrossRef]
- Zhao, J.; Kim, K.D.; Yang, X.; Auh, S.; Fu, Y.-X.; Tang, H. Hyper innate responses in neonates lead to increased morbidity and mortality after infection. Proc. Natl. Acad. Sci. USA 2008, 105, 7528–7533. [Google Scholar] [CrossRef]
- Green, J.; Petty, J.; Bromley, P.; Walker, K.; Jones, L. COVID-19 in babies: Knowledge for neonatal care. J. Neonatal Nurs. 2020, 26, 239–246. [Google Scholar] [CrossRef]
- Mangalmurti, N.; Hunter, C.A. Cytokine storms: Understanding COVID-19. Immunity 2020, 53, 19–25. [Google Scholar] [CrossRef]
- Bell, M.J.; Ternberg, J.L.; Feigin, R.D.; Keating, J.P.; Marshall, R.; Barton, L.; Brotherton, T. Neonatal necrotizing enterocolitis. Therapeutic decisions based upon clinical staging. Ann. Surg. 1978, 187, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Sankar, M.J.; Agarwal, R.; Deorari, A.K.; Paul, V.K. Sepsis in the newborn. Indian J. Pediatr. 2008, 75, 261–266. [Google Scholar] [CrossRef] [PubMed]
- Manganaro, R.; Mamì, C.; Gemelli, M. The validity of the Apgar scores in the assessment of asphyxia at birth. Eur. J. Obstet. Gynecol. Reprod. Biol. 1994, 54, 99–102. [Google Scholar] [CrossRef] [PubMed]
- Gilstrap III, L.C.; Leveno, K.J.; Burris, J.; Williams, M.L.; Little, B.B. Diagnosis of birth asphyxia on the basis of fetal pH, Apgar score, and newborn cerebral dysfunction. Am. J. Obstet. Gynecol. 1989, 161, 825–830. [Google Scholar] [CrossRef]
- Aggarwal, R.; Singhal, A.; Deorari, A.K.; Paul, V.K. Apnea in the newborn. Indian J. Pediatr. 2001, 68, 959–962. [Google Scholar] [CrossRef]
- Silverman, W.A.; Andersen, D.H. A controlled clinical trial of effects of water mist on obstructive respiratory signs, death rate and necropsy findings among premature infants. Pediatrics 1956, 17, 1–10. [Google Scholar]
- Ghasemi, A.; Zahediasl, S. Normality tests for statistical analysis: A guide for non-statisticians. Int. J. Endocrinol. Metab. 2012, 10, 486–489. [Google Scholar] [CrossRef]
- Ostertagova, E.; Ostertag, O.; Kováč, J. Methodology and application of the Kruskal-Wallis test. In Applied Mechanics and Materials; Trans Tech Publications Ltd.: Wollerau, Switzerland, 2016; pp. 115–120. [Google Scholar] [CrossRef]
- Hart, A. Mann-Whitney test is not just a test of medians: Differences in spread can be important. BMJ 2001, 323, 391–393. [Google Scholar] [CrossRef]
- Carlos, W.G.; Dela Cruz, C.S.; Cao, B.; Pasnick, S.; Jamil, S. Novel Wuhan (2019-nCoV) Coronavirus. Am. J. Respir. Crit. Care Med. 2020, 201, P7–P8. [Google Scholar] [CrossRef]
- Song, F.; Shi, N.; Shan, F.; Zhang, Z.; Shen, J.; Lu, H.; Ling, Y.; Jiang, Y.; Shi, Y. Emerging 2019 novel coronavirus (2019-nCoV) pneumonia. Radiology 2020, 295, 210–217. [Google Scholar] [CrossRef]
- Jamilloux, Y.; Henry, T.; Belot, A.; Viel, S.; Fauter, M.; El Jammal, T.; Walzer, T.; François, B.; Sève, P. Should we stimulate or suppress immune responses in COVID-19? Cytokine and anti-cytokine interventions. Autoimmun. Rev. 2020, 19, 102567. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, T.; Ellingson, M.K.; Wong, P.; Israelow, B.; Lucas, C.; Klein, J.; Silva, J.; Mao, T.; Oh, J.E.; Tokuyama, M. Sex differences in immune responses that underlie COVID-19 disease outcomes. Nature 2020, 588, 315–320. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Wang, T.; Cai, D.; Hu, Z.; Liao, H.; Zhi, L.; Wei, H.; Zhang, Z.; Qiu, Y.; Wang, J. Cytokine storm intervention in the early stages of COVID-19 pneumonia. Cytokine Growth Factor Rev. 2020, 53, 38–42. [Google Scholar] [CrossRef] [PubMed]
- Hu, B.; Huang, S.; Yin, L. The cytokine storm and COVID-19. J. Med. Virol. 2021, 93, 250–256. [Google Scholar] [CrossRef] [PubMed]
- Hong, H.; Wang, Y.; Chung, H.-T.; Chen, C.-J. Clinical characteristics of novel coronavirus disease 2019 (COVID-19) in newborns, infants and children. Pediatr. Neonatol. 2020, 61, 131–132. [Google Scholar] [CrossRef]
- Mirbeyk, M.; Saghazadeh, A.; Rezaei, N. A systematic review of pregnant women with COVID-19 and their neonates. Arch. Gynecol. Obstet. 2021, 304, 5–38. [Google Scholar] [CrossRef]
- Aleebrahim-Dehkordi, E.; Soveyzi, F.; Deravi, N.; Rabbani, Z.; Saghazadeh, A.; Rezaei, N. Human coronaviruses SARS-CoV, MERS-CoV, and SARS-CoV-2 in children. J. Pediatr. Nurs. 2021, 56, 70–79. [Google Scholar] [CrossRef] [PubMed]
- Zeng, L.; Xia, S.; Yuan, W.; Yan, K.; Xiao, F.; Shao, J.; Zhou, W. Neonatal early-onset infection with SARS-CoV-2 in 33 neonates born to mothers with COVID-19 in Wuhan, China. JAMA Pediatr. 2020, 174, 722–725. [Google Scholar] [CrossRef]
- Cook, J.; Harman, K.; Zoica, B.; Verma, A.; D’Silva, P.; Gupta, A. Horizontal transmission of severe acute respiratory syndrome coronavirus 2 to a premature infant: Multiple organ injury and association with markers of inflammation. Lancet Child. Adolesc. Health 2020, 4, 548–551. [Google Scholar] [CrossRef]
- Rezaei, N. COVID-19 affects healthy pediatricians more than pediatric patients. Infect. Control. Hosp. Epidemiol. 2020, 41, 1106–1107. [Google Scholar] [CrossRef]
- Yang, H.; Wang, C.; Poon, L. Novel coronavirus infection and pregnancy. Ultrasound Obstet. Gynecol. 2020, 55, 435. [Google Scholar] [CrossRef] [PubMed]
- Liu, P.; Zheng, J.; Yang, P.; Wang, X.; Wei, C.; Zhang, S.; Feng, S.; Lan, J.; He, B.; Zhao, D. The immunologic status of newborns born to SARS-CoV-2–infected mothers in Wuhan, China. J. Allergy Clin. Immunol. 2020, 146, 101–109.e101. [Google Scholar] [CrossRef] [PubMed]
- Kam, K.; Yung, C.; Cui, L. A well infant with coronavirusdisease 2019 (COVID-19) with high viral load. Clin. Infect. Dis. 2020, 71, 847–849. [Google Scholar] [CrossRef] [PubMed]
- Dashraath, P.; Wong, J.L.J.; Lim, M.X.K.; Lim, L.M.; Li, S.; Biswas, A.; Choolani, M.; Mattar, C.; Su, L.L. Coronavirus disease 2019 (COVID-19) pandemic and pregnancy. Am. J. Obstet. Gynecol. 2020, 222, 521–531. [Google Scholar] [CrossRef]
- Berger, A. Th1 and Th2 responses: What are they? BMJ 2000, 321, 424. [Google Scholar] [CrossRef]
- Melendi, G.A.; Laham, F.R.; Monsalvo, A.C.; Casellas, J.M.; Israele, V.; Polack, N.R.; Kleeberger, S.R.; Polack, F.P. Cytokine profiles in the respiratory tract during primary infection with human metapneumovirus, respiratory syncytial virus, or influenza virus in infants. Pediatrics 2007, 120, e410–e415. [Google Scholar] [CrossRef]
- Pancham, K.; Perez, G.F.; Huseni, S.; Jain, A.; Kurdi, B.; Rodriguez-Martinez, C.E.; Preciado, D.; Rose, M.C.; Nino, G. Premature infants have impaired airway antiviral IFNγ responses to human metapneumovirus compared to respiratory syncytial virus. Pediatr. Res. 2015, 78, 389–394. [Google Scholar] [CrossRef]
- Broughton, S.; Sylvester, K.P.; Fox, G.; Zuckerman, M.; Smith, M.; Milner, A.D.; Rafferty, G.F.; Greenough, A. Lung function in prematurely born infants after viral lower respiratory tract infections. Pediatr. Infect. Dis. J. 2007, 26, 1019–1024. [Google Scholar] [CrossRef]
- García-García, M.; Calvo, C.; Casas, I.; Bracamonte, T.; Rellán, A.; Gozalo, F.; Tenorio, T.; Perez-Brena, P. Human metapneumovirus bronchiolitis in infancy is an important risk factor for asthma at age 5. Pediatr. Pulmonol. 2007, 42, 458–464. [Google Scholar] [CrossRef]
- Drysdale, S.B.; Prendergast, M.; Alcazar, M.; Wilson, T.; Smith, M.; Zuckerman, M.; Broughton, S.; Rafferty, G.F.; Johnston, S.L.; Hodemaekers, H.M. Genetic predisposition of RSV infection-related respiratory morbidity in preterm infants. Eur. J. Pediatr. 2014, 173, 905–912. [Google Scholar] [CrossRef]
- Hadjadj, J.; Yatim, N.; Barnabei, L.; Corneau, A.; Boussier, J.; Smith, N.; Péré, H.; Charbit, B.; Bondet, V.; Chenevier-Gobeaux, C. Impaired type I interferon activity and inflammatory responses in severe COVID-19 patients. Science 2020, 369, 718–724. [Google Scholar] [CrossRef] [PubMed]
- Karki, R.; Sharma, B.R.; Tuladhar, S.; Williams, E.P.; Zalduondo, L.; Samir, P.; Zheng, M.; Sundaram, B.; Banoth, B.; Malireddi, R.S. Synergism of TNF-α and IFN-γ triggers inflammatory cell death, tissue damage, and mortality in SARS-CoV-2 infection and cytokine shock syndromes. Cell 2021, 184, 149–168.e117. [Google Scholar] [CrossRef]
- King, A.; Vail, A.; O’Leary, C.; Hannan, C.; Brough, D.; Patel, H.; Galea, J.; Ogungbenro, K.; Wright, M.; Pathmanaban, O. Anakinra in COVID-19: Important considerations for clinical trials. Lancet Rheumatol. 2020, 2, e379–e381. [Google Scholar] [CrossRef] [PubMed]
- Christiaansen, A.F.; Syed, M.A.; Ten Eyck, P.P.; Hartwig, S.M.; Durairaj, L.; Kamath, S.S.; Varga, S.M. Altered Treg and cytokine responses in RSV-infected infants. Pediatr. Res. 2016, 80, 702–709. [Google Scholar] [CrossRef] [PubMed]
- Chung, J.-Y.; Han, T.H.; Kim, J.-S.; Kim, S.W.; Park, C.-G.; Hwang, E.-S. Th1 and Th2 cytokine levels in nasopharyngeal aspirates from children with human bocavirus bronchiolitis. J. Clin. Virol. 2008, 43, 223–225. [Google Scholar] [CrossRef]
- Pala, P.; Bjarnason, R.; Sigurbergsson, F.; Metcalfe, C.; Sigurs, N.; Openshaw, P. Enhanced IL-4 responses in children with a history of respiratory syncytial virus bronchiolitis in infancy. Eur. Respir. J. 2002, 20, 376–382. [Google Scholar] [CrossRef]
- Legg, J.P.; Hussain, I.R.; Warner, J.A.; Johnston, S.L.; Warner, J.O. Type 1 and type 2 cytokine imbalance in acute respiratory syncytial virus bronchiolitis. Am. J. Respir. Crit. Care Med. 2003, 168, 633–639. [Google Scholar] [CrossRef]
- Tripp, R.A.; Moore, D.; Barskey, A.T.; Jones, L.; Moscatiello, C.; Keyserling, H.; Anderson, L.J. Peripheral blood mononuclear cells from infants hospitalized because of respiratory syncytial virus infection express T helper-1 and T helper-2 cytokines and CC chemokine messenger RNA. J. Infect. Dis. 2002, 185, 1388–1394. [Google Scholar] [CrossRef]
- Brandenburg, A.H.; Kleinjan, A.; van Het Land, B.; Moll, H.A.; Timmerman, H.H.; de Swart, R.L.; Neijens, H.J.; Fokkens, W.; Osterhaus, A.D. Type 1-like immune response is found in children with respiratory syncytial virus infection regardless of clinical severity. J. Med. Virol. 2000, 62, 267–277. [Google Scholar] [CrossRef]
- Vennemann, M.M.; Loddenkotter, B.; Fracasso, T.; Mitchell, E.A.; Debertin, A.S.; Larsch, K.P.; Sperhake, J.P.; Brinkmann, B.; Sauerland, C.; Lindemann, M.; et al. Cytokines and sudden infant death. Int. J. Leg. Med. 2012, 126, 279–284. [Google Scholar] [CrossRef]
- Berdat, P.A.; Wehrle, T.J.; Kung, A.; Achermann, F.; Sutter, M.; Carrel, T.P.; Nydegger, U.E. Age-specific analysis of normal cytokine levels in healthy infants. Clin. Chem. Lab. Med. 2003, 41, 1335–1339. [Google Scholar] [CrossRef] [PubMed]
- Saha, S.; Pal, P.; Mukherjee, D. Neonatal MIS-C: Managing the Cytokine Storm. Pediatrics 2021, 148, e2020042093. [Google Scholar] [CrossRef] [PubMed]
- Debock, I.; Flamand, V. Unbalanced Neonatal CD4(+) T-Cell Immunity. Front. Immunol. 2014, 5, 393. [Google Scholar] [CrossRef] [PubMed]
- Heinonen, S.; Rodriguez-Fernandez, R.; Diaz, A.; Oliva Rodriguez-Pastor, S.; Ramilo, O.; Mejias, A. Infant Immune Response to Respiratory Viral Infections. Immunol. Allergy Clin. N. Am. 2019, 39, 361–376. [Google Scholar] [CrossRef]
- Yan, L.; Zhang, H.-T.; Goncalves, J.; Xiao, Y.; Wang, M.; Guo, Y.; Sun, C.; Tang, X.; Jing, L.; Zhang, M. An interpretable mortality prediction model for COVID-19 patients. Nat. Mach. Intell. 2020, 2, 283–288. [Google Scholar] [CrossRef]
- Reeve, J.L.; Twomey, P.J. Consider laboratory aspects in developing patient prediction models. Nat. Mach. Intell. 2021, 3, 18. [Google Scholar] [CrossRef]
- Can, E.; Oğlak, S.C.; Ölmez, F.; Bulut, H. Serum neutrophil gelatinase-associated lipocalin concentrations are significantly associated with the severity of COVID-19 in pregnant patients. Saudi Med. J. 2022, 43, 559–566. [Google Scholar] [CrossRef]
- Göklü, M.R.; Şeyhmus, T.; Aksin, Ş.; Andan, C. Effects of pro-inflammatory interleukin-6 and anti-inflammatory interleukin-10 cytokines in pregnant women diagnosed with coronavirus disease 2019. Anatol. Clin. J. Med. Sci. 2022, 27, 268–274. [Google Scholar]

| Sex | Total (%) |
|---|---|
| Male | 45% |
| Female | 55% |
| Clinical manifestations | Total (%) |
| Enterocolitis | 5% |
| Early-onset neonatal sepsis | 20% |
| Asphyxia | 10% |
| Cough | 5% |
| Fever | 10% |
| Apnea | 20% |
| Respiratory discomfort | 35% |
| Need for NICU | 100% |
| Deaths | 0% |
| Laboratory tests | Average (min. and max.) |
| Aspartate aminotransferase | 56 (26–78) |
| Alanine aminotransferase | 11.77 (0.8–21) |
| Total bilirubin | 6.04 (2.8–11.8) |
| Direct bilirubin | 1.16 (0.3–5.7) |
| Urea | 24.64 (7–42) |
| Creatinine | 0.76 (0.5–1.3) |
| Creatinophosphokinase | 445.64 (109–1.197) |
| CK-MB | 44.81 (23–55) |
| Troponin | All negative |
| Lactate | 33.77 (4.1–72.6) |
| D-dimer | 585 (1.06–1066) |
| PCR | 0.22 (0.1–0.7) |
| Lactate dehydrogenase | 596.9 (457–1036) |
| Ferritin | 245.65 (150.32–558.71) |
| Triglycerides | 118.72 (43–239) |
| RT-qPCR | All negative |
| Oxygen saturation | 98.97 (98.2–99.7) |
| Total leukocytes | 10.790 (5.500–18,600) |
| Neutrophils | 5.433 (1.975–12,640) |
| Lymphocytes | 3.975 (1.637–9.343) |
| Platelets | 205.384 (14.500–372.000) |
| Quantification of Cytokines (pg/mL) | |||||
|---|---|---|---|---|---|
| Patient ID | IL-2 | IL-6 | IL-10 | TNF-α | IFN-γ |
| 1 | 74.51 | 136.26 | 51.21 | 103.25 | 100.2 |
| 2 | 79.69 | 141.95 | 46.63 | 101.22 | 96.36 |
| 3 | 77.94 | 127.59 | 38.85 | 95.38 | 92.14 |
| 4 | 73.15 | 126.83 | 38.73 | 94.71 | 91.16 |
| 5 | 71.43 | 128.19 | 43.43 | 99.06 | 91.37 |
| 6 | 79.98 | 113.69 | 47.97 | 97.73 | 91.13 |
| 7 | 68.17 | 114.84 | 35.14 | 92.36 | 91.15 |
| 8 | 74.00 | 131.53 | 44.31 | 99.74 | 92.11 |
| 9 | 74.17 | 119.49 | 59.92 | 100.97 | 87.29 |
| 10 | 181.14 | 405.94 | 153.85 | 188.58 | 127.17 |
| 11 | 105.13 | 163.6 | 50.98 | 116.16 | 117.28 |
| 12 | 70.54 | 116.88 | 43.47 | 96.23 | 86.70 |
| 13 | 69.51 | 115.16 | 39.85 | 94.23 | 89.18 |
| 14 | 74.22 | 119.4 | 41.34 | 99.75 | 95.17 |
| 15 | 72.78 | 122.55 | 43.23 | 94.69 | 89.43 |
| 16 | 90.59 | 142.12 | 44.04 | 101.38 | 95.72 |
| 17 | 72.01 | 119.04 | 39.25 | 93.33 | 88.66 |
| 18 | 72.76 | 121.61 | 41.28 | 96.34 | 95.51 |
| 19 | 82.15 | 152.78 | 51.35 | 103.13 | 97.33 |
| 20 | 81.85 | 140.45 | 49.78 | 99.96 | 93.84 |
| Quantification of Cytokines (pg/mL) | |||||
|---|---|---|---|---|---|
| Patient ID | IL-2 | IL-6 | IL-10 | TNF-α | IFN-γ |
| 1 | 15.86 | 30.41 | 20.02 | ND | 20.01 |
| 2 | ND | 60.08 | 40.18 | 6.17 | 12.16 |
| 3 | 20.08 | ND | ND | 20.13 | ND |
| 4 | ND | 40.15 | ND | ND | ND |
| 5 | ND | ND | 18.29 | ND | 3.73 |
| 6 | 30.26 | 20.18 | 13.21 | 2.26 | 25.42 |
| 7 | 40.05 | 8.17 | ND | ND | ND |
| 9 | ND | ND | ND | ND | 5.05 |
| 10 | 8.13 | ND | ND | 3.15 | ND |
| SW Normality Test | CR (IG) | HR (IG) | CR (CG) | HR (CG) |
|---|---|---|---|---|
| n | 20 | 20 | 9 | 9 |
| M | 281,143 | 193,226 | 23,606 | 27,854 |
| SD | 54,272 | 87,916 | 20,468 | 33,054 |
| W | 0.494 | 0.402 | 0.908 | 0.840 |
| p | 0.0057 | 0.0046 | 0.3626 | 0.0693 |
| Mann–Whitney U Test | Cellular Response (Infected Group) | Cellular Response (Control Group) |
|---|---|---|
| Sample size | 20 | 9 |
| Sum of ranks (Ri) | 390 | 45 |
| Median | 265.66 | 18.33 |
| U | 0 | - |
| z (U) | 42.426 | - |
| p (one-tailed) | <0.0001 | - |
| p (two-tailed) | <0.0001 | - |
| Mann–Whitney U Test | Humoral Response (Infected Group) | Humoral Response (Control Group) |
|---|---|---|
| Sample size | 20 | 9 |
| Sum of ranks (Ri) | 390 | 45 |
| Median | 169.03 | 18.29 |
| U | 0 | - |
| z (U) | 42.426 | - |
| p (one-tailed) | <0.0001 | - |
| p (two-tailed) | <0.0001 | - |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Moreira, A.L.E.; Silva, P.A.N.d.; Gomes, R.S.; Santos, M.d.O.; Ito, C.R.M.; Barbosa, L.C.G.; Souza, P.P.d.; Peixoto, F.A.d.O.; Wastowski, I.J.; Carneiro, L.C.; et al. Characterization of T Helper 1 and 2 Cytokine Profiles in Newborns of Mothers with COVID-19. Biomedicines 2023, 11, 910. https://doi.org/10.3390/biomedicines11030910
Moreira ALE, Silva PANd, Gomes RS, Santos MdO, Ito CRM, Barbosa LCG, Souza PPd, Peixoto FAdO, Wastowski IJ, Carneiro LC, et al. Characterization of T Helper 1 and 2 Cytokine Profiles in Newborns of Mothers with COVID-19. Biomedicines. 2023; 11(3):910. https://doi.org/10.3390/biomedicines11030910
Chicago/Turabian StyleMoreira, André Luís Elias, Paulo Alex Neves da Silva, Rodrigo Saar Gomes, Mônica de Oliveira Santos, Célia Regina Malveste Ito, Lucas Candido Gonçalves Barbosa, Paula Pires de Souza, Fernanda Aparecida de Oliveira Peixoto, Isabela Jubé Wastowski, Lilian Carla Carneiro, and et al. 2023. "Characterization of T Helper 1 and 2 Cytokine Profiles in Newborns of Mothers with COVID-19" Biomedicines 11, no. 3: 910. https://doi.org/10.3390/biomedicines11030910
APA StyleMoreira, A. L. E., Silva, P. A. N. d., Gomes, R. S., Santos, M. d. O., Ito, C. R. M., Barbosa, L. C. G., Souza, P. P. d., Peixoto, F. A. d. O., Wastowski, I. J., Carneiro, L. C., & Avelino, M. A. G. (2023). Characterization of T Helper 1 and 2 Cytokine Profiles in Newborns of Mothers with COVID-19. Biomedicines, 11(3), 910. https://doi.org/10.3390/biomedicines11030910







