Nanomechanical and Morphological AFM Mapping of Normal Tissues and Tumors on Live Brain Slices Using Specially Designed Embedding Matrix and Laser-Shaped Cantilevers
Abstract
1. Introduction
2. Materials and Methods
2.1. Tissue Sample Preparation
2.2. Embedding Matrix
2.3. Vibratome Sectioning of Brain Samples
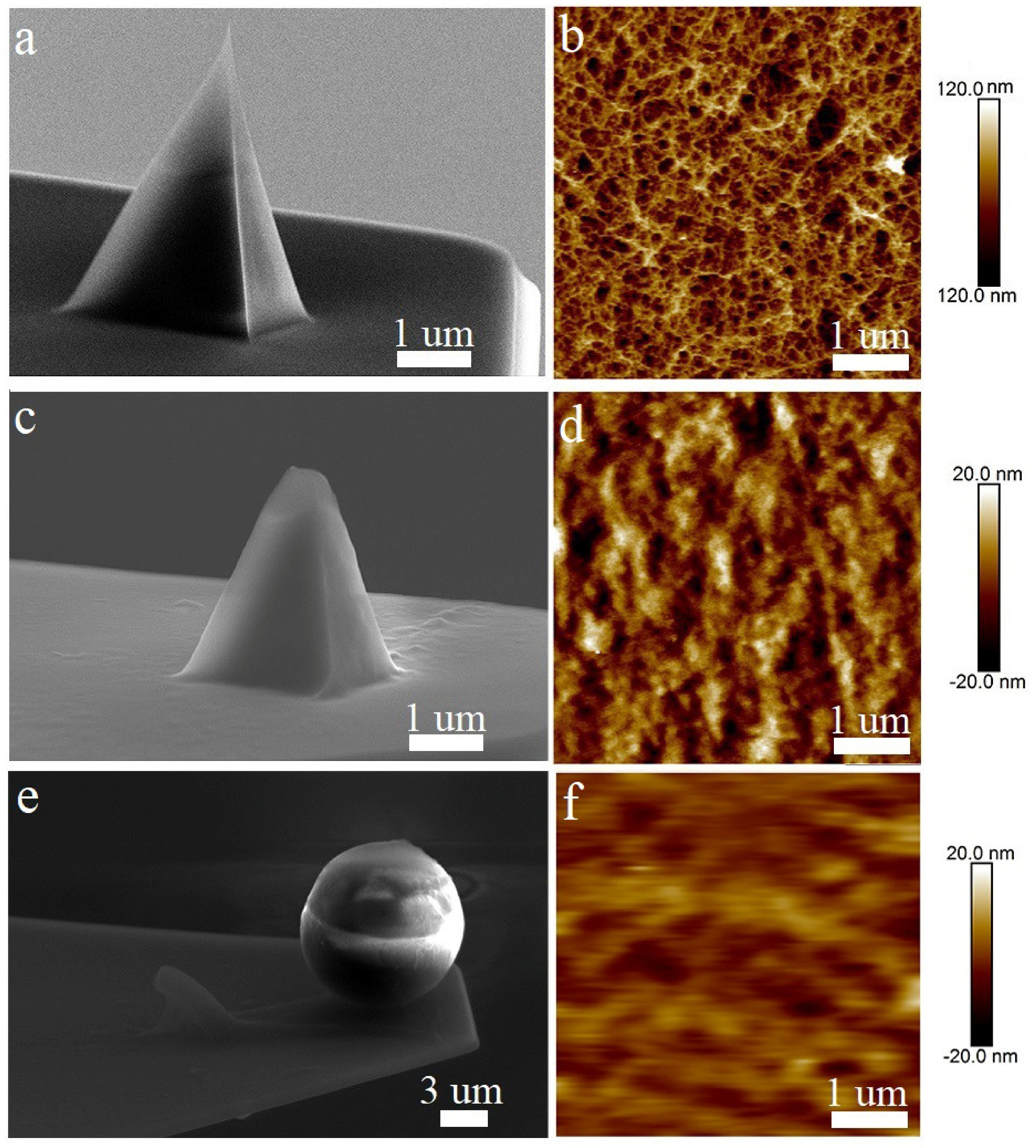
2.4. Modification and Evaluation of AFM Probes
2.5. Atomic Force Microscopy
2.6. Laser Scanning Microscopy of Live Slices Stained with FDA-PI Fluorescent Tracers
2.7. Immunohistochemical Analysis Combined with AFM
3. Results
3.1. Obtaining Live Slices of Brain Specimens Embedded in the Matrix
3.2. AFM Probe Characterisation
3.3. Combined AFM and LSM Studies of Brain Live Slices
3.4. Quick Mapping and Characterization of Live Tissue Nanomechanics
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Migliorini, E.; Ban, J.; Grenci, G.; Andolfi, L.; Pozzato, A.; Tormen, M.; Torre, V.; Lazzarino, M. Nanomechanics controls neuronal precursors adhesion and differentiation. Biotechnol. Bioeng. 2013, 110, 2301–2310. [Google Scholar] [CrossRef] [PubMed]
- Lv, H.; Li, L.; Sun, M.; Zhang, Y.; Chen, L.; Rong, Y.; Li, Y. Mechanism of regulation of stem cell differentiation by matrix stiffness. Stem Cell Res. Ther. 2015, 6, 103. [Google Scholar] [CrossRef] [PubMed]
- Belousov, A.; Titov, S.; Shved, N.; Garbuz, M.; Malykin, G.; Gulaia, V.; Kagansky, A.; Kumeiko, V. The Extracellular Matrix and Biocompatible Materials in Glioblastoma Treatment. Front. Bioeng. Biotechnol. 2019, 7, 341. [Google Scholar] [CrossRef] [PubMed]
- Huml, M.; Silye, R.; Zauner, G.; Hutterer, S.; Schilcher, K. Brain tumor classification using AFM in combination with data mining techniques. Biomed Res. Int. 2013, 2013, 176519. [Google Scholar] [CrossRef] [PubMed]
- Yallapu, M.M.; Katti, K.S.; Katti, D.R.; Mishra, S.R.; Khan, S.; Jaggi, M.; Chauhan, S.C. The roles of cellular nanomechanics in cancer. Med. Res. Rev. 2015, 35, 198–223. [Google Scholar] [CrossRef] [PubMed]
- Yilmaz, M.; Christofori, G. Mechanisms of motility in metastasizing cells. Mol. Cancer Res. 2010, 8, 629–642. [Google Scholar] [CrossRef] [PubMed]
- Huttenlocher, A.; Horwitz, A.R. Integrins in Cell Migration. Cold Spring Harb. Perspect. Biol. 2011, 3, a005074. [Google Scholar] [CrossRef]
- Grigorieva, E.V. Radiation Effects on Brain Extracellular Matrix. Front. Oncol. 2020, 10, 576701. [Google Scholar] [CrossRef]
- Louca, M.; Stylianou, A.; Minia, A.; Pliaka, V.; Alexopoulos, L.G.; Gkretsi, V.; Stylianopoulos, T. Ras suppressor-1 (RSU-1) promotes cell invasion in aggressive glioma cells and inhibits it in non-aggressive cells through STAT6 phospho-regulation. Sci. Rep. 2019, 9, 7782. [Google Scholar] [CrossRef]
- Walter, C.; Crawford, L.; Lai, M.; Toonen, J.A.; Pan, Y.; Sakiyama-Elbert, S.; Gutmann, D.H.; Pathak, A. Increased Tissue Stiffness in Tumors from Mice with Neurofibromatosis-1 Optic Glioma. Biophys. J. 2017, 112, 1535–1538. [Google Scholar] [CrossRef]
- Stewart, D.C.; Rubiano, A.; Dyson, K.; Simmons, C.S. Mechanical characterization of human brain tumors from patients and comparison to potential surgical phantoms. PLoS ONE 2017, 12, e0177561. [Google Scholar] [CrossRef] [PubMed]
- Yubero, M.L.; Kosaka, P.M.; San Paulo, Á.; Malumbres, M.; Calleja, M.; Tamayo, J. Effects of energy metabolism on the mechanical properties of breast cancer cells. Commun. Biol. 2020, 3, 590. [Google Scholar] [CrossRef] [PubMed]
- Stylianou, A.; Gkretsi, V.; Stylianopoulos, T. Transforming growth factor-β modulates pancreatic cancer associated fibroblasts cell shape, stiffness and invasion. Biochim. Biophys. Acta Gen. Subj. 2018, 1862, 1537–1546. [Google Scholar] [CrossRef]
- Alibert, C.; Goud, B.; Manneville, J.-B. Are cancer cells really softer than normal cells? Biol. Cell 2017, 109, 167–189. [Google Scholar] [CrossRef]
- Smith, J.R.; Maherally, Z.; Ghoneim, M.K.; Dickson, J.L.; An, Q.; Fillmore, H.L.; Pilkington, G.J. P48AFM Stiffness Measurements Of Glioma Cells And Cytoskeletal Protein Analysis Following CD44 Knockdown: Implications For Glioma Cell Invasion. Neuro-Oncology 2014, 16 (Suppl. 6), vi8. [Google Scholar] [CrossRef][Green Version]
- Miletić, M.; Aškrabić, S.; Rüger, J.; Vasić, B.; Korićanac, L.; Mondol, A.S.; Dellith, J.; Popp, J.; Schie, I.W.; Dohčević-Mitrović, Z. Combined Raman and AFM detection of changes in HeLa cervical cancer cells induced by CeO2 nanoparticles—Molecular and morphological perspectives. Analyst 2020, 145, 3983–3995. [Google Scholar] [CrossRef] [PubMed]
- Mikulska-Ruminska, K.; Kulik, A.J.; Benadiba, C.; Bahar, I.; Dietler, G.; Nowak, W. Nanomechanics of multidomain neuronal cell adhesion protein contactin revealed by single molecule AFM and SMD. Sci. Rep. 2017, 7, 8852. [Google Scholar] [CrossRef]
- Chtcheglova, L.A.; Hinterdorfer, P. Simultaneous AFM topography and recognition imaging at the plasma membrane of mammalian cells. Semin. Cell Dev. Biol. 2018, 73, 45–56. [Google Scholar] [CrossRef]
- Viji Babu, P.K.; Radmacher, M. Mechanics of Brain Tissues Studied by Atomic Force Microscopy: A Perspective. Front. Neurosci. 2019, 13, 600. [Google Scholar] [CrossRef]
- Stolz, M.; Raiteri, R.; Daniels, A.U.; VanLandingham, M.R.; Baschong, W.; Aebi, U. Dynamic elastic modulus of porcine articular cartilage determined at two different levels of tissue organization by indentation-type atomic force microscopy. Biophys. J. 2004, 86, 3269–3283. [Google Scholar] [CrossRef]
- Menal, M.J.; Jorba, I.; Torres, M.; Montserrat, J.M.; Gozal, D.; Colell, A.; Piñol-Ripoll, G.; Navajas, D.; Almendros, I.; Farré, R. Alzheimer’s Disease Mutant Mice Exhibit Reduced Brain Tissue Stiffness Compared to Wild-type Mice in both Normoxia and following Intermittent Hypoxia Mimicking Sleep Apnea. Front. Neurol. 2018, 9, 1. [Google Scholar] [CrossRef] [PubMed]
- Holtzmann, K.; Gautier, H.O.B.; Christ, A.F.; Guck, J.; Káradóttir, R.T.; Franze, K. Brain tissue stiffness is a sensitive marker for acidosis. J. Neurosci. Methods 2016, 271, 50–54. [Google Scholar] [CrossRef]
- Moeendarbary, E.; Weber, I.P.; Sheridan, G.K.; Koser, D.E.; Soleman, S.; Haenzi, B.; Bradbury, E.J.; Fawcett, J.; Franze, K. The soft mechanical signature of glial scars in the central nervous system. Nat. Commun. 2017, 8, 14787. [Google Scholar] [CrossRef]
- Smolyakov, G.; Dague, E.; Roux, C.; Seguelas, M.H.; Galés, C.; Senard, J.M.; Arvanitis, D.N. Nanoscale structural mapping as a measure of maturation in the murine frontal cortex. Brain Struct. Funct. 2018, 223, 255–265. [Google Scholar] [CrossRef] [PubMed]
- Andolfi, L.; Bourkoula, E.; Migliorini, E.; Palma, A.; Pucer, A.; Skrap, M.; Scoles, G.; Beltrami, A.P.; Cesselli, D.; Lazzarino, M. Investigation of Adhesion and Mechanical Properties of Human Glioma Cells by Single Cell Force Spectroscopy and Atomic Force Microscopy. PLoS ONE 2014, 9, e112582. [Google Scholar] [CrossRef] [PubMed]
- Roy, R.; Chen, W.; Goodell, L.A.; Jun, H.; Foran, D.J.; Desai, J.P. Microarray-facilitated mechanical characterization of breast tissue pathology samples using contact-mode Atomic Force Microscopy (AFM). In Proceedings of the 2010 3rd IEEE RAS & EMBS International Conference on Biomedical Robotics and Biomechatronics, Tokyo, Japan, 26–29 September 2010; pp. 710–715. [Google Scholar]
- Graham, H.K.; Hodson, N.W.; Hoyland, J.A.; Millward-Sadler, S.J.; Garrod, D.; Scothern, A.; Griffiths, C.E.M.; Watson, R.E.B.; Cox, T.R.; Erler, J.T.; et al. Tissue section AFM: In situ ultrastructural imaging of native biomolecules. Matrix Biol. 2010, 29, 254–260. [Google Scholar] [CrossRef]
- Efremov, Y.M.; Pukhlyakova, E.A.; Bagrov, D.V.; Shaitan, K.V. Atomic force microscopy of living and fixed Xenopus laevis embryos. Micron 2011, 42, 840–852. [Google Scholar] [CrossRef]
- Ciasca, G.; Sassun, T.E.; Minelli, E.; Antonelli, M.; Papi, M.; Santoro, A.; Giangaspero, F.; Delfini, R.; De Spirito, M. Nano-mechanical signature of brain tumours. Nanoscale 2016, 8, 19629–19643. [Google Scholar] [CrossRef]
- Deptuła, P.; Łysik, D.; Pogoda, K.; Cieśluk, M.; Namiot, A.; Mystkowska, J.; Król, G.; Głuszek, S.; Janmey, P.A.; Bucki, R. Tissue Rheology as a Possible Complementary Procedure to Advance Histological Diagnosis of Colon Cancer. ACS Biomater. Sci. Eng. 2020, 6, 5620–5631. [Google Scholar] [CrossRef]
- Antonovaite, N.; Beekmans, S.V.; Hol, E.M.; Wadman, W.J.; Iannuzzi, D. Regional variations in stiffness in live mouse brain tissue determined by depth-controlled indentation mapping. Sci. Rep. 2018, 8, 12517. [Google Scholar] [CrossRef]
- Aouassa, M.; Mitsai, E.; Syubaev, S.; Pavlov, D.; Zhizhchenko, A.; Jadli, I.; Hassayoun, L.; Zograf, G.; Makarov, S.; Kuchmizhak, A. Temperature-feedback direct laser reshaping of silicon nanostructures. Appl. Phys. Lett. 2017, 111, 243103. [Google Scholar] [CrossRef]
- Bouchonville, N.; Meyer, M.; Gaude, C.; Gay, E.; Ratel, D.; Nicolas, A. AFM mapping of the elastic properties of brain tissue reveals kPa μm−1 gradients of rigidity. Soft Matter 2016, 12, 6232–6239. [Google Scholar] [CrossRef] [PubMed]
- Finan, J.D.; Elkin, B.S.; Pearson, E.M.; Kalbian, I.L.; Morrison, B., 3rd. Viscoelastic properties of the rat brain in the sagittal plane: Effects of anatomical structure and age. Ann. Biomed. Eng. 2012, 40, 70–78. [Google Scholar] [CrossRef]
- Tao, B.; Song, Y.; Wu, Y.; Yang, X.; Peng, T.; Peng, L.; Xia, K.; Xia, X.; Chen, L.; Zhong, C. Matrix stiffness promotes glioma cell stemness by activating BCL9L/Wnt/β-catenin signaling. Aging 2021, 13, 5284–5296. [Google Scholar] [CrossRef]
- Tsitlakidis, A.; Tsingotjidou, A.S.; Kritis, A.; Cheva, A.; Selviaridis, P.; Aifantis, E.C.; Foroglou, N. Atomic Force Microscope Nanoindentation Analysis of Diffuse Astrocytic Tumor Elasticity: Relation with Tumor Histopathology. Cancers 2021, 13, 4539. [Google Scholar] [CrossRef] [PubMed]
- Lekka, M.; Gil, D.; Pogoda, K.; Dulińska-Litewka, J.; Jach, R.; Gostek, J.; Klymenko, O.; Prauzner-Bechcicki, S.; Stachura, Z.; Wiltowska-Zuber, J.; et al. Cancer cell detection in tissue sections using AFM. Arch. Biochem. Biophys. 2012, 518, 151–156. [Google Scholar] [CrossRef]
- Takano, T.; He, W.; Han, X.; Wang, F.; Xu, Q.; Wang, X.; Oberheim Bush, N.A.; Cruz, N.; Dienel, G.A.; Nedergaard, M. Rapid manifestation of reactive astrogliosis in acute hippocampal brain slices. Glia 2014, 62, 78–95. [Google Scholar] [CrossRef]
- Sallee, C.J.; Russell, D.F. Embedding of neural tissue in agarose or glyoxyl agarose for vibratome sectioning. Biotech. Histochem. 1993, 68, 360–368. [Google Scholar] [CrossRef]
- Hayaran, A.; Bijlani, V. Polyacrylamide as an infiltrating and embedding medium for vibratome sectioning of human fetal cerebellum containing Dil-filled axons. J. Neurosci. Methods 1992, 42, 65–68. [Google Scholar] [CrossRef]
- Grunow, B.; Kirchhoff, T.; Lange, T.; Moritz, T.; Harzsch, S. Histochemistry on vibratome sections of fish tissue: A comparison of fixation and embedding methods. Aquat. Biol. 2015, 23, 251–263. [Google Scholar] [CrossRef]
- Nayak, K.K.; Gupta, P. In vitro biocompatibility study of keratin/agar scaffold for tissue engineering. Int. J. Biol. Macromol. 2015, 81, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Selestina, G.; Vanja, K. Collagen- vs. Gelatine-Based Biomaterials and Their Biocompatibility: Review and Perspectives. In Biomaterials: Applications for Nanomedicine; Pignatello, R., Ed.; IntechOpen: London, UK, 2011; pp. 17–40. [Google Scholar]
- Smiley, J.F.; Bleiwas, C. Embedding matrix for simultaneous processing of multiple histological samples. J. Neurosci. Methods 2012, 209, 195–198. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Shen, Y.; Schmidt, T.; Diz-Muñoz, A. Protocol on Tissue Preparation and Measurement of Tumor Stiffness in Primary and Metastatic Colorectal Cancer Samples with an Atomic Force Microscope. STAR Protoc. 2020, 1, 100167. [Google Scholar] [CrossRef] [PubMed]
- Vanický, I.; Blaško, J.; Končeková, J.; Dzurjašková, Z.; Michalová, Z.; Székiová, E. Formaldehyde–hardened albumin as a non-penetrating embedding matrix for frozen and vibratome sectioning. Acta Histochem. 2022, 124, 151838. [Google Scholar] [CrossRef]
- Jones, M.V.; Calabresi, P.A. Agar-gelatin for embedding tissues prior to paraffin processing. BioTechniques 2007, 42, 569–570. [Google Scholar] [CrossRef]
- Dos Santos Riccardi, C.; Kranz, C.; Kowalik, J.; Yamanaka, H.; Mizaikoff, B.; Josowicz, M. Label-free DNA detection of hepatitis C virus based on modified conducting polypyrrole films at microelectrodes and atomic force microscopy tip-integrated electrodes. Anal. Chem. 2008, 80, 237–245. [Google Scholar] [CrossRef]
- Stylianou, A.; Lekka, M.; Stylianopoulos, T. AFM assessing of nanomechanical fingerprints for cancer early diagnosis and classification: From single cell to tissue level. Nanoscale 2018, 10, 20930–20945. [Google Scholar] [CrossRef]
- Pepin, K.M.; McGee, K.P.; Arani, A.; Lake, D.S.; Glaser, K.J.; Manduca, A.; Parney, I.F.; Ehman, R.L.; Huston, J., 3rd. MR Elastography Analysis of Glioma Stiffness and IDH1-Mutation Status. AJNR. Am. J. Neuroradiol. 2018, 39, 31–36. [Google Scholar] [CrossRef]
- Canovic, E.P.; Qing, B.; Mijailovic, A.S.; Jagielska, A.; Whitfield, M.J.; Kelly, E.; Turner, D.; Sahin, M.; Van Vliet, K.J. Characterizing Multiscale Mechanical Properties of Brain Tissue Using Atomic Force Microscopy, Impact Indentation, and Rheometry. J. Vis. Exp. 2016, 115, e54201. [Google Scholar] [CrossRef]
- Vappou, J.; Breton, E.; Choquet, P.; Goetz, C.; Willinger, R.; Constantinesco, A. Magnetic resonance elastography compared with rotational rheometry for in vitro brain tissue viscoelasticity measurement. Magn. Reson. Mater. Phys. Biol. Med. 2007, 20, 273. [Google Scholar] [CrossRef]
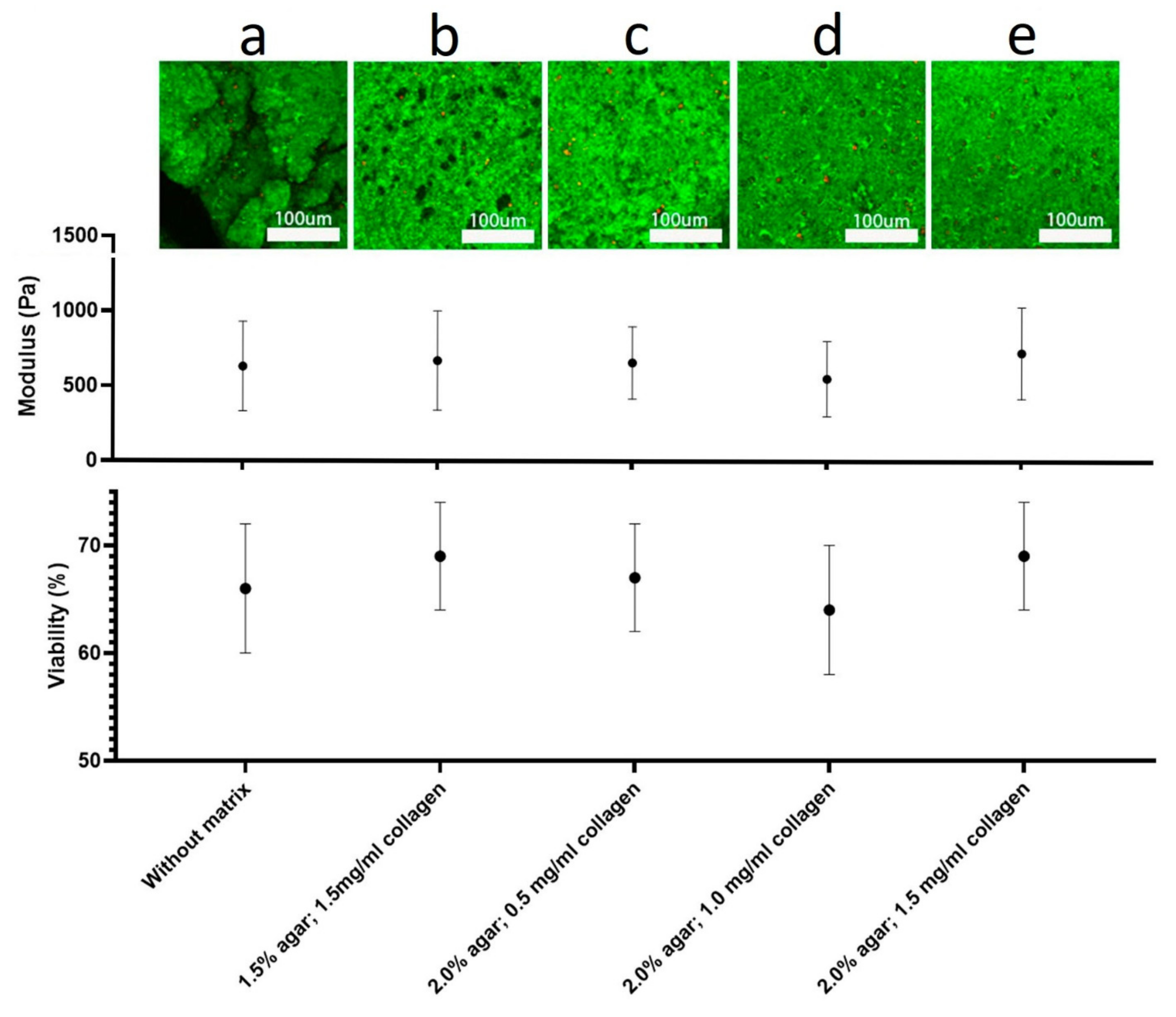
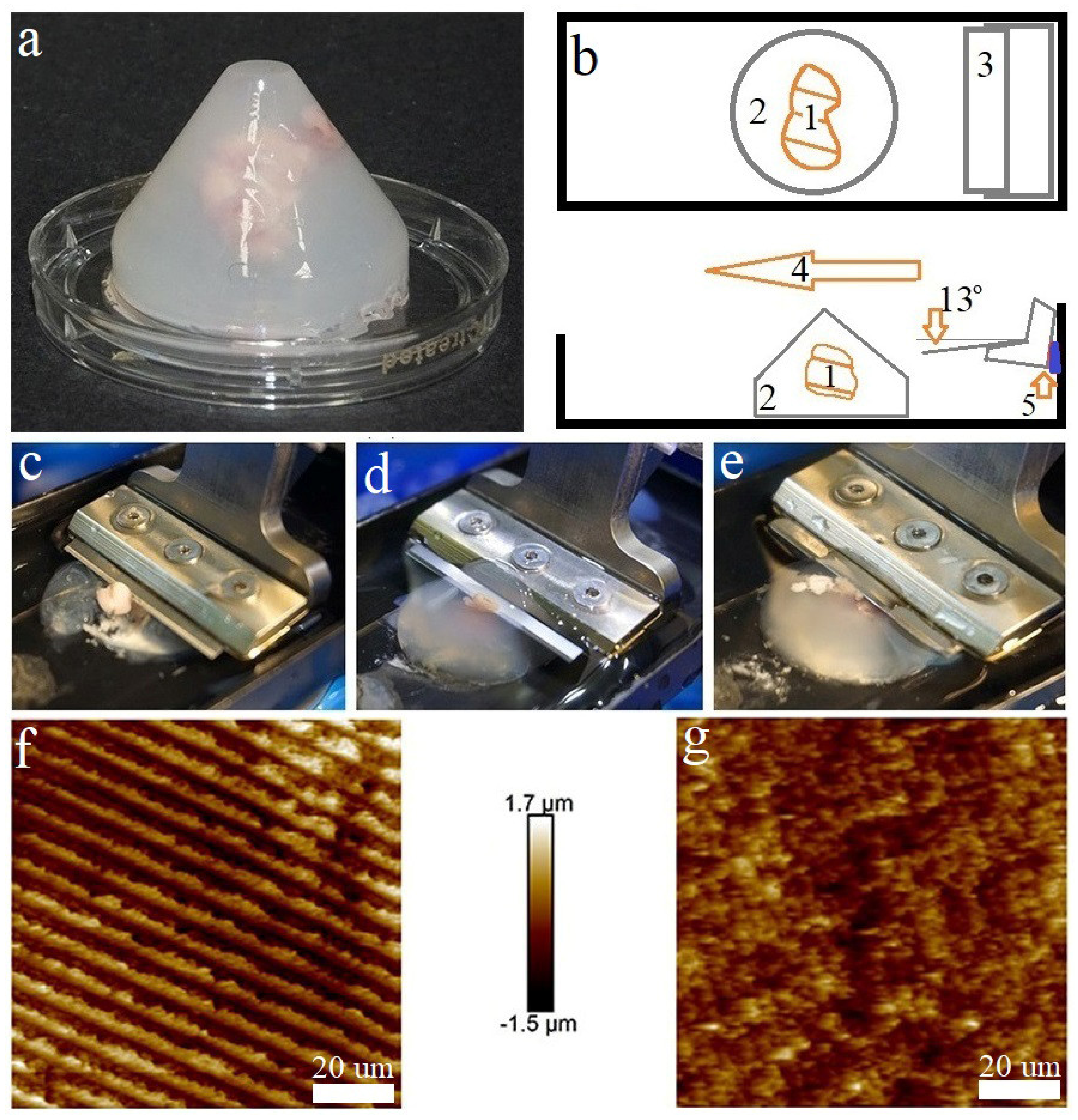

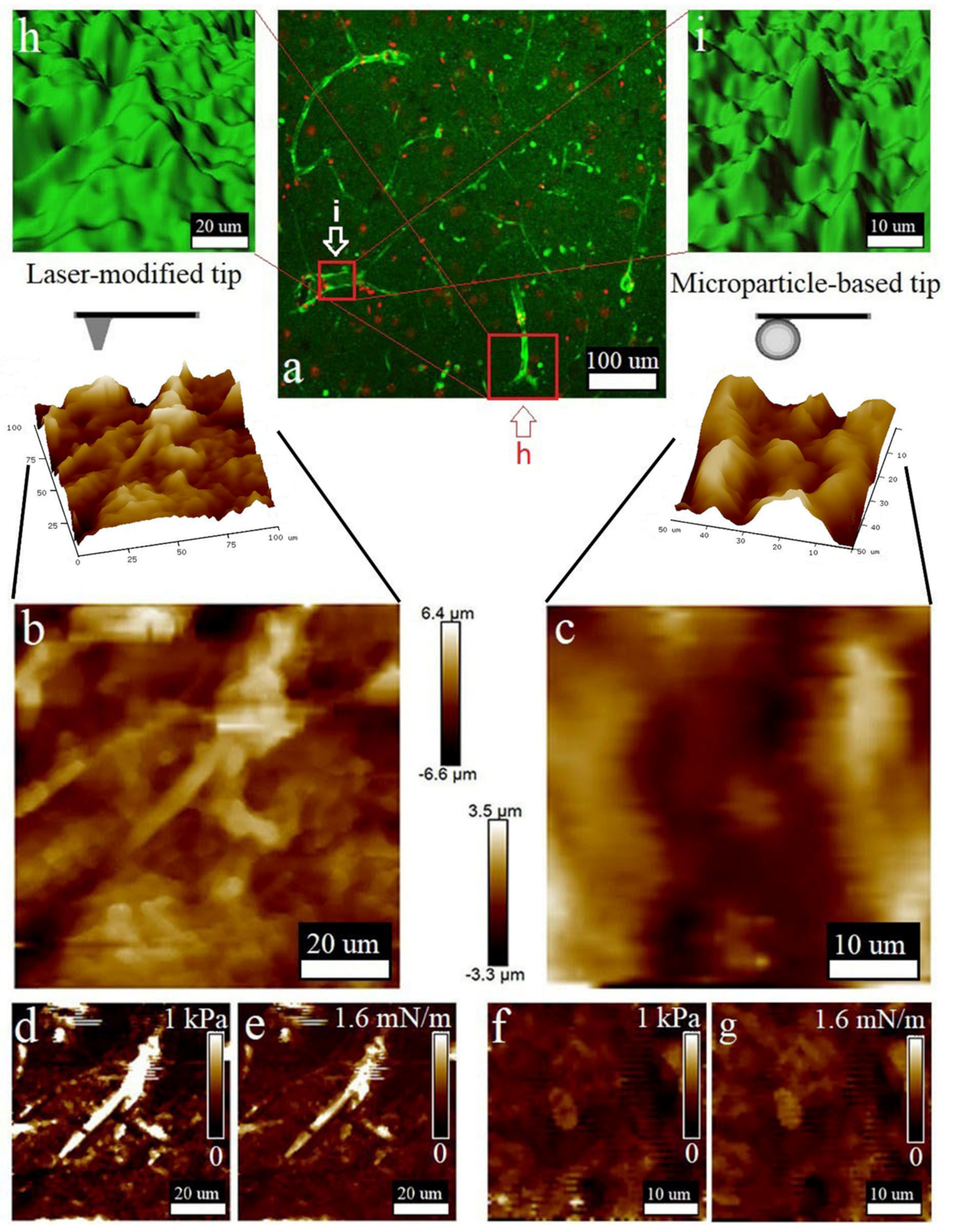
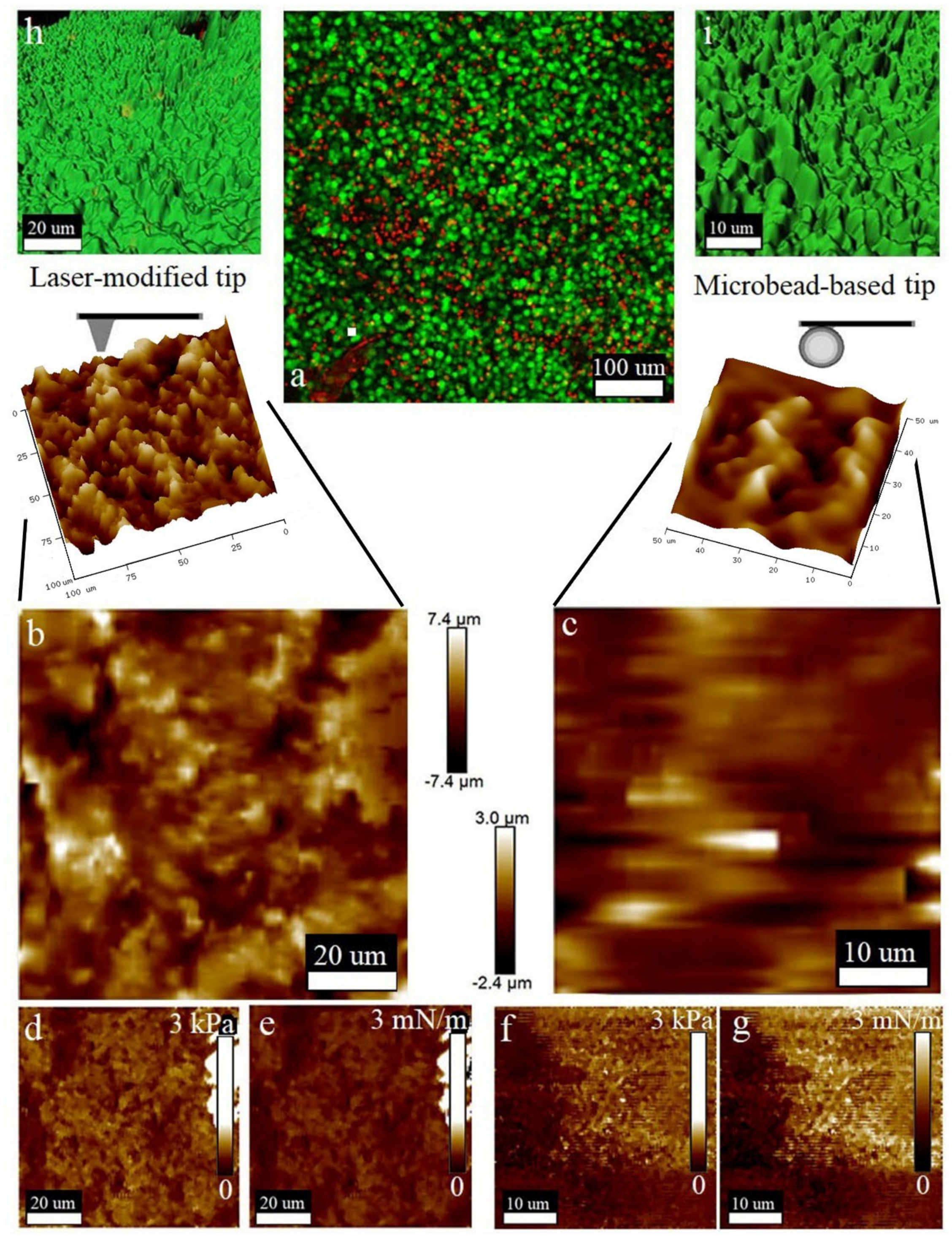
| Variable Components | Stiffness, µN/m | Elasticity, kPa |
|---|---|---|
| 0.5% agar 0.5 mg/mL collagen | 0.37 ± 0.03 | 02.83 ± 0.09 |
| 1.0% agar; 0.5 mg/mL collagen | 0.8 ± 0.05 | 03.89 ± 0.22 |
| 0.5% agar; 1.0 mg/mL collagen | 0.8 ± 0.05 | 04.32 ± 0.26 |
| 0.5% agar | 0.64 ± 0.05 | 06.23 ± 0.27 |
| 0.5% agar; 1.5 mg/mL collagen | 1.19 ± 0.07 | 07.63 ± 0.24 |
| 1.0% agar | 1.05 ± 0.06 | 07.96 ± 0.36 |
| 1.5% agar | 1.74 ± 0.09 | 09.38 ± 0.38 |
| 1.0% agar; 1.5 mg/mL collagen | 1.68 ± 0.09 | 11.93 ± 0.32 |
| 1.0% agar; 1.0 mg/mL collagen | 2.07 ± 0.1 | 13.97 ± 0.337 |
| 1.5% agar; 0.5 mg/mL collagen | 2.26 ± 0.08 | 16.16 ± 0.35 |
| 1.5% agar; 1.0 mg/mL collagen | 2.68 ± 0.09 | 20.86 ± 0.65 |
| 2.0% agar | 2.69 ± 0.17 | 21.35 ± 0.67 |
| 2.5% agar | 3.41 ± 0.18 | 24.15 ± 0.71 |
| 1.5% agar; 1.5 mg/mL collagen * | 2.58 ± 0.1 | 27.62 ± 0.85 |
| 2.0% agar; 0.5 mg/mL collagen * | 2.93 ± 0.1 | 27.46 ± 0.7 |
| 2.0% agar; 1.0 mg/mL collagen * | 3.21 ± 0.17 | 27.62 ± 0.73 |
| 2.0% agar; 1.5 mg/mL collagen * | 3.07 ± 0.18 | 27.82 ± 0.91 |
| 2.5% agar; 0.5 mg/mL collagen | 3.73 ± 0.18 | 35.59 ± 0.94 |
| 2.5% agar; 1.0 mg/mL collagen | 4.24 ± 0.26 | 46.38 ± 1.1 |
| 2.5% agar; 1.5 mg/mL collagen | 4.35 ± 0.29 | 46.59 ± 0.27 |
| Action Needed | Timing |
|---|---|
| Embedding matrix and mold preparation: Prepare: 1. 2.5–3.3% agar on 200 мM NaCl solution (A) 2. 2.5–7.5 mg/mL Collagen I on 30 мM acetic acid (B) 3. Gelation initiator (150 mM NaCl, 120 mM HEPES-NaOH, pH 7.4, 1.0 мM NaOH (C) 4. Prepare the embedding mold by cutting off the conical bottom of the 50 mL Falcon tube | Prepare and keep in advance |
| AFM tip preparation: Equip your AFM instrument with a fluid imaging probe that has a tip radius of about 400 nm. The originally modified probe can be prepared by laser ablation using the standard ScanAsyst Fluid High Resolution Probe. The tip radius of the cantilever can be increased via thermal reshaping under direct multi-pulse laser irradiation with sub-nanosecond laser pulses (a central wavelength of 533 nm, a pulse duration of 0.5 ns, a pulse repetition rate of 1.5 KHz, and an average power of 20 μW) generated by a passively Q-switched laser. To laterally confine the melting region, the laser radiation should be focused on the cantilever tip with a high numerical aperture (NA) objective of 0.9, which provides a full width at half the maximum of the focused beam of about 0.4 μm. | Prepare and keep in advance |
| Biopsy embedding: Melt the agar (A) in the microwave and put it in water bath at a temperature of 37 °C for 5 min. Select an appropriate biopsy fragment under a stereomicroscope and transfer this with tweezers into the ice-cold DMEM. Put 3 mL of melted agar (A) in the embedding mold, add 1 mL of collagen I (B), add 1 mL of gelation initiator (C), and mix all the components by gentle aspiration. Immediately place the piece of tissue inside the mold, submerging it by the matrix. Place it in the center and avoid tissue contact with mold wall. Put the mold in ice bath for 5 min. | 15 min |
| Embedding conemounting: Take the mold from ice bath. Turn the cone over and gently remove the cured embedding matrix. Stick the embedding matrix cone with cyanoacrylate glue onto a glass slide and fix it in the vibratome tray. | 5 min |
| Tissue vibrosectioning: Attach the tray to the vibratome. Set the blade angle to 13°, the vibration frequency to 50Hz, and the movement speed to 0.2 mm/sec. Make several test sections with a thickness of 50 to 500 µm. | 5 min |
| Tissue section mounting: Apply a drop of cyanoacrylate glue on Petri dish using a pipette and immediately remove the glue. A thin film of glue should remain on the surface of the dish. Using tweezers, place the tissue section on the glue. Wait for 2 min and add DNase solution prepared on PBS (1000 U/mL) to the dish for 5 min at room temperature. Remove DNase; add ice-cold DMEM. The tissue section is ready for AFM analysis. | 10 min |
| AFM Scanning: Calibrate the probe properly. Select the region of interest on the tissue section using an optical channel. Set it to PeakForce Quantitative Nanomechanical mapping mode and adjust the scan parameters to obtain a high-quality force curve. | ~20 min (depends on resolution) |
| Additional steps: This may include some other procedures with tissue slices, already mapped by AFM: live cell tracing, live/dead assay, tissue fixation, and immunostaining. | depends on type of procedure |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Farniev, V.M.; Shmelev, M.E.; Shved, N.A.; Gulaia, V.S.; Biktimirov, A.R.; Zhizhchenko, A.Y.; Kuchmizhak, A.A.; Kumeiko, V.V. Nanomechanical and Morphological AFM Mapping of Normal Tissues and Tumors on Live Brain Slices Using Specially Designed Embedding Matrix and Laser-Shaped Cantilevers. Biomedicines 2022, 10, 1742. https://doi.org/10.3390/biomedicines10071742
Farniev VM, Shmelev ME, Shved NA, Gulaia VS, Biktimirov AR, Zhizhchenko AY, Kuchmizhak AA, Kumeiko VV. Nanomechanical and Morphological AFM Mapping of Normal Tissues and Tumors on Live Brain Slices Using Specially Designed Embedding Matrix and Laser-Shaped Cantilevers. Biomedicines. 2022; 10(7):1742. https://doi.org/10.3390/biomedicines10071742
Chicago/Turabian StyleFarniev, Vladislav M., Mikhail E. Shmelev, Nikita A. Shved, Valeriia S. Gulaia, Arthur R. Biktimirov, Alexey Y. Zhizhchenko, Aleksandr A. Kuchmizhak, and Vadim V. Kumeiko. 2022. "Nanomechanical and Morphological AFM Mapping of Normal Tissues and Tumors on Live Brain Slices Using Specially Designed Embedding Matrix and Laser-Shaped Cantilevers" Biomedicines 10, no. 7: 1742. https://doi.org/10.3390/biomedicines10071742
APA StyleFarniev, V. M., Shmelev, M. E., Shved, N. A., Gulaia, V. S., Biktimirov, A. R., Zhizhchenko, A. Y., Kuchmizhak, A. A., & Kumeiko, V. V. (2022). Nanomechanical and Morphological AFM Mapping of Normal Tissues and Tumors on Live Brain Slices Using Specially Designed Embedding Matrix and Laser-Shaped Cantilevers. Biomedicines, 10(7), 1742. https://doi.org/10.3390/biomedicines10071742







