PARP Inhibitors: A New Horizon for Patients with Prostate Cancer
Abstract
1. Introduction
2. Rationale of Use of PARP Inhibitors in PCa
2.1. DNA Repair Pathway
2.2. Role of DNA Damage Repair Genes in Prostate Cancer
2.2.1. DDR Mutations in Prostate Cancer
2.2.2. Germline Mutations
2.2.3. Somatic Mutations
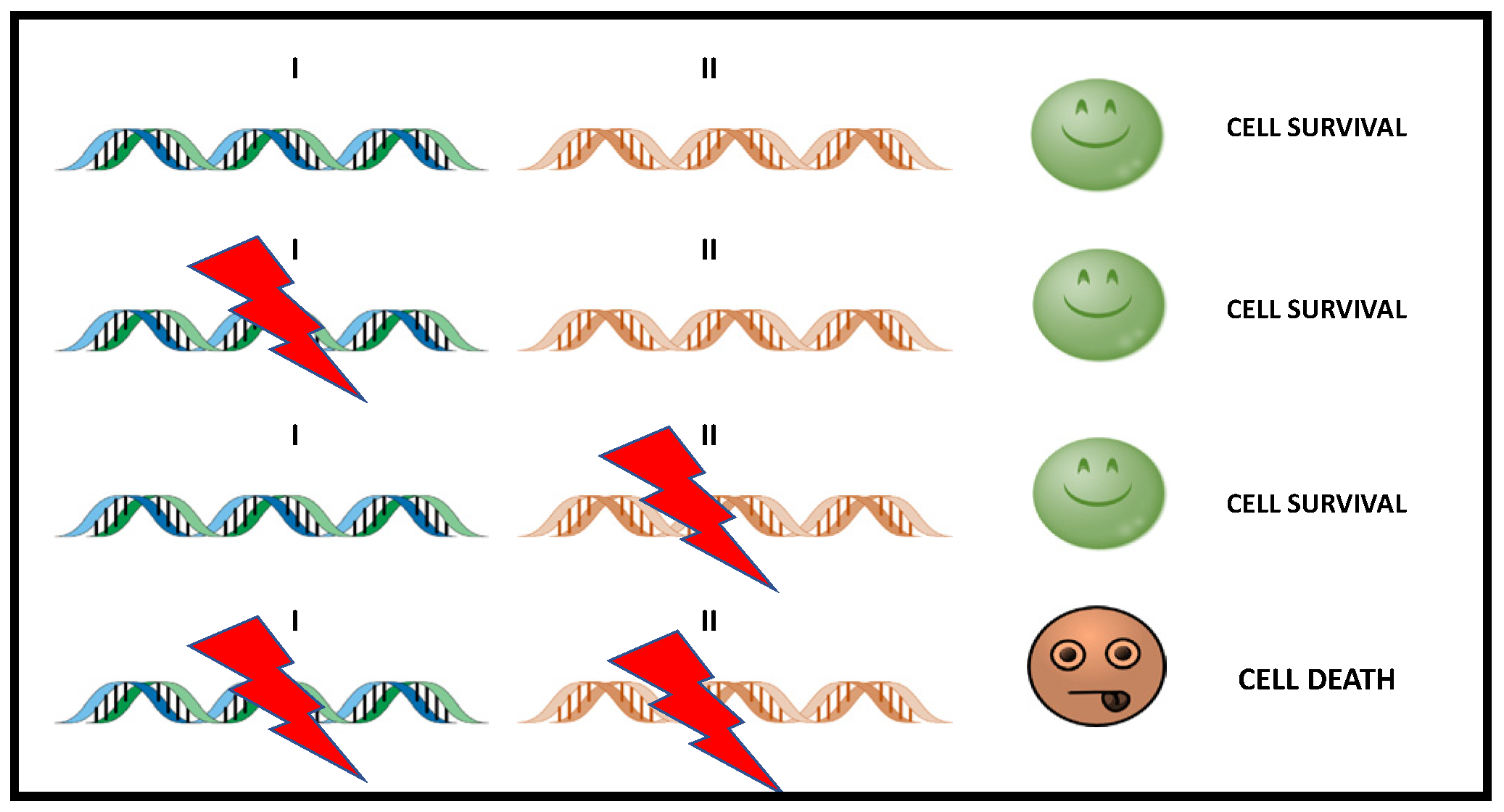
2.3. Mechanism of Action of PARP Inhibitors
3. Clinical Development of PARP Inhibitors in Prostate Cancer
3.1. PARPi in Monotherapy
3.1.1. OLAPARIB
3.1.2. RUCAPARIB
3.1.3. NIRAPARIB
3.1.4. TALAZOPARIB
3.2. Adverse Events and Tolerability
3.3. Mechanisms of Intrinsic and Acquired Resistance to PARP Inhibitors
- Resistance mechanisms that restore the pathway of PARP-independent homologous recombination. There are multiple pathways leading to the restoration of HR function, including:
- Minor mutations of the genes involved in HR (BRCA1/2, PALB2, RAD51C/D) that restore the ability to repair errors in DNA. This mechanism is also responsible for platinum resistance [62];
- Expression of hypomorphic variants BRCA1/2;
- Epigenetic changes in the genes that take part in HR, such as the promoter demethylation of BRCA1 and RAD51C genes [63]; and
- Resistance mechanisms independent of the HR pathway:
- Reduced expression of PARP enzymes, which contributes to the reduction of drug activity and reduction of PARP1 entrapment [64];
- Patients with reverse mutations in the BRCA1 gene exhibit MMEJ signals, suggesting the POLQ as a PARPi resistance vehicle. Thus, POLQ inhibitors can suppress resistance acquired from PARPi, conferring synthetic lethality in tumors with deficits in HR and NHEJ [67]; and
- Mutations in the poly (ADP-ribose)-glycohydrolase enzymes (PARG) can lead to resistance through a mechanism that does not restore HR. The loss of PARG results in the accumulation of chains PAR which, by not being degraded, maintain the activity of the enzymes.
3.4. PARPi in Combination
3.4.1. Combinations with Antiandrogen Therapy
- PARP promotes AR transcription, so inhibition of this pathway potentiates the antiandrogenic effect [70];
- ADT promotes PARP overexpression, increasing its sensitivity to PARP inhibitors; and
3.4.2. Combinations with Immune Checkpoint Inhibitors (ICI)
3.4.3. Combination with Anti-VEGF Therapies
3.4.4. Combination with AKT Inhibitors/ATR Inhibitors
3.4.5. Combination with DNA-Damaging and Other DDR-Targeting Agents
3.4.6. Combination with Radionuclides
3.5. PARPi and Radiotherapy
4. Mutation Testing.: Whom, When, and How
5. Conclusions and Future Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Dyba, T.; Randi, G.; Bray, F.; Martos, C.; Giusti, F.; Nicholson, N.; Gavin, A.; Flego, M.; Neamtiu, L.; Dimitrova, N.; et al. The European cancer burden in 2020: Incidence and mortality estimates for 40 countries and 25 major cancers. Eur. J. Cancer 2021, 157, 308–347. [Google Scholar] [CrossRef] [PubMed]
- Siegel, R.; Miller, K.D.; Fuchs, H.E.; Jemal, A. Cancer statistics, 2022. ACS J. 2022, 72, 7–33. [Google Scholar] [CrossRef] [PubMed]
- Mottet, N.; Van den Bergh, R.C.N.; Briers, E.; Expert Patient Advocate (European Prostate Cancer Coalition/Europa UOMO); De Santis, M.; Gillessen, S.; Grummet, J.; Henry, A.M.; van der Kwast, T.H.; Lam, T.B.; et al. AU-EANM-ESTRO-ESUR-ISUP-SIOG Guidelines on Prostate Cancer 2022. Available online: https://uroweb.org/guideline/prostate-cancer/ (accessed on 25 April 2022).
- Haffner, M.C.; Zwart, W.; Roudier, M.P.; True, L.D.; Nelson, W.G.; Epstein, J.I.; De Marzo, A.M.; Nelson, P.S.; Yegnasubramanian, S. Genomic and phenotypic heterogeneity in prostate cancer. Nat. Rev. Urol. 2021, 18, 79–92. [Google Scholar] [CrossRef] [PubMed]
- Tan, M.H.E.; Li, J.; Xu, H.E.; Melcher, K.; Yong, E.-L. Androgen receptor: Structure, role in prostate cancer and drug discovery. Acta Pharmacol. Sin. 2015, 36, 3–23. [Google Scholar] [CrossRef]
- Mollica, V.; Di Nunno, V.; Cimadamore, A.; Lopez-Beltran, A.; Cheng, L.; Santoni, M.; Scarpelli, M.; Montironi, R.; Massari, F. Molecular Mechanisms Related to Hormone Inhibition Resistance in Prostate Cancer. Cells 2019, 8, 43. [Google Scholar] [CrossRef]
- Leung, D.K.W.; Chiu, P.K.F.; Ng, C.-F.; Teoh, J.Y.C. Novel Strategies for Treating Castration-Resistant Prostate Cancer. Biomedicines 2021, 9, 339. [Google Scholar] [CrossRef]
- Cheng, H.H.; Sokolova, A.O.; Schaeffer, E.M.; Small, E.J.; Higano, C.S. Germline and Somatic Mutations in Prostate Cancer for the Clinician. J. Natl. Compr. Cancer Netw. 2019, 17, 515–521. [Google Scholar] [CrossRef]
- Henríquez, I.; Roach, M.; Morgan, T.M.; Bossi, A.; Gómez, J.A.; Abuchaibe, O.; Couñago, F. Current and Emerging Therapies for Metastatic Castration-Resistant Prostate Cancer (mCRPC). Biomedicines 2021, 9, 1247. [Google Scholar] [CrossRef]
- Lord, C.J.; Ashworth, A. The DNA damage response and cancer therapy. Nature 2012, 481, 287–294. [Google Scholar] [CrossRef]
- Lord, C.J.; Ashworth, A. PARP inhibitors: The First Synthetic Lethal Targeted Therapy. Science 2017, 355, 1152–1158. [Google Scholar] [CrossRef]
- Mirza, M.; Pignata, S.; Ledermann, J. Latest clinical evidence and further development of PARP inhibitors in ovarian cancer. Ann. Oncol. 2018, 29, 1366–1376. [Google Scholar] [CrossRef] [PubMed]
- Robinson, D.; Van Allen, E.M.; Wu, Y.M.; Schultz, N.; Lonigro, R.J.; Mosquera, J.; Montgomery, B.; Taplin, M.; Pritchard, C.C.; Attard, G.; et al. Integrative clinical genomics of advanced prostate cancer. Cell 2015, 161, 1215. [Google Scholar] [CrossRef] [PubMed]
- Pritchard, C.C.; Mateo, J.; Walsh, M.F.; De Sarkar, N.; Abida, W.; Beltran, H.; Garofalo, A.; Gulati, R.; Carreira, S.; Eeles, R.; et al. Inherited DNA-Repair Gene Mutations in Men with Metastatic Prostate Cancer. N. Engl. J. Med. 2016, 375, 443–453. [Google Scholar] [CrossRef] [PubMed]
- Nombela, P.; Lozano, R.; Aytes, A.; Mateo, J.; Olmos, D.; Castro, E. BRCA2 and Other DDR Genes in Prostate Cancer. Cancers 2019, 11, 352. [Google Scholar] [CrossRef]
- Abida, W.; Armenia, J.; Gopalan, A.; Brennan, R.; Walsh, M.; Barron, D.; Danila, D.; Rathkopf, D.; Morris, M.; Slovin, S.; et al. Prospective Genomic Profiling of Prostate Cancer Across Disease States Reveals Germline and Somatic Alterations That May Affect Clinical Decision Making. JCO Precis. Oncol. 2017, 1, 1–32. [Google Scholar] [CrossRef]
- Kim, E., Jr.; Kim, S.; Srivastava, A.; Saraiya, B.; Mayer, T.; Kim, W.J.; Kim, I.Y. Similar Incidence of DNA Damage Response Pathway Alterations Between Clinically Localized and Metastatic Prostate Cancer. BMC Urol. 2019, 19, 33. [Google Scholar] [CrossRef]
- Cancer Genome Atlas Research Network. The Molecular Taxonomy of Primary Prostate Cancer. Cell 2015, 163, 1011–1025. [Google Scholar] [CrossRef]
- Marshall, C.H.; Fu, W.; Wang, H.; Baras, A.S.; Lotan, T.; Antonarakis, E.S. Prevalence of DNA repair gene mutations in localized prostate cancer according to clinical and pathologic features: Association of Gleason score and tumor stage. Prostate Cancer Prostatic Dis. 2019, 22, 59–65. [Google Scholar] [CrossRef]
- Lang, S.H.; Swift, S.L.; White, H.; Misso, K.; Kleijnen, J.; Quek, R.G. A systematic review of the prevalence of DNA damage response gene mutations in prostate cancer. Int. J. Oncol. 2019, 55, 597–616. [Google Scholar] [CrossRef]
- Castro, E.; Goh, C.; Olmos, D.; Saunders, E.; Leongamornlert, D.; Tymrakiewicz, M.; Mahmud, N.; Dadaev, T.; Govindasami, K.; Guy, M.; et al. Germline BRCA mutations are associated with higher risk of nodal involvement, distant metastasis, and poor survival outcomes in prostate cancer. J. Clin. Oncol. 2013, 31, 1748–1757. [Google Scholar] [CrossRef]
- Kote-Jarai, Z.; Leongamornlert, D.; Saunders, E.; Tymrakiewicz, M.; Castro, E.; Mahmud, N.; Guy, M.; Edwards, S.; O’Brien, L.; Sawyer, E.; et al. BRCA2 is a moderate penetrance gene contributing to young-onset prostate cancer: Implications for genetic testing in prostate cancer patients. Br. J. Cancer 2011, 105, 1230–1234. [Google Scholar] [CrossRef] [PubMed]
- Carter, H.B.; Helfand, B.; Mamawala, M.; Wu, Y.; Landis, P.; Yu, H.; Wiley, K.; Na, R.; Shi, Z.; Petkewicz, J.; et al. Germline mutations in ATM and BRCA1/2 are associated with grade reclassification in men on active surveillance for prostate cancer. Eur Urol. 2019, 75, 743–749. [Google Scholar] [CrossRef] [PubMed]
- Castro, E.; Goh, C.; Leongamornlert, D.; Saunders, E.; Tymrakiewicz, M.; Dadaev, T.; Govindasami, K.; Guy, M.; Ellis, S.; Frost, D.; et al. Effect of BRCA Mutations on Metastatic Relapse and Cause-specific Survival After Radical Treatment for Localised Prostate Cancer. Eur. Urol. 2015, 68, 186–193. [Google Scholar] [CrossRef] [PubMed]
- Castro, E.; Romero-Laorden, N.; Del Pozo, A.; Lozano, R.; Medina, A.; Puente, J.; Piulats, J.M.; Lorente, D.; Saez, M.I.; Morales-Barrera, R.; et al. PROREPAIR-B: A Prospective Cohort Study of the Impact of Germline DNA Repair Mutations on the Outcomes of Patients With Metastatic Castration-Resistant Prostate Cancer. J. Clin. Oncol. 2019, 37, 490–503. [Google Scholar] [CrossRef] [PubMed]
- Nicolosi, P.; Ledet, E.; Yang, S.; Michalski, S.; Freschi, B.; O’Leary, E.; Esplin, E.D.; Nussbaum, R.L.; Sartor, O. Prevalence of Germline Variants in Prostate Cancer and Implications for Current Genetic Testing Guidelines. JAMA Oncol. 2019, 5, 523–528. [Google Scholar] [CrossRef] [PubMed]
- Koochekpour, S. Genetic and Epigenetic Changes in Human Prostate Cancer. Iran. Red Crescent Med. J. 2011, 13, 80–98. [Google Scholar]
- Beltran, H.; Yelensky, R.; Frampton, G.; Park, K.; Downing, S.R.; MacDonald, T.Y.; Jarosz, M.; Lipson, D.; Tagawa, S.T.; Nanus, D.M.; et al. Targeted Next-generation Sequencing of Advanced Prostate Cancer Identifies Potential Therapeutic Targets and Disease Heterogeneity. Eur. Urol. 2012, 63, 920–926. [Google Scholar] [CrossRef]
- Bramco, C.; Joana Paredes, J. PARP Inhibitors: From the Mechanism of Action to Clinical Practice. Acta Med. Port. 2022, 35, 135–143. [Google Scholar]
- Farmer, H.; McCabe, N.; Lord, C.J.; Tutt, A.N.J.; Johnson, D.A.; Richardson, T.B.; Santarosa, M.; Dillon, K.J.; Hickson, I.; Knights, C.; et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 2005, 434, 917–921. [Google Scholar] [CrossRef]
- Bryant, H.E.; Schultz, N.; Thomas, H.D.; Parker, K.M.; Flower, D.; Lopez, E.; Kyle, S.; Meuth, M.; Curtin, N.J.; Helleday, T. Specific killing of BRCA2-deficient tumours with inhibitors of poly(ADP-ribose) polymerase. Nature 2005, 434, 913–917. [Google Scholar] [CrossRef]
- King, M.C. “The race” to clone BRCA1. Science 2014, 343, 462–1465. [Google Scholar] [CrossRef] [PubMed]
- Ashworth, A.; Lord, C.J.; Reis-Filho, J.S. Genetic Interactions in Cancer Progression and Treatment. Cell 2011, 145, 30–38. [Google Scholar] [CrossRef] [PubMed]
- Murai, J.; Huang, S.-Y.N.; Das, B.B.; Renaud, A.; Zhang, Y.; Doroshow, J.H.; Ji, J.; Takeda, S.; Pommier, Y. Trapping of PARP1 and PARP2 by Clinical PARP Inhibitors. Cancer Res. 2012, 72, 5588–5599. [Google Scholar] [CrossRef] [PubMed]
- Murai, J.; Huang, S.-Y.N.; Renaud, A.; Zhang, Y.; Ji, J.; Takeda, S.; Morris, J.; Teicher, B.; Doroshow, J.H.; Pommier, Y. Stereospecific PARP Trapping by BMN 673 and Comparison with Olaparib and Rucaparib. Mol. Cancer Ther. 2014, 13, 433–443. [Google Scholar] [CrossRef]
- Pommier, Y.; O’Connor, M.J.; de Bono, J. Laying a trap to kill cancer cells: PARP inhibitors and their mechanisms of action. Sci. Transl. Med. 2016, 8, 362–379. [Google Scholar] [CrossRef]
- Mirza, M.R.; Monk, B.J.; Herrstedt, J.; Oza, A.M.; Mahner, S.; Redondo, A.; Fabbro, M.; Ledermann, J.A.; Lorusso, D.; Vergote, I.; et al. Niraparib Maintenance Therapy in Platinum-Sensitive, Recurrent Ovarian Cancer. N. Engl. J. Med. 2016, 375, 2154–2164. [Google Scholar] [CrossRef]
- González-Martín, A.; Pothuri, B.; Vergote, I.; DePont Christensen, R.; Graybill, W.; Mirza, M.R.; McCormick, C.; Lorusso, D.; Hoskins, P.; Freyer, G.; et al. Niraparib in Patients with Newly Diagnosed Advanced Ovarian Cancer. N. Engl. J. Med. 2019, 381, 2391–2402. [Google Scholar] [CrossRef]
- Díaz-Mejía, N.; García-Illescas, D.; Morales-Barrera, R.; Suarez, C.; Planas, J.; Maldonado, X.; Carles, J.; Mateo, J. PARP inhibitors in advanced prostate cancer: When to use them? Endocr.-Relat. Cancer 2021, 28, 79–93. [Google Scholar] [CrossRef]
- Yi, M.; Dong, B.; Qin, S.; Chu, Q.; Wu, K.; Luo, S. Advances and perspectives of PARP inhibitors. Exp. Hematol. Oncol. 2019, 8, 29–41. [Google Scholar] [CrossRef]
- Mateo, J.; Carreira, S.; Sandhu, S.; Miranda, S.; Mossop, H.; Perez-Lopez, R.; Nava Rodrigues, D.; Robinson, D.; Omlin, A.; Tunariu, N.; et al. DNA-Repair Defects and Olaparib in Metastatic Prostate Cancer. N. Engl. J. Med. 2015, 373, 1697–1708. [Google Scholar] [CrossRef]
- Mateo, J.; Porta, N.; Bianchini, D.; McGovern, U.; Elliott, T.; Jones, R.; Syndikus, I.; Ralph, C.; Jain, S.; Varughese, M.; et al. Olaparib in patients with metastatic castration-resistant prostate cancer with DNA repair gene aberrations (TOPARP-B): A multicentre, open-label, randomised, phase 2 trial. Lancet Oncol. 2020, 21, 162–174. [Google Scholar] [CrossRef]
- De Bono, J.; Mateo, J.; Fizazi, K.; Saad, F.; Shore, N.; Sandhu, S.; Chi, K.N.; Sartor, O.; Agarwal, N.; Olmos, D.; et al. Olaparib for Metastatic Castration-Resistant Prostate Cancer. N. Engl. J. Med. 2020, 382, 2091–2102. [Google Scholar] [CrossRef] [PubMed]
- Polo, S.H.; Muñoz, D.M.; Rodríguez, A.R.; Ruiz, J.S.; Rodríguez, D.R.; Couñago, F. Changing the History of Prostate Cancer with New Targeted Therapies. Biomedicines 2022, 9, 392. [Google Scholar] [CrossRef] [PubMed]
- Antonarakis, E.S.; Wang, H.; Teply, B.A.; Kelly, W.K.; Willms, J.; Sullivan, R.; King, S.; Marshall, C.H.; Lotan, T. Interim results from a phase 2 study of olaparib (without ADT) in men with biochemically-recurrent prostate cancer after prostatectomy, with integrated biomarker analysis. J. Clin. Oncol. 2019, 37 (Suppl. 15), 5045. [Google Scholar] [CrossRef]
- Abida, W.; Campbell, D.; Patnaik, A.; Sautois, B.; Shapiro, J.; Vogelzang, N.; Bryce, A.; McDermott, R.; Ricci, F.; Rowe, J.; et al. Preliminary results from the TRITON2 study of rucaparib in patients with DNA damage repair (DDR)-deficient metastatic castration-resistant prostate cancer (mCRPC): Updated analyses. Ann. Oncol. 2019, 30, 327–328. [Google Scholar] [CrossRef]
- Abida, W.; Campbell, D.; Patnaik, A.; Shapiro, J.D.; Sautois, B.; Vogelzang, N.J.; Voog, E.G.; Bryce, A.H.; McDermott, R.; Ricci, F.; et al. Non-BRCA DNA Damage Repair Gene Alterations and Response to the PARP Inhibitor Rucaparib in Metastatic Castration-Resistant Prostate Cancer: Analysis From the Phase II TRITON2 Study. Clin. Cancer Res. 2020, 26, 2487–2496. [Google Scholar] [CrossRef]
- Markowski, M.C.; Antonarakis, E.S. PARP inhibitors in prostate cancer: Time to narrow patient selection? Expert Rev. Anticancer. Ther. 2020, 20, 523–526. [Google Scholar] [CrossRef]
- Ryan, C.J.; Abida, W.; Bryce, A.H.; Balar, A.V.; Dumbadze, I.; Given, R.W.; Morris, D.; Petrylak, D.P.; Redfern, C.H.; Scher, H.I.; et al. TRITON3: An international, randomized, open-label, phase III study of the PARP inhibitor Rucaparib vs. physician’s choice of therapy for patients with metastatic castration-resistant prostate cancer (mCRPC) associated with homologous recombination deficiency (HRD). J. Clin. Oncol. 2018, 36 (Suppl. 6), 6. [Google Scholar]
- Markowski, M.C.; Wang, H.; Sullivan, R.; Haffner, M.; De Marzo, A.M.; Lotan, T.L.; Antonarakis, E.S. Phase II trial of rucaparib (Without ADT) in patients with metastatic hormone-sensitive prostate cancer harboring germline DNA repair gene mutations (TRIUMPH). J. Clin. Oncol. 2018, 36, 35. [Google Scholar] [CrossRef]
- ClinicalTrials. Available online: https://clinicaltrials.gov/ct2/show/NCT03533946?term=NCT03533946&draw=2&rank=1 (accessed on 27 April 2022).
- Smith, M.R.; Sandhu, S.K.; Kelly, W.K.; Scher, H.I.; Efstathiou, E.; Lara, P.N.; Yu, E.Y.; George, D.J.; Chi, K.N.; Saad, F.; et al. Pre-specified interim analysis of GALAHAD: A phase II study of niraparib in patients (pts) with metastatic castration-resistant prostate cancer (mCRPC) and biallelic DNA-repair gene defects (DRD). Ann. Oncol. 2019, 30, 884–885. [Google Scholar] [CrossRef]
- Smith, M.R.; Fizazi, K.; Sandhu, S.K.; Kelly, W.K.; Efstathiou, E.; Lara, P.; Yu, E.Y.; George, D.J.; Chi, K.N.; Saad, F.; et al. Niraparib in patients (pts) with metastatic castration-resistant prostate cancer (mCRPC) and biallelic DNA-repair gene defects (DRD): Correlative measures of tumor response in phase II GALAHAD study. J. Clin. Oncol. 2020, 38, 118. [Google Scholar] [CrossRef]
- ClinicalTrials. Available online: https://clinicaltrials.gov/ct2/show/NCT04030559?term=NCT04030559&draw=2&rank=1 (accessed on 27 April 2022).
- Litton, J.K.; Rugo, H.S.; Ettl, J.; Hurvitz, S.A.; Gonçalves, A.; Lee, K.-H.; Fehrenbacher, L.; Yerushalmi, R.; Mina, L.A.; Martin, M.; et al. Talazoparib in Patients with Advanced Breast Cancer and a Germline BRCA Mutation. N. Engl. J. Med. 2018, 379, 753–763. [Google Scholar] [CrossRef] [PubMed]
- de Bono, J.S.; Mehra, N.; Scagliotti, G.V.; Castro, E.; Dorff, T.; Stirling, A.; Stenzl, A.; Fleming, M.T.; Higano, C.S.; Saad, F.; et al. Talazoparib monotherapy in metastatic castration-resistant prostate cancer with DNA repair alterations (TALAPRO-1): An open-label, phase 2 trial. Lancet Oncol. 2021, 22, 1250–1264. [Google Scholar] [CrossRef]
- Pujade-Lauraine, E.; Ledermann, J.A.; Selle, F.; Gebski, V.; Penson, R.T.; Oza, A.M.; Korach, J.; Huzarski, T.; Poveda, A.; Pignata, S.; et al. Olaparib tablets as maintenance therapy in patients with platinum-sensitive, relapsed ovarian cancer and a BRCA1/2 mutation (SOLO2/ENGOT-Ov21): A double-blind, randomised, placebo-controlled, phase 3 trial. Lancet Oncol. 2017, 18, 1274–1284. [Google Scholar] [CrossRef]
- Coleman, R.L.; Oza, A.M.; Lorusso, D.; Aghajanian, C.; Oaknin, A.; Dean, A.; Colombo, N.; Weberpals, J.I.; Clamp, A.; Scambia, G.; et al. Rucaparib maintenance treatment for recurrent ovarian carcinoma after response to platinum therapy (ARIEL3): A randomised, double-blind, placebo-controlled, phase 3 trial. Lancet 2017, 390, 1949–1961. [Google Scholar] [CrossRef]
- Abida, W.; Patnaik, A.; Campbell, D.; Shapiro, J.; Bryce, A.H.; McDermott, R.; Sautois, B.; Vogelzang, N.J.; Bambury, R.M.; Voog, E.; et al. Rucaparib in Men With Metastatic Castration-Resistant Prostate Cancer Harboring a BRCA1 or BRCA2 Gene Alteration. J. Clin. Oncol. 2020, 38, 3763–3772. [Google Scholar] [CrossRef] [PubMed]
- Nitecki, R.; Melamed, A.; Gockley, A.A.; Floyd, J.; Krause, K.J.; Coleman, R.L.; Matulonis, U.A.; Giordano, S.H.; Lu, K.H.; Rauh-Hain, J.A. Incidence of myelodysplastic syndrome and acute myeloid leukemia in patients receiving poly-ADP ribose polymerase inhibitors for the treatment of solid tumors: A meta-analysis of randomized trials. Gynecol. Oncol. 2021, 161, 653–659. [Google Scholar] [CrossRef]
- Teyssonneau, D.; Margot, H.; Cabart, M.; Anonnay, M.; Sargos, P.; Vuong, N.-S.; Soubeyran, I.; Sevenet, N.; Roubaud, G. Prostate cancer and PARP inhibitors: Progress and challenges. J. Hematol. Oncol. 2021, 14, 51–70. [Google Scholar] [CrossRef]
- Lord, C.J.; Ashworth, A. Mechanisms of resistance to therapies targeting BRCA-mutant cancers. Nat. Med. 2013, 19, 1381–1388. [Google Scholar] [CrossRef]
- D’Andrea, A.D. Mechanisms of PARP inhibitor sensitivity and resistance. DNA Repair 2018, 71, 172–176. [Google Scholar] [CrossRef]
- Du, Y.; Yamaguchi, H.; Hsu, J.L.; Hung, M.-C. PARP inhibitors as precision medicine for cancer treatment. Natl. Sci. Rev. 2017, 4, 576–592. [Google Scholar] [CrossRef][Green Version]
- Sachdev, E.; Tabatabai, R.; Roy, V.; Rimel, B.J.; Mita, M.M. PARP Inhibition in Cancer: An Update on Clinical Development. Target. Oncol. 2019, 14, 657–679. [Google Scholar] [CrossRef] [PubMed]
- Ganesan, S. Tumor Suppressor Tolerance: Reversion Mutations in BRCA1 and BRCA2 and Resistance to PARP Inhibitors and Platinum. JCO Precis. Oncol. 2018, 2, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Gourley, C.; Balmaña, J.; Ledermann, J.A.; Serra, V.; Dent, R.; Loibl, S.; Pujade-Lauraine, E.; Boulton, S.J. Moving From Poly (ADP-Ribose) Polymerase Inhibition to Targeting DNA Repair and DNA Damage Response in Cancer Therapy. J. Clin. Oncol. 2019, 37, 2257–2269. [Google Scholar] [CrossRef] [PubMed]
- Sharifi, N.; Gulley, J.L.; Dahut, W.L. An update on androgen deprivation therapy for prostate cancer. Endocr.-Relat. Cancer 2010, 17, 305–315. [Google Scholar] [CrossRef] [PubMed]
- Lepor, H.; Shore, N.D. LHRH Agonists for the Treatment of Prostate Cancer. Rev. Urol. 2012, 14, 1–12. [Google Scholar]
- Li, L.; Karanika, S.; Yang, G.; Wang, J.; Park, S.; Broom, B.M.; Manyam, G.C.; Wu, W.; Luo, Y.; Basourakos, S.; et al. Androgen receptor inhibitor–induced “BRCAness” and PARP inhibition are synthetically lethal for castration-resistant prostate cancer. Sci. Signal. 2017, 10, 7479–7503. [Google Scholar] [CrossRef]
- Asim, M.; Tarish, F.; Zecchini, H.I.; Sanjiv, K.; Gelali, E.; Massie, C.E.; Baridi, A.; Warren, A.Y.; Zhao, W.; Ogris, C.; et al. Synthetic lethality between androgen receptor signalling and the PARP pathway in prostate cancer. Nat. Commun. 2017, 8, 374–402. [Google Scholar] [CrossRef]
- Polkinghorn, W.R.; Parker, J.S.; Lee, M.X.; Kass, E.M.; Spratt, D.; Iaquinta, P.J.; Arora, V.K.; Yen, W.-F.; Cai, L.; Zheng, D.; et al. Androgen Receptor Signaling Regulates DNA Repair in Prostate Cancers. Cancer Discov. 2013, 3, 1245–1253. [Google Scholar] [CrossRef]
- Schiewer, M.J.; Goodwin, J.F.; Han, S.; Brenner, J.C.; Augello, M.A.; Dean, J.L.; Liu, F.; Planck, J.L.; Ravindranathan, P.; Chinnaiyan, A.M.; et al. Dual Roles of PARP-1 Promote Cancer Growth and Progression. Cancer Discov. 2012, 2, 1134–1149. [Google Scholar] [CrossRef]
- Hussain, M.; Daignault-Newton, S.; Twardowski, P.W.; Albany, C.; Stein, M.N.; Kunju, L.P.; Siddiqui, J.; Wu, Y.-M.; Robinson, D.; Lonigro, R.J.; et al. Targeting Androgen Receptor and DNA Repair in Metastatic Castration-Resistant Prostate Cancer: Results From NCI 9012. J. Clin. Oncol. 2018, 36, 991–999. [Google Scholar] [CrossRef]
- Clarke, N.; Wiechno, P.; Alekseev, B.; Sala, N.; Jones, R.; Kocak, I.; Chiuri, V.E.; Jassem, J.; Fléchon, A.; Redfern, C.; et al. Olaparib combined with abiraterone in patients with metastatic castration-resistant prostate cancer: A randomised, double-blind, placebo-controlled, phase 2 trial. Lancet Oncol. 2018, 19, 975–986. [Google Scholar] [CrossRef]
- Saad, F.; Chi, K.; Shore, N.; Graff, J.; Posadas, E.; Freeman, S.; Tryon, J.; Trudel, G.; De Jong, J.; Meltzer, J.; et al. Interim results of a phase Ib study of niraparib plus androgen receptor-targeted therapy in men with metastatic castration-resistant prostate cancer. Ann. Oncol. 2018, 29, 8. [Google Scholar] [CrossRef]
- Chi, K.N.; Rathkopf, D.E.; Smith, M.R.; Efstathiou, E.; Attard, G.; Olmos, D.; Lee, J.Y.; Small, E.J.; Gomes, A.J.; Roubaud, G.; et al. Phase 3 MAGNITUDE study: First results of niraparib (NIRA) with abiraterone acetate and prednisone (AAP) as first-line therapy in patients (pts) with metastatic castration-resistant prostate cancer (mCRPC) with and without homologous recombination repair (HRR) gene alterations. J. Clin. Oncol. 2022, 40, 12. [Google Scholar] [CrossRef]
- Saad, F.; Armstrong, A.J.; Thiery-Vuillemin, A.; Oya, M.; Loredo, E.; Procopio, G.; de Menezes, J.J.; Girotto, G.C.; Arslan, C.; Mehra, N.; et al. PROpel: Phase III trial of olaparib (ola) and abiraterone (abi) versus placebo (pbo) and abi as first-line (1L) therapy for patients (pts) with metastatic castration-resistant prostate cancer (mCRPC). J. Clin. Oncol. 2022, 40, 11. [Google Scholar] [CrossRef]
- Virtanen, V.; Paunu, K.; Ahlskog, J.K.; Varnai, R.; Sipeky, C.; Sundvall, M. PARP Inhibitors in Prostate Cancer–the Preclinical Rationale and Current Clinical Development. Genes 2019, 10, 565. [Google Scholar] [CrossRef]
- Jiao, S.; Xia, W.; Yamaguchi, H.; Wei, Y.; Chen, M.K.; Hsu, J.M.; Hsu, J.L.; Yu, W.H.; Du, Y.; Lee, H.H.; et al. PARP Inhibitor Upregulates PD-L1 Expression and Enhances Cancer-Associated Immunosuppression. Clin. Cancer Res. 2017, 23, 3711–3720. [Google Scholar] [CrossRef]
- McGrail, D.J.; Federico, L.; Li, Y.; Dai, H.; Lu, Y.; Mills, G.B.; Yi, S.; Lin, S.-Y.; Sahni, N. Multi-omics analysis reveals neoantigen-independent immune cell infiltration in copy-number driven cancers. Nat. Commun. 2018, 9, 1317. [Google Scholar] [CrossRef]
- Ablasser, A.; Goldeck, M.; Cavlar, T.; Deimling, T.; Witte, G.; Röhl, I.; Hopfner, K.; Ludwig, J.; Hornung, V. CGAS produces a 2′-5′-linked cyclic dinucleotide second messenger that activates STING. Nature 2013, 498, 380–384. [Google Scholar] [CrossRef]
- Karzai, F.; VanderWeele, D.; Madan, R.A.; Owens, H.; Cordes, L.M.; Hankin, A.; Couvillon, A.; Nichols, E.; Bilusic, M.; Beshiri, M.; et al. Activity of durvalumab plus olaparib in metastatic castration-resistant prostate cancer in men with and without DNA damage repair mutations. J. Immunother. Cancer 2018, 6, 141. [Google Scholar] [CrossRef]
- Yu, E.; Piulats, J.; Gravis, G.; Fong, P.; Todenhöfer, T.; Laguerre, B.; Arranz, J.; Oudard, S.; Massard, C.; Stoeckle, M.; et al. Association between homologous recombination repair mutations and response to pembrolizumab (pembro) plus olaparib (ola) in metastatic castration-resistant prostate cancer (mCRPC): KEYNOTE-365 Cohort A biomarker analysis. Ann. Oncol. 2021, 32, 387. [Google Scholar] [CrossRef]
- Study of Pembrolizumab (MK-3475) Plus Olaparib Versus Abiraterone Acetate or Enzalutamide in Metastatic Castration-Resistant Prostate Cancer (mCRPC) (MK-7339-010/KEYLYNK-010) (KEYLYNK-010). Available online: Clinicaltrials.gov/ct2/show/NCT03834519 (accessed on 27 April 2022).
- Chan, N.; Koritzinsky, M.; Zhao, H.; Bindra, R.; Glazer, P.; Powell, S.; Belmaaza, A.; Wouters, B.; Bristow, R. Chronic Hypoxia Decreases Synthesis of Homologous Recombination Proteins to Offset Chemoresistance and Radioresistance. Cancer Res. 2008, 68, 605–614. [Google Scholar] [CrossRef] [PubMed]
- McKay, R.R.; Radke, M.R.; Shyr, Y.; Zhao, S.; Taplin, M.-E.; Davis, N.B.; Monk, P.; Appleman, L.J.; Lara, P.N.; Vaishampayan, U.N.; et al. Biomarker analysis from a randomized phase II study of olaparib with or without cediranib in men with metastatic castration-resistant prostate cancer (mCRPC). J. Clin. Oncol. 2021, 39 (Suppl. 6), 7. [Google Scholar] [CrossRef]
- Dias, M.P.; Moser, S.C.; Ganesan, S.; Jonkers, J. Understanding and overcoming resistance to PARP inhibitors in cancer therapy. Nat. Rev. Clin. Oncol. 2021, 18, 773–791. [Google Scholar] [CrossRef] [PubMed]
- Jette, N.R.; Kumar, M.; Radhamani, S.; Arthur, G.; Goutam, S.; Yip, S.; Kolinsky, M.; Williams, G.J.; Bose, P.; Lees-Miller, S.P. ATM-Deficient Cancers Provide New Opportunities for Precision Oncology. Cancers 2020, 12, 687. [Google Scholar] [CrossRef] [PubMed]
- Hussain, M.; Carducci, M.A.; Slovin, S.; Cetnar, J.; Qian, J.; McKeegan, E.M.; Refici-Buhr, M.; Chyla, B.; Shepherd, S.P.; Giranda, V.L.; et al. Targeting DNA repair with combination veliparib (ABT-888) and temozolomide in patients with metastatic castration-resistant prostate cancer. Inv. New Drugs 2014, 32, 904–912. [Google Scholar] [CrossRef]
- Kelly, W.K.; Leiby, B.; Einstein, D.J.; Szmulewitz, R.Z.; Sartor, A.O.; Yang, E.S.-H.; Sonpavde, G. Radium-223 (Rad) and niraparib (Nira) treatment (tx) in castrate-resistant prostate cancer (CRPC) patients (pts) with and without prior chemotherapy (chemo). J. Clin. Oncol. 2020, 38, 5540. [Google Scholar] [CrossRef]
- Lin, M.; Ta, R.T.; Kairemo, K.; Le, D.B.; Ravizzini, G.C. Prostate-Specific Membrane Antigen-Targeted Radiopharmaceuticals in Diagnosis and Therapy of Prostate Cancer: Current Status and Future Perspectives. Cancer Biother. Radiopharm. 2021, 36, 237–251. [Google Scholar] [CrossRef]
- Wurzer, A.; di Carlo, D.; Schmidt, A.; Beck, R.; Eiber, M.; Schwaiger, M.; Wester, H.-J. Radiohybrid Ligands: A Novel Tracer Concept Exemplified by 18F- or 68Ga-Labeled rhPSMA Inhibitors. J. Nucl. Med. 2020, 61, 735–742. [Google Scholar] [CrossRef]
- Hofman, M.S.; Violet, J.; Hicks, R.J.; Ferdinandus, J.; Thang, S.P.; Akhurst, T.; Iravani, A.; Kong, G.; Kumar, A.R.; Murphy, D.G.; et al. [177 Lu]-PSMA-617 radionuclide treatment in patients with metastatic castration-resistant prostate cancer (LuPSMA trial): A single-centre, single-arm, phase 2 study. Lancet Oncol. 2018, 19, 825–833. [Google Scholar] [CrossRef]
- Rahbar, K.; Boegemann, M.; Yordanova, A.; Eveslage, M.; Schäfers, M.; Essler, M.; Ahmadzadehfar, H. PSMA targeted radioligandtherapy in metastatic castration resistant prostate cancer after chemotherapy, abiraterone and/or enzalutamide. A retrospective analysis of overall survival. Eur. J. Pediatr. 2018, 45, 12–19. [Google Scholar] [CrossRef] [PubMed]
- Marchetti, A.; Rosellini, M.; Nuvola, G.; Tassinari, E.; Mollica, V.; Rizzo, A.; Santoni, M.; Cimadamore, A.; Farolfi, A.; Montironi, R.; et al. PARP Inhibitors and Radiometabolic Approaches in Metastatic Castration-Resistant Prostate Cancer: What’s Now, What’s New, and What’s Coming? Cancers 2022, 14, 907. [Google Scholar] [CrossRef] [PubMed]
- Powell, C.; Mikropoulos, C.; Kaye, S.; Nutting, C.; Bhide, S.; Newbold, K.; Harrington, K. Pre-clinical and clinical evaluation of PARP inhibitors as tumour-specific radiosensitisers. Cancer Treat. Rev. 2010, 36, 566–575. [Google Scholar] [CrossRef] [PubMed]
- Qiu, Z.; Oleinick, N.L.; Zhang, J. ATR/CHK1 inhibitors and cancer therapy. Radiother. Oncol. 2018, 126, 450–464. [Google Scholar] [CrossRef] [PubMed]
- Dungey, F.A.; Löser, D.A.; Chalmers, A.J. Replication-Dependent Radiosensitization of Human Glioma Cells by Inhibition of Poly(ADP-Ribose) Polymerase: Mechanisms and Therapeutic Potential. Int. J. Radiat. Oncol. 2008, 72, 1188–1197. [Google Scholar] [CrossRef]
- Jia, G.; Fu, Y.; He, C. Reversible RNA adenosine methylation in biological regulation. Trends Genet. 2013, 29, 108–115. [Google Scholar] [CrossRef]
- Mansour, W.Y.; Rhein, T.; Dahm-Daphi, J. The alternative end-joining pathway for repair of DNA double-strand breaks requires PARP1 but is not dependent upon microhomologies. Nucleic Acids Res. 2010, 38, 6065–6077. [Google Scholar] [CrossRef]
- Wang, M.; Wu, W.; Wu, W.; Rosidi, B.; Zhang, L.; Wang, H.; Iliakis, G. PARP-1 and Ku compete for repair of DNA double strand breaks by distinct NHEJ pathways. Nucleic Acids Res. 2006, 34, 6170–6182. [Google Scholar] [CrossRef]
- Toulany, M. Targeting DNA Double-Strand Break Repair Pathways to Improve Radiotherapy Response. Genes 2019, 10, 25. [Google Scholar] [CrossRef]
- Rae, C.; Mairs, R.J. Evaluation of the radiosensitizing potency of chemotherapeutic agents in prostate cancer cells. Int. J. Radiat. Biol. 2017, 93, 194–203. [Google Scholar] [CrossRef]
- Zumsteg, Z.; Karrison, T.; Michaelson, M.; Tran, P.; Kudchadker, R.; Feng, F. 689TiP NRG Oncology’s GU007 (NADIR): A randomized phase II trial of niraparib with standard combination androgen deprivation therapy (ADT) and radiotherapy (RT) in high-risk prostate cancer (PC) (with initial phase I). Ann. Oncol. 2020, 31, 546. [Google Scholar] [CrossRef]
- Messina, C.; Cattrini, C.; Soldato, D.; Vallome, G.; Caffo, O.; Castro, E.; Olmos, D.; Boccardo, F.; Zanardi, E. BRCA Mutations in Prostate Cancer: Prognostic and Predictive Implications. J. Oncol. 2020, 49, 863–865. [Google Scholar] [CrossRef] [PubMed]
- Page, E.C.; Bancroft, E.K.; Brook, M.N.; Assel, M.; Hassan Al Battat, M.; Thomas, S.; Taylor, N.; Chamberlain, A.; Pope, J.; Ni Raghallaigh, H.; et al. Interim Results from the IMPACT Study: Evidence for Prostate-specific Antigen Screening in BRCA2 Mutation Carriers. Eur. Urol. 2019, 76, 831–842. [Google Scholar] [CrossRef] [PubMed]
- Bancroft, E.; Page, E.C.; Castro, E.; Lilja, H.; Vickers, A.; Sjoberg, D.; Assel, M.; Foster, C.S.; Mitchell, G.; Drew, K.; et al. Targeted Prostate Cancer Screening in BRCA1 and BRCA2 Mutation Carriers: Results from the Initial Screening Round of the IMPACT Study. Eur. Urol. 2014, 66, 489–499. [Google Scholar] [CrossRef] [PubMed]
- Giri, V.N.; Knudsen, K.E.; Kelly, W.K.; Abida, W.; Andriole, G.L.; Bangma, C.H.; Bekelman, J.E.; Benson, M.C.; Blanco, A.; Burnett, A.; et al. Role of Genetic Testing for Inherited Prostate Cancer Risk: Philadelphia Prostate Cancer Consensus Conference 2017. J. Clin. Oncol. 2018, 36, 414–424. [Google Scholar] [CrossRef] [PubMed]
- Sokolova, A.O.; Cheng, H.H. Genetic Testing in Prostate Cancer. Curr. Oncol. Rep. 2020, 22, 5. [Google Scholar] [CrossRef]
- Teyssonneau, D.; Thiery-Vuillemin, A.; Dariane, C.; Barret, E.; Beauval, J.-B.; Brureau, L.; Créhange, G.; Fiard, G.; Fromont, G.; Gauthé, M.; et al. PARP Inhibitors as Monotherapy in Daily Practice for Advanced Prostate Cancers. J. Clin. Med. 2022, 11, 1734. [Google Scholar] [CrossRef]
- Shah, S.; Rachmat, R.; Enyioma, S.; Ghose, A.; Revythis, A.; Boussios, S. BRCA Mutations in Prostate Cancer: Assessment, Implications and Treatment Considerations. Int. J. Mol. Sci. 2021, 22, 12628. [Google Scholar] [CrossRef]
- National Comprehensive Cancer Network Clinical Guidelines in Oncology (NCCN Guidelines): Prostate Cancer Early Detection (Version 1.2022). Available online: https://www.nccn.org/professionals/physician_gls/pdf/prostate_detection (accessed on 27 April 2022).
- National Comprehensive Cancer Network Clinical Guidelines in Oncology (NCCN Guidelines): Prostate Cancer (Version 4.2022). Available online: https://www.nccn.org/guidelines/guidelines-detail?category=1&id=1459 (accessed on 27 April 2022).
- Parker, C.; Castro, E.; Fizazi, K.; Heidenreich, A.; Ost, P.; Procopio, G.; Tombal, B.; Gillessen, S.; ESMO Guidelines Committee. Prostate cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann. Oncol. 2020, 31, 1119–1134. [Google Scholar] [CrossRef]
- Bieńkowski, M.; Tomasik, B.; Braun, M.; Jassem, J. PARP inhibitors for metastatic castration-resistant prostate cancer: Biological rationale and current evidence. Cancer Treat. Rev. 2022, 104, 620–631. [Google Scholar] [CrossRef]
- Antonarakis, E.S.; Gomella, L.G.; Petrylak, D.P. When and How to Use PARP Inhibitors in Prostate Cancer: A Systematic Review of the Literature with an Update on On-Going Trials. Eur. Urol. Oncol. 2020, 3, 594–611. [Google Scholar] [CrossRef] [PubMed]
- Mohyuddin, G.R.; Aziz, M.; Britt, A.; Wade, L.; Sun, W.; Baranda, J.; Al-Rajabi, R.; Saeed, A.; Kasi, A. Similar response rates and survival with PARP inhibitors for patients with solid tumors harboring somatic versus Germline BRCA mutations: A Meta-analysis and systematic review. BMC Cancer 2020, 20, 507. [Google Scholar] [CrossRef] [PubMed]

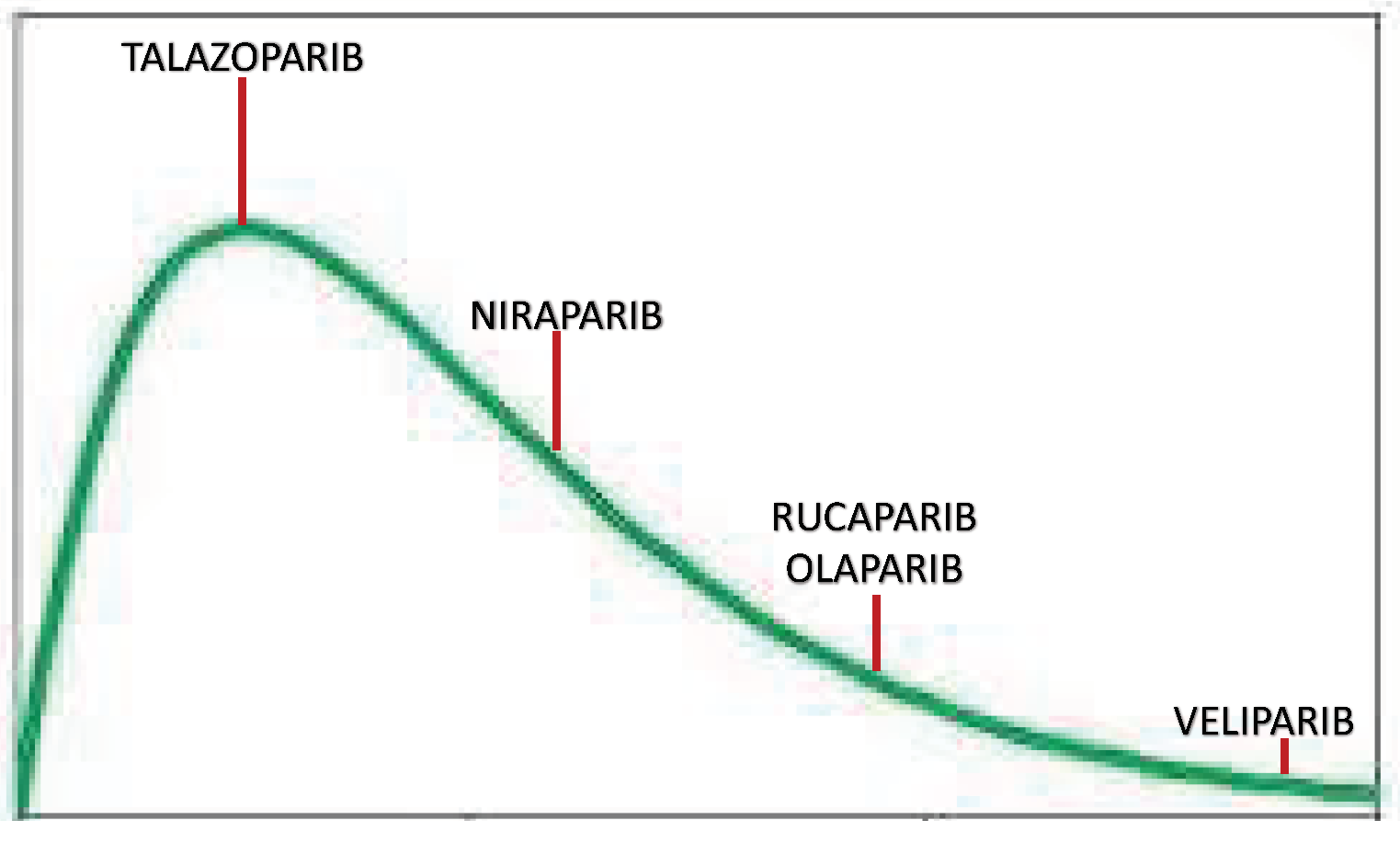
| Structural Formula | Dose [mg] | Cmax [ng/mL] | Tmax [h] | Half-Life [h] | PARP1 Trapping Ability | Direct Off-Targets | Indirect Off-Targets | |
|---|---|---|---|---|---|---|---|---|
| Olaparib (AZD-2281, MK-7339) | 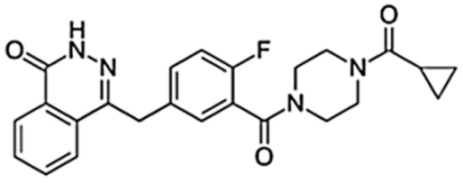 | 300/12 h | 7700 | 1.5 | 14.9 | Moderate | PARP1, PARP2, PARP3 | |
| Rucaparib (AG014699) | 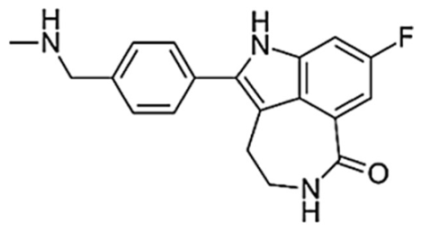 | 600/12 h | 1940 | 1.9 | 25.9 | Moderate | ARTD5, ARTD6, PARP1,2,3 TNKS1, TNKS2 | ALDH2, H6PD, CDK16, PIM3, DYRK1B |
| Niraparib (MK-4827) |  | 300/24 h | 2232 | 3.0 | 36 | Moderate–high | PARP2, PARP3, PARP4, PARP12 | ALDH2, CIT, DCK, DYRK1A, DYRK1B |
| Talazoparib (BMN-673) | 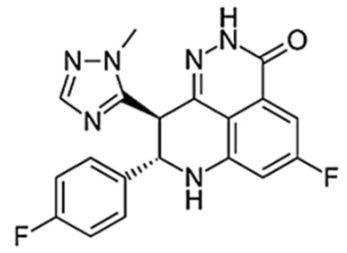 | 1/24 h | 16.4 | 1–2 | 90 | Very high | PARP2 PARP1 | |
| Veliparib (ABT-888) | 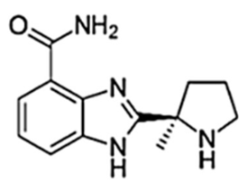 | 40/12 h | 410 | 1.0 | 6 | Very low | PARP2, PARP3, PARP10 |
| STUDY | DESIGN | Estimated Enrollment | Setting | Agent(s) | Homologous Recombination Repair Mutations | Primary Endpoint(s) |
|---|---|---|---|---|---|---|
|
TALAPRO-2 NCT03395197 | Phase III, randomized | 872 | mCRPC treatment naïve | Talazoparib + enza vs. placebo + enza | Selected | Safety, PFS |
| TALAPRO-3 NCT04821622 | Phase III, randomized | 550 | mHSPC | Talazoparib + enza vs. placebo + enza | Selected | rPFS |
| NCT03012321 | Phase II, randomized | 70 | mCRPC treatment naïve | Olaparib vs. AAP vs. olaparib + AAP | Selected | PFS |
| PROpel NCT03732820 | Phase III, randomized | 720 | mCRPC treatment naïve | Olaparib + AAP vs. placebo + AAP | Unselected | rPFS |
| CASPAR NCT04455750 | Phase III, randomized | 1002 | mCRPC treatment naive | Rucaparib + enza vs. placebo + enza | Unselected | rPFS, OS |
| MAGNITUDE NCT03748641 | Phase III, randomized | 1000 | mCRPC treatment naïve | Niraparib + AAP vs. placebo + AAP | Selected | rPFS |
| NADIR NCT04037254 | Phase II, randomized | 180 | High risk localized or locally advanced PCa (no prior treatment) | Niraparib + RT + ADT vs. niraparib alone vs. RT + ADT | Unselected | Maintenance of disease-free state |
| ASCLEPIuS NCT04194554 | Phase I/II, single arm, open label | 100 | High risk locally advanced PCa (cN+) | Niraparib + AAP + leuprolide + RT | Unselected | DLT, biochemical failure (% of pts) |
| AMPLITUDE NCT04497844 | Phase III, randomized | 788 | mHSPC | NIRAPARIB+AAP vs. PLACEBO+AAP | Selected | rPFS |
| ZZ-First (NCT04332744) | Phase II, randomized, open label | 54 | mHSPC | Enzalutamide vs. enzalutamide + talazoparib | Unselected | PSA-Complete Response |
| STUDY | DESIGN | Estimated Enrollment | Setting | Agent(s) | Homologous Recombination Repair Mutations | Primary Endpoint(s) |
|---|---|---|---|---|---|---|
| KEYNOTE- 365, NCT02861573 | Phase Ib/II, nonrandomized | 1000 (104 in COHORT A) | mCRPC after docetaxel and one prior ARSi | Olaparib + pembrolizumab (cohort A) | Unselected | PSA response rate, ORR, safety |
| KEYLYNK-010, NCT03834519 | Phase III, randomized | 780 | mCRPC after docetaxel and one prior ARSi | Olaparib + pembrolizumab vs. enza/AAP | Unselected | OS, rPFS |
| NCT03810105 | Phase II, single arm | 32 | Biochemically recurrent nmCRPC | Olaparib + durvalumab | Selected | Number of pts with undetectable PSA |
| CheckMate 9KD, NCT03338790 | Phase II, nonrandomized | 330 | mCRPC chemotherapynaïve | Nivolumab + rucaparib/ enza/docetaxel | Selected | ORR, PSA response rate |
| QUEST, NCT03431350 | Phase Ib/II, multiarm, nonrandomized | 150 | mCRPC after prior CT and ARSi (depending on cohorts) | Niraparib + AAP vs. niraparib + JNJ-63723283 (anti-PD1) | Both selected and unselected | ORR, incidence of AEs |
| STUDY | DESIGN | Estimated Enrollment | Setting | Agent (s) | Homologous Recombination Repair Mutations | Primary Endpoint(s) |
|---|---|---|---|---|---|---|
| LuPARP, NCT03874884 | Phase I, single arm | 52 | mCRPC after prior CT and ARSis | Olaparib + 177Lu-PSMA | Not available | Dose-limiting toxicity (DLT) recommended phase II dose |
| COMRADE, NCT03317392 | Phase I/II, randomized | 112 | mCRPC after prior CT and ARSis | Olaparib + Radium-223 vs. Radium-223 | Not available | rPFS, maximum tolerated dose |
| NiraRad, NCT03076203 | Phase Ib, single-arm | 14 | mCRPC after at least one prior ARSi, with or without prior CT | Niraparib + Radium-223 | Unselected | DLT |
| TRAP, NCT03787680 | Phase II, nonrandomized | 47 | mCRCP after prior ARSi | Olaparib + AZD6738 (ATRinhibitor) | Selected | Rate of response (CR or PR), PSA response >50% decline |
| NCT03840200 | Phase Ib, nonrandomized | 51 | mCRPC after one prior ARSi | Rucaparib + ipatasertib (AKTinhibitor) | Unselected | DLT, PSA response rate |
| NCT02893917 | Phase II, randomized | 90 | mCRPC after at least one prior therapy | Olaparib + cediranib (VEGF-R TKI) vs. olaparib | Not available | rPFS |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Congregado, B.; Rivero, I.; Osmán, I.; Sáez, C.; Medina López, R. PARP Inhibitors: A New Horizon for Patients with Prostate Cancer. Biomedicines 2022, 10, 1416. https://doi.org/10.3390/biomedicines10061416
Congregado B, Rivero I, Osmán I, Sáez C, Medina López R. PARP Inhibitors: A New Horizon for Patients with Prostate Cancer. Biomedicines. 2022; 10(6):1416. https://doi.org/10.3390/biomedicines10061416
Chicago/Turabian StyleCongregado, Belén, Inés Rivero, Ignacio Osmán, Carmen Sáez, and Rafael Medina López. 2022. "PARP Inhibitors: A New Horizon for Patients with Prostate Cancer" Biomedicines 10, no. 6: 1416. https://doi.org/10.3390/biomedicines10061416
APA StyleCongregado, B., Rivero, I., Osmán, I., Sáez, C., & Medina López, R. (2022). PARP Inhibitors: A New Horizon for Patients with Prostate Cancer. Biomedicines, 10(6), 1416. https://doi.org/10.3390/biomedicines10061416






