Temporal and Locational Values of Images Affecting the Deep Learning of Cancer Stem Cell Morphology
Abstract
:1. Introduction
2. Materials and Methods
2.1. Cell Culture
2.2. Microscopy
2.3. Image Processing and AI
2.4. Statistical Analysis
3. Results
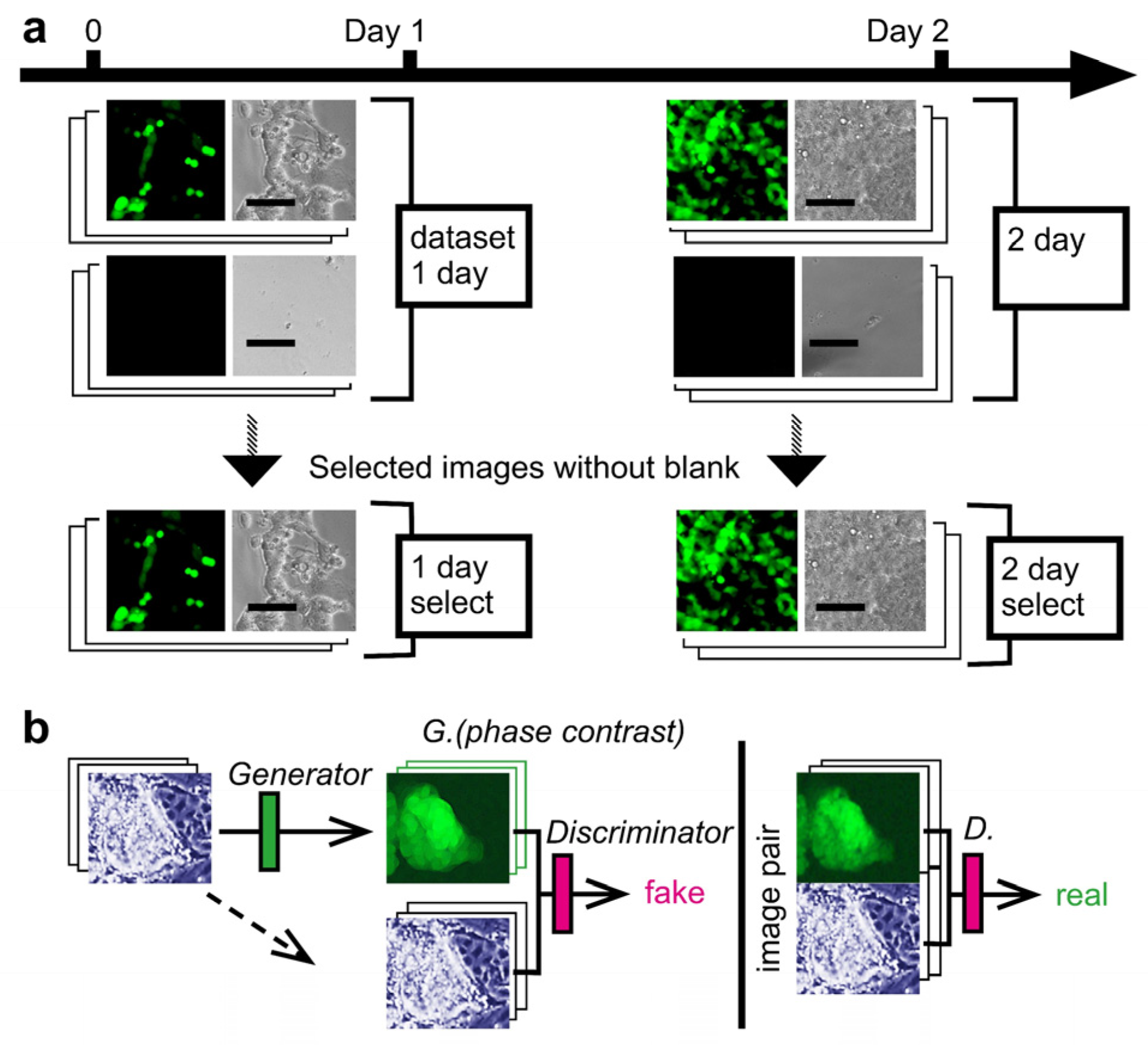
3.1. Image Acquisition of Cultured CSC and Deep Learning
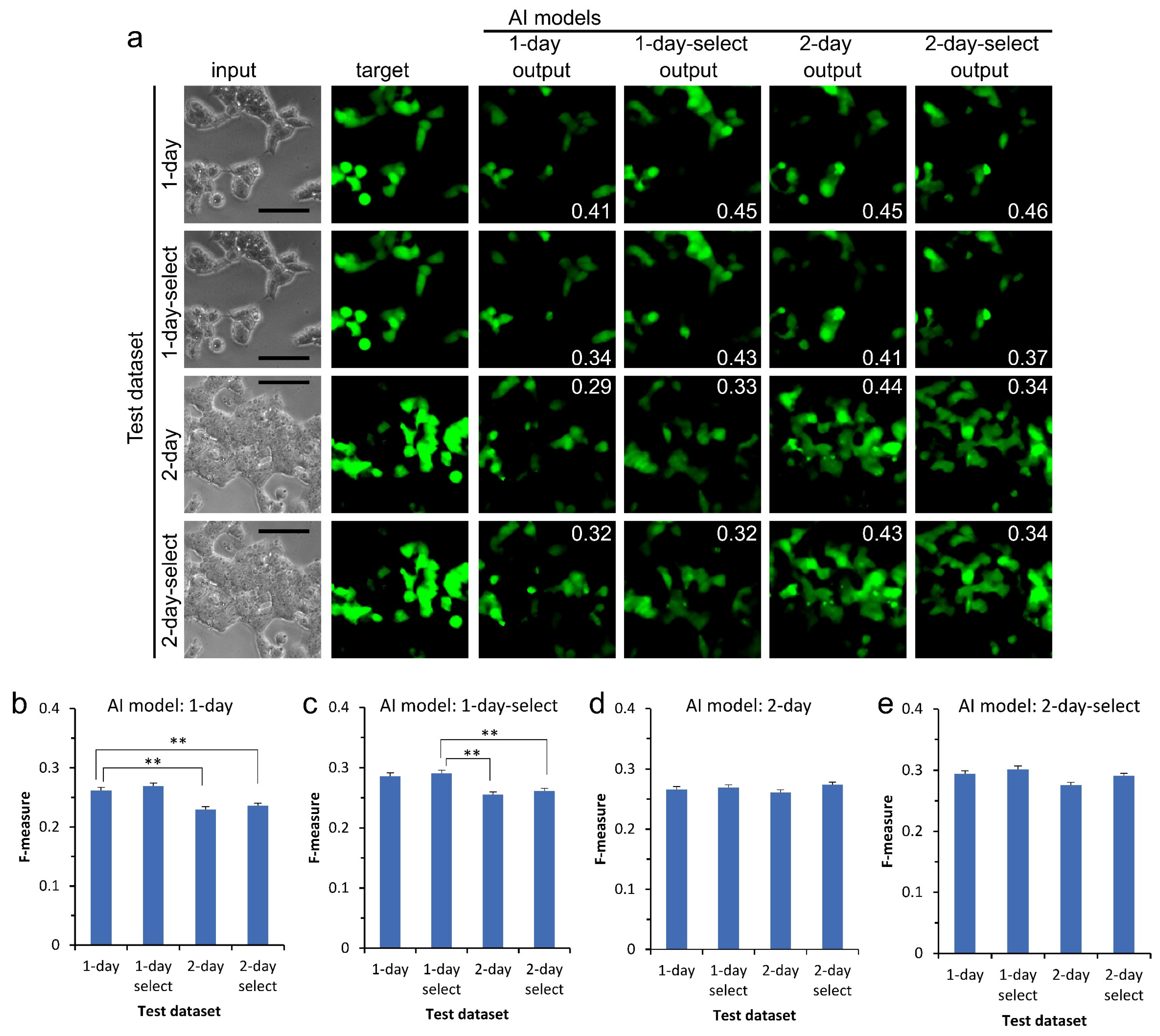
3.2. Versatility of Application of AI to Datasets Other Than the Training Dataset
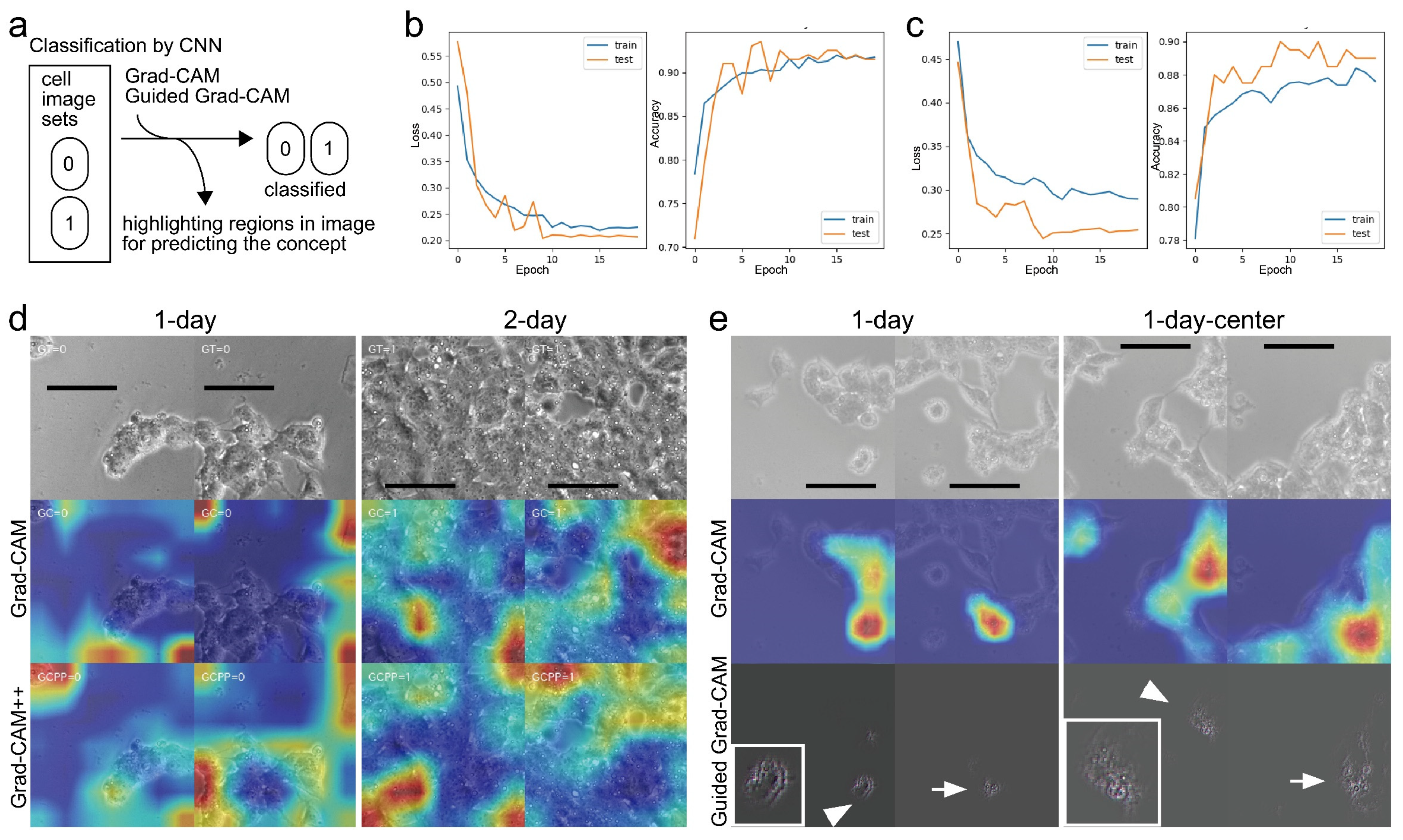
3.3. CSC Object Recognized by Deep Learning
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
References
- Batlle, E.; Clevers, H. Cancer stem cells revisited. Nat. Med. 2017, 23, 1124–1134. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, K.; Yamanaka, S. Induction of Pluripotent Stem Cells from Mouse Embryonic and Adult Fibroblast Cultures by Defined Factors. Cell 2006, 126, 663–676. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, L.; Kasai, T.; Li, Y.; Sugii, Y.; Jin, G.; Okada, M.; Vaidyanath, A.; Mizutani, A.; Satoh, A.; Kudoh, T.; et al. A model of cancer stem cells derived from mouse induced pluripotent stem cells. PLoS ONE 2012, 7, e33544. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Y.M.; Liu, Y.Q.; Liu, D.; Zhang, L.; Qin, J.; Zhang, Z.; Su, Y.; Yan, C.; Luo, Y.L.; Li, J.; et al. The Effects of Astragalus Polysaccharide on Bone Marrow-Derived Mesenchymal Stem Cell Proliferation and Morphology Induced by A549 Lung Cancer Cells. Med. Sci. Monit. 2019, 25, 4110–4121. [Google Scholar] [CrossRef] [PubMed]
- Aida, S.; Okugawa, J.; Fujisaka, S.; Kasai, T.; Kameda, H.; Sugiyama, T. Deep Learning of Cancer Stem Cell Morphology Using Conditional Generative Adversarial Networks. Biomolecules 2020, 10, 931. [Google Scholar] [CrossRef] [PubMed]
- Orita, K.; Sawada, K.; Koyama, R.; Ikegaya, Y. Deep learning-based quality control of cultured human-induced pluripotent stem cell-derived cardiomyocytes. J. Pharmacol. Sci. 2019, 140, 313–316. [Google Scholar] [CrossRef] [PubMed]
- Waisman, A.; La Greca, A.; Mobbs, A.M.; Scarafia, M.A.; Santin Velazque, N.L.; Neiman, G.; Moro, L.N.; Luzzani, C.; Sevlever, G.E.; Guberman, A.S.; et al. Deep Learning Neural Networks Highly Predict Very Early Onset of Pluripotent Stem Cell Differentiation. Stem. Cell Rep. 2019, 12, 845–859. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Boldu, L.; Merino, A.; Acevedo, A.; Molina, A.; Rodellar, J. A deep learning model (ALNet) for the diagnosis of acute leukaemia lineage using peripheral blood cell images. Comput. Methods Progr. Biomed. 2021, 202, 105999. [Google Scholar] [CrossRef] [PubMed]
- Piotrowski, T.; Rippel, O.; Elanzew, A.; Niessing, B.; Stucken, S.; Jung, S.; Konig, N.; Haupt, S.; Stappert, L.; Brustle, O.; et al. Deep-learning-based multi-class segmentation for automated, non-invasive routine assessment of human pluripotent stem cell culture status. Comput. Biol. Med. 2021, 129, 104172. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Huang, R.; Wu, Z.; Song, S.; Cheng, L.; Zhu, R. Deep learning-based predictive identification of neural stem cell differentiation. Nat. Commun. 2021, 12, 2614. [Google Scholar] [CrossRef] [PubMed]
- Pratapa, A.; Doron, M.; Caicedo, J.C. Image-based cell phenotyping with deep learning. Curr. Opin. Chem. Biol. 2021, 65, 9–17. [Google Scholar] [CrossRef] [PubMed]
- Okita, K.; Ichisaka, T.; Yamanaka, S. Generation of germline-competent induced pluripotent stem cells. Nature 2007, 448, 313–317. [Google Scholar] [CrossRef] [PubMed]
- Isola, P.; Zhu, J.; Zhou, T.; Efros, A.A. Image-to-Image Translation with Conditional Adversarial Networks. In Proceedings of the 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 21–26 July 2017; pp. 5967–5976. [Google Scholar]
- Selvaraju, R.R.; Cogswell, M.; Das, A.; Vedantam, R.; Parikh, D.; Batra, D. Grad-CAM: Visual Explanations from Deep Networks via Gradient-Based Localization. Int. J. Comput. Vis. 2020, 128, 336–359. [Google Scholar] [CrossRef] [Green Version]
- Chattopadhay, A.; Sarkar, A.; Howlader, P.; Balasubramanian, V.N. Grad-CAM++: Generalized Gradient-Based Visual Explanations for Deep Convolutional Networks. In Proceedings of the 2018 IEEE Winter Conference on Applications of Computer Vision (WACV), Lake Tahoe, NV, USA, 12–15 March 2018; pp. 839–847. [Google Scholar]
- Liu, Z.; Jin, L.; Chen, J.; Fang, Q.; Ablameyko, S.; Yin, Z.; Xu, Y. A survey on applications of deep learning in microscopy image analysis. Comput. Biol. Med. 2021, 134, 104523. [Google Scholar] [CrossRef] [PubMed]
- Edlund, C.; Jackson, T.R.; Khalid, N.; Bevan, N.; Dale, T.; Dengel, A.; Ahmed, S.; Trygg, J.; Sjogren, R. LIVECell-A large-scale dataset for label-free live cell segmentation. Nat. Methods 2021, 18, 1038–1045. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Guo, L.-P.; Chen, L.-Z.; Zeng, Y.-X.; Lu, S.H. Identification of Cancer Stem Cell–Like Side Population Cells in Human Nasopharyngeal Carcinoma Cell Line. Cancer Res. 2007, 67, 3716–3724. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Meshorer, E.; Misteli, T. Chromatin in pluripotent embryonic stem cells and differentiation. Nat. Rev. Mol. Cell Biol. 2006, 7, 540–546. [Google Scholar] [CrossRef] [PubMed]
- Wiblin, A.E.; Cui, W.; Clark, A.J.; Bickmore, W.A. Distinctive nuclear organisation of centromeres and regions involved in pluripotency in human embryonic stem cells. J. Cell Sci. 2005, 118, 3861–3868. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tokunaga, K.; Saitoh, N.; Goldberg, I.G.; Sakamoto, C.; Yasuda, Y.; Yoshida, Y.; Yamanaka, S.; Nakao, M. Computational image analysis of colony and nuclear morphology to evaluate human induced pluripotent stem cells. Sci. Rep. 2014, 4, 6996. [Google Scholar] [CrossRef] [PubMed] [Green Version]


| Transfer-Learning Model | Class * | Precision | Recall | F-Measure |
|---|---|---|---|---|
| ResNet50 | 0 | 0.888 | 0.950 | 0.918 |
| 1 | 0.946 | 0.880 | 0.912 | |
| VGG16 | 0 | 0.862 | 0.940 | 0.900 |
| 1 | 0.934 | 0.850 | 0.890 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hanai, Y.; Ishihata, H.; Zhang, Z.; Maruyama, R.; Kasai, T.; Kameda, H.; Sugiyama, T. Temporal and Locational Values of Images Affecting the Deep Learning of Cancer Stem Cell Morphology. Biomedicines 2022, 10, 941. https://doi.org/10.3390/biomedicines10050941
Hanai Y, Ishihata H, Zhang Z, Maruyama R, Kasai T, Kameda H, Sugiyama T. Temporal and Locational Values of Images Affecting the Deep Learning of Cancer Stem Cell Morphology. Biomedicines. 2022; 10(5):941. https://doi.org/10.3390/biomedicines10050941
Chicago/Turabian StyleHanai, Yumi, Hiroaki Ishihata, Zaijun Zhang, Ryuto Maruyama, Tomonari Kasai, Hiroyuki Kameda, and Tomoyasu Sugiyama. 2022. "Temporal and Locational Values of Images Affecting the Deep Learning of Cancer Stem Cell Morphology" Biomedicines 10, no. 5: 941. https://doi.org/10.3390/biomedicines10050941
APA StyleHanai, Y., Ishihata, H., Zhang, Z., Maruyama, R., Kasai, T., Kameda, H., & Sugiyama, T. (2022). Temporal and Locational Values of Images Affecting the Deep Learning of Cancer Stem Cell Morphology. Biomedicines, 10(5), 941. https://doi.org/10.3390/biomedicines10050941






