Epidemiological, Clinical, and Genomic Profile in Head and Neck Cancer Patients and Their Families
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Participants
2.2. DNA Extraction and Array-CGH and the Data Analysis
3. Results
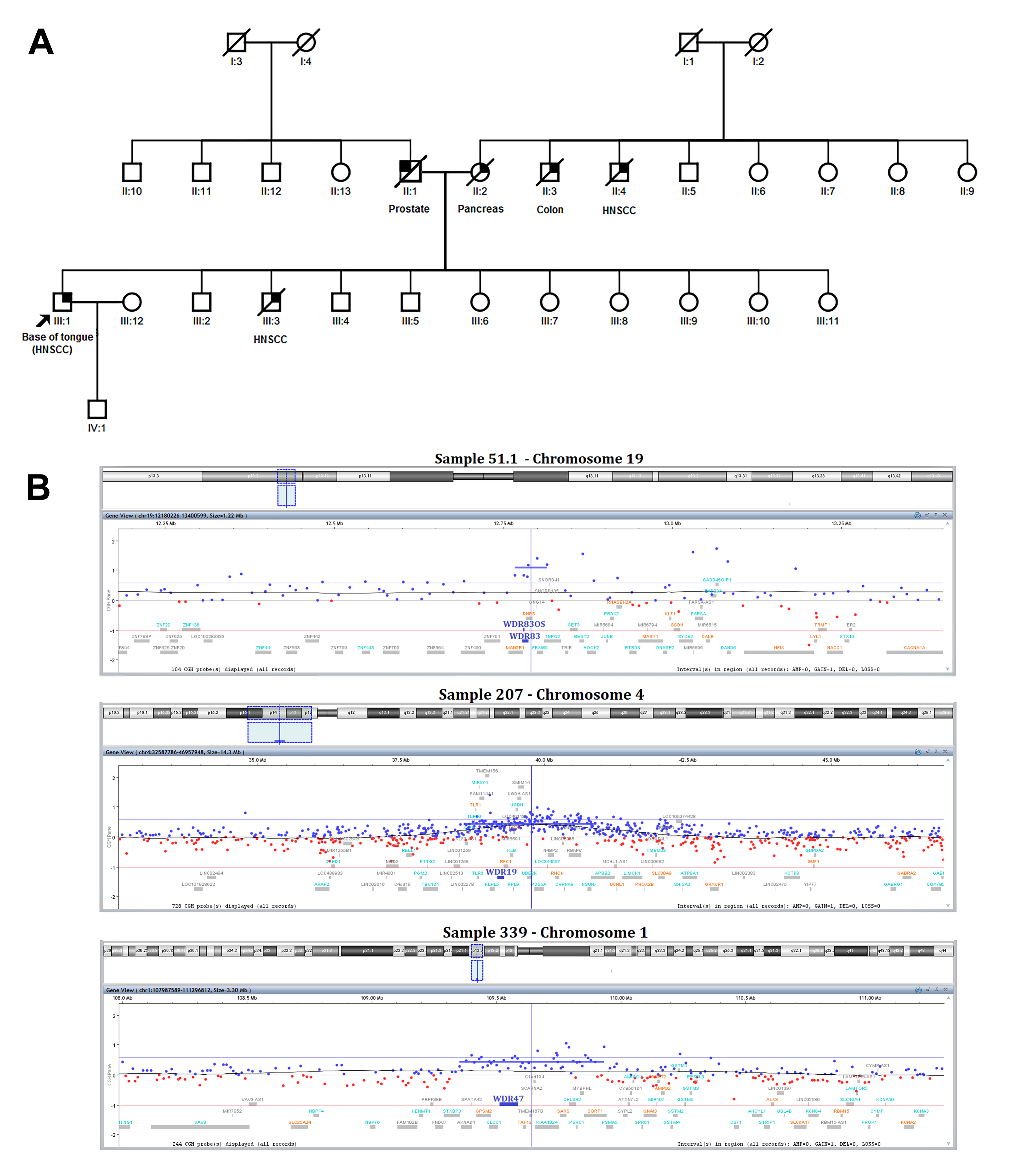
Copy Number Variations
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- Shanmugam, A.; Hariharan, A.K.; Hasina, R.; Nair, J.R.; Katragadda, S.; Irusappan, S.; Ravichandran, A.; Veeramachaneni, V.; Bettadapura, R.; Bhati, M.; et al. Ultrasensitive detection of tumor-specific mutations in saliva of patients with oral cavity squamous cell carcinoma. Cancer 2021, 127, 1576–1589. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, A.L.; Nishimoto, I.N.; Califano, J.A.; Kowalski, L.P. Trends in incidence and prognosis for head and neck cancer in the United States: A site-specific analysis of the SEER database. Int. J. Cancer 2005, 114, 806–816. [Google Scholar] [CrossRef] [PubMed]
- Fuller, C.D.; Wang, S.J.; Thomas, C.R.; Hoffman, H.T.; Weber, R.S.; Rosenthal, D.I. Conditional survival in head and neck squamous cell carcinoma: Results from the SEER dataset 1973–1998. Cancer 2007, 109, 1331–1343. [Google Scholar] [CrossRef] [PubMed]
- Blot, W.J.; McLaughlin, J.K.; Winn, D.M.; Austin, D.F.; Greenberg, R.S.; Preston-Martin, S.; Bernstein, L.; Schoenberg, J.B.; Stemhagen, A.; Fraumeni, J.F., Jr. Smoking and Drinking in Relation to Oral and Pharyngeal Cancer. Cancer Res. 1988, 48, 3282–3287. [Google Scholar] [CrossRef]
- Furniss, C.S.; McClean, M.D.; Smith, J.F.; Bryan, J.; Nelson, H.H.; Peters, E.S.; Posner, M.R.; Clark, J.R.; Eisen, E.A.; Kelsey, K.T. Human papillomavirus 16 and head and neck squamous cell carcinoma. Int. J. Cancer 2007, 120, 2386–2392. [Google Scholar] [CrossRef]
- Franco, E.L.; Kowalski, L.P.; Oliveira, B.V.; Curado, M.P.; Pereira, R.N.; Silva, M.E.; Fava, A.S.; Torloni, H. Risk factors for oral cancer in Brazil: A case-control study. Int. J. Cancer 1989, 43, 992–1000. [Google Scholar] [CrossRef]
- Hashibe, M.; Brennan, P.; Chuang, S.-C.; Boccia, S.; Castellsague, X.; Chen, C.; Curado, M.P.; Maso, L.D.; Daudt, A.W.; Fabianova, E.; et al. Interaction between Tobacco and Alcohol Use and the Risk of Head and Neck Cancer: Pooled Analysis in the International Head and Neck Cancer Epidemiology Consortium. Cancer Epidemiol. Biomark. Prev. 2009, 18, 541–550. [Google Scholar] [CrossRef]
- Rodriguez, T.; Altieri, A.; Chatenoud, L.; Gallus, S.; Bosetti, C.; Negri, E.; Franceschi, S.; Levi, F.; Talamini, R.; La Vecchia, C. Risk factors for oral and pharyngeal cancer in young adults. Oral Oncol. 2003, 40, 207–213. [Google Scholar] [CrossRef]
- Zhang, Z.F.; Morgenstern, H.; Spitz, M.R.; Tashkin, D.P.; Yu, G.P.; Hsu, T.C.; Schantz, S.P. Environmental tobacco smoking, mutagen sensitivity, and head and neck squamous cell carcinoma. Cancer Epidemiol. Biomark. Prev. 2000, 9, 1043–1049. [Google Scholar]
- Toner, M.; O’Regan, E.M. Head and Neck Squamous Cell Carcinoma in the Young: A Spectrum or a Distinct Group? Part 1. Head Neck Pathol. 2009, 3, 246–248. [Google Scholar] [CrossRef] [PubMed]
- Copper, M.P.; Jovanovic, A.; Nauta, J.J.P.; Braakhuis, B.J.M.; de Vries, N.; van der Waal, I.; Snow, G.B. Role of genetic factors in the etiology of squamous cell carcinoma of the head and neck. Arch. Otolaryngol. Head Neck Surg. 1995, 121, 157–160. [Google Scholar] [CrossRef] [PubMed]
- Foulkes, W.; Brunet, J.; Kowalski, L.P.; Narod, S.A.; Franco, E.L. Family history of cancer is a risk factor for squamous cell carcinoma of the head and neck in Brazil: A case-control study. Int. J. Cancer 1995, 63, 769–773. [Google Scholar] [CrossRef]
- Foulkes, W.D.; Brunet, J.S.; Sieh, W.; Black, M.J.; Shenouda, G.; Narod, S.A. Familial risks of squamous cell carcinoma of the head and neck: Retrospective case-control study. BMJ 1996, 313, 716–721. [Google Scholar] [CrossRef] [PubMed]
- Brown, L.M.; Gridley, G.; Diehl, S.R.; Winn, D.M.; Harty, L.C.; Otero, E.B.; Fraumeni, J.F., Jr.; Hayes, R.B. Family cancer history and susceptibility to oral carcinoma in Puerto Rico. Cancer 2001, 92, 2102–2108. [Google Scholar] [CrossRef] [PubMed]
- Garavello, W.; Foschi, R.; Talamini, R.; La Vecchia, C.; Rossi, M.; Maso, L.D.; Tavani, A.; Levi, F.; Barzan, L.; Ramazzotti, V.; et al. Family history and the risk of oral and pharyngeal cancer. Int. J. Cancer 2008, 122, 1827–1831. [Google Scholar] [CrossRef] [PubMed]
- Negri, E.; Boffetta, P.; Berthiller, J.; Castellsagué, X.; Curado, M.P.; Dal Maso, L.; Daudt, A.W.; Fabianova, E.; Fernandez, L.; Wünsch-Filho, V.; et al. Family history of cancer: Pooled analysis in the International Head and Neck Cancer Epidemiology Consortium. Int. J. Cancer 2009, 124, 394–401. [Google Scholar] [CrossRef] [PubMed]
- Yu, G.; Zhang, Z.; Hsu, T.; Spitz, M.; Schantz, S. Family history of cancer, mutagen sensitivity, and increased risk of head and neck cancer. Cancer Lett. 1999, 146, 93–101. [Google Scholar] [CrossRef]
- Monroe, M.M.; Hashibe, M.; Orb, Q.; Alt, J.; Buchmann, L.; Hunt, J.; Cannon-Albright, L.A. Familial clustering of oropharyngeal squamous cell carcinoma in the Utah population. Head Neck 2018, 40, 384–393. [Google Scholar] [CrossRef]
- Bertonha, F.B.; Filho, M.D.C.B.; Kuasne, H.; dos Reis, P.P.; Prando, E.D.C.; Muñoz, J.J.A.M.; Roffé, M.; Hajj, G.N.M.; Kowalski, L.P.; Rainho, C.A.; et al. PHF21B as a candidate tumor suppressor gene in head and neck squamous cell carcinomas. Mol. Oncol. 2015, 9, 450–462. [Google Scholar] [CrossRef]
- Yu, K.K.; Zanation, A.M.; Moss, J.R.; Yarbrough, W.G. Familial head and neck cancer: Molecular analysis of a new clinical entity. Laryngoscope 2002, 112, 1587–1593. [Google Scholar] [CrossRef] [PubMed]
- Fostira, F.; Koutsodontis, G.; Vagia, E.; Economopoulou, P.; Kotsantis, I.; Sasaki, C.; Rontogianni, D.; Perisanidis, C.; Psyrri, A. Predisposing Germline Mutations in Young Patients with Squamous Cell Cancer of the Oral Cavity. JCO Precis. Oncol. 2018, 2, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Cury, S.S.; de Miranda, P.M.; Marchi, F.A.; Canto, L.M.D.; Chulam, T.C.; Petersen, A.H.; Aagaard, M.M.; Pinto, C.A.L.; Kowalski, L.P.; Rogatto, S.R. Germline variants in DNA repair genes are associated with young-onset head and neck cancer. Oral Oncol. 2021, 122, 105545. [Google Scholar] [CrossRef] [PubMed]
- Cancer Genome Atlas Network. Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature 2015, 517, 576–582. [Google Scholar] [CrossRef] [PubMed]
- Ribeiro, I.; Carreira, I.; Esteves, L.; Caramelo, F.; Liehr, T.; Melo, J. Chromosomal breakpoints in a cohort of head and neck squamous cell carcinoma patients. Genomics 2020, 112, 297–303. [Google Scholar] [CrossRef] [PubMed]
- Hu, L.; Yao, X.; Huang, H.; Guo, Z.; Cheng, X.; Xu, Y.; Shen, Y.; Xu, B.; Li, D. Clinical significance of germline copy number variation in susceptibility of human diseases. J. Genet. Genom. 2018, 45, 3–12. [Google Scholar] [CrossRef] [PubMed]
- Krepischi, A.C.; Achatz, M.I.; Santos, E.M.; Costa, S.S.; Lisboa, B.C.; Brentani, H.; Santos, T.M.; Goncalves, A.; Nobrega, A.F.; Pearson, P.L.; et al. Germline DNA copy number variation in familial and early-onset breast cancer. Breast Cancer Res. 2012, 14, R24. [Google Scholar] [CrossRef] [PubMed]
- Hara, H.; Ozeki, S.; Shiratsuchi, Y.; Tashiro, H.; Jingu, K. Familial occurrence of oral cancer: Report of cases. J. Oral Maxillofac. Surg. 1988, 46, 1098–1102. [Google Scholar] [CrossRef]
- Tashiro, H.; Abe, K.; Tanioka, H. Familial occurrence of cancer of the mouth. J. Oral Maxillofac. Surg. 1986, 44, 322–323. [Google Scholar] [CrossRef]
- Leemans, C.R.; Snijders, P.J.F.; Brakenhoff, R.H. The molecular landscape of head and neck cancer. Nat. Rev. Cancer 2018, 18, 269–282. [Google Scholar] [CrossRef]
- Pires, R.C.; Carvalho, R.; Gama, R.R.; Carvalho, A.L.; Santos, C.R.; Capuzzo, R.D.C. Progressive increase trend in HPV-related oropharyngeal squamous cell carcinoma in Brazil. Int. Arch. Otorhinolaryngol. 2021, 26, e132–e136. [Google Scholar] [CrossRef] [PubMed]
- Lacko, M.; Braakhuis, B.J.; Sturgis, E.M.; Boedeker, C.C.; Suárez, C.; Rinaldo, A.; Ferlito, A.; Takes, R.P. Genetic Susceptibility to Head and Neck Squamous Cell Carcinoma. Int. J. Radiat. Oncol. 2014, 89, 38–48. [Google Scholar] [CrossRef] [PubMed]
- Hu, D.; Shi, W.; Yu, M.; Zhang, L. High WDR34 mRNA expression as a potential prognostic biomarker in patients with breast cancer as determined by integrated bioinformatics analysis. Oncol. Lett. 2019, 18, 3177–3187. [Google Scholar] [CrossRef] [PubMed]
- Schapira, M.; Tyers, M.; Torrent, M.; Arrowsmith, C. WD40 repeat domain proteins: A novel target class? Nat. Rev. Drug Discov. 2017, 16, 773–786. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, J.I.; Kasamatsu, A.; Okubo, Y.; Nakashima, D.; Fushimi, K.; Minakawa, Y.; Kasama, H.; Shiiba, M.; Tanzawa, H.; Uzawa, K. Evaluation of tryptophan-aspartic acid repeat-containing protein 34 as a novel tumor-suppressor molecule in human oral cancer. Biochem. Biophys. Res. Commun. 2018, 495, 2469–2474. [Google Scholar] [CrossRef]
- Su, W.Y.; Li, J.T.; Cui, Y.; Hong, J.; Du, W.; Wang, Y.C.; Lin, Y.W.; Xiong, H.; Wang, J.L.; Kong, X.; et al. Bidirectional regulation between WDR83 and its natural antisense transcript DHPS in gastric cancer. Cell Res. 2012, 22, 1374–1389. [Google Scholar] [CrossRef][Green Version]
- Lin, B.; Utleg, A.G.; Gravdal, K.; White, J.T.; Halvorsen, O.J.; Lu, W.; True, L.D.; Vessella, R.; Lange, P.H.; Nelson, P.S.; et al. WDR19 Expression is Increased in Prostate Cancer Compared with Normal Cells, but Low-Intensity Expression in Cancers is Associated with Shorter Time to Biochemical Failures and Local Recurrence. Clin. Cancer Res. 2008, 14, 1397–1406. [Google Scholar] [CrossRef]
- Chatrath, A.; Przanowska, R.; Kiran, S.; Su, Z.; Saha, S.; Wilson, B.; Tsunematsu, T.; Ahn, J.H.; Lee, K.Y.; Paulsen, T.; et al. The pan-cancer landscape of prognostic germline variants in 10,582 patients. Genome Med. 2020, 12, 15. [Google Scholar] [CrossRef]
- Karmakar, M.; Lai, P.C.; Sinha, S.; Glaser, S.; Chakraborty, S. Identification of miR-203a, mir-10a, and miR-194 as predictors for risk of lymphovascular invasion in head and neck cancers. Oncotarget 2021, 12, 1499–1519. [Google Scholar] [CrossRef]
- Xiang, Z.; Song, J.; Zhuo, X.; Li, Q.; Zhang, X. MiR-146a rs2910164 polymorphism and head and neck carcinoma risk: A meta-analysis based on 10 case-control studies. Oncotarget 2017, 8, 1226–1233. [Google Scholar] [CrossRef]
- Chu, C.; Liu, X.; Zhao, Z.; Shi, Z. Circ_0008035 promotes the progression of gastric cancer via the regulation of miR-1256/CEACAM6 axis. Cell Cycle 2022, 21, 1091–1102. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.J.; Xiao, Q.; Li, X.-Y. NF-κB–Activated miR-574 Promotes Multiple Malignant and Metastatic Phenotypes by Targeting BNIP3 in Thyroid Carcinoma. Mol. Cancer Res. 2020, 18, 955–967. [Google Scholar] [CrossRef] [PubMed]
- Calhoun, M.A.; Cui, Y.; Elliott, E.E.; Mo, X.; Otero, J.J.; Winter, J.O. MicroRNA-mRNA Interactions at Low Levels of Compressive Solid Stress Implicate mir-548 in Increased Glioblastoma Cell Motility. Sci. Rep. 2020, 10, 311. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Liu, Z.; Tong, H.; Peng, H.; Xian, Z.; Li, L.; Hu, B.; Xie, S. Linc01194 acts as an oncogene in colorectal carcinoma and is associated with poor survival outcome. Cancer Manag. Res. 2019, 11, 2349–2362. [Google Scholar] [CrossRef]
- Zhai, D.; Zhang, M.; Li, Y.; Bi, J.; Kuang, X.; Shan, Z.; Shao, N.; Lin, Y. LINC01194 recruits NUMA1 to promote ubiquitination of RYR2 to enhance malignant progression in triple-negative breast cancer. Cancer Lett. 2022, 544, 215797. [Google Scholar] [CrossRef]
- Liu, D.M.; Yang, H.; Yuan, Z.N.; Yang, X.G.; Pei, R.; He, H.J. Long noncoding RNA LINC01194 enhances the malignancy of laryngeal squamous cell carcinoma by sponging miR-655 to increase SOX18 expression. Biochem. Biophys. Res. Commun. 2020, 529, 148–155. [Google Scholar] [CrossRef]
- Liu, L.; Liu, J.; Lin, Q. Histone demethylase KDM2A: Biological functions and clinical values (Review). Exp. Ther. Med. 2021, 22, 723. [Google Scholar] [CrossRef]
- Greco, A.; Pierotti, M.A.; Bongarzone, I.; Pagliardini, S.; Lanzi, C.; Della Porta, G. TRK-T1 is a novel oncogene formed by the fusion of TPR and TRK genes in human papillary thyroid carcinomas. Oncogene 1992, 7, 237–242. [Google Scholar]
- Li, F.; Zhai, Y.-P.; Tang, Y.-M.; Wang, L.-P.; Wan, P.-J. Identification of a novel partner gene, TPR, fused to FGFR1 in 8p11 myeloproliferative syndrome. Genes Chromosom. Cancer 2012, 51, 890–897. [Google Scholar] [CrossRef]
- León, J.; Casado, J.; Ruiz, S.M.J.; Zurita, M.S.; González-Puga, C.; Rejón, J.D.; Gila, A.; de Rueda, P.M.; Pavón, E.J.; Reiter, R.J.; et al. Melatonin reduces endothelin-1 expression and secretion in colon cancer cells through the inactivation of FoxO-1 and NF-κβ. J. Pineal Res. 2014, 56, 415–426. [Google Scholar] [CrossRef]
- Lamprou, I.; Tsolou, A.; Kakouratos, C.; Mitrakas, A.G.; Xanthopoulou, E.T.; Kassela, K.; Karakasiliotis, I.; Zois, C.E.; Giatromanolaki, A.; Koukourakis, M.I. Suppressed PLIN3 frequently occurs in prostate cancer, promoting docetaxel resistance via intensified autophagy, an event reversed by chloroquine. Med Oncol. 2021, 38, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Seya, T.; Oshiumi, H.; Sasai, M.; Akazawa, T.; Matsumoto, M. TICAM-1 and TICAM-2: Toll-like receptor adapters that participate in induction of type 1 interferons. Int. J. Biochem. Cell Biol. 2005, 37, 524–529. [Google Scholar] [CrossRef] [PubMed]
| Variable | Category | Frequency (%) (n = 74) | Array-CGH (n = 18) |
|---|---|---|---|
| Gender | Male | 56 (75.7) | 15 |
| Female | 18 (24.3) | 3 | |
| Age (years) | ≤45 | 3 | 1 |
| >45 | 71 | 17 | |
| Race | Caucasian | 63 (85.2) | 18 |
| Not Caucasian | 11 (14.8) | 0 | |
| Smoking | No | 13 (18.9) | 4 |
| Yes | 61 (81.1) | 14 | |
| Passive Smoking | No | 63 (82.5) | 15 |
| Yes | 13 (17.5) | 3 | |
| Alcohol Consumption | No | 15 (20.3) | 4 |
| Yes | 59 (79.7) | 14 | |
| Tumor Site | Oral Cavity | 31 (41.9) | 10 |
| Oropharynx | 15 (20.3) | 6 | |
| Larynx | 24 (32.4) | 0 | |
| Hypopharynx | 4 (5.4) | 2 | |
| Clinical Stage | I | 12 (16.2) | 2 |
| II | 11 (14.9) | 3 | |
| III | 15 (20.3) | 4 | |
| IV | 36 (48.6) | 9 | |
| HPV Status | Yes (HPV16) | 5 (6.7) | 2 |
| No | 60 (81.1) | 14 | |
| NA | 9 (12.2) | 2 | |
| Status | Alive without disease | 65 (87.8) | 11 |
| Alive with disease | 5 (6.8) | 2 | |
| Died by disease or other causes | 5 (6.8) | 4 |
| Tumor | Father n = 31 (%) | Mother n = 28 (%) | Sibling n = 58 (%) | Child n = 4 (%) | 1st Degree Relatives (%) | 2nd Degree Relatives (%) |
|---|---|---|---|---|---|---|
| Head and Neck | 5 (16.1) | 1 (3.6) | 13 (22.4) | - | 19 (15.7) | 11 (27.5) |
| Breast | - | 7 (25.0) | 14 (24.1) | - | 21 (17.4) | 5 (12.5) |
| Stomach | 7 (22.6) | 3 (10.7) | 2 (3.4) | 1 (25.0) | 13 (10.7) | 5 (12.5) |
| Esophagus | 2 (6.5) | - | 4 (6.9) | - | 6 (5.0) | 3 (7.5) |
| Colon | 5 (16.1) | 4 (14.3) | 5 (8.6) | - | 14 (11.6) | 7 (17.5) |
| Prostate | 5 (16.1) | - | 2 (3.4) | - | 7 (5.8) | 1 (2.5) |
| Melanoma | 1 (3.2) | - | - | - | 1 (0.8) | - |
| Lung | 2 (6.5) | 3 (10.7) | 3 (5.2) | - | 8 (6.6) | 3 (7.5) |
| CNS | - | - | - | - | 1 (2.5) | |
| Liver and Biliary Tract | 2 (6.5) | - | 1 (1.7) | - | 3 (2.5) | - |
| Thyroid | - | 2 (7.1) | 1 (1.7) | - | 3 (2.5) | - |
| Uterine Cervix | - | 7 (25.0) | 7 (12.1) | 1 (25.0) | 15 (12.4) | 3 (7.5) |
| Pancreas | - | 1 (3.6) | 3 (5.2) | - | 4 (3.3) | 1 (2.5) |
| Skin | - | - | 1 (1.7) | - | 1 (0.8) | - |
| Leukemia | - | - | - | 1 (25.0) | 1 (0.8) | - |
| Kidney | - | - | 1 (1.7) | - | 1 (0.8) | - |
| Bladder | 1 (3.2) | - | 1 (1.7) | - | 2 (1.7) | - |
| Testicle | - | - | - | 1 (25.0) | 1 (0.8) | - |
| Sample | Chr | Cytoband | Start | End | #Probes | Event | Log2 Ratio | p-Value | miRNAs/lncRNAs | Genes | Note |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 2.1 | 19 | q12 | 29,816,969 | 29,877,356 | 4 | loss | −0.729499 | 6.89 × 10−11 | -/VSTM2B-DT | - | Only intronic region (non-coding RNA) |
| X | q22.2 | 103,186,126 | 103,353,105 | 13 | gain | 0.596946 | 8.82 × 10−23 | MIR1256/- | TMSB15B, H2BFXP, H2BFWT, H2BFM, SLC25A53 | ||
| 14.1 | 2 | p23.3 | 25,875,529 | 25,930,894 | 6 | loss | −0.727414 | 2.15 × 10−11 | - | DTNB | Encompasses the first exon of the main isoform |
| 26.1 | 1 | q31.1 | 186,322,925 | 186,565,261 | 17 | gain | 0.877082 | 2.90 × 10−64 | MIR548F1/- | TPR, ODR4, OCLM, PDC | Fully covers all genes, except the miRNA (1/2 exons) |
| 27.1 | 5 | p15.2 | 12,592,363 | 12,669,790 | 4 | loss | −0.940583 | 1.18 × 10−11 | -/LINC01194 | - | Only intronic region |
| 27.2 | 5 | p15.2 | 12,592,363 | 12,669,790 | 4 | loss | −1.072796 | 3.07 × 10−17 | -/LINC01194 | - | Only intronic region |
| 11 | q13.2 | 66,831,965 | 66,984,794 | 13 | gain | 0.86627 | 1.79 × 10−25 | - | RHOD, KDM2A | Encompasses 4/5 exons of RHOD and 8/21 exons of KDM2A | |
| 51.1 | 2 | p25.1 | 9,802,238 | 9,886,004 | 4 | gain | 0.957729 | 1.59 × 10−10 | - | - | |
| 19 | p13.2 | 12,766,741 | 12,814,116 | 6 | gain | 1.046386 | 4.80 × 10-11 | - | MAN2B1, WDR83, WDR83OS, DHPS, FBXW9, TNPO2 | Fully covers WDR83, WDR83OS, DHPS, FBXW9, 13/24 exons of MAN2B1, and 6/26 exons of TNPO2 | |
| 53 | X | p22.33 | 1,731,671 | 1,746,762 | 4 | gain | 0.750151 | 1.12 × 10−15 | - | ASMT | Encompasses 4/10 exons |
| 58.1 | 22 | q13.31-q13.32 | 46,613,498 | 48,444,626 | 156 | gain | 0.394413 | 2.94 × 10−61 | -/GTSE1,KLF3-AS1 | PPARA, CDPF1, PKDREJ, TTC38, TRMU, CELSR1, GRAMD4, CERK, TBC1D22A | |
| 66 | 4 | q25 | 110,754,346 | 110,812,252 | 5 | loss | −0.713604 | 3.13 × 10−10 | - | RRH, LRIT3 | Encompasses 5/7 exons of RRH and the whole LRIT3 gene |
| 7 | q21.13 | 89,865,734 | 89,917,070 | 6 | loss | −0.66345 | 1.47 × 10−10 | - | STEAP2, CFAP69 | Encompasses 1/6 exons of STEAP2 and 14/23 exons of CFAP69 | |
| 84.1 | 17 | p12 | 14,100,118 | 15,442,066 | 68 | loss | −0.794377 | 4.01 × 10−102 | -/MGC12916, CDRT7 | COX10, CDRT15, HS3ST3B1, PMP22, TEKT3, CDRT4, TVP23C-CDRT4, TVP23C | Fully covers all genes, except COX10 (1/6 exons), TVP23C 1/6 exons), and TVP23C-CDRT4 (2/7 exons) |
| 20 | q13.12 | 44,125,965 | 44,408,171 | 25 | gain | 0.518682 | 4.41 × 10−23 | - | SPINT3, WFDC6, SPINLW1-WFDC6, SPINLW1, WFDC8, WFDC9, WFDC10A, WFDC11, WFDC10B, WFDC13, SPINT4, WFDC3 | Encompasses all of the genes, except WFDC3 (3/7 exons) | |
| 162 | 8 | q21.13 | 81,838,414 | 82,051,934 | 19 | gain | 0.559769 | 7.35 × 10−23 | - | PAG1 | |
| 207 | 2 | q11.2 | 101,488,408 | 101,727,546 | 22 | gain | 0.709334 | 2.37 × 10−40 | - | NPAS2, RPL31, TBC1D8 | |
| 4 | p14 | 38,637,640 | 40,908,108 | 178 | gain | 0.397618 | 9.56 × 10−123 | MIR574/KLF3-AS1 | KLF3, TLR10, TLR1, TLR6, FAM114A1, TMEM156, KLHL5, WDR19, RFC1, KLB, RPL9, LIAS, LOC401127, UGDH, SMIM14, UBE2K, PDS5A, LOC344967, N4BP2, RHOH, CHRNA9, RBM47, NSUN7, APBB2 | ||
| 229 | 12 | q14.1 | 61,562,860 | 61,638,882 | 4 | gain | 0.720912 | 2.17 × 10−10 | - | ||
| 19 | p13.3 | 4,809,281 | 4,889,734 | 10 | loss | −0.619179 | 7.88 × 10−13 | - | TICAM1, PLIN3 | Fully covers both genes | |
| 339 | 1 | p13.3 | 109,351,734 | 109,932,682 | 48 | gain | 0.396988 | 6.11 × 10−35 | -/SCARNA2 | STXBP3, AKNAD1, LOC642864, GPSM2, CLCC1, WDR47, TAF13, TMEM167B, CFAP276, ELAPOR13, SARS1, CELSR2, PSRC1, MYBPHL, SORT1 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chulam, T.C.; Bertonha, F.B.; Villacis, R.A.R.; Filho, J.G.; Kowalski, L.P.; Rogatto, S.R. Epidemiological, Clinical, and Genomic Profile in Head and Neck Cancer Patients and Their Families. Biomedicines 2022, 10, 3278. https://doi.org/10.3390/biomedicines10123278
Chulam TC, Bertonha FB, Villacis RAR, Filho JG, Kowalski LP, Rogatto SR. Epidemiological, Clinical, and Genomic Profile in Head and Neck Cancer Patients and Their Families. Biomedicines. 2022; 10(12):3278. https://doi.org/10.3390/biomedicines10123278
Chicago/Turabian StyleChulam, Thiago Celestino, Fernanda Bernardi Bertonha, Rolando André Rios Villacis, João Gonçalves Filho, Luiz Paulo Kowalski, and Silvia Regina Rogatto. 2022. "Epidemiological, Clinical, and Genomic Profile in Head and Neck Cancer Patients and Their Families" Biomedicines 10, no. 12: 3278. https://doi.org/10.3390/biomedicines10123278
APA StyleChulam, T. C., Bertonha, F. B., Villacis, R. A. R., Filho, J. G., Kowalski, L. P., & Rogatto, S. R. (2022). Epidemiological, Clinical, and Genomic Profile in Head and Neck Cancer Patients and Their Families. Biomedicines, 10(12), 3278. https://doi.org/10.3390/biomedicines10123278






