High-Resolution Magic-Angle-Spinning NMR in Revealing Hepatoblastoma Hallmarks
Abstract
1. Introduction
2. Materials and Methods
2.1. Samples
2.2. Metabolomics by NMR Spectroscopy
2.3. Statistical Analysis of NMR Data
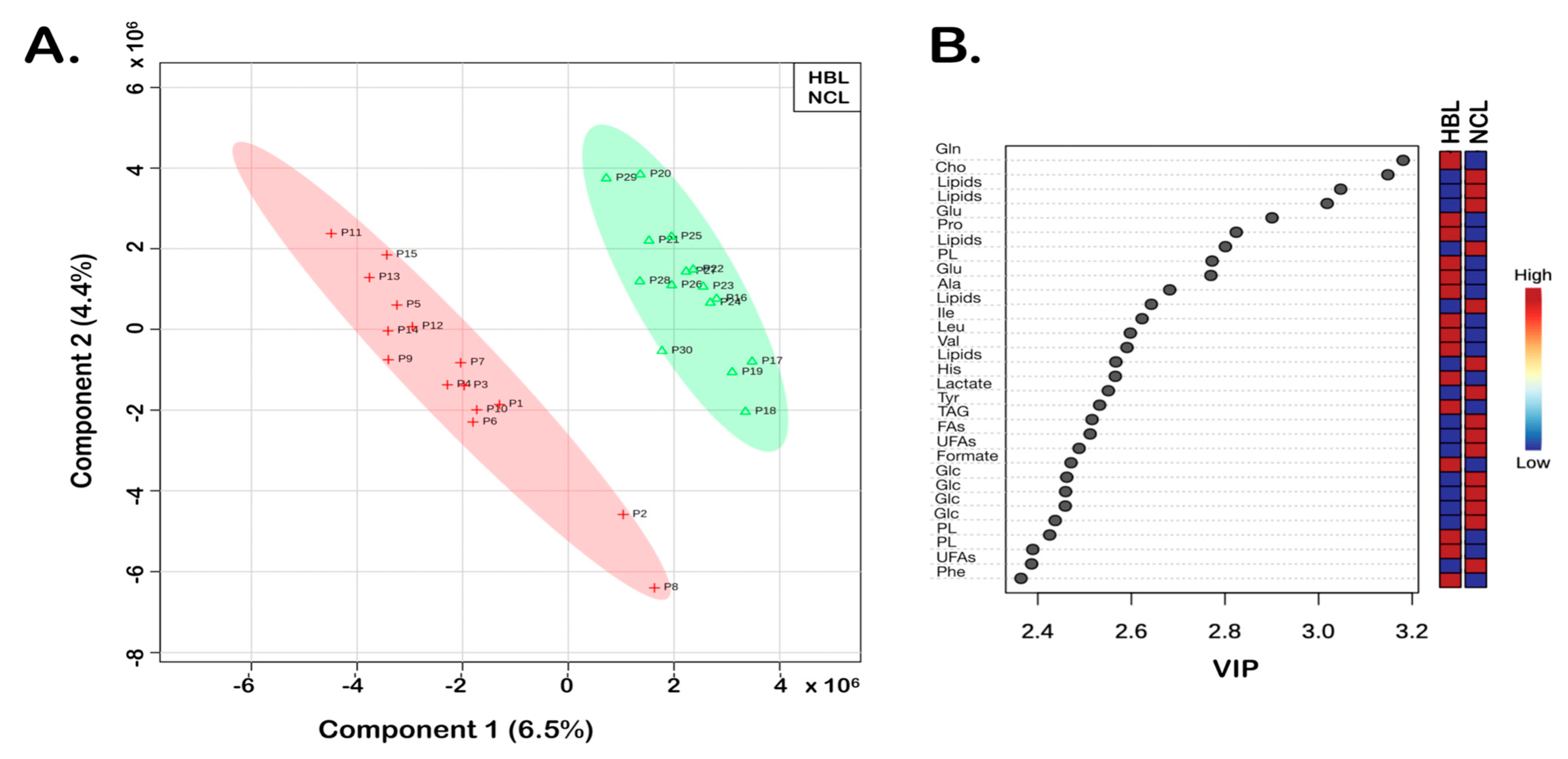
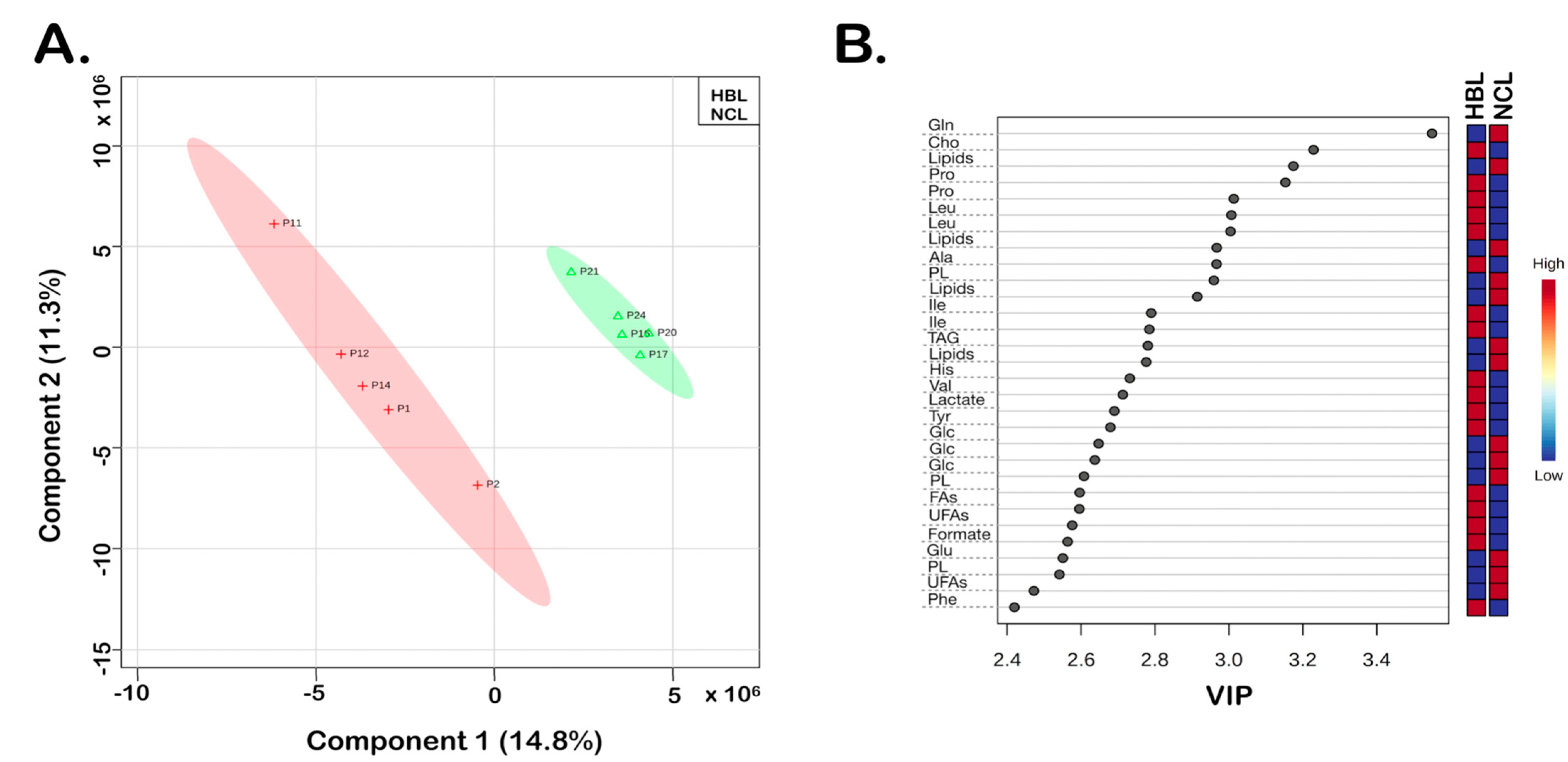
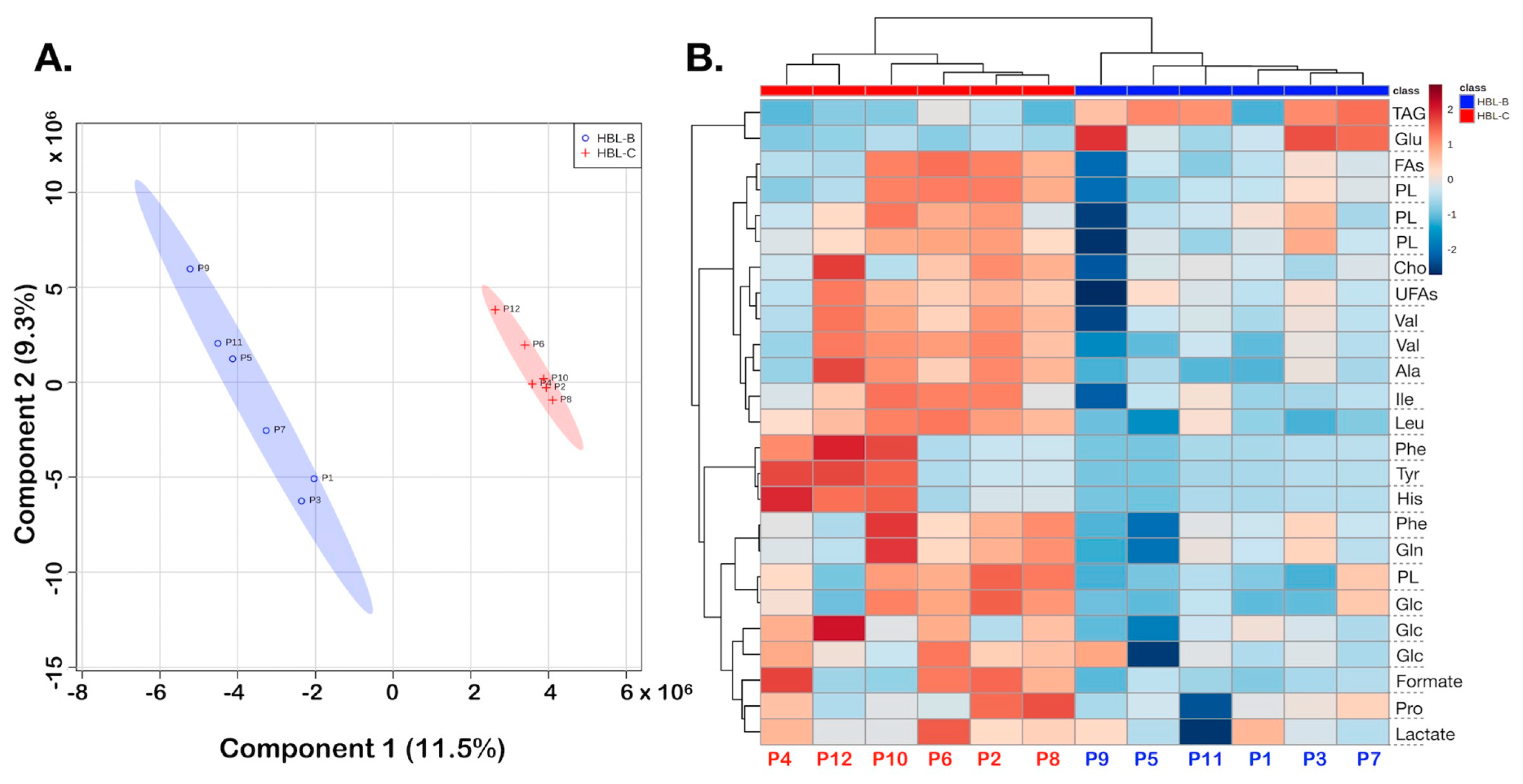
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Johnson, C.; Ivanisevic, J.; Siuzdak, G. Metabolomics: Beyond biomarkers and towards mechanisms. Nat. Rev. Mol. Cell Biol. 2016, 17, 451–459. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Tian, R.; Shan, Y.; Li, J.; Gao, H.; Xie, C.; Ma, Y.; Wu, Y.; Ji, B.; Gu, S.; et al. Metabolomics study of the metabolic changes in hepatoblastoma cells in response to NTCP/SLC10A1 overexpression. Int. J. Biochem. Cell Biol. 2020, 125, 105773. [Google Scholar] [CrossRef] [PubMed]
- Emwas, A.H.; Roy, R.; McKay, R.T.; Tenori, L.; Saccenti, E.; Gowda, G.A.N.; Raftery, D.; Alahmari, F.; Jaremko, L.; Jaremko, M.; et al. NMR spectroscopy for metabolomics research. Metabolites 2019, 9, 123. [Google Scholar] [CrossRef] [PubMed]
- Pontes, J.G.M.; Brasil, A.J.M.; Cruz, G.C.F.; de Souza, R.N.; Tasic, L. NMR-based metabolomics strategies: Plants, animals and humans. Anal. Methods 2017, 9, 1078. [Google Scholar] [CrossRef]
- Quintero Escobar, M.; Costa, T.B.B.C.; Martins, L.G.; Costa, S.S.; VanHelvoort Lengert, A.; Boldrini, E.; Morini da Silva, S.R.; Lopes, L.F.; Vidal, D.O.; Krepischi, A.C.V.; et al. Insights in osteosarcoma by proton nuclear magnetic resonance serum metabonomics. Front. Oncol. 2020, 10, 506959. [Google Scholar] [CrossRef]
- Quintero Escobar, M.; Maschietto, M.; Krepischi, A.C.V.; Avramovic, N.; Tasic, L. Insights into the Chemical Biology of Childhood Embryonal Solid Tumors by NMR-Based Metabolomics. Biomolecules 2019, 9, 843. [Google Scholar] [CrossRef]
- Stanisic, D.; Martins, L.G.; Tasic, L. Nuclear Magnetic Resonance Spectroscopy in Analyses of Biological Samples. In Tools and Trends in Bioanalytical Chemistry; Kubota, L.T., da Silva, J.A.F., Sena, M.M., Alves, W.A., Eds.; Springer: Cham, Switzerland, 2022; pp. 203–221. [Google Scholar] [CrossRef]
- Takis, P.G.; Ghini, V.; Tenori, L.; Turano, P.; Luchinat, C. Uniqueness of the NMR approach to metabolomics. TrAC Trends Anal. Chem. 2019, 120, 115300. [Google Scholar] [CrossRef]
- Le Guennec, A.; Giraudeau, P.; Caldarelli, S. Evaluation of fast 2D NMR for metabolomics. Anal. Chem. 2014, 86, 5946–5954. [Google Scholar] [CrossRef]
- Andrisic, L.; Dudzik, D.; Barbas, C.; Milkovic, L.; Grune, T.; Zarkovic, N. Short overview on metabolomics approach to study pathophysiology of oxidative stress in cancer. Redox Biol. 2018, 14, 47–58. [Google Scholar] [CrossRef]
- Wishart, D. Emerging applications of metabolomics in drug discovery and precision medicine. Nat. Rev. Drug Discov. 2016, 15, 473–484. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. The hallmarks of cancer. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef]
- Abdelahamid, S.; Khedr, R.; Wakeel, E.L.; Younes, M.A.; Ahmed, G.; Elkinaai, N.; Tantawy, M.; Hafez, H. Hepatoblastoma in developing countries; eight years of single Centre experience. J. Cancer Ther. 2018, 9, 793–806. [Google Scholar] [CrossRef][Green Version]
- Aronson, D.C.; Meyers, R.L. Malignant tumors of the liver in children. Semin. Pediatric Surg. 2016, 25, 265–275. [Google Scholar] [CrossRef]
- Cairo, S.; Armengol, C.; De Reyniès, A.; Wei, Y.; Thomas, E.; Renard, C.A.; Goga, A.; Balakrishnan, A.; Semeraro, M.; Gresh, L.; et al. Hepatic stem-like phenotype and interplay of Wnt/β-catenin and Myc signaling in aggressive childhood liver cancer. Cancer Cell 2008, 14, 471–484. [Google Scholar] [CrossRef]
- Eichenmüller, M.; Trippel, F.; Kreuder, M.; Beck, A.; Schwarzmayr, T.; Häberle, B.; Cairo, S.; Leuschner, I.; Von Schweinitz, D.; Strom, T.M.; et al. The genomic landscape of hepatoblastoma and their progenies with HCC-like features. J. Hepatol. 2014, 61, 1312–1320. [Google Scholar] [CrossRef]
- Koch, A.; Denkhaus, D.; Albrecht, S.; Leuschner, I.; Von Schweinitz, D.; Pietsch, T. Childhood hepatoblastomas frequently carry a mutated degradation targeting box of the beta-catenin gene. Cancer Res. 1999, 59, 269–273. [Google Scholar]
- López-Terrada, D.; Gunaratne, P.H.; Adesina, A.M.; Pulliam, J.; Hoang, D.M.; Nguyen, Y.; Mistretta, T.A.; Margolin, J.; Finegold, M.J. Histologic subtypes of hepatoblastoma are characterized by differential canonical Wnt and Notch pathway activation in DLK+ precursors. Hum. Pathol. 2009, 40, 783–794. [Google Scholar] [CrossRef]
- Sumazin, P.; Chen, Y.; Treviño, L.R.; Sarabia, S.F.; Hampton, O.A.; Patel, K.; Mistretta, T.A.; Zorman, B.; Thompson, P.; Heczey, A.; et al. Genomic analysis of hepatoblastoma identifies distinct molecular and prognostic subgroups. Hepatology 2017, 65, 104–121. [Google Scholar] [CrossRef]
- Kalish, J.M.; Doros, L.; Helman, L.J.; Hennekam, R.C.; Kuiper, R.P.; Maas, S.M.; Maher, E.R.; Nichols, K.E.; Plon, S.E.; Porter, C.C.; et al. Surveillance recommendations for children with overgrowth syndromes and predisposition to Wilms tumors and hepatoblastoma. Clin. Cancer Res. 2017, 23, e115–e122. [Google Scholar] [CrossRef]
- Chen, H.; Guan, Q.; Guo, H.; Miao, L.; Zhuo, Z. The genetic changes of hepatoblastoma. Front. Oncol. 2021, 11, 690641. [Google Scholar] [CrossRef]
- Marchese, S.; Sorice, A.; Ariano, A.; Florio, S.; Budillon, A.; Costantini, S.; Severino, L. Evaluation of aflatoxin M1 effects on the metabolomic and cytokinomic profiling of a hepatoblastoma cell line. Toxins 2018, 10, 436. [Google Scholar] [CrossRef] [PubMed]
- Wishart, D.S.; Knox, C.; Guo, A.C.; Eisner, R.; Young, N.; Gautam, B.; Hau, D.D.; Psychogios, N.; Dong, E.; Bouatra, S.; et al. HMDB: A knowledgebase for the human metabolome. Nucleic. Acids Res. 2009, 37, 603–610. [Google Scholar] [CrossRef] [PubMed]
- Ulrich, E.L.; Akutsu, H.; Doreleijers, J.F.; Harano, Y.; Ioannidis, Y.E.; Lin, J.; Livny, M.; Mading, S.; Maziuk, D.; Miller, Z.; et al. BioMagResBank. Nucleic. Acids Res. 2008, 36, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Meyers, R.L.; Maibach, R.; Hiyama, E.; Häberle, B.; Krailo, M.; Rangaswami, A.; Aronson, D.C.; Malogolowkin, M.H.; Perilongo, G.; Von Schweinitz, D.; et al. Risk-stratified staging in paediatric hepatoblastoma: A unified analysis from the Children’s Hepatic tumors International Collaboration. Lancet Oncol. 2017, 18, 122–131. [Google Scholar] [CrossRef] [PubMed]
- Baenke, F.; Peck, B.; Miess, H.; Schulze, A. Hooked on fat: The role of lipid synthesis in cancer metabolism and tumour development. Dis. Model. Mech. 2013, 6, 1353–1363. [Google Scholar] [CrossRef]
- Butler, L.M.; Perone, Y.; Dehairs, J.; Lupien, L.E.; de Laat, V.; Talebi, A.; Loda, M.; Kinla, W.B.; Swinnen, J.V. Lipids, and cancer: Emerging roles in pathogenesis, diagnosis and therapeutic intervention. Adv. Drug Deliv. Rev. 2020, 159, 245–293. [Google Scholar] [CrossRef]
- Warburg, O.; Wind, F.; Negelein, E. The metabolism of tumors in the body. J. Gen. Physiol. 1927, 8, 519–530. [Google Scholar] [CrossRef]
- Luo, X.; Cheng, C.; Tan, Z.; Li, N.; Tang, M.; Yang, L.; Cao, Y. Emerging roles of lipid metabolism in cancer metastasis. Mol. Cancer 2017, 16, 76. [Google Scholar] [CrossRef]
- Currie, E.; Schulze, A.R.; Zechner, T.C.; Walther, R.; Farese, V., Jr. Cell metabolism perspective cellular fatty acid metabolism and cancer. Cell Metab. 2013, 18, 153–161. [Google Scholar] [CrossRef]
- Stephenson, D.J.; Hoeferlin, L.A.; Chalfant, C.E. Lipidomics in translational research and the clinical significance of lipid-based biomarkers. Transl. Res. 2017, 189, 13–29. [Google Scholar] [CrossRef]
- Rivas, M.P.; Aguiar, T.F.M.; Maschietto, M.; Lemes, R.B.; Caires-Júnior, L.C.; Goulart, E.; Telles-Silva, K.A.; Novak, E.; Cristofani, L.M.; Odone, V.; et al. Hepatoblastomas exhibit marked NNMT downregulation driven by promoter DNA hypermethylation. Tumour Biol. 2020, 42, 1010428320977124. [Google Scholar] [CrossRef]
- Schmidt, A.; Armento, A.; Bussolati, O.; Chiu, M.; Ellerkamp, V.; Scharpf, M.O.; Sander, P.; Schmid, E.; Warmann, S.W.; Fuchs, J. Hepatoblastoma: Glutamine depletion hinders cell viability in the embryonal subtype but high GLUL expression is associated with better overall survival. J. Cancer. Res. Clin. Oncol. 2021, 147, 3169–3181. [Google Scholar] [CrossRef]
- Cadoret, A.; Ovejero, C.; Terris, B.; Souil, E.; Lévy, L.; Lamers, W.H.; Kitajewski, J.; Kahn, A.; Perret, C. New targets of beta-catenin signaling in the liver are involved in the glutamine metabolism. Oncogene 2002, 21, 8293–8301. [Google Scholar] [CrossRef]
- Shen, G.; Shen, H.; Zhang, J.; Yan, Q.; Liu, H. DNA methylation in hepatoblastoma—A literature review. Ital. J. Pediatrics 2020, 46, 113. [Google Scholar] [CrossRef]
- Crippa, S.; Ancey, P.B.; Vazquez, J.; Angelino, P.; Rougemont, A.L.; Guettier, C.; Zoete, V.; Delorenzi, M.; Michielin, O.; Meylan, E. Mutant CTNNB1 and histological heterogeneity define metabolic subtypes of hepatoblastoma. EMBO Mol. Med. 2017, 9, 1589–1604. [Google Scholar] [CrossRef]

| Metabolites | Chemical Shifts (ppm), Peak Multiplicities, and Coupling Constants | |
|---|---|---|
| Lipids (-CH3) | 1a | 0.83 m |
| Lipids (-CH2-) | 1b | 1.28 m |
| Ile (Isoleucine) | 2 | 0.93 t (J = 7.4 Hz); 1.00 d (J = 7 Hz); 1.25 m; 1.46 m; 1.97 m; 3.67 d (J = 3.97 Hz) |
| Leu (Leucine) | 3 | 0.95 t (J = 8 Hz; 9 Hz); 1.70 m; 3.72 m |
| Val (Valine) | 4 | 0.98 d (J = 8 Hz); 1.03 (J = 7 Hz); 2.26 m; 3.60 d (J = 4 Hz) |
| Lactate | 5 | 1.33 d (J = 7 Hz); 4.10 q (J = 7 Hz) |
| Ala (Alanine) | 6 | 1.46 d; 3.77 q |
| Pro (Proline) | 7 | 2.00 m; 2.07 m; 2.35 m; 3.34 m; 3.42 m; 4.13 m |
| Glu (Glutamate) | 8 | 2.04 m; 2.12 m; 2.34 m; 3.75 dd (J = 7.19, 4.72 Hz) |
| Gln (Glutamine) | 9 | 2.13 m; 2.45 m; 3.77 t (J = 6.18 Hz) |
| His (Histidine) | 10 | 3.16 dd (J = 7.75 Hz); 3.23 (J = 4.93 Hz); 3.98 (J = 4.98 Hz); 7.10 d (J = 5 Hz); 7.90 d (J = 2 Hz) |
| Cho (Choline) | 11 | 3.19 s; 3.51 dd (J = 5.816 Hz, 4.162 Hz); 4.05 ddd |
| PL (Glycerophosphocholine) | 12 | 3.20 s; 3.62 m; 3.90 m; 4.30 m |
| Glc (Glucose) | 13 | 3.23 dd (J = 9.41 Hz, 7.98 Hz); 3.40 m; 3.46 m; 3.52 dd (J = 9.82 Hz, 3.77 Hz); 3.73 m; 3.82 m; 3.88 dd (J = 12.30 Hz, 2.23 Hz); 4.63 d (J = 7.98 Hz); 5.22 d (J = 3.80 Hz) |
| Tyr (Tyrosine) | 14 | 3.03 dd (J = 14.55 Hz, 8.01 Hz); 3.34 dd (J = 14.53 Hz, 4.68 Hz); 4.04 dd (J = 8.03 Hz, 4.68 Hz); 6.94 m; 7.20 dd (J = 7.95 Hz, 1.51 Hz); 7.24 td (J = 7.76 Hz, 1.71 Hz) |
| Phe (Phenylalanine) | 15 | 3.19 m; 3.98 dd (J = 7.88, 5.31 Hz); 7.32 d (J = 6.96 Hz); 7.34 m; 7.42 m |
| Formate | 16 | 8.40 s |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tasic, L.; Avramović, N.; Jadranin, M.; Quintero, M.; Stanisic, D.; Martins, L.G.; Costa, T.B.B.C.; Novak, E.; Odone, V.; Rivas, M.; et al. High-Resolution Magic-Angle-Spinning NMR in Revealing Hepatoblastoma Hallmarks. Biomedicines 2022, 10, 3091. https://doi.org/10.3390/biomedicines10123091
Tasic L, Avramović N, Jadranin M, Quintero M, Stanisic D, Martins LG, Costa TBBC, Novak E, Odone V, Rivas M, et al. High-Resolution Magic-Angle-Spinning NMR in Revealing Hepatoblastoma Hallmarks. Biomedicines. 2022; 10(12):3091. https://doi.org/10.3390/biomedicines10123091
Chicago/Turabian StyleTasic, Ljubica, Nataša Avramović, Milka Jadranin, Melissa Quintero, Danijela Stanisic, Lucas G. Martins, Tássia Brena Barroso Carneiro Costa, Estela Novak, Vicente Odone, Maria Rivas, and et al. 2022. "High-Resolution Magic-Angle-Spinning NMR in Revealing Hepatoblastoma Hallmarks" Biomedicines 10, no. 12: 3091. https://doi.org/10.3390/biomedicines10123091
APA StyleTasic, L., Avramović, N., Jadranin, M., Quintero, M., Stanisic, D., Martins, L. G., Costa, T. B. B. C., Novak, E., Odone, V., Rivas, M., Aguiar, T., Carraro, D. M., Werneck da Cunha, I., Lima da Costa, C. M., Maschietto, M., & Krepischi, A. (2022). High-Resolution Magic-Angle-Spinning NMR in Revealing Hepatoblastoma Hallmarks. Biomedicines, 10(12), 3091. https://doi.org/10.3390/biomedicines10123091







